Precise Regulation of the TAA1/TAR-YUCCA Auxin Biosynthesis Pathway in Plants
Abstract
:1. Introduction
2. Small Chemical Inhibitors Target TAA1/TARs and YUCCA to Modulate Auxin Synthesis
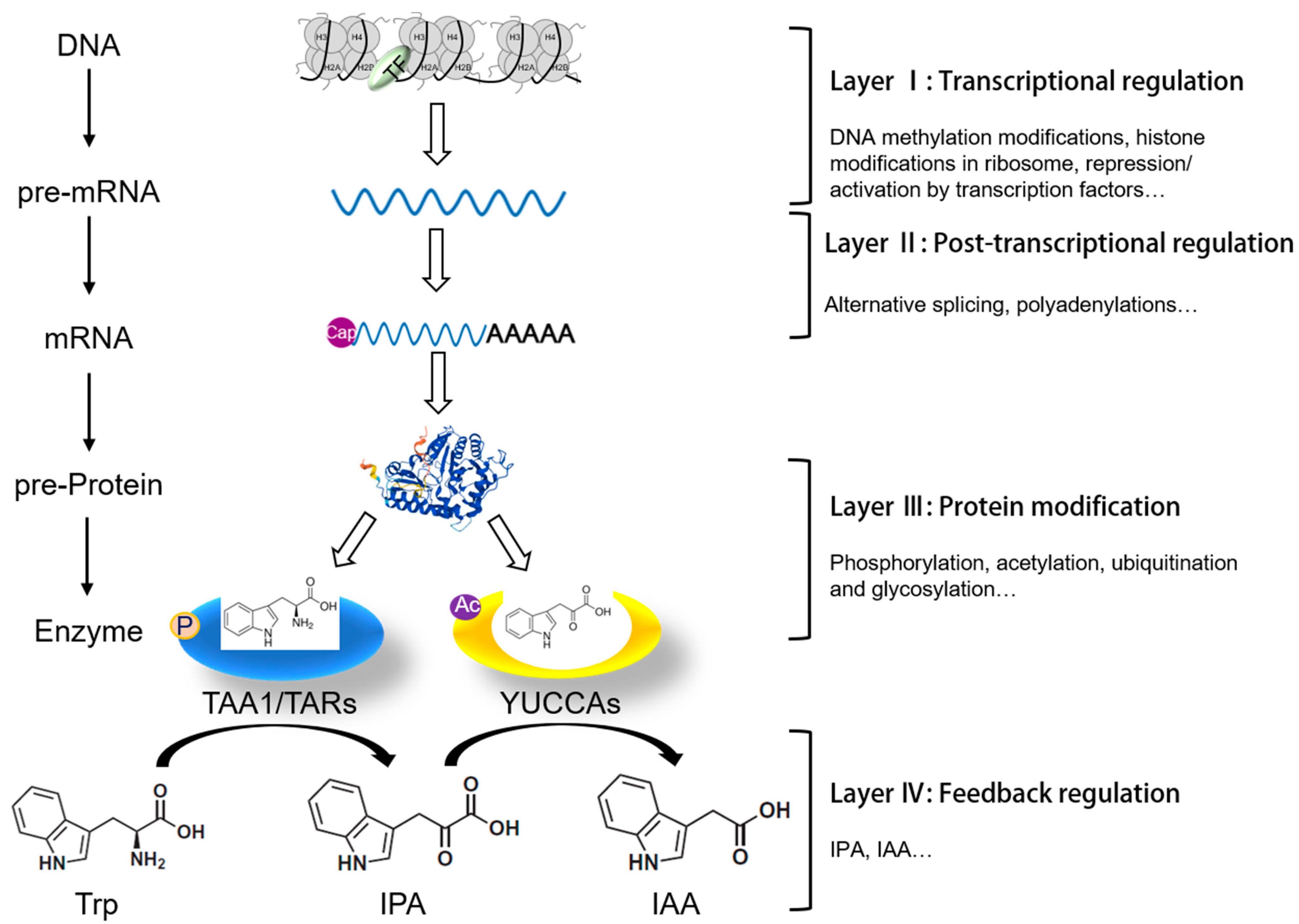
3. Layer Ⅰ: Finely Tuned Transcriptional Regulation of IPA-Dependent Auxin Biosynthesis
3.1. Epigenetic Modification of Genes Involved in IPA-Dependent Auxin Biosynthesis Pathway
3.2. Complex Transcriptional Regulatory Mechanisms of the TAA1/TAR and YUCCA Genes
3.2.1. Vegetative Stage
3.2.2. Reproductive Stage
4. Layer II: Post-Transcriptional Regulation of TAA1/TAR and YUC Genes in Plants
5. Layer III: Precise Control of IPA-Dependent Auxin Biosynthesis through Post-Translational Protein Modification
6. Layer IV: Negative Feedback Regulation of IPA Pathway
7. Concluding Remarks
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Blakeslee, J.J.; Spatola Rossi, T.; Kriechbaumer, V. Auxin biosynthesis: Spatial regulation and adaptation to stress. J. Exp. Bot. 2019, 70, 5041–5049. [Google Scholar] [CrossRef] [PubMed]
- Smolko, A.; Bauer, N.; Pavlovic, I.; Pencik, A.; Novak, O.; Salopek-Sondi, B. Altered Root Growth, Auxin Metabolism and Distribution in Arabidopsis thaliana Exposed to Salt and Osmotic Stress. Int. J. Mol. Sci. 2021, 22, 7993. [Google Scholar] [CrossRef]
- Tiwari, M.; Kumar, R.; Subramanian, S.; Doherty, C.J.; Jagadish, S.V.K. Auxin-cytokinin interplay shapes root functionality under low-temperature stress. Trends Plant Sci. 2023, 28, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Negi, N.P.; Pareek, S.; Mudgal, G.; Kumar, D. Auxin response factors in plant adaptation to drought and salinity stress. Physiol. Plant. 2022, 174, e13714. [Google Scholar] [CrossRef] [PubMed]
- Korasick, D.A.; Enders, T.A.; Strader, L.C. Auxin biosynthesis and storage forms. J. Exp. Bot. 2013, 64, 2541–2555. [Google Scholar] [CrossRef]
- Di, D.W.; Zhang, C.; Luo, O.; An, C.-W.; Guo, G.-Q. The biosynthesis of auxin: How many paths truly lead to IAA? Plant Growth Regul. 2016, 78, 275–285. [Google Scholar] [CrossRef]
- Morffy, N.; Strader, L.C. Old Town Roads: Routes of auxin biosynthesis across kingdoms. Curr. Opin. Plant Biol. 2020, 55, 21–27. [Google Scholar] [CrossRef]
- Tao, Y.; Ferrer, J.L.; Ljung, K.; Pojer, F.; Hong, F.; Long, J.A.; Li, L.; Moreno, J.E.; Bowman, M.E.; Ivans, L.J.; et al. Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 2008, 133, 164–176. [Google Scholar] [CrossRef]
- Stepanova, A.N.; Robertson-Hoyt, J.; Yun, J.; Benavente, L.M.; Xie, D.Y.; Dolezal, K.; Schlereth, A.; Jurgens, G.; Alonso, J.M. TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 2008, 133, 177–191. [Google Scholar] [CrossRef]
- Yamada, M.; Greenham, K.; Prigge, M.J.; Jensen, P.J.; Estelle, M. The TRANSPORT INHIBITOR RESPONSE2 gene is required for auxin synthesis and diverse aspects of plant development. Plant Physiol. 2009, 151, 168–179. [Google Scholar] [CrossRef]
- Zhou, Z.Y.; Zhang, C.G.; Wu, L.; Zhang, C.G.; Chai, J.; Wang, M.; Jha, A.; Jia, P.F.; Cui, S.J.; Yang, M.; et al. Functional characterization of the CKRC1/TAA1 gene and dissection of hormonal actions in the Arabidopsis root. Plant J. 2011, 66, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.; Jang, S.; Yoon, E.K.; Lee, S.A.; Dhar, S.; Kim, J.; Lee, M.M.; Lim, J. Involvement of Pyridoxine/Pyridoxamine 5′-Phosphate Oxidase (PDX3) in Ethylene-Induced Auxin Biosynthesis in the Arabidopsis Root. Mol. Cells 2018, 41, 1033–1044. [Google Scholar] [CrossRef] [PubMed]
- Sato, A.; Soeno, K.; Kikuchi, R.; Narukawa-Nara, M.; Yamazaki, C.; Kakei, Y.; Nakamura, A.; Shimada, Y. Indole-3-pyruvic acid regulates TAA1 activity, which plays a key role in coordinating the two steps of auxin biosynthesis. Proc. Natl. Acad. Sci. USA 2022, 119, e2203633119. [Google Scholar] [CrossRef] [PubMed]
- Mashiguchi, K.; Tanaka, K.; Sakai, T.; Sugawara, S.; Kawaide, H.; Natsume, M.; Hanada, A.; Yaeno, T.; Shirasu, K.; Yao, H.; et al. The main auxin biosynthesis pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 18512–18517. [Google Scholar] [CrossRef] [PubMed]
- Stepanova, A.N.; Yun, J.; Robles, L.M.; Novak, O.; He, W.; Guo, H.; Ljung, K.; Alonso, J.M. The Arabidopsis YUCCA1 flavin monooxygenase functions in the indole-3-pyruvic acid branch of auxin biosynthesis. Plant Cell 2011, 23, 3961–3973. [Google Scholar] [CrossRef]
- Zhao, Y.; Christensen, S.K.; Fankhauser, C.; Cashman, J.R.; Cohen, J.D.; Weigel, D.; Chory, J. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 2001, 291, 306–309. [Google Scholar] [CrossRef]
- Di, D.W.; Wu, L.; Zhang, L.; An, C.W.; Zhang, T.Z.; Luo, P.; Gao, H.H.; Kriechbaumer, V.; Guo, G.Q. Functional roles of Arabidopsis CKRC2/YUCCA8 gene and the involvement of PIF4 in the regulation of auxin biosynthesis by cytokinin. Sci. Rep. 2016, 6, 36866. [Google Scholar] [CrossRef]
- Dai, X.; Mashiguchi, K.; Chen, Q.; Kasahara, H.; Kamiya, Y.; Ojha, S.; DuBois, J.; Ballou, D.; Zhao, Y. The biochemical mechanism of auxin biosynthesis by an arabidopsis YUCCA flavin-containing monooxygenase. J. Biol. Chem. 2013, 288, 1448–1457. [Google Scholar] [CrossRef]
- Cao, X.; Yang, H.; Shang, C.; Ma, S.; Liu, L.; Cheng, J. The Roles of Auxin Biosynthesis YUCCA Gene Family in Plants. Int. J. Mol. Sci. 2019, 20, 6343. [Google Scholar] [CrossRef]
- Barbazuk, W.B.; Fu, Y.; McGinnis, K.M. Genome-wide analyses of alternative splicing in plants: Opportunities and challenges. Genome Res. 2008, 18, 1381–1392. [Google Scholar] [CrossRef]
- Di, D.W.; Zhang, C.G.; Guo, G.Q. Involvement of secondary messengers and small organic molecules in auxin perception and signaling. Plant Cell Rep. 2015, 34, 895–904. [Google Scholar] [CrossRef] [PubMed]
- Gallego-Bartolome, J. DNA methylation in plants: Mechanisms and tools for targeted manipulation. New Phytol. 2020, 227, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Hong, L.W.; Ye, C.T.; Lin, J.C.; Fu, H.H.; Wu, X.H.; Li, Q.S.Q. Alternative polyadenylation is involved in auxin-based plant growth and development. Plant J. 2018, 93, 246–258. [Google Scholar] [CrossRef] [PubMed]
- Mateo-Bonmati, E.; Casanova-Saez, R.; Ljung, K. Epigenetic Regulation of Auxin Homeostasis. Biomolecules 2019, 9, 623. [Google Scholar] [CrossRef]
- Wang, J.L.; Di, D.W.; Luo, P.; Zhang, L.; Li, X.F.; Guo, G.Q.; Wu, L. The roles of epigenetic modifications in the regulation of auxin biosynthesis. Front. Plant Sci. 2022, 13, 959053. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin biosynthesis and its role in plant development. Annu. Rev. Plant Biol. 2010, 61, 49–64. [Google Scholar] [CrossRef]
- Aizezi, Y.; Xie, Y.P.; Guo, H.W.; Jiang, K. New Wine in an Old Bottle: Utilizing Chemical Genetics to Dissect Apical Hook Development. Life 2022, 12, 1285. [Google Scholar] [CrossRef]
- Hayashi, K.I. Chemical Biology in Auxin Research. Cold Spring Harb. Perspect. Biol. 2021, 13, a040105. [Google Scholar] [CrossRef]
- Jiang, K.; Asami, T. Chemical regulators of plant hormones and their applications in basic research and agriculture. Biosci. Biotechnol. Biochem. 2018, 82, 1265–1300. [Google Scholar] [CrossRef]
- Xie, Y.; Zhu, Y.; Wang, N.; Luo, M.; Ota, T.; Guo, R.; Takahashi, I.; Yu, Z.; Aizezi, Y.; Zhang, L.; et al. Chemical genetic screening identifies nalacin as an inhibitor of GH3 amido synthetase for auxin conjugation. Proc. Natl. Acad. Sci. USA 2022, 119, e2209256119. [Google Scholar] [CrossRef]
- Ruegger, M.; Dewey, E.; Hobbie, L.; Brown, D.; Bernasconi, P.; Turner, J.; Muday, G.; Estelle, M. Reduced naphthylphthalamic acid binding in the tir3 mutant of Arabidopsis is associated with a reduction in polar auxin transport and diverse morphological defects. Plant Cell 1997, 9, 745–757. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, K.; Neve, J.; Hirose, M.; Kuboki, A.; Shimada, Y.; Kepinski, S.; Nozaki, H. Rational design of an auxin antagonist of the SCF(TIR1) auxin receptor complex. ACS Chem. Biol. 2012, 7, 590–598. [Google Scholar] [CrossRef] [PubMed]
- He, W.R.; Brumos, J.; Li, H.J.; Ji, Y.S.; Ke, M.; Gong, X.Q.; Zeng, Q.L.; Li, W.Y.; Zhang, X.Y.; An, F.Y.; et al. A Small-Molecule Screen Identifies L-Kynurenine as a Competitive Inhibitor of TAA1/TAR Activity in Ethylene-Directed Auxin Biosynthesis and Root Growth in Arabidopsis. Plant Cell 2011, 23, 3944–3960. [Google Scholar] [CrossRef] [PubMed]
- Kakei, Y.; Nakamura, A.; Yamamoto, M.; Ishida, Y.; Yamazaki, C.; Sato, A.; Narukawa-Nara, M.; Soeno, K.; Shimada, Y. Biochemical and Chemical Biology Study of Rice OsTAR1 Revealed that Tryptophan Aminotransferase is Involved in Auxin Biosynthesis: Identification of a Potent OsTAR1 Inhibitor, Pyruvamine2031. Plant Cell Physiol. 2017, 58, 598–606. [Google Scholar] [CrossRef]
- Narukawa-Nara, M.; Nakamura, A.; Kikuzato, K.; Kakei, Y.; Sato, A.; Mitani, Y.; Yamasaki-Kokudo, Y.; Ishii, T.; Hayashi, K.; Asami, T.; et al. Aminooxy-naphthylpropionic acid and its derivatives are inhibitors of auxin biosynthesis targeting l-tryptophan aminotransferase: Structure-activity relationships. Plant J. 2016, 87, 245–257. [Google Scholar] [CrossRef]
- Soeno, K.; Goda, H.; Ishii, T.; Ogura, T.; Tachikawa, T.; Sasaki, E.; Yoshida, S.; Fujioka, S.; Asami, T.; Shimada, Y. Auxin biosynthesis inhibitors, identified by a genomics-based approach, provide insights into auxin biosynthesis. Plant Cell Physiol. 2010, 51, 524–536. [Google Scholar] [CrossRef]
- Kakei, Y.; Yamazaki, C.; Suzuki, M.; Nakamura, A.; Sato, A.; Ishida, Y.; Kikuchi, R.; Higashi, S.; Kokudo, Y.; Ishii, T.; et al. Small-molecule auxin inhibitors that target YUCCA are powerful tools for studying auxin function. Plant J. 2015, 84, 827–837. [Google Scholar] [CrossRef]
- Eswaramoorthy, S.; Bonanno, J.B.; Burley, S.K.; Swaminathan, S. Mechanism of action of a flavin-containing monooxygenase. Proc. Natl. Acad. Sci. USA 2006, 103, 9832–9837. [Google Scholar] [CrossRef]
- Nishimura, T.; Hayashi, K.; Suzuki, H.; Gyohda, A.; Takaoka, C.; Sakaguchi, Y.; Matsumoto, S.; Kasahara, H.; Sakai, T.; Kato, J.; et al. Yucasin is a potent inhibitor of YUCCA, a key enzyme in auxin biosynthesis. Plant J. 2014, 77, 352–366. [Google Scholar] [CrossRef]
- Tsugafune, S.; Mashiguchi, K.; Fukui, K.; Takebayashi, Y.; Nishimura, T.; Sakai, T.; Shimada, Y.; Kasahara, H.; Koshiba, T.; Hayashi, K.I. Yucasin DF, a potent and persistent inhibitor of auxin biosynthesis in plants. Sci. Rep. 2017, 7, 13992. [Google Scholar] [CrossRef]
- Zhu, Y.; Li, H.J.; Su, Q.; Wen, J.; Wang, Y.F.; Song, W.; Xie, Y.P.; He, W.R.; Yang, Z.; Jiang, K.; et al. A phenotype-directed chemical screen identifies ponalrestat as an inhibitor of the plant flavin monooxygenase YUCCA in auxin biosynthesis. J. Biol. Chem. 2019, 294, 19923–19933. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lang, Z.; Zhu, J.K. Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Stroud, H.; Do, T.; Du, J.; Zhong, X.; Feng, S.; Johnson, L.; Patel, D.J.; Jacobsen, S.E. Non-CG methylation patterns shape the epigenetic landscape in Arabidopsis. Nat. Struct. Mol. Biol. 2014, 21, 64–72. [Google Scholar] [CrossRef]
- Markulin, L.; Skiljaica, A.; Tokic, M.; Jagic, M.; Vuk, T.; Bauer, N.; Leljak Levanic, D. Taking the Wheel—De novo DNA Methylation as a Driving Force of Plant Embryonic Development. Front. Plant Sci. 2021, 12, 764999. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Hu, X.; Zhu, M.; Xu, M.; Wang, L. Transcription factors NF-YA2 and NF-YA10 regulate leaf growth via auxin signaling in Arabidopsis. Sci. Rep. 2017, 7, 1395. [Google Scholar] [CrossRef]
- Xu, Y.; Prunet, N.; Gan, E.S.; Wang, Y.; Stewart, D.; Wellmer, F.; Huang, J.; Yamaguchi, N.; Tatsumi, Y.; Kojima, M.; et al. SUPERMAN regulates floral whorl boundaries through control of auxin biosynthesis. Embo J. 2018, 37, e97499. [Google Scholar] [CrossRef]
- Gyula, P.; Baksa, I.; Toth, T.; Mohorianu, I.; Dalmay, T.; Szittya, G. Ambient temperature regulates the expression of a small set of sRNAs influencing plant development through NF-YA2 and YUC2. Plant Cell Environ. 2018, 41, 2404–2417. [Google Scholar] [CrossRef]
- Li, C.; Gu, L.; Gao, L.; Chen, C.; Wei, C.Q.; Qiu, Q.; Chien, C.W.; Wang, S.; Jiang, L.; Ai, L.F.; et al. Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat. Genet. 2016, 48, 687–693. [Google Scholar] [CrossRef]
- Poulios, S.; Vlachonasios, K.E. Synergistic action of GCN5 and CLAVATA1 in the regulation of gynoecium development in Arabidopsis thaliana. New Phytol. 2018, 220, 593–608. [Google Scholar] [CrossRef]
- Yamaguchi, N.; Huang, J.; Tatsumi, Y.; Abe, M.; Sugano, S.S.; Kojima, M.; Takebayashi, Y.; Kiba, T.; Yokoyama, R.; Nishitani, K.; et al. Chromatin-mediated feed-forward auxin biosynthesis in floral meristem determinacy. Nat. Commun. 2018, 9, 5290. [Google Scholar] [CrossRef]
- Lin, X.; Yuan, C.; Zhu, B.; Yuan, T.; Li, X.; Yuan, S.; Cui, S.; Zhao, H. LFR Physically and Genetically Interacts With SWI/SNF Component SWI3B to Regulate Leaf Blade Development in Arabidopsis. Front. Plant Sci. 2021, 12, 717649. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Luo, P.; Bai, J.; Wu, L.; Di, D.W.; Liu, H.Q.; Li, J.J.; Liu, Y.L.; Khaskheli, A.J.; Zhao, C.M.; et al. Function of histone H2B monoubiquitination in transcriptional regulation of auxin biosynthesis in Arabidopsis. Commun. Biol. 2021, 4, 206. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Jung, J.H.; Cortes Llorca, L.; Kim, S.G.; Lee, S.; Baldwin, I.T.; Park, C.M. FCA mediates thermal adaptation of stem growth by attenuating auxin action in Arabidopsis. Nat. Commun. 2014, 5, 5473. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Li, Z.; Zhou, N.; Ma, M.; Jiang, Y.; Dong, A.; Shen, W.H.; Li, L. Linking PHYTOCHROME-INTERACTING FACTOR to Histone Modification in Plant Shade Avoidance. Plant Physiol. 2018, 176, 1341–1351. [Google Scholar] [CrossRef]
- Tasset, C.; Singh Yadav, A.; Sureshkumar, S.; Singh, R.; van der Woude, L.; Nekrasov, M.; Tremethick, D.; van Zanten, M.; Balasubramanian, S. POWERDRESS-mediated histone deacetylation is essential for thermomorphogenesis in Arabidopsis thaliana. PLoS Genet. 2018, 14, e1007280. [Google Scholar] [CrossRef]
- van der Woude, L.C.; Perrella, G.; Snoek, B.L.; van Hoogdalem, M.; Novak, O.; van Verk, M.C.; van Kooten, H.N.; Zorn, L.E.; Tonckens, R.; Dongus, J.A.; et al. HISTONE DEACETYLASE 9 stimulates auxin-dependent thermomorphogenesis in Arabidopsis thaliana by mediating H2A.Z depletion. Proc. Natl. Acad. Sci. USA 2019, 116, 25343–25354. [Google Scholar] [CrossRef]
- Xue, M.; Zhang, H.; Zhao, F.; Zhao, T.; Li, H.; Jiang, D. The INO80 chromatin remodeling complex promotes thermomorphogenesis by connecting H2A.Z eviction and active transcription in Arabidopsis. Mol. Plant 2021, 14, 1799–1813. [Google Scholar] [CrossRef]
- Cui, X.; Lu, F.; Qiu, Q.; Zhou, B.; Gu, L.; Zhang, S.; Kang, Y.; Cui, X.; Ma, X.; Yao, Q.; et al. REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat. Genet. 2016, 48, 694–699. [Google Scholar] [CrossRef]
- Lee, K.; Seo, P.J. Coordination of matrix attachment and ATP-dependent chromatin remodeling regulate auxin biosynthesis and Arabidopsis hypocotyl elongation. PLoS ONE 2017, 12, e0181804. [Google Scholar] [CrossRef]
- Figueiredo, D.D.; Batista, R.A.; Roszak, P.J.; Kohler, C. Auxin production couples endosperm development to fertilization. Nat. Plants 2015, 1, 15184. [Google Scholar] [CrossRef]
- Milutinovic, M.; Lindsey, B.E., 3rd; Wijeratne, A.; Hernandez, J.M.; Grotewold, N.; Fernandez, V.; Grotewold, E.; Brkljacic, J. Arabidopsis EMSY-like (EML) histone readers are necessary for post-fertilization seed development, but prevent fertilization-independent seed formation. Plant Sci. 2019, 285, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Rizzardi, K.; Landberg, K.; Nilsson, L.; Ljung, K.; Sundas-Larsson, A. TFL2/LHP1 is involved in auxin biosynthesis through positive regulation of YUCCA genes. Plant J. 2011, 65, 897–906. [Google Scholar] [CrossRef] [PubMed]
- Manuela, D.; Xu, M. Juvenile Leaves or Adult Leaves: Determinants for Vegetative Phase Change in Flowering Plants. Int. J. Mol. Sci. 2020, 21, 9753. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.; Harrar, Y.; Lin, C.; Reinhart, B.; Newell, N.R.; Talavera-Rauh, F.; Hokin, S.A.; Barton, M.K.; Kerstetter, R.A. Arabidopsis KANADI1 acts as a transcriptional repressor by interacting with a specific cis-element and regulates auxin biosynthesis, transport, and signaling in opposition to HD-ZIPIII factors. Plant Cell 2014, 26, 246–262. [Google Scholar] [CrossRef] [PubMed]
- Ohashi-Ito, K.; Iwamoto, K.; Nagashima, Y.; Kojima, M.; Sakakibara, H.; Fukuda, H. A Positive Feedback Loop Comprising LHW-TMO5 and Local Auxin Biosynthesis Regulates Initial Vascular Development in Arabidopsis Roots. Plant Cell Physiol. 2019, 60, 2684–2691. [Google Scholar] [CrossRef] [PubMed]
- Khanday, I.; Santos-Medellin, C.; Sundaresan, V. Somatic embryo initiation by rice BABY BOOM1 involves activation of zygote-expressed auxin biosynthesis genes. New Phytol. 2023, 238, 673–687. [Google Scholar] [CrossRef]
- Lorrain, S.; Allen, T.; Duek, P.D.; Whitelam, G.C.; Fankhauser, C. Phytochrome-mediated inhibition of shade avoidance involves degradation of growth-promoting bHLH transcription factors. Plant J. 2008, 53, 312–323. [Google Scholar] [CrossRef]
- Gangappa, S.N.; Kumar, S.V. DET1 and HY5 Control PIF4-Mediated Thermosensory Elongation Growth through Distinct Mechanisms. Cell Rep. 2017, 18, 344–351. [Google Scholar] [CrossRef]
- Huai, J.L.; Zhang, X.Y.; Li, J.L.; Ma, T.T.; Zha, P.; Jing, Y.J.; Lin, R.C. SEUSS and PIF4 Coordinately Regulate Light and Temperature Signaling Pathways to Control Plant Growth. Mol. Plant 2018, 11, 928–942. [Google Scholar] [CrossRef]
- Ma, D.; Li, X.; Guo, Y.; Chu, J.; Fang, S.; Yan, C.; Noel, J.P.; Liu, H. Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc. Natl. Acad. Sci. USA 2016, 113, 224–229. [Google Scholar] [CrossRef]
- Zhu, J.Y.; Oh, E.; Wang, T.; Wang, Z.Y. TOC1-PIF4 interaction mediates the circadian gating of thermoresponsive growth in Arabidopsis. Nat. Commun. 2016, 7, 13692. [Google Scholar] [CrossRef]
- de Lucas, M.; Daviere, J.M.; Rodriguez-Falcon, M.; Pontin, M.; Iglesias-Pedraz, J.M.; Lorrain, S.; Fankhauser, C.; Blazquez, M.A.; Titarenko, E.; Prat, S. A molecular framework for light and gibberellin control of cell elongation. Nature 2008, 451, 480–484. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Liu, X.; Ljung, K.; Li, S.; Zhao, W.; Yang, F.; Wang, M.; Tao, Y. Type B Response Regulators Act As Central Integrators in Transcriptional Control of the Auxin Biosynthesis Enzyme TAA1. Plant Physiol 2017, 175, 1438–1454. [Google Scholar] [CrossRef]
- Fiorucci, A.S.; Galvao, V.C.; Ince, Y.C.; Boccaccini, A.; Goyal, A.; Allenbach Petrolati, L.; Trevisan, M.; Fankhauser, C. PHYTOCHROME INTERACTING FACTOR 7 is important for early responses to elevated temperature in Arabidopsis seedlings. New Phytol. 2020, 226, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Rawat, R.; Schwartz, J.; Jones, M.A.; Sairanen, I.; Cheng, Y.F.; Andersson, C.R.; Zhao, Y.D.; Ljung, K.; Harmer, S.L. REVEILLE1, a Myb-like transcription factor, integrates the circadian clock and auxin pathways. Proc. Natl. Acad. Sci. USA 2009, 106, 16883–16888. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Li, Y.; Pan, Y.; Zhong, S. Activation of HLS1 by Mechanical Stress via Ethylene-Stabilized EIN3 Is Crucial for Seedling Soil Emergence. Front. Plant Sci. 2016, 7, 1571. [Google Scholar] [CrossRef] [PubMed]
- Kwon, Y.; Kim, J.H.; Nguyen, H.N.; Jikumaru, Y.; Kamiya, Y.; Hong, S.W.; Lee, H. A novel Arabidopsis MYB-like transcription factor, MYBH, regulates hypocotyl elongation by enhancing auxin accumulation. J. Exp. Bot. 2013, 64, 3911–3922. [Google Scholar] [CrossRef]
- Saitoh, A.; Takase, T.; Abe, H.; Watahiki, M.; Hirakawa, Y.; Kiyosue, T. ZEITLUPE enhances expression of PIF4 and YUC8 in the upper aerial parts of Arabidopsis seedlings to positively regulate hypocotyl elongation. Plant Cell Rep. 2021, 40, 479–489. [Google Scholar] [CrossRef]
- Challa, K.R.; Aggarwal, P.; Nath, U. Activation of YUCCA5 by the Transcription Factor TCP4 Integrates Developmental and Environmental Signals to Promote Hypocotyl Elongation in Arabidopsis. Plant Cell 2016, 28, 2117–2130. [Google Scholar] [CrossRef]
- Cai, X.T.; Xu, P.; Zhao, P.X.; Liu, R.; Yu, L.H.; Xiang, C.B. Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nat. Commun. 2014, 5, 5833. [Google Scholar] [CrossRef]
- Perez-Alonso, M.M.; Sanchez-Parra, B.; Ortiz-Garcia, P.; Santamaria, M.E.; Diaz, I.; Pollmann, S. Jasmonic Acid-Dependent MYC Transcription Factors Bind to a Tandem G-Box Motif in the YUCCA8 and YUCCA9 Promoters to Regulate Biotic Stress Responses. Int. J. Mol. Sci. 2021, 22, 9768. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Gao, S.; Tian, H.; Wu, W.; Robert, H.S.; Ding, Z. Local Transcriptional Control of YUCCA Regulates Auxin Promoted Root-Growth Inhibition in Response to Aluminium Stress in Arabidopsis. PLoS Genet. 2016, 12, e1006360. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; Zhang, Z.; Wang, J.; Chen, X.; Wei, P.; Huang, R. The activation of OsEIL1 on YUC8 transcription and auxin biosynthesis is required for ethylene-inhibited root elongation in rice early seedling development. PLoS Genet. 2017, 13, e1006955. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Ma, B.; Tao, J.J.; Yin, C.C.; Hu, Y.; Huang, Y.H.; Wei, W.; Xin, P.Y.; Chu, J.F.; Zhang, W.K.; et al. Rice EIL1 interacts with OsIAAs to regulate auxin biosynthesis mediated by the tryptophan aminotransferase MHZ10/OsTAR2 during root ethylene responses. Plant Cell 2022, 34, 4366–4387. [Google Scholar] [CrossRef]
- Qin, H.; Wang, J.; Zhou, J.; Qiao, J.; Li, Y.; Quan, R.; Huang, R. Abscisic acid promotes auxin biosynthesis to inhibit primary root elongation in rice. Plant Physiol. 2023, 191, 1953–1967. [Google Scholar] [CrossRef]
- Tang, L.P.; Zhou, C.; Wang, S.S.; Yuan, J.; Zhang, X.S.; Su, Y.H. FUSCA3 interacting with LEAFY COTYLEDON2 controls lateral root formation through regulating YUCCA4 gene expression in Arabidopsis thaliana. New Phytol. 2017, 213, 1740–1754. [Google Scholar] [CrossRef]
- Yu, L.H.; Miao, Z.Q.; Qi, G.F.; Wu, J.; Cai, X.T.; Mao, J.L.; Xiang, C.B. MADS-box transcription factor AGL21 regulates lateral root development and responds to multiple external and physiological signals. Mol. Plant 2014, 7, 1653–1669. [Google Scholar] [CrossRef]
- Meng, W.J.; Cheng, Z.J.; Sang, Y.L.; Zhang, M.M.; Rong, X.F.; Wang, Z.W.; Tang, Y.Y.; Zhang, X.S. Type-B ARABIDOPSIS RESPONSE REGULATORs Specify the Shoot Stem Cell Niche by Dual Regulation of WUSCHEL. Plant Cell 2017, 29, 1357–1372. [Google Scholar] [CrossRef]
- Cui, D.; Zhao, J.; Jing, Y.; Fan, M.; Liu, J.; Wang, Z.; Xin, W.; Hu, Y. The arabidopsis IDD14, IDD15, and IDD16 cooperatively regulate lateral organ morphogenesis and gravitropism by promoting auxin biosynthesis and transport. PLoS Genet. 2013, 9, e1003759. [Google Scholar] [CrossRef]
- Eklund, D.M.; Staldal, V.; Valsecchi, I.; Cierlik, I.; Eriksson, C.; Hiratsu, K.; Ohme-Takagi, M.; Sundstrom, J.F.; Thelander, M.; Ezcurra, I.; et al. The Arabidopsis thaliana STYLISH1 protein acts as a transcriptional activator regulating auxin biosynthesis. Plant Cell 2010, 22, 349–363. [Google Scholar] [CrossRef]
- Gao, F.; Wang, K.; Liu, Y.; Chen, Y.; Chen, P.; Shi, Z.; Luo, J.; Jiang, D.; Fan, F.; Zhu, Y.; et al. Blocking miR396 increases rice yield by shaping inflorescence architecture. Nat. Plants 2015, 2, 15196. [Google Scholar] [CrossRef] [PubMed]
- Reyes-Olalde, J.I.; Zuniga-Mayo, V.M.; Serwatowska, J.; Chavez Montes, R.A.; Lozano-Sotomayor, P.; Herrera-Ubaldo, H.; Gonzalez-Aguilera, K.L.; Ballester, P.; Ripoll, J.J.; Ezquer, I.; et al. The bHLH transcription factor SPATULA enables cytokinin signaling, and both activate auxin biosynthesis and transport genes at the medial domain of the gynoecium. PLoS Genet. 2017, 13, e1006726. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Chen, Y.; Liu, L.; See, Y.H.B.; Mao, C.; Gan, Y.; Yu, H. OsFTIP7 determines auxin-mediated anther dehiscence in rice. Nat. Plants 2018, 4, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Stone, S.L.; Braybrook, S.A.; Paula, S.L.; Kwong, L.W.; Meuser, J.; Pelletier, J.; Hsieh, T.F.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. Arabidopsis LEAFY COTYLEDON2 induces maturation traits and auxin activity: Implications for somatic embryogenesis. Proc. Natl. Acad. Sci. USA 2008, 105, 3151–3156. [Google Scholar] [CrossRef] [PubMed]
- Balcerowicz, M. Filling the grain: Transcription factor OsNF-YB1 triggers auxin biosynthesis to boost rice grain size. Plant Physiol. 2021, 185, 757–758. [Google Scholar] [CrossRef]
- Li, Y.J.; Yu, Y.; Liu, X.; Zhang, X.S.; Su, Y.H. The Arabidopsis MATERNAL EFFECT EMBRYO ARREST45 protein modulates maternal auxin biosynthesis and controls seed size by inducing AINTEGUMENTA. Plant Cell 2021, 33, 1907–1926. [Google Scholar] [CrossRef]
- Zhang, M.; Zheng, H.; Jin, L.; Xing, L.; Zou, J.; Zhang, L.; Liu, C.; Chu, J.; Xu, M.; Wang, L. miR169o and ZmNF-YA13 act in concert to coordinate the expression of ZmYUC1 that determines seed size and weight in maize kernels. New Phytol. 2022, 235, 2270–2284. [Google Scholar] [CrossRef]
- Kriechbaumer, V.; Wang, P.; Hawes, C.; Abell, B.M. Alternative splicing of the auxin biosynthesis gene YUCCA4 determines its subcellular compartmentation. Plant J. 2012, 70, 292–302. [Google Scholar] [CrossRef]
- Kashkan, I.; Hrtyan, M.; Retzer, K.; Humpolickova, J.; Jayasree, A.; Filepova, R.; Vondrakova, Z.; Simon, S.; Rombaut, D.; Jacobs, T.B.; et al. Mutually opposing activity of PIN7 splicing isoforms is required for auxin-mediated tropic responses in Arabidopsis thaliana. New Phytol. 2022, 233, 329–343. [Google Scholar] [CrossRef]
- Kashkan, I.; Timofeyenko, K.; Kollarova, E.; Ruzicka, K. In Vivo Reporters for Visualizing Alternative Splicing of Hormonal Genes. Plants 2020, 9, 868. [Google Scholar] [CrossRef]
- Millevoi, S.; Vagner, S. Molecular mechanisms of eukaryotic pre-mRNA 3′ end processing regulation. Nucleic Acids Res. 2010, 38, 2757–2774. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zeng, L. Crosstalk between Ubiquitination and Other Post-translational Protein Modifications in Plant Immunity. Plant Communities 2020, 1, 100041. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Qin, G.; Cao, M.; Chen, R.; He, Y.; Yang, L.; Zeng, Z.; Yu, Y.; Gu, Y.; Xing, W.; et al. A phosphorylation-based switch controls TAA1-mediated auxin biosynthesis in plants. Nat. Commun. 2020, 11, 679. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.Q.; Pu, Z.X.; Di, D.W.; Zou, Y.J.; Guo, Y.M.; Wang, J.L.; Zhang, L.; Tian, P.; Fei, Q.H.; Li, X.F.; et al. Significance of NatB-mediated N-terminal acetylation of auxin biosynthetic enzymes in maintaining auxin homeostasis in Arabidopsis thaliana. Commun. Biol. 2022, 5, 1410. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Zhang, C.; Sha, H.; Wang, X.; Yu, Y.; Liu, J.; Zhao, G.; Wang, J.; Qiu, G.; Xu, X.; et al. Ubiquitin-Conjugating Enzyme OsUBC11 Affects the Development of Roots via Auxin Pathway. Rice 2023, 16, 9. [Google Scholar] [CrossRef]
- Tan, S.; Luschnig, C.; Friml, J. Pho-view of Auxin: Reversible Protein Phosphorylation in Auxin Biosynthesis, Transport and Signaling. Mol. Plant 2021, 14, 151–165. [Google Scholar] [CrossRef]
- Suzuki, M.; Yamazaki, C.; Mitsui, M.; Kakei, Y.; Mitani, Y.; Nakamura, A.; Ishii, T.; Soeno, K.; Shimada, Y. Transcriptional feedback regulation of YUCCA genes in response to auxin levels in Arabidopsis. Plant Cell Rep. 2015, 34, 1343–1352. [Google Scholar] [CrossRef]
- Eliot, A.C.; Kirsch, J.F. Pyridoxal phosphate enzymes: Mechanistic, structural, and evolutionary considerations. Annu. Rev. Biochem. 2004, 73, 383–415. [Google Scholar] [CrossRef]
- Chen, L.; Huang, X.X.; Zhao, S.M.; Xiao, D.W.; Xiao, L.T.; Tong, J.H.; Wang, W.S.; Li, Y.J.; Ding, Z.; Hou, B.K. IPyA glucosylation mediates light and temperature signaling to regulate auxin-dependent hypocotyl elongation in Arabidopsis. Proc. Natl. Acad. Sci. USA 2020, 117, 6910–6917. [Google Scholar] [CrossRef]
- Zheng, Z.; Guo, Y.; Novak, O.; Dai, X.; Zhao, Y.; Ljung, K.; Noel, J.P.; Chory, J. Coordination of auxin and ethylene biosynthesis by the aminotransferase VAS1. Nat. Chem. Biol. 2013, 9, 244–246. [Google Scholar] [CrossRef]
- Di, D.W.; Sun, L.; Wang, M.; Wu, J.; Kronzucker, H.J.; Fang, S.; Chu, J.; Shi, W.; Li, G. WRKY46 promotes ammonium tolerance in Arabidopsis by repressing NUDX9 and indole-3-acetic acid-conjugating genes and by inhibiting ammonium efflux in the root elongation zone. New Phytol. 2021, 232, 190–207. [Google Scholar] [CrossRef] [PubMed]
- Luo, P.; Di, D.; Wu, L.; Yang, J.; Lu, Y.; Shi, W. MicroRNAs Are Involved in Regulating Plant Development and Stress Response through Fine-Tuning of TIR1/AFB-Dependent Auxin Signaling. Int. J. Mol. Sci. 2022, 23, 510. [Google Scholar] [CrossRef] [PubMed]
- Forgione, I.; Woloszynska, M.; Pacenza, M.; Chiappetta, A.; Greco, M.; Araniti, F.; Abenavoli, M.R.; Van Lijsebettens, M.; Bitonti, M.B.; Bruno, L. Hypomethylated drm1 drm2 cmt3 mutant phenotype of Arabidopsis thaliana is related to auxin pathway impairment. Plant Sci. 2019, 280, 383–396. [Google Scholar] [CrossRef] [PubMed]
- Gaddam, S.R.; Bhatia, C.; Sharma, A.; Badola, P.K.; Saxena, G.; Trivedi, P.K. miR775 integrates light, sucrose and auxin associated pathways to regulate root growth in Arabidopsis thaliana. Plant Sci. 2021, 313, 111073. [Google Scholar] [CrossRef]
- Li, J.Y.; Ren, J.J.; Zhang, T.X.; Cui, J.H.; Gong, C.M. CkREV Enhances the Drought Resistance of Caragana korshinskii through Regulating the Expression of Auxin Synthetase Gene CkYUC5. Int. J. Mol. Sci. 2022, 23, 5902. [Google Scholar] [CrossRef]
- Li, Q.; Yin, M.; Li, Y.; Fan, C.; Yang, Q.; Wu, J.; Zhang, C.; Wang, H.; Zhou, Y. Expression of Brassica napus TTG2, a regulator of trichome development, increases plant sensitivity to salt stress by suppressing the expression of auxin biosynthesis genes. J. Exp. Bot. 2015, 66, 5821–5836. [Google Scholar] [CrossRef]
- Shao, A.; Ma, W.; Zhao, X.; Hu, M.; He, X.; Teng, W.; Li, H.; Tong, Y. The Auxin Biosynthetic TRYPTOPHAN AMINOTRANSFERASE RELATED TaTAR2.1-3A Increases Grain Yield of Wheat. Plant Physiol. 2017, 174, 2274–2288. [Google Scholar] [CrossRef]
- Di, D.W.; Sun, L.; Zhang, X.N.; Li, G.J.; Kronzucker, H.J.; Shi, W.M. Involvement of auxin in the regulation of ammonium tolerance in rice (Oryza sativa L.). Plant Soil 2018, 432, 373–387. [Google Scholar] [CrossRef]
- Zhao, X.; Wen, B.; Li, C.; Liu, L.; Chen, X.; Li, D.; Li, L.; Fu, X. PpEBB1 directly binds to the GCC box-like element of auxin biosynthesis related genes. Plant Sci. 2021, 306, 110874. [Google Scholar] [CrossRef]
- Min, L.; Hu, Q.; Li, Y.; Xu, J.; Ma, Y.; Zhu, L.; Yang, X.; Zhang, X. LEAFY COTYLEDON1-CASEIN KINASE I-TCP15-PHYTOCHROME INTERACTING FACTOR4 network regulates somatic embryogenesis by regulating auxin homeostasis. Plant Physiol. 2015, 169, 2805–2821. [Google Scholar] [CrossRef]
- Schiessl, K.; Lilley, J.L.S.; Lee, T.; Tamvakis, I.; Kohlen, W.; Bailey, P.C.; Thomas, A.; Luptak, J.; Ramakrishnan, K.; Carpenter, M.D.; et al. NODULE INCEPTION recruits the lateral root developmental program for symbiotic nodule organogenesis in Medicago truncatula. Curr. Biol. 2019, 29, 3657–3668 e5. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Wang, R.; Kong, K.; Begum, N.; Almakas, A.; Liu, J.; Li, H.; Liu, B.; Zhao, T.; Zhao, T. An APETALA2/ethylene responsive factor transcription factor GmCRF4a regulates plant height and auxin biosynthesis in soybean. Front. Plant Sci. 2022, 13, 983650. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhou, Z.; Wang, L.; Yan, S.; Cheng, Z.; Liu, X.; Han, L.; Chen, G.; Wang, S.; Song, W.; et al. The CsHEC1-CsOVATE module contributes to fruit neck length variation via modulating auxin biosynthesis in cucumber. Proc. Natl. Acad. Sci. USA 2022, 119, e2209717119. [Google Scholar] [CrossRef] [PubMed]
- Li, L.L.; Zheng, T.C.; Li, P.; Liu, W.C.; Qiu, L.K.; Wang, J.; Cheng, T.R.; Zhang, Q.X. Integrative analysis of HD-Zip III gene PmHB1 contribute to the plant architecture in Prunus mume. Sci. Horticamsterdam 2022, 293, 110664. [Google Scholar] [CrossRef]
- Wang, H.L.; Yang, Q.; Tan, S.Y.; Wang, T.; Zhang, Y.; Yang, Y.L.; Yin, W.L.; Xia, X.L.; Guo, H.W.; Li, Z.H. Regulation of cytokinin biosynthesis using PtRD26pro-IPT module improves drought tolerance through PtARR10-PtYUC4/5-mediated reactive oxygen species removal in Populus. J. Integr. Plant Biol. 2022, 64, 771–786. [Google Scholar] [CrossRef]



| Gene | Regulated by | Related Developmental Process | Ref. |
|---|---|---|---|
| YUC1 | SUP-LHP1-PRC2 complex | Floral patterning | [46] |
| YUC2 | DRM1 and DRM2 | Leaf growth and thermomorphogenesis | [45,47] |
| YUC3 | BRM and REF6 | n.s. | [48] |
| YUC4 | GCN5/HAG1 | Gynoecium development | [49] |
| YUC4 | SUP-LHP1-PRC2 complex | Floral whorl boundaries | [46] |
| YUC4 | CLF, LHP1, CHR11, and CHR17 | Floral patterning and floral determinacy | [46,50] |
| YUC6 | SWI3B | Leaf blade development | [51] |
| YUC7 | HUB complex | Root gravitropism | [52] |
| YUC8 | FCA | Thermal adaptation of stem growth | [53] |
| YUC8 | PIF7-MRG2 complex | Shade-induced hypocotyl elongation | [54] |
| YUC8 | SWR1 chromatin remodeling complex | Thermomorphogenesis | [55] |
| YUC8 | HDA9-PWR complex | Thermomorphogenesis | [56] |
| YUC8 | INO80 chromatin remodeling complex | Thermomorphogenesis | [57] |
| YUC8 | JMJ14, JMJ15, and JMJ18 | Response to high temperature | [58] |
| YUC9 | ARP4 | Shade-induced hypocotyl elongation | [59] |
| YUC10 | FIS2-PRC2 complex | Endosperm development | [60] |
| YUC10 | EML1 and EML3 | Seed development | [61] |
| YUCs | TFL2/LHP1 | n.s. | [62] |
| YUCs | JMJ12/REF6 | Thermomorphogenesis | [58] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, P.; Di, D.-W. Precise Regulation of the TAA1/TAR-YUCCA Auxin Biosynthesis Pathway in Plants. Int. J. Mol. Sci. 2023, 24, 8514. https://doi.org/10.3390/ijms24108514
Luo P, Di D-W. Precise Regulation of the TAA1/TAR-YUCCA Auxin Biosynthesis Pathway in Plants. International Journal of Molecular Sciences. 2023; 24(10):8514. https://doi.org/10.3390/ijms24108514
Chicago/Turabian StyleLuo, Pan, and Dong-Wei Di. 2023. "Precise Regulation of the TAA1/TAR-YUCCA Auxin Biosynthesis Pathway in Plants" International Journal of Molecular Sciences 24, no. 10: 8514. https://doi.org/10.3390/ijms24108514
APA StyleLuo, P., & Di, D. -W. (2023). Precise Regulation of the TAA1/TAR-YUCCA Auxin Biosynthesis Pathway in Plants. International Journal of Molecular Sciences, 24(10), 8514. https://doi.org/10.3390/ijms24108514







