Trimethyllysine: From Carnitine Biosynthesis to Epigenetics
Abstract
:1. Introduction
2. Carnitine Biosynthesis
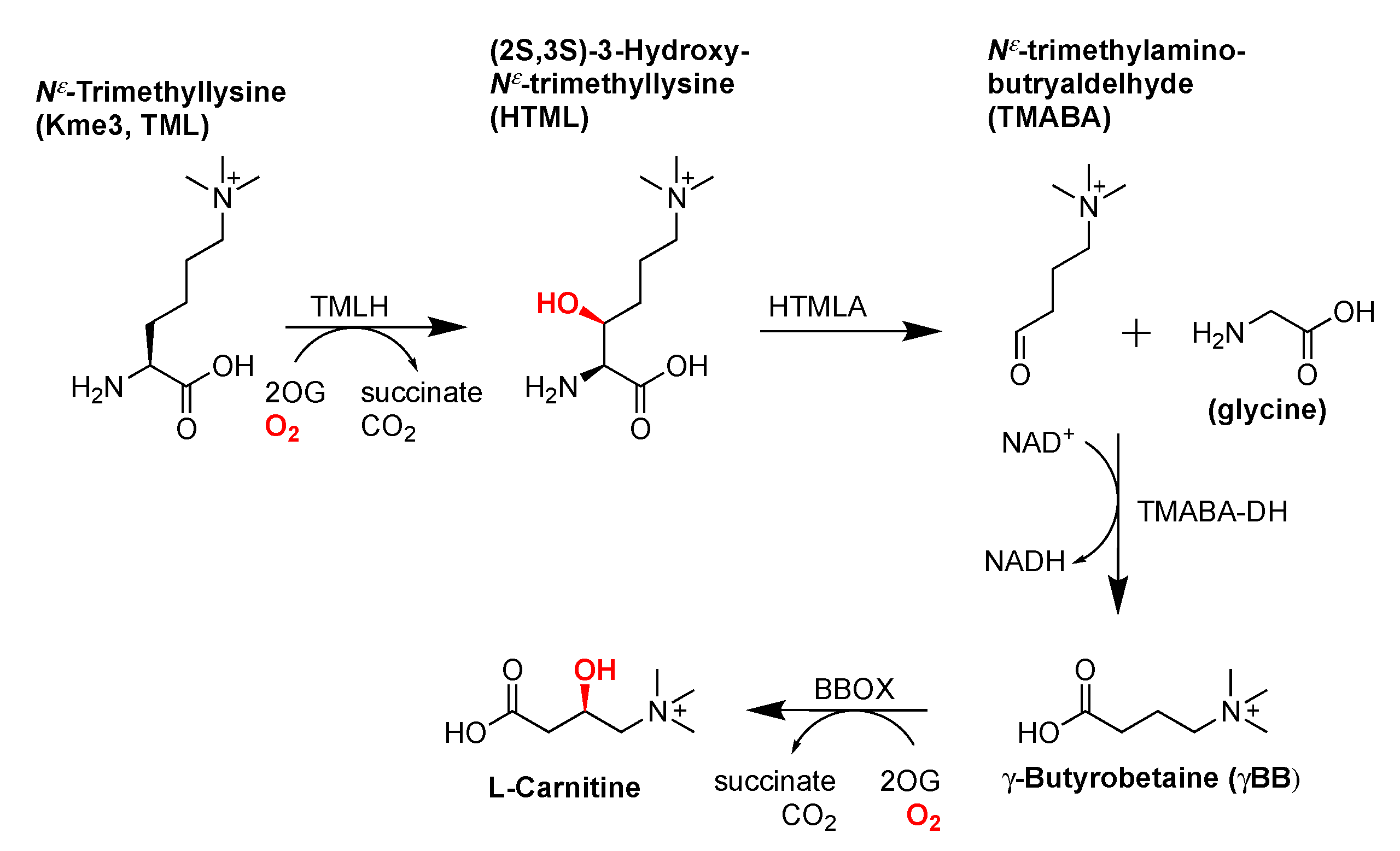
2.1. TMLH-Catalyzed C3 Hydroxylation of Kme3 and Its Role in the Carnitine Biosynthesis Pathway
2.2. Hydroxylation of Trimethyllysine Analogues by TMLH
3. Writing Kme3
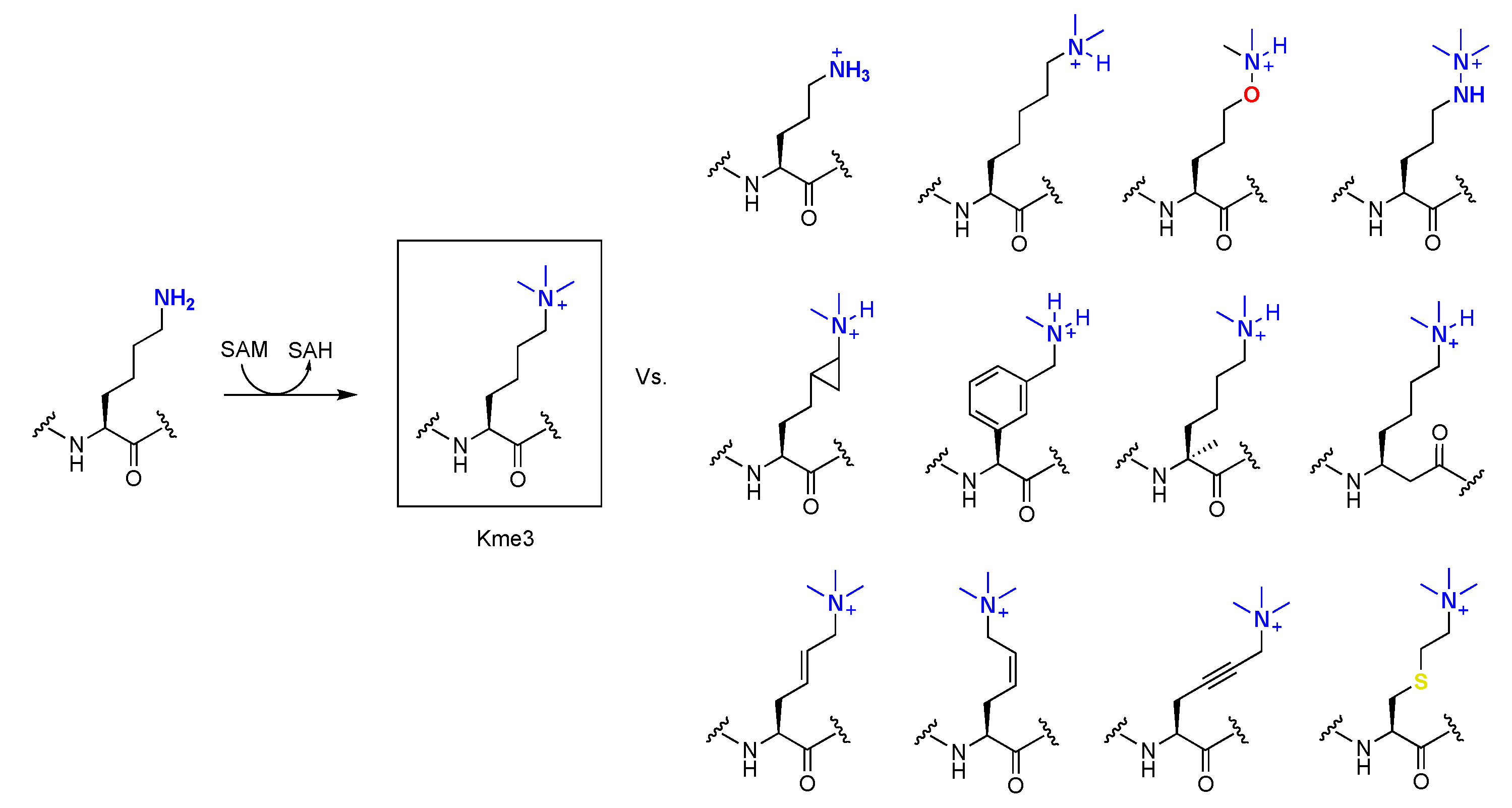
3.1. KMT-Catalyzed Formation of Kme3 Analogues
3.2. KMT-Catalyzed Formation of Kme3 Mimics in the Presence of SAM Analogues
4. Erasing Kme3
4.1. FAD-Dependent Lysine-Specific Demethylases Remove Methyl Group in Kme1 and Kme2
4.2. JmjC-Dependent Oxygenases Catalyse Removal of Methyl Group in Kme3
4.3. Demethylation of Kme3 Analogues
5. Reading Kme3
5.1. Kme3 Reader Domains
5.1.1. Recognition of Kme3 by Readers of the Royal Superfamily
5.1.2. Recognition of Kme3 by PHD Zinc Fingers
5.1.3. Recognition of Kme3 by WD40 Zinc Fingers
5.2. Modification of Reader Proteins to Investigate Biomolecular Recognition
5.3. Recognition of Kme3 Analogues by Epigenetic Readers
6. The Role of Kme3 on the Nucleosome Assembly
7. Summary and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Bischoff, R.; Schlüter, H. Amino acids: Chemistry, functionality and selected non-enzymatic post-translational modifications. J. Proteomics 2012, 75, 2275–2296. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sheehan, D. Introduction to Proteins: Structure, Function and Motion. By Amit Kessel and Nir Ben-Tal. ChemBioChem 2011, 12, 1603–1604. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information. PubChem Compound Summary for CID 5962, Lysine. 2020. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/Lysine (accessed on 13 November 2020).
- Tomé, D.; Bos, C. Lysine Requirement through the Human Life Cycle. J. Nutr. 2007, 137, 1642S–1645S. [Google Scholar] [CrossRef] [Green Version]
- Hudson, A.O.; Bless, C.; Macedo, P.; Chatterjee, S.P.; Singh, B.K.; Gilvarg, C.; Leustek, T. Biosynthesis of lysine in plants: Evidence for a variant of the known bacterial pathways. Biochim. Biophys. Acta 2005, 1721, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Fazius, F.; Shelest, E.; Gebhardt, P.; Brock, M. The fungal α-aminoadipate pathway for lysine biosynthesis requires two enzymes of the aconitase family for the isomerization of homocitrate to homoisocitrate. Mol. Microbiol. 2012, 86, 1508–1530. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Velasco, A.M.; Leguina, J.I.; Lazcano, A. Molecular Evolution of the Lysine Biosynthetic Pathways. J. Mol. Evol. 2002, 55, 445–449. [Google Scholar] [CrossRef] [PubMed]
- Luo, M. Chemical and Biochemical Perspectives of Protein Lysine Methylation. Chem. Rev. 2018, 118, 6656–6705. [Google Scholar] [CrossRef]
- Wagner, T.; Robaa, D.; Sippl, W.; Jung, M. Mind the methyl: Methyllysine binding proteins in epigenetic regulation. ChemMedChem 2014, 9, 466–483. [Google Scholar] [CrossRef]
- Andrews, F.H.; Strahl, B.D.; Kutateladze, T.G. Insights into newly discovered marks and readers of epigenetic information. Nat. Chem. Biol. 2016, 12, 662–668. [Google Scholar] [CrossRef]
- Kamps, J.J.A.G.; Huang, J.; Poater, J.; Xu, C.; Pieters, B.J.G.E.; Dong, A.; Min, J.; Sherman, W.; Beuming, T.; Matthias Bickelhaupt, F.; et al. Chemical basis for the recognition of trimethyllysine by epigenetic reader proteins. Nat. Commun. 2015, 6, 8911. [Google Scholar] [CrossRef]
- Buuh, Z.Y.; Lyu, Z.; Wang, R.E. Interrogating the Roles of Post-Translational Modifications of Non-Histone Proteins. J. Med. Chem. 2018, 61, 3239–3252. [Google Scholar] [CrossRef]
- Bannister, A.J.; Kouzarides, T. Regulation of chromatin by histone modifications. Cell Res. 2011, 21, 381–395. [Google Scholar] [CrossRef] [PubMed]
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strahl, B.D.; Allis, C.D. The language of covalent histone modifications. Nature 2000, 403, 41–45. [Google Scholar] [CrossRef]
- Taverna, S.D.; Li, H.; Ruthenburg, A.J.; Allis, C.D.; Patel, D.J. How chromatin-binding modules interpret histone modifications: Lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 2007, 14, 1025–1040. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qian, C.; Zhou, M.-M. SET domain protein lysine methyltransferases: Structure, specificity and catalysis. Cell. Mol. Life Sci. 2006, 63, 2755–2763. [Google Scholar] [CrossRef] [PubMed]
- Boriack-Sjodin, A.P.; Swinger, K.K. Protein Methyltransferases: A Distinct, Diverse, and Dynamic Family of Enzymes. Biochemistry 2016, 55, 1557–1569. [Google Scholar] [CrossRef]
- Min, J.; Feng, Q.; Li, Z.; Zhang, Y.; Xu, R.-M. Structure of the Catalytic Domain of Human DOT1L, a Non-SET Domain Nucleosomal Histone Methyltransferase. Cell 2003, 112, 711–723. [Google Scholar] [CrossRef] [Green Version]
- Husmann, D.; Gozani, O. Histone lysine methyltransferases in biology and disease. Nat. Struct. Mol. Biol. 2019, 26, 880–889. [Google Scholar] [CrossRef]
- Greer, E.L.; Shi, Y. Histone methylation: A dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 2012, 13, 343. [Google Scholar] [CrossRef] [Green Version]
- Shi, Y.; Whetstine, J.R. Dynamic Regulation of Histone Lysine Methylation by Demethylases. Mol. Cell 2007, 25, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Chinenov, Y. A second catalytic domain in the Elp3 histone acetyltransferases: A candidate for histone demethylase activity? Trends Biochem. Sci. 2002, 27, 115–117. [Google Scholar] [CrossRef]
- Hou, H.; Yu, H. Structural insights into histone lysine demethylation. Curr. Opin. Struct. Biol. 2010, 20, 739–748. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yun, M.; Wu, J.; Workman, J.L.; Li, B. Readers of histone modifications. Cell Res. 2011, 21, 564–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Porzberg, M.R.B.; Pieters, B.J.G.E.; Mecinović, J. Biomolecular Recognition of Methylated Histones. In The DNA, RNA, and Histone Methylomes; Jurga, S., Barciszewski, J., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 435–451. ISBN 978-3-030-14792-1. [Google Scholar]
- Servillo, L.; Giovane, A.; Cautela, D.; Castaldo, D.; Balestrieri, M.L. Where does Nε-trimethyllysine for the carnitine biosynthesis in mammals come from? PLoS ONE 2014, 9, e84589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strijbis, K.; Vaz, F.M.; Distel, B. Enzymology of the carnitine biosynthesis pathway. IUBMB Life 2010, 62, 357–362. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, A.; Arimondo, P.B.; Rots, M.G.; Jeronimo, C.; Berdasco, M. The timeline of epigenetic drug discovery: From reality to dreams. Clin. Epigenetics 2019, 11, 1–17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, J.R.; Polik, W.F. WebMO Enterprise, Version 19.0; WebMO LLC: Holland, MI, USA, 2019; Available online: http://www.webmo.net (accessed on 15 November 2019).
- Vaz, F.M.; Wanders, R.J.A. Carnitine biosynthesis in mammals. Biochem. J. 2002, 361, 417–429. [Google Scholar] [CrossRef]
- Reddy, Y.V.; Al Temimi, A.H.K.K.; White, P.B.; Mecinović, J. Evidence That Trimethyllysine Hydroxylase Catalyzes the Formation of (2S,3S)-3-Hydroxy-Nε-trimethyllysine. Org. Lett. 2017, 19, 400–403. [Google Scholar] [CrossRef]
- Islam, M.S.; Leissing, T.M.; Chowdhury, R.; Hopkinson, R.J.; Schofield, C.J. 2-Oxoglutarate-Dependent Oxygenases. Annu. Rev. Biochem. 2018, 87, 585–620. [Google Scholar] [CrossRef]
- McDonough, M.A.; Loenarz, C.; Chowdhury, R.; Clifton, I.J.; Schofield, C.J. Structural studies on human 2-oxoglutarate dependent oxygenases. Curr. Opin. Struct. Biol. 2010, 20, 659–672. [Google Scholar] [CrossRef] [PubMed]
- Clifton, I.J.; McDonough, M.A.; Ehrismann, D.; Kershaw, N.J.; Granatino, N.; Schofield, C.J. Structural studies on 2-oxoglutarate oxygenases and related double-stranded β-helix fold proteins. J. Inorg. Biochem. 2006, 100, 644–669. [Google Scholar] [CrossRef] [PubMed]
- Leung, I.K.H.; Krojer, T.J.; Kochan, G.T.; Henry, L.; von Delft, F.; Claridge, T.D.W.; Oppermann, U.; McDonough, M.A.; Schofield, C.J. Structural and Mechanistic Studies on γ-Butyrobetaine Hydroxylase. Chem. Biol. 2010, 17, 1316–1324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Reddy, Y.V.; Al Temimi, A.H.K.; Venselaar, H.; Nelissen, F.H.T.; Lenstra, D.C.; Mecinović, J. Investigating the active site of human trimethyllysine hydroxylase. Biochem. J. 2019, 476, 1109–1119. [Google Scholar] [CrossRef]
- Hulse, J.D.; Ellis, S.R.; Henderson, L.M. Carnitine biosynthesis. beta-Hydroxylation of trimethyllysine by an alpha-ketoglutarate-dependent mitochondrial dioxygenase. J. Biol. Chem. 1978, 253, 1654–1659. [Google Scholar]
- Sachan, D.S.; Hoppel, C.L. Carnitine biosynthesis. Hydroxylation of N6-trimethyl-lysine to 3-hydroxy-N6-trimethyl-lysine. Biochem. J. 1980, 188, 529–534. [Google Scholar] [CrossRef] [Green Version]
- Novak, R.F.; Swift, T.J.; Hoppel, C.L. N6-Trimethyl-lysine metabolism. Structural identification of the metabolite 3-hydroxy-N6-trimethyl-lysine. Biochem. J. 1980, 188, 521–527. [Google Scholar] [CrossRef] [Green Version]
- Vaz, F.M.; Ofman, R.; Westinga, K.; Back, J.W.; Wanders, R.J.A. Molecular and Biochemical Characterization of Rat ε-N-Trimethyllysine Hydroxylase, the First Enzyme of Carnitine Biosynthesis. J. Biol. Chem. 2001, 276, 33512–33517. [Google Scholar] [CrossRef] [Green Version]
- Kazaks, A.; Makrecka-Kuka, M.; Kuka, J.; Voronkova, T.; Akopjana, I.; Grinberga, S.; Pugovics, O.; Tars, K. Expression and purification of active, stabilized trimethyllysine hydroxylase. Protein Expr. Purif. 2014, 104, 1–6. [Google Scholar] [CrossRef]
- Leśniak, R.K.; Markolovic, S.; Tars, K.; Schofield, C.J. Human carnitine biosynthesis proceeds via (2S,3S )-3-hydroxy- N ε -trimethyllysine. Chem. Commun. 2017, 53, 440–442. [Google Scholar] [CrossRef] [Green Version]
- Al Temimi, A.H.; Pieters, B.J.G.; Reddy, Y.V.; White, P.B.; Mecinović, J. Substrate scope for trimethyllysine hydroxylase catalysis. Chem. Commun. 2016, 52, 12849–12852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reddy, Y.V.; Al Temimi, A.H.K.; Mecinović, J. Fluorinated trimethyllysine as a 19 F NMR probe for trimethyllysine hydroxylase catalysis. Org. Biomol. Chem. 2017, 15, 1350–1354. [Google Scholar] [CrossRef] [PubMed]
- Rydzik, A.M.; Leung, I.K.H.; Kochan, G.T.; Thalhammer, A.; Oppermann, U.; Claridge, T.D.W.; Schofield, C.J. Development and Application of a Fluoride-Detection-Based Fluorescence Assay for γ-Butyrobetaine Hydroxylase. ChemBioChem 2012, 13, 1559–1563. [Google Scholar] [CrossRef] [PubMed]
- Rydzik, A.M.; Leung, I.K.H.; Thalhammer, A.; Kochan, G.T.; Claridge, T.D.W.; Schofield, C.J. Fluoromethylated derivatives of carnitine biosynthesis intermediates—Synthesis and applications. Chem. Commun. 2014, 50, 1175–1177. [Google Scholar] [CrossRef] [Green Version]
- Lanouette, S.; Mongeon, V.; Figeys, D.; Couture, J.-F. The functional diversity of protein lysine methylation. Mol. Syst. Biol. 2014, 10. [Google Scholar] [CrossRef]
- Linscott, J.A.; Kapilashrami, K.; Wang, Z.; Senevirathne, C.; Bothwell, I.R.; Blum, G.; Luo, M. Kinetic isotope effects reveal early transition state of protein lysine methyltransferase SET8. Proc. Natl. Acad. Sci. USA 2016, 113, E8369–E8378. [Google Scholar] [CrossRef] [Green Version]
- Poulin, M.B.; Schneck, J.L.; Matico, R.E.; McDevitt, P.J.; Huddleston, M.J.; Hou, W.; Johnson, N.W.; Thrall, S.H.; Meek, T.D.; Schramm, V.L. Transition state for the NSD2-catalyzed methylation of histone H3 lysine 36. Proc. Natl. Acad. Sci. USA 2016, 113, 1197–1201. [Google Scholar] [CrossRef] [Green Version]
- Dillon, S.C.; Zhang, X.; Trievel, R.C.; Cheng, X. The SET-domain protein superfamily: Protein lysine methyltransferases. Genome Biol. 2005, 6, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Raunser, S.; Magnani, R.; Huang, Z.; Houtz, R.L.; Trievel, R.C.; Penczek, P.A.; Walz, T. Rubisco in complex with Rubisco large subunit methyltransferase. Proc. Natl. Acad. Sci. USA 2009, 106, 3160–3165. [Google Scholar] [CrossRef] [Green Version]
- Friso, G.; van Wijk, K.J. Posttranslational Protein Modifications in Plant Metabolism. Plant Physiol. 2015, 169, 1469–1487. [Google Scholar] [CrossRef] [Green Version]
- Trievel, R.C.; Flynn, E.M.; Houtz, R.L.; Hurley, J.H. Mechanism of multiple lysine methylation by the SET domain enzyme Rubisco LSMT. Nat. Struct. Mol. Biol. 2003, 10, 545–552. [Google Scholar] [CrossRef] [PubMed]
- Feng, Q.; Wang, H.; Ng, H.H.; Erdjument-Bromage, H.; Tempst, P.; Struhl, K.; Zhang, Y. Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr. Biol. 2002, 12, 1052–1058. [Google Scholar] [CrossRef] [Green Version]
- Schuhmacher, M.K.; Kudithipudi, S.; Kusevic, D.; Weirich, S.; Jeltsch, A. Activity and specificity of the human SUV39H2 protein lysine methyltransferase. Biochim. Biophys. Acta Gene Regul. Mech. 2015, 1849, 55–63. [Google Scholar] [CrossRef]
- Zhang, Y.; Mittal, A.; Reid, J.; Reich, S.; Gamblin, S.J.; Wilson, J.R. Evolving Catalytic Properties of the MLL Family SET Domain. Structure 2015, 23, 1921–1933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, H.; Min, J.; Lunin, V.V.; Antoshenko, T.; Dombrovski, L.; Zeng, H.; Allali-Hassani, A.; Campagna-Slater, V.; Vedadi, M.; Arrowsmith, C.H.; et al. Structural biology of human H3K9 methyltransferases. PLoS ONE 2010, 5, e8570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piao, L.; Nakakido, M.; Suzuki, T.; Dohmae, N.; Nakamura, Y.; Hamamoto, R. Automethylation of SUV39H2, an oncogenic histone lysine methyltransferase, regulates its binding affinity to substrate proteins. Oncotarget 2016, 7, 22846–22856. [Google Scholar] [CrossRef] [Green Version]
- Lenstra, D.C.; Al Temimi, A.H.K.; Mecinović, J. Inhibition of histone lysine methyltransferases G9a and GLP by ejection of structural Zn(II). Bioorganic Med. Chem. Lett. 2018, 28, 1234–1238. [Google Scholar] [CrossRef]
- Zhang, X.; Yang, Z.; Khan, S.I.; Horton, J.R.; Tamaru, H.; Selker, E.U.; Cheng, X. Structural Basis for the Product Specificity of Histone Lysine Methyltransferases. Mol. Cell 2003, 12, 177–185. [Google Scholar] [CrossRef] [Green Version]
- Iglesias, N.; Currie, M.A.; Jih, G.; Paulo, J.A.; Siuti, N.; Kalocsay, M.; Gygi, S.P.; Moazed, D. Automethylation-induced conformational switch in Clr4 (Suv39h) maintains epigenetic stability. Nature 2018, 560, 504–508. [Google Scholar] [CrossRef]
- Zhang, P.; Chaturvedi, C.-P.; Tremblay, V.; Cramet, M.; Brunzelle, J.S.; Skiniotis, G.; Brand, M.; Shilatifard, A.; Couture, J.-F. A phosphorylation switch on RbBP5 regulates histone H3 Lys4 methylation. Genes Dev. 2015, 29, 123–128. [Google Scholar] [CrossRef] [Green Version]
- Hsu, P.L.; Li, H.; Lau, H.-T.; Leonen, C.; Dhall, A.; Ong, S.-E.; Chatterjee, C.; Zheng, N. Crystal Structure of the COMPASS H3K4 Methyltransferase Catalytic Module. Cell 2018, 174, 1106–1116.e9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valencia-Sánchez, M.I.; De Ioannes, P.; Wang, M.; Vasilyev, N.; Chen, R.; Nudler, E.; Armache, J.-P.; Armache, K.-J. Structural Basis of Dot1L Stimulation by Histone H2B Lysine 120 Ubiquitination. Mol. Cell 2019, 74, 1010–1019. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Jing, C.; Wilson, J.R.; Walker, P.A.; Vasisht, N.; Kelly, G.; Howell, S.; Taylor, I.A.; Blackburn, M.G.; Gamblin, S.J. Structure and catalytic mechanism of the human histone methyltransferase SET7/9. Nature 2003, 421, 652–656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schapira, M. Structural chemistry of human SET domain protein methyltransferases. Curr. Chem. Genom. 2011, 5, 85–94. [Google Scholar] [CrossRef] [Green Version]
- Belle, R.; Al Temimi, A.H.K.; Kumar, K.; Pieters, B.J.G.E.; Tumber, A.; Dunford, J.E.; Johansson, C.; Oppermann, U.; Brown, T.; Schofield, C.J.; et al. Investigating D-lysine stereochemistry for epigenetic methylation, demethylation and recognition. Chem. Commun. 2017, 53, 13264–13267. [Google Scholar] [CrossRef]
- Al Temimi, A.H.K.; Reddy, Y.V.; White, P.B.; Guo, H.; Qian, P.; Mecinović, J. Lysine Possesses the Optimal Chain Length for Histone Lysine Methyltransferase Catalysis. Sci. Rep. 2017, 7, 16148. [Google Scholar] [CrossRef] [Green Version]
- Al Temimi, A.H.K.; Amatdjais-Groenen, H.I.V.; Reddy, Y.V.; Blaauw, R.H.; Guo, H.; Qian, P.; Mecinović, J. The nucleophilic amino group of lysine is central for histone lysine methyltransferase catalysis. Commun. Chem. 2019, 2, 112. [Google Scholar] [CrossRef] [Green Version]
- Al Temimi, A.H.K.; Tran, V.; Teeuwen, R.S.; Altunc, A.J.; Amatdjais-Groenen, H.I.V.; White, P.B.; Lenstra, D.C.; Proietti, G.; Wang, Y.; Wegert, A.; et al. Examining sterically demanding lysine analogs for histone lysine methyltransferase catalysis. Sci. Rep. 2020, 10, 3671. [Google Scholar] [CrossRef]
- Al Temimi, A.H.K.; Teeuwen, R.S.; Tran, V.; Altunc, A.J.; Lenstra, D.C.; Ren, W.; Qian, P.; Guo, H.; Mecinović, J. Importance of the main chain of lysine for histone lysine methyltransferase catalysis. Org. Biomol. Chem. 2019, 17, 5693–5697. [Google Scholar] [CrossRef]
- Al Temimi, A.H.K.; White, P.B.; Mulders, M.J.M.; Van Der Linden, N.G.A.; Blaauw, R.H.; Wegert, A.; Rutjes, F.P.J.T.; Mecinović, J. Methylation of geometrically constrained lysine analogues by histone lysine methyltransferases. Chem. Commun. 2020, 56, 3039–3042. [Google Scholar] [CrossRef]
- Dadová, J.; Galan, S.R.; Davis, B.G. Synthesis of modified proteins via functionalization of dehydroalanine. Curr. Opin. Chem. Biol. 2018, 46, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.A.; Liu, W.R. Proteins with Site-Specific Lysine Methylation. Chem. A Eur. J. 2017, 23, 11732–11737. [Google Scholar] [CrossRef] [PubMed]
- Al Temimi, A.H.K.; van der Wekken-de Bruijne, R.; Proietti, G.; Guo, H.; Qian, P.; Mecinović, J. γ-Thialysine versus Lysine: An Insight into the Epigenetic Methylation of Histones. Bioconjug. Chem. 2019, 30, 1798–1804. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zheng, Y.G. SAM/SAH Analogs as Versatile Tools for SAM-Dependent Methyltransferases. ACS Chem. Biol. 2016, 11, 583–597. [Google Scholar] [CrossRef] [Green Version]
- Klimasauskas, S.; Weinhold, E. A new tool for biotechnology: AdoMet-dependent methyltransferases. Trends Biotechnol. 2007, 25, 99–104. [Google Scholar] [CrossRef]
- Huber, T.D.; Johnson, B.R.; Zhang, J.; Thorson, J.S. AdoMet analog synthesis and utilization: Current state of the art. Curr. Opin. Biotechnol. 2016, 42, 189–197. [Google Scholar] [CrossRef] [Green Version]
- Peters, W.; Willnow, S.; Duisken, M.; Kleine, H.; Macherey, T.; Duncan, K.E.; Litchfield, D.W.; Lüscher, B.; Weinhold, E. Enzymatic site-specific functionalization of protein methyltransferase substrates with alkynes for click labeling. Angew. Chem. Int. Ed. 2010, 49, 5170–5173. [Google Scholar] [CrossRef]
- Wang, R.; Islam, K.; Liu, Y.; Zheng, W.; Tang, H.; Lailler, N.; Blum, G.; Deng, H.; Luo, M. Profiling Genome-wide Chromatin Methylation with Engineered Posttranslation Apparatus within Living Cells. J. Am. Chem. Soc. 2013, 135, 1048–1056. [Google Scholar] [CrossRef] [Green Version]
- Islam, K.; Zheng, W.; Yu, H.; Deng, H.; Luo, M. Expanding cofactor repertoire of protein lysine methyltransferase for substrate labeling. ACS Chem. Biol. 2011, 6, 679–684. [Google Scholar] [CrossRef]
- Islam, K.; Bothwell, I.; Chen, Y.; Sengelaub, C.; Wang, R.; Deng, H.; Luo, M. Bioorthogonal profiling of protein methylation using azido derivative of S-adenosyl-L-methionine. J. Am. Chem. Soc. 2012, 134, 5909–5915. [Google Scholar] [CrossRef] [Green Version]
- Willnow, S.; Martin, M.; Lüscher, B.; Weinhold, E. A selenium-based click AdoMet analogue for versatile substrate labeling with wild-type protein methyltransferases. ChemBioChem 2012, 13, 1167–1173. [Google Scholar] [CrossRef] [PubMed]
- Bothwell, I.R.; Luo, M. Large-Scale, Protection-Free Synthesis of Se-Adenosyl-l-selenomethionine Analogues and Their Application as Cofactor Surrogates of Methyltransferases. Org. Lett. 2014, 16, 3056–3059. [Google Scholar] [CrossRef] [PubMed]
- Bothwell, I.R.; Islam, K.; Chen, Y.; Zheng, W.; Blum, G.; Deng, H.; Luo, M. Se-adenosyl-L-selenomethionine cofactor analogue as a reporter of protein methylation. J. Am. Chem. Soc. 2012, 134, 14905–14912. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al Temimi, A.H.K.; Martin, M.; Meng, Q.; Lenstra, D.C.; Qian, P.; Guo, H.; Weinhold, E.; Mecinovic, J. Lysine Ethylation by Histone Lysine Methyltransferases. ChemBioChem 2020, 21, 392–400. [Google Scholar] [CrossRef]
- Walport, L.J.; Hopkinson, R.J.; Schofield, C.J. Mechanisms of human histone and nucleic acid demethylases. Curr. Opin. Chem. Biol. 2012, 16, 525–534. [Google Scholar] [CrossRef]
- Shi, Y.; Lan, F.; Matson, C.; Mulligan, P.; Whetstine, J.R.; Cole, P.A.; Casero, R.A.; Shi, Y. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 2004, 119, 941–953. [Google Scholar] [CrossRef] [Green Version]
- Mosammaparast, N.; Shi, Y. Reversal of Histone Methylation: Biochemical and Molecular Mechanisms of Histone Demethylases. Annu. Rev. Biochem. 2010, 79, 155–179. [Google Scholar] [CrossRef]
- Kooistra, S.M.; Helin, K. Molecular mechanisms and potential functions of histone demethylases. Nat. Rev. Mol. Cell Biol. 2012, 13, 297–311. [Google Scholar] [CrossRef]
- Wojcieszyńska, D.; Hupert-Kocurek, K.; Guzik, U. Flavin-dependent enzymes in cancer prevention. Int. J. Mol. Sci. 2012, 13, 16751–16768. [Google Scholar] [CrossRef] [Green Version]
- Baron, R.; Binda, C.; Tortorici, M.; McCammon, J.A.; Mattevi, A. Molecular mimicry and ligand recognition in binding and catalysis by the histone demethylase LSD1-CoREST complex. Structure 2011, 19, 212–220. [Google Scholar] [CrossRef] [Green Version]
- Yang, M.; Gocke, C.B.; Luo, X.; Borek, D.; Tomchick, D.R.; Machius, M.; Otwinowski, Z.; Yu, H. Structural basis for CoREST-dependent demethylation of nucleosomes by the human LSD1 histone demethylase. Mol. Cell 2006, 23, 377–387. [Google Scholar] [CrossRef] [PubMed]
- Stavropoulos, P.; Blobel, G.; Hoelz, A. Crystal structure and mechanism of human lysine-specific demethylase-1. Nat. Struct. Mol. Biol. 2006, 13, 626–632. [Google Scholar] [CrossRef]
- Ciccone, D.N.; Su, H.; Hevi, S.; Gay, F.; Lei, H.; Bajko, J.; Xu, G.; Li, E.; Chen, T. KDM1B is a histone H3K4 demethylase required to establish maternal genomic imprints. Nature 2009, 461, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Sengupta, R.; Espejo, A.B.; Lee, M.G.; Dorsey, J.A.; Richter, M.; Opravil, S.; Shiekhattar, R.; Bedford, M.T.; Jenuwein, T.; et al. p53 is regulated by the lysine demethylase LSD1. Nature 2007, 449, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Kontaki, H.; Talianidis, I. Lysine methylation regulates E2F1-induced cell death. Mol. Cell 2010, 39, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hevi, S.; Kurash, J.K.; Lei, H.; Gay, F.; Bajko, J.; Su, H.; Sun, W.; Chang, H.; Xu, G.; et al. The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat. Genet. 2009, 41, 125–129. [Google Scholar] [CrossRef]
- Cloos, P.A.C.C.; Christensen, J.; Agger, K.; Helin, K. Erasing the methyl mark: Histone demethylases at the center of cellular differentiation and disease. Genes Dev. 2008, 22, 1115–1140. [Google Scholar] [CrossRef] [Green Version]
- Culhane, J.C.; Cole, P.A. LSD1 and The Chemistry of Histone Demethylation. Curr. Opin. Chem. Biol. 2007, 11, 561–568. [Google Scholar] [CrossRef] [Green Version]
- Forneris, F.; Battaglioli, E.; Mattevi, A.; Binda, C. New roles of flavoproteins in molecular cell biology: Histone demethylase LSD1 and chromatin. FEBS J. 2009, 276, 4304–4312. [Google Scholar] [CrossRef]
- Yang, M.; Culhane, J.C.; Szewczuk, L.M.; Gocke, C.B.; Brautigam, C.A.; Tomchick, D.R.; Machius, M.; Cole, P.A.; Yu, H. Structural basis of histone demethylation by LSD1 revealed by suicide inactivation. Nat. Struct. Mol. Biol. 2007, 14. [Google Scholar] [CrossRef]
- Tsukada, Y.; Fang, J.; Erdjument-Bromage, H.; Warren, M.E.; Borchers, C.H.; Tempst, P.; Zhang, Y. Histone demethylation by a family of JmjC domain-containing proteins. Nature 2006, 439, 811–816. [Google Scholar] [CrossRef] [PubMed]
- Thinnes, C.C.; England, K.S.; Kawamura, A.; Chowdhury, R.; Schofield, C.J.; Hopkinson, R.J. Targeting histone lysine demethylases—Progress, challenges, and the future. Biochim. Biophys. Acta 2014, 1839, 1416–1432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walport, L.J.; Hopkinson, R.J.; Chowdhury, R.; Schiller, R.; Ge, W.; Kawamura, A.; Schofield, C.J. Arginine demethylation is catalysed by a subset of JmjC histone lysine demethylases. Nat. Commun. 2016, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Jing, L.; Li, M.; He, L.; Guo, Z. Regulation of histone arginine methylation/demethylation by methylase and demethylase (Review). Mol. Med. Rep. 2019, 49, 3963–3971. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, B.; Chen, Y.; Zhao, Y.; Bruick, R.K. JMJD6 is a histone arginine demethylase. Science 2007, 318, 444–447. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johansson, C.; Tumber, A.; Che, K.; Cain, P.; Nowak, R.; Gileadi, C.; Oppermann, U. The roles of Jumonji-type oxygenases in human disease. Epigenomics 2014, 6, 89–120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ng, S.S.; Kavanagh, K.L.; McDonough, M.A.; Butler, D.; Pilka, E.S.; Lienard, B.M.; Bray, J.E.; Savitsky, P.; Gileadi, O.; von Delft, F.; et al. Crystal structures of histone demethylase JMJD2A reveal basis for substrate specificity. Nature 2007, 448, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, A.K.; Horton, J.R.; Zhang, X.; Cheng, X. Coordinated methyl-lysine erasure: Structural and functional linkage of a Jumonji demethylase domain and a reader domain. Curr. Opin. Struct. Biol. 2011, 21, 750–760. [Google Scholar] [CrossRef] [Green Version]
- Martinez, S.; Hausinger, R.P. Catalytic Mechanisms of Fe(II)- and 2-Oxoglutarate-dependent Oxygenases. J. Biol. Chem. 2015, 290, 20702–20711. [Google Scholar] [CrossRef] [Green Version]
- Couture, J.-F.; Collazo, E.; Ortiz-Tello, P.A.; Brunzelle, J.S.; Trievel, R.C. Specificity and mechanism of JMJD2A, a trimethyllysine-specific histone demethylase. Nat. Struct. Mol. Biol. 2007, 14, 689–695. [Google Scholar] [CrossRef]
- Smith, B.C.; Denu, J.M. Chemical mechanisms of histone lysine and arginine modifications. Biochim. Biophys. Acta 2009, 1789, 45–57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hopkinson, R.J.; Walport, L.J.; Münzel, M.; Rose, N.R.; Smart, T.J.; Kawamura, A.; Claridge, T.D.W.; Schofield, C.J. Is JmjC Oxygenase Catalysis Limited to Demethylation? Angew. Chem. Int. Ed. 2013, 52, 7709–7713. [Google Scholar] [CrossRef] [Green Version]
- Hopkinson, R.J.; Langley, G.W.; Belle, R.; Walport, L.J.; Dunne, K.; Münzel, M.; Salah, E.; Kawamura, A.; Claridge, T.D.W.; Schofield, C.J. Human histone demethylase KDM6B can catalyse sequential oxidations. Chem. Commun. 2018, 54, 7975–7978. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langley, G.W.; Brinkø, A.; Münzel, M.; Walport, L.J.; Schofield, C.J.; Hopkinson, R.J. Analysis of JmjC Demethylase-Catalyzed Demethylation Using Geometrically-Constrained Lysine Analogues. ACS Chem. Biol. 2016, 11, 755–762. [Google Scholar] [CrossRef] [PubMed]
- Hagel, J.M.; Facchini, P.J. Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy. Nat. Chem. Biol. 2010, 6, 273–275. [Google Scholar] [CrossRef]
- Shiau, C.; Trnka, M.J.; Bozicevic, A.; Torres, I.O.; Al-Sady, B.; Burlingame, A.L.; Narlikar, G.J.; Fujimori, D.G. Reconstitution of nucleosome demethylation and catalytic properties of a jumonji histone demethylase. Chem. Biol. 2013, 20, 494–499. [Google Scholar] [CrossRef] [Green Version]
- Xu, C.; Cui, G.; Botuyan, M.V.; Mer, G. Methyllysine Recognition by the Royal Family Modules: Chromo, Tudor, MBT, Chromo Barrel, and PWWP Domains. In Histone Recognition; Zhou, M.-M., Ed.; Springer International Publishing: Cham, Switzerland, 2015; pp. 49–82. ISBN 978-3-319-18102-8. [Google Scholar]
- Daze, K.; Hof, F. Molecular Interaction and Recognition. In Encyclopedia of Physical Organic Chemistry; 5 Volume Set; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2016; pp. 1–51. [Google Scholar]
- Musselman, C.A.; Lalonde, M.-E.; Côté, J.; Kutateladze, T.G. Perceiving the epigenetic landscape through histone readers. Nat. Struct. Mol. Biol. 2012, 19, 1218–1227. [Google Scholar] [CrossRef] [Green Version]
- Sims, R.J.; Reinberg, D. Histone H3 Lys 4 methylation: Caught in a bind? Genes Dev. 2006, 20, 2779–2786. [Google Scholar] [CrossRef] [Green Version]
- Gallivan, J.P.; Dougherty, D.A. A Computational Study of Cation−π Interactions vs Salt Bridges in Aqueous Media: Implications for Protein Engineering. J. Am. Chem. Soc. 2000, 122, 870–874. [Google Scholar] [CrossRef]
- Musselman, C.A.; Khorasanizadeh, S.; Kutateladze, T.G. Towards understanding methyllysine readout. Biochim. Biophys. Acta 2014, 1839, 686–693. [Google Scholar] [CrossRef] [Green Version]
- Blus, B.J.; Wiggins, K.; Khorasanizadeh, S. Epigenetic virtues of chromodomains. Crit. Rev. Biochem. Mol. Biol. 2011, 46, 507–526. [Google Scholar] [CrossRef] [Green Version]
- Teske, K.A.; Hadden, M.K. Methyllysine binding domains: Structural insight and small molecule probe development. Eur. J. Med. Chem. 2017, 136, 14–35. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, P.R.; Nietlispach, D.; Buscaino, A.; Warner, R.J.; Akhtar, A.; Murzin, A.G.; Murzina, N.V.; Laue, E.D. Structure of the Chromo Barrel Domain from the MOF Acetyltransferase. J. Biol. Chem. 2005, 280, 32326–32331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eisert, R.J.; Waters, M.L. Tuning HP1α chromodomain selectivity for di- and trimethyllysine. ChemBioChem 2011, 12, 2786–2790. [Google Scholar] [CrossRef] [PubMed]
- Pieters, B.J.G.E.; Meulenbroeks, E.; Belle, R.; Mecinović, J. The Role of Electrostatic Interactions in Binding of Histone H3K4me2/3 to the Sgf29 Tandem Tudor Domain. PLoS ONE 2015, 10, e0139205. [Google Scholar] [CrossRef] [Green Version]
- Lu, R.; Wang, G.G. Tudor: A versatile family of histone methylation “readers”. Trends Biochem. Sci. 2013, 38, 546–555. [Google Scholar] [CrossRef] [Green Version]
- Pieters, B.; Belle, R.; Mecinović, J. The effect of the length of histone H3K4me3 on recognition by reader proteins. ChemBioChem 2013, 14, 2408–2412. [Google Scholar] [CrossRef]
- Bian, C.; Xu, C.; Ruan, J.; Lee, K.K.; Burke, T.L.; Tempel, W.; Barsyte, D.; Li, J.; Wu, M.; Zhou, B.O.; et al. Sgf29 binds histone H3K4me2/3 and is required for SAGA complex recruitment and histone H3 acetylation. Embo J. 2011, 30, 2829–2842. [Google Scholar] [CrossRef] [Green Version]
- Su, X.; Zhu, G.; Ding, X.; Lee, S.Y.; Dou, Y.; Zhu, B.; Wu, W.; Li, H. Molecular basis underlying histone H3 lysine-arginine methylation pattern readout by Spin/Ssty repeats of Spindlin1. Genes Dev. 2014, 28, 622–636. [Google Scholar] [CrossRef] [Green Version]
- Rona, G.B.; Eleutherio, E.C.A.; Pinheiro, A.S. PWWP domains and their modes of sensing DNA and histone methylated lysines. Biophys. Rev. 2016, 8, 63–74. [Google Scholar] [CrossRef] [Green Version]
- Qin, S.; Min, J. Structure and function of the nucleosome-binding PWWP domain. Trends Biochem. Sci. 2014, 39, 536–547. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Zeng, H.; Lam, R.; Tempel, W.; Amaya, M.F.; Xu, C.; Dombrovski, L.; Qiu, W.; Wang, Y.; Min, J. Structural and histone binding ability characterizations of human PWWP domains. PLoS ONE 2011, 6, e18919. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacobs, S.A.; Khorasanizadeh, S. Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 2002, 295, 2080–2083. [Google Scholar] [CrossRef] [PubMed]
- Musselman, C.A.; Avvakumov, N.; Watanabe, R.; Abraham, C.G.; Lalonde, M.-E.; Hong, Z.; Allen, C.; Roy, S.; Nuñez, J.K.; Nickoloff, J.; et al. Molecular basis for H3K36me3 recognition by the Tudor domain of PHF1. Nat. Struct. Mol. Biol. 2012, 19, 1266–1272. [Google Scholar] [CrossRef]
- Vezzoli, A.; Bonadies, N.; Allen, M.D.; Freund, S.M.V.; Santiveri, C.M.; Kvinlaug, B.T.; Huntly, B.J.P.; Göttgens, B.; Bycroft, M. Molecular basis of histone H3K36me3 recognition by the PWWP domain of Brpf1. Nat. Struct. Mol. Biol. 2010, 17, 617–619. [Google Scholar] [CrossRef]
- Peña, P.V.; Davrazou, F.; Shi, X.; Walter, K.L.; Verkhusha, V.V.; Gozani, O.; Zhao, R.; Kutateladze, T.G. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 2006, 442, 100. [Google Scholar] [CrossRef]
- Iwase, S.; Xiang, B.; Ghosh, S.; Ren, T.; Lewis, P.W.; Cochrane, J.C.; Allis, C.D.; Picketts, D.J.; Patel, D.J.; Li, H.; et al. ATRX ADD domain links an atypical histone methylation recognition mechanism to human mental-retardation syndrome. Nat. Struct. Mol. Biol. 2011, 18, 769–776. [Google Scholar] [CrossRef]
- Xu, C.; Bian, C.; Yang, W.; Galka, M.; Ouyang, H.; Chen, C.; Qiu, W.; Liu, H.; Jones, A.E.; MacKenzie, F.; et al. Binding of different histone marks differentially regulates the activity and specificity of polycomb repressive complex 2 (PRC2). Proc. Natl. Acad. Sci. USA 2010, 107, 19266–19271. [Google Scholar] [CrossRef] [Green Version]
- Champagne, K.S.; Kutateladze, T.G. Structural insight into histone recognition by the ING PHD fingers. Curr. Drug Targets 2009, 10, 432–441. [Google Scholar] [CrossRef] [Green Version]
- Sanchez, R.; Zhou, M.-M. The PHD Finger: A Versatile Epigenome Reader. Trends Biochem. Sci. 2011, 36, 364–372. [Google Scholar] [CrossRef] [Green Version]
- Bortoluzzi, A.; Amato, A.; Lucas, X.; Blank, M.; Ciulli, A. Structural basis of molecular recognition of helical histone H3 tail by PHD finger domains. Biochem. J. 2017, 474, 1633–1651. [Google Scholar] [CrossRef] [Green Version]
- Eustermann, S.; Yang, J.-C.; Law, M.J.; Amos, R.; Chapman, L.M.; Jelinska, C.; Garrick, D.; Clynes, D.; Gibbons, R.J.; Rhodes, D.; et al. Combinatorial readout of histone H3 modifications specifies localization of ATRX to heterochromatin. Nat. Struct. Mol. Biol. 2011, 18, 777–782. [Google Scholar] [CrossRef] [PubMed]
- Gatchalian, J.; Kutateladze, T.G. PHD fingers as histone readers. In Histone Recognition; Springer International Publishing: Cham, Switzerland, 2015; pp. 27–48. ISBN 9783319181028. [Google Scholar]
- Wilson, J.R.; Justin, N. Histone recognition by WD40 proteins. In Histone Recognition; Springer International Publishing: Cham, Switzerland, 2015; pp. 83–100. ISBN 9783319181028. [Google Scholar]
- Lee, Y.-J.; Schmidt, M.J.; Tharp, J.M.; Weber, A.; Koenig, A.L.; Zheng, H.; Gao, J.; Waters, M.L.; Summerer, D.; Liu, W.R. Genetically encoded fluorophenylalanines enable insights into the recognition of lysine trimethylation by an epigenetic reader. Chem. Commun. 2016, 52, 12606–12609. [Google Scholar] [CrossRef]
- Pieters, B.J.G.E.; Wuts, M.H.M.; Poater, J.; Kumar, K.; White, P.B.; Kamps, J.J.A.G.; Sherman, W.; Pruijn, G.J.M.; Paton, R.S.; Beuming, T.; et al. Mechanism of biomolecular recognition of trimethyllysine by the fluorinated aromatic cage of KDM5A PHD3 finger. Commun. Chem. 2020, 3, 69. [Google Scholar] [CrossRef]
- Baril, S.A.; Koenig, A.L.; Krone, M.W.; Albanese, K.I.; He, C.Q.; Lee, G.Y.; Houk, K.N.; Waters, M.L.; Brustad, E.M. Investigation of Trimethyllysine Binding by the HP1 Chromodomain via Unnatural Amino Acid Mutagenesis. J. Am. Chem. Soc. 2017, 139, 17253–17256. [Google Scholar] [CrossRef] [PubMed]
- Krone, M.W.; Albanese, K.I.; Leighton, G.O.; He, C.Q.; Lee, G.Y.; Garcia-Borràs, M.; Guseman, A.J.; Williams, D.C.; Houk, K.N.; Brustad, E.M.; et al. Thermodynamic consequences of Tyr to Trp mutations in the cation–π-mediated binding of trimethyllysine by the HP1 chromodomain. Chem. Sci. 2020, 11, 3495–3500. [Google Scholar] [CrossRef] [Green Version]
- Al Temimi, A.H.K.; Belle, R.; Kumar, K.; Poater, J.; Betlem, P.; Pieters, B.J.G.E.; Paton, R.S.; Bickelhaupt, F.M.; Mecinović, J. Recognition of shorter and longer trimethyllysine analogues by epigenetic reader proteins. Chem. Commun. 2018, 54, 2409–2412. [Google Scholar] [CrossRef] [Green Version]
- Seeliger, D.; Soeroes, S.; Klingberg, R.; Schwarzer, D.; Grubmüller, H.; Fischle, W. Quantitative Assessment of Protein Interaction with Methyl-Lysine Analogues by Hybrid Computational and Experimental Approaches. ACS Chem. Biol. 2012, 7, 150–154. [Google Scholar] [CrossRef]
- Chen, Z.; Notti, R.Q.; Ueberheide, B.; Ruthenburg, A.J. Quantitative and Structural Assessment of Histone Methyllysine Analogue Engagement by Cognate Binding Proteins Reveals Affinity Decrements Relative to Those of Native Counterparts. Biochemistry 2018, 57, 300–304. [Google Scholar] [CrossRef]
- Simon, M.D.; Chu, F.; Racki, L.R.; de la Cruz, C.C.; Burlingame, A.L.; Panning, B.; Narlikar, G.J.; Shokat, K.M. The site-specific installation of methyl-lysine analogs into recombinant histones. Cell 2007, 128, 1003–1012. [Google Scholar] [CrossRef] [Green Version]
- Munari, F.; Soeroes, S.; Zenn, H.M.; Schomburg, A.; Kost, N.; Schröder, S.; Klingberg, R.; Rezaei-Ghaleh, N.; Stützer, A.; Gelato, K.A.; et al. Methylation of lysine 9 in histone H3 directs alternative modes of highly dynamic interaction of heterochromatin protein hHP1β with the nucleosome. J. Biol. Chem. 2012, 287, 33756–33765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richards, E.J.; Elgin, S.C.R. Epigenetic codes for heterochromatin formation and silencing: Rounding up the usual suspects. Cell 2002, 108, 489–500. [Google Scholar] [CrossRef] [Green Version]
- Hintzen, J.C.J.; Poater, J.; Kumar, K.; Al Temimi, A.H.K.; Pieters, B.J.G.E.; Paton, R.S.; Bickelhaupt, F.M.; Mecinović, J. Comparison of Molecular Recognition of Trimethyllysine and Trimethylthialysine by Epigenetic Reader Proteins. Molecules 2020, 25, 1918. [Google Scholar] [CrossRef] [Green Version]
- Pieters, B.J.G.E.; Hintzen, J.C.J.; Grobben, Y.; Al Temimi, A.H.K.; Kamps, J.J.A.G.; Mecinović, J. Installation of Trimethyllysine Analogs on Intact Histones via Cysteine Alkylation. Bioconjug. Chem. 2019, 30, 952–958. [Google Scholar] [CrossRef] [Green Version]
- Lu, X.; Simon, M.D.; Chodaparambil, J.V.; Hansen, J.C.; Shokat, K.M.; Luger, K. The effect of H3K79 dimethylation and H4K20 trimethylation on nucleosome and chromatin structure. Nat. Struct. Mol. Biol. 2008, 15, 1122–1124. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.; Zhang, Y. The diverse functions of histone lysine methylation. Nat. Rev. Mol. Cell Biol. 2005, 6, 838. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.M.; Muir, T.W. Histones: At the crossroads of peptide and protein chemistry. Chem. Rev. 2015, 115, 2296–2349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, J.; Fan, J.Y.; Rangasamy, D.; Tremethick, D.J. The nucleosome surface regulates chromatin compaction and couples it with transcriptional repression. Nat. Struct. Mol. Biol. 2007, 14, 1070–1076. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.P.; Garcia Alai, M.M.; Kapadnis, P.B.; Neumann, H.; Chin, J.W. Genetically encoding Nε-methyl-L-lysine in recombinant histones. J. Am. Chem. Soc. 2009, 131, 14194–14195. [Google Scholar] [CrossRef]
- Nguyen, D.P.; Alai, M.M.G.; Virdee, S.; Chin, J.W. Genetically directing ε-N, N-dimethyl-l-lysine in recombinant histones. Chem. Biol. 2010, 17, 1072–1076. [Google Scholar] [CrossRef] [Green Version]
- He, S.; Bauman, D.; Davis, J.S.; Loyola, A.; Nishioka, K.; Gronlund, J.L.; Reinberg, D.; Meng, F.; Kelleher, N.; McCafferty, D.G. Facile synthesis of site-specifically acetylated and methylated histone proteins: Reagents for evaluation of the histone code hypothesis. Proc. Natl. Acad. Sci. USA 2003, 100, 12033–12038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muir, T.W. Semisynthesis of proteins by expressed protein ligation. Annu. Rev. Biochem. 2003, 72, 249–289. [Google Scholar] [CrossRef] [PubMed]
- Dawson, P.E.; Muir, T.W.; Clark-Lewis, I.; Kent, S.B.H. Synthesis of proteins by native chemical ligation. Science 1994, 266, 776–779. [Google Scholar] [CrossRef] [PubMed]
- Cistrone, P.A.; Bird, M.J.; Flood, D.T.; Silvestri, A.P.; Hintzen, J.C.J.; Thompson, D.A.; Dawson, P.E. Native Chemical Ligation of Peptides and Proteins. Curr. Protoc. Chem. Biol. 2019, 11, e61. [Google Scholar] [CrossRef] [Green Version]
- Kawakami, T.; Akai, Y.; Fujimoto, H.; Kita, C.; Aoki, Y.; Konishi, T.; Waseda, M.; Takemura, L.; Aimoto, S. Sequential peptide ligation by combining the CysPro ester (CPE) and thioester methods and its application to the synthesis of histone H3 containing a trimethyl lysine residue. Bull. Chem. Soc. Jpn. 2013, 86, 690–697. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Li, Y.Y.; Liu, L.; Li, Y.Y.; He, Q.; Li, H. One-pot native chemical ligation of peptide hydrazides enables total synthesis of modified histones. Org. Biomol. Chem. 2014, 12, 5435–5441. [Google Scholar] [CrossRef]
- Ruthenburg, A.J.; Li, H.; Milne, T.A.; Dewell, S.; McGinty, R.K.; Yuen, M.; Ueberheide, B.; Dou, Y.; Muir, T.W.; Patel, D.J.; et al. Recognition of a Mononucleosomal Histone Modification Pattern by BPTF via Multivalent Interactions. Cell 2011, 145, 692–706. [Google Scholar] [CrossRef] [Green Version]
- Tars, K.; Leitans, J.; Kazaks, A.; Zelencova, D.; Liepinsh, E.E.; Kuka, J.; Makrecka, M.; Lola, D.; Andrianovs, V.; Gustina, D.; et al. Targeting carnitine biosynthesis: Discovery of new inhibitors against γ-butyrobetaine hydroxylase. J. Med. Chem. 2014, 57, 2213–2236. [Google Scholar] [CrossRef]
- Dambrova, M.; Liepinsh, E.; Kalvinsh, I. Mildronate: Cardioprotective action through carnitine-lowering effect. Trends Cardiovasc. Med. 2002, 12, 275–279. [Google Scholar] [CrossRef]
- Gruber, T. Synthetic Receptors for the Recognition and Discrimination of Post-Translationally Methylated Lysines. ChemBioChem 2018, 19, 2324–2340. [Google Scholar] [CrossRef]
- Hof, F. Host-guest chemistry that directly targets lysine methylation: Synthetic host molecules as alternatives to bio-reagents. Chem. Commun. 2016, 52, 10093–10108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beaver, J.E.; Waters, M.L. Molecular Recognition of Lys and Arg Methylation. ACS Chem. Biol. 2016, 11, 643–653. [Google Scholar] [CrossRef] [PubMed]
- Rasool, M.; Malik, A.; Naseer, M.I.; Manan, A.; Ansari, S.A.; Begum, I.; Qazi, M.H.; Pushparaj, P.N.; Abuzenadah, A.M.; Al-Qahtani, M.H.; et al. The role of epigenetics in personalized medicine: Challenges and opportunities. BMC Genomics 2015, 8, S5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patnaik, S. Anupriya Drugs targeting epigenetic modifications and plausible therapeutic strategies against colorectal cancer. Front. Pharmacol. 2019, 10. [Google Scholar] [CrossRef] [Green Version]
- Cheng, Y.; He, C.; Wang, M.; Ma, X.; Mo, F.; Yang, S.; Han, J.; Wei, X. Targeting epigenetic regulators for cancer therapy: Mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 2019, 4. [Google Scholar] [CrossRef] [Green Version]
















Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maas, M.N.; Hintzen, J.C.J.; Porzberg, M.R.B.; Mecinović, J. Trimethyllysine: From Carnitine Biosynthesis to Epigenetics. Int. J. Mol. Sci. 2020, 21, 9451. https://doi.org/10.3390/ijms21249451
Maas MN, Hintzen JCJ, Porzberg MRB, Mecinović J. Trimethyllysine: From Carnitine Biosynthesis to Epigenetics. International Journal of Molecular Sciences. 2020; 21(24):9451. https://doi.org/10.3390/ijms21249451
Chicago/Turabian StyleMaas, Marijn N., Jordi C. J. Hintzen, Miriam R. B. Porzberg, and Jasmin Mecinović. 2020. "Trimethyllysine: From Carnitine Biosynthesis to Epigenetics" International Journal of Molecular Sciences 21, no. 24: 9451. https://doi.org/10.3390/ijms21249451




