Abstract
Hepatitis B (HB) vaccination effectively reduces the risks of chronic infection with the hepatitis B virus (HBV). It is unknown whether there is a common genetic determinant for response to the HB vaccine and susceptibility to chronic HBV infection. This case-control study, which included 193 chronic HBV carriers and 495 non-carriers, aimed to explore the effects of the most significant single nucleotide polymorphisms (SNPs) in response to the HB vaccine on the risks of chronic HBV infection. Out of 13 tested SNPs, the genotype distributions of four SNPs at the human leukocyte antigen (HLA) class II region, including rs34039593, rs614348, rs7770370, and rs9277535, were significantly different between HBV carriers and non-carriers. The age-sex-adjusted odds ratios (OR) of chronic HBV infection for rs34039593 TG, rs614348 TC, rs7770370 AA, and rs9277535 AA genotypes were 0.51 (95% confidence interval [CI], 0.33–0.79; p = 0.0028), 0.49 (95% CI, 0.32–0.75; p = 6.5 × 10−4), 0.33 (95% CI, 0.18–0.63; p = 7.4 × 10−4), and 0.31 (95% CI, 0.14–0.70; p = 0.0043), respectively. Multivariable analyses showed that rs614348 TC and rs7770370 AA genotypes were significantly independent protectors against chronic HBV infection. The multivariable-adjusted ORs for subjects with none, either one, or both of the protective genotypes were 1.00 (referent), 0.47 (95% CI: 0.32–0.71; p = 3.0 × 10−4), and 0.16 (95% CI: 0.05–0.54; p = 0.0032), respectively. Among eight HBeAg-positive carriers, only one of them carried a protective genotype. This study shows that response to the HB vaccine and susceptibility to chronic HBV infection share common genetic determinants and indicates that HLA class II members are the main responsible host genetic factors.
1. Introduction
Chronic hepatitis B (HB) virus (HBV) is common, with a global prevalence of 3.9%, and is more frequent in East Asian populations [1]. It is one of the main causes of deaths and diseases worldwide and is linked to a significant increase in the risk of developing end-stage liver diseases, such as cirrhosis and liver cancer [2,3]. In 2017, approximately 800,000 deaths were caused by HBV, including 90,000 cases of acute hepatitis, 325,400 cases of liver cancer, and 384,000 cases of cirrhosis and other chronic liver diseases [4]. Chronic HBV infection also caused a loss of 28.3 million disability-adjusted life years and 24.4 million years of life in 2017 [5]. Thus, reducing HBV infections in susceptible individuals and reducing disease progression in HBV-infected individuals are crucial for global health.
The implementation of the universal HB vaccination program in Taiwan has led to a decrease in the rates of chronic HBV infection, from over 10% in the 1980s to less than 2% in recent years [6,7]. This has also resulted in a notable decrease in the incidence of liver cancer in children and young adults in Taiwan. As compared with subjects born before the universal HB vaccination program at ages 6–9, 10–14, 15–19, and 20–26 years, the rate ratios of hepatocellular carcinoma for subjects born after the program were 0.26, 0.34, 0.37, and 0.42, respectively [8]. Unfortunately, not many countries have adopted such programs until recently, and only half of the countries and territories have reached the Global Vaccine Action Plan target for HB vaccination coverage [9]. Furthermore, approximately 300 million individuals globally are HBV carriers [1]. As a result, new HBV infections inevitably occur, and an etiologic study on HBV infection seems relevant and urgent.
The universal HB vaccination program in Taiwan provides evidence that HB vaccines effectively induce host immunological responses against chronic HBV infection when contracting HBV and subsequently block the associated pathogenesis. Accordingly, studies on responses to HB vaccines may provide critical clues for the prevention of chronic HBV infection. Recently, several genome-wide association studies (GWAS) have identified a few single nucleotide polymorphisms (SNPs) that are correlated with responses to HB vaccines [10,11,12,13,14]. It is plausible that these genetic markers may also be associated with susceptibility to chronic HBV infection. Considering this, we conducted this community-based genetic association study to explore the relationships between these common genetic variants and the susceptibility to chronic infection in HBV-infected individuals.
2. Results
2.1. Clinical Characteristics of Study Subjects
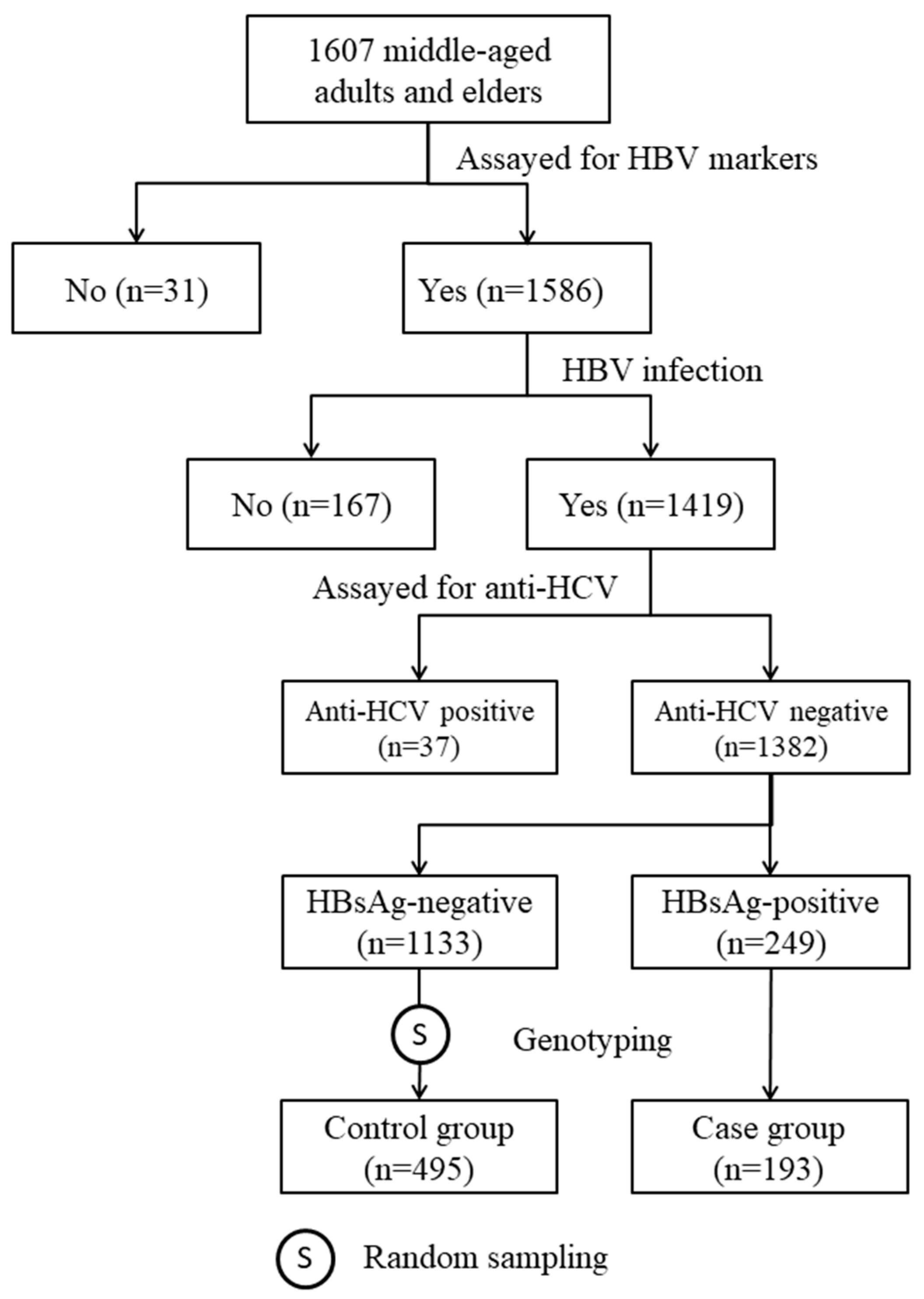
The subjects of the study were selected from participants in our previous community-based cohort study [15]. In the cohort, there were 1382 anti-hepatitis C virus (anti-HCV)-negative HBV-infected individuals. All HBV carriers with DNA samples available for genotyping (n = 193) and 495 randomly selected non-carriers were included in this case-control study (Figure 1). Among the HBV carriers, 191 (99.0%) of them were also positive for anti-body to HB core antigen (HBcAb; Table 1). The positive rates of HB e antigen (HBeAg) and anti-body to HBeAg (HBeAb) in HBV carriers were 4.3% and 85.3%, respectively. Among the non-carrier controls, the seropositive rates of anti-body to HB surface antigen (HBsAb) and HBcAb were 97.8% and 95.0%, respectively. Table 1 also shows that, as compared with the non-carrier controls, HBV carriers were significantly younger (50.1 ± 8.0 vs. 56.4 ± 9.1 years, p < 0.0001) and had a significantly higher mean level of ALT (27.1 ± 17.3 vs. 24.1 ± 14.4 IU/L, p = 0.030). There was no significant difference in other baseline clinical and anthropometric measurements between the cases and the controls.
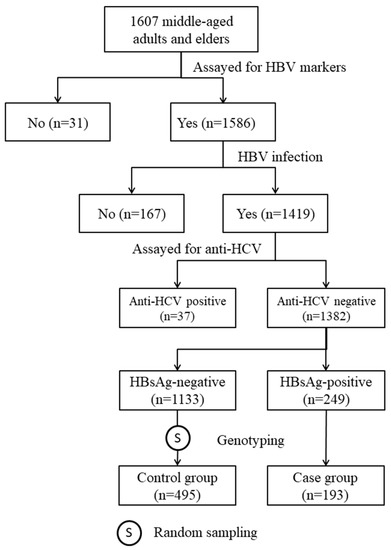
Figure 1.
Steps of subject selection.

Table 1.
Baseline clinical characteristics of HBV carriers and non-carrier controls.
2.2. Genotype Distributions and Genotypic Effects of the Tested SNPs
All 13 tested SNPs had a call rate >95%, a minor allele frequency >10.0%, and a p-value of the Hardy–Weinberg equilibrium test >0.05 (Table 2). The genotype distributions of rs34039593, rs614348, rs7770370, and rs9277535 were significantly different between the case and control groups. As compared with the non-carrier controls, the relative frequencies of the minor alleles of these four SNPs were significantly lower in the HBV carriers (rs34039593 G: 10.6% vs. 15.9%, p = 0.013; rs614348 C: 11.7% vs. 18.6%, p = 0.0022; rs7770370 A: 32.1% vs. 40.2%, p = 0.0068; rs9277535 A: 26.9% vs. 33.2%, p = 0.029). There was no significant difference in the genotype distributions of the remaining nine SNPs between the cases and the controls. Further genotypic effect analyses of the four significant SNPs showed that rs34039593 TG, rs614348 TC, rs7770370 AA, and rs9277535 AA genotypes were associated with significantly decreased risks of chronic HBV infection. The corresponding crude ORs were 0.52 (95% CI: 0.34–0.79; p = 0.0023), 0.51 (95% CI: 0.34–0.75; p = 7.0 × 10−4), 0.38 (95% CI: 0.21–0.67; p = 0.0020), and 0.34 (95% CI: 0.16–0.74; p = 0.0063), respectively.

Table 2.
Genotype distributions of 13 tested SNPs between HBV carriers and non-carrier controls.
2.3. Association Analyses with Outcomes of HBV Infection
After controlling for the effects of age and sex, these four genotypes remained correlated with significantly lower risks of chronic HBV infection (Table 3). Further multivariable analyses revealed that the rs614348 TC and rs7770370 AA genotypes showed significantly protective effects against chronic HBV infection. The corresponding adjusted ORs were 0.50 (95% CI: 0.33–0.77; p = 0.0016) and 0.35 (95% CI: 0.19–0.68; p = 0.0016), respectively.

Table 3.
Association analyses with risks of chronic HBV infection.
Table 4 shows that the HBV carriers had a significantly lower mean of the 4-locus genetic protective score (4-GPS) than the non-carrier controls (0.47 ± 0.88 vs. 0.88 ± 1.15, p = 8.2 × 10−7). As compared with individuals who had no protective genotype, the multivariable-adjusted ORs of chronic HBV infection for individuals who had 1–2 and 3–4 protective genotypes were 0.45 (95% CI: 0.30–0.67; p = 1.1 × 10−5) and 0.17 (95% CI: 0.05–0.58; p = 0.0047), respectively. The multivariable-adjusted OR of chronic HBV infection per 1.0 increase in 4-GPS was 0.65 (95% CI: 0.54–0.79; p = 1.1 × 10−5). Similar results were observed for the 2-locus GPS (2-GPS). The multivariable-adjusted OR of chronic HBV infection per 1.0 increase in the 2-GPS was 0.45 (95% CI: 0.32–0.63; p = 5.6 × 10−6).

Table 4.
Association analyses for chronic HBV infection with multi-locus genetic protective scores (GPS).
2.4. Genotypes of HBeAg-Positive Carriers
Among HBV carriers, eight (4.3%) of them were also positive for HBeAg. Table 5 shows that only one HBeAg-positive carrier was also positive for HBsAb, and none of them was positive for HBeAb. One HBeAg-positive carrier had an abnormal ALT level, and another two HBeAg-positive carriers had elevated ALT levels close to the upper limit of the normal range. Only one HBeAg-positive carrier (subject no. 629) carried protective genotypes at rs7770370 and rs9277535. The other HBeAg-positive carriers carried no protective genotype at all four polymorphic sites.

Table 5.
Genotypes and HBV markers of HBeAg-positive carriers.
3. Discussion
In this study, we examined the relationships between the top 13 significant SNPs in response to the HB vaccine and the risks of chronic HBV infection. We found that rs614348 C, rs7770370 A, rs34039593 G, and rs9277356 A alleles were correlated with significantly lower risks of being HBV carriers. Multivariable analysis showed that rs614348 TC and rs7770370 AA genotypes were significantly independent protectors against chronic HBV infection. Additionally, among eight HBeAg-positive carriers, only one of them carried the rs7770370 AA genotype. As far as we know, there is no prior research connecting the polymorphisms of rs614348 and rs34039593 to human health outcomes.
Our previous GWAS showed that human leukocyte antigen-DP (HLA-DP) is the key region for genetic determinants of response to the HB vaccine [10]. We searched the GWAS catalog database [16] and found there were only four associated GWAS studies [11,12,13,14]. These studies reported 26 tagged SNPs, 18 of which had a p-value < 10−6 and are located in chromosome 6 30.80–33.08 M bp. More importantly, 14 of these SNPs are located in the HLA class II region. Our [10] and previous GWAS [11,12,13,14] findings demonstrated the central roles of HLA class II members in responses to HB vaccines. It is also noted that HLA class II members may also play critical roles in HBV chronic infection, clearance, and responses to anti-viral therapies [17].
In the study, four SNPs were found to be significantly correlated with the risk of chronic HBV infection. The ORs of HBV carriers were significantly lower for the rs7770370 AA and rs9277535 AA genotypes. The directions of effects of the rs7770370 A and rs9277535 A alleles were consistent with both our [10] and a Korean study on the response to the HB vaccine [18]. SNP rs614348 is nearly completely linked with rs477515 in Southern Han Chinese [19]. SNP rs477515 was reported as the most significant SNP for response to the HB vaccine in Chinese Han populations. As compared with the C allele, the T allele was associated with a significantly higher OR (2.05; 95% CI: 1.95–2.41) of non-response to the HB vaccine [12]. Additionally, rs34039593 was reported as one of the lead SNPs for response to a HB vaccine in the Japanese, with the G allele linked to higher anti-HBs levels [13]. Our findings are consistent with previous genetic association studies on response to the HB vaccine.
Previous genetic studies support the significance of rs9277535 and rs7770370 polymorphisms with HBV chronic infection or clearance. Kamatani et al. were the first to report that rs9277535 was the most significantly associated SNP with chronic HBV infection, with the A allele correlated with a significantly lower risk (OR = 0.57; 95% CI: 0.52–0.62) [20]. This finding was consistently replicated in multiple GWAS studies [21,22,23,24]. A meta-analysis including 22,065 chronic HBV infection cases and 23,500 controls from 32 datasets showed that the rs9277535 A allele had a summary OR of 0.60 (95% CI: 0.57–0.63) for chronic HBV infection compared to the G allele [25]. A recent GWAS with 1943 chronic HBV infection cases and 9571 controls identified a cluster of 450 significant SNPs in the HLA class II region [26], among which rs7770370 was the tagged genetic marker and the A allele was correlated with a significantly lower risk of chronic HBV infections [26]. More recently, a community-based study that enrolled more than 0.5 million adults aged 30–79 years found that the rs7770370 A allele and the rs9277535 A allele were both correlated with significantly lower risks of HBsAg positivity [27]. There is no report regarding the health effects of the rs614348 and rs34039593 polymorphisms, except for their relationships with responses to the HB vaccine.
There are three lines of evidence to support the notion that polymorphisms of rs614348, rs7770370, rs34039593, and rs9277535 may contribute to the susceptibility to chronic HBV infection. First, we used the Ensemble Genome Browser [28] to retrieve the linkage disequilibrium (LD) data of these significant SNPs in the 1000 Human Genome Project Phase 3 Southern Han Chinese [19]. We found that all four of these SNPs are closely linked with nearby functional SNPs (Supplementary Table S1). SNP rs614348 is located in the intergenic region of the HLA-DRB1 and -DQA1 genes. It is closely linked with two missense variants (HLA-DQA1 rs75983419 and rs74379225), one synonymous variant (HLA-DRB1 rs201125976), and another thirteen functional SNPs. SNP rs7770370 is a non-coding transcript exon variant of the HLA-DPB1 gene and is also closely linked with another non-coding transcript exon variant of the HLA-DPB1 gene (rs7770371). SNP rs9277535 is located at the 3′-untranslated region (3′-UTR) of the HLA-DPB1 gene. It is closely linked with two splice region variants (HLA-DPB1 rs9277453 and rs9277454), seven missense variants (HLA-DPB1 rs9277471, rs1042335, rs1042187, rs9277356, rs9277355, rs9277354, and rs1042169), and three synonymous variants (HLA-DPB1 rs1042212, rs1042331, and rs1071597). Additionally, there are close linkages between the rs9277535 polymorphism and 60 3′-UTR variants of HLA-DPB1 [19]. SNP rs34039593 is located in the intergenic region of HLA-DRB1 and -DQA1 genes and is closely linked with one HLA-DRB1 missense variant (rs17879995; r2 = 0.95) and four regulatory region variants.
Second, we accessed the expression data in human immune cells [29] using the Ensemble Genome Browser [28] and found that the expression levels of the associated HLA class II genes are significantly influenced by these four significant SNPs. There is a perfect linkage between rs614348 C/T and rs9272545 G/T polymorphisms in Southern Han Chinese. SNP rs9272545 is a 3′-UTR variant of HLA-DQA1, and the G allele was correlated with significantly higher expression levels of HLA-DQA1 in multiple B cells, T cells, and monocytes compared to the C allele [29]. Furthermore, expression levels of HLA-DQB1, -DQB1-AS1, -DQB2, and -DRB5 in multiple immune cells were also significantly correlated with the rs9272545 polymorphism ([29]; Supplementary Table S2). The rs7770370 A allele is correlated with significantly higher expression levels of HLA-DPA1 in memory regulatory T cells, naïve B cells, and several monocytes. The rs7770370 A allele also correlates with a higher expression level of HLA-DPB1 in monocytes. Additionally, as compared with the rs7770370 G allele, the expression levels of HLA-DMA, -DMB, -DQA2, -DQB1, -DQB1-AS1, -DQB2, -DRB1, and -DRB5 are also significantly increased for the A allele in multiple B cells, T cells, and monocytes ([29]; Supplementary Table S2).
The rs9277535 A allele is correlated with higher expression levels of HLA-DPB1 in multiple B cells, T cells, and monocytes. In addition, the expressions of HLA-DMB, -DPA1, -DQA1, -DQB1, and -DQB1-AS1 in many immune cells are also influenced by the rs9277535 polymorphism ([29]; Supplementary Table S2). When using the nearest regulatory region variant rs3129753 as the surrogate for rs34039593 (r2 = 0.95), significant correlations with the expression levels of several HLA class II members were observed. For example, as compared with the rs3129753 G allele, the expression levels of HLA-DRB1 and -DQA1 were significantly increased in Th1, Th2, Th17, regulatory, CD4+, and CD8+ T cells, B cells, and several monocytes ([29]; Supplementary Table S2). It is therefore reasonable to hypothesize that the rs34039593 polymorphism may influence the expression of many HLA class II genes in immune cells.
Our previous study showed that both rs777037 A and rs9277535 A alleles are positively correlated with HLA-DPB1*protective alleles, including 02:01, 02:02, 03:01, and 04:01, and are negatively correlated with HLA-DPB1*risk alleles, including 05:01 and 09:01 [10]. Moreover, we also found that the HLA-DPA1*02:02:02-DPB1*risk haplotype was positively correlated and the HLA-DPA1*01:03:01-DPB1*protective haplotype was negatively correlated with the risks of non-response to the HB vaccine [30]. Recent studies found that the HLA-DPA1*01:03-DPB1*04:02 haplotype is correlated with high binding affinities for a large proportion of HBV epitopes [13,26]. On the contrary, the HLA-DPA1*02:02-DPB1*05:01 haplotype is correlated with poor binding affinities [13].
This study found no significant correlation between nine candidate SNPs and the risk of chronic HBV infection. One reason could be that except for rs3135363 and rs383006, the other seven SNPs were identified by the same Japanese GWAS study that enrolled subjects who received a HB vaccine designed based on HBV genotype C [13]. However, the most common HBV genotype in Taiwan, with a prevalence of 71%, is genotype B [31]. This suggests that the host genetic factors affecting immune responses to HBV genotypes B and C are likely different. The major antigenic determinants of these genotypes may also vary. Conversely, rs34039593 was identified as a tagged SNP by the same Japanese GWAS study [13] and was found to be related to the risk of chronic HBV infection in this study. It seems that the rs34039593 polymorphism confers immune responses to both HBV genotype B and genotype C. However, blood samples from more than half of the HBV carriers in this study were not enough for HBV genotyping. Further research is needed to test this hypothesis.
In conclusion, we found that four of thirteen candidate SNPs, which showed significant or promising associations with responses to HB vaccines, were significantly correlated with the risks of chronic HBV infection. Notably, the frequencies of the minor alleles of these four significant SNPs were significantly lower in the HBV carriers, and the rs614348 TC and rs7770370 AA genotypes showed significantly protective effects against chronic HBV infection. The results indicate that both the response to HB vaccination and the susceptibility to chronic HBV infection have shared genetic determinants.
4. Materials and Methods
4.1. Study Subjects
The subjects of this study were selected from participants in our previous community-based study [15]. From September 2010 to May 2012, a total of 1607 middle-aged adults and elders voluntarily gave informed consent and received physical examinations. At the date of enrollment, a fasting blood sample was drawn from each participant for clinical evaluation and to determine their HBV markers, including HBsAg, HBcAb, HBsAb, and antibody to hepatitis C virus (anti-HCV). Out of the 1607 participants, 1382 individuals who were positive for at least one of the three HBV markers and negative for anti-HCV were eligible for the genetic association study (Figure 1). Among 249 HBsAg-positive subjects, 193 had DNA samples for genotyping and were included as cases. The controls were 495 subjects randomly selected from those who had DNA samples and tested negative for HBsAg but positive for HBcAb and/or HBsAb (Figure 1).
4.2. Serologic Testing
In the study, the serostatus of HBsAg and HBcAb was determined using microparticle enzyme immunoassays (MEIA) with commercial kits AxSYN HBsAg (V2) and CORE 2.0 (Abbott Diagnostics, North Chicago, IL, USA), respectively. The titer of HBsAb in the blood samples was quantified by a commercial kit, AxSYN AUSAB (Abbott Diagnostics), with a detection limit of 1.0 mIU/mL. HBeAg, HBeAb, and anti-HCV were assayed by chemiluminescent microparticle immunoassay (CMIA) with commercial kits ARCHITECT HBeAg, Anti-HBe, and Anti-HCV (Abbott Core Laboratories, North Chicago, Illinois A, USA), respectively.
In the study, all subjects were born before 1972 and were 12 or older before the implementation of the universal HB vaccination program in Taiwan. It can be assumed that none of them received the HB vaccination. Accordingly, HBV infection was defined as a positive result for any of the three HBV markers (HBsAg, HBsAb, and HBcAb). HBV chronic carriers were defined as subjects who were seropositive for HBsAg, regardless of the serostatus of HBsAb or HBcAb. HBV non-carriers were defined as subjects who were seropositive for either HBsAb or HBcAb but negative for HBsAg.
4.3. SNP Selections
We searched the GWAS catalog [16] and identified 4 GWAS on responses to HB vaccines [11,12,13,14]. These studies reported 18 SNPs with a p-value less than 10−6, including rs12527394, rs9267665, rs3132969, rs9268202, rs4248166, rs3135363, rs2395179, rs9268831, rs477515, rs34039593, rs9273062, rs7745040, rs1015166, rs9277176, rs9277356, rs9277464, rs9277535, and rs9277549. Additionally, our previous GWAS study identified 3 SNPs (rs35953215, rs3830066, and rs7770370) that showed promising associations with responses to HB vaccines [10].
We used the Ensemble Genome Browser [29] to retrieve the linkage disequilibrium (LD) data of these candidate SNPs in the 1000 Human Genome Project Phase 3 Southern Han Chinese [19]. The LD data show that there are close linkages between rs3132969 and rs9268202 (r2 = 0.83), between rs4248166 and rs2395179 (r2 = 0.86), and among rs9277356, rs9277464, rs9277535, and rs9277549 (all pairwise r2 > 0.94). Our previous GWAS study also showed that SNP rs35953215 is completely linked, with an r2 of 1.00, with rs3830066 [10]. To reduce the influence of multiple comparisons, we used the SNP with the lowest p-value in each LD block as the candidate SNP, resulting in 15 independent SNPs (rs12527394, rs9267665, rs9268202, rs3135363, rs2395179, rs9268831, rs477515, rs34039593, rs9273062, rs7745040, rs1015166, rs9277176, rs3830066, rs9277356, and rs7770370) eligible for the genetic association study.
SNPs rs3135363, rs9268831, rs34039593, rs1015166, rs3830066, and rs7770370 were the designed SNPs of the Axiom CHB1 plate (Thermo Fisher Scientific, Waltham, MA, USA). For the other SNPs, we used their highest and nearest LD SNPs as surrogates. The surrogate SNPs for rs12527394, rs9268202, rs2395179, rs477515, rs7745040, rs9277176, and rs9277356 were rs4947302, rs9268176, rs3129846, rs614348, rs6457620, rs3097662, and 9277535, respectively. SNPs rs9267665 and rs9273062 were excluded due to the lack of a suitable surrogate. As a result, 13 SNPs were eligible for the genetic association study. Information about these SNPs and the characteristics of the typed SNPs are shown in Table 6.

Table 6.
Basic characteristics of candidate and tested SNPs.
4.4. Genotyping
The genomic DNA of each subject was extracted from EDTA-containing whole blood samples using a semi-automated extraction system, Smart LabAssist (Taiwan Advanced Nanotech Inc., Tau-Yuan County, Taiwan), with a TANBead Blood DNA plate (Taiwan Advanced Nanotech Inc.). The genotyping of all samples was performed at the National Center for Genomic Medicine, Academia Sinica, Taiwan, using the Axiom CHB1 array plate.
The genotype information for the 13 tested SNPs in HBV carriers and non-carriers is displayed in Table 2. The eligibility of SNPs for association analyses was a call rate >95%, a p-value of the Hardy–Weinberg equilibrium (HWE) test in the controls >0.001, and a minor allele frequency >3%.
4.5. Statistical Analyses
We used Pearson’s chi-square test to assess whether there was a significant difference in the genotype distributions of the candidate SNPs between HBV carriers and non-carrier controls. All significant SNPs were subject to analysis for their genotypic effects by four models, including major allele dominant, minor allele dominant, co-dominant, and additive. The most significant genotypic effect model of each significant SNP was subject to multivariable analyses. We used unconditional logistic regression to calculate crude and adjusted odds ratios (OR) and their 95% confidence intervals (CI) of chronic HBV infection for specific alleles or genotypes. The significance level of this study was set at 0.05 divided by the number of candidate SNPs. We further generated multi-locus genetic protective scores (GPSs) by summing the number of protective alleles or genotypes for each individual and then assessed their associations with chronic HBV infection. All statistical analyses were performed using SAS 9.4 (SAS Institute Inc., Cary, NC, USA).
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms24119741/s1.
Author Contributions
Conceptualization, T.-W.W., C.-L.C., C.-F.C. and L.-Y.W.; methodology, T.-W.W. and L.-Y.W.; validation, L.-Y.W.; formal analysis, L.-Y.W.; investigation, T.-W.W., C.-L.C., C.-F.C. and L.-Y.W.; resources, L.-Y.W.; data curation, T.-W.W. and C.-F.C.; writing—original draft preparation, T.-W.W. and L.-Y.W.; writing—review and editing, C.-L.C. and C.-F.C.; visualization, L.-Y.W.; supervision, L.-Y.W.; project administration, L.-Y.W.; funding acquisition, L.-Y.W. All authors have read and agreed to the published version of the manuscript.
Funding
This work was funded by research grants from the Council of Science and Technology of Taiwan (MOST 108-2314-B-715-005-MY3 and MOST 111-2314-B-715-007) and MacKay Medical College (MMC-RD-110-1B-P010 and MMC-RD-111-1B-P007). The funding agencies played no role in the research.
Institutional Review Board Statement
The study was conducted in accordance with the Declaration of Helsinki and approved by the Institutional Review Board of Mackay Memorial Hospital (No. 14MMHIS075, granted date: 23 May 2014).
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The datasets used and/or analyzed in the current study are available from the corresponding author (Li-Yu Wang) on reasonable request.
Acknowledgments
We thank the staff in the district health stations of Sanzhi District, Tamsui District, and Shimen District, New Taipei City, for their administrative support.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study.
References
- Polaris Observatory Collaborators. Global prevalence, treatment, and prevention of hepatitis B virus infection in 2016: A modelling study. Lancet Gastroenterol. Hepatol. 2018, 3, 383–403. [Google Scholar] [CrossRef]
- Chen, C.J.; Yang, H.I.; Su, J.U.N.; Jen, C.L.; You, S.L.; Lu, S.N.; Huang, G.T.; Iloeje, U.H.; Reveal-HBV Study Group. Risk of hepatocellular carcinoma across a biological gradient of serum hepatitis B virus DNA level. JAMA 2006, 295, 65–73. [Google Scholar] [CrossRef]
- Lee, M.H.; Yang, H.I.; Liu, J.; Batrla-Utermann, R.; Jen, C.L.; Iloeje, U.H.; Lu, S.N.; You, S.L.; Wang, L.Y.; Chen, C.J.; et al. Prediction models of long-term cirrhosis and hepatocellular carcinoma risk in chronic hepatitis B patients: Risk scores integrating host and virus profiles. Hepatology 2013, 58, 546–554. [Google Scholar] [CrossRef] [PubMed]
- GBD 2017 Causes of Death Collaborators. Global, regional, and national age-sex-specific mortality for 282 causes of death in 195 countries and territories, 1980–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1736–1788. [Google Scholar] [CrossRef] [PubMed]
- GBD 2017 DALYs and HALE Collaborators. Global, regional, and national disability-adjusted life-years (DALYs) for 359 diseases and injuries and healthy life expectancy (HALE) for 195 countries and territories, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1859–1922. [Google Scholar] [CrossRef]
- Chen, H.L.; Chang, M.H.; Ni, Y.H.; Hsu, H.Y.; Lee, P.L.; Lee, C.Y.; Chen, D.S. Seroepidemiology of hepatitis B virus infection in children: Ten years of mass vaccination in Taiwan. JAMA 1996, 276, 906–908. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.W.; Lin, H.H.; Wang, L.Y. Chronic hepatitis B infection in adolescents who received primary infantile vaccination. Hepatology 2013, 57, 37–45. [Google Scholar] [CrossRef]
- Chang, M.H.; You, S.L.; Chen, C.J.; Liu, C.J.; Lai, M.W.; Wu, T.C.; Wu, S.F.; Lee, C.M.; Yang, S.S.; Chu, H.C.; et al. Long-term effects of hepatitis B immunization of infants in preventing liver cancer. Gastroenterology 2016, 151, 472–480. [Google Scholar] [CrossRef]
- Galles, N.C.; Liu, P.Y.; Updike, R.L.; Fullman, N.; Nguyen, J.; Rolfe, S.; Sbarra, A.N.; Schipp, M.F.; Marks, A.; Abady, G.G.; et al. Measuring routine childhood vaccination coverage in 204 countries and territories, 1980-2019: A systematic analysis for the Global Burden of Disease Study 2020, Release 1. Lancet 2021, 398, 503–521. [Google Scholar]
- Wu, T.W.; Chen, C.F.; Lai, S.K.; Lin, H.H.; Chu, C.C.; Wang, L.Y. SNP rs7770370 in HLA-DPB1 loci as a major genetic determinant of response to booster hepatitis B vaccination: Results of a genome-wide association study. J. Gastroenterol. Hepatol. 2015, 30, 891–899. [Google Scholar] [CrossRef]
- Png, E.; Thalamuthu, A.; Ong, R.T.; Snippe, H.; Boland, G.J.; Seielstad, M. A genome-wide association study of hepatitis B vaccine response in an Indonesian population reveals multiple independent risk variants in the HLA region. Hum. Mol. Genet. 2011, 20, 3893–3898. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Zhang, L.; Zhang, W.; Wu, X.; Li, Y.; Yan, B.; Zhu, X.; Liu, X.; Yang, C.; Xu, J.; et al. A genome-wide association study identifies polymorphisms in the HLA-DR region associated with non-response to hepatitis B vaccination in Chinese Han populations. Hum. Mol. Genet. 2014, 23, 2210–2219. [Google Scholar] [CrossRef] [PubMed]
- Nishida, N.; Sugiyama, M.; Sawai, H.; Nishina, S.; Sakai, A.; Ohashi, J.; Khor, S.S.; Kakisaka, K.; Tsuchiura, T.; Hino, K.; et al. Key HLA-DRB1-DQB1 haplotypes and role of the BTNL2 gene for response to a hepatitis B vaccine. Hepatology 2018, 68, 848–858. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.; Roh, E.Y.; Park, B.; Lee, Y.; Shin, S.; Yoon, J.H.; Song, E.Y. GWAS identifying HLA-DPB1 gene variants associated with responsiveness to hepatitis B virus vaccination in Koreans: Independent association of HLA-DPB1*04:02 possessing rs1042169 G-rs9277355 C-rs9277356 A. J. Viral Hepat. 2019, 26, 1318–1329. [Google Scholar] [CrossRef]
- Wu, T.W.; Chan, H.L.; Hung, C.L.; Lu, I.J.; Wang, S.D.; Wang, S.W.; Wu, Y.J.; Wang, L.Y.; Yeh, H.I.; Wei, Y.H. Differential patterns of effects of age and sex on metabolic syndrome in Taiwan: Implication for the inadequate internal consistency of the current criteria. Diabetes Res. Clin. Pract. 2014, 105, 239–244. [Google Scholar] [CrossRef]
- Buniello, A.; MacArthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef]
- Zhang, Z.; Wang, C.; Liu, Z.; Zou, G.; Li, J.; Lu, M. Host Genetic Determinants of Hepatitis B Virus Infection. Front. Genet. 2019, 10, 696. [Google Scholar] [CrossRef]
- Roh, E.Y.; Yoon, J.H.; In, J.W.; Lee, N.; Shin, S.; Song, E.Y. Association of HLA-DP variants with the responsiveness to hepatitis B virus vaccination in Korean infants. Vaccine 2016, 34, 2602–2607. [Google Scholar] [CrossRef]
- Yates, A.; Akanni, W.; Amode, M.R.; Barrell, D.; Billis, K.; Carvalho-Silva, D.; Cummins, C.; Clapham, P.; Fitzgerald, S.; Gil, L.; et al. Ensembl 2016. Nucleic Acids Res. 2016, 44, D710–D716. [Google Scholar] [CrossRef]
- Kamatani, Y.; Wattanapokayakit, S.; Ochi, H.; Kawaguchi, T.; Takahashi, A.; Hosono, N.; Kubo, M.; Tsunoda, T.; Kamatani, N.; Kumada, H.; et al. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat. Genet. 2009, 41, 591–595. [Google Scholar] [CrossRef]
- Mbarek, H.; Ochi, H.; Urabe, Y.; Kumar, V.; Kubo, M.; Hosono, N.; Takahashi, A.; Kamatani, Y.; Miki, D.; Abe, H.; et al. A genome-wide association study of chronic hepatitis B identified novel risk locus in a Japanese population. Hum. Mol. Genet. 2011, 20, 3884–3892. [Google Scholar]
- Hu, L.; Zhai, X.; Liu, J.; Chu, M.; Pan, S.; Jiang, J.; Zhang, Y.; Wang, H.; Chen, J.; Shen, H.; et al. Genetic variants in human leukocyte antigen/DP-DQ influence both hepatitis B virus clearance and hepatocellular carcinoma development. Hepatology 2012, 55, 1426–1431. [Google Scholar] [CrossRef] [PubMed]
- Nishida, N.; Sawai, H.; Matsuura, K.; Sugiyama, M.; Ahn, S.H.; Park, J.Y.; Hige, S.; Kang, J.-H.; Suzuki, K.; Kurosaki, M.; et al. Genome-wide association study confirming association of HLA-DP with protection against chronic hepatitis B and viral clearance in Japanese and Korean. PLoS ONE 2012, 7, e39175. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.-W.; Fann, C.S.-J.; Su, W.-H.; Wang, Y.C.; Weng, C.C.; Yu, C.-J.; Hsu, C.-L.; Hsieh, A.-R.; Chien, R.-N.; Chu, C.-M.; et al. A genome-wide association study on chronic HBV infection and its clinical progression in male Han-Taiwanese. PLoS ONE 2014, 9, e99724. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Cheng, Y.-j.; Cheng, M.-l.; Yao, Y.-m.; Zhang, Q.; Zhao, X.-k.; Liu, H.-j.; Hu, Y.-x.; Mu, M.; Wang, B.; et al. Quantitative assessment of common genetic variations in HLA-DP with hepatitis B virus infection, clearance and hepatocellular carcinoma development. Sci. Rep. 2015, 5, 14933. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Huang, Y.H.; Liao, S.F.; Khor, S.S.; Lin, Y.J.; Chen, H.Y.; Chang, Y.H.; Huang, Y.H.; Lu, S.N.; Lee, H.W.; Ko, W.Y.; et al. Large-scale genome-wide association study identifies HLA class II variants associated with chronic HBV infection: A study from Taiwan Biobank. Aliment. Pharmacol. Ther. 2020, 52, 682–691. [Google Scholar] [CrossRef]
- Hamilton, E.; Yang, L.; Mentzer, A.J.; Guo, Y.; Chen, Y.; Lv, J.; Fletcher, R.; Wright, N.; Lin, K.; Walters, R.; et al. Conventional and genetic risk factors for chronic Hepatitis B virus infection in a community-based study of 0.5 million Chinese adults. Sci. Rep. 2022, 12, 12075. [Google Scholar] [CrossRef]
- Cunningham, F.; Allen, J.E.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Austine-Orimoloye, O.; Azov, A.G.; Barnes, I.; Bennett, R.; et al. Ensembl 2022. Nucleic Acids Res. 2022, 50, D988–D995. [Google Scholar] [CrossRef]
- Schmiedel, B.J.; Singh, D.; Madrigal, A.; Valdovino-Gonzalez, A.G.; White, B.M.; Zapardiel-Gonzalo, J.; Ha, B.; Altay, G.; Greenbaum, J.A.; McVicker, G.; et al. Impact of genetic polymorphisms on human immune cell gene expression. Cell 2018, 175, 1701–1715. [Google Scholar] [CrossRef]
- Wang, L.Y.; Chen, C.F.; Wu, T.W.; Lai, S.K.; Chu, C.C.; Lin, H.H. Response to hepatitis B vaccination is co-determined by HLA-DPA1 and -DPB1. Vaccine 2019, 37, 6435–6440. [Google Scholar] [CrossRef]
- Kao, J.H. Molecular epidemiology of hepatitis B virus. Korean J. Intern. Med. 2011, 26, 255–261. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).