ALK, ROS1, RET and NTRK1–3 Gene Fusions in Colorectal and Non-Colorectal Microsatellite-Unstable Cancers
Abstract
:1. Introduction
2. Results
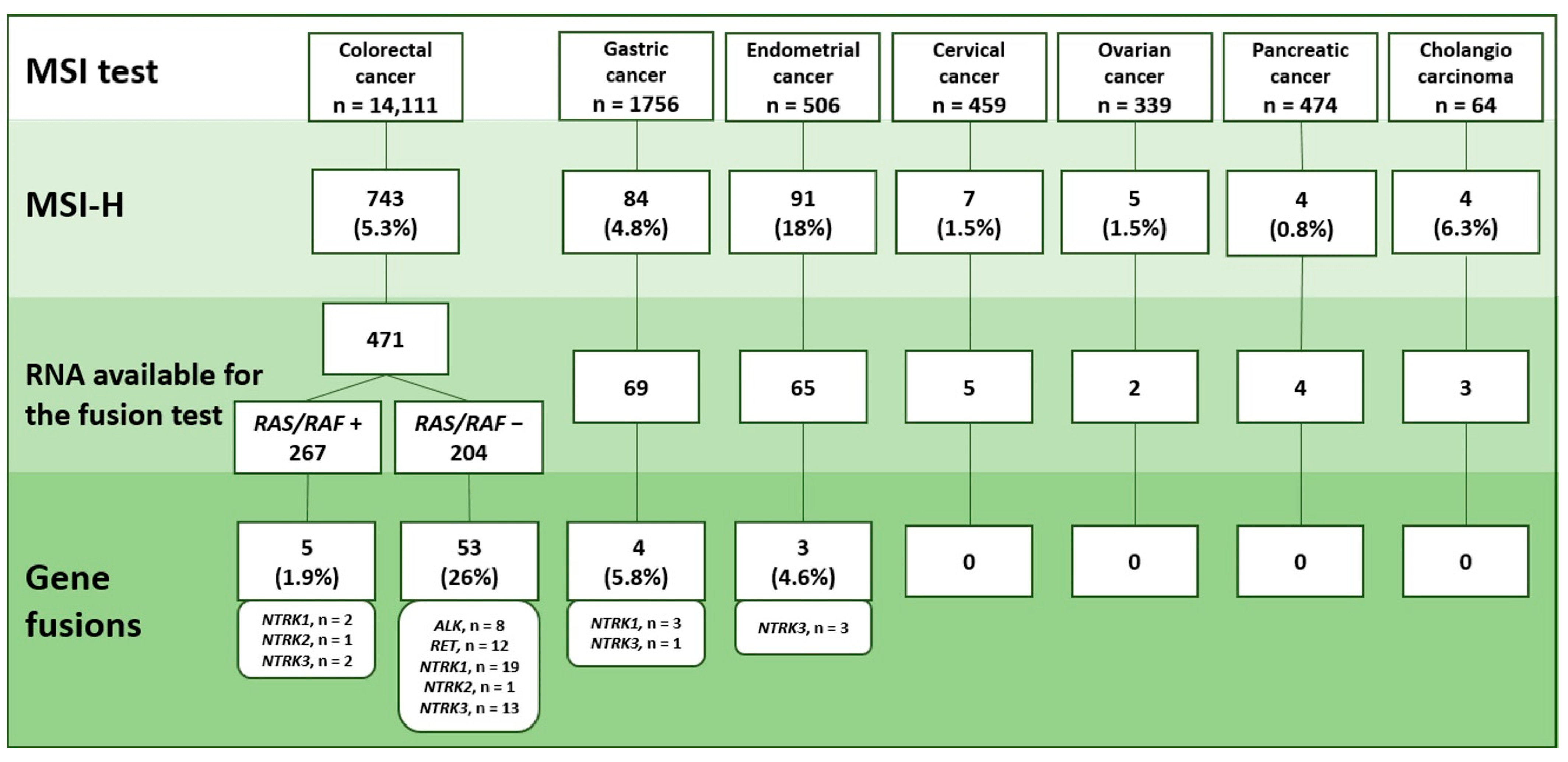
2.1. MSI Detection
2.2. Analysis of Gene Rearrangements
2.3. Pattern of Gene Rearrangement in Various Categories of Microsatellite-Unstable Tumors
2.4. MSI Analysis in Non-Colorectal Tumors Carrying ALK/ROS/RET/NTRK Rearrangements
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Shia, J. The diversity of tumours with microsatellite instability: Molecular mechanisms and impact upon microsatellite instability testing and mismatch repair protein immunohistochemistry. Histopathology 2020, 78, 485–497. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, R.; Li, J.; Yi, Y.; Liu, X.; Chen, J.; Zhang, H.; Lu, J.; Li, C.; Wu, H.; et al. Comprehensive analysis of oncogenic fusions in mismatch repair deficient colorectal carcinomas by sequential DNA and RNA next generation sequencing. J. Transl. Med. 2021, 19, 1–13. [Google Scholar] [CrossRef]
- Gallon, R.; Gawthorpe, P.; Phelps, R.L.; Hayes, C.; Borthwick, G.M.; Santibanez-Koref, M.; Jackson, M.S.; Burn, J. How Should We Test for Lynch Syndrome? A Review of Current Guidelines and Future Strategies. Cancers 2021, 13, 406. [Google Scholar] [CrossRef] [PubMed]
- Taieb, J.; Svrcek, M.; Cohen, R.; Basile, D.; Tougeron, D.; Phelip, J.M. Deficient mismatch repair/microsatellite unstable colorectal cancer: Diagnosis, prognosis and treatment. Eur. J. Cancer 2022, 175, 136–157. [Google Scholar] [CrossRef] [PubMed]
- Eso, Y.; Shimizu, T.; Takeda, H.; Takai, A.; Marusawa, H. Microsatellite instability and immune checkpoint inhibitors: Toward precision medicine against gastrointestinal and hepatobiliary cancers. J. Gastroenterol. 2019, 55, 15–26. [Google Scholar] [CrossRef]
- Jin, Z.; Sinicrope, F.A. Mismatch Repair-Deficient Colorectal Cancer: Building on Checkpoint Blockade. J. Clin. Oncol. 2022, 40, 2735–2750. [Google Scholar] [CrossRef] [PubMed]
- Imyanitov, E.; Kuligina, E. Molecular testing for colorectal cancer: Clinical applications. World J. Gastrointest. Oncol. 2021, 13, 1288–1301. [Google Scholar] [CrossRef]
- Vaňková, B.; Vaněček, T.; Ptáková, N.; Hájková, V.; Dušek, M.; Michal, M.; Švajdler, P.; Daum, O.; Daumová, M.; Mezencev, R.; et al. Targeted next generation sequencing of MLH1 -deficient, MLH1 promoter hypermethylated, and BRAF/RAS -wild-type colorectal adenocarcinomas is effective in detecting tumors with actionable oncogenic gene fusions. Genes Chromosom. Cancer 2020, 59, 562–568. [Google Scholar] [CrossRef] [PubMed]
- Bocciarelli, C.; Caumont, C.; Samaison, L.; Cariou, M.; Aline-Fardin, A.; Doucet, L.; Roudié, J.; Terris, B.; Merlio, J.P.; Marcorelles, P.; et al. MSI-High RAS-BRAF wild-type colorectal adenocarcinomas with MLH1 loss have a high frequency of targetable oncogenic gene fusions whose diagnoses are feasible using methods easy-to-implement in pathology laboratories. Hum. Pathol. 2021, 114, 99–109. [Google Scholar] [CrossRef]
- Singh, H.; Li, Y.Y.; Spurr, L.F.; Shinagare, A.B.; Abhyankar, R.; Reilly, E.; Brais, L.K.; Nag, A.; Ducar, M.D.; Thorner, A.R.; et al. Molecular Characterization and Therapeutic Targeting of Colorectal Cancers Harboring Receptor Tyrosine Kinase Fusions. Clin. Cancer Res. 2021, 27, 1695–1705. [Google Scholar] [CrossRef]
- Madison, R.W.; Hu, X.; Ramanan, V.; Xu, Z.; Huang, R.S.; Sokol, E.S.; Frampton, G.M.; Schrock, A.B.; Ali, S.M.; Ganesan, S.; et al. Clustered 8-Oxo-Guanine Mutations and Oncogenic Gene Fusions in Microsatellite-Unstable Colorectal Cancer. JCO Precis. Oncol. 2022, 6, e2100477. [Google Scholar] [CrossRef] [PubMed]
- Okano, S.; Yamashiro, Y.; Onagi, H.; Sasa, K.; Hayashi, T.; Takahashi, M.; Sugimoto, K.; Sakamoto, K.; Yao, T.; Saito, T. Tyrosine kinase alterations in colorectal cancer with emphasis on the distinct clinicopathological characteristics. Histopathology 2023. [Google Scholar] [CrossRef]
- Kurnit, K.C.; Westin, S.N.; Coleman, R.L. Microsatellite instability in endometrial cancer: New purpose for an old test. Cancer 2019, 125, 2154–2163. [Google Scholar] [CrossRef] [PubMed]
- Puliga, E.; Corso, S.; Pietrantonio, F.; Giordano, S. Microsatellite instability in Gastric Cancer: Between lights and shadows. Cancer Treat. Rev. 2021, 95, 102175. [Google Scholar] [CrossRef] [PubMed]
- Preobrazhenskaya, E.V.; Suleymanova, A.M.; Bizin, I.V.; Zagrebin, F.A.; Romanko, A.A.; Saitova, E.S.; Mulkidzhan, R.S.; Imyanitov, E.N. Spectrum of kinase gene rearrangements in a large series of paediatric inflammatory myofibroblastic tumours. Histopathology 2023, 83, 109–115. [Google Scholar] [CrossRef]
- Tiurin, V.I.; Preobrazhenskaya, E.V.; Mitiushkina, N.V.; Romanko, A.A.; Anuskina, A.A.; Mulkidjan, R.S.; Saitova, E.S.; Krivosheyeva, E.A.; Kharitonova, E.D.; Shevyakov, M.P.; et al. Rapid and Cost-Efficient Detection of RET Rearrangements in a Large Consecutive Series of Lung Carcinomas. Int. J. Mol. Sci. 2023, 24, 10530. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Matsumoto, S.; Liu, J.; Tanaka, K.; Mori, S.; Hayashi, K.; Kumagai, S.; Shibata, Y.; Hayashida, T.; Watanabe, K.; et al. The CLIP1–LTK fusion is an oncogenic driver in non-small-cell lung cancer. Nature 2021, 600, 319–323. [Google Scholar] [CrossRef]
- Imyanitov, E.; Zagrebin, F.; Mulkidzhan, R.; Krivosheeva, E.; Saitova, E.; Iyevleva, A.; Bizin, I.; Preobrazhenskaya, E. 25P Identification of novel kinase-activating fusions in non-small cell lung carcinomas (NSCLCs). Ann. Oncol. 2022, 33, S553–S554. [Google Scholar] [CrossRef]
- Martianov, A.S.; Mitiushkina, N.V.; Ershova, A.N.; Martynenko, D.E.; Bubnov, M.G.; Amankwah, P.; Yanus, G.A.; Aleksakhina, S.N.; Tiurin, V.I.; Venina, A.R.; et al. KRAS, NRAS, BRAF, HER2 and MSI Status in a Large Consecutive Series of Colorectal Carcinomas. Int. J. Mol. Sci. 2023, 24, 4868. [Google Scholar] [CrossRef]
- Afrăsânie, V.A.; Gafton, B.; Marinca, M.V.; Alexa-Stratulat, T.; Miron, L.; Rusu, C.; Ivanov, A.V.; Balan, G.G.; Croitoru, A.E. The Coexistence of RAS and BRAF Mutations in Metastatic Colorectal Cancer: A Case Report and Systematic Literature Review. J. Gastrointest. Liver Dis. 2020, 29, 251–256. [Google Scholar] [CrossRef]
- Uchida, S.; Kojima, T.; Sugino, T. Frequency and Clinicopathological Characteristics of Patients With KRAS/BRAF Double-Mutant Colorectal Cancer: An In Silico Study. Pathol. Oncol. Res. 2022, 28, 1610206. [Google Scholar] [CrossRef] [PubMed]
- Mustachio, L.M.; Roszik, J. Single-Cell Sequencing: Current Applications in Precision Onco-Genomics and Cancer Therapeutics. Cancers 2022, 14, 657. [Google Scholar] [CrossRef]
- O’haire, S.; Franchini, F.; Kang, Y.J.; Steinberg, J.; Canfell, K.; Desai, J.; Fox, S.; Ijzerman, M. Systematic review of NTRK 1/2/3 fusion prevalence pan-cancer and across solid tumours. Sci. Rep. 2023, 13, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, A.; Ferro, B.; Pasciuto, M.P.; Zampacorta, C.; Buttitta, F.; D’angelo, E. NTRK gene fusions in solid tumors: Agnostic relevance, prevalence and diagnostic strategies. Pathologica 2022, 114, 199–216. [Google Scholar] [CrossRef] [PubMed]
- Qi, C.; Zhou, T.; Bai, Y.; Chen, H.; Yuan, J.; Zhao, F.; Liu, C.; Ma, M.; Bei, T.; Chen, S.; et al. China special issue on gastrointestinal tumor—NTRK fusion in a large real-world population and clinical utility of circulating tumor DNA genotyping to guide TRK inhibitor treatment. Int. J. Cancer 2023. [Google Scholar] [CrossRef] [PubMed]
- Fois, S.S.; Paliogiannis, P.; Zinellu, A.; Fois, A.G.; Cossu, A.; Palmieri, G. Molecular Epidemiology of the Main Druggable Genetic Alterations in Non-Small Cell Lung Cancer. Int. J. Mol. Sci. 2021, 22, 612. [Google Scholar] [CrossRef]
- Wang, C.; Zhang, L.; Vakiani, E.; Shia, J. Detecting mismatch repair deficiency in solid neoplasms: Immunohistochemistry, microsatellite instability, or both? Mod. Pathol. 2022, 35, 1515–1528. [Google Scholar] [CrossRef] [PubMed]
- Marchiò, C.; Scaltriti, M.; Ladanyi, M.; Iafrate, A.; Bibeau, F.; Dietel, M.; Hechtman, J.; Troiani, T.; López-Rios, F.; Douillard, J.Y.; et al. ESMO recommendations on the standard methods to detect NTRK fusions in daily practice and clinical research. Ann. Oncol. 2019, 30, 1417–1427. [Google Scholar] [CrossRef] [PubMed]
- Nagtegaal, I.D.; Odze, R.D.; Klimstra, D.; Paradis, V.; Rugge, M.; Schirmacher, P.; Washington, K.M.; Carneiro, F.; Cree, I.A.; the WHO Classification of Tumours Editorial Board. The 2019 WHO classification of tumours of the digestive system. Histopathology 2020, 76, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.P.; Hoang, J.M.; Li, Y.J.; Seruca, R.; Carneiro, F.; Sobrinho-Simoes, M.; Lothe, R.A.; Gleeson, C.M.; Russell, S.H.; Muzeau, F.; et al. Determination of the replication error phenotype in human tumors without the requirement for matching normal DNA by analysis of mononucleotide repeat microsatellites. Genes Chromosom. Cancer 1998, 21, 101–107. [Google Scholar] [CrossRef]
- Suraweera, N.; Duval, A.; Reperant, M.; Vaury, C.; Furlan, D.; Leroy, K.; Seruca, R.; Iacopetta, B.; Hamelin, R. Evaluation of tumor microsatellite instability using five quasimonomorphic mononucleotide repeats and pentaplex PCR. Gastroenterology 2002, 123, 1804–1811. [Google Scholar] [CrossRef] [PubMed]

| ID | Sex | Age, Years | Diagnosis | RAS/RAF Mutations | 5′/3′-End Expression Imbalance | Fusion | Method of Fusion Identification | |
|---|---|---|---|---|---|---|---|---|
| 1. | A9365 | m | 74 | CRC | negative | ALK | STRN::ALK (S5;ins53A20) | NGS |
| 2. | C1182 | f | 71 | CRC | negative | ALK | EML4::ALK (E6;A20) | PCR |
| 3. | C3234 | f | 49 | CRC | negative | ALK | SPTBN1::ALK (S7;A20) | NGS |
| 4. | D0934 | m | 70 | CRC | negative | ALK | RBBP8::ALK (R8;A20) | NGS |
| 5. | P28580 | f | 66 | CRC | negative | ALK | SPTBN1::ALK (S7;A20) | NGS |
| 6. | P30811 | m | 80 | CRC | negative | ALK | EML4::ALK (E20;A20) | PCR |
| 7. | B9775 | f | 62 | CRC | negative | - | CCDC6::RET (C1;RE12) | PCR |
| 8. | C7622 | m | 58 | CRC | negative | - | CCDC6::RET (C1;RE12) | PCR |
| 9. | A7310 | m | 65 | CRC | negative | - | CCDC6::RET (C1;RE12) | PCR |
| 10. | P22395 | m | 62 | CRC | negative | - | CCDC6::RET (C1;RE12) | PCR |
| 11. | B8013 | f | 63 | CRC | negative | RET | NCOA4::RET (N8;R12) | PCR |
| 12. | C2542 | f | 67 | CRC | negative | RET | NCOA4::RET (N9del501;R12) | NGS |
| 13. | C4286 | m | 54 | CRC | negative | RET | NCOA4::RET (N9;R12) | NGS |
| 14. | P16667 | f | 65 | CRC | negative | RET | NCOA4::RET (N9;R12) | NGS |
| 15. | P24158 | f | 66 | CRC | negative | RET | CCDC6::RET (C1;RE12) | PCR |
| 16. | A4695 | f | 64 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 17. | B2240 | f | 60 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 18. | B9472 | m | 52 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 19. | C1589 | f | 64 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 20. | C6335 | f | 69 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 21. | C6755 | m | 72 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 22. | D0273 | m | 68 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 23. | D1407 | f | 58 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 24. | E2169 | m | 58 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 25. | P18891 | f | 59 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N12) | PCR |
| 26. | P21693 | m | 86 | CRC | negative | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 27. | E4114 | f | 69 | CRC | negative | NTRK1 | LMNA::NTRK1 (L3;N10) | PCR |
| 28. | B9540 | m | 72 | CRC | negative | NTRK1 | TPR::NTRK1 (T15;N10) | NGS |
| 29. | C4763 | m | 65 | CRC | negative | NTRK1 | TPR::NTRK1 (T20;N11) | NGS |
| 30. | C7112 | f | 65 | CRC | negative | NTRK1 | ZKSCAN1::NTRK1 (Z3;N12) | NGS |
| 31. | D0155 | f | 81 | CRC | negative | NTRK1 | LMNA::NTRK1 (L9ins44;del74N11) | NGS |
| 32. | B9665 | m | 74 | CRC | negative | NTRK2 | ETV6::NTRK2 (E5;N15) | NGS |
| 33. | A4890 | f | 65 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 34. | B1263 | f | 58 | CRC | negative | - | EML4::NTRK3 (E2;N14) | PCR |
| 35. | B4631 | f | 68 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 36. | C3968 | f | 70 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 37. | C5878 | m | 57 | CRC | negative | NTRK3 | ETV6::NTRK3 (E5;N15) | PCR |
| 38. | C6430 | f | 72 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 39. | C7912 | f | 54 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 40. | D7461 | f | 61 | CRC | negative | NTRK3 | ETV6::NTRK3 (E5;N15) | PCR |
| 41. | P22904 | f | 68 | CRC | negative | NTRK3 | EML4::NTRK3 (E2;N14) | PCR |
| 42. | P23972 | f | 67 | CRC | negative | NTRK3 | ETV6::NTRK3 (E5;N15) | PCR |
| 43. | B4561 | m | 31 | CRC | KRAS p.A146T | NTRK2 | ETV6::NTRK2 (E5;N15) | NGS |
| 44. | C1274 | f | 73 | CRC | BRAF p.V600E | - | ETV6::NTRK3 (E5;N15) | PCR |
| 45. | C3231 | f | 76 | CRC | BRAF p.V600E | NTRK1 | LMNA::NTRK1 (L3;N10) | PCR |
| 46. | P29082 | f | 85 | CRC | NRAS p.Q61K | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 47. | C7527 | f | 71 | GC | KRAS p.G12C | NTRK1 | TPM3::NTRK1 (T8;N10) | PCR |
| 48. | C4292 | f | 72 | CRC | Negative | ALK | not defined | n/a for NGS |
| 49. | C8576 | m | 66 | CRC | Negative | ALK | not defined | n/a for NGS |
| 50. | C2716 | m | 80 | CRC | Negative | RET | not defined | n/a for NGS |
| 51. | C5821 | m | 83 | CRC | Negative | RET | not defined | n/a for NGS |
| 52. | P15387 | m | 68 | CRC | Negative | RET | not defined | n/a for NGS |
| 53. | B9814 | f | 65 | CRC | Negative | NTRK1 | not defined | n/a for NGS |
| 54. | C4831 | f | 60 | CRC | Negative | NTRK1 | not defined | n/a for NGS |
| 55. | D0795 | m | 74 | CRC | Negative | NTRK1 | not defined | n/a for NGS |
| 56. | C5665 | f | 68 | CRC | Negative | NTRK3 | not defined | n/a for NGS |
| 57. | C9396 | m | 32 | CRC | Negative | NTRK3 | not defined | n/a for NGS |
| 58. | D3390 | f | 80 | CRC | Negative | NTRK3 | not defined | n/a for NGS |
| 59. | D5144 | m | 66 | CRC | KRAS p.G12D | NTRK3 | not defined | n/a for NGS |
| 60. | A9663 | m | 66 | GC | Negative | NTRK1 | not defined | n/a for NGS |
| 61. | C1642 | m | 83 | GC | Negative | NTRK1 | not defined | n/a for NGS |
| 62. | E3046 | m | 34 | GC | Negative | NTRK3 | not defined | n/a for NGS |
| 63. | C8708 | f | 67 | EC | Negative | NTRK3 | not defined | n/a for NGS |
| 64. | D5486 | f | 71 | EC | Negative | NTRK3 | not defined | n/a for NGS |
| 65. | D6096 | f | 64 | EC | Negative | NTRK3 | not defined | n/a for NGS |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mulkidjan, R.S.; Saitova, E.S.; Preobrazhenskaya, E.V.; Asadulaeva, K.A.; Bubnov, M.G.; Otradnova, E.A.; Terina, D.M.; Shulga, S.S.; Martynenko, D.E.; Semina, M.V.; et al. ALK, ROS1, RET and NTRK1–3 Gene Fusions in Colorectal and Non-Colorectal Microsatellite-Unstable Cancers. Int. J. Mol. Sci. 2023, 24, 13610. https://doi.org/10.3390/ijms241713610
Mulkidjan RS, Saitova ES, Preobrazhenskaya EV, Asadulaeva KA, Bubnov MG, Otradnova EA, Terina DM, Shulga SS, Martynenko DE, Semina MV, et al. ALK, ROS1, RET and NTRK1–3 Gene Fusions in Colorectal and Non-Colorectal Microsatellite-Unstable Cancers. International Journal of Molecular Sciences. 2023; 24(17):13610. https://doi.org/10.3390/ijms241713610
Chicago/Turabian StyleMulkidjan, Rimma S., Evgeniya S. Saitova, Elena V. Preobrazhenskaya, Karimat A. Asadulaeva, Mikhail G. Bubnov, Ekaterina A. Otradnova, Darya M. Terina, Sofia S. Shulga, Darya E. Martynenko, Maria V. Semina, and et al. 2023. "ALK, ROS1, RET and NTRK1–3 Gene Fusions in Colorectal and Non-Colorectal Microsatellite-Unstable Cancers" International Journal of Molecular Sciences 24, no. 17: 13610. https://doi.org/10.3390/ijms241713610






