Plant Defensins from a Structural Perspective
Abstract
:1. Introduction
2. Primary Sequence
3. Secondary and Tertiary Structures
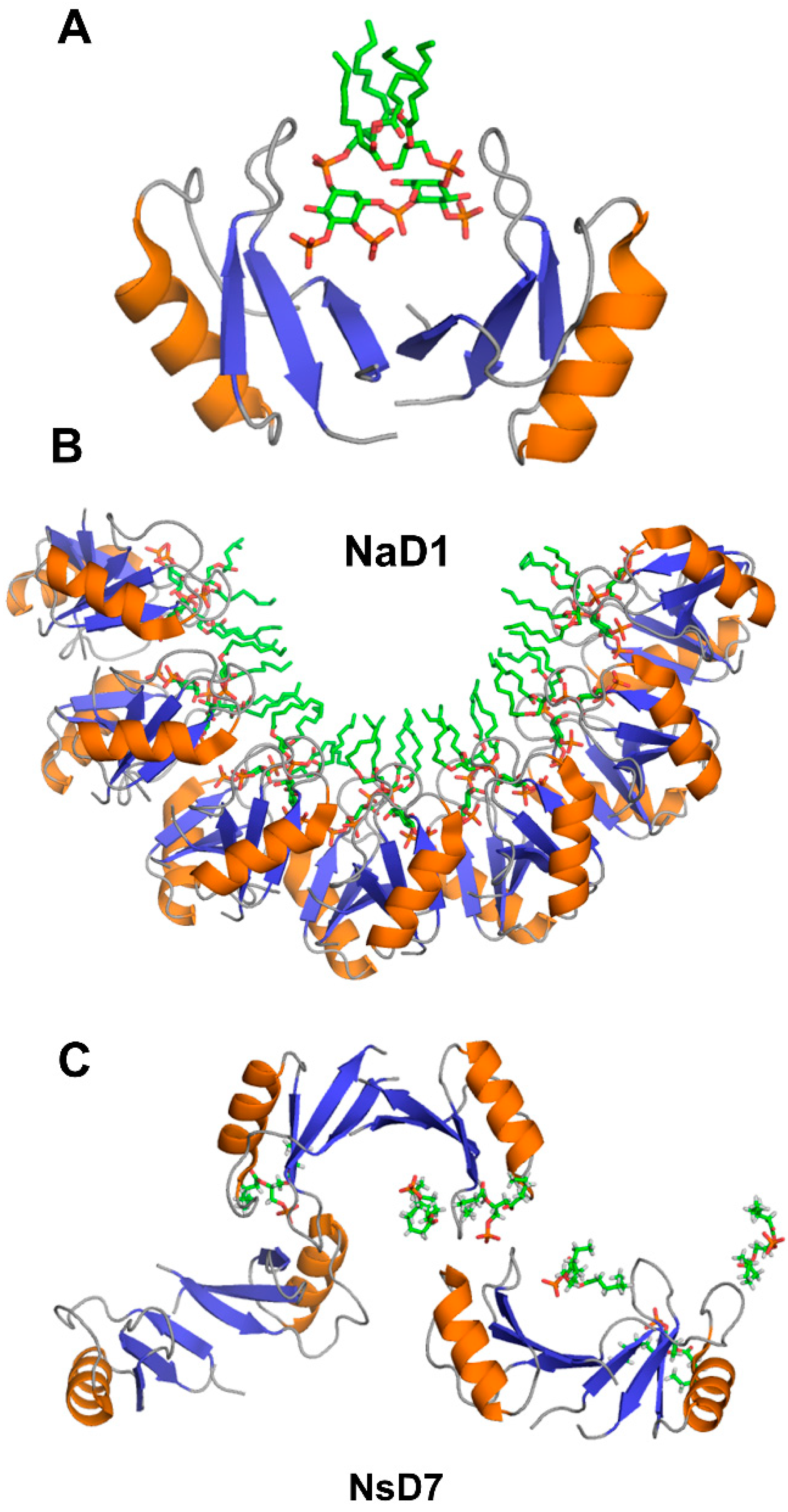
4. The Oligomerization of Plant Defensins and Cell Membrane Targeting
5. Structural Motifs of Plant Defensins Important for Antifungal Activity
5.1. γ-core Motif
5.2. Loop L3 of the γ-core Motif
5.3. Loops L1 and L2
6. Structural Features of Plant Defensins Important for α-Amylase Activity
7. Intracellular Targets of Plant Defensins
8. Dynamics Properties of Plant Defensins
Author Contributions
Funding
Conflicts of Interest
References
- Hammami, R.; Ben Hamida, J.; Vergoten, G.; Fliss, I. PhytAMP: A database dedicated to antimicrobial plant peptides. Nucleic Acids Res. 2009, 37, D963–D968. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Broekaert, W.F.; Terras, F.R.; Cammue, B.P.; Osborn, R.W. Plant defensins: Novel antimicrobial peptides as components of the host defense system. Plant Physiol. 1995, 108, 1353–1358. [Google Scholar] [CrossRef] [PubMed]
- Lay, F.T.; Anderson, M.A. Defensins--components of the innate immune system in plants. Curr. Protein Pept. Sci. 2005, 6, 85–101. [Google Scholar] [CrossRef] [PubMed]
- Terras, F.R.; Eggermont, K.; Kovaleva, V.; Raikhel, N.V.; Osborn, R.W.; Kester, A.; Rees, S.B.; Torrekens, S.; Van Leuven, F.; Vanderleyden, J. Small cysteine-rich antifungal proteins from radish: Their role in host defense. Plant Cell. 1995, 7, 573–588. [Google Scholar]
- Lay, F.T.; Brugliera, F.; Anderson, M.A. Isolation and properties of floral defensins from ornamental tobacco and petunia. Plant Physiol. 2003, 131, 1283–1293. [Google Scholar] [CrossRef] [Green Version]
- Kragh, K.M.; Nielsen, J.E.; Nielsen, K.K.; Dreboldt, S.; Mikkelsen, J.D. Characterization and localization of new antifungal cysteine-rich proteins from Beta vulgaris. Mol. Plant Microbe Interact. 1995, 8, 424–434. [Google Scholar] [CrossRef]
- Mir Drikvand, R.; Sohrabi, S.M.; Samiei, K. Molecular cloning and characterization of six defensin genes from lentil plant (Lens culinaris L.). 3 Biotech. 2019, 9, 104. [Google Scholar] [CrossRef]
- Meyer, B.; Houlne, G.; Pozueta-Romero, J.; Schantz, M.L.; Schantz, R. Fruit-specific expression of a defensin-type gene family in bell pepper. Upregulation during ripening and upon wounding. Plant Physiol. 1996, 112, 615–622. [Google Scholar] [CrossRef] [Green Version]
- Carvalho Ade, O.; Gomes, V.M. Plant defensins and defensin-like peptides—Biological activities and biotechnological applications. Curr. Pharm. Des. 2011, 17, 4270–4293. [Google Scholar] [CrossRef]
- Ermakova, E.A.; Faizullin, D.A.; Idiyatullin, B.Z.; Khairutdinov, B.I.; Mukhamedova, L.N.; Tarasova, N.B.; Toporkova, Y.Y.; Osipova, E.V.; Kovaleva, V.; Gogolev, Y.V.; et al. Structure of Scots pine defensin 1 by spectroscopic methods and computational modeling. Int. J. Biol. Macromol. 2016, 84, 142–152. [Google Scholar] [CrossRef] [PubMed]
- Kerenga, B.K.; McKenna, J.A.; Harvey, P.J.; Quimbar, P.; Garcia-Ceron, D.; Lay, F.T.; Phan, T.K.; Veneer, P.K.; Vasa, S.; Parisi, K.; et al. Salt-Tolerant Antifungal and Antibacterial Activities of the Corn Defensin ZmD32. Front Microbiol. 2019, 10, 795. [Google Scholar] [CrossRef] [PubMed]
- Lacerda, A.; Vasconcelos, É.; Pelegrini, P.; Grossi-de-Sa, M.F. Antifungal defensins and their role in plant defense. Front. Microbiol. 2014, 5, 116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vriens, K.; Cammue, B.P.; Thevissen, K. Antifungal plant defensins: Mechanisms of action and production. Molecules 2014, 19, 12280–12303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomma, B.P.; Cammue, B.P.; Thevissen, K. Plant defensins. Planta 2002, 216, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Mendez, E.; Rocher, A.; Calero, M.; Girbes, T.; Citores, L.; Soriano, F. Primary structure of omega-hordothionin, a member of a novel family of thionins from barley endosperm, and its inhibition of protein synthesis in eukaryotic and prokaryotic cell-free systems. Eur. J. Biochem. 1996, 239, 67–73. [Google Scholar] [CrossRef]
- Pelegrini, P.B.; Lay, F.T.; Murad, A.M.; Anderson, M.A.; Franco, O.L. Novel insights on the mechanism of action of alpha-amylase inhibitors from the plant defensin family. Proteins 2008, 73, 719–729. [Google Scholar] [CrossRef]
- Melo, F.R.; Rigden, D.J.; Franco, O.L.; Mello, L.V.; Ary, M.B.; Grossi de Sa, M.F.; Bloch, C., Jr. Inhibition of trypsin by cowpea thionin: Characterization, molecular modeling, and docking. Proteins 2002, 48, 311–319. [Google Scholar] [CrossRef]
- Spelbrink, R.G.; Dilmac, N.; Allen, A.; Smith, T.J.; Shah, D.M.; Hockerman, G.H. Differential antifungal and calcium channel-blocking activity among structurally related plant defensins. Plant Physiol. 2004, 135, 2055–2067. [Google Scholar] [CrossRef] [Green Version]
- Osborn, R.W.; De Samblanx, G.W.; Thevissen, K.; Goderis, I.; Torrekens, S.; Van Leuven, F.; Attenborough, S.; Rees, S.B.; Broekaert, W.F. Isolation and characterisation of plant defensins from seeds of Asteraceae, Fabaceae, Hippocastanaceae and Saxifragaceae. FEBS Lett. 1995, 368, 257–262. [Google Scholar] [CrossRef] [Green Version]
- Terras, F.R.; Schoofs, H.M.; De Bolle, M.F.; Van Leuven, F.; Rees, S.B.; Vanderleyden, J.; Cammue, B.P.; Broekaert, W.F. Analysis of two novel classes of plant antifungal proteins from radish (Raphanus sativus L.) seeds. J. Biol. Chem. 1992, 267, 15301–15309. [Google Scholar]
- Segura, A.; Moreno, M.; Molina, A.; Garcia-Olmedo, F. Novel defensin subfamily from spinach (Spinacia oleracea). FEBS Lett. 1998, 435, 159–162. [Google Scholar] [CrossRef] [Green Version]
- van der Weerden, N.L.; Anderson, M.A. Plant defensins: Common fold, multiple functions. Fungal Biol. Rev. 2013, 26, 121–131. [Google Scholar] [CrossRef]
- Lay, F.T.; Poon, S.; McKenna, J.A.; Connelly, A.A.; Barbeta, B.L.; McGinness, B.S.; Fox, J.L.; Daly, N.L.; Craik, D.J.; Heath, R.L.; et al. The C-terminal propeptide of a plant defensin confers cytoprotective and subcellular targeting functions. BMC Plant Biol. 2014, 14, 41. [Google Scholar] [CrossRef] [Green Version]
- Shafee, T.M.; Lay, F.T.; Phan, T.K.; Anderson, M.A.; Hulett, M.D. Convergent evolution of defensin sequence, structure and function. Cell Mol. Life Sci. 2017, 74, 663–682. [Google Scholar] [CrossRef]
- Koehbach, J. Structure-Activity Relationships of Insect Defensins. Front. Chem. 2017, 5, 45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Landon, C.; Sodano, P.; Hetru, C.; Hoffmann, J.; Ptak, M. Solution structure of drosomycin, the first inducible antifungal protein from insects. Protein Sci. 1997, 6, 1878–1884. [Google Scholar] [CrossRef] [Green Version]
- Blanc, E.; Sabatier, J.M.; Kharrat, R.; Meunier, S.; el Ayeb, M.; Van Rietschoten, J.; Darbon, H. Solution structure of maurotoxin, a scorpion toxin from Scorpio maurus, with high affinity for voltage-gated potassium channels. Proteins 1997, 29, 321–333. [Google Scholar] [CrossRef]
- Bontems, F.; Gilquin, B.; Roumestand, C.; Menez, A.; Toma, F. Analysis of side-chain organization on a refined model of charybdotoxin: Structural and functional implications. Biochemistry 1992, 31, 7756–7764. [Google Scholar] [CrossRef]
- Ming, D.; Hellekant, G. Brazzein, a new high-potency thermostable sweet protein from Pentadiplandra brazzeana B. FEBS Lett. 1994, 355, 106–108. [Google Scholar] [CrossRef] [Green Version]
- Janssen, B.J.; Schirra, H.J.; Lay, F.T.; Anderson, M.A.; Craik, D.J. Structure of Petunia hybrida defensin 1, a novel plant defensin with five disulfide bonds. Biochemistry 2003, 42, 8214–8222. [Google Scholar] [CrossRef]
- De Bolle, M.F.C.; Terras, F.R.G.; Cammue, B.P.A.; Rees, S.B.; Broekaert, W.F. Mirabilis Jalapa Antibacterial Peptides and Raphanus Sativus Antifungal Proteins: A Comparative Study of their Structure and Biological Activities. In Mechanisms of Plant Defense Responses Developments in Plant Pathology; Fritig, B., Legrand, M., Eds.; Springer: Dordrecht, The Netherlands, 1993. [Google Scholar]
- Ramamoorthy, V.; Zhao, X.; Snyder, A.K.; Xu, J.R.; Shah, D.M. Two mitogen-activated protein kinase signalling cascades mediate basal resistance to antifungal plant defensins in Fusarium graminearum. Cell Microbiol. 2007, 9, 1491–1506. [Google Scholar] [CrossRef] [PubMed]
- Thevissen, K.; Osborn, R.W.; Acland, D.P.; Broekaert, W.F. Specific binding sites for an antifungal plant defensin from Dahlia (Dahlia merckii) on fungal cells are required for antifungal activity. Mol. Plant Microbe. Interact. 2000, 13, 54–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Almeida, M.S.; Cabral, K.M.; Zingali, R.B.; Kurtenbach, E. Characterization of two novel defense peptides from pea (Pisum sativum) seeds. Arch Biochem. Biophys. 2000, 378, 278–286. [Google Scholar] [CrossRef]
- Chen, J.J.; Chen, G.H.; Hsu, H.C.; Li, S.S.; Chen, C.S. Cloning and functional expression of a mungbean defensin VrD1 in Pichia pastoris. J. Agric. Food Chem. 2004, 52, 2256–2261. [Google Scholar] [CrossRef]
- Song, X.; Wang, J.; Wu, F.; Li, X.; Teng, M.; Gong, W. cDNA cloning, functional expression and antifungal activities of a dimeric plant defensin SPE10 from Pachyrrhizus erosus seeds. Plant Mol. Biol. 2005, 57, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.H.; Hsu, M.P.; Tan, C.H.; Sung, H.Y.; Kuo, C.G.; Fan, M.J.; Chen, H.M.; Chen, S.; Chen, C.S. Cloning and characterization of a plant defensin VaD1 from azuki bean. J. Agric. Food Chem. 2005, 53, 982–988. [Google Scholar] [CrossRef]
- Vijayan, S.; Imani, J.; Tanneeru, K.; Guruprasad, L.; Kogel, K.H.; Kirti, P.B. Enhanced antifungal and insect alpha-amylase inhibitory activities of Alpha-TvD1, a peptide variant of Tephrosia villosa defensin (TvD1) generated through in vitro mutagenesis. Peptides 2012, 33, 220–229. [Google Scholar] [CrossRef]
- Kant, P.; Liu, W.Z.; Pauls, K.P. PDC1, a corn defensin peptide expressed in Escherichia coli and Pichia pastoris inhibits growth of Fusarium graminearum. Peptides 2009, 30, 1593–1599. [Google Scholar] [CrossRef]
- Portieles, R.; Ayra, C.; Gonzalez, E.; Gallo, A.; Rodriguez, R.; Chacon, O.; Lopez, Y.; Rodriguez, M.; Castillo, J.; Pujol, M.; et al. NmDef02, a novel antimicrobial gene isolated from Nicotiana megalosiphon confers high-level pathogen resistance under greenhouse and field conditions. Plant Biotechnol. J. 2010, 8, 678–690. [Google Scholar] [CrossRef]
- Kovaleva, V.; Krynytskyy, H.; Gout, I.; Gout, R. Recombinant expression, affinity purification and functional characterization of Scots pine defensin 1. Appl. Microbiol. Biotechnol. 2011, 89, 1093–1101. [Google Scholar] [CrossRef]
- Sagaram, U.S.; Pandurangi, R.; Kaur, J.; Smith, T.J.; Shah, D.M. Structure-activity determinants in antifungal plant defensins MsDef1 and MtDef4 with different modes of action against Fusarium graminearum. PLoS ONE 2011, 6, e18550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Slavokhotova, A.A.; Odintsova, T.I.; Rogozhin, E.A.; Musolyamov, A.K.; Andreev, Y.A.; Grishin, E.V.; Egorov, T.A. Isolation, molecular cloning and antimicrobial activity of novel defensins from common chickweed (Stellaria media L.) seeds. Biochimie 2011, 93, 450–456. [Google Scholar] [PubMed]
- Picart, P.; Pirttilä, A.M.; Raventos, D.; Kristensen, H.-H.; Sahl, H.-G. Identification of defensin-encoding genes of Picea glauca: Characterization of PgD5, a conserved spruce defensin with strong antifungal activity. BMC Plant Biol. 2012, 12, 180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nanni, V.; Zanetti, M.; Bellucci, M.; Moser, C.; Bertolini, P.; Guella, G.; Dalla Serra, M.; Baraldi, E. The peach (Prunus persica) defensin PpDFN1 displays antifungal activity through specific interactions with the membrane lipids. Plant Pathol. 2013, 62, 393–403. [Google Scholar] [CrossRef]
- Dracatos, P.M.; Van der Weerden, N.L.; Carroll, K.T.; Johnson, E.D.; Plummer, K.M.; Anderson, M.A. Inhibition of cereal rust fungi by both class I and II defensins derived from the flowers of Nicotiana alata. Mol. Plant Pathol. 2014, 15, 67–79. [Google Scholar] [CrossRef] [PubMed]
- Shenkarev, Z.O.; Gizatullina, A.K.; Finkina, E.I.; Alekseeva, E.A.; Balandin, S.V.; Mineev, K.S.; Arseniev, A.S.; Ovchinnikova, T.V. Heterologous expression and solution structure of defensin from lentil Lens culinaris. Biochem. Biophys. Res. Commun. 2014, 451, 252–257. [Google Scholar] [CrossRef]
- Omidvar, R.; Xia, Y.; Porcelli, F.; Bohlmann, H.; Veglia, G. NMR structure and conformational dynamics of AtPDFL2.1, a defensin-like peptide from Arabidopsis thaliana. Biochim. Biophys. Acta 2016, 1864, 1739–1747. [Google Scholar] [CrossRef]
- Li, H.; Velivelli, S.L.S.; Shah, D.M. Antifungal Potency and Modes of Action of a Novel Olive Tree Defensin Against Closely Related Ascomycete Fungal Pathogens. Mol. Plant Microbe. Interact 2019, 32, 1649–1664. [Google Scholar] [CrossRef]
- Pinheiro-Aguiar, R.; do Amaral, V.S.G.; Pereira, I.B.; Kurtenbach, E.; Almeida, F.C.L. Nuclear magnetic resonance solution structure of Pisum sativum defensin 2 provides evidence for the presence of hydrophobic surface-clusters. Proteins 2020, 88, 242–246. [Google Scholar] [CrossRef]
- Khairutdinov, B.I.; Ermakova, E.A.; Yusypovych, Y.M.; Bessolicina, E.K.; Tarasova, N.B.; Toporkova, Y.Y.; Kovaleva, V.; Zuev, Y.F.; Nesmelova, I.V. NMR structure, conformational dynamics, and biological activity of PsDef1 defensin from Pinus sylvestris. Biochim. Biophys. Acta 2017, 1865, 1085–1094. [Google Scholar] [CrossRef]
- Liu, Y.J.; Cheng, C.S.; Lai, S.M.; Hsu, M.P.; Chen, C.S.; Lyu, P.C. Solution structure of the plant defensin VrD1 from mung bean and its possible role in insecticidal activity against bruchids. Proteins 2006, 63, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Vriens, K.; Cools, T.L.; Harvey, P.J.; Craik, D.J.; Braem, A.; Vleugels, J.; De Coninck, B.; Cammue, B.P.; Thevissen, K. The radish defensins RsAFP1 and RsAFP2 act synergistically with caspofungin against Candida albicans biofilms. Peptides 2016, 75, 71–79. [Google Scholar] [CrossRef] [PubMed]
- De Samblanx, G.W.; Goderis, I.J.; Thevissen, K.; Raemaekers, R.; Fant, F.; Borremans, F.; Acland, D.P.; Osborn, R.W.; Patel, S.; Broekaert, W.F. Mutational analysis of a plant defensin from radish (Raphanus sativus L.) reveals two adjacent sites important for antifungal activity. J. Biol. Chem. 1997, 272, 1171–1179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fant, F.; Vranken, W.; Broekaert, W.; Borremans, F. Determination of the three-dimensional solution structure of Raphanus sativus antifungal protein 1 by 1H NMR. J. Mol. Biol. 1998, 279, 257–270. [Google Scholar] [CrossRef] [PubMed]
- Bruix, M.; Jimenez, M.A.; Santoro, J.; Gonzalez, C.; Colilla, F.J.; Mendez, E.; Rico, M. Solution structure of gamma 1-H and gamma 1-P thionins from barley and wheat endosperm determined by 1H-NMR: A structural motif common to toxic arthropod proteins. Biochemistry 1993, 32, 715–724. [Google Scholar] [CrossRef]
- Fant, F.; Vranken, W.F.; Borremans, F.A. The three-dimensional solution structure of Aesculus hippocastanum antimicrobial protein 1 determined by 1H nuclear magnetic resonance. Proteins 1999, 37, 388–403. [Google Scholar] [CrossRef]
- Almeida, M.S.; Cabral, K.M.; Kurtenbach, E.; Almeida, F.C.; Valente, A.P. Solution structure of Pisum sativum defensin 1 by high resolution NMR: Plant defensins, identical backbone with different mechanisms of action. J. Mol. Biol. 2002, 315, 749–757. [Google Scholar] [CrossRef]
- Lin, K.F.; Lee, T.R.; Tsai, P.H.; Hsu, M.P.; Chen, C.S.; Lyu, P.C. Structure-based protein engineering for alpha-amylase inhibitory activity of plant defensin. Proteins 2007, 68, 530–540. [Google Scholar] [CrossRef]
- de Paula, V.S.; Razzera, G.; Barreto-Bergter, E.; Almeida, F.C.; Valente, A.P. Portrayal of complex dynamic properties of sugarcane defensin 5 by NMR: Multiple motions associated with membrane interaction. Structure 2011, 19, 26–36. [Google Scholar] [CrossRef] [Green Version]
- Sagaram, U.S.; El-Mounadi, K.; Buchko, G.W.; Berg, H.R.; Kaur, J.; Pandurangi, R.S.; Smith, T.J.; Shah, D.M. Structural and functional studies of a phosphatidic acid-binding antifungal plant defensin MtDef4, identification of an RGFRRR motif governing fungal cell entry. PLoS ONE 2013, 8, e82485. [Google Scholar]
- Meindre, F.; Lelievre, D.; Loth, K.; Mith, O.; Aucagne, V.; Berthomieu, P.; Marques, L.; Delmas, A.F.; Landon, C.; Paquet, F. The nuclear magnetic resonance solution structure of the synthetic AhPDF1.1b plant defensin evidences the structural feature within the gamma-motif. Biochemistry 2014, 53, 7745–7754. [Google Scholar] [CrossRef] [PubMed]
- Vriens, K.; Cools, T.L.; Harvey, P.J.; Craik, D.J.; Spincemaille, P.; Cassiman, D.; Braem, A.; Vleugels, J.; Nibbering, P.H.; Drijfhout, J.W.; et al. Synergistic Activity of the Plant Defensin HsAFP1 and Caspofungin against Candida albicans Biofilms and Planktonic Cultures. PLoS ONE 2015, 10, e0132701. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lay, F.T.; Schirra, H.J.; Scanlon, M.J.; Anderson, M.A.; Craik, D.J. The three-dimensional solution structure of NaD1, a new floral defensin from Nicotiana alata and its application to a homology model of the crop defense protein alfAFP. J. Mol. Biol. 2003, 325, 175–188. [Google Scholar] [CrossRef]
- Lay, F.T.; Mills, G.D.; Poon, I.K.; Cowieson, N.P.; Kirby, N.; Baxter, A.A.; van der Weerden, N.L.; Dogovski, C.; Perugini, M.A.; Anderson, M.A.; et al. Dimerization of plant defensin NaD1 enhances its antifungal activity. J. Biol. Chem. 2012, 287, 19961–19972. [Google Scholar] [CrossRef] [Green Version]
- Poon, I.; Baxter, A.A.; Lay, F.T.; Mills, G.D.; Adda, C.G.; Payne, J.A.; Phan, T.K.; Ryan, G.F.; White, J.A.; Veneer, P.K.; et al. Phosphoinositide-mediated oligomerization of a defensin induces cell lysis. Elife 2014, 3, e01808. [Google Scholar] [CrossRef]
- Jarva, M.; Lay, F.T.; Phan, T.K.; Humble, C.; Poon, I.K.H.; Bleackley, M.R.; Anderson, M.A.; Hulett, M.D.; Kvansakul, M. X-ray structure of a carpet-like antimicrobial defensin-phospholipid membrane disruption complex. Nat. Commun. 2018, 9, 1962–1972. [Google Scholar] [CrossRef]
- Kvansakul, M.; Lay, F.T.; Adda, C.G.; Veneer, P.K.; Baxter, A.A.; Phan, T.K.; Poon, I.K.; Hulett, M.D. Binding of phosphatidic acid by NsD7 mediates the formation of helical defensin-lipid oligomeric assemblies and membrane permeabilization. Proc. Natl. Acad. Sci. USA 2016, 113, 11202–11207. [Google Scholar] [CrossRef] [Green Version]
- Jarva, M.; Lay, F.T.; Hulett, M.D.; Kvansakul, M. Structure of the defensin NsD7 in complex with PIP2 reveals that defensin: Lipid oligomer topologies are dependent on lipid type. FEBS Lett. 2017, 591, 2482–2490. [Google Scholar] [CrossRef] [Green Version]
- Lay, F.T.; Ryan, G.F.; Caria, S.; Phan, T.K.; Veneer, P.K.; White, J.A.; Kvansakul, M.; Hulett, M.D. Structural and functional characterization of the membrane-permeabilizing activity of Nicotiana occidentalis defensin NoD173 and protein engineering to enhance oncolysis. FASEB J. 2019, 33, 6470–6482. [Google Scholar] [CrossRef]
- Song, X.; Zhang, M.; Zhou, Z.; Gong, W. Ultra-high resolution crystal structure of a dimeric defensin SPE10. FEBS Lett. 2011, 585, 300–306. [Google Scholar] [CrossRef] [Green Version]
- Baxter, A.A.; Richter, V.; Lay, F.T.; Poon, I.K.; Adda, C.G.; Veneer, P.K.; Phan, T.K.; Bleackley, M.R.; Anderson, M.A.; Hulett, M.D.; et al. The Tomato Defensin TPP3 Binds Phosphatidylinositol (4,5)-Bisphosphate via a Conserved Dimeric Cationic Grip Conformation To Mediate Cell Lysis. Mol. Cell Biol. 2015, 35, 1964–1978. [Google Scholar] [CrossRef] [Green Version]
- Ochiai, A.; Ogawa, K.; Fukuda, M.; Suzuki, M.; Ito, K.; Tanaka, T.; Sagehashi, Y.; Taniguchi, M. Crystal structure of rice defensin OsAFP1 and molecular insight into lipid-binding. J. Biosci. Bioeng. 2020, 130, 6–13. [Google Scholar] [CrossRef]
- Shafee, T.M.; Lay, F.T.; Hulett, M.D.; Anderson, M.A. The Defensins Consist of Two Independent, Convergent Protein Superfamilies. Mol. Biol. Evol. 2016, 33, 2345–2356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shafee, T.; Anderson, M.A. A quantitative map of protein sequence space for the cis-defensin superfamily. Bioinformatics 2019, 35, 743–752. [Google Scholar] [CrossRef] [PubMed]
- Betz, S.F. Disulfide bonds and the stability of globular proteins. Protein Sci. Publ. Protein Soc. 1993, 2, 1551–1558. [Google Scholar] [CrossRef]
- Tam, J.P.; Wang, S.; Wong, K.H.; Tan, W.L. Antimicrobial Peptides from Plants. Pharmaceuticals (Basel) 2015, 8, 711–757. [Google Scholar] [CrossRef] [PubMed]
- Broekaert, W.F.; Cammue, B.P.A.; De Bolle, M.F.C.; Thevissen, K.; De Samblanx, G.W.; Osborn, R.W.; Nielson, K. Antimicrobial Peptides from Plants. Crit. Rev. Plant Sci. 1997, 16, 297–323. [Google Scholar] [CrossRef]
- Ochiai, A.; Ogawa, K.; Fukuda, M.; Ohori, M.; Kanaoka, T.; Tanaka, T.; Taniguchi, M.; Sagehashi, Y. Rice Defensin OsAFP1 is a New Drug Candidate against Human Pathogenic Fungi. Sci. Rep. 2018, 8, 11434. [Google Scholar] [CrossRef] [PubMed]
- Shwaiki, L.N.; Arendt, E.K.; Lynch, K.M. Anti-yeast activity and characterisation of synthetic radish peptides Rs-AFP1 and Rs-AFP2 against food spoilage yeast. Food Control 2020, 113, 107178. [Google Scholar] [CrossRef]
- Chan, Y.S.; Ng, T.B. Northeast Red Beans Produce a Thermostable and pH-Stable Defensin-Like Peptide with Potent Antifungal Activity. Cell Biochem. Biophys. 2013, 66, 637–648. [Google Scholar] [CrossRef]
- Chan, Y.S.; Wong, J.H.; Fang, E.F.; Pan, W.L.; Ng, T.B. An antifungal peptide from Phaseolus vulgaris cv. brown kidney bean. Acta Biochim. Biophys. Sin. (Shanghai) 2012, 44, 307–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leung, E.H.; Wong, J.H.; Ng, T.B. Concurrent purification of two defense proteins from French bean seeds: A defensin-like antifungal peptide and a hemagglutinin. J. Pept. Sci. 2008, 14, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, M.; Arendt, E.K.; Thery, T.L.C. Isolation and characterisation of the antifungal activity of the cowpea defensin Cp-thionin II. Food Microbiol. 2019, 82, 504–514. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Decuadro, S.; Dans, P.D.; Borba, M.A.; Benko-Iseppon, A.M.; Cecchetto, G. Gene isolation and structural characterization of a legume tree defensin with a broad spectrum of antimicrobial activity. Planta 2019, 250, 1757–1772. [Google Scholar] [CrossRef]
- Song, X.; Zhou, Z.; Wang, J.; Wu, F.; Gong, W. Purification, characterization and preliminary crystallographic studies of a novel plant defensin from Pachyrrhizus erosus seeds. Acta Crystallogr. D Biol. Crystallogr. 2004, 60, 1121–1124. [Google Scholar] [CrossRef]
- Guillen-Chable, F.; Arenas-Sosa, I.; Islas-Flores, I.; Corzo, G.; Martinez-Liu, C.; Estrada, G. Antibacterial activity and phospholipid recognition of the recombinant defensin J1-1 from Capsicum genus. Protein Expr. Purif. 2017, 136, 45–51. [Google Scholar] [CrossRef]
- Tantong, S.; Pringsulaka, O.; Weerawanich, K.; Meeprasert, A.; Rungrotmongkol, T.; Sarnthima, R.; Roytrakul, S.; Sirikantaramas, S. Two novel antimicrobial defensins from rice identified by gene coexpression network analyses. Peptides 2016, 84, 7–16. [Google Scholar] [CrossRef]
- Islam, K.T.; Velivelli, S.L.S.; Berg, R.H.; Oakley, B.; Shah, D.M. A novel bi-domain plant defensin MtDef5 with potent broad-spectrum antifungal activity binds to multiple phospholipids and forms oligomers. Sci. Rep. 2017, 7, 16157. [Google Scholar] [CrossRef]
- Parisi, K.; Shafee, T.M.A.; Quimbar, P.; van der Weerden, N.L.; Bleackley, M.R.; Anderson, M.A. The evolution, function and mechanisms of action for plant defensins. Semin. Cell Dev. Biol. 2019, 88, 107–118. [Google Scholar] [CrossRef]
- Shai, Y. Mechanism of the binding, insertion and destabilization of phospholipid bilayer membranes by alpha-helical antimicrobial and cell non-selective membrane-lytic peptides. Biochim. Biophys. Acta 1999, 1462, 55–70. [Google Scholar] [CrossRef] [Green Version]
- Ermakova, E.; Zuev, Y. Interaction of Scots Pine Defensin with Model Membrane by Coarse-Grained Molecular Dynamics. J. Membr. Biol. 2017, 250, 205–216. [Google Scholar] [CrossRef]
- Beccaccioli, M.; Reverberi, M.; Scala, V. Fungal lipids: Biosynthesis and signalling during plant-pathogen interaction. Front. Biosci. (Landmark Ed) 2019, 24, 172–185. [Google Scholar] [PubMed]
- Bowman, S.M.; Free, S.J. The structure and synthesis of the fungal cell wall. Bioessays 2006, 28, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Schoffelmeer, E.A.; Klis, F.M.; Sietsma, J.H.; Cornelissen, B.J. The cell wall of Fusarium oxysporum. Fungal Genet. Biol. 1999, 27, 275–282. [Google Scholar] [CrossRef] [Green Version]
- Thevissen, K.; Terras, F.R.; Broekaert, W.F. Permeabilization of fungal membranes by plant defensins inhibits fungal growth. Appl. Environ. Microbiol. 1999, 65, 5451–5458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yount, N.Y.; Yeaman, M.R. Multidimensional signatures in antimicrobial peptides. Proc. Natl. Acad. Sci. USA 2004, 101, 7363–7368. [Google Scholar] [CrossRef] [Green Version]
- Yount, N.Y.; Yeaman, M.R. Structural congruence among membrane-active host defense polypeptides of diverse phylogeny. Biochim. Biophys. Acta 2006, 1758, 1373–1386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Oliveira Mello, E.; Taveira, G.B.; de Oliveira Carvalho, A.; Gomes, V.M. Improved smallest peptides based on positive charge increase of the gamma-core motif from PnuD1 and their mechanism of action against Candida species. Int. J. Nanomed. 2019, 14, 407–420. [Google Scholar] [CrossRef] [Green Version]
- Sathoff, A.E.; Velivelli, S.; Shah, D.M.; Samac, D.A. Plant Defensin Peptides have Antifungal and Antibacterial Activity Against Human and Plant Pathogens. Phytopathology 2019, 109, 402–408. [Google Scholar] [CrossRef] [Green Version]
- Avitabile, C.; Capparelli, R.; Rigano, M.M.; Fulgione, A.; Barone, A.; Pedone, C.; Romanelli, A. Antimicrobial peptides from plants: Stabilization of the gamma core of a tomato defensin by intramolecular disulfide bond. J. Pept. Sci. 2013, 19, 240–245. [Google Scholar] [CrossRef] [Green Version]
- Rigano, M.M.; Romanelli, A.; Fulgione, A.; Nocerino, N.; D’Agostino, N.; Avitabile, C.; Frusciante, L.; Barone, A.; Capuano, F.; Capparelli, R. A novel synthetic peptide from a tomato defensin exhibits antibacterial activities against Helicobacter pylori. J. Pept. Sci. 2012, 18, 755–762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Souza, G.S.; de Carvalho, L.P.; de Melo, E.J.T.; da Silva, F.C.V.; Machado, O.L.T.; Gomes, V.M.; de Oliveira Carvalho, A. A synthetic peptide derived of the beta2-beta3 loop of the plant defensin from Vigna unguiculata seeds induces Leishmania amazonensis apoptosis-like cell death. Amino Acids. 2019, 51, 1633–1648. [Google Scholar] [CrossRef] [PubMed]
- Kaewklom, S.; Wongchai, M.; Petvises, S.; Hanpithakphong, W.; Aunpad, R. Structural and biological features of a novel plant defensin from Brugmansia x candida. PLoS ONE 2018, 13, e0201668. [Google Scholar] [CrossRef]
- Munoz, A.; Chu, M.; Marris, P.I.; Sagaram, U.S.; Kaur, J.; Shah, D.M.; Read, N.D. Specific domains of plant defensins differentially disrupt colony initiation, cell fusion and calcium homeostasis in Neurospora crassa. Mol. Microbiol. 2014, 92, 1357–1374. [Google Scholar] [CrossRef]
- Bleackley, M.R.; Payne, J.A.; Hayes, B.M.; Durek, T.; Craik, D.J.; Shafee, T.M.; Poon, I.K.; Hulett, M.D.; van der Weerden, N.L.; Anderson, M.A. Nicotiana alata Defensin Chimeras Reveal Differences in the Mechanism of Fungal and Tumor Cell Killing and an Enhanced Antifungal Variant. Antimicrob. Agents Chemother. 2016, 60, 6302–6312. [Google Scholar] [CrossRef] [Green Version]
- Velivelli, S.L.S.; Islam, K.T.; Hobson, E.; Shah, D.M. Modes of Action of a Bi-domain Plant Defensin MtDef5 Against a Bacterial Pathogen Xanthomonas campestris. Front. Microbiol. 2018, 9, 934. [Google Scholar] [CrossRef]
- de Medeiros, L.N.; Angeli, R.; Sarzedas, C.G.; Barreto-Bergter, E.; Valente, A.P.; Kurtenbach, E.; Almeida, F.C. Backbone dynamics of the antifungal Psd1 pea defensin and its correlation with membrane interaction by NMR spectroscopy. Biochim. Biophys. Acta. 2010, 1798, 105–113. [Google Scholar] [CrossRef] [Green Version]
- Chen, K.C.; Lin, C.Y.; Kuan, C.C.; Sung, H.Y.; Chen, C.S. A novel defensin encoded by a mungbean cDNA exhibits insecticidal activity against bruchid. J. Agric. Food Chem. 2002, 50, 7258–7263. [Google Scholar] [CrossRef]
- Kovaleva, V.; Kiyamova, R.; Cramer, R.; Krynytskyy, H.; Gout, I.; Filonenko, V.; Gout, R. Purification and molecular cloning of antimicrobial peptides from Scots pine seedlings. Peptides 2009, 30, 2136–2143. [Google Scholar] [CrossRef]
- Bloch, C., Jr.; Richardson, M. A new family of small (5 kDa) protein inhibitors of insect alpha-amylases from seeds or sorghum (Sorghum bicolar (L.) Moench) have sequence homologies with wheat gamma-purothionins. FEBS Lett. 1991, 279, 101–104. [Google Scholar] [CrossRef] [Green Version]
- Mendez, E.; Moreno, A.; Colilla, F.; Pelaez, F.; Limas, G.G.; Mendez, R.; Soriano, F.; Salinas, M.; de Haro, C. Primary structure and inhibition of protein synthesis in eukaryotic cell-free system of a novel thionin, gamma-hordothionin, from barley endosperm. Eur. J. Biochem. 1990, 194, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Buonocore, V.; Poerio, E. Interaction of Tenebrio molitor L. alpha-amylase with a wheat flour protein inhibitor. FEBS Lett. 1976, 67, 202–206. [Google Scholar] [CrossRef] [Green Version]
- Franco, O.L.; Rigden, D.J.; Melo, F.R.; Grossi-De-Sa, M.F. Plant alpha-amylase inhibitors and their interaction with insect alpha-amylases. Eur. J. Biochem. 2002, 269, 397–412. [Google Scholar] [CrossRef] [PubMed]
- Nahoum, V.; Farisei, F.; Le-Berre-Anton, V.; Egloff, M.P.; Rouge, P.; Poerio, E.; Payan, F. A plant-seed inhibitor of two classes of alpha-amylases: X-ray analysis of Tenebrio molitor larvae alpha-amylase in complex with the bean Phaseolus vulgaris inhibitor. Acta Crystallogr. D Biol. Crystallogr. 1999, 55, 360–362. [Google Scholar] [CrossRef] [PubMed]
- Pereira, P.J.; Lozanov, V.; Patthy, A.; Huber, R.; Bode, W.; Pongor, S.; Strobl, S. Specific inhibition of insect alpha-amylases: Yellow meal worm alpha-amylase in complex with the amaranth alpha-amylase inhibitor at 2.0 A resolution. Structure 1999, 7, 1079–1088. [Google Scholar] [CrossRef] [Green Version]
- Strobl, S.; Maskos, K.; Wiegand, G.; Huber, R.; Gomis-Ruth, F.X.; Glockshuber, R. A novel strategy for inhibition of alpha-amylases: Yellow meal worm alpha-amylase in complex with the Ragi bifunctional inhibitor at 2.5 A resolution. Structure 1998, 6, 911–921. [Google Scholar] [CrossRef] [Green Version]
- Strobl, S.; Maskos, K.; Betz, M.; Wiegand, G.; Huber, R.; Gomis-Ruth, F.X.; Glockshuber, R. Crystal structure of yellow meal worm alpha-amylase at 1.64 A resolution. J. Mol. Biol. 1998, 278, 617–628. [Google Scholar] [CrossRef]
- Kozakov, D.; Hall, D.R.; Xia, B.; Porter, K.A.; Padhorny, D.; Yueh, C.; Beglov, D.; Vajda, S. The ClusPro web server for protein-protein docking. Nat. Protoc. 2017, 12, 255–278. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Jablonska, J.; Pravda, L.; Varekova, R.S.; Thornton, J.M. PDBsum: Structural summaries of PDB entries. Protein Sci. 2018, 27, 129–134. [Google Scholar] [CrossRef]
- do Nascimento, V.V.; Mello Éde, O.; Carvalho, L.P.; de Melo, E.J.; Carvalho Ade, O.; Fernandes, K.V.; Gomes, V.M. PvD1 defensin, a plant antimicrobial peptide with inhibitory activity against Leishmania amazonensis. Biosci. Rep. 2015, 35, e00248. [Google Scholar] [CrossRef] [Green Version]
- Cools, T.L.; Vriens, K.; Struyfs, C.; Verbandt, S.; Ramada, M.H.S.; Brand, G.D.; Bloch, C., Jr.; Koch, B.; Traven, A.; Drijfhout, J.W.; et al. The Antifungal Plant Defensin HsAFP1 Is a Phosphatidic Acid-Interacting Peptide Inducing Membrane Permeabilization. Front. Microbiol. 2017, 8, 2295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lobo, D.S.; Pereira, I.B.; Fragel-Madeira, L.; Medeiros, L.N.; Cabral, L.M.; Faria, J.; Bellio, M.; Campos, R.C.; Linden, R.; Kurtenbach, E. Antifungal Pisum sativum defensin 1 interacts with Neurospora crassa cyclin F related to the cell cycle. Biochemistry 2007, 46, 987–996. [Google Scholar] [CrossRef]
- Sugiarto, H.; Yu, P.L. Mechanisms of action of ostrich beta-defensins against Escherichia coli. FEMS Microbiol. Lett. 2007, 270, 195–200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kovaleva, V.; Cramer, R.; Krynytskyy, H.; Gout, I.; Gout, R. Analysis of tyrosine phosphorylation and phosphotyrosine-binding proteins in germinating seeds from Scots pine. Plant Physiol. Biochem. 2013, 67, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Valente, A.P.; de Paula, V.S.; Almeida, F.C. Revealing the properties of plant defensins through dynamics. Molecules 2013, 18, 11311–11326. [Google Scholar] [CrossRef] [PubMed]
- Ikeya, T.; Ban, D.; Lee, D.; Ito, Y.; Kato, K.; Griesinger, C. Solution NMR views of dynamical ordering of biomacromolecules. Biochim. Biophys. Acta 2018, 1862, 287–306. [Google Scholar] [CrossRef]
- Sekhar, A.; Kay, L.E. An NMR View of Protein Dynamics in Health and Disease. Annu. Rev. Biophys. 2019, 48, 297–319. [Google Scholar] [CrossRef]
- Lisi, G.P.; Loria, J.P. Using NMR spectroscopy to elucidate the role of molecular motions in enzyme function. Prog. Nucl. Mag. Res. Sp. 2016, 92, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Palmer, A.G., 3rd. NMR characterization of the dynamics of biomacromolecules. Chem. Rev. 2004, 104, 3623–3640. [Google Scholar] [CrossRef]
- Palmer, A.G., 3rd. A dynamic look backward and forward. J. Magn. Reson. 2016, 266, 73–80. [Google Scholar] [CrossRef] [Green Version]
- Dishman, A.F.; Lee, M.W.; de Anda, J.; Lee, E.Y.; He, J.; Huppler, A.R.; Wong, G.C.L.; Volkman, B.F. Switchable Membrane Remodeling and Antifungal Defense by Metamorphic Chemokine XCL1. ACS Infect. Dis. 2020, 6, 1204–1213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harvey, S.R.; Porrini, M.; Konijnenberg, A.; Clarke, D.J.; Tyler, R.C.; Langridge-Smith, P.R.; MacPhee, C.E.; Volkman, B.F.; Barran, P.E. Dissecting the dynamic conformations of the metamorphic protein lymphotactin. J. Phys. Chem. B 2014, 118, 12348–12359. [Google Scholar] [CrossRef] [PubMed]
- Nevins, A.M.; Subramanian, A.; Tapia, J.L.; Delgado, D.P.; Tyler, R.C.; Jensen, D.R.; Ouellette, A.J.; Volkman, B.F. A Requirement for Metamorphic Interconversion in the Antimicrobial Activity of Chemokine XCL1. Biochemistry 2016, 55, 3784–3793. [Google Scholar] [CrossRef] [PubMed]






| Defensin and Source | pI | ΔN | IC50 (µM) | Experimental Conditions | Reference | ||
|---|---|---|---|---|---|---|---|
| Fusarium Oxysporum | Fusarium Graminearum | Botrytis Cinerea | |||||
| Rs-AFP1 Raphanus sativus | 8.3 | +3 | 2.6 2.6–5.2 | 1.4 1.4 1.4 | ½ PDB, T = 24 °C SM, T = 28 °C SM, T = 28 °C | [31] [4] [20] | |
| Rs-AFP2 Raphanus sativus | 8.7 | +6 | 0.4 0.4 | 0.2–0.5 0.4 | 0.4 1.7 0.4 0.4 | ½ PDB, T = 24 °C ½ PDB, T = 24 °C SM, T = 28 °C SM, T = 28 °C PDB, T = 24 °C SM, T = 24 °C | [31] [19] [4] [20] [18] [32] |
| Ah-AMP1 Aesculus hippocastanum | 7.6 | +1 | 4.3 | ½ PDB, T = 24 °C | [19] | ||
| Dm-AMP1 Dahlia merckii | 7.6 | +1 | 2.2 14 | ½ PDB, T = 24 °C PDB, T = 22 °C | [19] [33] | ||
| Ct-AMP1 Clitoria ternatea | 8.1 | +3 | 3.5 | ½ PDB, T = 24 °C | [19] | ||
| Hs-AFP1 Heuchera sanguinea | 8.2 | +3 | 1.0 | ½ PDB, T = 24 °C | [19] | ||
| Psd1 Pisum sativum | 7.6 | +1 | >19 | PDB, T = 25 °C | [34] | ||
| Psd2 Pisum sativum | 8.1 | +3 | >19 | PDB, T= 25 °C | [34] | ||
| PhD1 Petunia hybrida | 8.5 | +6 | 0.4 | 1.4 | ½ PDB, T = 24 °C | [5] | |
| PhD2 Petunia hybrida | 8.3 | +5 | 1.4 | 1.9 | ½ PDB, T = 24 °C | [5] | |
| VrD1 Vigna radiata | 8.7 | +6 | 1.0–3.4 | T = 28 °C | [35] | ||
| SPE10 Pachyrrhizus erosus | 7.5 | +1 | >18 | >18 | PDB, T = 28 °C | [36] | |
| VaD1 Azuki Bean | 8.9 | +7 | 5.8–10 | not listed | [37] | ||
| TvD1 Tephrosia villosa | 7.8 | +2 | 1.2 | PDB, T = 28 °C | [38] | ||
| PDC1 Zea mays | 8.1 | +3 | ~1.1 | PDB, T = 28 °C | [39] | ||
| NmDef02 Nicotiana megalosiphon | 8.1 | +3 | 1 | PDB, T = 28 °C | [40] | ||
| PsDef1 Pinus sylvestris | 8.9 | +7 | 0.5–0.7 | ~0.1 | PDB, T = 23 °C | [41] | |
| MsDef1 Medicago sativa | 8.1 | +3 | 1.2–2.3 2-4 1.5–3.0 | PDB, T = 24 °C SM SM, T = 24 °C | [18] [42] [32] | ||
| MtDef2 Medicago truncatula | 6.8 | -1 | >19 6-9 | PDB, T = 24 °C SM, T = 24 °C | [18] [32] | ||
| MtDef4 Medicago truncatula | 8.5 | +6 | 0.75–1 0.75–1.0 | SM SM, T = 24 °C | [42] [32] | ||
| Sm-AMP D1 Sm-AMP D2 Stellaria media | 7.0 7.5 | +1 +1 | 0.4 | 0.5 | 1.0 | T = 22 °C | [43] |
| PgD5 Picea glauca | 8.5 | +5 | 1.9 | 0.7 | PDB, T = 22 °C | [44] | |
| PpDFN1 Prunus persica | 2.9 | 1% glucose in H2O, T = 20 °C | [45] | ||||
| NaD1 Nicotiana alata | 8.7 | +6 | 0.4 1 | 1 ± 0.5 0.5 | 1 | PDB, T = 22 °C ½ PDB, T = 25 °C ½ PDB, T = 25 °C | [5] [11] [46] |
| NaD2 Nicotiana alata | 8.5 | +5 | 5 | 2 | [46] | ||
| Lc-def Lens culinaris | 7.8 | +2 | 9.25 | ½ PGB, T = 25 °C | [47] | ||
| AtPDFL2.1 Arabidopsis thaliana | 7.5 | +1 | 4 | ¼ PDB, T = 25 °C | [48] | ||
| ZmD32 Zea mays | 11 | +10 | 1 ± 0.7 | ½ PDB, T = 25 °C | [11] | ||
| OefDef1.1 Olea europaea | 9.1 | +8 | 0.4 ± 0.1 | 1.6 ± 0.6 | 0.7 ± 0.3 | PDB, T = 24 °C | [49] |
| Defensin and Source | PDB Code | Oligomeric State | Method | Experimental Conditions | Reference |
|---|---|---|---|---|---|
| γ1-P thionin Triticum aestivum | 1GPS | Monomer | NMR | H2O/D2O pH = 4.0, T = 22, 32 °C 1–1.5 mM | [56] |
| γ1-H thionin Hordeum vulgare | 1GPT | Monomer | NMR | H2O/D2O pH = 4.0, T = 22, 32 °C 1–1.5 mM | [56] |
| Rs-AFP1 Raphanus sativus | 1AYJ | Monomer | NMR | H2O/D2O pH = 4.2, T = 32.3 °C 1.3 mM | [55] |
| Rs-AFP2 Raphanus sativus | 2N2R | Monomer | NMR | H2O/D2O pH = 4.0, T = 15−35 °C 2 mg/mL (~0.3 mM) | [53] |
| Ah-AMP1 Aesculus hippocastanum | 1BK8 | Monomer | NMR | H2O/D2O pH = 4.1, T = 19.8 °C 3.1 mM | [57] |
| Psd1 Pisum sativum | 1JKZ | Monomer | NMR | 10 mM sodium phosphate pH = 4.0, T = 17, 27, 37 °C 1.8 mM | [58] |
| Psd2 Pisum sativum | 6NOM | Monomer | NMR | 20 mM phosphate 10 mM NaCl pH = 5.0, T = 25 °C 1 mM | [50] |
| PhD1 Petunia hybrida | 1N4N | Monomer | NMR | H2O/D2O pH = 3.1, T = 7, 37 °C 0.94 mM | [30] |
| VrD1 Vigna radiata | 1TI5 | Monomer | NMR | 50 mM phosphate pH = 6.0 T = 10, 15, 20, 25 °C 2 mM | [52] |
| VrD2 Vigna radiata | 2GL1 | Monomer | NMR | 50 mM phosphate pH = 6.0 T = 15, 20, 25 °C 2 mM | [59] |
| Sd5 Saccharum officinarum | 2KSK | Monomer | NMR | 5 mM sodium phosphate pH = 4.0 T = 25 °C 0.2–1.0 mM | [60] |
| MtDef4 Medicago truncatula | 2LR3 | Monomer | NMR | 20 mM Tris pH 4.4 T = 20 °C 1 mM | [61] |
| Lc-def Lens culinaris | 2LJ7 | Monomer | NMR | H2O/D2O pH = 5.0, T = 27, 55 °C 0.5–1.0 mM | [47] |
| AhPDF1.1 Arabidopsis halleri | 2M8B | Monomer | NMR | H2O/D2O pH = 4.5, T = 25, 35 °C 1.3 mM | [62] |
| AtPDFL2.1 Arabidopsis thaliana | 2MZ0 | Monomer | NMR | H2O/D2O pH = 6.5, T = 25 °C 0.2 mM | [48] |
| HsAFP1 Heuchera sanguinea | 2N2Q | Monomer | NMR | H2O/D2O pH = 4.0, T = 25 °C 2 mg/mL (~3 mM) | [63] |
| PsDef1 Pinus sylvestris | 5NCE | Monomer | NMR | 10 mM acetate pH 4.5, T = 33 °C 0.4-0.7 mM | [51] |
| ZmD32 Zea mays | 6DMZ | Monomer | NMR | H2O/D2O pH = 3.5, T = 25 °C 1 mM | [11] |
| NaD1 Nicotiana alata | 1MR4 4AAZ 4AB0 4CQK 6B55 | Monomer Dimer 7-mer of dimers | NMR x-ray x-ray x-ray x-ray | H2O/D2O T = 25, 30, 37 °C 1.5 mM complex with PIP2complex with PA | [64] [65] [65] [66] [67] |
| NsD7 Nicotiana suaveolens | 5KK4 5VYP | Dimer 6-mer of dimers | x-ray | complex with PA complex with PIP2 | [68] [69] |
| NoD173 Nicotiana occidentalis | 6MRY | 6-mer of dimers | x-ray | [70] | |
| SPE10 from Pachyrhizus erosus | 3PSM | Dimer | x-ray | [71] | |
| TPP3 Solanum lycopersicum | 4UJ0 | Dimer | x-ray | [72] | |
| OsAFP1 Oryza sativa | 6LCQ | Dimer | x-ray | [73] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kovaleva, V.; Bukhteeva, I.; Kit, O.Y.; Nesmelova, I.V. Plant Defensins from a Structural Perspective. Int. J. Mol. Sci. 2020, 21, 5307. https://doi.org/10.3390/ijms21155307
Kovaleva V, Bukhteeva I, Kit OY, Nesmelova IV. Plant Defensins from a Structural Perspective. International Journal of Molecular Sciences. 2020; 21(15):5307. https://doi.org/10.3390/ijms21155307
Chicago/Turabian StyleKovaleva, Valentina, Irina Bukhteeva, Oleg Y. Kit, and Irina V. Nesmelova. 2020. "Plant Defensins from a Structural Perspective" International Journal of Molecular Sciences 21, no. 15: 5307. https://doi.org/10.3390/ijms21155307






