Apolipoprotein-CIII O-Glycosylation, a Link between GALNT2 and Plasma Lipids
Abstract
:1. Introduction
2. Results
2.1. Cohort Characteristics
2.2. Apo-CIII O-Glycosylation, Type 2 Diabetes and Lipid Parameters
2.3. GWASs of Apo-CIII O-Glycosylation Proteoforms
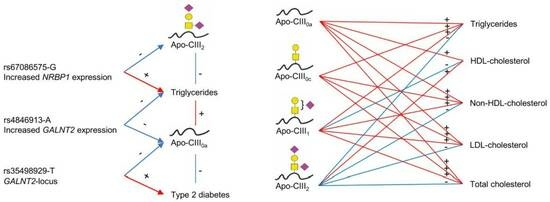
3. Discussion
4. Material and Methods
4.1. DiaGene Study
4.2. The Hoorn Diabetes Care System Cohort
4.3. Apo-CIII Glycosylation Measurements
4.4. GWAS of Apo-CIII Glycosylation
4.5. Statistical Methods
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Declaration of Generative AI and AI-assisted technologies in the writing process
References
- Kyu, H.H.; Abate, D.; Abate, K.H.; Abay, S.M.; Abbafati, C.; Abbasi, N.; Abbastabar, H.; Abd-Allah, F.; Abdela, J.; Abdelalim, A.; et al. Global, regional, and national disability-adjusted life-years (DALYs) for 359 diseases and injuries and healthy life expectancy (HALE) for 195 countries and territories, 1990-2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1859–1922. [Google Scholar] [CrossRef] [PubMed]
- Sone, H.; Tanaka, S.; Tanaka, S.; Iimuro, S.; Oida, K.; Yamasaki, Y.; Oikawa, S.; Ishibashi, S.; Katayama, S.; Ohashi, Y.; et al. Serum level of triglycerides is a potent risk factor comparable to LDL cholesterol for coronary heart disease in Japanese patients with type 2 diabetes: Subanalysis of the Japan Diabetes Complications Study (JDCS). J. Clin. Endocrinol. Metab. 2011, 96, 3448–3456. [Google Scholar] [CrossRef] [PubMed]
- Borén, J.; Packard, C.J.; Taskinen, M.R. The Roles of ApoC-III on the Metabolism of Triglyceride-Rich Lipoproteins in Humans. Front. Endocrinol. 2020, 11, 474. [Google Scholar] [CrossRef] [PubMed]
- Ginsberg, H.N.; Le, N.A.; Goldberg, I.J.; Gibson, J.C.; Rubinstein, A.; Wang-Iverson, P.; Norum, R.; Brown, W.V. Apolipoprotein B metabolism in subjects with deficiency of apolipoproteins CIII and AI. Evidence that apolipoprotein CIII inhibits catabolism of triglyceride-rich lipoproteins by lipoprotein lipase in vivo. J. Clin. Investig. 1986, 78, 1287–1295. [Google Scholar] [CrossRef]
- Kinnunen, P.K.J.; Ehnholm, C. Effect of serum and C-apoproteins from very low density lipoproteins on human postheparin plasma hepatic lipase. FEBS Lett. 1976, 65, 354–357. [Google Scholar] [CrossRef]
- Zhang, J.; de Albuquerque Rocha, N.; McCullough, P.A. Contribution of ApoCIII to Diabetic Dyslipidemia and Treatment With Volanesorsen. Rev. Cardiovasc. Med. 2018, 19, 13–19. [Google Scholar] [CrossRef]
- Sacks, F.M.; Zheng, C.; Cohn, J.S. Complexities of plasma apolipoprotein C-III metabolism. J. Lipid Res. 2011, 52, 1067–1070. [Google Scholar] [CrossRef]
- Fredenrich, A.; Giroux, L.M.; Tremblay, M.; Krimbou, L.; Davignon, J.; Cohn, J.S. Plasma lipoprotein distribution of apoC-III in normolipidemic and hypertriglyceridemic subjects: Comparison of the apoC-III to apoE ratio in different lipoprotein fractions. J. Lipid Res. 1997, 38, 1421–1432. [Google Scholar] [CrossRef]
- Valladolid-Acebes, I.; Berggren, P.-O.; Juntti-berggren, L. Apolipoprotein CIII Is an Important Piece in the Type-1 Diabetes Jigsaw Puzzle. Int. J. Mol. Sci. 2021, 22, 932. [Google Scholar] [CrossRef]
- Jørgensen, A.B.; Frikke-Schmidt, R.; Nordestgaard, B.G.; Tybjærg-Hansen, A. Loss-of-Function Mutations in APOC3 and Risk of Ischemic Vascular Disease. N. Engl. J. Med. 2014, 371, 32–41. [Google Scholar] [CrossRef]
- Kawakami, A.; Aikawa, M.; Alcaide, P.; Luscinskas, F.W.; Libby, P.; Sacks, F.M. Apolipoprotein CIII induces expression of vascular cell adhesion molecule-1 in vascular endothelial cells and increases adhesion of monocytic cells. Circulation 2006, 114, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Juntti-Berggren, L.; Refai, E.; Appelskog, I.; Andersson, M.; Imreh, G.; Dekki, N.; Uhles, S.; Yu, L.; Griffiths, W.J.; Zaitsev, S.; et al. Apolipoprotein CIII promotes Ca2+-dependent β cell death in type 1 diabetes. Proc. Natl. Acad. Sci. USA 2004, 101, 10090–10094. [Google Scholar] [CrossRef] [PubMed]
- Digenio, A.; Dunbar, R.L.; Alexander, V.J.; Hompesch, M.; Morrow, L.; Lee, R.G.; Graham, M.J.; Hughes, S.G.; Yu, R.; Singleton, W.; et al. Antisense-Mediated Lowering of Plasma Apolipoprotein C-III by Volanesorsen Improves Dyslipidemia and Insulin Sensitivity in Type 2 Diabetes. Diabetes Care 2016, 39, 1408–1415. [Google Scholar] [CrossRef]
- Van Hoek, M.; Van Herpt, T.W.; Dehghan, A.; Hofman, A.; Lieverse, A.G.; Van Duijn, C.M.; Witteman, J.C.M.; Sijbrands, E.J.G. Association of an APOC3 promoter variant with type 2 diabetes risk and need for insulin treatment in lean persons. Diabetologia 2011, 54, 1360–1367. [Google Scholar] [CrossRef] [PubMed]
- Schjoldager, K.T.B.G.; Vakhrushev, S.Y.; Kong, Y.; Steentoft, C.; Nudelman, A.S.; Pedersen, N.B.; Wandall, H.H.; Mandel, U.; Bennett, E.P.; Levery, S.B.; et al. Probing isoform-specific functions of polypeptide GalNAc-transferases using zinc finger nuclease glycoengineered SimpleCells. Proc. Natl. Acad. Sci. USA 2012, 109, 9893–9898. [Google Scholar] [CrossRef]
- Wada, Y.; Okamoto, N. Apolipoprotein C-III O-glycoform profiling of 500 serum samples by matrix-assisted laser desorption/ionization mass spectrometry for diagnosis of congenital disorders of glycosylation. J. Mass Spectrom. 2020, 56, e4597. [Google Scholar] [CrossRef] [PubMed]
- Surakka, I.; Horikoshi, M.; Mägi, R.; Sarin, A.-P.; Mahajan, A.; Lagou, V.; Marullo, L.; Ferreira, T.; Miraglio, B.; Timonen, S.; et al. The impact of low-frequency and rare variants on lipid levels. Nat. Genet. 2015, 47, 589–597. [Google Scholar] [CrossRef]
- Yassine, H.N.; Trenchevska, O.; Ramrakhiani, A.; Parekh, A.; Koska, J.; Walker, R.W.; Billheimer, D.; Reaven, P.D.; Yen, F.T.; Nelson, R.W.; et al. The association of human apolipoprotein C-III sialylation proteoforms with plasma triglycerides. PLoS ONE 2015, 10, e0144138. [Google Scholar] [CrossRef]
- Nicolardi, S.; Van Der Burgt, Y.E.M.; Dragan, I.; Hensbergen, P.J.; Deelder, A.M. Identification of new apolipoprotein-CIII glycoforms with ultrahigh resolution MALDI-FTICR mass spectrometry of human sera. J. Proteome Res. 2013, 12, 2260–2268. [Google Scholar] [CrossRef]
- Dos Santos Seckler, H.; Park, H.M.; Lloyd-Jones, C.M.; Melani, R.D.; Camarillo, J.M.; Wilkins, J.T.; Compton, P.D.; Kelleher, N.L. New Interface for Faster Proteoform Analysis: Immunoprecipitation Coupled with SampleStream-Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2021, 32, 1659–1670. [Google Scholar] [CrossRef]
- Kegulian, N.C.; Ramms, B.; Horton, S.; Trenchevska, O.; Nedelkov, D.; Graham, M.J.; Lee, R.G.; Esko, J.D.; Yassine, H.N.; Gordts, P.L.S.M. ApoC-III glycoforms are differentially cleared by hepatic TRL (Triglyceride-Rich Lipoprotein) receptors. Arterioscler. Thromb. Vasc. Biol. 2019, 39, 2145–2156. [Google Scholar] [CrossRef] [PubMed]
- Holleboom, A.G.; Karlsson, H.; Lin, R.S.; Beres, T.M.; Sierts, J.A.; Herman, D.S.; Stroes, E.S.G.; Aerts, J.M.; Kastelein, J.J.P.; Motazacker, M.M.; et al. Heterozygosity for a loss-of-function mutation in GALNT2 improves plasma triglyceride clearance in man. Cell Metab. 2011, 14, 811–818. [Google Scholar] [CrossRef] [PubMed]
- Mendoza, S.; Trenchevska, O.; King, S.M.; Nelson, R.W.; Nedelkov, D.; Krauss, R.M.; Yassine, H.N. Changes in low-density lipoprotein size phenotypes associate with changes in apolipoprotein C-III glycoforms after dietary interventions. J. Clin. Lipidol. 2017, 11, 224–233.e2. [Google Scholar] [CrossRef] [PubMed]
- Koska, J.; Yassine, H.; Trenchevska, O.; Sinar, S.; Schwenke, D.C.; Yen, F.T.; Billheimer, D.; Nelson, R.W.; Nedelkov, D.; Reaven, P.D. Disialylated apolipoprotein C-III proteoform is associated with improved lipids in prediabetes and type 2 diabetes1. J. Lipid Res. 2016, 57, 894–905. [Google Scholar] [CrossRef]
- Demus, D.; Naber, A.; Dotz, V.; Jansen, B.C.; Bladergroen, M.R.; Nouta, J.; Sijbrands, E.J.G.; Van Hoek, M.; Nicolardi, S.; Wuhrer, M. Large-Scale Analysis of Apolipoprotein CIII Glycosylation by Ultrahigh Resolution Mass Spectrometry. Front. Chem. 2021, 9, 322. [Google Scholar] [CrossRef]
- Elsworth, B.; Lyon, M.; Alexander, T.; Liu, Y.; Matthews, P.; Hallett, J.; Bates, P.; Palmer, T.; Haberland, V.; Smith, G.D.; et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv 2020. [Google Scholar] [CrossRef]
- Type 2 Diabetes Knowledge Portal., n.d. rs4846913, rs67086575, and rs10842926 variant pages. Available online: https://hugeamp.org/ (accessed on 25 May 2022).
- Van Den Boogert, M.A.W.; Rader, D.J.; Holleboom, A.G. New insights into the role of glycosylation in lipoprotein metabolism. Curr. Opin. Lipidol. 2017, 28, 502–506. [Google Scholar] [CrossRef]
- Schjoldager, K.T.B.G.; Vester-Christensen, M.B.; Bennett, E.P.; Levery, S.B.; Schwientek, T.; Yin, W.; Blixt, O.; Clausen, H. O-glycosylation modulates proprotein convertase activation of angiopoietin-like protein 3: Possible role of polypeptide GalNAc-transferase-2 in regulation of concentrations of plasma lipids. J. Biol. Chem. 2010, 285, 36293–36303. [Google Scholar] [CrossRef]
- Roman, T.S.; Marvelle, A.F.; Fogarty, M.P.; Vadlamudi, S.; Gonzalez, A.J.; Buchkovich, M.L.; Huyghe, J.R.; Fuchsberger, C.; Jackson, A.U.; Wu, Y.; et al. Multiple Hepatic Regulatory Variants at the GALNT2 GWAS Locus Associated with High-Density Lipoprotein Cholesterol. Am. J. Hum. Genet. 2015, 97, 801–815. [Google Scholar] [CrossRef]
- Cavalli, M.; Pan, G.; Nord, H.; Wadelius, C. Looking beyond GWAS: Allele-specific transcription factor binding drives the association of GALNT2 to HDL-C plasma levels. Lipids Health Dis. 2016, 15, 18. [Google Scholar] [CrossRef]
- Guo, T.; Yin, R.X.; Huang, F.; Yao, L.M.; Lin, W.X.; Pan, S.L. Association between the DOCK7, PCSK9 and GALNT2 Gene Polymorphisms and Serum Lipid levels. Sci. Rep. 2016, 6, 19079. [Google Scholar] [CrossRef] [PubMed]
- Ridker, P.M.; Paré, G.; Parker, A.N.; Zee, R.Y.L.; Miletich, J.P.; Chasman, D.I. Polymorphism in the CETP gene region, HDL cholesterol, and risk of future myocardial infarction: Genomewide analysis among 18 245 initially healthy women from the Women’s Genome Health Study. Circ. Cardiovasc. Genet. 2009, 2, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Taschner, M.; Bhogaraju, S.; Lorentzen, E. Architecture and function of IFT complex proteins in ciliogenesis. Differentiation 2012, 83, S12. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Song, J.; Jung, J.H.; Ko, H.W. Primary cilia in energy balance signaling and metabolic disorder. BMB Rep. 2015, 48, 647. [Google Scholar] [CrossRef] [PubMed]
- Võsa, U.; Claringbould, A.; Westra, H.-J.; Bonder, M.J.; Deelen, P.; Zeng, B.; Kirsten, H.; Saha, A.; Kreuzhuber, R.; Yazar, S.; et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 2021, 53, 1300–1310. [Google Scholar] [CrossRef]
- De Langhe, S.; Haataja, L.; Senadheera, D.; Groffen, J.; Heisterkamp, N. Interaction of the small GTPase Rac3 with NRBP, a protein with a kinase-homology domain. Int. J. Mol. Med. 2002, 9, 451–459. [Google Scholar] [CrossRef]
- Harduin-Lepers, A.; Vallejo-Ruiz, V.; Krzewinski-Recchi, M.A.; Samyn-Petit, B.; Julien, S.; Delannoy, P. The human sialyltransferase family. Biochimie 2001, 83, 727–737. [Google Scholar] [CrossRef]
- Hiukka, A.; Ståhlman, M.; Pettersson, C.; Levin, M.; Adiels, M.; Teneberg, S.; Leinonen, E.S.; Hultén, L.M.; Wiklund, O.; Orešič, M.; et al. ApoCIII-enriched LDL in type 2 diabetes displays altered lipid composition, increased susceptibility for sphingomyelinase, and increased binding to biglycan. Diabetes 2009, 58, 2018–2026. [Google Scholar] [CrossRef]
- Rodríguez, M.; Rehues, P.; Iranzo, V.; Mora, J.; Balsells, C.; Guardiola, M.; Ribalta, J. Distribution of seven ApoC-III glycoforms in plasma, VLDL, IDL, LDL and HDL of healthy subjects. J. Proteom. 2022, 251, 104398. [Google Scholar] [CrossRef]
- Van Herpt, T.T.W.; Lemmers, R.F.H.; van Hoek, M.; Langendonk, J.G.; Erdtsieck, R.J.; Bravenboer, B.; Lucas, A.; Mulder, M.T.; Haak, H.R.; Lieverse, A.G.; et al. Introduction of the DiaGene study: Clinical characteristics, pathophysiology and determinants of vascular complications of type 2 diabetes. Diabetol. Metab. Syndr. 2017, 9, 47. [Google Scholar] [CrossRef]
- Van Der Heijden, A.A.W.A.; Rauh, S.P.; Dekker, J.M.; Beulens, J.W.; Elders, P.; T’Hart, L.M.; Rutters, F.; Van Leeuwen, N.; Nijpels, G. The Hoorn Diabetes Care System (DCS) cohort. A prospective cohort of persons with type 2 diabetes treated in primary care in the Netherlands. BMJ Open 2017, 7, e015599. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, S.; Das, S.; Kretzschmar, W.; Delaneau, O.; Wood, A.R.; Teumer, A.; Kang, H.M.; Fuchsberger, C.; Danecek, P.; Sharp, K.; et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 2016, 48, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Loh, P.; Danecek, P.; Palamara, P.F.; Fuchsberger, C.; Reshef, A.; Finucane, H.K.; Schoenherr, S.; Forer, L.; Mccarthy, S.; Abecasis, G.R.; et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 2016, 48, 1443–1448. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Forer, L.; Schönherr, S.; Sidore, C.; Locke, A.E.; Kwong, A.; Vrieze, S.I.; Chew, E.Y.; Levy, S.; McGue, M.; et al. Next-generation genotype imputation service and methods. Nat. Genet. 2016, 48, 1284–1287. [Google Scholar] [CrossRef]
- Goldstein, J.I.; Crenshaw, A.; Carey, J.; Grant, G.B.; Maguire, J.; Fromer, M.; O’dushlaine, C.; Moran, J.L.; Chambert, K.; Stevens, C.; et al. Zcall: A rare variant caller for array-based genotyping. Bioinformatics 2012, 28, 2543–2545. [Google Scholar] [CrossRef]
- Watanabe, K.; Taskesen, E.; Van Bochoven, A.; Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 2017, 8, 1826. [Google Scholar] [CrossRef]
- Zhan, X.; Hu, Y.; Li, B.; Abecasis, G.R.; Liu, D.J. RVTESTS: An efficient and comprehensive tool for rare variant association analysis using sequence data. Bioinformatics 2016, 32, 1423–1426. [Google Scholar] [CrossRef]
- Winkler, T.W.; Day, F.R.; Croteau-Chonka, D.C.; Wood, A.R.; Locke, A.E.; Mägi, R.; Ferreira, T.; Fall, T.; Graff, M.; Justice, A.E.; et al. Quality control and conduct of genome-wide association meta-analyses. Nat. Protoc. 2014, 9, 1192–1212. [Google Scholar] [CrossRef]

| DiaGene | Hoorn DCS | |
|---|---|---|
| Number of participants | 2318 | 5409 |
| Female sex, n (%) | 1175 (50.7) | 2414 (44.6) |
| Age, year, mean (±SD) | 65.3 (9.5) | 61.1 (11,0) |
| BMI, kg/m2, median (IQR) | 28.0 (25.2–31.6) | 29.4 (26.6–33.1) |
| Type 2 diabetes, n (%) | 1572 (67.8) | 5409 (100) |
| HDL-cholesterol, mmol/L, median (IQR) | 1.2 (1.0–1.5) | 1.2 (1–1.4) |
| Non-HDL cholesterol, mmol/L, mean (±SD) | 3.4 (1.0) | 3.8 (1.5) |
| LDL-cholesterol, mmol/L, mean (±SD) | 2.8 (1.0) | 2.9 (1.5) |
| Triglycerides, mmol/L, median (IQR) | 1.4 (1.0–1.9) | 1.7 (1.2–2.3) |
| Total cholesterol, mmol/L, mean (±SD) | 4.7 (1.1) | 5.0 (1.6) |
| Use of any lipid-lowering therapy, n (%) | 1113 (50.3) | 2276 (42.1) |
| Use of statins, n (%) | 1090 (47.0) | 2182 (40.3) |
| Use of fibrates, n (%) | 33 (1.4) | 68 (1.3) |
| Use of bile acid sequestrants, n (%) | 1 (0.04) | 6 (0.1) |
| Use of nicotinic acid and derivates, n (%) | 5 (0.2) | 7 (0.1) |
| Use of other lipid modifying agents, n (%) | 56 (2.4) | 56 (1.0) |
| HDL-c | |||
|---|---|---|---|
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | 0.007 | 0.005 to 0.009 | 5.81 × 10−9 |
| Apo-CIII0c | −0.002 | −0.005 to 0.001 | 0.232 |
| Apo-CIII1 | −0.011 | −0.015 to −0.007 | 3.24 × 10−8 |
| Apo-CIII2 | 0.013 | 0.008 to 0.018 | 3.20 × 10−7 |
| Apo-CIII1/apo-CIII2 ratio | −0.321 | −0.422 to −0.221 | 4.00 × 10−10 |
| Non-HDL-c | |||
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | 0.004 | 0.003 to 0.004 | 1.48 × 10−18 |
| Apo-CIII0c | 0.002 | 0.001 to 0.003 | 3.09 × 10−4 |
| Apo-CIII1 | 0.006 | 0.004 to 0.007 | 9.04 × 10−16 |
| Apo-CIII2 | −0.007 | −0.009 to −0.006 | 3.60 × 10−17 |
| Apo-CIII1/apo-CIII2 ratio | 0.133 | 0.099 to 0.167 | 2.82 × 10−14 |
| LDL-c | |||
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | 0.004 | 0.003 to 0.005 | 3.86 × 10−20 |
| Apo-CIII0c | 0.001 | 0.000 to 0.002 | 0.041 |
| Apo-CIII1 | 0.004 | 0.003 to 0.005 | 4.33 × 10−8 |
| Apo-CIII2 | −0.005 | −0.007 to −0.003 | 5.74 × 10−8 |
| Apo-CIII1/apo-CIII2 ratio | 0.080 | 0.045 to 0.116 | 9.65 × 10−6 |
| Total cholesterol | |||
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | 0.004 | 0.003 to 0.005 | 4.37 × 10−24 |
| Apo-CIII0c | 0.002 | 0.001 to 0.003 | 0.003 |
| Apo-CIII1 | 0.004 | 0.002 to 0.005 | 8.99 × 10−9 |
| Apo-CIII2 | −0.005 | −0.007 to −0.004 | 4.17 × 10−10 |
| Apo-CIII1/apo-CIII2 ratio | 0.084 | 0.052 to 0.116 | 2.99 × 10−7 |
| Triglycerides | |||
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | 0.001 | 0.000 to 0.001 | 0.045 |
| Apo-CIII0c | 0.003 | 0.002 to 0.004 | 2.08 × 10−8 |
| Apo-CIII1 | 0.008 | 0.007 to 0.009 | 9.66 × 10−49 |
| Apo-CIII2 | −0.011 | −0.012 to −0.009 | 1.03 × 10−49 |
| Apo-CIII1/apo-CIII2 ratio | 0.225 | 0.193 to 0.257 | 2.26 × 10−51 |
| Type 2 diabetes | |||
| Proteoform | Beta | 95% CI | p-Value |
| Apo-CIII0a | −29.59 | −34.43 to −24.74 | 5.42 × 10−33 |
| Apo-CIII0c | −0.963 | −4.338 to 2.412 | 0.576 |
| Apo-CIII1 | 1.275 | −1.399 to 3.950 | 0.350 |
| Apo-CIII2 | −0.296 | −2.377 to 1.786 | 0.781 |
| Apo-CIII1/apo-CIII2 ratio | 0.110 | 0.002 to 0.217 | 0.046 |
| Proteoform | rsID | Chr | Pos | EA | RA | EAF | Beta | SE | p-Value | Locus |
|---|---|---|---|---|---|---|---|---|---|---|
| Apo-CIII0a | rs35498929 | 1 | 230286016 | T | C | 0.1249 | −0.1223 | 0.02472 | 7.44 × 10−7 | GALNT2 (intronic) |
| rs4846913 | 1 | 230294715 | A | C | 0.5964 | −0.2027 | 0.01624 | 9.77 × 10−36 | GALNT2 (intronic) | |
| rs3213497 | 1 | 230416320 | T | C | 0.1282 | −0.1271 | 0.02407 | 1.30 × 10−7 | GALNT2:RP5-956O18.3 (ncRNA exonic) | |
| Apo-CIII0c | rs9378785 | 6 | 3316862 | C | T | 0.0508 | −0.0918 | 0.01705 | 7.34 × 10−8 | SLC22A23 (intronic) |
| Apo-CIII1 | rs10842926 | 12 | 27689893 | C | G | 0.0826 | −0.0145 | 0.002928 | 7.07 × 10−7 | PPFIBP1 (intronic) |
| Apo-CIII2 | rs67086575 | 2 | 27686480 | G | A | 0.1502 | −0.0469 | 0.009473 | 7.32 × 10−7 | IFT172 (intronic) |
| rs2493926 | 6 | 148614267 | C | T | 0.1112 | 0.0520 | 0.01058 | 9.12 × 10−7 | SASH1 (intronic) | |
| rs2481968 | 13 | 28567172 | C | A | 0.4874 | 0.0334 | 0.006647 | 4.90 × 10−7 | RN7SL272P (intergenic) | |
| rs7175584 | 15 | 97500494 | T | C | 0.5638 | −0.0339 | 0.00671 | 4.48 × 10−7 | RN7SKP181 (intergenic) | |
| rs10412211 | 19 | 13591542 | T | G | 0.4579 | −0.0336 | 0.006713 | 5.62 × 10−7 | CACNA1A (intronic) | |
| Apo-CIII1/apo-CIII2 ratio | rs67086575 | 2 | 27686480 | G | A | 0.1502 | 0.0561 | 0.01133 | 7.37 × 10−7 | IFT172 (intronic) |
| rs9462715 | 6 | 12368221 | C | A | 0.0571 | −0.0826 | 0.01676 | 8.35 × 10−7 | RN7SKP293 (intergenic) | |
| rs2493926 | 6 | 148614267 | C | T | 0.1112 | −0.0627 | 0.01266 | 7.46 × 10−7 | SASH1 (intronic) | |
| rs7175584 | 15 | 97500494 | T | C | 0.5638 | 0.0419 | 0.008028 | 1.80 × 10−7 | RN7SKP181 (intergenic) | |
| rs10412211 | 19 | 13591542 | T | G | 0.4579 | 0.0396 | 0.008031 | 8.24 × 10−7 | CACNA1A (intronic) |
| HDL-c | DiaGene | Hoorn DCS | Meta-Analysis | |||||
|---|---|---|---|---|---|---|---|---|
| rsID | EA | Beta | 95% C.I. | p-Value | Beta | 95% C.I. | p-Value | p-Value |
| rs4846913 | A | 0.006 | −0.017 to 0.030 | 0.606 | 0.011 | −0.001 to 0.023 | 0.076 | 0.065 |
| rs9378785 | C | −0.058 | −0.113 to −0.003 | 0.040 | 0.005 | −0.025 to 0.033 | 0.751 | 0.472 |
| Non-HDL-c | DiaGene | Hoorn DCS | Meta-analysis | |||||
| rsID | EA | Beta | 95% C.I. | p-Value | Beta | 95% C.I. | p-Value | p-Value |
| rs67086575 | G | 0.120 | 0.025 to 0.215 | 0.013 | 0.001 | −0.056 to 0.059 | 0.961 | 0.207 |
| rs2493926 | C | −0.157 | −0.266 to −0.048 | 0.005 | 0.030 | −0.041 to 0.101 | 0.412 | 0.522 |
| LDL-c | DiaGene | Hoorn DCS | Meta-analysis | |||||
| rsID | EA | Beta | 95% C.I. | p-Value | Beta | 95% C.I. | p-Value | p-Value |
| rs67086575 | G | 0.109 | 0.018 to 0.201 | 0.019 | −0.045 | −0.097 to 0.006 | 0.086 | 0.707 |
| rs2493926 | C | −0.183 | −0.288 to −0.078 | 0.001 | 0.035 | −0.029 to 0.099 | 0.289 | 0.464 |
| rs10842926 | C | 0.038 | −0.084 to 0.160 | 0.547 | 0.089 | 0.016 to 0.162 | 0.016 | 0.016 |
| Total cholesterol | DiaGene | Hoorn DCS | Meta-analysis | |||||
| rsID | EA | Beta | 95% C.I. | p-Value | Beta | 95% C.I. | p-Value | p-Value |
| rs67086575 | G | 0.115 | 0.014 to 0.216 | 0.025 | 0.007 | −0.051 to 0.065 | 0.817 | 0.190 |
| rs2493926 | C | −0.171 | −0.286 to −0.055 | 0.004 | 0.017 | −0.055 to 0.088 | 0.650 | 0.311 |
| Triglycerides | DiaGene | Hoorn DCS | Meta-analysis | |||||
| rsID | EA | Beta | 95% C.I. | p-Value | Beta | 95% C.I. | p-Value | p-Value |
| rs4846913 | A | −0.063 | −0.135 to 0.008 | 0.032 | −0.052 | −0.097 to −0.006 | 0.034 | 0.006 |
| rs67086575 | G | 0.084 | −0.014 to 0.183 | 0.017 | 0.084 | 0.023 to 0.145 | 2.20 × 10−4 | 1.07 × 10−5 |
| rs10842926 | C | −0.081 | −0.212 to 0.051 | 0.061 | −0.061 | −0.147 to 0.025 | 0.185 | 0.039 |
| rs2481968 | C | −0.076 | −0.146 to −0.007 | 0.025 | 0.017 | −0.028 to 0.062 | 0.759 | 0.391 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naber, A.; Demus, D.; Slieker, R.; Nicolardi, S.; Beulens, J.W.J.; Elders, P.J.M.; Lieverse, A.G.; Sijbrands, E.J.G.; ’t Hart, L.M.; Wuhrer, M.; et al. Apolipoprotein-CIII O-Glycosylation, a Link between GALNT2 and Plasma Lipids. Int. J. Mol. Sci. 2023, 24, 14844. https://doi.org/10.3390/ijms241914844
Naber A, Demus D, Slieker R, Nicolardi S, Beulens JWJ, Elders PJM, Lieverse AG, Sijbrands EJG, ’t Hart LM, Wuhrer M, et al. Apolipoprotein-CIII O-Glycosylation, a Link between GALNT2 and Plasma Lipids. International Journal of Molecular Sciences. 2023; 24(19):14844. https://doi.org/10.3390/ijms241914844
Chicago/Turabian StyleNaber, Annemieke, Daniel Demus, Roderick Slieker, Simone Nicolardi, Joline W. J. Beulens, Petra J. M. Elders, Aloysius G. Lieverse, Eric J. G. Sijbrands, Leen M. ’t Hart, Manfred Wuhrer, and et al. 2023. "Apolipoprotein-CIII O-Glycosylation, a Link between GALNT2 and Plasma Lipids" International Journal of Molecular Sciences 24, no. 19: 14844. https://doi.org/10.3390/ijms241914844