Effect and Mechanism Analysis of Pig FUT8 Gene on Resistance to Escherichia coli F18 Infection
Abstract
1. Introduction
2. Results
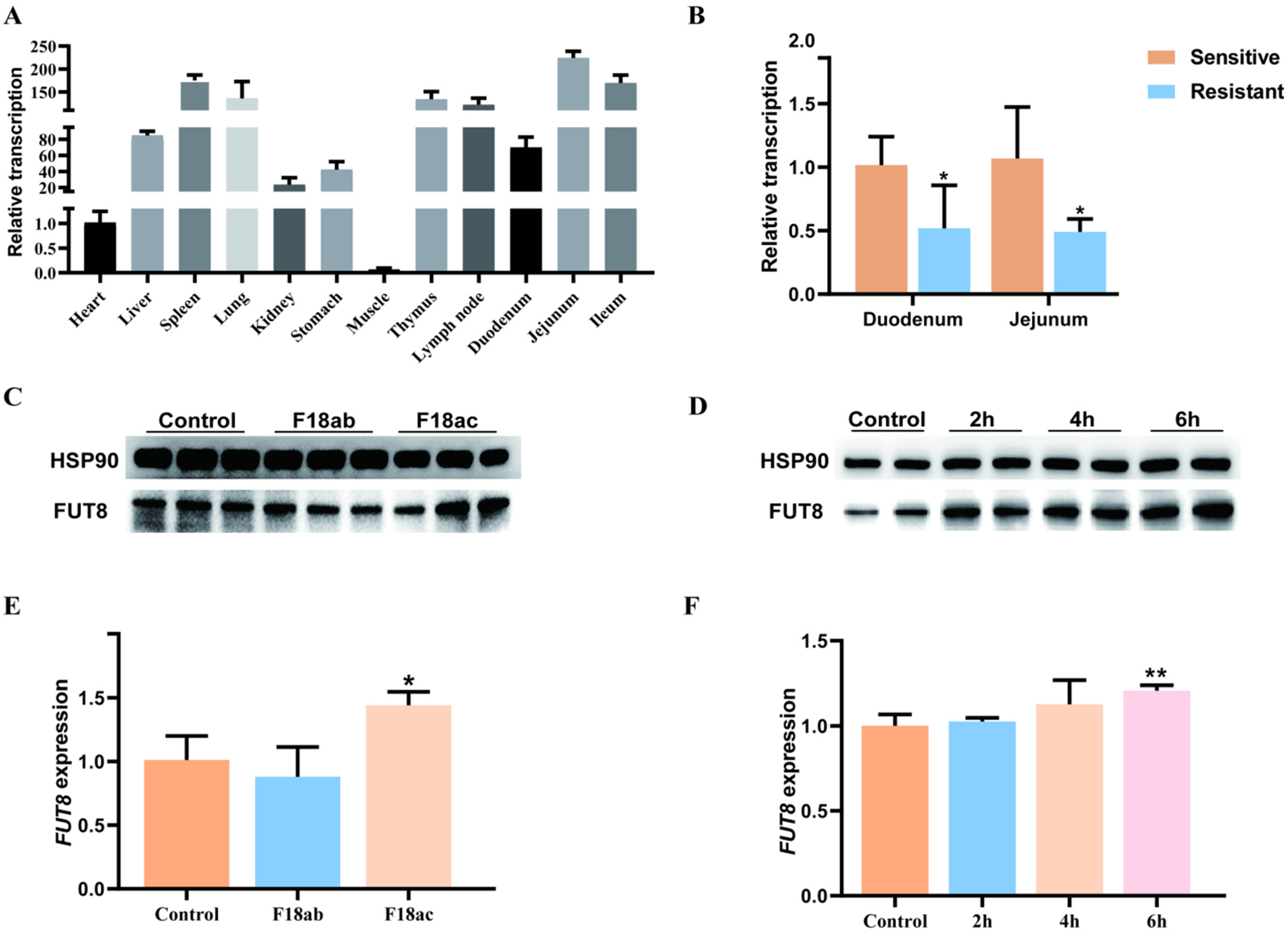
2.1. Association Analysis of FUT8 Expression and E. coli F18 Infection
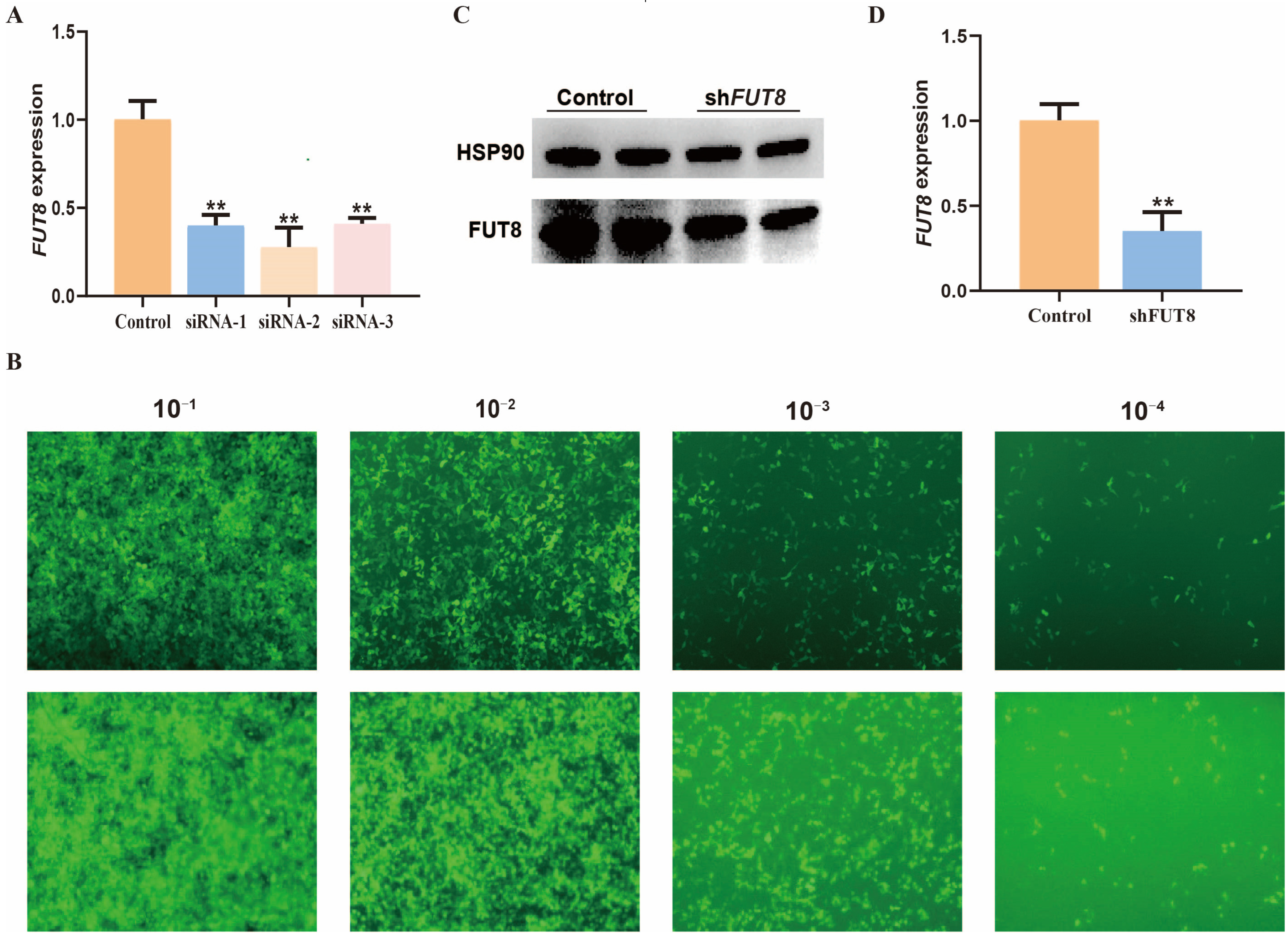
2.2. Establishment of IPEC-J2 Cell Lines with FUT8 Interference
2.3. Effect of FUT8 Knockdown on the Adhesion Ability of E. coli F18
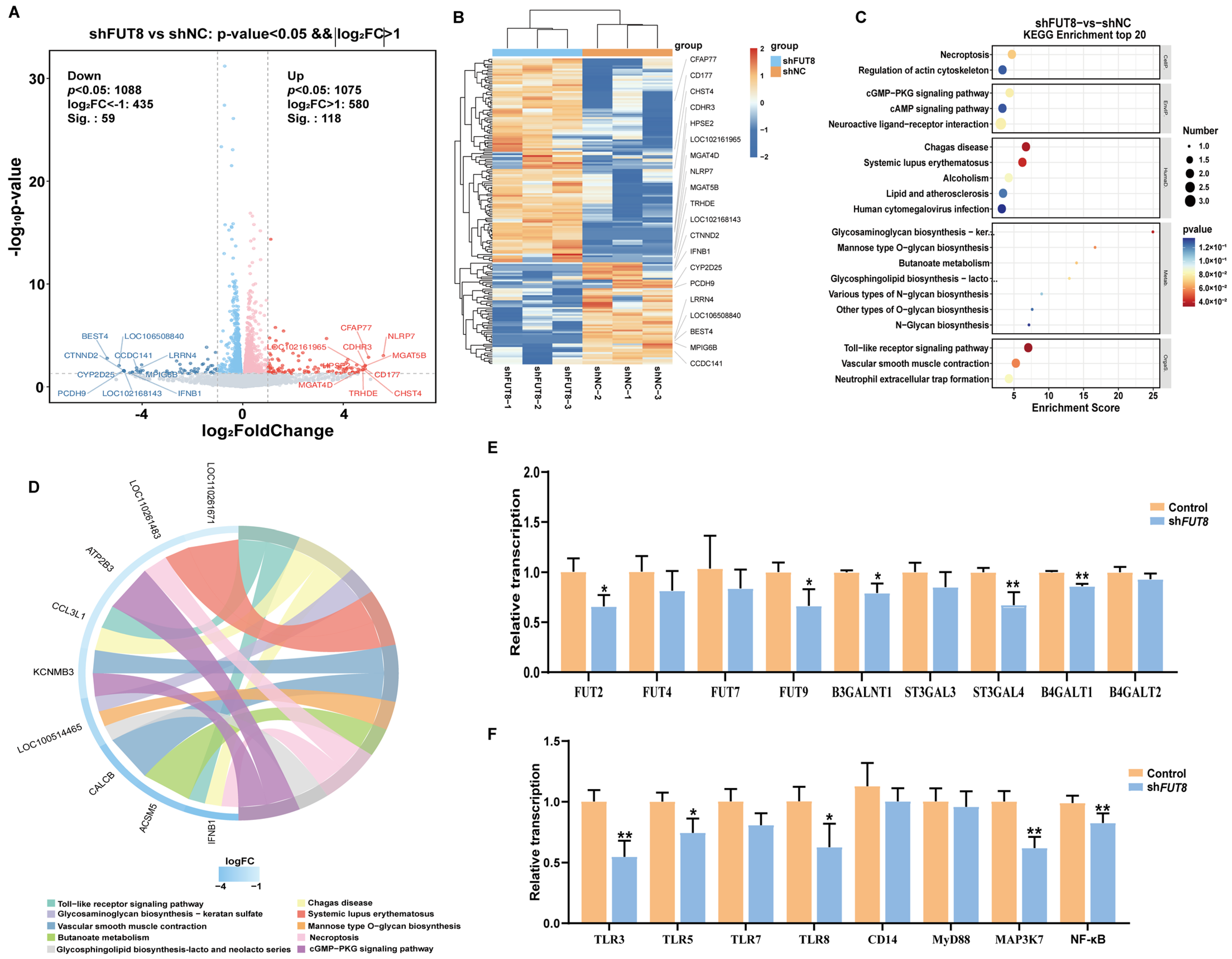
2.4. Transcriptome Sequencing Analysis of IPEC-J2 Cells after FUT8 Knockdown by RNA-seq
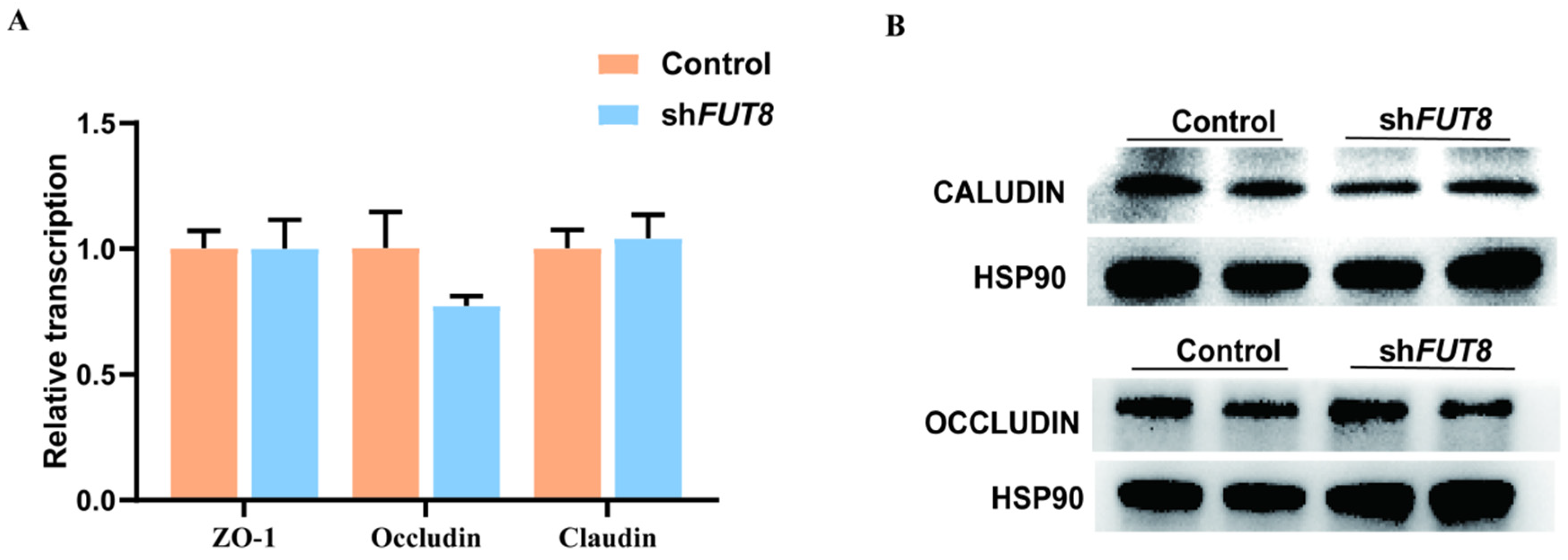
2.5. Analysis of the Effect of FUT8 on Tight Junction Genes Expression
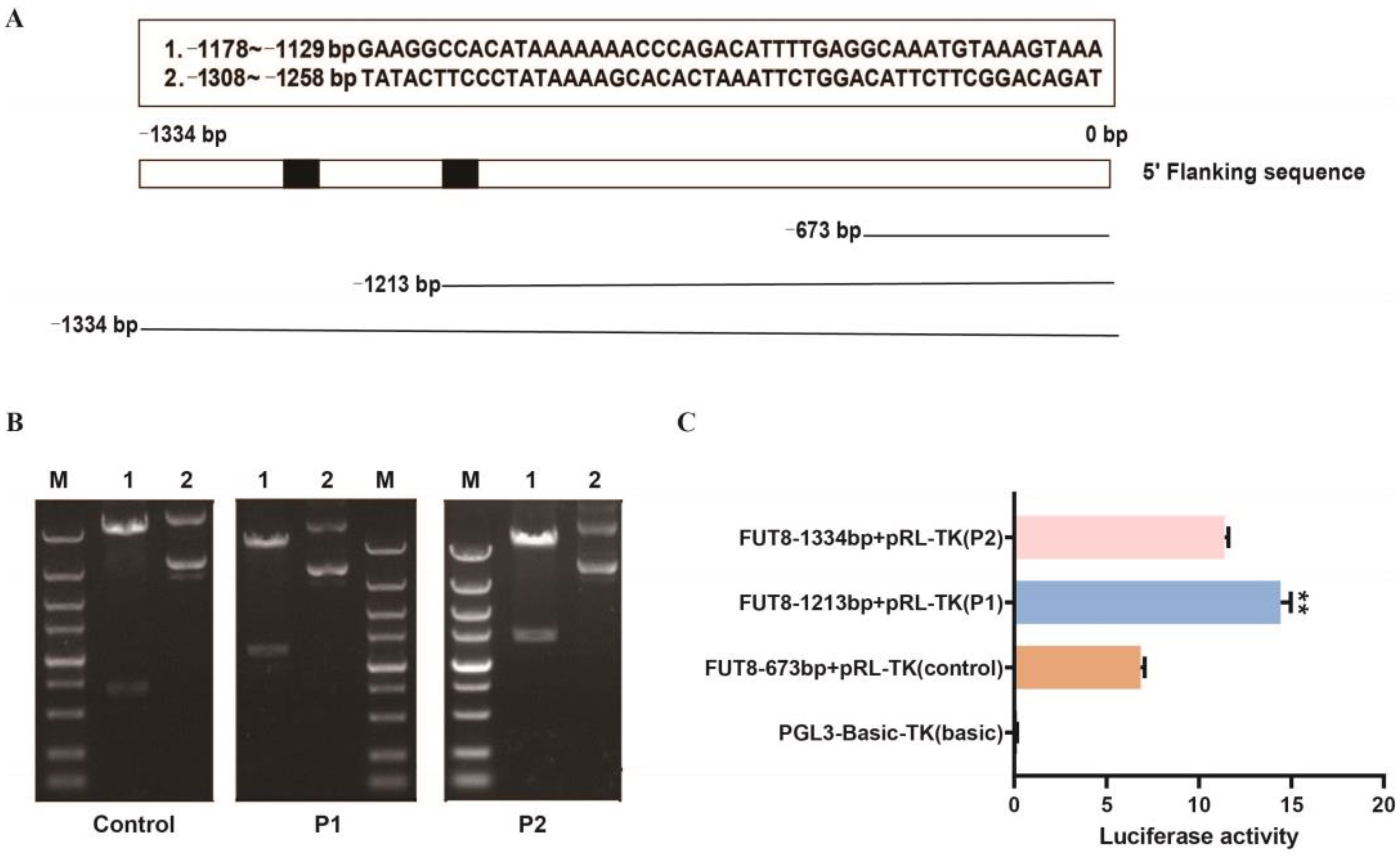
2.6. Identification Analysis of Pig FUT8 Core Promoter Region
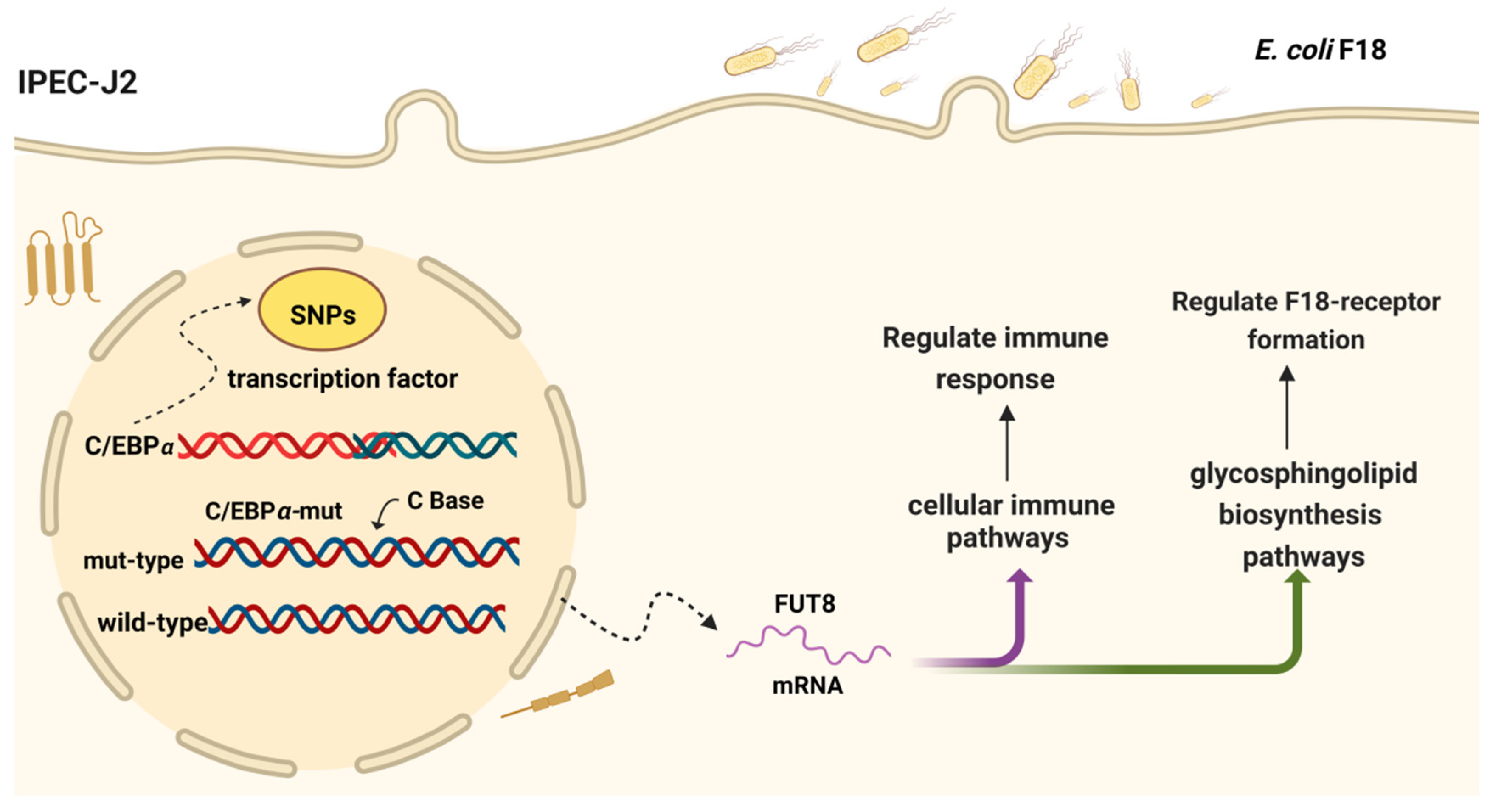
2.7. Important SNP and Transcription Factor Identification Analysis of Pig the FUT8 Gene Promoters
3. Discussion
4. Materials and Methods
4.1. Ethics Statement
4.2. Reagents and Animal Material
4.3. E. coli F18 Exposure and LPS Induction
4.4. Primer Design and Sequence Synthesis
4.5. Cell Culture
4.6. RT-qPCR Analysis
4.7. Western Blot Analysis
4.8. Subcellular Fraction Extraction
4.9. Immunohistochemical Analysis
4.10. In vitro Adherence Assays with E. coli F18
4.11. Indirect Immunofluorescence (IFA)
4.12. Single-Nucleotide Polymorphisms in the Porcine FUT8 Gene Promote
4.13. Dual-luciferase Reporter Assays
4.14. Transcription Factor Prediction
4.15. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| FUT8 | Fucosyltransferase 8 |
| PWD | Post-weaning diarrhea |
| E. coli | Escherichia coli |
| IPEC-J2 | Intestinal porcine epithelial cell |
| IFA | Indirect immunofluorescence assay |
| SEM | Scanning electron microscopy |
| LPS | Lipopolysaccharide |
| FBS | Fetal bovine serum |
| DMEM | Dulbecco’s modified Eagle’s medium |
| NF-κB | Nuclear factor κB |
| SNPs | Single-nucleotide polymorphisms |
| DEGs | Differentially expressed genes |
| TSSs | Transcription start sites |
| KEGG | Kyoto Encyclopedia of Genes and Genome |
| EMSA | Electrophoretic mobility shift assay |
| CoIP | Co-immunoprecipitation |
| TU | Transducing units |
References
- Boldin, B. Persistence and spread of gastro-intestinal infections: The case of enterotoxigenic Escherichia coli in piglets. Bull. Math. Biol. 2008, 70, 2077–2101. [Google Scholar] [CrossRef] [PubMed]
- Imberechts, H.; Bertschinger, H.U.; Stamm, M.; Sydler, T.; Pohl, P.; De Greve, H.; Hernalsteens, J.P.; Van Montagu, M.; Lintermans, P. Prevalence of F107 fimbriae on Escherichia coli isolated from pigs with oedema disease or postweaning diarrhoea. Vet. Microbiol. 1994, 40, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Fairbrother, J.M.; Nadeau, É.; Gyles, C.L. Escherichia coli in postweaning diarrhea in pigs: An update on bacterial types, pathogenesis, and prevention strategies. Anim. Health Res. Rev. 2005, 6, 17–39. [Google Scholar] [CrossRef] [PubMed]
- Sinha, R.; Sahoo, N.R.; Shrivastava, K.; Kumar, P.; Qureshi, S.; De Kumar, U.; Kumar, A.; Kumar, G.V.P.P.S.R.; Bhushan, B. Resistance to ETEC F4/F18-mediated piglet diarrhoea: Opening the gene black box. Trop. Anim. Health Prod. 2019, 51, 1307–1320. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, P.; Fontanals-Cirera, B.; Sokolova, E.; Jacob, S.; Vaiana, C.A.; Argibay, D.; Davalos, V.; McDermott, M.; Nayak, S.; Darvishian, F.; et al. A Systems Biology Approach Identifies FUT8 as a Driver of Melanoma Metastasis. Cancer Cell. 2017, 31, 804–819. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Fukuda, T.; Isaji, T.; Duan, C.; Song, W.; Wang, Y.; Gu, J. 1,6-Fucosyltransferase contributes to cell migration and proliferation as well as to cancer stemness features in pancreatic carcinoma. Biochim. Biophys. Acta Gen. Subj. 2021, 1865, 129870. [Google Scholar] [CrossRef]
- Bastian, K.; Scott, E.; Elliott, D.J.; Munkley, J. FUT8 Alpha-(1,6)-Fucosyltransferase in Cancer. Int. J. Mol. Sci. 2021, 22, 455. [Google Scholar] [CrossRef]
- Zahid, D.; Zhang, N.; Fang, H.; Gu, J.; Li, M.; Li, W. Loss of core fucosylation suppressed the humoral immune response in Salmonella typhimurium infected mice. J. Microbiol. Immunol. Infect. 2021, 54, 606–615. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Yu, R.; Ma, B.; Yang, Y.; Jiao, X.; Liu, Y.; Cao, H.; Dong, W.; Liu, L.; Ma, K.; et al. Core fucosylation of IgG B cell receptor is required for antigen recognition and antibody production. J. Immunol. 2015, 194, 2596–2606. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Bai, Y.Q.; He, C.Y.; Yan, Q.L.; Fang, H.; Li, Z. Intestinal microbial structure of core fucosyltransferase FUT8 gene knockout mice. Chin. J. Microecol. 2017, 29, 5. [Google Scholar]
- Osumi, D.; Takahashi, M.; Miyoshi, E.; Yokoe, S.; Taniguchi, N. Core fucosylation of E-cadherin enhances cell–cell adhesion in human colon carcinoma WiDr cells. Cancer Sci. 2009, 100, 888–895. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Guo, D.; Tan, Z.; Du, J.; Guan, F.; Li, X. Fucosyltransferase 8 regulation and breast cancer suppression by transcription factor activator protein 2γ. Cancer Sci. 2021, 112, 3190–3204. [Google Scholar] [CrossRef] [PubMed]
- Ihara, H.; Okada, T.; Taniguchi, N.; Ikeda, Y. Involvement of the α-helical and Src homology 3 domains in the molecular assembly and enzymatic activity of human α1, 6-fucosyltransferase, FUT8. Biochim. Biophys. Acta. Gen. Subj. 2020, 1864, 129596. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Shinzaki, S.; Iijima, H.; Wakamatsu, K.; Iwamoto, C.; Sobajima, T.; Kuwahara, R.; Hiyama, S.; Hayashi, Y.; Takamatsu, S. Core Fucosylation on T Cells, Required for Activation of T-Cell Receptor Signaling and Induction of Colitis in Mice, Is Increased in Patients with Inflammatory Bowel Disease. Gastroenterology 2016, 150, 1620–1632. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, K.; Wakamatsu, K.; Fujii, H.; Shinzaki, S.; Takamatsu, S.; Kitazume, S.; Kamada, Y.; Takehara, T.; Taniguchi, N.; Miyoshi, E. Core fucose is essential glycosylation for CD14-dependent Toll-like receptor 4 and Toll-like receptor 2 signalling in macrophages. J. Biochem. 2019, 165, 227–237. [Google Scholar] [CrossRef] [PubMed]
- Xiaoli, L.; Fengbin, H.; Shihui, H.; Xi, N.; Sheng, L.; Zhou, W.; Xueqin, R.; Jiafu, W. Detection of genomic structure variants associated with wrinkled skin in Xiang pig by next generation sequencing. Aging 2021, 13, 24710–24739. [Google Scholar] [CrossRef] [PubMed]
- Zi, C.; Wu, Z.; Wang, J.; Huo, Y.B.; Zhu, G.; Wu, S.; Bao, W. Transcriptional activity of the FUT1 gene promoter region in pigs. Int. J. Mol. Sci. 2013, 14, 24126–24134. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, K. PCR-Based Detection Methods for Single-Nucleotide Polymorphism or Mutation: Real-Time PCR and Its Substantial Contribution Toward Technological Refinement. Adv. Clin. Chem. 2017, 80, 45–72. [Google Scholar]
- Shastry, B.S. SNPs: Impact on gene function and phenotype. Methods Mol. Biol. 2009, 578, 3–22. [Google Scholar] [PubMed]
- Dong, W.H.; Dai, C.H.; Sun, L.; Wang, J.; Sun, S.Y.; Zhu, G.Q.; Wu, S.L.; Bao, W.B. Expression of key glycosphingolipid biosynthesis-globo series pathway genes in Escherichia coli F18-resistant and Escherichia coli F18-sensitive piglets. Anim. Genet. 2016, 47, 428–435. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Fan, H.; Jin, J.; Gao, S.; Huang, R.; Wu, S.; Bao, W. Insight into mechanisms of pig lncRNA FUT3-AS1 regulating E. coli F18-bacterial diarrhea. PLoS Pathog. 2022, 18, e1010584. [Google Scholar] [CrossRef] [PubMed]
- Aarestrup, F.M.; Jorsal, S.E.; Ahrens, P.; Jensen, N.E.; Meyling, A. Molecular characterization of Escherichia coli strains isolated from pigs with edema disease. J. Clin. Microbiol. 1997, 35, 20–24. [Google Scholar] [CrossRef] [PubMed]
- Batisson, I.; Guimond, M.P.; Girard, F.; An, H.; Zhu, C.; Oswald, E.; Fairbrother, J.M.; Jacques, M.; Harel, J. Characteriza-tion of the novel factor paa involved in the early steps of the adhesion mechanism of attaching and effacing Escherichia coli. Infect. Immun. 2003, 71, 4516–4525. [Google Scholar] [CrossRef] [PubMed]
- Nagy, B.; Fekete, P.Z. Enterotoxigenic Escherichia coli (ETEC) in farm animals. Vet. Res. 1999, 30, 259–284. [Google Scholar] [PubMed]
- Sham, H.P.; Shames, S.R.; Croxen, M.A.; Ma, C.; Chan, J.M.; Khan, M.A.; Wickham, M.E.; Deng, W.; Finlay, B.B.; Vallance, B.A. Attaching and effacing bacterial effector NleC suppresses epithelial inflammatory responses by inhibiting NF-κB and p38 mitogen-activated protein kinase activation. Infect. Immun. 2011, 79, 3552–3562. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fukuda, T.; Hashimoto, H.; Okayasu, N.; Kameyama, A.; Onogi, H.; Nakagawasai, O.; Nakazawa, T.; Kurosawa, T.; Hao, Y.; Isaji, T.; et al. a1,6-Fucosyltransferase-deficient mice exhibit multiple behavioral abnormalities associated with a schizophrenia-like phenotype: Importance of the balance between the dopamine and serotonin systems. J. Biol. Chem. 2011, 286, 18434–18443. [Google Scholar] [CrossRef]
- Keeley, T.S.; Yang, S.; Lau, E. The Diverse Contributions of Fucose Linkages in Cancer. Cancers 2019, 11, 1241. [Google Scholar] [CrossRef]
- Li, S.; Liu, X.Y.; Pan, Q.; Wu, J.; Liu, Z.H.; Wang, Y.; Liu, M.; Zhang, X.L. Hepatitis C Virus-Induced FUT8 Causes 5-FU Drug Resistance in Human Hepatoma Huh7.5.1 Cells. Viruses 2019, 11, 378. [Google Scholar] [CrossRef]
- Tu, C.F.; Wu, M.Y.; Lin, Y.C.; Kannagi, R.; Yang, R.B. FUT8 promotes breast cancer cell invasiveness by remodeling TGF-β receptor core fucosylation. Breast Cancer Rev. 2017, 19, 111. [Google Scholar] [CrossRef]
- Li, F.; Zhao, S.; Cui, Y.; Guo, T.; Qiang, J.; Xie, Q.; Yu, W.; Guo, W.; Deng, W.; Gu, C.; et al. a1,6-Fucosyltransferase (FUT8) regulates the cancer-promoting capacity of cancer-associated fibroblasts (CAFs) by modifying EGFR core fucosylation (CF) in non-small cell lung cancer (NSCLC). Am. J. Cancer Res. 2020, 10, 816–837. [Google Scholar] [PubMed]
- Sonneveld, M.E.; van der Schoot, C.E.; Vidarsson, G. The Elements Steering Pathogenesis in IgG-Mediated Alloimmune Diseases. J. Clin. Immunol. 2016, 36, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Pham, D.; Vincentz, J.W.; Firulli, A.B.; Kaplan, M.H. Twist1 regulates Ifng expression in Th1 cells by interfering with Runx3 function. J. Immunol. 2012, 189, 832–840. [Google Scholar] [CrossRef] [PubMed]
- Shinzaki, S.; Iijima, H.; Fujii, H.; Kamada, Y.; Naka, T.; Takehara, T.; Miyoshi, E. A novel pathogenesis of inflammatory bowel disease from the perspective of glyco-immunology. J. Biochem. 2017, 161, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Fermín, S.D.M.; Isabel, R.C.; Cristina, M.; Olga, M.A. Intestinal inflammation and mucosal barrier function. Inflamm. Bowel Dis. 2014, 20, 2394–2404. [Google Scholar]
- Moonens, K.; Bouckaert, J.; Coddens, A.; Tran, T.; Panjikar, S.; De Kerpel, M.; Cox, E.; Remaut, H.; De Greve, H. Structural insight in histo-blood group binding by the F18 fimbrial adhesin FedF. Mol. Microbiol. 2012, 86, 82–95. [Google Scholar] [CrossRef]
- Theodoratou, E.; Campbell, H.; Ventham, N.T.; Kolarich, D.; Pučić-Baković, M.; Zoldoš, V.; Fernandes, D.; Pemberton, I.K.; Rudan, I.; Kennedy, N.A. The role of glycosylation in IBD. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 588–600. [Google Scholar] [CrossRef]
- Schadendorf, D.; Hodi, F.S.; Robert, C.; Weber, J.S.; Margolin, K.; Hamid, O.; Patt, D.; Chen, T.T.; Berman, D.M.; Wolchok, J.D. Pooled Analysis of Long-Term Survival Data from Phase II and Phase III Trials of Ipilimumab in Unresectable or Metastatic Melanoma. J. Clin. Oncol. 2015, 33, 1889–1894. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.; Latchman, Y.; Buhlmann, J.; Tomczak, M.; Horwitz, B.; Freeman, G.; Sharpe, A. Regulation of PD-1, PD-L1, and PD-L2 expression during normal and autoimmune responses. Eur. J. Immunol. 2010, 33, 2706–2716. [Google Scholar] [CrossRef] [PubMed]
- Kashiwazaki, H.; Kakizaki, M.; Ikehara, Y.; Togayachi, A.; Narimatsu, H.; Watanabe, R. Mice lacking α1,3-fucosyltransferase 9 exhibit modulation of in vivo immune responses against pathogens. Pathol. Int. 2014, 64, 199–208. [Google Scholar] [CrossRef]
- Cai, Y.J.; Zheng, X.F.; Lu, C.H.; Jiang, Q.; Liu, Q.; Xin, Y.H. Effect of FUT3 gene silencing with miRNA on proliferation, invasion and migration abilities of human KATO-III gastric cancer cell line. Cell Mol. Biol. 2016, 62, 15–20. [Google Scholar]
- Orntoft, T.F.; Vestergaard, E.M.; Holmes, E.; Jakobsen, J.S.; Grunnet, N.; Mortensen, M.; Johnson, P.; Bross, P.; Gregersen, N.; Skorstengaard, K. Influence of Lewis α1-3/4-L-Fucosyltransferase (FUT3) Gene Mutations on Enzyme Activity, Erythrocyte Phenotyping, and Circulating Tumor Marker Sialyl-Lewis a Levels. J. Biol. Chem. 1996, 271, 32260–32268. [Google Scholar] [CrossRef] [PubMed]
- Faghfuri, E.; Faramarzi, M.A.; Nikfar, S.; Abdollahi, M. Nivolumab and pembrolizumab as immune-modulating monoclonal antibodies targeting the PD-1 receptor to treat melanoma. Expert. Rev. Anticancer Ther. 2015, 15, 981–993. [Google Scholar] [CrossRef] [PubMed]
- Munkley, J.; Mills, I.G.; Elliott, D.J. The role of glycans in the development and progression of prostate cancer. Nat. Rev. Urol. 2016, 13, 324–333. [Google Scholar] [CrossRef]
- Chen, J.Q.; Szodoray, P.; Zeher, M. Toll-Like Receptor Pathways in Autoimmune Diseases. Clin. Rev. Allergy. Immunol. 2016, 50, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Sanjo, H.; Takeda, K.; Ninomiya-Tsuji, J.; Yamamoto, M.; Kawai, T.; Matsumoto, K.; Takeuchi, O.; Akira, S. Essential function for the kinase TAK1 in innate and adaptive immune responses. Nat. Immunol. 2005, 6, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- Kravchenko, V.V.; Kaufmann, G.F. Bacterial inhibition of inflammatory responses via TLR-independent mechanisms. Cell. Microbiol. 2013, 15, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Yang, L.; Jin, J.; Wang, H.; Wu, S.L.; Bao, W.B. Regulation and Molecular Mechanism of TLR5 on Resistance to Escherichia coli F18 in Weaned Piglets. Animals 2019, 9, 735. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J. IPC: Professional type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annu. Rev. Immunol. 2005, 23, 275–306. [Google Scholar] [CrossRef] [PubMed]
- Umek, R.M.; Friedman, A.D.; McKnight, S.L. CCAAT-enhancer binding protein: A component of a differentiation switch. Science 1991, 251, 288–292. [Google Scholar] [CrossRef]
- Rosenbauer, F.; Tenen, D.G. Transcription factors in myeloid development: Balancing differentiation with transformation. Nat. Rev. Immunol. 2007, 7, 105–117. [Google Scholar] [CrossRef]
- Zhang, P.; Iwasaki-Arai, J.; Iwasaki, H.; Fenyus, M.L.; Dayaram, T.; Owens, B.M.; Shigematsu, H.; Levantini, E.; Huettner, C.S.; Lekstrom-Himes, J.A.; et al. Enhancement of hematopoietic stem cell repopulating capacity and self-renewal in the absence of the transcription factor C/EBP alpha. Immunity 2004, 21, 853–863. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, A.; Sarker, D.; Reebye, V.; Jarvis, S.; Sodergren, M.H.; Kossenkov, A.; Sanseviero, E.; Raulf, N.; Vasara, J.; Andrikakou, P. Upregulation of C/EBPα Inhibits Suppressive Activity of Myeloid Cells and Potentiates Antitumor Response in Mice and Patients with Cancer. Clin. Cancer. Res. 2021, 27, 5961–5978. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.F.; Cleaves, R.; Kummalue, T.; Nerlov, C.; Friedman, A.D. Cell cycle inhibition mediated by the outer surface of the C/EBPalpha basic region is required but not sufficient for granulopoiesis. Oncogene 2003, 22, 2548–2557. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wu, Z.C.; Dong, W.H.; Liu, Y.; Yang, J.S.; Zhu, G.Q.; Wu, S.L.; Bao, W.B. Attack Experiment and Phenotype Analysis of Meishan piglets by E. coli F18 Strain. Chin. J. Anim. Vet. Sci. 2014, 45, 1608–1615. (In Chinese) [Google Scholar]
- Liu, L.; Wang, J.; Zhao, Q.; Zi, C.; Wu, Z.; Su, X.; Huo, Y.; Zhu, G.Q.; Wu, S.L.; Bao, W.B. Genetic variation in exon 10 of the BPI gene is associated with Escherichia coli F18 susceptibility in Sutai piglets. Gene 2013, 523, 70–75. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Gan, L.; Qin, W.; Zi, C.; Zhu, G.Q.; Wu, S.L.; Bao, W.B. Use of fluorescence quantitative polymerase chain reaction (PCR) for the detection of Escherichia coli adhesion to pig intestinal epithelial cells. Pol. J. Vet. Sci. 2016, 19, 619–625. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, L.; Wang, Y.; Wu, S.; Wu, Z.; Bao, W. Effect and Mechanism Analysis of Pig FUT8 Gene on Resistance to Escherichia coli F18 Infection. Int. J. Mol. Sci. 2022, 23, 14713. https://doi.org/10.3390/ijms232314713
Wu L, Wang Y, Wu S, Wu Z, Bao W. Effect and Mechanism Analysis of Pig FUT8 Gene on Resistance to Escherichia coli F18 Infection. International Journal of Molecular Sciences. 2022; 23(23):14713. https://doi.org/10.3390/ijms232314713
Chicago/Turabian StyleWu, Lisi, Yifu Wang, Shenglong Wu, Zhengchang Wu, and Wenbin Bao. 2022. "Effect and Mechanism Analysis of Pig FUT8 Gene on Resistance to Escherichia coli F18 Infection" International Journal of Molecular Sciences 23, no. 23: 14713. https://doi.org/10.3390/ijms232314713
APA StyleWu, L., Wang, Y., Wu, S., Wu, Z., & Bao, W. (2022). Effect and Mechanism Analysis of Pig FUT8 Gene on Resistance to Escherichia coli F18 Infection. International Journal of Molecular Sciences, 23(23), 14713. https://doi.org/10.3390/ijms232314713






