Novel Insights to Assess Climate Resilience in Goats Using a Holistic Approach of Skin-Based Advanced NGS Technologies
Abstract
:1. Introduction
2. Results
2.1. Simulation of Comfort and Heat-Stress Environment within the Climate Chambers
2.2. Hair Fiber Analysis
2.3. Hair Cortisol Estimation
2.4. Hair Follicle qPCR Analysis
2.5. Sweating Rate and Active Sweat Gland Estimation
2.6. Skin Histology
2.7. Infrared Thermography of Caprine Skin
2.8. Skin 16S rRNA V3-V4 Metagenomics
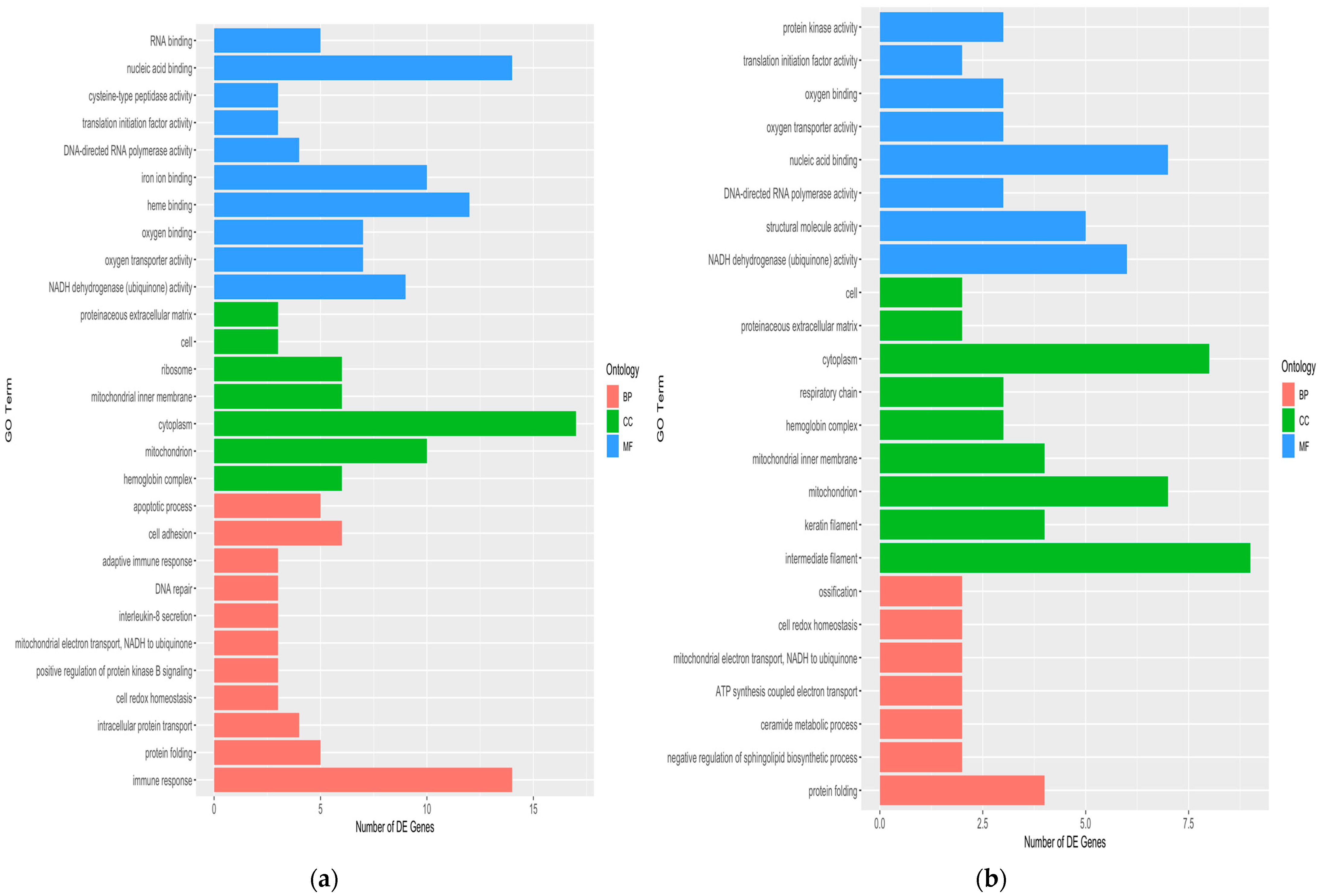
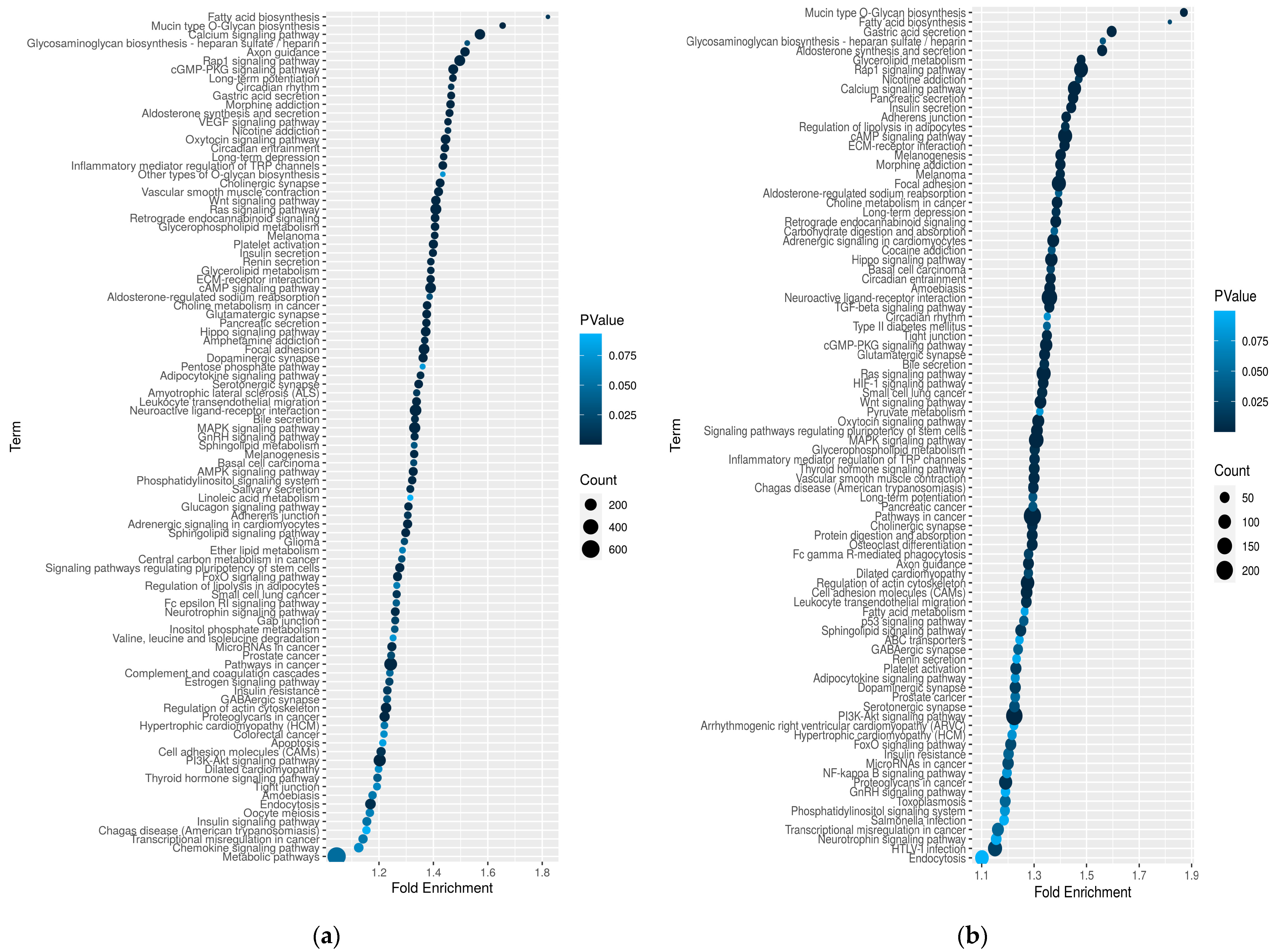
2.9. Skin Transcriptomics Analysis
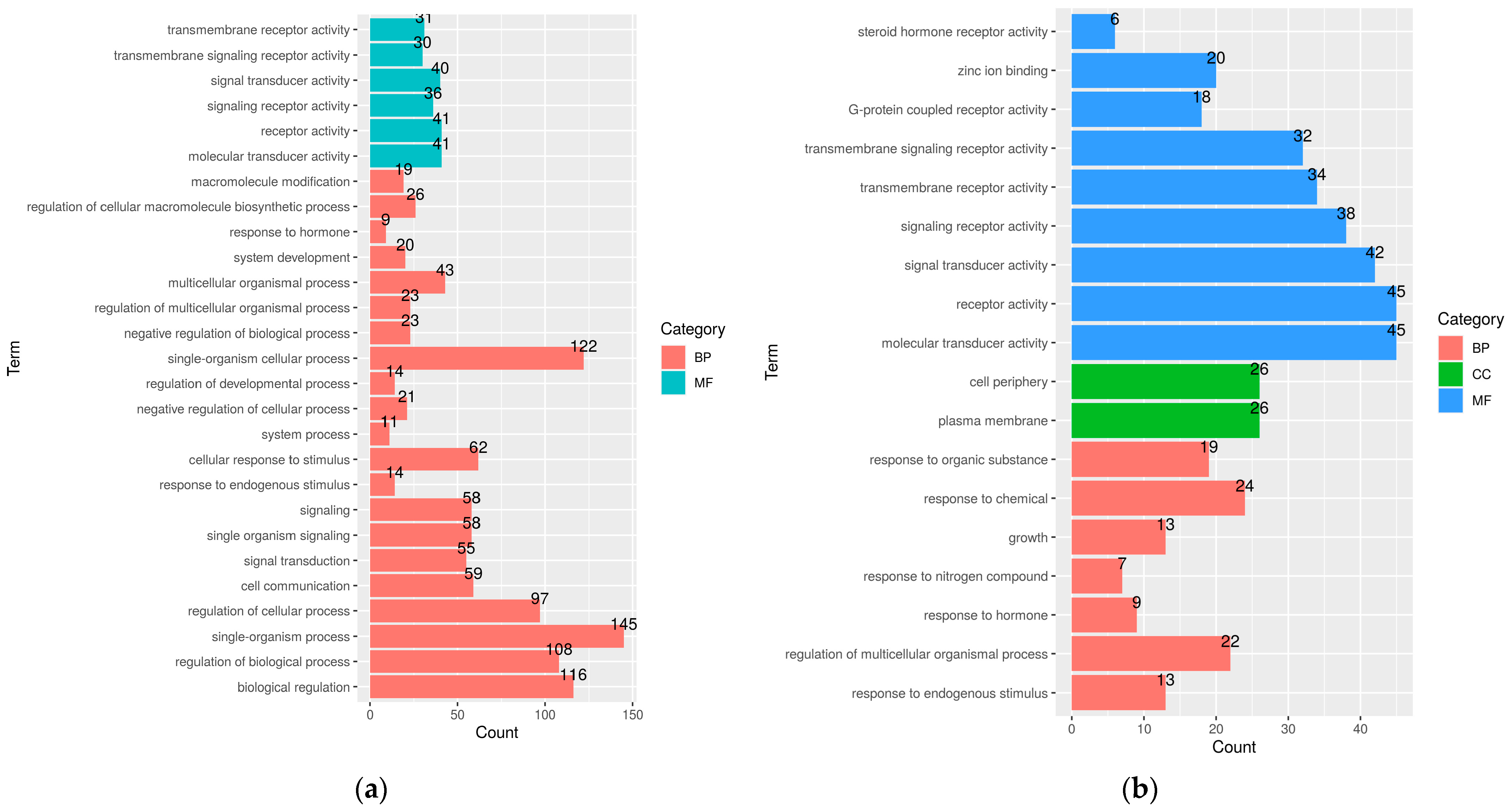
2.10. Skin Bisulfite Sequencing
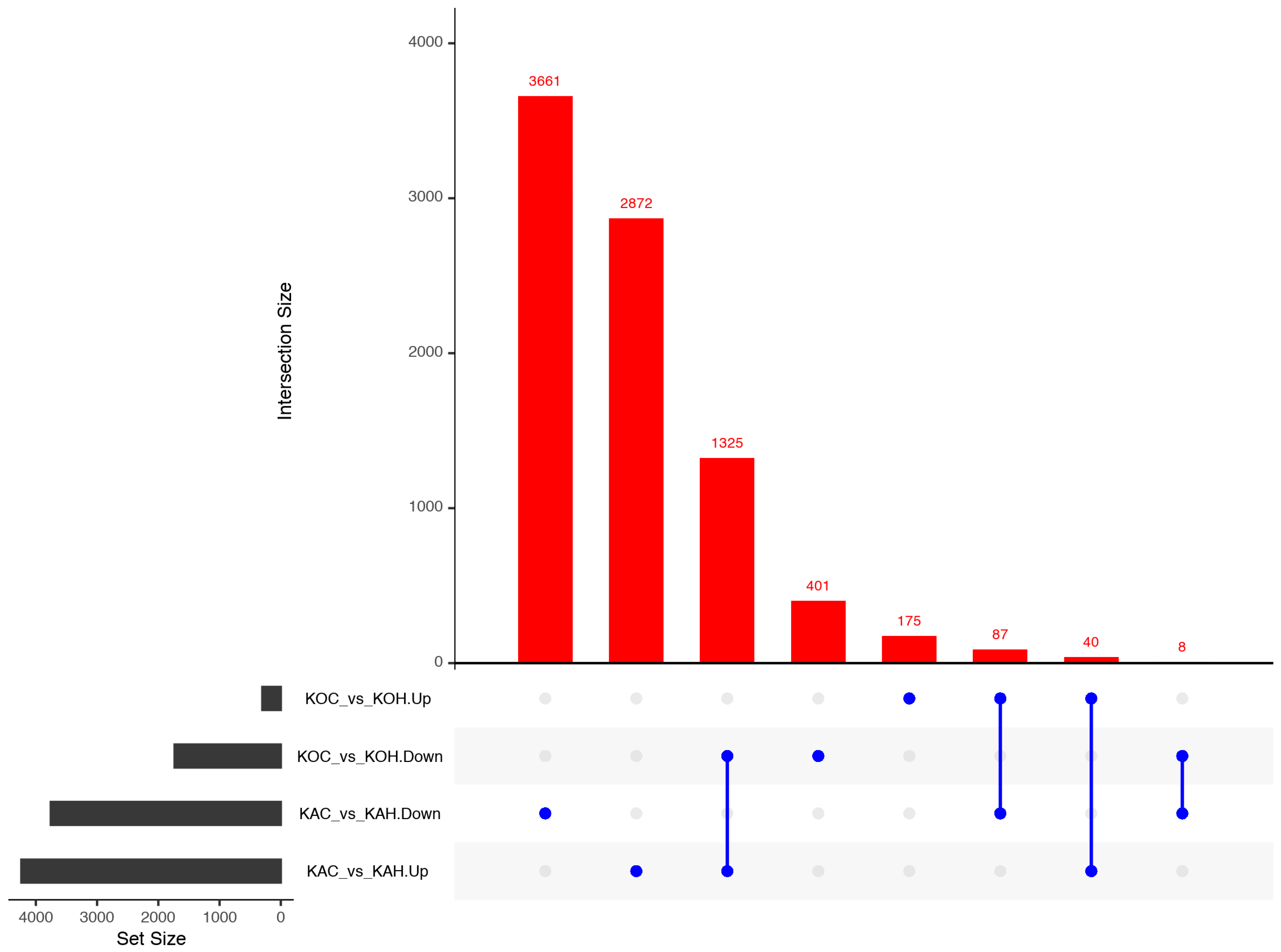
2.11. Linking Skin Transcriptomics and Epigenetics Analysis
3. Discussion
4. Materials and Methods
4.1. Location of the Study
4.2. Animal Details
4.3. Experimental Design
4.4. Hair Fiber Analysis
4.5. Hair Cortisol Estimation
4.6. Hair Follicle qPCR Analysis
4.7. Sweating Rate and Active Sweat Gland Estimation
4.8. Skin Histology
4.9. Infrared Thermography of Caprine Skin
4.10. Skin 16S rRNA V3-V4 Metagenomics
4.11. Skin Transcriptomics Analysis
4.12. Skin Bisulfite Sequencing
4.13. Statiscial Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sequence Availability
References
- Thornton, P.; Nelson, G.; Mayberry, D.; Herrero, M. Increases in Extreme Heat Stress in Domesticated Livestock Species during the Twenty-First Century. Glob. Chang. Biol. 2021, 27, 5762–5772. [Google Scholar] [CrossRef] [PubMed]
- Sejian, V.; Bhatta, R.; Gaughan, J.B.; Dunshea, F.R.; Lacetera, N. Review: Adaptation of Animals to Heat Stress. Animal 2018, 12, s431–s444. [Google Scholar] [CrossRef] [Green Version]
- Carvajal, M.A.; Alaniz, A.J.; Gutiérrez-Gómez, C.; Vergara, P.M.; Sejian, V.; Bozinovic, F. Increasing Importance of Heat Stress for Cattle Farming under Future Global Climate Scenarios. Sci. Total Environ. 2021, 801, 149661. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, J.; Mutua, J.Y.; Notenbaert, A.M.O.; Marshall, K.; Butterbach-Bahl, K. Heat Stress Will Detrimentally Impact Future Livestock Production in East Africa. Nat. Food 2021, 2, 88–96. [Google Scholar] [CrossRef]
- Sejian, V.; Silpa, M.V.; Reshma Nair, M.R.; Devaraj, C.; Devapriya, A.; Ramachandran, N.; Awachat, V.B.; Shashank, C.G.; Fonsêca, V.F.C.; Bhatta, R. Goat as the Ideal Future Climate Resilient Animal Model. In Climate Change and Livestock Production: Recent Advances and Future Perspectives; Sejian, V., Chauhan, S.S., Devaraj, C., Malik, P.K., Bhatta, R., Eds.; Springer: Singapore, 2021; pp. 279–292. ISBN 978-981-16-9836-1. [Google Scholar]
- Aleena, J.; Sejian, V.; Krishnan, G.; Bagath, M.; Pragna, P.; Bhatta, R. Heat Stress Impact on Blood Biochemical Response and Plasma Aldosterone Level in Three Different Indigenous Goat Breeds. J. Anim. Behav. Biometeorol. 2020, 8, 266–275. [Google Scholar] [CrossRef]
- Aleena, J.; Sejian, V.; Bagath, M.; Krishnan, G.; Beena, V.; Bhatta, R. Resilience of Three Indigenous Goat Breeds to Heat Stress Based on Phenotypic Traits and PBMC HSP70 Expression. Int. J. Biometeorol. 2018, 62, 1995–2005. [Google Scholar] [CrossRef]
- Archana, P.R.; Sejian, V.; Ruban, W.; Bagath, M.; Krishnan, G.; Aleena, J.; Manjunathareddy, G.B.; Beena, V.; Bhatta, R. Comparative Assessment of Heat Stress Induced Changes in Carcass Traits, Plasma Leptin Profile and Skeletal Muscle Myostatin and HSP70 Gene Expression Patterns between Indigenous Osmanabadi and Salem Black Goat Breeds. Meat Sci. 2018, 141, 66–80. [Google Scholar] [CrossRef] [PubMed]
- Devapriya, A.; Sejian, V.; Ruban, W.; Devaraj, C.; Spandan, P.V.; Silpa, M.V.; Reshma Nair, M.R.; Nameer, P.O.; Bhatta, R. Analysis of Carcass Traits and Quantitative Expression Patterns of Different Meat Quality Governing Genes during Heat Stress Exposure in Indigenous Goats. Food Chem. Mol. Sci. 2021, 3, 100052. [Google Scholar] [CrossRef]
- Contreras-Jodar, A.; Salama, A.A.; Hamzaoui, S.; Vailati-Riboni, M.; Caja, G.; Loor, J.J. Effects of Chronic Heat Stress on Lactational Performance and the Transcriptomic Profile of Blood Cells in Lactating Dairy Goats. J. Dairy Res. 2018, 85, 423–430. [Google Scholar] [CrossRef]
- Rashamol, V.P.; Sejian, V.; Bagath, M.; Krishnan, G.; Beena, V.; Bhatta, R. Effect of Heat Stress on the Quantitative Expression Patterns of Different Cytokine Genes in Malabari Goats. Int. J. Biometeorol. 2019, 63, 1005–1013. [Google Scholar] [CrossRef]
- McDowell, R.E. Improvement of Livestock Production in Warm Climates; WH Freeman and Co.: San Fransisco, CA, USA, 1972. [Google Scholar]
- Ribeiro, N.L.; Germano Costa, R.; Pimenta Filho, E.C.; Ribeiro, M.N.; Bozzi, R. Effects of the Dry and the Rainy Season on Endocrine and Physiologic Profiles of Goats in the Brazilian Semi-Arid Region. Ital. J. Anim. Sci. 2018, 17, 454–461. [Google Scholar] [CrossRef] [Green Version]
- da Silva, R.G.; Scala, N.L., Jr.; Pocay, P.L.B. Transmission of Ultraviolet Radiation Through the Haircoat and the Skin of Cattle. R. Bras. Zootec. 2001, 30, 1939–1947. [Google Scholar] [CrossRef] [Green Version]
- Maia, A.S.C.; daSilva, R.G.; Battiston Loureiro, C.M. Sensible and Latent Heat Loss from the Body Surface of Holstein Cows in a Tropical Environment. Int. J. Biometeorol. 2005, 50, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Dulude-de Broin, F.; Côté, S.D.; Whiteside, D.P.; Mastromonaco, G.F. Faecal Metabolites and Hair Cortisol as Biological Markers of HPA-Axis Activity in the Rocky Mountain Goat. Gen. Comp. Endocrinol. 2019, 280, 147–157. [Google Scholar] [CrossRef]
- Hecker, J.G.; McGarvey, M. Heat Shock Proteins as Biomarkers for the Rapid Detection of Brain and Spinal Cord Ischemia: A Review and Comparison to Other Methods of Detection in Thoracic Aneurysm Repair. Cell Stress Chaperones 2011, 16, 119–131. [Google Scholar] [CrossRef] [Green Version]
- Dangi, S.S.; Gupta, M.; Nagar, V.; Yadav, V.P.; Dangi, S.K.; Shankar, O.; Chouhan, V.S.; Kumar, P.; Singh, G.; Sarkar, M. Impact of Short-Term Heat Stress on Physiological Responses and Expression Profile of HSPs in Barbari Goats. Int. J. Biometeorol. 2014, 58, 2085–2093. [Google Scholar] [CrossRef] [PubMed]
- Afsal, A.; Bagath, M.; Sejian, V.; Krishnan, G.; Beena, V.; Bhatta, R. Effect of Heat Stress on HSP70 Gene Expression Pattern in Different Vital Organs of Malabari Goats. Biol. Rhythm Res. 2021, 52, 380–394. [Google Scholar] [CrossRef]
- Kim, W.S.; Nejad, J.G.; Peng, D.Q.; Jung, U.S.; Kim, M.J.; Jo, Y.H.; Jo, J.H.; Lee, J.S.; Lee, H.G. Identification of Heat Shock Protein Gene Expression in Hair Follicles as a Novel Indicator of Heat Stress in Beef Calves. Animal 2020, 14, 1502–1509. [Google Scholar] [CrossRef] [PubMed]
- Madhusoodan, A.P.; Bagath, M.; Sejian, V.; Krishnan, G.; Rashamol, V.P.; Savitha, S.T.; Awachat, V.B.; Bhatta, R. Summer Season Induced Changes in Quantitative Expression Patterns of Different Heat Shock Response Genes in Salem Black Goats. Trop. Anim. Health Prod. 2020, 52, 2725–2730. [Google Scholar] [CrossRef]
- Ferreira, J.; Silveira, R.M.F.; de Sousa, J.E.R.; Façanha, D.A.E. Locally Adapted Goats Efficiently Gain and Lose Heat in an Equatorial Semi-Arid Environment. Int. J. Biometeorol. 2020, 64, 1777–1782. [Google Scholar] [CrossRef]
- Maurya, V.P.; Sejian, V.; Kumar, D.; Naqvi, S.M.K. Impact of Heat Stress, Nutritional Stress and Their Combinations on the Adaptive Capability of Malpura Sheep under Hot Semi-Arid Tropical Environment. J. Anim. Behav. Biometeorol. 2019, 7, 31–38. [Google Scholar] [CrossRef]
- Byrd, A.L.; Belkaid, Y.; Segre, J.A. The Human Skin Microbiome. Nat. Rev. Microbiol. 2018, 16, 143–155. [Google Scholar] [CrossRef]
- Morenikeji, O.B.; Ajayi, O.O.; Peters, S.O.; Mujibi, F.D.; De Donato, M.; Thomas, B.N.; Imumorin, I.G. RNA-Seq Profiling of Skin in Temperate and Tropical Cattle. J. Anim. Sci. Technol. 2020, 62, 141–158. [Google Scholar] [CrossRef] [Green Version]
- Acharya, R.M.; Gupta, U.D.; Sehgal, J.P.; Singh, M. Coat Characteristics of Goats in Relation to Heat Tolerance in the Hot Tropics. Small Rumin. Res. 1995, 18, 245–248. [Google Scholar] [CrossRef]
- Stuart-Fox, D.; Newton, E.; Clusella-Trullas, S. Thermal Consequences of Colour and Near-Infrared Reflectance. Philos. Trans. R. Soc. B 2017, 372, 20160345. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.-J. HRI Stress Signaling and HbF Production. Blood 2020, 135, 2113–2114. [Google Scholar] [CrossRef]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The Integrated Stress Response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef] [Green Version]
- Hjerpe, R.; Bett, J.S.; Keuss, M.J.; Solovyova, A.; McWilliams, T.G.; Johnson, C.; Sahu, I.; Varghese, J.; Wood, N.; Wightman, M.; et al. UBQLN2 Mediates Autophagy-Independent Protein Aggregate Clearance by the Proteasome. Cell 2016, 166, 935–949. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Downs, C.A.; Heckathorn, S.A. The mitochondrial small heat-shock protein protects NADH: Ubiquinone oxidoreductase of the electron transport chain during heat stress in plants. FEBS Lett. 1998, 430, 246–250. [Google Scholar] [CrossRef] [Green Version]
- Barros, S.P.; Offenbacher, S. Epigenetics: Connecting Environment and Genotype to Phenotype and Disease. J. Dent. Res. 2009, 88, 400–408. [Google Scholar] [CrossRef] [PubMed]
- Del Corvo, M.; Lazzari, B.; Capra, E.; Zavarez, L.; Milanesi, M.; Utsunomiya, Y.T.; Utsunomiya, A.T.H.; Stella, A.; de Paula Nogueira, G.; Garcia, J.F.; et al. Methylome Patterns of Cattle Adaptation to Heat Stress. Front. Genet. 2021, 12, 633132. [Google Scholar] [CrossRef]
- Ahlawat, S.; Arora, R.; Sharma, R.; Sharma, U.; Kaur, M.; Kumar, A.; Singh, K.V.; Singh, M.K.; Vijh, R.K. Skin Transcriptome Profiling of Changthangi Goats Highlights the Relevance of Genes Involved in Pashmina Production. Sci. Rep. 2020, 10, 6050. [Google Scholar] [CrossRef] [Green Version]
- Peñagaricano, F.; Zorrilla, P.; Naya, H.; Robello, C.; Urioste, J.I. Gene Expression Analysis Identifies New Candidate Genes Associated with the Development of Black Skin Spots in Corriedale Sheep. J. Appl. Genet. 2012, 53, 99–106. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Schleger, A.V.; Turner, H.G. Sweating Rates of Cattle in the Field and Their Reaction to Diurnal and Seasonal Changes. Aust. J. Agric. Res. 1965, 16, 92–106. [Google Scholar] [CrossRef]
- Gagnon, D.; Ganio, M.S.; Lucas, R.A.I.; Pearson, J.; Crandall, C.G.; Kenny, G.P. Modified Iodine-Paper Technique for the Standardized Determination of Sweat Gland Activation. J. Appl. Physiol. 2012, 112, 1419–1425. [Google Scholar] [CrossRef] [PubMed]
- Rajesh, T.; Anthony, T.; Saranya, S.; Pushpam, P.L.; Gunasekaran, P. Functional Characterization of a New Holin-like Antibacterial Protein Coding Gene Tmp1 from Goat Skin Surface Metagenome. Appl. Microbiol. Biotechnol. 2011, 89, 1061–1073. [Google Scholar] [CrossRef]




| Variable | Kanni Aadu | Kodi Aadu | |||
|---|---|---|---|---|---|
| KAC | KAH | KOC | KOH | ||
| Hair characteristics | Fiber diameter (µm) | 68.00 ± 2.43 a | 64.33 ± 1.49 a | 82.93 ± 3.18 b | 84.24 ± 3.97 b |
| p-value: 0.00 | |||||
| Staple length (cm) | 3.77 ± 0.03 b | 3.53 ± 0.12 ab | 3.72 ± 0.27 b | 3.11 ± 0.17 a | |
| p-value: 0.05 | |||||
| Hair cortisol | Hair cortisol (ng/mL) | 6.89 ± 0.16 a | 6.67 ± 0.16 a | 6.92 ± 0.22 a | 6.62 ± 0.16 a |
| p-value: 0.54 | |||||
| Hair follicle qPCR | HSP70 | 1.00 ± 0.08 a | 0.99 ± 0.12 a | 1.00 ± 0.13 a | 0.52 ± 0.19 b |
| p-value: 0.89 | p-value: 0.02 | ||||
| HSP90 | 1.00 ± 0.11 a | 0.63 ± 0.07 b | 1.00 ± 0.01 a | 0.26 ± 0.10 b | |
| p-value: 0.01 | p-value: 0.00 | ||||
| HSP110 | 1.00 ± 0.09 a | 0.98 ± 0.06 a | 1.00 ± 0.15 a | 0.68 ± 0.07 b | |
| p-value: 0.81 | p-value: 0.05 | ||||
| Sweating | Sweating rate (g/m2/h) | 0.00 ± 0.00 a | 2.80 ± 0.37 b | 0.00 ± 0.00 a | 2.82 ± 0.37 b |
| p-value: 0.00 | |||||
| Active sweat gland measurement (No of gland/cm2) | 0.00 ± 0.00 a | 0.03 ± 0.01 b | 0.00 ± 0.00 a | 0.06 ± 0.01 b | |
| p-value: 0.00 | |||||
| Skin histometry | Epithelial height (µm) | 15.62 ± 0.23 a | 16.84 ± 0.18 b | 18.15 ± 0.49 c | 22.94 ± 0.60 d |
| p-value: 0.00 | |||||
| No. of sweat glands/cm2 | 421.67 ± 3.53 a | 422.75 ± 1.83 ab | 429.33 ± 2.14 bc | 431.42 ± 1.97 c | |
| p-value: 0.00 | |||||
| Skin-surface infra-red thermal imaging | Eye (°C) | 36.93 ± 0.11 a | 40.96 ± 0.12 b | 36.88 ± 0.15 a | 41.14 ± 0.09 b |
| p-value: 0.00 | |||||
| Forehead (°C) | 29.39 ± 0.10 a | 40.44 ± 0.47 b | 29.83 ± 0.24 a | 40.28 ± 0.13 b | |
| p-value: 0.00 | |||||
| Flank (°C) | 30.68 ± 0.26 a | 40.78 ± 0.22 b | 31.34 ± 0.27 a | 40.43 ± 0.13 b | |
| p-value: 0.00 | |||||
| Back (°C) | 28.73 ± 0.47 a | 40.41 ± 0.16 c | 29.63 ± 0.24 b | 39.83 ± 0.13 a | |
| p-value: 0.00 | |||||
| Front leg (°C) | 27.06 ± 0.34 a | 40.71 ± 0.25 b | 27.30 ± 0.13 a | 40.38 ± 0.20 b | |
| p-value: 0.00 | |||||
| Skin 16S rRNA V3-V4 metagenomics | Relative abundance of microbes at phylum level (%) | ||||
| Bacteroidetes | 31.86 | 33.23 | 37.10 | 31.38 | |
| Firmicutes | 21.40 | 29.00 | 28.13 | 27.76 | |
| Proteobacteria | 31.86 | 18.54 | 20.21 | 17.12 | |
| Actinobacteria | 7.67 | 12.44 | 6.02 | 17.48 | |
| Spirochaetes | 2.25 | 2.27 | 2.52 | 1.58 | |
| Fibrobacteres | 2.40 | 1.04 | 1.68 | 1.56 | |
| Verrucomicrobia | 1.26 | 1.52 | 2.30 | 1.49 | |
| TM7 | 0.61 | 1.23 | 1.04 | 0.69 | |
| Hair Characteristics | Hair Cortisol | Hair Follicle qPCR | Sweating | Skin Histometry | Skin-Surface Infra-Red Thermal Imaging | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Breed | Group | THI | FD | SL | HCC | HSP70 | HSP90 | HSP110 | SR | ASG | SEH | SG | Eye | FH | Back | Flank | L | |
| Breed | 1 | |||||||||||||||||
| Group | 0.89 ** | 1 | ||||||||||||||||
| THI | 0.00 | 0.45 * | 1 | |||||||||||||||
| FD | 0.79 ** | 0.69 ** | −0.05 | 1 | ||||||||||||||
| SL | −0.26 | −0.43 * | −0.46 * | −0.27 | 1 | |||||||||||||
| HCC | −0.01 | −0.15 | −0.31 | 0.23 | 0.01 | 1 | ||||||||||||
| HSP70 | 0.84 ** | 0.90 ** | 0.34 | 0.68 ** | −0.43 * | −0.05 | 1 | |||||||||||
| HSP90 | 0.74 ** | 0.93 ** | 0.60 ** | 0.58 ** | −0.52 ** | −0.25 | 0.91 ** | 1 | ||||||||||
| HSP110 | 0.86 ** | 0.91 ** | 0.31 | 0.76 ** | −0.35 | −0.13 | 0.91 ** | 0.91 ** | 1 | |||||||||
| SR | 0.00 | 0.44 * | 0.99 ** | −0.07 | −0.44 * | −0.34 | 0.34 | 0.60 ** | 0.31 | 1 | ||||||||
| ASG | 0.20 | 0.55 ** | 0.82 ** | 0.23 | −0.59 ** | −0.25 | 0.50 * | 0.72 ** | 0.48 * | 0.83 ** | 1 | |||||||
| SEH | 0.69 ** | 0.83 ** | 0.48 * | 0.58 ** | −0.45 * | −0.175 | 0.78 ** | 0.88 ** | 0.83 ** | 0.47 * | 0.51 * | 1 | ||||||
| SG | 0.57 ** | 0.56 ** | 0.11 | 0.38 | −0.30 | −0.19 | 0.59 ** | 0.49 * | 0.55 ** | 0.12 | 0.14 | 0.41 * | 1 | |||||
| Eye | 0.02 | 0.46 * | 0.99 ** | −0.02 | −0.45 * | −0.28 | 0.37 | 0.62 ** | 0.33 | 0.97 ** | 0.83 ** | 0.49 * | 0.12 | 1 | ||||
| FH | 0.01 | 0.46 * | 0.99 ** | −0.04 | −0.47 * | −0.34 | 0.34 | 0.60 ** | 0.30 | 0.98 ** | 0.83 ** | 0.49 * | 0.1 | 0.99 ** | 1 | |||
| B | 0.01 | 0.46 * | 0.99 ** | −0.02 | −0.42 * | −0.28 | 0.33 | 0.59 ** | 0.29 | 0.98 ** | 0.87 ** | 0.47 * | 0.09 | 0.99 ** | 0.99 ** | 1 | ||
| Flank | 0.02 | 0.46 * | 0.99 ** | −0.02 | −0.42 * | −0.29 | 0.34 | 0.59 ** | 0.31 | 0.98 ** | 0.82 ** | 0.48 * | 0.06 | 0.99 ** | 0.99 ** | 0.997 ** | 1 | |
| L | −0.00 | 0.44 * | 0.997 ** | −0.04 | −0.45 * | −0.31 | 0.33 | 0.59 ** | 0.3 | 0.98 ** | 0.82 ** | 0.48 * | 0.10 | 0.99 ** | 0.995 ** | 0.99 ** | 0.99 ** | 1 |
| Variable | Kanni Aadu (KAC vs. KAH) | Kodi Aadu (KOC vs. KOH) | |
|---|---|---|---|
| Skin transcriptomics | DEGs | 7993 | 2036 |
| Up-regulated DEGs | 4237 | 302 | |
| Down-regulated DEGs | 3756 | 1734 | |
| Skin whole-genome bisulfite sequencing | DMR | 50,560 | 40,648 |
| Hyper-methylated DMR | 25,178 | 19,657 | |
| Hypo-methylated DMR | 25,382 | 20,991 | |
| DMG | 14,646 | 13,388 | |
| Hyper-methylated DMG | 7336 | 6507 | |
| Hypo-methylated DMG | 7310 | 6904 |
| KAC_vs_KAH | KOC_vs_KOH | ||
|---|---|---|---|
| Gene | Log2FC | Gene | Log2FC |
| EIF–ATF pathway | |||
| EIF2A | 1.775 | ||
| EIF2B1 | −1.343 | ||
| EIF2B2 | 1.398 | ||
| EIF2B4 | 2.026 | EIF2B4 | −1.337 |
| EIF2B5 | 1.547 | EIF2B5 | −1.77 |
| EIF2S1 | 1.702 | ||
| EIF2S2 | −1.416 | ||
| EIF2S3 | 1.752 | EIF2S3 | −1.493 |
| ATF4 | 2.369 | ATF4 | −1.875 |
| ATF5 | 2.519 | ATF5 | −1.367 |
| Stress-associated molecular chaperones | |||
| UBQLN2 | 1.593 | UBQLN2 | −1.437 |
| UBQLN3 | −7.974 | ||
| HSF1 | 2.078 | ||
| HSP70.1 | 3.035 | HSP70.1 | −1.492 |
| HSP90AB1 | 2.285 | ||
| HSP90B1 | 1.863 | HSP90B1 | −1.392 |
| HSBP1L1 | −2.111 | ||
| HSPA13 | −1.108 | ||
| HSPA14 | −1.344 | ||
| HSPA4 | −1.1 | ||
| HSPA5 | 2.021 | HSPA5 | −1.591 |
| HSPA8 | 2.098 | HSPA8 | −1.756 |
| HSPA9 | 1.805 | ||
| HSPB1 | 4.01 | HSPB1 | −1.898 |
| HSPB3 | −2.07 | ||
| HSPB6 | 3.04 | HSPB6 | −1.665 |
| HSPB8 | 2.799 | HSPB8 | −1.901 |
| HSPBP1 | 1.771 | ||
| HSPD1 | 1.555 | ||
| KAC_vs_KAH | KOC_vs_KOH | ||||
|---|---|---|---|---|---|
| Description | No. of DEG | p-Value | Description | No. of DEG | p-Value |
| Metabolic pathways | 499 | 0.00 | Metabolic pathways | 189 | 0.00 |
| Neuroactive ligand–receptor interaction | 124 | 0.00 | Huntington’s disease | 64 | 0.00 |
| Ribosome | 113 | 0.00 | Oxidative phosphorylation | 62 | 0.00 |
| Huntington’s disease | 109 | 0.00 | Parkinson’s disease | 61 | 0.00 |
| Biosynthesis of antibiotics | 105 | 0.00 | Alzheimer’s disease | 58 | 0.00 |
| Endocytosis | 97 | 0.02 | Biosynthesis of antibiotics | 51 | 0.00 |
| Parkinson’s disease | 93 | 0.00 | Spliceosome | 35 | 0.00 |
| Oxidative phosphorylation | 90 | 0.00 | Endocytosis | 34 | 0.03 |
| Protein processing in endoplasmic reticulum | 81 | 0.00 | RNA transport | 28 | 0.01 |
| Spliceosome | 77 | 0.00 | Protein processing in endoplasmic reticulum | 28 | 0.01 |
| RNA transport | 71 | 0.01 | Carbon metabolism | 24 | 0.00 |
| Lysosome | 54 | 0.02 | Proteasome | 23 | 0.00 |
| Carbon metabolism | 53 | 0.01 | Lysosome | 22 | 0.01 |
| Epstein–Barr virus infection | 48 | 0.05 | Cardiac muscle contraction | 20 | 0.00 |
| Retrograde endocannabinoid signaling | 44 | 0.05 | Ribosome biogenesis in eukaryotes | 18 | 0.00 |
| Antigen processing and presentation | 41 | 0.00 | Biosynthesis of amino acids | 16 | 0.00 |
| Cardiac muscle contraction | 41 | 0.00 | Pyrimidine metabolism | 16 | 0.04 |
| GABAergic synapse | 41 | 0.01 | Glutathione metabolism | 11 | 0.02 |
| Proteasome | 37 | 0.00 | Citrate cycle (TCA cycle) | 9 | 0.01 |
| Biosynthesis of amino acids | 36 | 0.01 | Pyruvate metabolism | 9 | 0.03 |
| Bile secretion | 36 | 0.02 | RNA polymerase | 8 | 0.02 |
| Arginine and proline metabolism | 29 | 0.00 | Steroid biosynthesis | 7 | 0.02 |
| Fat digestion and absorption | 27 | 0.00 | |||
| Glutathione metabolism | 26 | 0.01 | |||
| Ether lipid metabolism | 23 | 0.04 | |||
| Phototransduction | 17 | 0.00 | |||
| alpha-Linolenic acid metabolism | 16 | 0.02 | |||
| RNA polymerase | 16 | 0.04 | |||
| Citrate cycle (TCA cycle) | 16 | 0.05 | |||
| Group | Total Reads | QC Passed Reads | % Mapped Reads | % Methylated Cs | % Methylated CpG | % Methylated CHG | % Methylated CHH |
|---|---|---|---|---|---|---|---|
| KAC | 49,128,009 | 48,056,118 | 50.35 | 5.97 | 85.54 | 3.34 | 11.12 |
| KAH | 54,655,288 | 53,458,502 | 52.66 | 5.90 | 85.71 | 3.22 | 11.07 |
| KOC | 56,831,837 | 55,694,741 | 52.21 | 5.69 | 84.84 | 3.40 | 11.75 |
| KOH | 48,747,713 | 47,743,401 | 51.62 | 5.61 | 85.03 | 3.32 | 11.65 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mullakkalparambil Velayudhan, S.; Sejian, V.; Devaraj, C.; Manjunathareddy, G.B.; Ruban, W.; Kadam, V.; König, S.; Bhatta, R. Novel Insights to Assess Climate Resilience in Goats Using a Holistic Approach of Skin-Based Advanced NGS Technologies. Int. J. Mol. Sci. 2023, 24, 10319. https://doi.org/10.3390/ijms241210319
Mullakkalparambil Velayudhan S, Sejian V, Devaraj C, Manjunathareddy GB, Ruban W, Kadam V, König S, Bhatta R. Novel Insights to Assess Climate Resilience in Goats Using a Holistic Approach of Skin-Based Advanced NGS Technologies. International Journal of Molecular Sciences. 2023; 24(12):10319. https://doi.org/10.3390/ijms241210319
Chicago/Turabian StyleMullakkalparambil Velayudhan, Silpa, Veerasamy Sejian, Chinnasamy Devaraj, Gundallahalli Bayyappa Manjunathareddy, Wilfred Ruban, Vinod Kadam, Sven König, and Raghavendra Bhatta. 2023. "Novel Insights to Assess Climate Resilience in Goats Using a Holistic Approach of Skin-Based Advanced NGS Technologies" International Journal of Molecular Sciences 24, no. 12: 10319. https://doi.org/10.3390/ijms241210319







