Transcriptome Analysis Reveals Candidate Genes Involved in Light-Induced Primordium Differentiation in Pleurotus eryngii
Abstract
:1. Introduction
2. Results
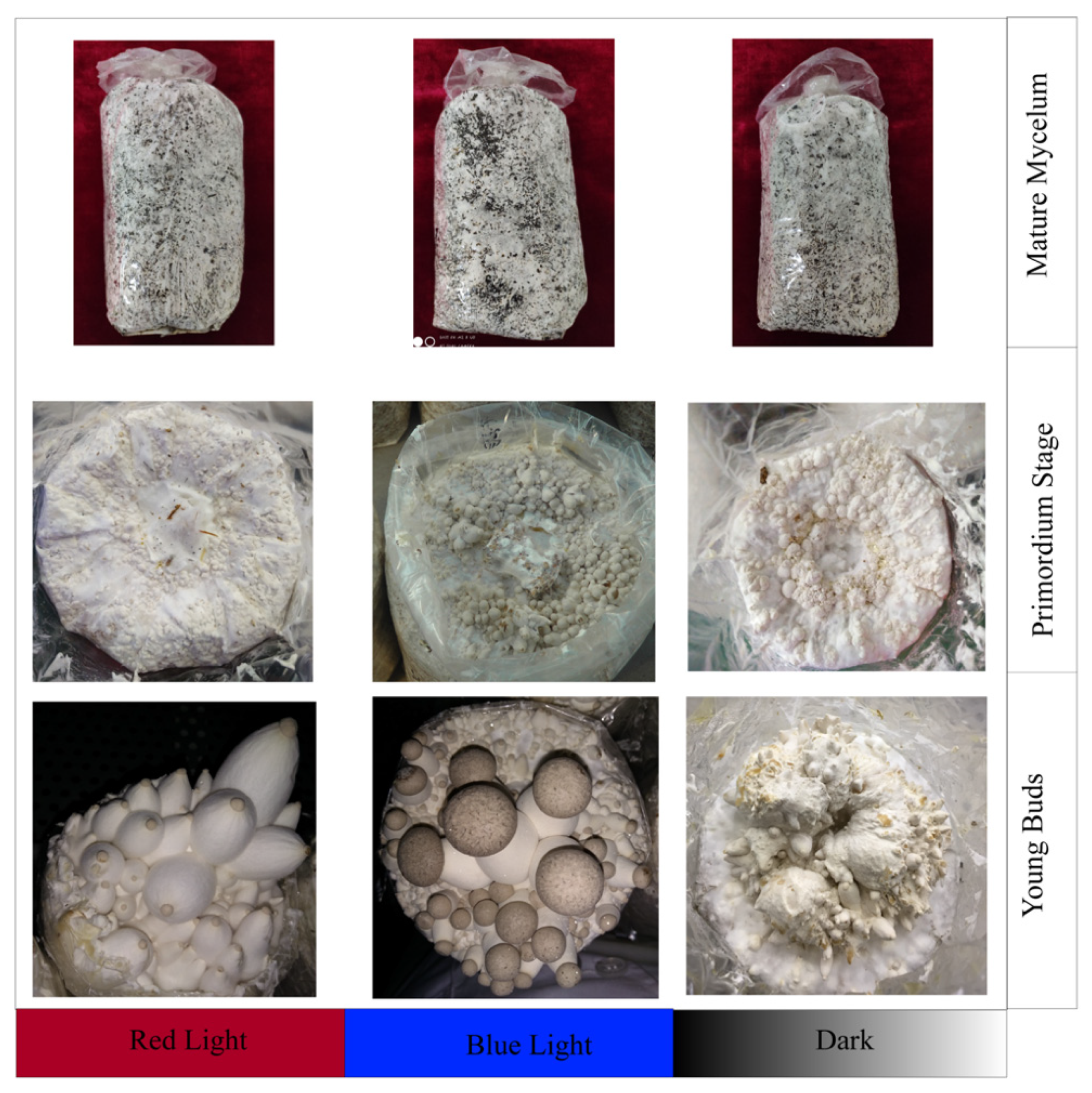
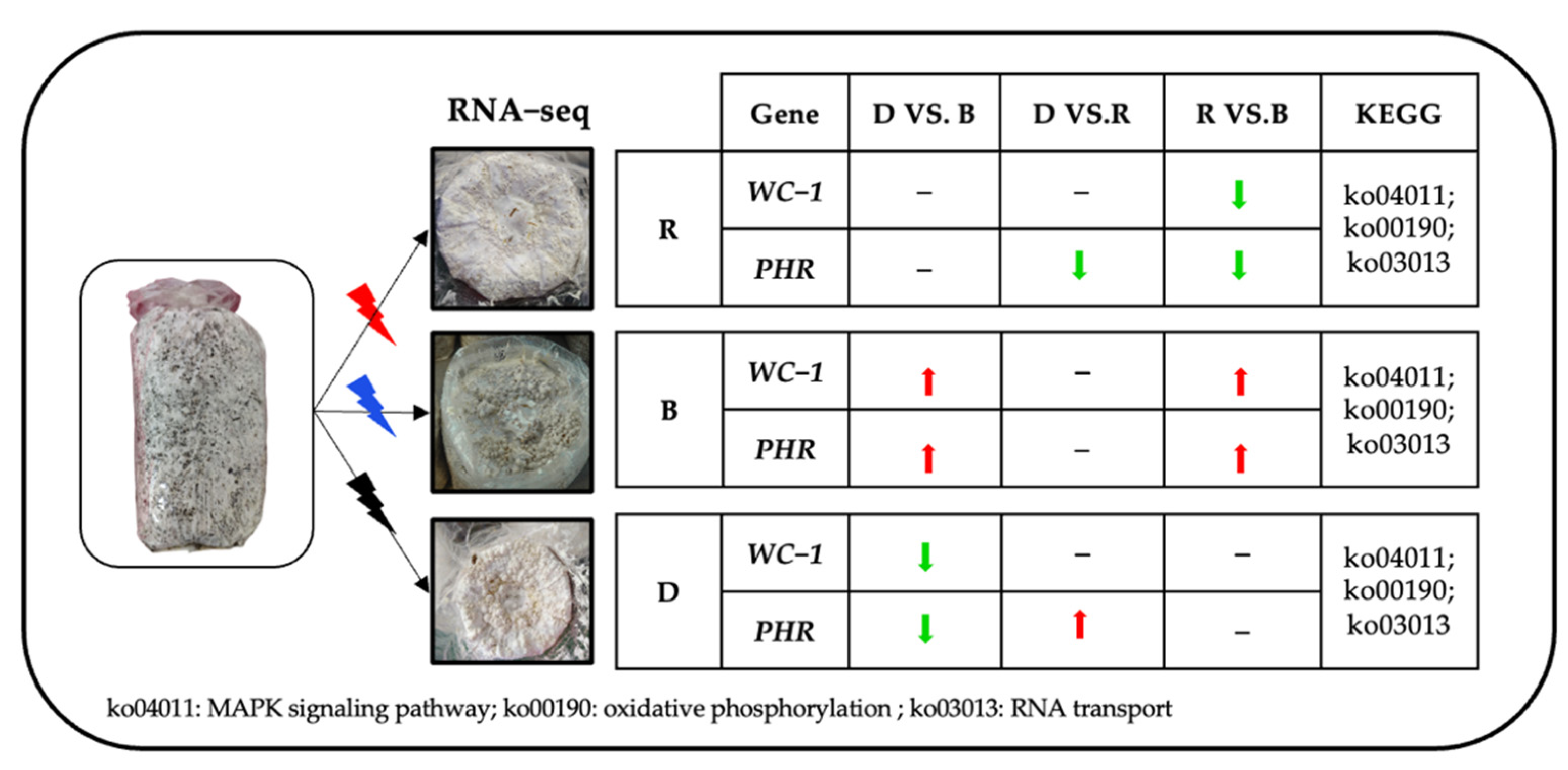
2.1. Morphological Features of Primordia under Different Light Conditions
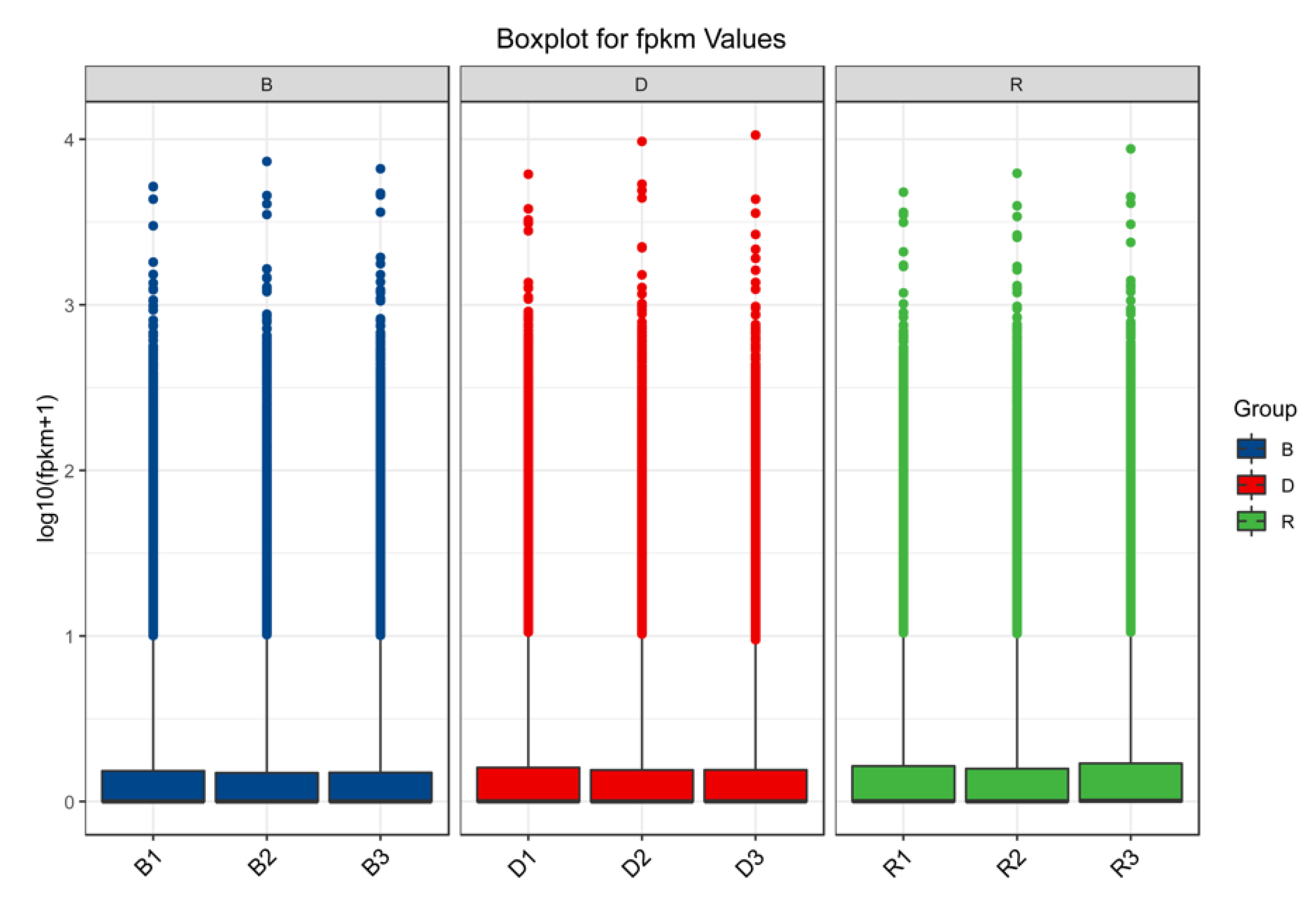
2.2. Sequencing
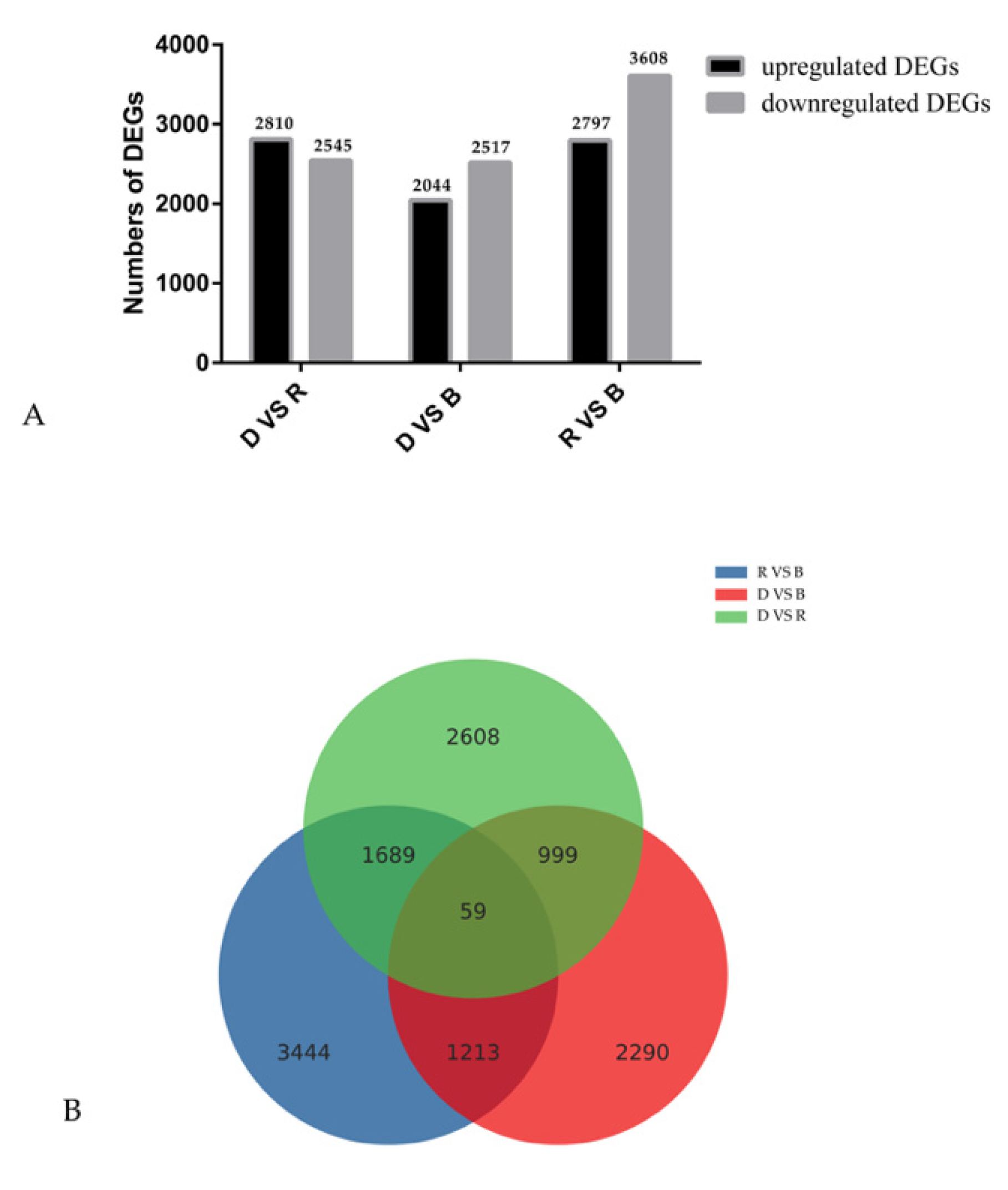
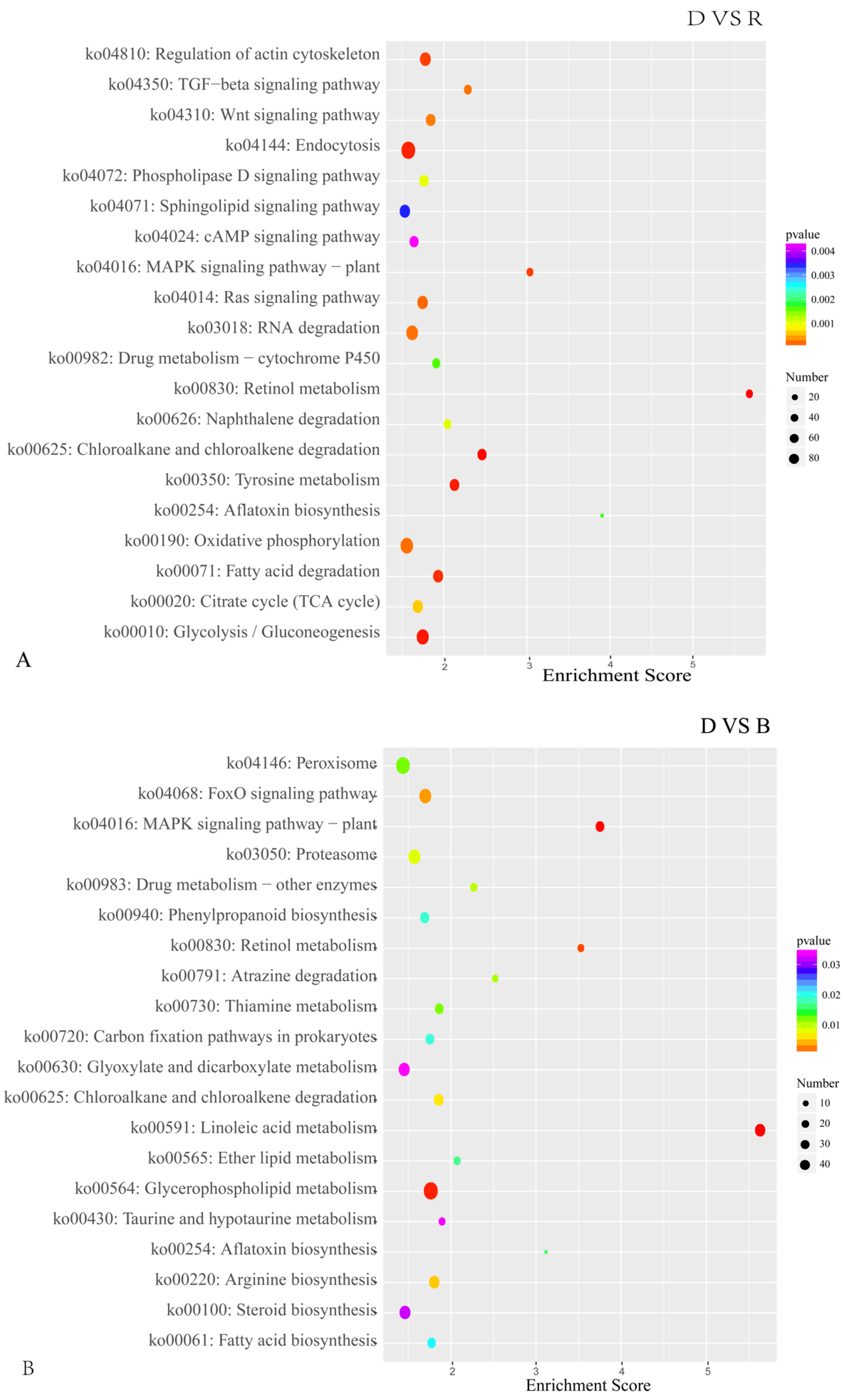
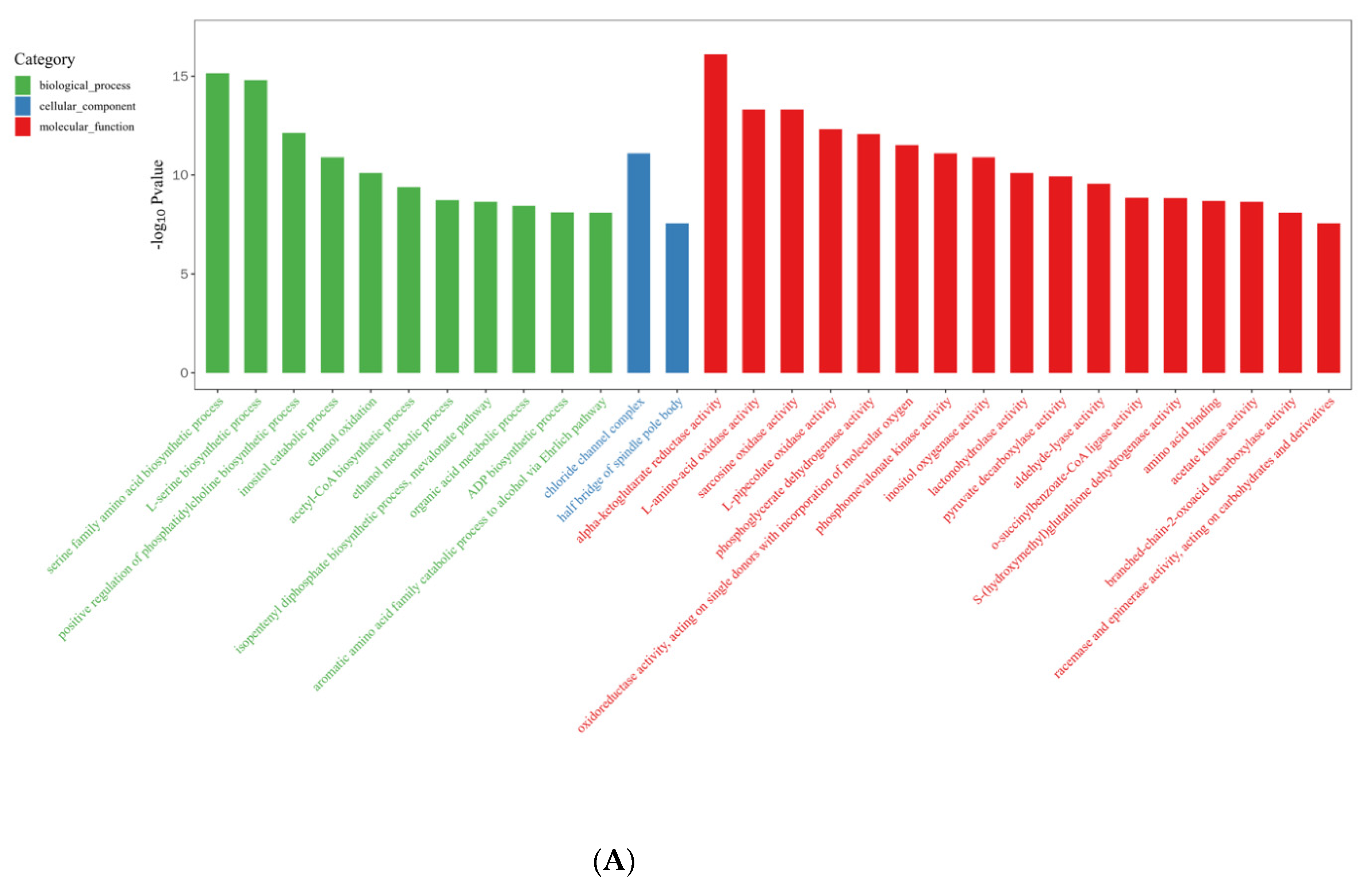
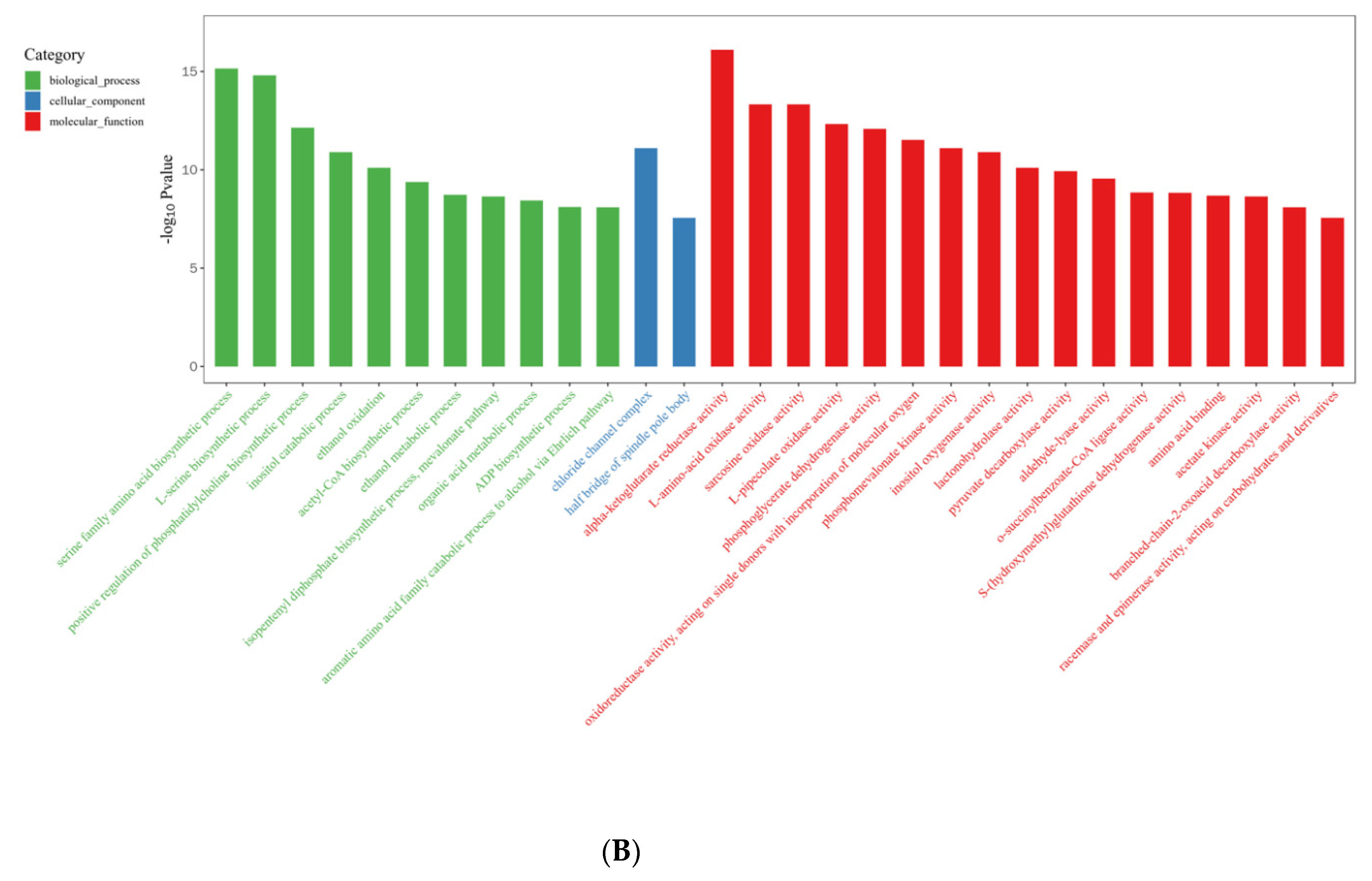
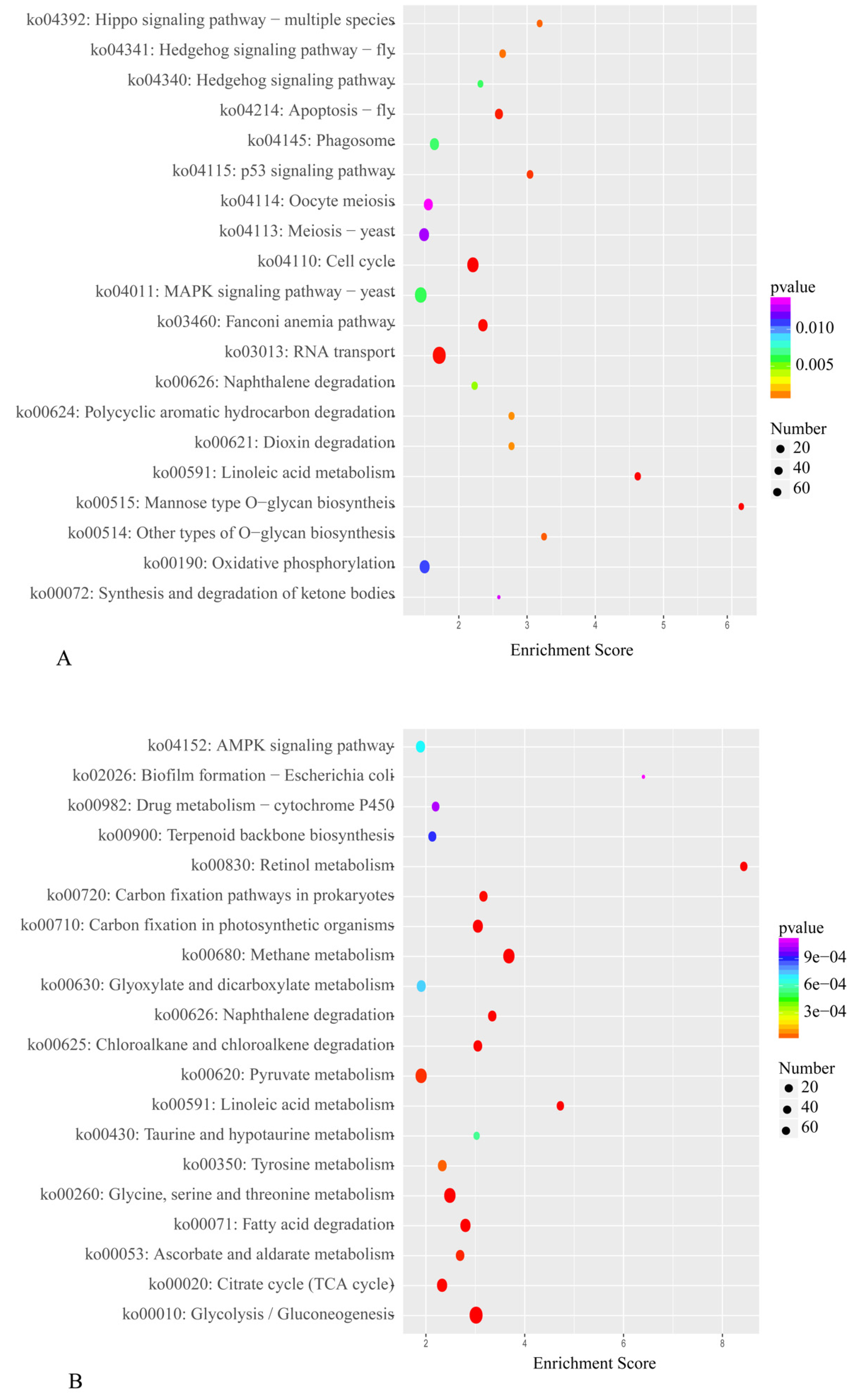
2.3. DEG Identification and Functional Annotation
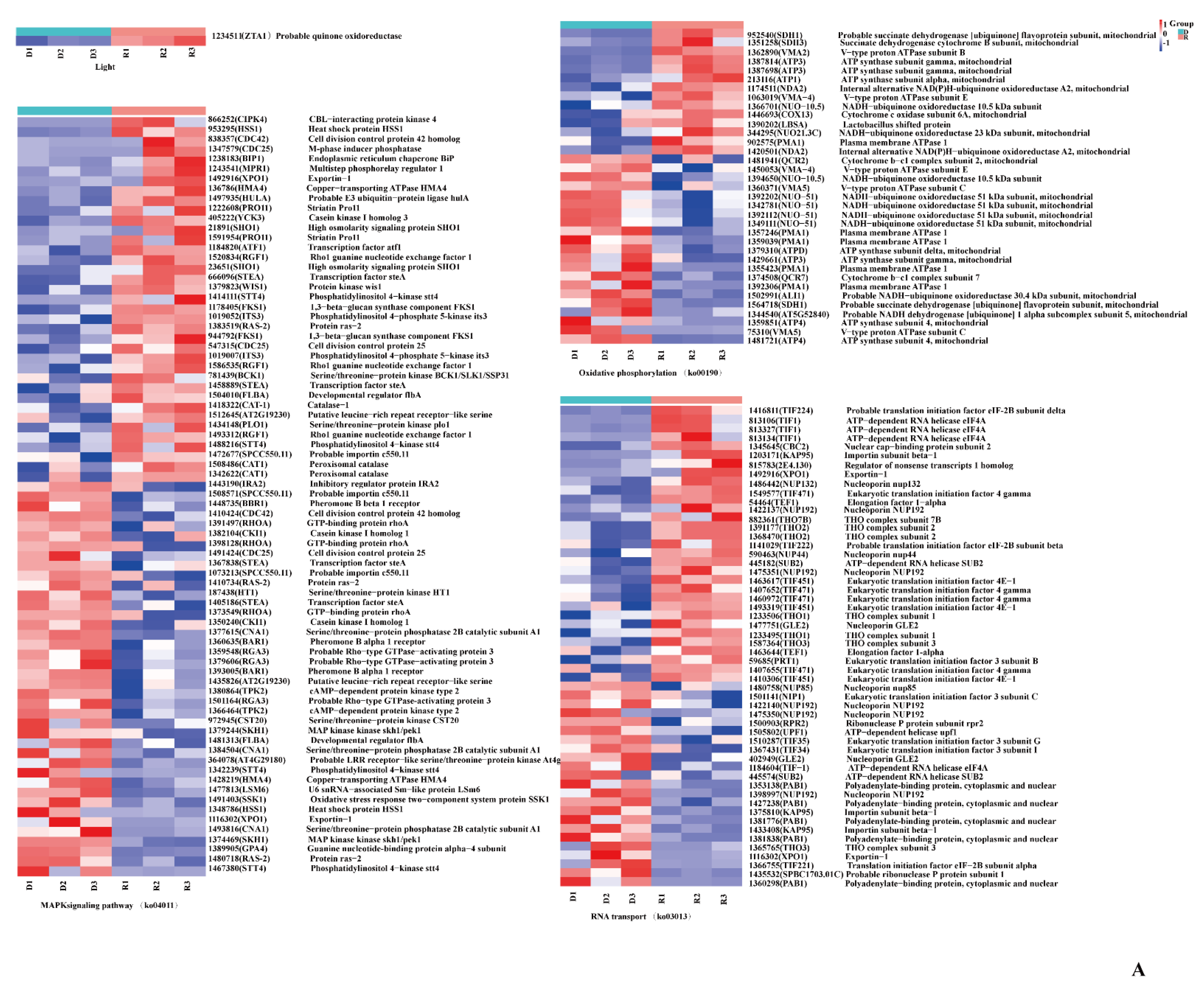
2.4. Analysis of DEGs Involved in Light-Induced Primordium Differentiation
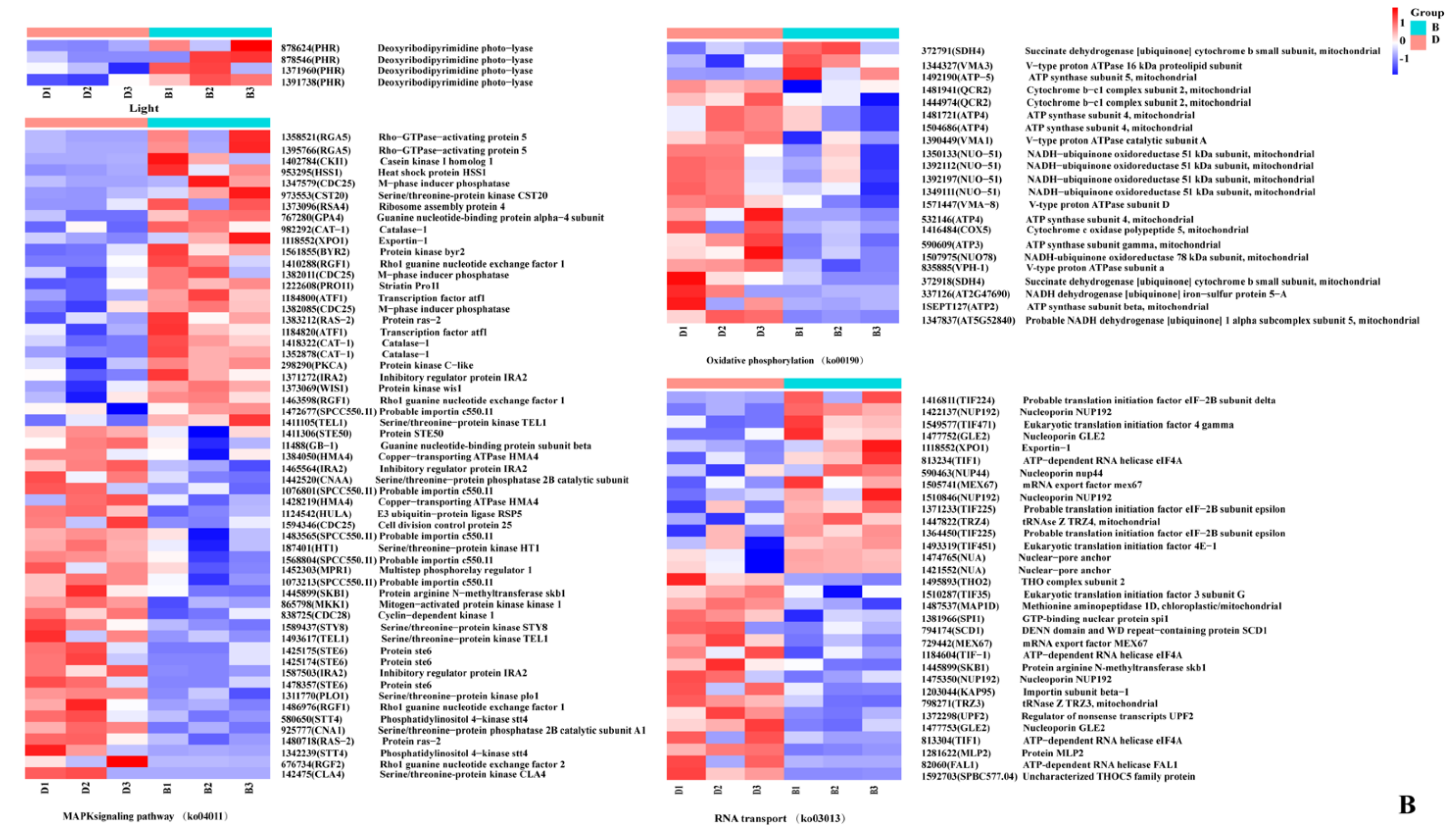
2.5. Analysis of DEGs during Primordium Differentiation Related to Red and Blue Light
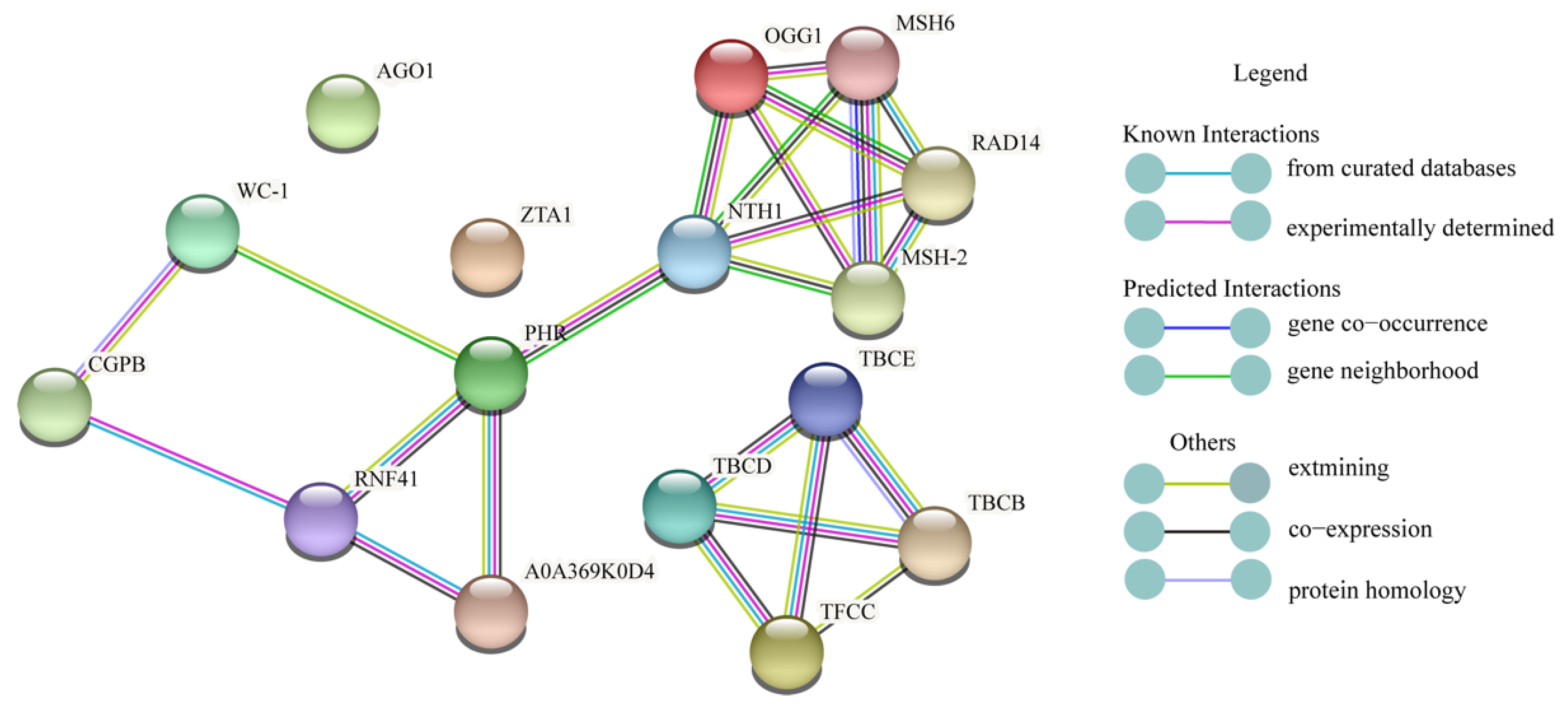
2.6. Protein-Protein Interaction (PPI) Network Prediction
2.7. Real-Time PCR Validation of Transcription Data
3. Discussion and Conclusions
4. Materials and Methods
4.1. Culture Conditions and Primordium Collection
4.2. RNA Extraction, cDNA Library Construction and Illumina Sequencing
4.3. Differentially Expressed Gene (DEG) Analysis and Functional Annotation
4.4. Real-Time Quantitative (RT-qPCR) Analysis
4.5. PPI Network
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nie, Y.; Zhang, P.; Deng, C.; Xu, L.; Yu, M.; Yang, W.; Zhao, R.; Li, B. Effects of Pleurotus eryngii (mushroom) powder and soluble polysaccharide addition on the rheological and microstructural properties of dough. Food Sci. Nutr. 2019, 7, 2113–2122. [Google Scholar] [CrossRef] [Green Version]
- Estrada, A.E.R.; Royse, D.J. Yield, size and bacterial blotch resistance of Pleurotus eryngii grown on cottonseed hulls/oak sawdust supplemented with manganese, copper and whole ground soybean. Bioresour. Technol. 2007, 98, 1898–1906. [Google Scholar] [CrossRef]
- Xie, C.; Gong, W.; Zhu, Z.; Yan, L.; Hu, Z.; Peng, Y. Comparative transcriptomics of Pleurotus eryngii reveals blue-light regulation of carbohydrate-active enzymes (CAZymes) expression at primordium differentiated into fruiting body stage. Genomics 2018, 110, 201–209. [Google Scholar] [CrossRef]
- Jang, M.J.; Lee, Y.H.; Kim, J.H.; Ju, Y.C. Effect of LED light on primordium formation, morphological properties, ergosterol content and antioxidant activity of fruit body in Pleurotus eryngii. Korean J. Mycol. 2011, 39, 175–179. [Google Scholar] [CrossRef]
- Ye, D.; Du, F.; Zou, Y.J.; Zhang, H.J.; Hou, Z.S.; Hu, Q.X. Effects of light conditions on the differentiation and physiological effects of Pleurotus eryngii primordium. China J. Appl. Environ. Biol. 2019, 25, 1107–1112. [Google Scholar]
- Wang, F.; Song, X.; Dong, X.; Zhang, J.; Dong, C. DASH-type cryptochromes regulate fruiting body development and secondary metabolism differently than CmWC-1 in the fungus Cordyceps militaris. Appl. Microbiol. Biotechnol. 2017, 101, 4645–4657. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Tong, X.; Tian, F.; Jia, C.; Li, C.; Li, Y. Transcriptomic profiling sheds light on the blue-light and red-light response of oyster mushroom (Pleurotus ostreatus). AMB Express 2020, 10, 10. [Google Scholar] [CrossRef] [Green Version]
- Song, T.; Shen, Y.; Jin, Q.; Feng, W.; Fan, L.; Cai, W. Comparative phosphoproteome analysis to identify candidate phosphoproteins involved in blue light-induced brown film formation in Lentinula edodes. PeerJ 2020, 8, e9859. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Zhang, R.; Qiu, Y.; Wu, H.; Xiang, Q.; Yu, X.; Zhao, K.; Zhang, X.; Chen, Q.; Penttinen, P.; et al. RNA-seq profiling showed divergent carbohydrate-active enzymes (CAZymes) expression patterns in Lentinula edodes at brown film formation stage under blue light induction. Front. Microbiol. 2020, 11, 1044. [Google Scholar] [CrossRef]
- Kumar, M.; Kherawat, B.S.; Dey, P.; Saha, D.; Singh, A.; Bhatia, S.K.; Ghodake, G.S.; Kadam, A.A.; Kim, H.U.; Chung, S.M.; et al. Genome-wide identification and characterization of PIN-FORMED (PIN) gene family reveals role in developmental and various stress conditions in Triticum aestivum L. Int. J. Mol. Sci. 2021, 22, 7396. [Google Scholar] [CrossRef] [PubMed]
- Kesawat, M.S.; Kherawat, B.S.; Singh, A.; Dey, P.; Kabi, M.; Debnath, D.; Saha, D.; Khandual, A.; Rout, S.; Ali, A.; et al. Genome-Wide Identification and Characterization of the Brassinazole-resistant (BZR) Gene Family and Its Expression in the Various Developmental Stage and Stress Conditions in Wheat (Triticum aestivum L.). Int. J. Mol. Sci. 2021, 22, 8743. [Google Scholar] [CrossRef]
- Wang, M.; Chen, B.; Zhou, W.; Xie, L.; Wang, L.; Zhang, Y.; Zhang, Q. Genome-wide identification and expression analysis of the AT-hook Motif Nuclear Localized gene family in soybean. BMC Genom. 2021, 22, 361. [Google Scholar] [CrossRef]
- Jiang, M.; Chen, H.; Liu, J.; Du, Q.; Lu, S.; Liu, C. Genome-wide identification and functional characterization of natural antisense transcripts in Salvia miltiorrhiza. Sci. Rep. 2021, 11, 4769. [Google Scholar] [CrossRef]
- Shell Cox, J.; Van Dyke, M.W. General and genomic DNA-Binding specificity for the Thermus thermophilus HB8 transcription factor TTHB023. Biomolecules 2020, 10, 94. [Google Scholar]
- Li, Q.; Huang, W.; Xiong, C.; Zhao, J. Transcriptome analysis reveals the role of nitric oxide in Pleurotus eryngii responses to Cd(2+) stress. Chemosphere 2018, 201, 294–302. [Google Scholar] [CrossRef]
- Du, F.; Ti, N.; Hu, Q.; Zou, Y.; Ye, D.; Zhang, A.H. A comparative transcriptome analysis reveals physiological maturation properties of mycelia in Pleurotus tuoliensis. Genes 2019, 10, 703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tan, X.; Sun, J.; Ning, H.; Qin, Z.; Miao, Y.; Sun, T.; Zhang, X. De novo transcriptome sequencing and comprehensive analysis of the heat stress response genes in the basidiomycetes fungus Ganoderma lucidum. Gene 2018, 661, 139–151. [Google Scholar] [CrossRef]
- Song, H.Y.; Kim, D.H.; Kim, J.M. Comparative transcriptome analysis of dikaryotic mycelia and mature fruiting bodies in the edible mushroom Lentinula Edodes. Sci. Rep. 2018, 8, 8983. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Dai, Y.; Yang, C.; Wei, P.; Song, B.; Yang, Y.; Sun, L.; Zhang, Z.W.; Li, Y. Comparative transcriptome analysis identified candidate genes related to Bailinggu mushroom formation and genetic markers for genetic analyses and breeding. Sci. Rep. 2017, 7, 9266. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, N.; Yoshida, Y.; Hasunuma, K. Loss of Catalase-1 (Cat-1) results in decreased conidial viability enhanced by exposure to light in Neurospora crassa. Mol. Genet. Genom. 2007, 277, 13–22. [Google Scholar] [CrossRef]
- Pena, A.; Babiker, R.; Chaduli, D.; Lipzen, A.; Wang, M.; Chovatia, M.; Rencoret, J.; Marques, G.; Sanchez-Ruiz, M.I.; Kijpornyongpan, T.; et al. A multiomic approach to understand how Pleurotus eryngii transforms non-woody lignocellulosic material. J. Fungi 2021, 7, 426. [Google Scholar] [CrossRef]
- Zhang, Q.; Zhao, L.; Li, Y.; Wang, F.; Li, S.; Shi, G.; Ding, Z. Comparative transcriptomics and transcriptional regulation analysis of enhanced laccase production induced by co-culture of Pleurotus eryngii var. ferulae with Rhodotorula mucilaginosa. Appl. Microbiol. Biotechnol. 2020, 104, 241–255. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.A. The Influence of temperature and light on growth and sporulation of Melanotus sp. Trans. Br. Mycol. Soc. 1968, 51, 737–747. [Google Scholar] [CrossRef]
- Liu, K.B.; Wang, F.; Liu, G.J.; Dong, C.H. Effect of environmental conditions on synnema formation and nucleoside production in Cicada Flower, Isaria cicadae (Ascomycetes). Int. J. Med. Mushrooms 2019, 21, 59–69. [Google Scholar] [CrossRef]
- Chao, S.C.; Chang, S.L.; Lo, H.C.; Hsu, W.K.; Lin, Y.T.; Hsu, T.H. Enhanced production of fruiting body and bioactive ingredients of Cordyceps militaris with LED light illumination optimization. J. Agric. Sci. Technol. 2019, 21, 451–462. [Google Scholar]
- Ellis, R.J.; Bragdon, G.A.; Schlosser, B.J. Properties of the blue light requirements for primordia initiation and basidiocarp maturation in Coprinus stercorarius. Mycol. Res. 1999, 103, 779–784. [Google Scholar] [CrossRef]
- Lin, M.; Bachman, K.; Cheng, Z.; Daugherty, S.C.; Nagaraj, S.; Sengamalay, N.; Ott, S.; Godinez, A.; Tallon, L.J.; Sadzewicz, L.; et al. Analysis of complete genome sequence and major surface antigens of Neorickettsia helminthoeca, causative agent of salmon poisoning disease. Microb. Biotechnol. 2017, 10, 933–957. [Google Scholar] [CrossRef] [Green Version]
- Liu, Z.Y.; Yishake, D.; Fang, A.P.; Zhang, D.M.; Liao, G.C.; Tan, X.Y.; Zhang, Y.J.; Zhu, H.L. Serum choline is associated with hepatocellular carcinoma survival: A prospective cohort study. Nutr. Metab. 2020, 17, 25. [Google Scholar] [CrossRef] [PubMed]
- Luo, Z.; Loja, M.; Farwell, D.G.; Luu, Q.C.; Donald, P.J.; Amott, D.; Gandour-Edwards, R.; Nitin, N. High-resolution optical molecular imaging of changes in choline metabolism in oral neoplasia. Transl. Oncol. 2013, 6, 33–41. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.Y.; Park, S.B.; Kim, Y.E.; Yoo, H.M.; Hong, J.; Choi, K.J.; Kim, K.Y.; Kang, D. iTRAQ-Based Quantitative Proteomic Comparison of 2D and 3D Adipocyte Cell Models Co-cultured with Macrophages Using Online 2D-nanoLC-ESI-MS/MS. Sci. Rep. 2019, 9, 16746. [Google Scholar] [CrossRef]
- Zhang, J.; Ren, A.; Chen, H.; Zhao, M.; Shi, L.; Chen, M.; Wang, H.; Feng, Z. Transcriptome analysis and its application in identifying genes associated with fruiting body development in basidiomycete Hypsizygus marmoreus. PLoS ONE 2015, 10, e0123025. [Google Scholar] [CrossRef] [Green Version]
- Xiang, L.; Li, Y.; Zhu, Y.; Luo, H.; Li, C.; Xu, X.; Sun, C.; Song, J.; Shi, L.; He, L.; et al. Transcriptome analysis of the Ophiocordyceps sinensis fruiting body reveals putative genes involved in fruiting body development and cordycepin biosynthesis. Genomics 2014, 103, 154–159. [Google Scholar] [CrossRef]
- Lara-Rojas, F.; Sanchez, O.; Kawasaki, L.; Aguirre, J. Aspergillus nidulans transcription factor AtfA interacts with the MAPK SakA to regulate general stress responses, development and spore functions. Mol. Microbiol. 2011, 80, 436–454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamada, M.; Kurano, M.; Inatomi, S.; Taguchi, G.; Okazaki, M.; Shimosaka, M. Isolation and characterization of a gene coding for chitin deacetylase specifically expressed during fruiting body development in the basidiomycete Flammulina velutipes and its expression in the yeast Pichia pastoris. FEMS Microbiol. Lett. 2008, 289, 130–137. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, Y.P.; Liang, Y.; Dai, Y.T.; Yang, C.T.; Duan, M.Z.; Zhang, Z.; Hu, S.N.; Zhang, Z.W.; Li, Y. De Novo Sequencing and Transcriptome Analysis of Pleurotus eryngii subsp. tuoliensis (Bailinggu) Mycelia in Response to Cold Stimulation. Molecules 2016, 21, 560. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Wang, F.; Yang, Y.; Wang, Y.; Dong, C. CmVVD is involved in fruiting body development and carotenoid production and the transcriptional linkage among three blue-light receptors in edible fungus Cordyceps militaris. Environ. Microbiol. 2020, 22, 466–482. [Google Scholar] [CrossRef]
- Wieser, J.; Yu, J.H.; Adams, T.H. Dominant mutations affecting both sporulation and sterigmatocystin biosynthesis in Aspergillus nidulans. Curr. Gent. 1997, 32, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Krober, A.; Etzrodt, S.; Bach, M.; Monod, M.; Kniemeyer, O.; Staib, P.; Brakhage, A.A. The transcriptional regulators SteA and StuA contribute to keratin degradation and sexual reproduction of the dermatophyte Arthroderma benhamiae. Curr Genet. 2017, 63, 103–116. [Google Scholar] [CrossRef]
- Shantappa, S.; Dhingra, S.; Hernandez-Ortiz, P.; Espeso, E.A.; Calvo, A.M. Role of the zinc finger transcription factor SltA in morphogenesis and sterigmatocystin biosynthesis in the fungus Aspergillus nidulans. PLoS ONE 2013, 8, e68492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, J.; Hulsman, M.; Arentshorst, M.; Breeman, M.; Alazi, E.; Lagendijk, E.L.; Rocha, M.C.; Malavazi, I.; Nitsche, B.M.; van den Hondel, C.A.; et al. Transcriptomic and molecular genetic analysis of the cell wall salvage response of Aspergillus niger to the absence of galactofuranose synthesis. Cell Microbiol. 2016, 18, 1268–1284. [Google Scholar] [CrossRef]
- Bencina, M.; Bagar, T.; Lah, L.; Krasevec, N. A comparative genomic analysis of calcium and proton signaling/homeostasis in Aspergillus species. Fungal Genet. Biol. 2009, 46 (Suppl. S1), S93–S104. [Google Scholar] [CrossRef] [PubMed]
- Porte, S.; Crosas, E.; Yakovtseva, E.; Biosca, J.A.; Farres, J.; Fernandez, M.R.; Pares, X. MDR quinone oxidoreductases: The human and yeast zeta-crystallins. Chem. Biol. Interact. 2009, 178, 288–294. [Google Scholar] [CrossRef]
- Sancar, A. Structure and Function of DNA Photolyase and Cryptochrome Blue-Light Photoreceptors. Chem. Rev. 2003, 103, 2203–2237. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Guo, M.; Yang, H.; Guo, S.; Dong, C. The blue-light receptor CmWC-1 mediates fruit body development and secondary metabolism in Cordyceps militaris. Appl. Microbiol. Biotechnol. 2016, 100, 743–755. [Google Scholar] [CrossRef]
- Xu, X.; Chen, X.; Yu, W.; Liu, Y.; Zhang, W.; Lan, J. Cloning and analysis of the Glwc-1 and Glwc-2 genes encoding putative blue light photoreceptor from Ganoderma lucidum. J. Basic Microbiol. 2017, 57, 705–711. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Sun, X.; Ma, L.; Wen, Q.; Qiu, L.; Shen, J. Identification of two Pleurotus ostreatus blue light receptor genes (PoWC-1 and PoWC-2) and in vivo confirmation of complex PoWC-12 formation through yeast two hybrid system. Fungal Biol. 2020, 124, 8–14. [Google Scholar] [CrossRef]
- Cervantes-Badillo, M.G.; Munoz-Centeno, T.; Uresti-Rivera, E.E.; Arguello-Astorga, G.R.; Casas-Flores, S. The Trichoderma atroviride photolyase-encoding gene is transcriptionally regulated by non-canonical light response elements. FEBS J. 2013, 280, 3697–3708. [Google Scholar] [CrossRef] [PubMed]
- Linden, H.; Ballario, P.; Macino, G. Blue Light Regulation in Neurospora crassa. Fungal Genet. Biol. 1997, 22, 141–150. [Google Scholar] [CrossRef]
- Lauter, F.R.; Yamashiro, C.T.; Yanofsky, C. Charles Yanofsky Light stimulation of conidiation in Neurospora crassa- studies with the wild-type strain and mutants wc-1, wc-2 and acon-2. J. Photochem. Photobiol. B Biol. 1997, 37, 9. [Google Scholar] [CrossRef]
- Olmedo, M.; Ruger-Herreros, C.; Luque, E.M.; Corrochano, L.M. A complex photoreceptor system mediates the regulation by light of the conidiation genes con-10 and con-6 in Neurospora crassa. Fungal Genet. Biol. 2010, 47, 352–363. [Google Scholar] [CrossRef]
- Ambra, R.; Grimaldi, B.; Zamboni, S.; Filetici, P.; Macino, G.; Ballario, P. Photomorphogenesis in the hypogeous fungus Tuber borchii: Isolation and characterization of TbWC-1, the homologue of the blue-light photoreceptor of Neurospora crassa. Fungal Genet. Biol. 2004, 41, 688–697. [Google Scholar] [CrossRef] [PubMed]
- Du, F.; Zou, Y.; Hu, Q.; Zhang, H.; Ye, D. Comparative transcriptomic analysis reveals molecular processes involved in pileus morphogenesis in Pleurotus eryngii under different light conditions. Genomics 2020, 112, 1707–1715. [Google Scholar] [CrossRef]
- Patel, R.K.; Jain, M. NGS QC Toolkit: A toolkit for quality control of next generation sequencing data. PLoS ONE 2012, 7, e30619. [Google Scholar]
- Harris, M.A.; Clark, J.; Ireland, A.; Lomax, J.; Ashburner, M.; Foulger, R.; Eilbeck, K.; Lewis, S.; Marshall, B.; Mungall, C.; et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 2004, 32, D258–D261. [Google Scholar] [PubMed] [Green Version]
- Ogata, H.; Goto, S.; Sato, K.; Fujibuchi, W.; Bono, H.; Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 1998, 27, 29–34. [Google Scholar]




| Sample | Raw Reads | Clean Reads | Valid Bases | Q30 | GC | Total Mapped | Uniquely Mapped |
|---|---|---|---|---|---|---|---|
| B1 | 53,102,878 | 51,075,114 | 92.98% | 93.61% | 53.35% | 46,565,408 (91.17%) | 46,088,648 (90.24%) |
| B2 | 52,458,130 | 50,371,972 | 93.00% | 93.59% | 53.41% | 45,845,104 (91.01%) | 45,378,994 (90.09%) |
| B3 | 52,809,116 | 50,727,940 | 93.13% | 93.54% | 53.16% | 46,107,974 (90.89%) | 45,632,257 (89.95%) |
| R1 | 53,888,394 | 51,658,590 | 90.86% | 91.57% | 53.39% | 47,324,146 (91.61%) | 46,851,732 (90.69%) |
| R2 | 51,919,842 | 51,342,892 | 92.10% | 93.07% | 53.47% | 47,083,554 (91.70%) | 46,617,500 (90.80%) |
| R3 | 52,589,620 | 50,549,472 | 92.29% | 93.47% | 53.44% | 45,918,518 (90.84%) | 45,453,630 (89.92%) |
| D1 | 54,244,320 | 52,057,380 | 92.87% | 93.37% | 53.31% | 47,507,321 (91.26%) | 47,010,123 (90.30%) |
| D2 | 53,015,062 | 51,179,330 | 93.53% | 94.05% | 53.21% | 46,807,561 (91.46%) | 46,289,612 (90.45%) |
| D3 | 53,783,370 | 51,820,220 | 93.42% | 94.38% | 53.39% | 47,417,325 (91.50%) | 46,916,526 (90.54%) |
| Gene ID | Gene Name | Foldchange | Pval | GO ID | GO Term | Description |
|---|---|---|---|---|---|---|
| 1391851 | PHR | 135.255861 | 0.00109307 | GO: 0003904, GO: 0018298 | deoxyribodipyrimidine photolyase activity, protein-chromophore linkage | Deoxyribodipyrimidine photolyase |
| 1391738 | PHR | 15.2406249 | 6.29 × 10−9 | GO: 0003904, GO: 0018298 | deoxyribodipyrimidine photolyase activity, protein-chromophore linkage | Deoxyribodipyrimidine photolyase |
| 1447773 | PHR | 13.401109 | 0.01898976 | GO: 0003904, GO: 0018298 | deoxyribodipyrimidine photolyase activity, protein-chromophore linkage | Deoxyribodipyrimidine photolyase |
| 1350178 | ZTA1 | 12.9653923 | 0.04674992 | GO: 0009644 | response to high light intensity | Probable quinone oxidoreductase |
| 1506953 | PHR | 8.97926851 | 0.03066054 | GO: 0003904, GO: 0018298 | deoxyribodipyrimidine photolyase activity, protein-chromophore linkage | Deoxyribodipyrimidine photolyase |
| 1432018 | TFCC | 5.78562604 | 0.02793471 | GO: 0032391 | photoreceptor connecting cilium | Tubulin-folding cofactor C |
| 1483280 | TFCC | 5.78562604 | 0.02793471 | GO: 0032391 | photoreceptor connecting cilium | Tubulin-folding cofactor C |
| 1507938 | TFCC | 5.78562604 | 0.02793471 | GO: 0032391 | photoreceptor connecting cilium | Tubulin-folding cofactor C |
| 1164413 | WC-1 | 2.40057781 | 0.01738034 | GO: 0009881, GO: 0018298, GO: 0003700 | photoreceptor activity, protein-chromophore linkage, transcription factor activity, sequence-specific DNA binding | White collar 1 protein |
| 1445782 | AGO1 | 0.47726612 | 0.03675021 | GO: 0010218 | response to far red light | Protein argonaute 1 |
| 1486735 | OGG1 | 0.31928656 | 0.04423718 | GO: 0009416 | response to light stimulus | N-glycosylase/DNA lyase |
| 1557381 | SR45A | 0.21401009 | 0.04503058 | GO: 0009644 | response to high light intensity | Serine/arginine-rich splicing factor SR45a |
| 1234511 | ZTA1 | 0.02342917 | 0.00013587 | GO: 0009644 | response to high light intensity | Probable quinone oxidoreductase |
| GENE | Description | Primer (5′-3′) |
|---|---|---|
| GAPDH | Glyceraldehyde-phosphate dehydrogenase | F: ACGATGTCCGACGATGAG |
| R: GACGGCGATGTTGGTGAA | ||
| THO3 | THO complex subunit 3 | F: ATGACGGTTCGAGAGACACC |
| R: TGGCGCAAATATCGATGTAA | ||
| PHR | Deoxyribodipyrimidine photolyase | F: GATCTCAGGGTTGCGGATAA |
| R: GGATGTGGAGTTCGGCTAAA | ||
| KAP95 | Importin subunit beta-1 | F: GTGTGAGGCAACCCAAAACT |
| R: CCGTAGTCCTGAGCCTCTTG | ||
| UPF1 | ATP-dependent helicase upf1 | F: CCCCATCCAATTGTAACCTG |
| R: AACCAGAATGGATGGCAGTC | ||
| MUP1 | High-affinity methionine permease | F: CTGGTGATAGCACCCTTCTTAC |
| R: CCCAGCACGATGATACCAATAC | ||
| ATPD | ATP synthase subunit delta | F: GTTAACATCTCGGCCGCTAC |
| R: AGGGCCTCTTGAAGATTGGT | ||
| RFX1 | Transcriptional regulator RFX1 | F: CATGCGAGACTTGACCATTAGA |
| R: TACTACACCGGATTGAGCTTTG | ||
| RGF1 | Rho1 guanine nucleotide exchange factor 1 | F: GCGTTTCCTGAACCACAAAT |
| R: CTTCGTCGTGATCTGCGTAA | ||
| RAD17 | Checkpoint protein rad17 | F: CTGTTCACCCAAAGAAGGTAGA |
| R: CGCCAGGATCTTTCGGTATT | ||
| CDC25 | M-phase inducer phosphatase | F: CCGAAGATCCACTACCCTGA |
| R: GATGCCATCGCCGTAAGTAT | ||
| STEA | Transcription factor steA | F: CGGCCAGTACGTAACATGAA |
| R: ATCGTGAGGCACTGAGAACC | ||
| WIS1 | Protein kinase wis1 | F: TCCATCGAGATGTGAAACCA |
| R: TATTTTGGGATTCGCCTTTG | ||
| BOA4 | Cytochrome P450 monooxygenase BOA4 | F:CTGGATAGGATGGACGAAGATTAG |
| R: GACGAAGAGACGAGTTGAAGAG | ||
| CLM2 | Cytochrome P450 monooxygenase CLM2 | F: GAAGGAGGTGTTGAGATGGAAT |
| R: CGATCCTTTCGGGATGAAGTAA | ||
| ZTA1 | quinone oxidoreductase | F: CCAGCAGTCTTGGGTAAAGAA |
| R: A GGTGTACTTGCGAGCTTGAT |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ye, D.; Du, F.; Hu, Q.; Zou, Y.; Bai, X. Transcriptome Analysis Reveals Candidate Genes Involved in Light-Induced Primordium Differentiation in Pleurotus eryngii. Int. J. Mol. Sci. 2022, 23, 435. https://doi.org/10.3390/ijms23010435
Ye D, Du F, Hu Q, Zou Y, Bai X. Transcriptome Analysis Reveals Candidate Genes Involved in Light-Induced Primordium Differentiation in Pleurotus eryngii. International Journal of Molecular Sciences. 2022; 23(1):435. https://doi.org/10.3390/ijms23010435
Chicago/Turabian StyleYe, Dou, Fang Du, Qingxiu Hu, Yajie Zou, and Xue Bai. 2022. "Transcriptome Analysis Reveals Candidate Genes Involved in Light-Induced Primordium Differentiation in Pleurotus eryngii" International Journal of Molecular Sciences 23, no. 1: 435. https://doi.org/10.3390/ijms23010435






