Hybrid Pharmacophore- and Structure-Based Virtual Screening Pipeline to Identify Novel EGFR Inhibitors That Suppress Non-Small Cell Lung Cancer Cell Growth
Abstract
:1. Introduction
2. Results
2.1. Workflow
2.2. 3D QSAR Pharmacophore Generation and Validation
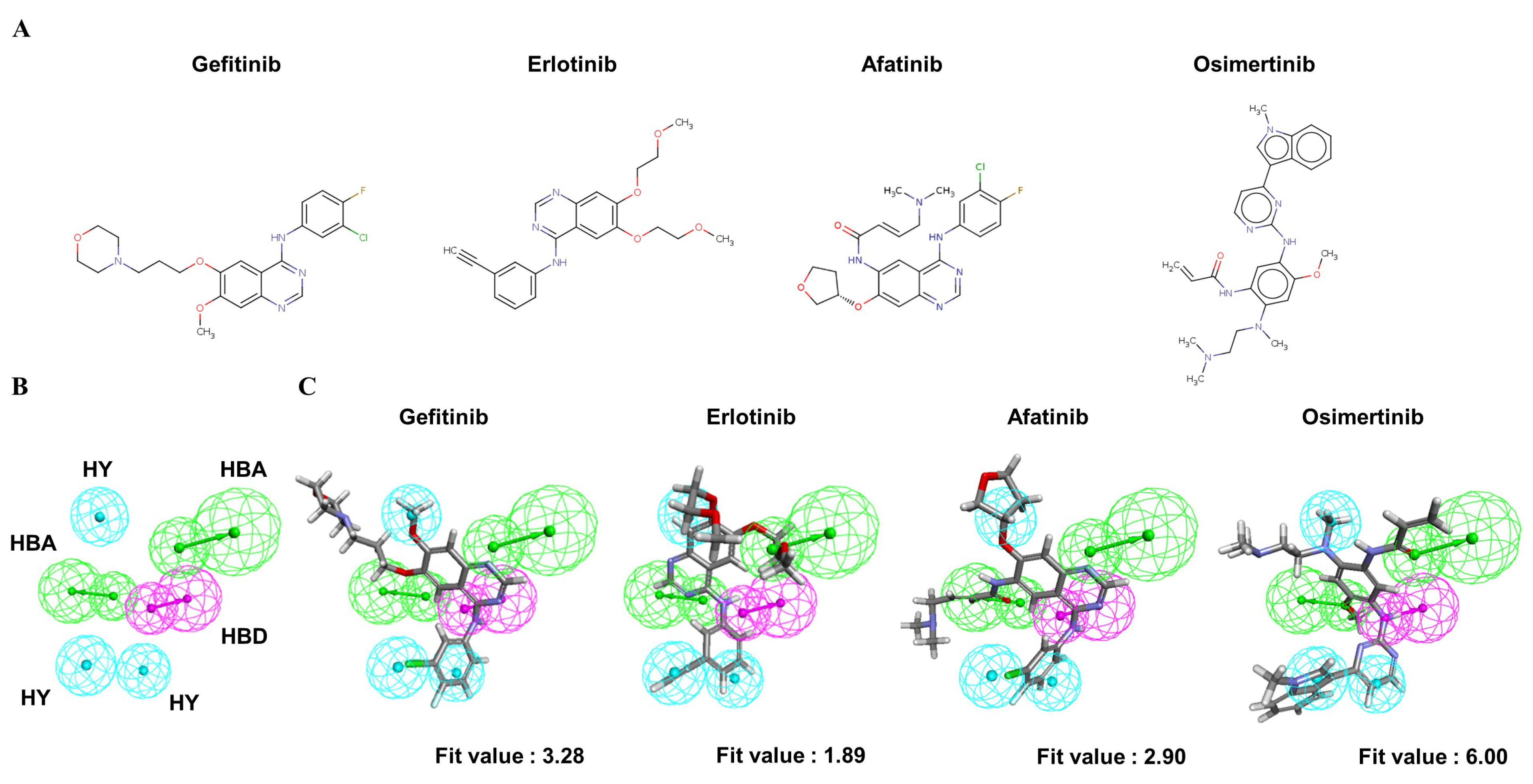
2.3. Identification of Common Feature Pharmacophores
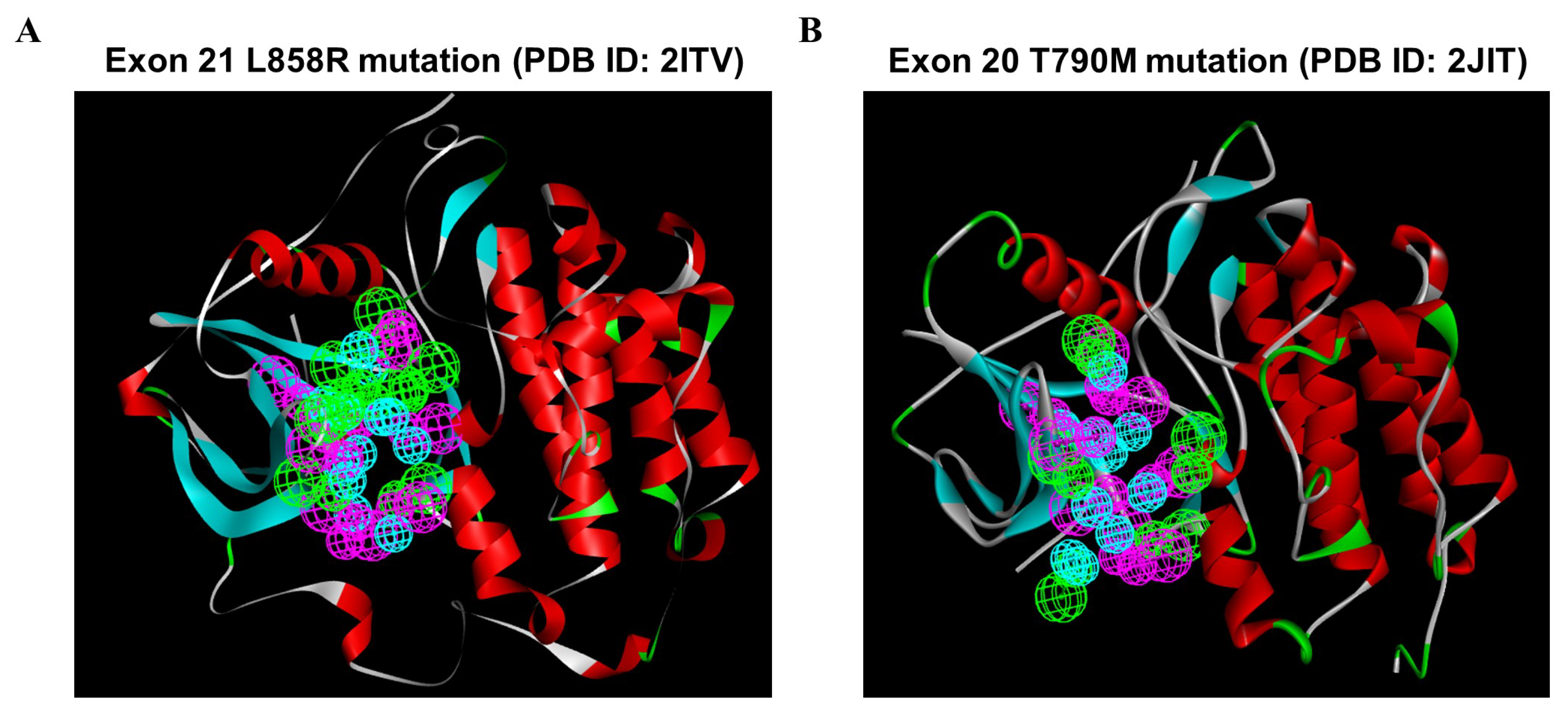
2.4. Generation of Structure-Based Pharmacophores
2.5. Identification of Candidate Compounds through Multiple-Stage PBVS
2.6. NSC609077 Inhibits the Anchorage-Dependent Growth and Motility of Lung Cancer Cells
2.7. Effects of NSC609077 on an EGFR-Associated Signaling Pathway
3. Discussion
4. Materials and Methods
4.1. 3D QSAR Pharmacophore Generation and Evaluation
4.2. Common Feature Pharmacophore Modeling
4.3. Structure-Based Pharmacophore Modeling
4.4. Pharmacophore-Based Virtual Screening (PBVS)
4.5. Molecular Docking Studies
4.6. Cell Culture and Drug Treatment
4.7. Enzyme-Linked Immunosorbent Assay (ELISA)
4.8. Cell Viability Assay
4.9. Clonogenicity Assay
4.10. Scratch-Wound Assay
4.11. Western Blot Analysis
4.12. Statistical Analysis
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A


| EGFR-TKI | 3D-QSAR Fit Value | CFP Fit Value | SBP Fit Value | LibDock Score | ||
|---|---|---|---|---|---|---|
| EGFR p.L858R (PDB: 2ITV) | EGFR p.T790M (PDB: 2JIT) | EGFR p.L858R (PDB: 2ITV) | EGFR p.T790M (PDB: 2JIT) | |||
| Erlotinib | 3.17 | 1.89 | 1.50 | 1.15 | 116.60 | 110.71 |
| Gefitinib | 3.92 | 3.28 | 2.49 | 2.29 | 113.73 | 106.28 |
| Afatinib | 5.89 | 2.90 | 2.32 | 2.49 | 111.59 | 107.47 |
| Osimertinib | 4.86 | 6.00 | 2.99 | 3.34 | 115.66 | 115.27 |
References
- Hubbard, S.R.; Miller, W.T. Receptor tyrosine kinases: Mechanisms of activation and signaling. Curr. Opin. Cell Biol. 2007, 19, 117–123. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lemmon, M.A.; Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 2010, 141, 1117–1134. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paul, M.K.; Mukhopadhyay, A.K. Tyrosine kinase—Role and significance in Cancer. Int. J. Med. Sci. 2004, 1, 101–115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takeuchi, K.; Ito, F. Receptor tyrosine kinases and targeted cancer therapeutics. Biol. Pharm. Bull. 2011, 34, 1774–1780. [Google Scholar] [CrossRef] [Green Version]
- Yarden, Y.; Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2001, 2, 127–137. [Google Scholar] [CrossRef]
- Lurje, G.; Lenz, H.J. EGFR signaling and drug discovery. Oncology 2009, 77, 400–410. [Google Scholar] [CrossRef]
- Ferguson, K.M.; Berger, M.B.; Mendrola, J.M.; Cho, H.S.; Leahy, D.J.; Lemmon, M.A. EGF activates its receptor by removing interactions that autoinhibit ectodomain dimerization. Mol. Cell 2003, 11, 507–517. [Google Scholar] [CrossRef]
- Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 2000, 103, 211–225. [Google Scholar] [CrossRef] [Green Version]
- Wieduwilt, M.J.; Moasser, M.M. The epidermal growth factor receptor family: Biology driving targeted therapeutics. Cell. Mol. Life Sci. 2008, 65, 1566–1584. [Google Scholar] [CrossRef] [Green Version]
- Wee, P.; Wang, Z. Epidermal Growth Factor Receptor Cell Proliferation Signaling Pathways. Cancers 2017, 9, 52. [Google Scholar] [CrossRef] [Green Version]
- Sunpaweravong, P.; Sunpaweravong, S.; Puttawibul, P.; Mitarnun, W.; Zeng, C.; Baron, A.E.; Franklin, W.; Said, S.; Varella-Garcia, M. Epidermal growth factor receptor and cyclin D1 are independently amplified and overexpressed in esophageal squamous cell carcinoma. J. Cancer Res. Clin. Oncol. 2005, 131, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Ohgaki, H.; Dessen, P.; Jourde, B.; Horstmann, S.; Nishikawa, T.; Di Patre, P.L.; Burkhard, C.; Schuler, D.; Probst-Hensch, N.M.; Maiorka, P.C.; et al. Genetic pathways to glioblastoma: A population-based study. Cancer Res. 2004, 64, 6892–6899. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, T.; Zhang, G.; Haura, E.B. Targeting epidermal growth factor receptor: Central signaling kinase in lung cancer. Biochem. Pharmacol. 2010, 80, 613–623. [Google Scholar] [CrossRef] [PubMed]
- Minder, P.; Zajac, E.; Quigley, J.P.; Deryugina, E.I. EGFR regulates the development and microarchitecture of intratumoral angiogenic vasculature capable of sustaining cancer cell intravasation. Neoplasia 2015, 17, 634–649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keller, S.; Schmidt, M.H.H. EGFR and EGFRvIII Promote Angiogenesis and Cell Invasion in Glioblastoma: Combination Therapies for an Effective Treatment. Int. J. Mol. Sci. 2017, 18, 1295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Normanno, N.; De Luca, A.; Bianco, C.; Strizzi, L.; Mancino, M.; Maiello, M.R.; Carotenuto, A.; De Feo, G.; Caponigro, F.; Salomon, D.S. Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 2006, 366, 2–16. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 7–34. [Google Scholar] [CrossRef] [Green Version]
- Molina, J.R.; Yang, P.; Cassivi, S.D.; Schild, S.E.; Adjei, A.A. Non-small cell lung cancer: Epidemiology, risk factors, treatment, and survivorship. Mayo Clin. Proc. 2008, 83, 584–594. [Google Scholar] [CrossRef]
- Barker, A.J.; Gibson, K.H.; Grundy, W.; Godfrey, A.A.; Barlow, J.J.; Healy, M.P.; Woodburn, J.R.; Ashton, S.E.; Curry, B.J.; Scarlett, L.; et al. Studies leading to the identification of ZD1839 (IRESSA): An orally active, selective epidermal growth factor receptor tyrosine kinase inhibitor targeted to the treatment of cancer. Bioorg. Med. Chem. Lett. 2001, 11, 1911–1914. [Google Scholar] [CrossRef]
- Bonomi, P. Erlotinib: A new therapeutic approach for non-small cell lung cancer. Expert Opin. Investig. Drugs 2003, 12, 1395–1401. [Google Scholar] [CrossRef]
- Kumar, S.; Agrawal, R. Next generation tyrosine kinase inhibitor (TKI): Afatinib. Recent Pat. Anti-Cancer Drug Discov. 2014, 9, 382–393. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Ambrogio, L.; Shimamura, T.; Kubo, S.; Takahashi, M.; Chirieac, L.R.; Padera, R.F.; Shapiro, G.I.; Baum, A.; Himmelsbach, F.; et al. BIBW2992, an irreversible EGFR/HER2 inhibitor highly effective in preclinical lung cancer models. Oncogene 2008, 27, 4702–4711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cross, D.A.; Ashton, S.E.; Ghiorghiu, S.; Eberlein, C.; Nebhan, C.A.; Spitzler, P.J.; Orme, J.P.; Finlay, M.R.; Ward, R.A.; Mellor, M.J.; et al. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 2014, 4, 1046–1061. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paez, J.G.; Janne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; et al. EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science 2004, 304, 1497–1500. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pao, W.; Girard, N. New driver mutations in non-small-cell lung cancer. Lancet Oncol. 2011, 12, 175–180. [Google Scholar] [CrossRef]
- Riely, G.J.; Pao, W.; Pham, D.; Li, A.R.; Rizvi, N.; Venkatraman, E.S.; Zakowski, M.F.; Kris, M.G.; Ladanyi, M.; Miller, V.A. Clinical course of patients with non-small cell lung cancer and epidermal growth factor receptor exon 19 and exon 21 mutations treated with gefitinib or erlotinib. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2006, 12, 839–844. [Google Scholar] [CrossRef] [Green Version]
- Gazdar, A.F. Activating and resistance mutations of EGFR in non-small-cell lung cancer: Role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 2009, 28 (Suppl. S1), S24–S31. [Google Scholar] [CrossRef] [Green Version]
- Park, K.; Tan, E.H.; O’Byrne, K.; Zhang, L.; Boyer, M.; Mok, T.; Hirsh, V.; Yang, J.C.; Lee, K.H.; Lu, S.; et al. Afatinib versus gefitinib as first-line treatment of patients with EGFR mutation-positive non-small-cell lung cancer (LUX-Lung 7): A phase 2B, open-label, randomised controlled trial. Lancet Oncol. 2016, 17, 577–589. [Google Scholar] [CrossRef]
- Ancevski Hunter, K.; Friedland, D.M.; Villaruz, L.C.; Burns, T.F. First-Line Osimertinib in Patients with Treatment-Naive Somatic or Germline EGFR T790M-Mutant Metastatic NSCLC. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2018, 13, e3–e5. [Google Scholar] [CrossRef] [Green Version]
- Van Der Steen, N.; Giovannetti, E.; Carbone, D.; Leonetti, A.; Rolfo, C.D.; Peters, G.J. Resistance to epidermal growth factor receptor inhibition in non-small cell lung cancer. Cancer Drug Resist. 2018, 1, 230–249. [Google Scholar] [CrossRef] [Green Version]
- Wang, S.; Tsui, S.T.; Liu, C.; Song, Y.; Liu, D. EGFR C797S mutation mediates resistance to third-generation inhibitors in T790M-positive non-small cell lung cancer. J. Hematol. Oncol. 2016, 9, 59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, D.; Hu, M.; Bai, Y.; Zhu, X.; Lu, X.; Wu, C.; Wang, J.; Liu, L.; Wang, Z.; Ni, J.; et al. EGFR G796D mutation mediates resistance to osimertinib. Oncotarget 2017, 8, 49671–49679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brooks, B.R.; Bruccoleri, R.E.; Olafson, B.D.; States, D.J.; Swaminathan, S.; Karplus, M. CHARMM: A program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 1983, 4, 187–217. [Google Scholar] [CrossRef]
- Gilson, M.K.; Liu, T.; Baitaluk, M.; Nicola, G.; Hwang, L.; Chong, J. BindingDB in 2015: A public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res. 2016, 44, D1045–D1053. [Google Scholar] [CrossRef]
- National Cancer Institute. DTP Data—Chemical Data. Available online: https://wiki.nci.nih.gov/display/NCIDTPdata/Chemical+Data (accessed on 1 September 2016).
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Veber, D.F.; Johnson, S.R.; Cheng, H.Y.; Smith, B.R.; Ward, K.W.; Kopple, K.D. Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 2002, 45, 2615–2623. [Google Scholar] [CrossRef]
- Li, H.; Schmid-Bindert, G.; Wang, D.; Zhao, Y.; Yang, X.; Su, B.; Zhou, C. Blocking the PI3K/AKT and MEK/ERK signaling pathways can overcome gefitinib-resistance in non-small cell lung cancer cell lines. Adv. Med. Sci. 2011, 56, 275–284. [Google Scholar] [CrossRef]
- Wang, B.; Jiang, H.; Wang, L.; Chen, X.; Wu, K.; Zhang, S.; Ma, S.; Xia, B. Increased MIR31HG lncRNA expression increases gefitinib resistance in non-small cell lung cancer cell lines through the EGFR/PI3K/AKT signaling pathway. Oncol. Lett. 2017, 13, 3494–3500. [Google Scholar] [CrossRef] [Green Version]
- Deng, Q.F.; Fang, Q.Y.; Ji, X.X.; Zhou, S.W. Cyclooxygenase-2 mediates gefitinib resistance in non-small cell lung cancer through the EGFR/PI3K/AKT axis. J. Cancer 2020, 11, 3667–3674. [Google Scholar] [CrossRef]
- Song, L.; Morris, M.; Bagui, T.; Lee, F.Y.; Jove, R.; Haura, E.B. Dasatinib (BMS-354825) selectively induces apoptosis in lung cancer cells dependent on epidermal growth factor receptor signaling for survival. Cancer Res. 2006, 66, 5542–5548. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Sutter, J.; Hoffmann, R. HypoGen: An automated system for generating 3D predictive pharmacophore models. In Pharmacophore Perception, Development and Use in Drug Design; Güner, O.F., Ed.; International University Line: La Jolla, CA, USA, 2000; pp. 171–189. [Google Scholar]
- Sakkiah, S.; Arullaperumal, V.; Hwang, S.; Lee, K.W. Ligand-based pharmacophore modeling and Bayesian approaches to identify c-Src inhibitors. J. Enzym. Inhib. Med. Chem. 2014, 29, 69–80. [Google Scholar] [CrossRef] [PubMed]
- Bommu, U.D.; Konidala, K.K.; Pamanji, R.; Yeguvapalli, S. Computational screening, ensemble docking and pharmacophore analysis of potential gefitinib analogues against epidermal growth factor receptor. J. Recept. Signal Transduct. Res. 2018, 38, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Xiang, M.; Lei, K.; Fan, W.; Lin, Y.; He, G.; Yang, M.; Chen, L.; Mo, Y. In silico identification of EGFR-T790M inhibitors with novel scaffolds: Start with extraction of common features. Drug Des. Dev. Ther. 2013, 7, 789–839. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scholz, S.W.; Mhyre, T.; Ressom, H.; Shah, S.; Federoff, H.J. Genomics and bioinformatics of Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2012, 2, a009449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, R.-Z.; Fan, X.-X.; Duan, F.-G.; Jiang, Z.-B.; Pan, H.-D.; Luo, L.-X.; Zhou, Y.-L.; Li, Y.; Yao, Y.-J.; Yao, X.-J.; et al. Proscillaridin A induces apoptosis and suppresses non-small-cell lung cancer tumor growth via calcium-induced DR4 upregulation. Cell Death Dis. 2018, 9, 696. [Google Scholar] [CrossRef]
- Engelman, J.A. Targeting PI3K signalling in cancer: Opportunities, challenges and limitations. Nat. Rev. Cancer 2009, 9, 550–562. [Google Scholar] [CrossRef]
- Ogino, A.; Kitao, H.; Hirano, S.; Uchida, A.; Ishiai, M.; Kozuki, T.; Takigawa, N.; Takata, M.; Kiura, K.; Tanimoto, M. Emergence of epidermal growth factor receptor T790M mutation during chronic exposure to gefitinib in a non small cell lung cancer cell line. Cancer Res. 2007, 67, 7807–7814. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Chang, A.Y.-C. Molecular mechanism of action and potential biomarkers of growth inhibition of synergistic combination of afatinib and dasatinib against gefitinib-resistant non-small cell lung cancer cells. Oncotarget 2018, 9, 16533–16546. [Google Scholar] [CrossRef] [Green Version]
- Sheng, Y.; Li, W.; Zhu, F.; Liu, K.; Chen, H.; Yao, K.; Reddy, K.; Lim, D.Y.; Oi, N.; Li, H.; et al. 3,6,2′,4′,5′-Pentahydroxyflavone, an orally bioavailable multiple protein kinase inhibitor, overcomes gefitinib resistance in non-small cell lung cancer. J. Biol. Chem. 2014, 289, 28192–28201. [Google Scholar] [CrossRef] [Green Version]
- Faber, A.C.; Li, D.; Song, Y.; Liang, M.C.; Yeap, B.Y.; Bronson, R.T.; Lifshits, E.; Chen, Z.; Maira, S.M.; Garcia-Echeverria, C.; et al. Differential induction of apoptosis in HER2 and EGFR addicted cancers following PI3K inhibition. Proc. Natl. Acad. Sci. USA 2009, 106, 19503–19508. [Google Scholar] [CrossRef] [Green Version]
- Barnum, D.; Greene, J.; Smellie, A.; Sprague, P. Identification of common functional configurations among molecules. J. Chem. Inf. Comput. Sci. 1996, 36, 563–571. [Google Scholar] [CrossRef] [PubMed]
- Yun, C.H.; Boggon, T.J.; Li, Y.; Woo, M.S.; Greulich, H.; Meyerson, M.; Eck, M.J. Structures of lung cancer-derived EGFR mutants and inhibitor complexes: Mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 2007, 11, 217–227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yun, C.H.; Mengwasser, K.E.; Toms, A.V.; Woo, M.S.; Greulich, H.; Wong, K.K.; Meyerson, M.; Eck, M.J. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. USA 2008, 105, 2070–2075. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bohm, H.J. The computer program LUDI: A new method for the de novo design of enzyme inhibitors. J. Comput.-Aided Mol. Des. 1992, 6, 61–78. [Google Scholar] [CrossRef]
- Bohm, H.J. LUDI: Rule-based automatic design of new substituents for enzyme inhibitor leads. J. Computer-Aided Mol. Des. 1992, 6, 593–606. [Google Scholar] [CrossRef]
- Rao, S.N.; Head, M.S.; Kulkarni, A.; LaLonde, J.M. Validation studies of the site-directed docking program LibDock. J. Chem. Inf. Model. 2007, 47, 2159–2171. [Google Scholar] [CrossRef] [Green Version]
- Rodriguez, L.G.; Wu, X.; Guan, J.L. Wound-healing assay. Methods Mol. Biol. 2005, 294, 23–29. [Google Scholar]
- Weng, C.W.; Li, J.H.; Tsai, J.Y.; Lin, S.H.; Chang, G.C.; Liu, C.C.; Chen, J.J. Pharmacophore-based virtual screening for the identification of the novel Src inhibitor SJG-136 against lung cancer cell growth and motility. Am. J. Cancer Res. 2020, 10, 1668–1690. [Google Scholar]




| Pharmacophore Hypothesis | Total Cost | Cost Difference | RMS | Correlation | Features |
|---|---|---|---|---|---|
| Hypo1 | 136.11 | 60.98 | 0.88 | 0.92 | HBA, HBD, HY |
| Hypo2 | 136.27 | 60.82 | 0.90 | 0.91 | HBA, HBD, HY |
| Hypo3 | 144.36 | 52.73 | 1.14 | 0.87 | HBA, HBD, HY |
| Hypo4 | 146.30 | 50.79 | 1.19 | 0.85 | HBA, HBD, HY |
| Hypo5 | 147.97 | 49.12 | 1.21 | 0.85 | HBD, HY, RA |
| Hypo6 | 150.20 | 46.89 | 1.28 | 0.83 | HBA, HBD, HY |
| Hypo7 | 150.36 | 46.73 | 1.25 | 0.84 | HBD, HY, RA |
| Hypo8 | 150.65 | 46.44 | 1.30 | 0.82 | HBA, HBD, HY |
| Hypo9 | 150.97 | 46.12 | 1.30 | 0.82 | HBD, HY, RA |
| Hypo10 | 151.64 | 45.45 | 1.27 | 0.84 | HBD, HY, RA |
| BindingDB Code | Experimental IC50 nM | Predicted IC50 nM | Error | Experimental Scale ‡ | Predicted Scale ‡ | 3D-QSAR Fit Value |
|---|---|---|---|---|---|---|
| BDBM3297 | 0.1 | 0.4 | 4.00 | +++ | +++ | 6.79 |
| BDBM3520 | 0.3 | 0.4 | 1.33 | +++ | +++ | 6.77 |
| BDBM3522 | 0.4 | 0.4 | 1.00 | +++ | +++ | 6.80 |
| BDBM3583 | 0.4 | 0.7 | 1.75 | +++ | +++ | 6.58 |
| BDBM3294 | 0.8 | 2.2 | 2.75 | +++ | +++ | 6.07 |
| BDBM3519 | 2.0 | 9.3 | 4.65 | +++ | +++ | 5.45 |
| BDBM3780 | 2.7 | 3.5 | 1.30 | +++ | +++ | 5.87 |
| BDBM3546 | 3.8 | 2.4 | −1.58 | +++ | +++ | 6.04 |
| BDBM3535 | 4.0 | 2.0 | −2.00 | +++ | +++ | 6.12 |
| BDBM3538 | 4.7 | 0.4 | −11.75 | +++ | +++ | 6.79 |
| BDBM3767 | 9.4 | 20.4 | 2.17 | +++ | +++ | 5.11 |
| BDBM3781 | 20.0 | 66.0 | 3.30 | +++ | +++ | 4.60 |
| BDBM3263 | 23.0 | 75.1 | 3.27 | +++ | +++ | 4.54 |
| BDBM3264 | 27.0 | 72.8 | 2.70 | +++ | +++ | 4.55 |
| BDBM3753 | 31.0 | 55.5 | 1.79 | +++ | +++ | 4.67 |
| BDBM3508 | 55.0 | 73.2 | 1.33 | +++ | +++ | 4.55 |
| BDBM3503 | 56.0 | 164.8 | 2.94 | +++ | ++ | 4.20 |
| BDBM3283 | 58.0 | 81.6 | 1.41 | +++ | +++ | 4.50 |
| BDBM3763 | 58.0 | 209.8 | 3.62 | +++ | ++ | 4.09 |
| BDBM3744 | 72.0 | 63.6 | −1.13 | +++ | +++ | 4.61 |
| BDBM3265 | 80.0 | 72.0 | −1.11 | +++ | +++ | 4.56 |
| BDBM3536 | 84.0 | 70.5 | −1.19 | +++ | +++ | 4.57 |
| BDBM3518 | 100.0 | 16.8 | −5.95 | ++ | +++ | 5.19 |
| BDBM3516 | 120.0 | 79.8 | −1.50 | ++ | +++ | 4.51 |
| BDBM3745 | 132.0 | 85.6 | −1.54 | ++ | +++ | 4.48 |
| BDBM3292 | 139.0 | 90.7 | −1.53 | ++ | +++ | 4.46 |
| BDBM3757 | 147.0 | 57.4 | −2.56 | ++ | +++ | 4.66 |
| BDBM3752 | 264.0 | 188.8 | −1.40 | ++ | ++ | 4.14 |
| BDBM3256 | 320.0 | 106.8 | −3.00 | ++ | ++ | 4.39 |
| BDBM3259 | 344.0 | 202.1 | −1.70 | ++ | ++ | 4.11 |
| BDBM3295 | 348.0 | 69.2 | −5.03 | ++ | +++ | 4.58 |
| BDBM3510 | 770.0 | 228.5 | −3.37 | ++ | ++ | 4.06 |
| BDBM3269 | 5500.0 | 3769.7 | −1.46 | + | + | 2.84 |
| Pharmacophore Hypothesis | Rank | Direct Hit | Partial Hit | Max Hit | Features |
|---|---|---|---|---|---|
| Pharm1 | 54.25 | 1101 | 0010 | 6 | RRHHAA |
| Pharm2 | 53.04 | 1111 | 0000 | 6 | HHHDAA |
| Pharm3 | 52.64 | 1101 | 0010 | 6 | RRHHAA |
| Pharm4 | 52.48 | 1111 | 0000 | 5 | RRHAA |
| Pharm5 | 52.32 | 1101 | 0010 | 6 | RRHHAA |
| Pharm6 | 52.32 | 1101 | 0010 | 6 | RRHHAA |
| Pharm7 | 52.25 | 1101 | 0010 | 6 | RRHHAA |
| Pharm8 | 52.25 | 1101 | 0010 | 6 | RRHHAA |
| Pharm9 | 52.20 | 1111 | 0000 | 5 | RRHAA |
| Pharm10 | 52.20 | 1111 | 0000 | 5 | RRHAA |
| NSC Number | 3D-QSAR Fit Value | CFP Fit Value | SBP Fit Value | LibDock Score | Phosphorylation Level (%) | Inhibitory Rate (%) | ||
|---|---|---|---|---|---|---|---|---|
| EGFR p.L858R (PDB: 2ITV) | EGFR p.T790M (PDB: 2JIT) | EGFR p.L858R (PDB: 2ITV) | EGFR p.T790M (PDB: 2JIT) | |||||
| NSC7521 | 6.00 | 3.18 | 2.89 | 2.57 | 124.05 | 122.70 | 75 ± 15 | 25 ± 15 |
| NSC342715 | 6.47 | 3.26 | 2.98 | 2.65 | 123.30 | 119.98 | 37 ± 6 * | 63 ± 6 * |
| NSC622394 | 6.05 | 3.37 | 1.97 | 2.54 | 148.73 | 123.90 | 32 ± 8 * | 68 ± 8 * |
| NSC609077 | 6.22 | 2.79 | 2.77 | 1.68 | 145.57 | 152.88 | 21 ± 6 * | 79 ± 6 * |
| NSC622442 | 6.68 | 3.73 | 3.12 | 2.49 | 129.85 | 124.22 | NA | NA |
| NSC623897 | 6.03 | 3.77 | 2.48 | 3.33 | 117.06 | 116.17 | NA | NA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Weng, C.-W.; Wei, C.-H.; Tsai, J.-Y.; Lai, Y.-H.; Chang, G.-C.; Chen, J.J.W. Hybrid Pharmacophore- and Structure-Based Virtual Screening Pipeline to Identify Novel EGFR Inhibitors That Suppress Non-Small Cell Lung Cancer Cell Growth. Int. J. Mol. Sci. 2022, 23, 3487. https://doi.org/10.3390/ijms23073487
Weng C-W, Wei C-H, Tsai J-Y, Lai Y-H, Chang G-C, Chen JJW. Hybrid Pharmacophore- and Structure-Based Virtual Screening Pipeline to Identify Novel EGFR Inhibitors That Suppress Non-Small Cell Lung Cancer Cell Growth. International Journal of Molecular Sciences. 2022; 23(7):3487. https://doi.org/10.3390/ijms23073487
Chicago/Turabian StyleWeng, Chia-Wei, Chi-Hsuan Wei, Jeng-Yuan Tsai, Yi-Hua Lai, Gee-Chen Chang, and Jeremy J. W. Chen. 2022. "Hybrid Pharmacophore- and Structure-Based Virtual Screening Pipeline to Identify Novel EGFR Inhibitors That Suppress Non-Small Cell Lung Cancer Cell Growth" International Journal of Molecular Sciences 23, no. 7: 3487. https://doi.org/10.3390/ijms23073487






