Abstract
Proton radiotherapy (PRT) has the potential to reduce the normal tissue toxicity associated with conventional photon-based radiotherapy (X-ray therapy, XRT) because the active dose can be more directly targeted to a tumor. Although this dosimetric advantage of PRT is well known, the molecular mechanisms affected by PRT remain largely elusive. Here, we combined the molecular toolbox of the eukaryotic model Saccharomyces cerevisiae with a systems biology approach to investigate the physiological effects of PRT compared to XRT. Our data show that the DNA damage response and protein stress response are the major molecular mechanisms activated after both PRT and XRT. However, RNA-Seq revealed that PRT treatment evoked a stronger activation of genes involved in the response to proteotoxic stress, highlighting the molecular differences between PRT and XRT. Moreover, inhibition of the proteasome resulted in decreased survival in combination with PRT compared to XRT, not only further confirming that protons induced a stronger proteotoxic stress response, but also hinting at the potential of using proteasome inhibitors in combination with proton radiotherapy in clinical settings.
1. Introduction
Today, up to 50% of cancer patients are treated with radiotherapy during their course of illness [1,2]. Radiotherapy strives for the optimal ratio between the dose delivered to the tumor and the lowest possible dose delivered to the surrounding healthy tissue. Current clinical practices mainly rely on classical X-rays or photon radiotherapy (XRT). However, because of the high entrance and exit dose of photons, damage to the healthy tissue surrounding the tumor remains a major problem associated with XRT [3].
Proton beam radiation (PRT) is an alternative form of radiation therapy that can be used to reduce the problem of the off-target toxicity associated with conventional XRT [4]. Due to their positive electrical charge and mass, protons are slowed down while penetrating tissues, and deposit most of their energy at the end of their path. These physical characteristics result in a unique, focused depth–dose distribution, the Bragg peak, allowing to spare the healthy tissues distal to the tumor from the toxic effects of the irradiation. The importance of PRT is well-recognized for the treatment of tumors close to critical structures; such as head-and-neck cancers; and is a treatment option for several cancers [5]. However, not all cancer patients benefit from the above described dosimetric advantage of protons, highlighting the need for understanding the underlying molecular mechanisms affected by PRT [6,7].
Studies comparing the cellular response to PRT and XRT on the level of survival, DNA damage induction or repair mechanisms affected, do not paint a clear picture [8,9,10,11,12,13,14,15,16,17]. One of the main reasons is the large variation in tumor biology which increases the complexity of interpreting these results. This hiatus in our understanding has limited the ability to achieve the true potential of PRT but also impeded the development of proton specific therapeutic and combinatorial strategies. Increased knowledge of the molecular pathways affected by PRT can lead to selective targeted therapy combinations. This is relevant in the field of personalized medicine because understanding the radiobiology of protons and its interaction with the complex biology of the tumor will allow for the integration of clinical, physical and biological parameters to adjust treatment to the specific needs of an individual patient. A system-wide approach, which is often missing, could help bring more insights into the molecular mechanisms affected by treatment with PRT.
Therefore, in this study, for the first time, we apply an integrated approach to identify the exact mechanisms and cellular pathways affected by both PRT and XRT using the unicellular eukaryote Saccharomyces cerevisiae as a model. The short life cycle, simple culture conditions and ease with which the genome can be manipulated make S. cerevisiae an extensively used model organism, not only in radiobiology but in medicine in general [18,19,20,21,22,23]. Moreover, the techniques and types of experiments available in this yeast make this model organism ideal for a systems biology approach, which is difficult to achieve in mammalian tumor cells because of the large heterogeneity between cell lines. Using Bar-Seq in combination with more detailed molecular assays, we found that genes involved in DNA repair determined the survival of cells exposed to XRT and PRT. Moreover, the DNA damage response was equally important for both irradiation types. Remarkably, in contrast to XRT, transcriptomic analysis after PRT showed a much stronger activation of genes involved in the response to proteotoxic stress. Additionally, inhibition of the proteasome resulted in decreased survival after PRT, but not after XRT. Altogether, our results offer a genome-wide view on the physiological effect of PRT and XRT and bridge the gap between current biological uncertainties and the translation of PRT to the clinic.
2. Results
2.1. Survival of S. cerevisiae Is Independent of the Radiation Type
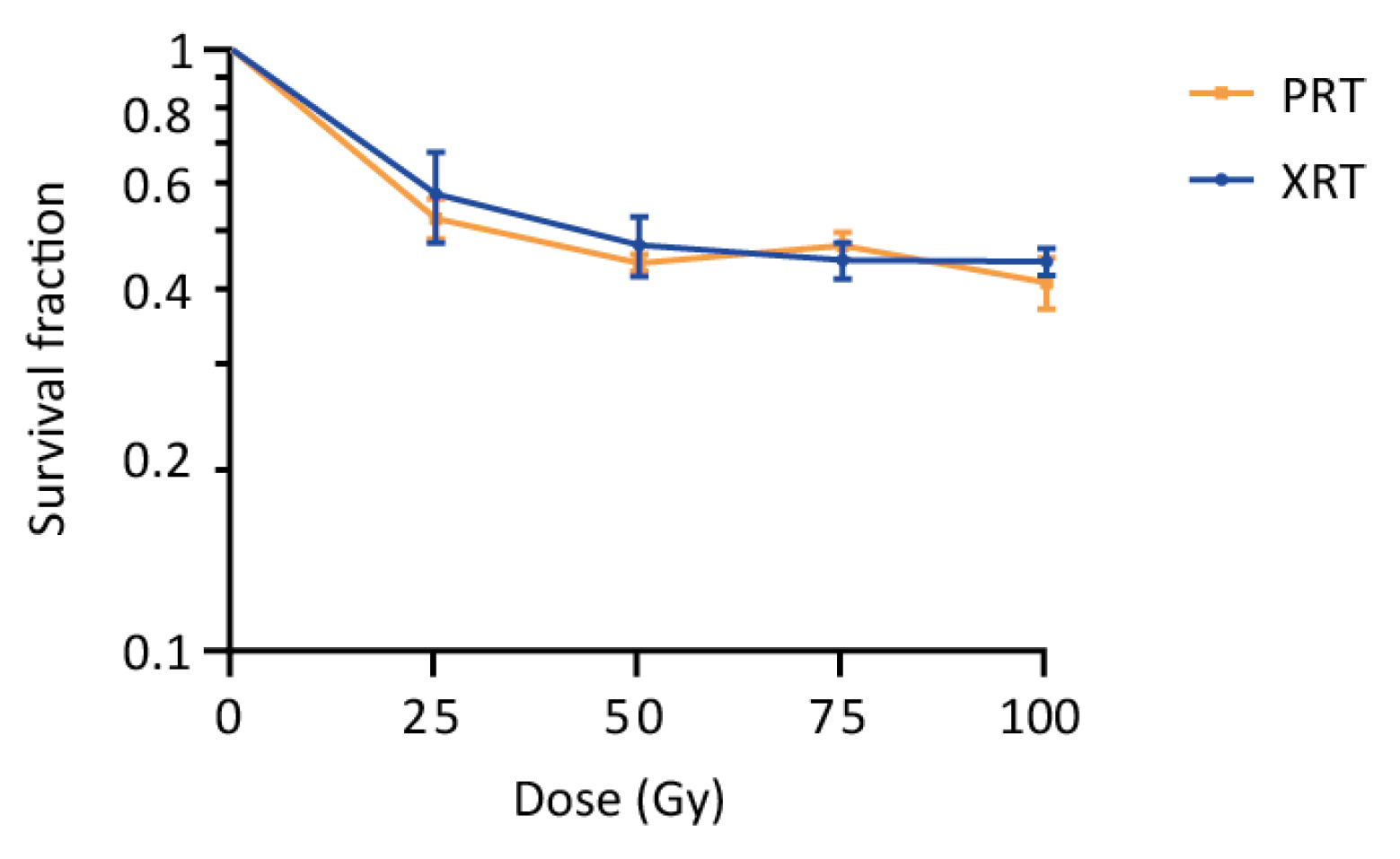
S. cerevisiae has previously been used as a model to study the effects of radiation on cells, and is known to tolerate high doses of radiation [24,25,26,27,28,29,30,31,32]. Our experiments confirmed this tolerance to both PRT and XRT, with cells showing survival fractions of 0.44 ± 0.014 and 0.47 ± 0.054 after receiving a dose of 50 Gy PRT or XRT, respectively (Figure 1). Increasing the dose to 100 Gy resulted in only a minor decrease in the survival (0.41 ± 0.04 for PRT and 0.45 ± 0.023 for XRT), further suggesting a largely similar survival in response to PRT and XRT. Strikingly, although non-significant, the survival fractions obtained for PRT are systematically lower compared to XRT (except at a dose of 75 Gy) with PRT resulting in, on average, 1.05 ± 0.07 more cell kill.
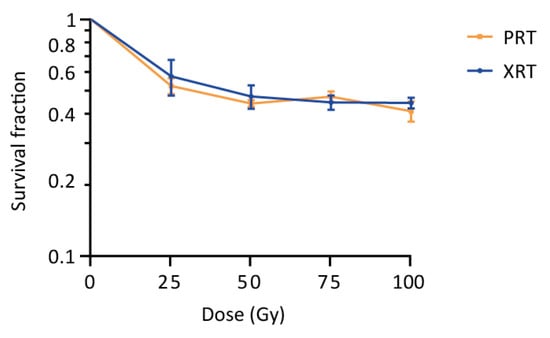
Figure 1.
Survival of wild type yeast after proton (PRT) and photon radiotherapy (XRT). Survival of a wild type yeast strain (KV447) after PRT and XRT was determined using plating assays. Survival fractions were calculated and a survival curve was plot on a logarithmic scale. Data are represented as the mean ± SEM for n = 3. p-values were calculated by multiple t-testing per dose, but no significant differences were detected (p-values of 0.66, 0.60, 0.56 and 0.50 were calculated at 25 Gy, 50 Gy, 75 Gy and 100 Gy, respectively).
2.2. Transcriptomic Response in S. cerevisiae Is Dependent on Radiation Type
The largely similar survival rates of S. cerevisiae in response to PRT or XRT do not exclude possible differences in the underlying physiological response to both types of radiation [33,34]. Therefore, we used RNA sequencing (RNA-Seq) to assess and compare the transcriptional response to a 50 Gy PRT and XRT treatment. As previously determined, this dose resulted in an average survival fraction of approximately 45%. The transcriptome (~6600 genes) was assessed at an early (30 min after radiation) and later (90 min after radiation) timepoint.
Principal component analysis (PCA) of the transcriptome data for the different treatments showed distinct clustering of the proton and photon samples and the respective non-irradiated controls (Figure S1A). PRT and XRT experiments were analyzed separately by calculating the Log2 fold changes (log2FC) of the irradiated condition compared to the corresponding control. To select significantly up- or downregulated genes, the false discovery rate (FDR) was set at 0.01 and a cut-off log2FC of 1.5 for upregulated and −1.5 for downregulated genes was used. Using these cut-offs, volcano plots showed that 68 and 104 genes were differentially expressed compared to cells that did not receive a treatment at 30 and 90 min after PRT, respectively. For XRT, only 37 genes showed differential expression patterns at 30 min, and this further decreased to only 24 genes at 90 min (Figure S1B), suggesting that the transcriptional response to PRT is different and stronger compared to the response to XRT. In line with these data, hierarchical clustering of the sets of differentially regulated genes showed that the samples were clustered per irradiation type (Figure 2A).
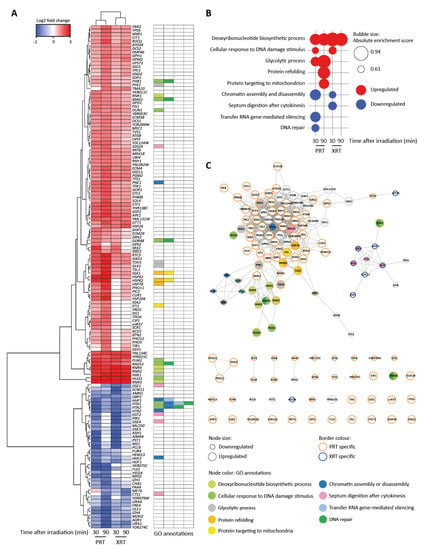
Figure 2.
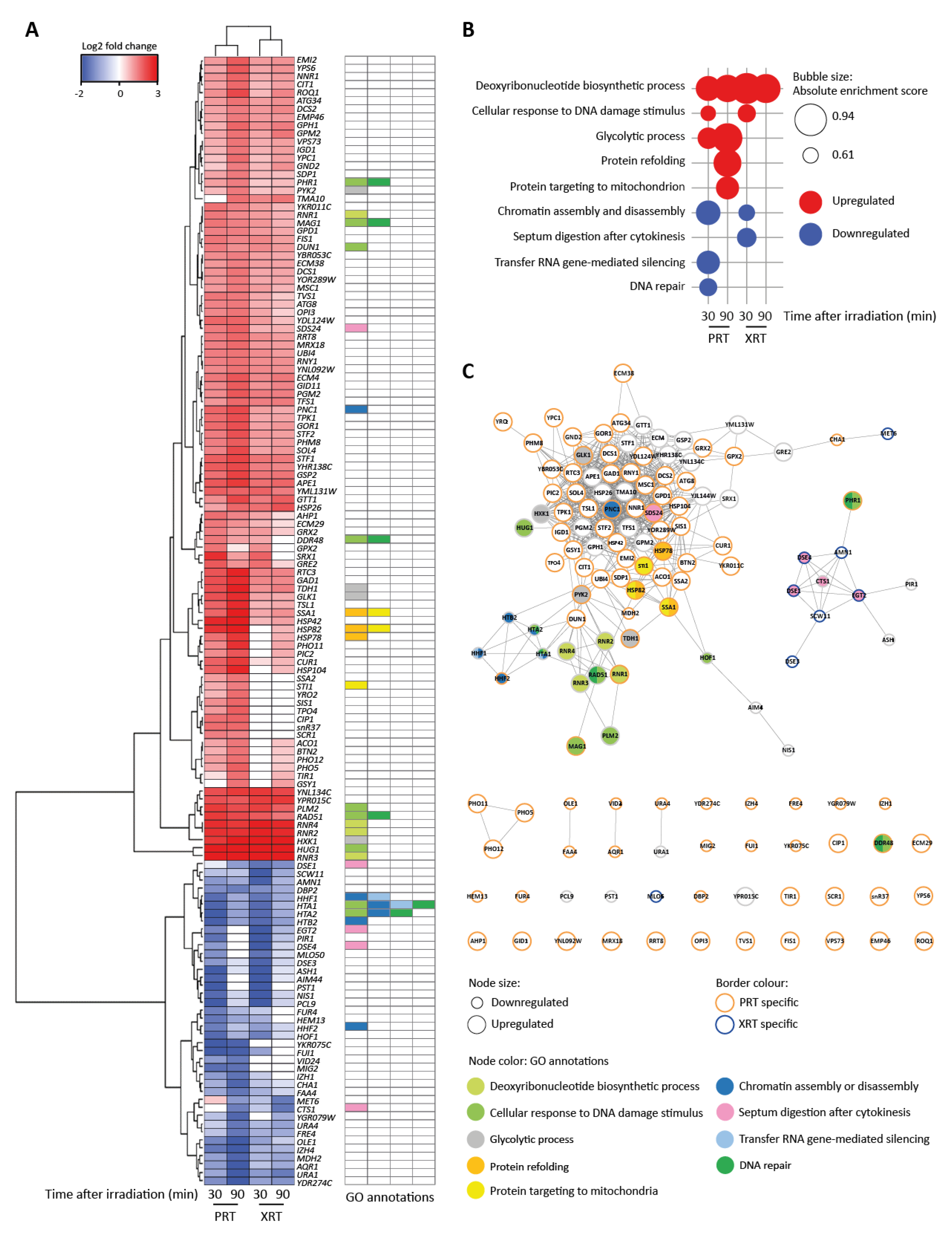
RNA-Seq reveals distinct transcriptional profiles after proton (PRT) and photon radiotherapy (XRT). (A) Heatmap for differentially regulated genes. Genes were selected based on two cut-offs: false discovery rate (FDR) < 0.01 and |log2FC| > 1.5 (log2 fold change) in at least one condition. Hierarchical clustering was performed by the Ward.D2 method based on the Euclidian distance. Gene ontology (GO) annotations for enriched GO terms found by gene set enrichment analysis (GSEA) in (B) are visualized. (B) Enriched GO terms identified using GSEA. GSEA was applied on a ranked gene list based on the log2FC calculated in edgeR. Genes were chosen per condition based on two cut-offs: FDR < 0.01 and log2FC < −1.5 for downregulated genes or log2FC > 1.5 for upregulated genes. For GSEA, the FDR was set at 0.25. (C) Interaction network of genes in (A). The network was built in String [43] and visualized in Cytoscape [44]. The node size corresponds to either up- or downregulated genes. Nodes with orange borders are genes only found to be deregulated after PRT. Nodes with blue borders are genes only found to be deregulated after XRT. Nodes with grey borders are genes deregulated after both PRT and XRT. Node colors represent the GO categories found in (B).
Gene Set Enrichment Analysis (GSEA) showed that deoxyribonucleotide biosynthetic process is highly enriched among the upregulated genes at both timepoints after irradiation and independent of the radiation type (Figure 2A,B). The upregulated genes in this gene ontology (GO) term are primarily involved in DNA metabolism. Among these genes are four genes that encode the subunits of the ribonucleotide reductase (RNR) complex which catalyzes the rate-limiting step in deoxyribonucleoside triphosphates (dNTP) synthesis: RNR1, RNR2, RNR3 and RNR4. dNTP levels are known to increase after DNA damage and are reported to indicate active DNA repair [35].
Although less pronounced, a second group of genes that were significantly upregulated, independent of radiation type, are involved in the cellular response to DNA damage stimulus (Figure 2A,B). This includes the genes encoding the cell cycle checkpoint kinase Dun1, the DNA glycosylase Mag1 that initiates the base excision repair pathway, and Rad51, which functions in strand exchange during homologous recombination. Ninety minutes after radiotherapy, this upregulation was less pronounced, which corresponds to the GSEA analysis (Figure 2B). PRT, but not XRT, also caused a slight induction of genes involved in glycolysis. This includes the genes encoding glucokinase GLX1, pyruvate kinase PYK2, hexokinase HXK1 and glyceraldehyde-3-phosphate dehydrogenase TDH1. It has previously been reported that yeast metabolism can be affected by various types of radiation treatment, but can also reflect the environmental stress response of yeast activated after various kinds of stresses [12,36,37,38].
Interestingly, a large set of genes was only or much stronger expressed after PRT, among which the genes SIS1, STI1, CUR1, SSA1, SSA2, BTN2, HSP42, HSP78, HSP82 and HSP104 which are involved in protein folding (Figure 2A). The upregulation of these genes was even stronger 90 min after irradiation. Indeed, GSEA revealed that genes that fall under the GO classes of “protein refolding” and “protein targeting to the mitochondria” were significantly enriched 90 min after PRT (Figure 2B). These GO classes encompass the chaperones HSP78, HSP82, HSP42, SSA1 and SSA2 and the disaggregase HSP104. These genes were upregulated with induction levels ranging from of 1.6 to 3.5 log2 fold changes, suggesting that some form of proteotoxic stress occurs after PRT. Additionally, XRT induced some genes involved in protein folding namely CUR1, SSA1 and HSP42 30 min after radiation. Additionally, 90 min after irradiation, BTN2, HSP42, HSP78, HSP82 and HSP104 were induced. However, the changes in expression observed after XRT were less strong compared to those elicited by PRT, with log2 fold changes after XRT ranging from 0.6 to 1.3.
A differential response was observed for the downregulated genes comparing the response 30 and 90 min after PRT and XRT. Hierarchical clustering for the downregulated genes showed four main groups (Figure 2A). The first group was enriched in genes categorized under the GO class “chromatin assembly or disassembly” (Figure 2B). This group is mainly composed of the histone-encoding genes HTA1, HTA2, HTB2, HHF1 and HHF2 that were highly downregulated at 30 min after PRT and XRT, and less strongly at 90 min after PRT and XRT. A second group of downregulated genes was involved in cell separation after mitosis and corresponds to the GO term “septum digestion after cytokinesis”, which was significantly enriched at 30 min after XRT. This group is composed of the genes CTS1, DSE1, DSE4, EGT2 and SDS24. For samples treated with PRT, two more significant GSEA enriched sets were identified, namely “transfer RNA gene-mediated silencing” and “DNA repair”. Both GO classes encompass the histone encoding genes HTA1, HTA2 and HHF1. Overall, these patterns of gene downregulation are in-line with reduced cell division. Histones are known to be coupled with DNA synthesis, and downregulation of histones is an indication of cell cycle checkpoint activation [39,40]. This is commonly seen after radiotherapy: when cells are coping with DNA damage, DNA damage checkpoints are activated and the progression through the cell cycle is slowed down to allow sufficient time for DNA repair.
Network analysis of the differentially expressed genes showed three distinct clusters (Figure 2C). Genes facilitating DNA repair, namely the genes involved in deoxyribonucleotide biosynthetic process, cellular response to DNA damage and chromatin assembly or disassembly cluster together. This part of the network consists of genes deregulated after both PRT and XRT. Genes involved in yeast metabolism, and the general stress response forms one big cluster together with the heat shock proteins. A separate cluster is formed by the genes categorized under the GO term “septum digestion after cytokinesis” that were deregulated after XRT. Network analysis also highlighted that although a large set of genes displayed changed expression levels after both PRT and XRT treatment (37 out of 139 genes), PRT caused deregulation of many more genes. An additional set of 94 genes were specifically deregulated after PRT compared to only 8 after XRT.
2.3. DNA Repair Determines the Survival of S. cerevisiae after PRT and XRT
RNA-Seq analysis strongly suggests that both PRT and XRT evoke a similar DNA damage response. However, the response after PRT seems slightly more complex, with the stronger induction of genes involved in proteotoxic stress. To further investigate these potential differences in molecular mechanisms affected after PRT and XRT, we performed a barcode sequencing (Bar-Seq) experiment to assess which genes are important to ensure survival after PRT or XRT. Using the haploid yeast deletion collection, we tested the complete collection of approximately 4800 mutants, each carrying a deletion for one of the 4800 non-essential S. cerevisiae genes. Importantly, each mutant also carries unique DNA barcodes, allowing to assess their sensitivity to PRT and XRT in a high-throughput assay termed “Bar-Seq” (for more details, see materials and methods and [41,42]). The deletion mutant pools were irradiated with a dose of 50 Gy PRT or XRT, while a control population received a mock treatment (i.e., undergoing a similar experimental procedure, but without actual radiation).
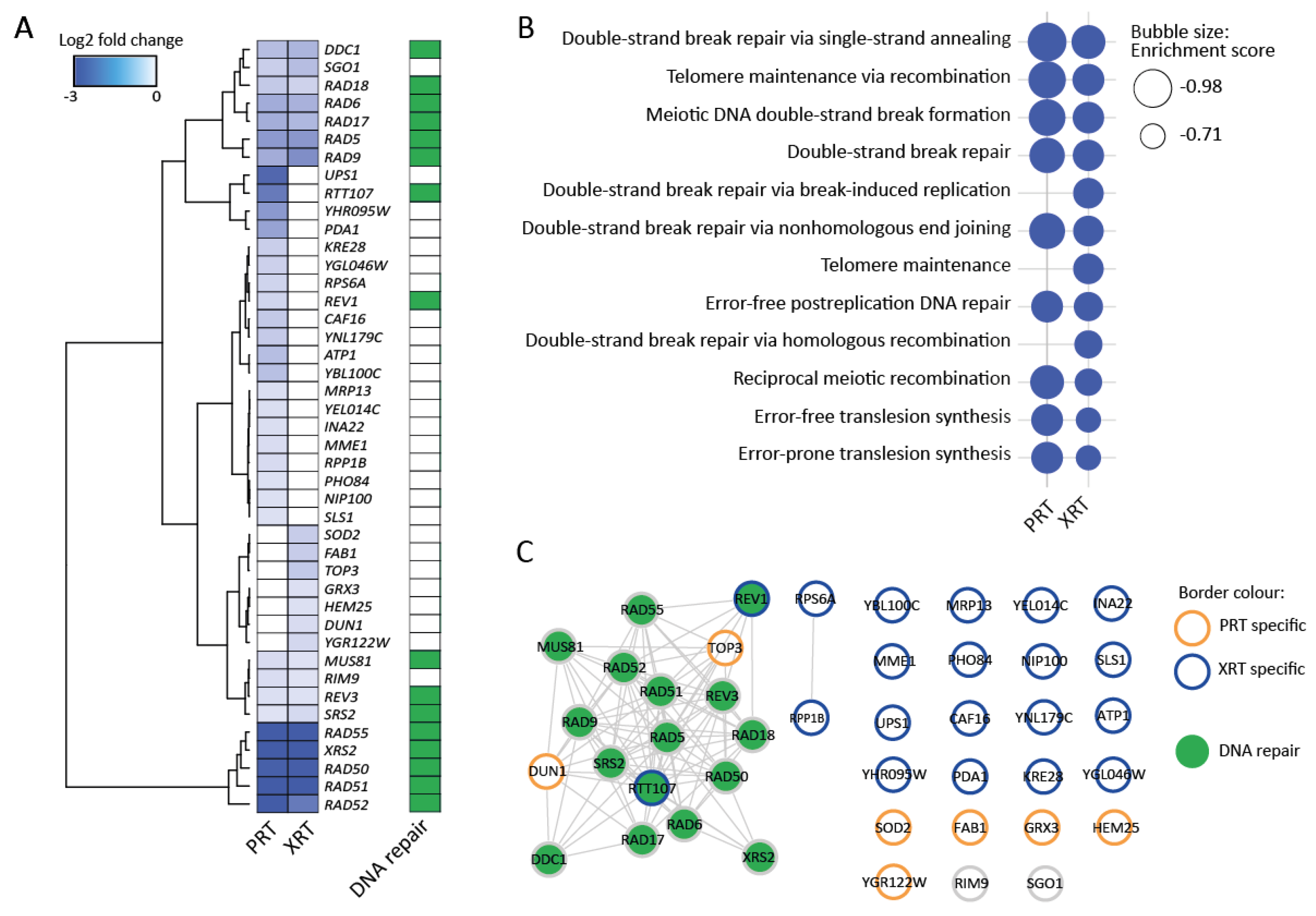
The results of the Bar-Seq experiment revealed 20 and 35 genes that, when deleted, caused sensitivity to PRT and XRT, respectively (FDR 0.05 and log2FC < −0.5). The most sensitive deletion strains were depleted with a log2FC of −3.42 and −3.51 relative to the control for the PRT and XRT, respectively. Similar as for the RNA-Seq experiment, multidimensional scaling (MDS) plots showed clustering according to the radiation type and within their respective controls (Figure S2). Figure 3A shows all the genes for which the cut-offs are met in either or both the PRT and XRT experiment. GSEA revealed an enrichment of DNA repair pathways for both PRT and XRT (Figure 3B). Moreover, the top five most sensitive deletion mutants were shared between PRT and XRT, namely deletions for the genes RAD50, RAD51, RAD52, RAD55 and XRS2. For the genes that are important for either PRT or XRT, no GO enrichment could be found. Network analysis showed a large cluster of genes involved in DNA repair shared between the PRT and XRT experiment (Figure 3C).
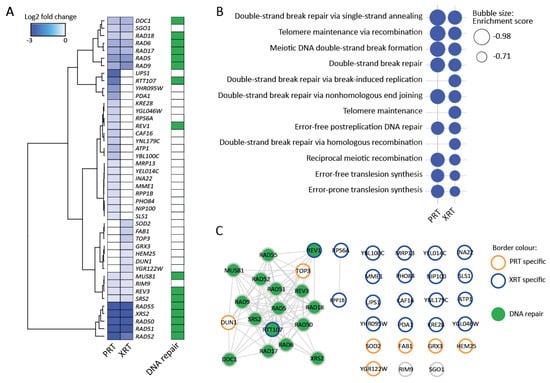
Figure 3.
Barcode sequencing (Bar-Seq) experiment identifies DNA repair genes as important for the response to proton (PRT) and photon radiotherapy (XRT). (A) Heatmap for sensitive gene deletions. Sensitive gene deletion mutants were selected based on two cut-offs: FDR < 0.05 and log2FC < −0.5 in either the PRT or XRT experiment. Hierarchical clustering was performed by the Ward.D2 method based on the Euclidian distance. Dark green squares indicate genes annotated with the GO term “DNA repair”. (B) Enriched GO terms identified using GSEA. GSEA was applied on a ranked gene list based on the log2FC calculated in edgeR. Genes were chosen based on two cut-offs: false discovery rate (FDR) < 0.05 and log2 fold change (log2FC) < −0.5 in either the proton or photon experiment. For GSEA, the FDR was set at 0.25. (C) Interaction network of genes in (A). The network was built in String [43] and visualized in Cytoscape [44]. Nodes with orange borders are deletions only found to be sensitive after PRT. Nodes with blue borders are deletions only found to be sensitive after XRT. Nodes with grey borders are deletions sensitive to both PRT and XRT. Nodes are colored in green when the gene is involved in DNA repair.
The sensitivity of these genes for PRT and XRT was confirmed by plating assays (Figure S3). Genes not involved in DNA repair also did not cluster in the network confirming the lack of a functional enrichment for these genes.
Bar-Seq also identified a few genes that when deleted reduced sensitivity to PRT or XRT. However, no functional enrichment for these genes could be identified. Moreover, these genes were often not shared between the PRT and XRT experiment. Additionally, the reduced sensitivity could not be validated by plating assays. Therefore, we only focused on the genes that, when deleted, cause sensitivity to PRT or XRT.
Constructing an interaction network for all genes found in either or both RNA-Seq and Bar-Seq highlighted that both experiments detected the importance of the DNA damage response after PRT or XRT; the experiments overlapped at the level of the GO term “response to DNA damage stimulus” (Figure S4). Two genes, RAD51 and DUN1, both involved in the DNA damage response, showed a differential expression in the RNA-Seq at 30 min after PRT and sensitivity in the Bar-Seq in after PRT. However, this does not imply that the other genes are less important, since not all key genes needed for DNA damage survival are induced by the stress [45]. Even though heat shock proteins were upregulated in the response to PRT and XRT, the Bar-Seq experiment revealed that none of the proteins by themselves are essential for survival following PRT or XRT. This is likely due to the redundancy among heat shock proteins [46,47,48,49]; where gene products are often functionally similar.
2.4. PRT Induces DNA Damage and Protein Damage in S. cerevisiae
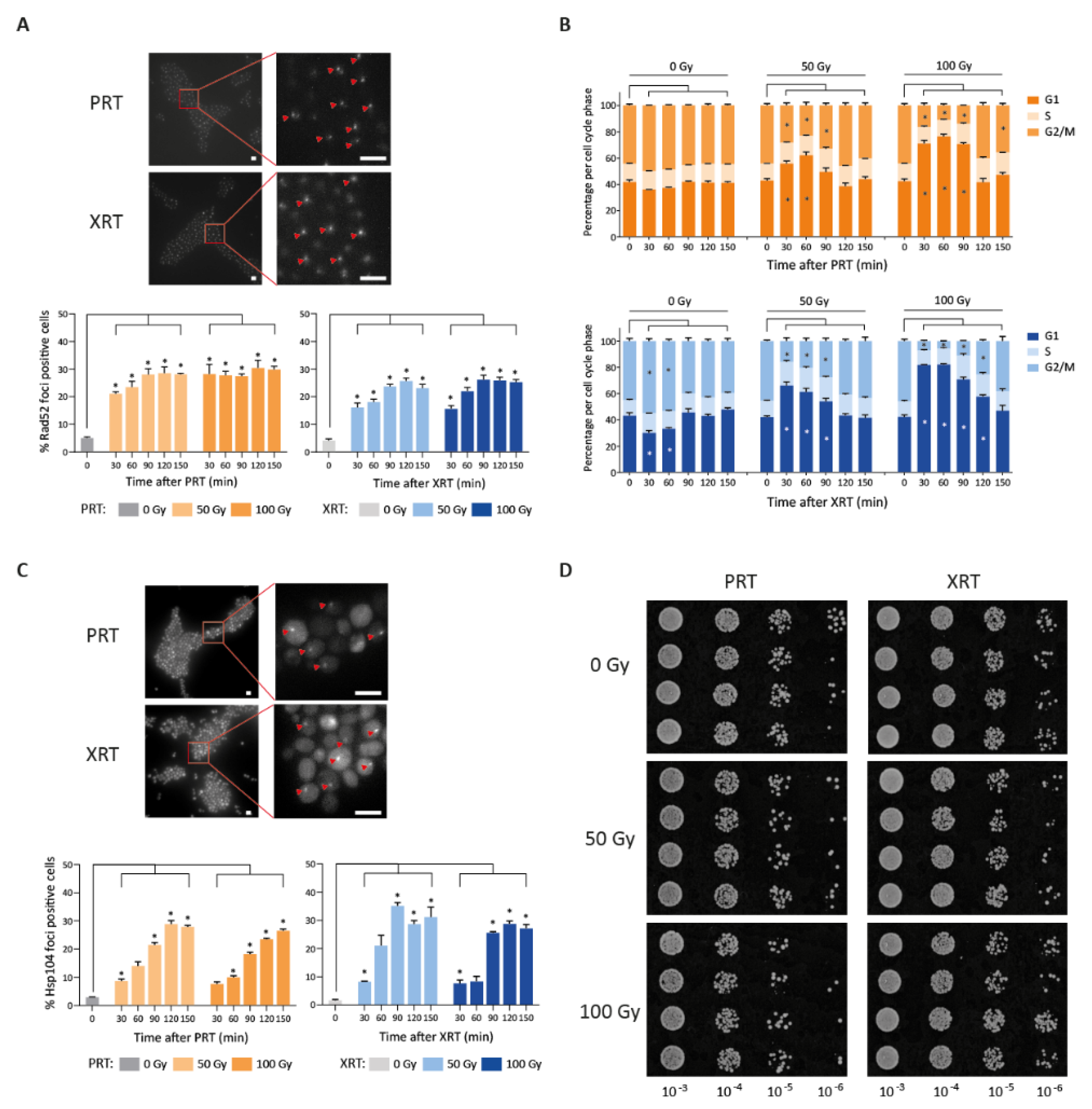
Our results so far confirmed the importance of DNA repair in response to XRT, and showed that the same response is also crucial for surviving PRT. Next, we set out to further investigate whether both radiation types induced a similar amount of DNA damage. First, we investigated the kinetics of double strand break repair by tracking cellular Rad52 foci under the microscope after radiotherapy, which serve as a marker for double strand breaks (Figure 4A) [50]. PRT and XRT both showed an induction of Rad52 foci with a maximum fold increase of 5.8 and 6.2 both at 120 min after 50 Gy PRT and XRT, respectively. A dose of 100 Gy resulted in a maximal 6.1-fold increase in foci 120 min after PRT, and a maximal increase of 6.3-fold 90 min after XRT (Figure 4A). Although the increase in Rad52 foci was comparable between the two radiation types, the dynamics appear slightly different. More Rad52 foci were observed at 30 min after PRT compared to XRT. PRT caused 4.33- and 5.88-fold induction after 50 Gy and 100 Gy, respectively, while XRT-induced Rad52 foci increased 4.19- and 4.03-fold after 50 Gy and 100 Gy, respectively. Moreover, after 150 min, more Rad52 foci have been cleared in cells subjected to XRT compared to PRT (23% versus 28% at 50 Gy and 25% versus 30% at 100 Gy after XRT and PRT, respectively), indicating PRT-induced DNA damage is more persistent.
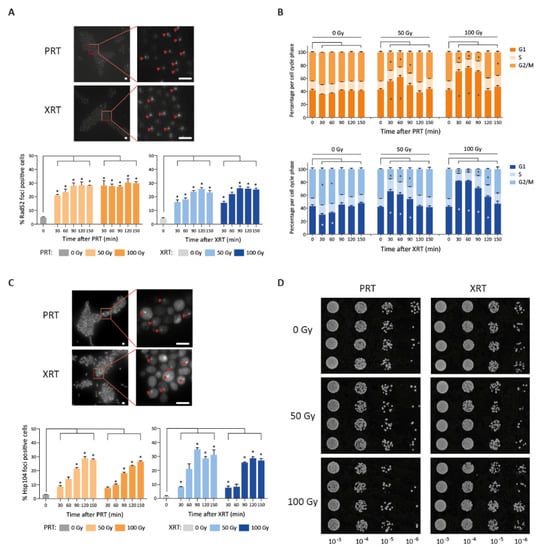
Figure 4.
Proton (PRT) and photon (XRT) radiotherapy induce DNA damage and protein damage in S. cerevisiae. (A) Repair kinetics after photon and proton irradiation. Example image of Rad52 foci 90 min after 100 Gy irradiation (strain LAV87) (top). Scale bars represent 5 µm. The percentage of Rad52 foci positive cells is depicted over time after 50 Gy or 100 Gy of PRT and XRT (bottom). All data are represented as the mean ± SEM for n = 3. For each replicate, at least 1500 cells were analyzed per condition. p-values were calculated using ANOVA with multiple comparisons test compared to the 0 min control. * p-values < 0.05. (B) Cell cycle distribution after proton (top) and photon (bottom) irradiation. The percentage of the population in G1, S and G2/M is depicted. p-values were calculated using ANOVA with multiple comparisons test compared to the 0 min control. * p-values < 0.05. (C) Example image of Hsp104 foci 90 min after 100 Gy irradiation (strain LAV91) (top). Scale bars represent 5 µm. The percentage of Hsp104 foci positive cells over time is depicted after 50 Gy and 100 Gy of PRT and XRT (bottom). All data are represented as the mean ± SEM for n = 3. For each replicate, at least 1500 cells were analyzed per condition. p-values were calculated using ANOVA with multiple comparisons test compared to the 0 min control. * p-values < 0.05. (D) Spotting assay with proteasome inhibitor MG-132 in combination with 50 Gy or 100 Gy proton or photon radiation. Additionally, 10-fold dilutions of mock irradiated or irradiated cells are spotted on YPD containing 50 µM MG-132. Images were taken after 48 h of incubation at 30 °C.
Next, we investigated possible changes in cell cycle dynamics after irradiation. Cell cycle arrest is generally recognized as an indispensable part of the DNA damage response. For example, previous studies showed that Gamma-irradiated synchronized yeast cultures show a prolonged G1 arrest that is dose-dependent and proportional to the amount of DNA damage [51]. In line with this previous result, we found that irradiation induced a dose-dependent increase in cells that are in the G1 phase, independent of the irradiation type (Figure 4B). The G1 arrest became less pronounced after 90 min for both PRT and XRT after 50 Gy and 100 Gy, although the 100 Gy treatment resulted in slightly more cells remaining in G1.
Besides the importance of the DNA damage response, RNA-Seq analysis revealed that genes involved in protein folding, mostly heat shock proteins, were more highly upregulated after PRT. To further investigate this, we examined the amount of protein aggregates after PRT and XRT by fluorescently tagging HSP104, a disaggregase that localizes to protein aggregates [52,53,54,55]. As S. cerevisiae is radioresistant, we expected protein aggregates to be cleared efficiently by the proteostasis machinery. Therefore, we additionally inhibited the central degradation machinery of protein control, the proteasome, using the potent proteasomal inhibitor MG-132. This is commonly performed to study protein aggregate formation [55,56]. Both PRT and XRT showed an increase in Hsp104 foci with a maximum of 29% foci positive cells at 120 min after 50 Gy PRT and 35% at 90 min after 50 Gy XRT (Figure 4C), indicating the presence of protein aggregates after both radiation types. This further suggests the induction of proteotoxic stress after both PRT and XRT, confirming the results from our RNA-Seq experiment. At 50 Gy the maximum amount of Hsp104 foci is reached at 90 min after XRT versus 120 min after PRT, indicating a delay of protein aggregate formation after PRT. Compared to the maximum, at 150 min after 50 Gy PRT the amount of Hsp104 foci has decreased from 29% to 28% while after XRT the decrease was more pronounced (from 35% to 31%). At 100 Gy a different profile was observed: while PRT-induced Hsp104 foci were still increasing from 90 to 150 min (from 18% to 23% to 26%), XRT-induced Hsp104 foci remained more constant (26% to 29% to 27%). As a result, similar to the kinetics observed for DNA damage markers, PRT-induced Hsp104 foci, indicative for protein aggregates, showed a more persistent pattern compared to XRT. We next investigated if differences in the aggregates formation could be linked to an increase in endoplasmic reticulum (ER) stress, since ER stress is one of the consequences of increased unfolded or misfolded proteins [57,58]. Moreover, transcriptome analysis revealed that genes involved in protein refolding were upregulated after 50 Gy PRT, strengthening the hypothesis of the presence of unfolded or misfolded proteins after PRT. We therefore measured the activation of the unfolded protein response (UPR), which is known to be a good marker for ER stress [59,60]. Specifically, we assayed the HAC1 mRNA splicing status by PCR across the intron of HAC1 mRNA [58,61,62]. The results showed activation of the UPR for both PRT and XRT, confirming that both PRT and XRT induce proteotoxic stress, although the extent of UPR activation varied between replicates at different timepoints (Figure S5). Next, to assess the effect of the radiation-induced protein damage on survival of S. cerevisiae, we performed spotting assays with the proteasomal inhibitor MG-132 in combination with RT. In line with the presence of more persistent protein aggregates, PRT resulted in stronger growth inhibition compared to XRT upon proteasome inhibition with MG-132 (Figure 4D).
3. Discussion
To gain more insight into the molecular mechanisms affected by PRT, we used S. cerevisiae as a model to investigate PRT radiobiology on a functional and transcriptional level and compared it with conventional XRT. To our knowledge, this is the first time an integrated systems biology approach has been used to shed light on the molecular mechanisms affected by PRT.
The ability to repair DNA damage caused by ionizing radiation has been shown to be a central factor in the response to XRT in several different organisms [26,63,64]. This agrees with our observations, which show an induction of genes involved in the DNA damage response following XRT. Moreover, our functional assays also showed that deleting certain central DNA damage response genes severely affects survival after XRT. Importantly, we find comparable results after PRT, suggesting that for both types of radiation, DNA repair genes are important for survival. Multiple studies in mammalian cells suggest that PRT-induced DNA lesions are more dependent on homologous recombination for repair, and less on the more common mechanism of non-homologous end joining [16,17,33,65]. Our results confirmed the involvement of homologous recombination for survival after PRT. However, it has to be noted that the pathway for non-homologous end joining is much less used by yeast cells [66,67], which implies that we cannot exclude that this pathway is also involved in repair after PRT in other organisms. Furthermore, the Bar-Seq experiment also identified several genes that affect survival but are not involved in DNA repair. However, we were unable to confirm their role through plating assays. This could be because, as any genome-wide screen, Bar-Seq often yields a few false positive hits, or because the plating assays are less suitable to detect subtle differences in survival after RT.
The importance of the DNA damage response after PRT and XRT was further examined by assessing the kinetics of the DNA damage response through tracking of Rad52 foci and characterizing cell cycle kinetics. Both PRT and XRT resulted in similar induction of DNA damage and G1 cell cycle phase arrest. However, clearance of DNA damage seemed slightly slower after PRT, indicating possible differences in the kinetics of DNA damage or repair. In mammalian cell research, contradictory results have been reported, with some studies detecting a prolonged cell cycle arrest after PRT [68,69,70], while others found no difference [13], or report a shorter cell cycle arrest for PRT [16].
Apart from DNA damage, radiation is known to also induce protein damage, which could make a substantial contribution to the radiobiological response that is elicited [71,72]. Strikingly, our transcriptional analysis reveals that PRT elicits a much stronger induction of genes involved in proteotoxic stress, for example genes encoding chaperones. Heat shock proteins play a significant role in the protein folding and the proteostasis machinery in general. However, they also help maintain efficient DNA repair, by stabilizing specific protein complexes (reviewed in Dubrez et al., 2020) [73]. Here, using proteasomal inhibition, we demonstrated a comparable induction of protein aggregates after PRT and XRT. Like the DNA damage response kinetics, we noted a slightly more persistent response for PRT, suggesting subtle differences in the induction and/or clearance of protein damage. Importantly, proteasomal inhibition in combination with PRT resulted in more extensive growth inhibition compared to XRT, suggesting that PRT elicits a stronger proteotoxic stress, resulting in a greater dependence on proper proteasomal functioning after PRT. It is interesting to note that in the study of Schultzhaus et al. the GO term “protein catabolism” was also slightly more upregulated in the particle irradiation group including PRT, deuterons and α-particles, compared to gamma-irradiation. However, in contrast to our study, the potential role and kinetic of this process was not further investigated.
Despite the stronger transcriptional and molecular response after PRT, our results show that irradiation of S. cerevisiae with medium to high doses (25 to 100 Gy) of PRT and XRT resulted in similar survival curves. However, it should be noted that, although non-significant, PRT resulted in on average 1.05 ± 0.07 times more cell killing compared to XRT.
We believe that our study offers more robust conclusions than previous work because the use of the eukaryotic model S. cerevisiae allowed for us to perform multiple replicates of very controlled genome-scale analyses, which allowed us to gain a complete view on the cellular response to PRT and XRT. In addition, we were able to verify the results of our genome-wide assays, RNA-Seq and Bar-Seq, by probing the consequences of deleting individual genes and by performing specific experiments looking in more detail at the affected molecular mechanisms. However, it should be underlined that our study also has some limitations. Along with the existing molecular toolbox, the large body of knowledge on its genetics and physiology, and the conserved nature of core processes such as the DNA damage response, caused S. cerevisiae to be a particularly suitable and popular model to study eukaryotic radiobiology [23,24,25,26,31,32]. However, as is the case for all studies using models, the results cannot be expanded to other, more complex, organisms without verification. In addition, a quantitative comparison between XRT and PRT is difficult because the treatments are carried out with different irradiation machines stationed at distinct locations, which implies the risk of minor differences that are independent of the radiation type. It also is important to note that PRT assays were performed in the plateau region of the depth–dose profile, i.e., at a depth located before the Bragg peak, to ensure that all cells would receive a uniform dose. This plateau region is characterized by a lower energy deposition and hence lower cell toxicity compared to the Bragg peak region, which could explain why we did not observe a statistical difference in biological effectiveness between PRT and XRT.
Thus, while the results clearly show a trend of similarities in the response to PRT and XRT, the more subtle differences we find in DNA and protein damage must further be investigated. However, the increased dependency of proper proteasomal functioning after PRT, in particular, offers novel therapeutic potential for proteasome inhibitors in combination with PRT. While the results need to be confirmed in higher organisms, our genome-scale analysis is difficult to achieve in mammalian tumor cells because the large heterogeneity between cell lines and the more restricted experimental possibilities. By offering a first in-depth view on the radiobiological response of S. cerevisiae to PRT in comparison to the more conventional XRT, our study provides a good basis for further, more targeted analyses and thus helps bridge the gap between the current biological uncertainties and translation of PRT to clinical settings.
4. Materials and Methods
4.1. Strains and Media
All strains used in this study were derived from the haploid auxotrophic S288c strain BY4741 (MATa). A full list of strains used can be found in Table S1. Primers used for strain construction and verification were ordered from IDT. Primer sequences are listed in Table S2.
Deletion strains were constructed with a HygB cassette amplified from the pCB1 plasmid using primers with at least 40 bp sequence homology to the target DNA [74]. For fluorescent tagging, yECitrine was amplified from the pKT140 plasmid using primers with 40 bp sequence homology to the target DNA [75]. A list of plasmids used in this study and their genotype can be found in Table S3. PCR products were used for yeast transformation using a LiAc-based procedure. Transformants were verified by PCR using specific check primers.
Yeast was cultured in peptone dextrose (YPD) medium containing 2% bacteriological peptone (Lab M, Heywood, UK), 1% yeast extract (Lab M, Heywood, UK) and 2% glucose (Sigma, St. Louis, MO, USA). For solid plates 2% agar (Lab M, Heywood, UK) was added. YPD containing 200 µg/L Hygromycin B (Invitrogen, Waltham, MA, USA) or 200 µg/mL G418 (Formedium, Swaffham, UK) was used for selection of yeast transformants.
4.2. Irradiations
Photon irradiations (6 MV, 5.3 Gy/min, LET = 2.36 keV/µm [76]) were performed on a TrueBeam medical linear accelerator (Varian Medical Systems, Palo Alto, CA, USA) placed at the Department of Radiation Oncology in University Hospitals Leuven. Proton irradiation (62 MeV, 17 Gy/min, mean LET = 10.51 keV/µm) was carried out at the Centre-de-Ressources du Cyclotron (Université Catholique de Louvain, Louvain-la-Neuve, Belgium). Samples were irradiated at the entrance of the proton depth–dose curve. Due to the separate locations of the two irradiation facilities, photon and proton experiments were never performed on the same day. For both irradiation types a physical dose of 50 Gy or 100 Gy was used. The biological effects of the physical radiation dose for both irradiation types were determined by investigation of relaxation patterns of 90% supercoiled phiX174 RF1 DNA (0.5 g/L DNA, Thermo Fisher Scientific, Waltham, MA, USA) as previously described [77,78]. The 5386 bp ϕDNA was diluted to a final concentration of 7.5 nM and mock-irradiated or irradiated with 5 and 15 Gy of protons and photons. Irradiated samples were analyzed on a 1% agarose gel. Densitometry was performed in ImageJ (Figure S6). The percentage of DNA damage, which was calculated as the ratio of circular DNA (cDNA) and linear DNA (lDNA) over the total amount of DNA (scDNA + cDNA + lDNA), showed a dose-dependent increase independent of the radiation type.
4.3. Plating Assays
Plating assays were performed in triplicate to determine the survival of yeast after PRT or XRT. Cells in exponential phase (OD ~0.2) were irradiated with indicated doses of PRT or XRT and were plated on YPD. The number of colonies was determined after 48 h of growth at 30 °C. Plates were considered countable when containing 30–300 colonies. The survival fraction was calculated by dividing the number of colonies from the irradiated conditions by the control condition.
4.4. RNA Sequencing and Analysis
Four replicate exponential phase cultures of the wild type strain (KV447) were irradiated with 50 Gy PRT or XRT. At 30 min and 90 min after irradiation samples were taken for RNA extraction. Therefore, 1.5 mL of the culture was centrifuged and after washing the pellet once in ice-cold DEPC water, the pellet was put on dry ice. Pellets were stored at −80 °C until RNA extraction. RNA was extracted using the MasterPure Yeast RNA Purification Kit (Lucigen, Middleton, WI, USA) with DNase treatment. RNA concentrations were measured with Qubit (Thermofisher). RNA sequencing was performed on a DNBseq platform at BGI Genomics. Sequences were assessed for quality control using FASTQC version 0.11.9 and were aligned against the S288c genome of S. cerevisiae using HISAT2 version 2.2.1 [79,80]. Mapped reads were counted using HTSeq version 0.13.5 [81]. Differential expression analysis was performed in edgeR 3.32.1 [82]. Heatmaps were visualized with the heatmap.2 function in the gplots R package [83]. Clustering was performed by the ward.D2 method using Euclidean distance as a distance measure. Enriched GO terms were identified using Gene Set Enrichment analysis (GSEA) version 4.1.0 [84,85]. GSEA was applied on a ranked gene list based on the log2FC calculated in edgeR. Networks were built in String version 11.5 and visualized in Cytoscape version 3.9.1. [43,44].
4.5. Barcode Sequencing and Analysis
For the Bar-Seq experiment the haploid MATa deletion collection was pooled as described by Perez-Samper et al. [86]. Shortly, the deletion collection in 96-well plates was thawed and 3 µL from each well was inoculated in YPD medium containing 200 µg/mL G418. All cultures were grown to stationary phase (OD600 > 1.0). 50 µL of each stationary phase culture was pooled and mixed. The pool was divided into 1 mL aliquots and frozen at −80 °C.
Two replicate deletion collection pool aliquots were thawed and pregrown to exponential phase (OD = 0.2) in 2% YPD and an initial sample was taken for each replicate and frozen as a 25% glycerol stock at −80 °C. For each replicate, the exponential phase cultures were diluted in YP in duplicate. One sample was irradiated with 50 Gy PRT or XRT, the other was mock irradiated as a control. After irradiation, the pools were grown for 8 generations in 2% YPD and at end of the growth cycle final samples were taken. Genomic DNA was extracted from all samples using a zymolyase-based protocol. UPTAGs and DNTAGs were amplified by separate PCR reactions using the primers described in Perez-Samper et al. [86]. DNA concentrations were measured with Qubit (Thermo Fisher Scientific, Waltham, MA, USA). Amplified UPTAGs and DNTAGs coming from the same sample were pooled at equal concentrations and sent for paired-end sequencing on an Illumina NextSeq 500 with a 1000× coverage at the Nucleomics Core facility in Leuven.
Sequences were assessed for quality control using FASTQC version 0.11.9 [79]. Barcodes were extracted from the raw reads using cutadapt version 2.3 [87]. Extracted barcodes were matched to the recharacterized deletion barcodes and counted using Barcas version 1.0 [42,88]. A maximum of two inexact matches was allowed. Differential expression analysis was performed using edgeR 3.32.1 [82]. The Log2 fold change in counted barcodes was calculated comparing the irradiated condition to the corresponding non-irradiated condition. Visualization of the results in a heatmap and network and GSEA was achieved similarly as for the RNA-Seq experiment.
4.6. Fluorescence Microscopy
Three replicate exponential phase cultures (LAV87 and LAV91) were mock irradiated or irradiated with 50 Gy or 100 Gy PRT or XRT. At indicated timepoints after irradiation, cells were fixed using 4% formaldehyde, washed in PBS and stored at 4 °C in PBS until imaging. In the case of Hsp104 fluorescent foci 50 µM MG-132 (Selleckchem, Houston, TX, USA) was added 2 h before irradiation. YFP fluorescence was imaged using an inverted automated Nikon TiE fluorescence microscope (Nikon, Tokio, Japan) at 60× magnification. In total, 11 fluorescent images were obtained at 0.3 µm intervals along the z-axis. Foci were counted from the fluorescent image using ImageJ. The number of foci positive cells was calculated by dividing the number of fluorescent foci by the total amount of cells as counted on the brightfield image. At least 1500 cells were analyzed per sample.
4.7. Cell Cycle Analysis
Three replicate exponential phase cultures of the wild type strain (KV447) were irradiated with 50 Gy or 100 Gy PRT or XRT. Samples were taken at the indicated timepoints, fixated with 70% fresh, ice-cold ethanol and stored at −20 °C. All samples were stained with propidium iodide (PI) for flow cytometry analysis. Briefly, fixated cells were pelleted and washed with sodium citrate buffer (0.035 g citric acid monohydrate (Sigma, St. Louis, MO, USA) and 7.3 g trisodium citrate dihydrate (VWR, Radnor, PA, USA) in 500 mL water). Next, pellets were resuspended in sodium citrate buffer containing 0.25 mg/mL RNase A (Thermo Fisher Scientific, Waltham, MA, USA) and incubated at 50 °C for one hour. Samples were centrifuged and pellets were resuspended in sodium citrate buffer containing 16 μg/mL PI (Sigma-Aldrich, St. Louis, MO, USA). The samples were analyzed with a flow cytometer at 488 nm (FACSverse, BD Biosciences, Franklin Lakes, NJ, USA). A total of 50,000 cells were analyzed per sample.
4.8. HAC1 mRNA Splicing Detection
Splicing status of HAC1 mRNA was detected using RT-PCR across the intron. Therefore, sampling and total RNA extraction was performed similar as described in for RNA-Seq samples. From these samples, cDNA was generated using the QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany) following the manufacturers guidelines. This cDNA was used a template for PCR amplification using an intron-flanking primer pair (Table S2). PCR products were analyzed on a 2% agarose gel.
4.9. Spot Assays with MG-132
Three replicate exponential phase cultures (LAV90) were incubated with 50 µM MG-132 (Selleckchem, Houston, TX, USA) for 2 h before mock irradiation or irradiation with 50 Gy or 100 Gy PRT or XRT. After irradiation, 10-fold serial dilutions (start OD = 1) were spotted on YPD containing 50 µM MG-132. Plates were incubated at 30 °C and imaged after 48 h of incubation.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms23105493/s1. References [74,75,89,90] are cited in the supplementary materials.
Author Contributions
Conceptualization, R.D., K.V., K.J.V. and S.N.; Formal analysis, L.V. (Laura Vanderwaeren) and L.V. (Laura Vandemaele); Funding acquisition, R.D.; Investigation, L.V. (Laura Vanderwaeren) and L.V. (Laura Vandemaele); Methodology, L.V. (Laura Vanderwaeren); Project administration, R.D.; Supervision, R.D., K.J.V. and S.N.; Visualization, L.V. (Laura Vanderwaeren); Writing—original draft, L.V. (Laura Vanderwaeren); Writing—review and editing, L.V. (Laura Vanderwaeren), R.D., K.V., K.J.V. and S.N.; L.V. (Laura Vanderwaeren) performed the sequencing experiments, strain construction, molecular assays and data analysis. L.V. (Laura Vandemaele) aided in plating assays and cell cycle experiments. R.D. and K.V. supervised the experimental work. S.N. and K.J.V. coordinated the project. L.V. (Laura Vanderwaeren) wrote the manuscript with input from all authors. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the KU Leuven Research Fund (C24/17/073). R.D. is supported by Emmanuel van der Schueren fellowship for postdoctoral researchers from Kom op Tegen Kanker and by Flemish Foundation of Scientific Research (FWO-Vlaanderen, 1521019N). Research in the laboratory of K.J.V. is supported by VIB, KU Leuven, FWO and European Research Council (ERC) Consolidator Grant CoG682009. S.N. is supported by clinical research mandate from the Flemish Foundation of Scientific Research (FWO-Vlaanderen, 18B4122N).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study can be made available upon request from the corresponding author.
Acknowledgments
The authors would like to thank Nancy Postiau at the Centre-de-Ressources du Cyclotron from Université Catholique de Louvain for her assistance in proton irradiations and Pierre Morsomme from Louvain Institute of Biomolecular Science and Technology at Université Catholique de Louvain for allowing us to use his lab space.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Delaney, G.; Jacob, S.; Featherstone, C.; Barton, M. The role of radiotherapy in cancer treatment: Estimating optimal utilization from a review of evidence-based clinical guidelines. Cancer 2005, 104, 1129–1137. [Google Scholar] [CrossRef] [PubMed]
- Barton, M.B.; Jacob, S.; Shafiq, J.; Wong, K.; Thompson, S.R.; Hanna, T.P.; Delaney, G.P. Estimating the demand for radiotherapy from the evidence: A review of changes from 2003 to 2012. Radiother. Oncol. 2014, 112, 140–144. [Google Scholar] [CrossRef]
- De Ruysscher, D.; Niedermann, G.; Burnet, N.G.; Siva, S.; Lee, A.W.M.; Hegi-Johnson, F. Radiotherapy toxicity. Nat. Rev. Dis. Prim. 2019, 5, 13. [Google Scholar] [CrossRef] [PubMed]
- Holliday, E.B.; Frank, S.J. Proton Radiation Therapy for Head and Neck Cancer: A Review of the Clinical Experience to Date. Int. J. Radiat. Oncol. 2014, 89, 292–302. [Google Scholar] [CrossRef]
- Brouwer, C.L.; Steenbakkers, R.J.H.M.; Bourhis, J.; Budach, W.; Grau, C.; Grégoire, V.; van Herk, M.; Lee, A.; Maingon, P.; Nutting, C.; et al. CT-based delineation of organs at risk in the head and neck region: DAHANCA, EORTC, GORTEC, HKNPCSG, NCIC CTG, NCRI, NRG Oncology and TROG consensus guidelines. Radiother. Oncol. 2015, 117, 83–90. [Google Scholar] [CrossRef]
- Lee, M.; Wynne, C.; Webb, S.; Nahum, A.E.; Dearnaley, D. A comparison of proton and megavoltage X-ray treatment planning for prostate cancer. Radiother. Oncol. 1994, 33, 239–253. [Google Scholar] [CrossRef]
- Liao, Z.; Lee, J.J.; Komaki, R.; Gomez, D.R.; O’Reilly, M.S.; Fossella, F.V.; Blumenschein, G.R.; Heymach, J.V.; Vaporciyan, A.A.; Swisher, S.G.; et al. Bayesian adaptive randomization trial of passive scattering proton therapy and intensity-modulated photon radiotherapy for locally advanced non-small-cell lung cancer. J. Clin. Oncol. 2018, 36, 1813–1822. [Google Scholar] [CrossRef]
- Finnberg, N.; Wambi, C.; Ware, J.H.; Kennedy, A.R.; El-Deiry, W.S. Gamma-radiation (GR) triggers a unique gene expression profile associated with cell death compared to proton radiation (PR) in mice in vivo. Cancer Biol. Ther. 2008, 7, 2023–2033. [Google Scholar] [CrossRef]
- Wang, L.; Yang, L.; Han, S.; Zhu, J.; Li, Y.; Wang, Z.; Fan, Y.H.; Lin, E.; Zhang, R.; Sahoo, N.; et al. Patterns of protein expression in human head and neck cancer cell lines differ after proton vs photon radiotherapy. Head Neck 2020, 42, 289–301. [Google Scholar] [CrossRef]
- Narang, H.; Bhat, N.; Gupta, S.K.; Santra, S.; Choudhary, R.K.; Kailash, S.; Krishna, M. Differential activation of mitogen-activated protein kinases following high and low LET radiation in murine macrophage cell line. Mol. Cell. Biochem. 2009, 324, 85–91. [Google Scholar] [CrossRef]
- Di Pietro, C.; Piro, S.; Tabbì, G.; Ragusa, M.; Di Pietro, V.; Zimmitti, V.; Cuda, F.; Anello, M.; Consoli, U.; Salinaro, E.T.; et al. Cellular and molecular effects of protons: Apoptosis induction and potential implications for cancer therapy. Apoptosis 2006, 11, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Schultzhaus, Z.; Chen, A.; Shuryak, I.; Wang, Z. The Transcriptomic and Phenotypic Response of the Melanized Yeast Exophiala dermatitidis to Ionizing Particle Exposure. Front. Microbiol. 2021, 0, 3442. [Google Scholar] [CrossRef]
- Alan Mitteer, R.; Wang, Y.; Shah, J.; Gordon, S.; Fager, M.; Butter, P.P.; Jun Kim, H.; Guardiola-Salmeron, C.; Carabe-Fernandez, A.; Fan, Y. Proton beam radiation induces DNA damage and cell apoptosis in glioma stem cells through reactive oxygen species. Sci. Rep. 2015, 5, 13961. [Google Scholar] [CrossRef] [PubMed]
- Oeck, S.; Szymonowicz, K.; Wiel, G.; Krysztofiak, A.; Lambert, J.; Koska, B.; Iliakis, G.; Timmermann, B.; Jendrossek, V. Relating linear energy transfer to the formation and resolution of DNA repair foci after irradiation with equal doses of X-ray photons, plateau, or bragg-peak protons. Int. J. Mol. Sci. 2018, 19, 3779. [Google Scholar] [CrossRef] [PubMed]
- Bracalente, C.; Ibañez, I.L.; Molinari, B.; Palmieri, M.; Kreiner, A.; Valda, A.; Davidson, J.; Durán, H. Induction and Persistence of Large γH2AX Foci by High Linear Energy Transfer Radiation in DNA-Dependent protein kinase–Deficient Cells. Int. J. Radiat. Oncol. 2013, 87, 785–794. [Google Scholar] [CrossRef] [PubMed]
- Grosse, N.; Fontana, A.O.; Hug, E.B.; Lomax, A.; Coray, A.; Augsburger, M.; Paganetti, H.; Sartori, A.A.; Pruschy, M. Deficiency in homologous recombination renders mammalian cells more sensitive to proton versus photon irradiation. Int. J. Radiat. Oncol. Biol. Phys. 2014, 88, 175–181. [Google Scholar] [CrossRef] [PubMed]
- Fontana, A.O.; Augsburger, M.A.; Grosse, N.; Guckenberger, M.; Lomax, A.J.; Sartori, A.A.; Pruschy, M.N. Differential DNA repair pathway choice in cancer cells after proton- and photon-irradiation. Radiother. Oncol. 2015, 116, 374–380. [Google Scholar] [CrossRef]
- Kachroo, A.H.; Laurent, J.M.; Yellman, C.M.; Meyer, A.G.; Wilke, C.O.; Marcotte, E.M. Systematic humanization of yeast genes reveals conserved functions and genetic modularity. Science 2015, 348, 921–925. [Google Scholar] [CrossRef]
- Pereira, C.; Coutinho, I.; Soares, J.; Bessa, C.; Leão, M.; Saraiva, L. New insights into cancer-related proteins provided by the yeast model. FEBS J. 2012, 279, 697–712. [Google Scholar] [CrossRef]
- Duina, A.A.; Miller, M.E.; Keeney, J.B. Budding yeast for budding geneticists: A primer on the Saccharomyces cerevisiae model system. Genetics 2014, 197, 33–48. [Google Scholar] [CrossRef]
- Botstein, D.; Fink, G.R. Yeast: An experimental organism for 21st Century biology. Genetics 2011, 189, 695–704. [Google Scholar] [CrossRef] [PubMed]
- Botstein, D.; Chervitz, S.A.; Cherry, J.M. Yeast as a model organism. Science 1997, 277, 1259–1260. [Google Scholar] [CrossRef] [PubMed]
- Resnick, M.A.; Cox, B.S. Yeast as an honorary mammal. Mutat. Res. Mol. Mech. Mutagen. 2000, 451, 1–11. [Google Scholar] [CrossRef]
- Dolling, J.A.; Boreham, D.R.; Bahen, M.E.; Mitchel, R.E.J. Role of RAD9-dependent cell-cycle checkpoints in the adaptive response to ionizing radiation in yeast, Saccharomyces cerevisiae. Int. J. Radiat. Biol. 2000, 76, 1273–1279. [Google Scholar] [CrossRef] [PubMed]
- Boreham, D.R.; Trivedi, A.; Weinberger, P.; Mitchel, R.E.J. The involvement of topoisomerases and DNA polymerase I in the mechanism of induced thermal and radiation resistance in yeast. Radiat. Res. 1990, 123, 203–212. [Google Scholar] [CrossRef] [PubMed]
- Game, J.C.; Birrell, G.W.; Brown, J.A.; Shibata, T.; Baccari, C.; Chu, A.M.; Williamson, M.S.; Brown, J.M. Use of a genome-wide approach to identify new genes that control resistance of Saccharomyces cerevisiae to ionizing radiation. Radiat. Res. 2003, 160, 14–24. [Google Scholar] [CrossRef]
- Lee, J.H.; Choi, I.Y.; Kil, I.S.; Kim, S.Y.; Yang, E.S.; Park, J.W. Protective role of superoxide dismutases against ionizing radiation in yeast. Biochim. Biophys. Acta 2001, 1526, 191–198. [Google Scholar] [CrossRef]
- Fasullo, M.; Bennett, T.; Ahching, P.; Koudelik, J. The Saccharomyces cerevisiae RAD9 checkpoint reduces the DNA damage-associated stimulation of directed translocations. Mol. Cell. Biol. 1998, 18, 1190. [Google Scholar] [CrossRef]
- Bennett, C.B.; Lewis, L.K.; Karthikeyan, G.; Lobachev, K.S.; Jin, Y.H.; Sterling, J.F.; Snipe, J.R.; Resnick, M.A. Genes required for ionizing radiation resistance in yeast. Nat. Genet. 2001, 29, 426–434. [Google Scholar] [CrossRef]
- Rostek, C.; Turner, E.L.; Robbins, M.; Rightnar, S.; Xiao, W.; Obenaus, A.; Harkness, T.A.A. Involvement of homologous recombination repair after proton-induced DNA damage. Mutagenesis 2008, 23, 119–129. [Google Scholar] [CrossRef]
- Aylon, Y.; Kupiec, M. DSB repair: The yeast paradigm. DNA Repair 2004, 3, 797–815. [Google Scholar] [CrossRef] [PubMed]
- James, A.P.; Werner, M.M. The radiobiology of yeast. Radiat. Bot. 1965, 5, 359–382. [Google Scholar] [CrossRef]
- Szymonowicz, K.; Krysztofiak, A.; van der Linden, J.; Kern, A.; Deycmar, S.; Oeck, S.; Squire, A.; Koska, B.; Hlouschek, J.; Vüllings, M.; et al. Proton Irradiation increases the necessity for homologous recombination repair along with the indispensability of non-homologous end joining. Cells 2020, 9, 889. [Google Scholar] [CrossRef] [PubMed]
- Winter, M.; Dokic, I.; Schlegel, J.; Warnken, U.; Debus, J.; Abdollahi, A.; Schnölzer, M. Deciphering the acute cellular phosphoproteome response to irradiation with X-rays, protons and carbon ions. Mol. Cell. Proteomics 2017, 16, 855–872. [Google Scholar] [CrossRef] [PubMed]
- Chabes, A.; Georgieva, B.; Domkin, V.; Zhao, X.; Rothstein, R.; Thelander, L. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by relaxed feedback inhibition of ribonucleotide reductase. Cell 2003, 112, 391–401. [Google Scholar] [CrossRef]
- Guo, X.; Zhang, M.; Gao, Y.; Lu, D.; Li, W.; Zhou, L. Repair characteristics and time-dependent effects in response to heavy-ion beam irradiation in Saccharomyces cerevisiae: A comparison with X-ray irradiation. Appl. Microbiol. Biotechnol. 2020, 104, 4043–4057. [Google Scholar] [CrossRef]
- Guo, X.; Zhang, M.; Liu, R.; Gao, Y.; Yang, Y.; Li, W.; Lu, D. Repair characteristics and time-dependent effects in Saccharomyces cerevisiae cells after X-ray irradiation. World J. Microbiol. Biotechnol. 2019, 35, 1. [Google Scholar] [CrossRef]
- Gasch, A.P.; Spellman, P.T.; Kao, C.M.; Carmel-Harel, O.; Eisen, M.B.; Storz, G.; Botstein, D.; Brown, P.O. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell. 2000, 11, 4241–4257. [Google Scholar] [CrossRef]
- Gasch, A.P.; Huang, M.; Metzner, S.; Botstein, D.; Elledge, S.J.; Brown, P.O. Genomic expression responses to DNA-damaging agents and the regulatory role of the yeast ATR Homolog Mec1p. Mol. Biol. Cell 2001, 12, 2987–3003. [Google Scholar] [CrossRef]
- Su, C.; Gao, G.; Schneider, S.; Helt, C.; Weiss, C.; O’Reilly, M.A.; Bohmann, D.; Zhao, J. DNA damage induces downregulation of histone gene expression through the G1 checkpoint pathway. EMBO J. 2004, 23, 1133. [Google Scholar] [CrossRef]
- Giaever, G.; Chu, A.M.; Ni, L.; Connelly, C.; Riles, L.; Véronneau, S.; Dow, S.; Lucau-Danila, A.; Anderson, K.; André, B.; et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002, 418, 387–391. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.M.; Heisler, L.E.; Mellor, J.; Kaper, F.; Thompson, M.J.; Chee, M.; Roth, F.P.; Giaever, G.; Nislow, C. Quantitative phenotyping via deep barcode sequencing. Genome Res. 2009, 19, 1836–1842. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Birrell, G.W.; Brown, J.A.; Wu, H.I.; Giaever, G.; Chu, A.M.; Davis, R.W.; Brown, J.M. Transcriptional response of Saccharomyces cerevisiae to DNA-damaging agents does not identify the genes that protect against these agents. Proc. Natl. Acad. Sci. USA 2002, 99, 8778–8783. [Google Scholar] [CrossRef]
- Werner-Washburne, M.; Stone, D.E.; Craig, E.A. Complex interactions among members of an essential subfamily of hsp70 genes in Saccharomyces cerevisiae. Mol. Cell. Biol. 1987, 7, 2568–2577. [Google Scholar] [CrossRef]
- Sahi, C.; Craig, E.A. Network of general and specialty J protein chaperones of the yeast cytosol. Proc. Natl. Acad. Sci. USA 2007, 104, 7163. [Google Scholar] [CrossRef]
- Youker, R.T.; Walsh, P.; Beilharz, T.; Lithgow, T.; Brodsky, J.L. Distinct roles for the Hsp40 and Hsp90 molecular chaperones during cystic fibrosis transmembrane conductance regulator degradation in yeast. Mol. Biol. Cell 2004, 15, 4787–4797. [Google Scholar] [CrossRef]
- Verghese, J.; Abrams, J.; Wang, Y.; Morano, K.A. Biology of the heat shock response and protein chaperones: Budding yeast (Saccharomyces cerevisiae) as a model system. Microbiol. Mol. Biol. Rev. 2012, 76, 115–158. [Google Scholar] [CrossRef]
- Lisby, M.; Rothstein, R.; Mortensen, U.H. Rad52 forms DNA repair and recombination centers during S phase. Proc. Natl. Acad. Sci. USA 2001, 98, 8276–8282. [Google Scholar] [CrossRef]
- Fitz Gerald, J.N.; Benjamin, J.M.; Kron, S.J. Robust G1 checkpoint arrest in budding yeast: Dependence on DNA damage signaling and repair. J. Cell Sci. 2002, 115, 1749–1757. [Google Scholar] [CrossRef] [PubMed]
- Glover, J.R.; Lindquist, S. Hsp104, Hsp70, and Hsp40: A novel chaperone system that rescues previously aggregated proteins. Cell 1998, 94, 73–82. [Google Scholar] [CrossRef]
- Tkach, J.M.; Glover, J.R. Amino acid substitutions in the C-terminal AAA+ module of Hsp104 prevent substrate recognition by disrupting oligomerization and cause high temperature inactivation. J. Biol. Chem. 2004, 279, 35692–35701. [Google Scholar] [CrossRef] [PubMed]
- Sathyanarayanan, U.; Musa, M.; Bou Dib, P.; Raimundo, N.; Milosevic, I.; Krisko, A. ATP hydrolysis by yeast Hsp104 determines protein aggregate dissolution and size in vivo. Nat. Commun. 2020, 11, 5226. [Google Scholar] [CrossRef] [PubMed]
- Kaganovich, D.; Kopito, R.; Frydman, J. Misfolded proteins partition between two distinct quality control compartments. Nature 2008, 454, 1088–1095. [Google Scholar] [CrossRef]
- Specht, S.; Miller, S.B.M.; Mogk, A.; Bukau, B. Hsp42 is required for sequestration of protein aggregates into deposition sites in Saccharomyces cerevisiae. J. Cell Biol. 2011, 195, 617–629. [Google Scholar] [CrossRef]
- Walter, P.; Ron, D. The unfolded protein response: From stress pathway to homeostatic regulation. Science 2011, 334, 1081–1086. [Google Scholar] [CrossRef]
- Hernández-Elvira, M.; Torres-Quiroz, F.; Escamilla-Ayala, A.; Domínguez-Martin, E.; Escalante, R.; Kawasaki, L.; Ongay-Larios, L.; Coria, R. The unfolded protein response pathway in the yeast Kluyveromyces lactis. A comparative View among yeast species. Cells 2018, 7, 106. [Google Scholar] [CrossRef]
- Labunskyy, V.M.; Gerashchenko, M.V.; Delaney, J.R.; Kaya, A.; Kennedy, B.K.; Kaeberlein, M.; Gladyshev, V.N. Lifespan extension conferred by endoplasmic reticulum secretory pathway deficiency requires induction of the unfolded protein response. PLoS Genet. 2014, 10, e1004019. [Google Scholar] [CrossRef]
- Bruch, A.; Klassen, R.; Schaffrath, R. Unfolded protein response suppression in yeast by loss of tRNA modifications. Genes 2018, 9, 516. [Google Scholar] [CrossRef]
- Wu, H.; Ng, B.S.H.; Thibault, G. Endoplasmic reticulum stress response in yeast and humans. Biosci. Rep. 2014, 34, 321–330. [Google Scholar] [CrossRef] [PubMed]
- Rüegsegger, U.; Leber, J.H.; Walter, P. Block of HAC1 mRNA translation by long-range base pairing is released by cytoplasmic splicing upon induction of the unfolded protein response. Cell 2001, 107, 103–114. [Google Scholar] [CrossRef]
- Jaklevic, B.R.; Su, T.T. Relative Contribution of DNA Repair, cell cycle checkpoints, and cell death to survival after DNA damage in Drosophila Larvae. Curr. Biol. 2004, 14, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Pollard, J.M.; Gatti, R.A. Clinical radiation sensitivity with DNA repair disorders: An overview. Int. J. Radiat. Oncol. Biol. Phys. 2009, 74, 1323. [Google Scholar] [CrossRef]
- Bright, S.J.; Flint, D.B.; Chakraborty, S.; McFadden, C.H.; Yoon, D.S.; Bronk, L.; Titt, U.; Mohan, R.; Grosshans, D.R.; Sumazin, P.; et al. Nonhomologous end joining is more important than proton linear energy transfer in dictating cell death. Int. J. Radiat. Oncol. Biol. Phys. 2019, 105, 1119–1125. [Google Scholar] [CrossRef]
- Shrivastav, M.; De Haro, L.P.; Nickoloff, J.A. Regulation of DNA double-strand break repair pathway choice. Cell Res. 2008, 18, 134–147. [Google Scholar] [CrossRef]
- Chapman, J.R.; Taylor, M.R.G.; Boulton, S.J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 2012, 47, 497–510. [Google Scholar] [CrossRef]
- Moertel, H.; Georgi, J.C.; Distel, L.; Eyrich, W.; Fritsch, M.; Grabenbauer, G.; Sauer, R. Effects of low energy protons on clonogenic survival, DSB repair and cell cycle in human glioblastoma cells and B14 fibroblasts. Radiother. Oncol. 2004, 73, S115–S118. [Google Scholar] [CrossRef]
- Antoccia, A.; Sgura, A.; Berardinelli, F.; Cavinato, M.; Cherubini, R.; Gerardi, S.; Tanzarella, C. Cell cycle perturbations and genotoxic effects in human primary fibroblasts induced by low-energy protons and X/γ-rays. J. Radiat. Res. 2009, 50, 457–468. [Google Scholar] [CrossRef]
- Narang, H.; Kumar, A.; Bhat, N.; Pandey, B.N.; Ghosh, A. Effect of proton and gamma irradiation on human lung carcinoma cells: Gene expression, cell cycle, cell death, epithelial-mesenchymal transition and cancer-stem cell trait as biological end points. Mutat. Res.-Fundam. Mol. Mech. Mutagen. 2015, 780, 35–46. [Google Scholar] [CrossRef]
- Daly, M.J. Death by protein damage in irradiated cells. DNA Repair 2012, 11, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Riley, P.A. Free radicals in biology: Oxidative stress and the effects of ionizing radiation. Int. J. Radiat. Biol. 1994, 65, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Dubrez, L.; Causse, S.; Borges Bonan, N.; Dumétier, B.; Garrido, C. Heat-shock proteins: Chaperoning DNA repair. Oncogene 2020, 39, 516–529. [Google Scholar] [CrossRef] [PubMed]
- Brown, C.A.; Murray, A.W.; Verstrepen, K.J. Rapid expansion and functional divergence of subtelomeric gene families in yeasts. Curr. Biol. 2010, 20, 895–903. [Google Scholar] [CrossRef] [PubMed]
- Sheff, M.A.; Thorn, K.S. Optimized cassettes for fluorescent protein tagging in Saccharomyces cerevisiae. Yeast 2004, 21, 661–670. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, H.; Kanai, T.; Kase, Y.; Matsumoto, Y.; Furusawa, Y.; Fujita, Y.; Saitoh, H.; Itami, J.; Kohno, T. Relation between lineal energy distribution and relative biological effectiveness for photon beams according to the microdosimetric kinetic model. J. Radiat. Res. 2011, 52, 75–81. [Google Scholar] [CrossRef]
- Folkard, M.; Prise, K.M.; Vojnovic, B.; Davies, S.; Roper, M.J.; Michael, B.D. Measurement of DNA damage by electrons with energies between 25 and 4000 EV. Int. J. Radiat. Biol. 1993, 64, 651–658. [Google Scholar] [CrossRef]
- Boudaïffa, B.; Cloutier, P.; Hunting, D.; Huels, M.A.; Sanche, L. Resonant formation of DNA strand breaks by low-energy (3 to 20 eV) electrons. Science 2000, 287, 1658–1660. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 18 October 2021).
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Warnes, G.R.; Bolker, B.; Bonebakker, L.; Gentleman, R.; Huber, W.; Liaw, A.; Lumley, T.; Maechler, M.; Magnusson, A.; Moeller, S.; et al. gplots: Various R Programming Tools for Plotting Data. R Package, Version 3.1.1. The Comprehensive R Archive Network. 2020. Available online: https://CRAN.R-project.org/package=gplots (accessed on 10 April 2022).
- Mootha, V.K.; Lindgren, C.M.; Eriksson, K.-F.; Subramanian, A.; Sihag, S.; Lehar, J.; Puigserver, P.; Carlsson, E.; Ridderstråle, M.; Laurila, E.; et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 2003, 34, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed]
- Perez-Samper, G.; Cerulus, B.; Jariani, A.; Vermeersch, L.; Barrajón Simancas, N.; Bisschops, M.M.M.; van den Brink, J.; Solis-Escalante, D.; Gallone, B.; De Maeyer, D.; et al. The crabtree effect shapes the Saccharomyces cerevisiae lag phase during the switch between different carbon sources. MBio 2018, 9, e01331-18. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.J. 2011, 17, 10. [Google Scholar] [CrossRef]
- Mun, J.; Kim, D.-U.; Hoe, K.-L.; Kim, S.-Y. Genome-wide functional analysis using the barcode sequence alignment and statistical analysis (Barcas) tool. BMC Bioinformatics 2016, 17, 475. [Google Scholar] [CrossRef] [PubMed]
- Kassambara, A.; Mundt, F. factoextra: Extract and Visualize the Results of Multivariate Data Analyses. R Package Version 1.0.7. 2017. [Google Scholar]
- Baker Brachmann, C.; Davies, A.; Cost, G.J.; Caputo, E.; Li, J.; Hieter, P.; Boeke, J.D. Designer Deletion Strains derived from Saccharomyces cerevisiae S288C: A Useful set of Strains and Plasmids for PCR-mediated Gene Disruption and Other Applications. Yeast 1998, 115–132. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).