Comparative Analyses of Chloroplast Genome Provide Effective Molecular Markers for Species and Cultivar Identification in Bougainvillea
Abstract
:1. Introduction
2. Results
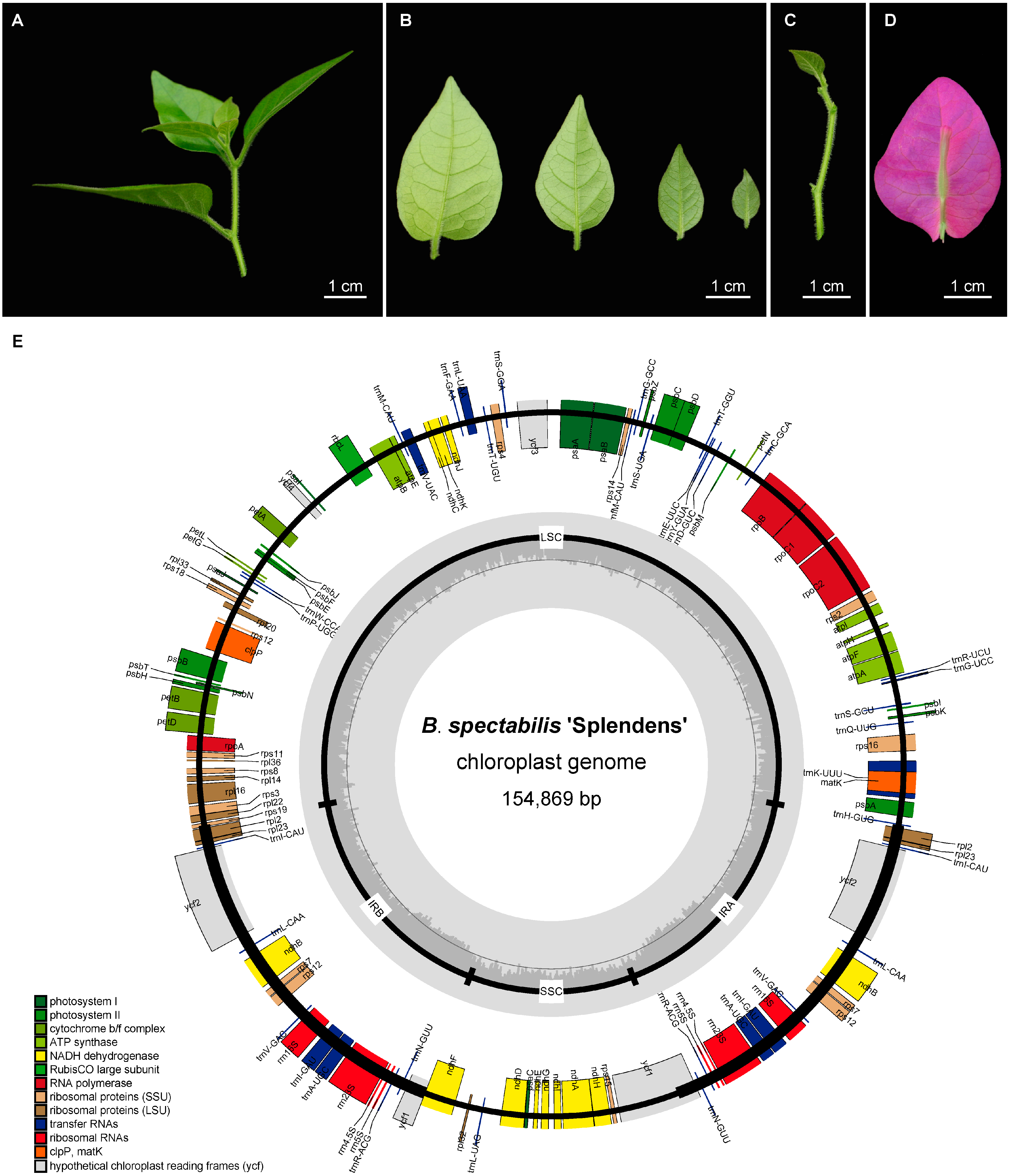
2.1. Morphological Characteristics and Chloroplast Genome Structure of B. spectabilis ‘Splendens’
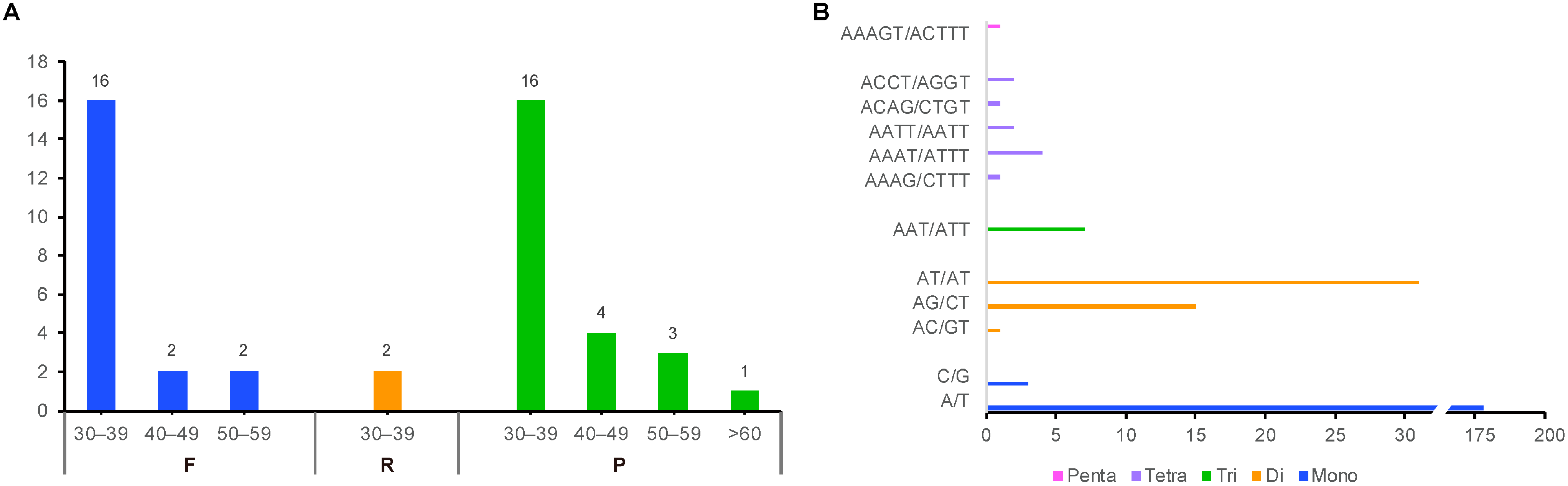
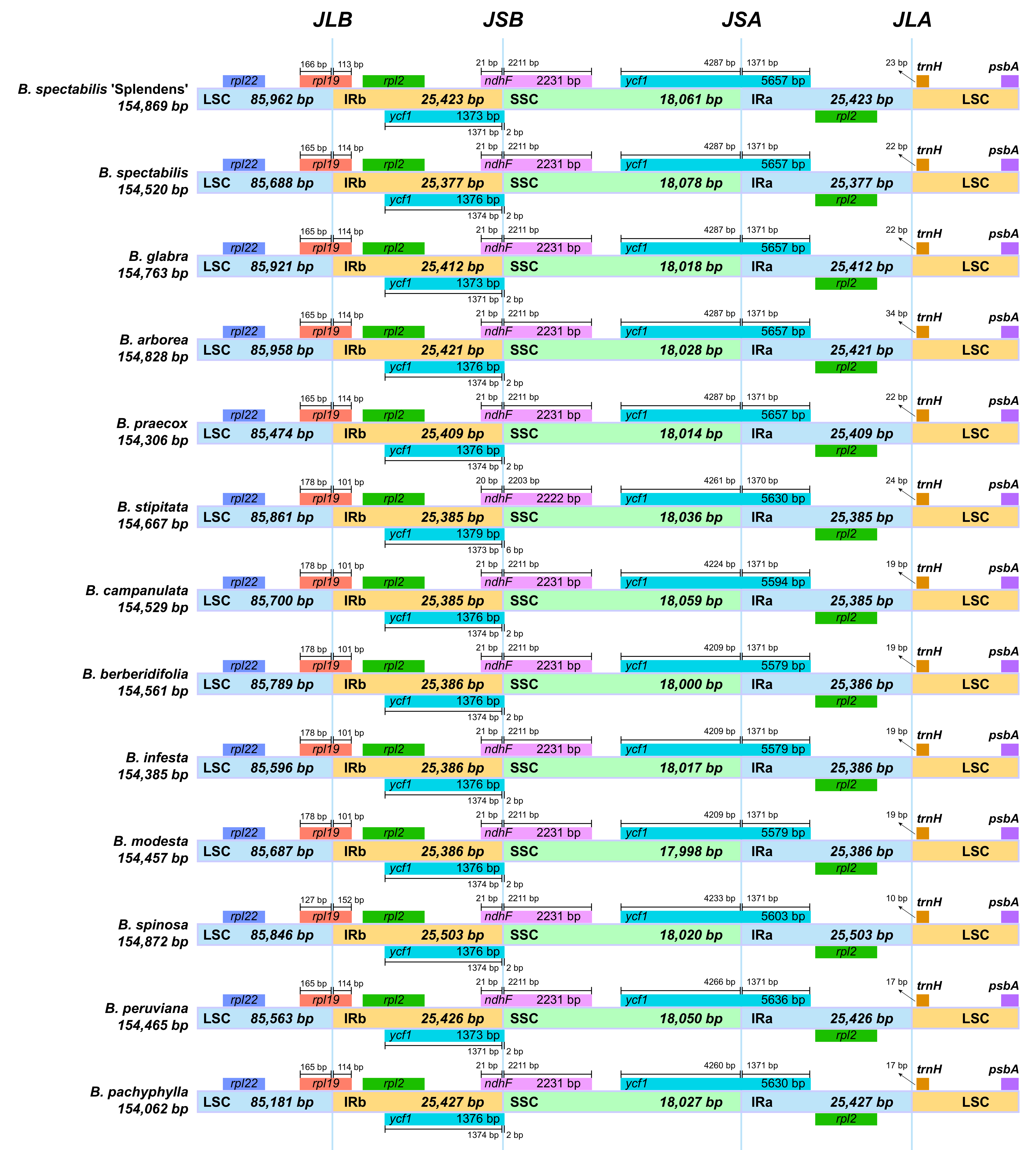
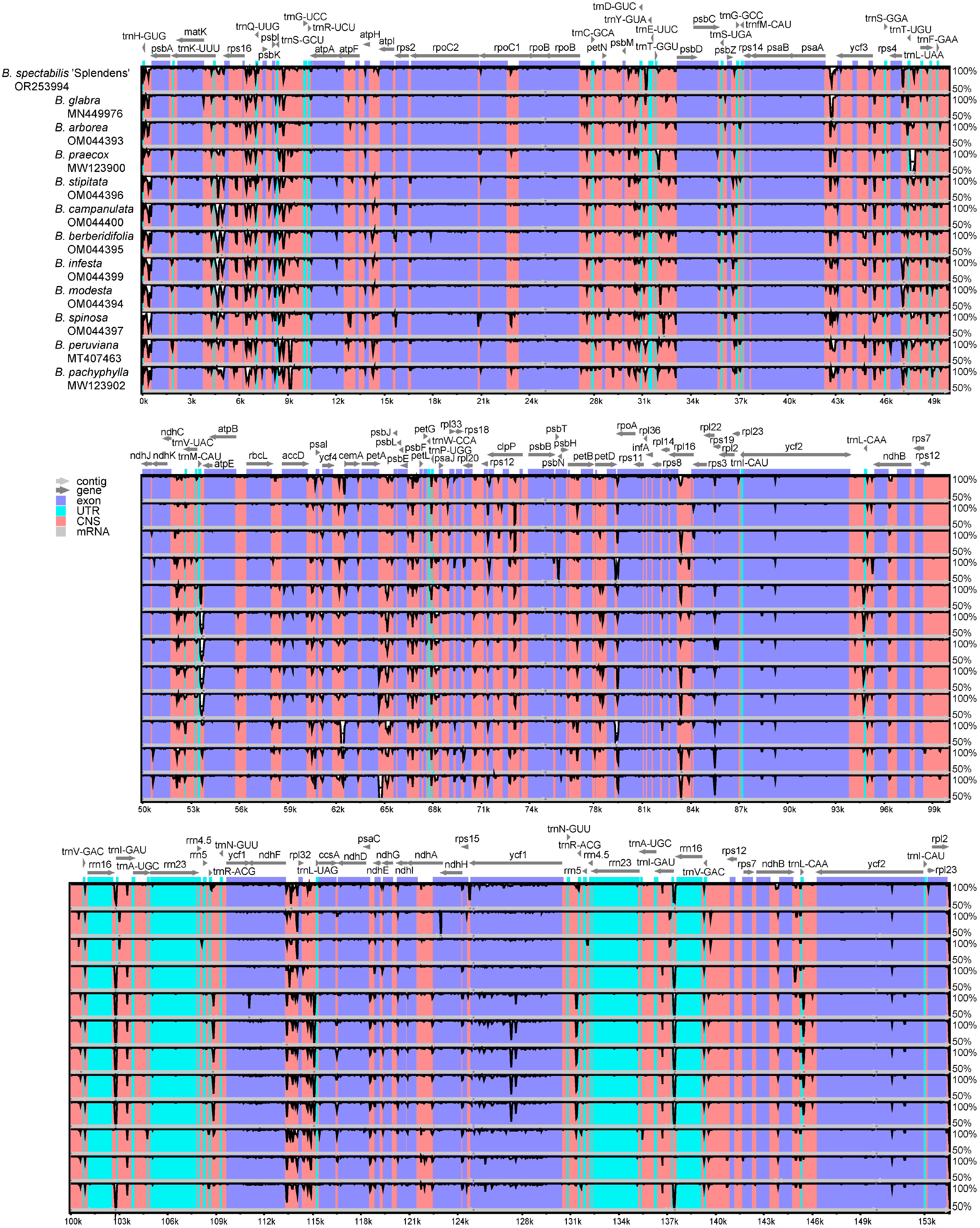
2.2. Comparative Genomic Divergence and Nucleotide Variability
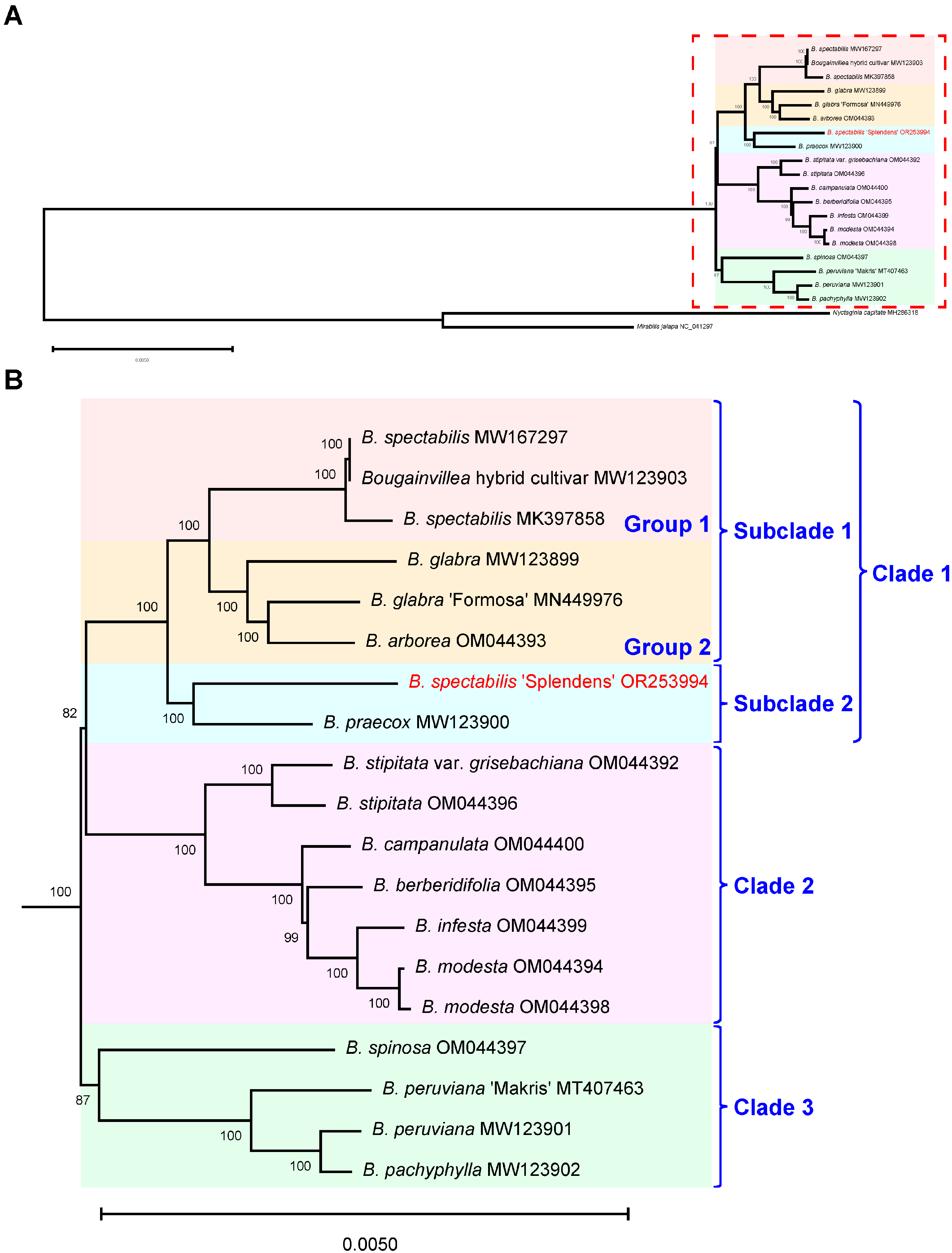
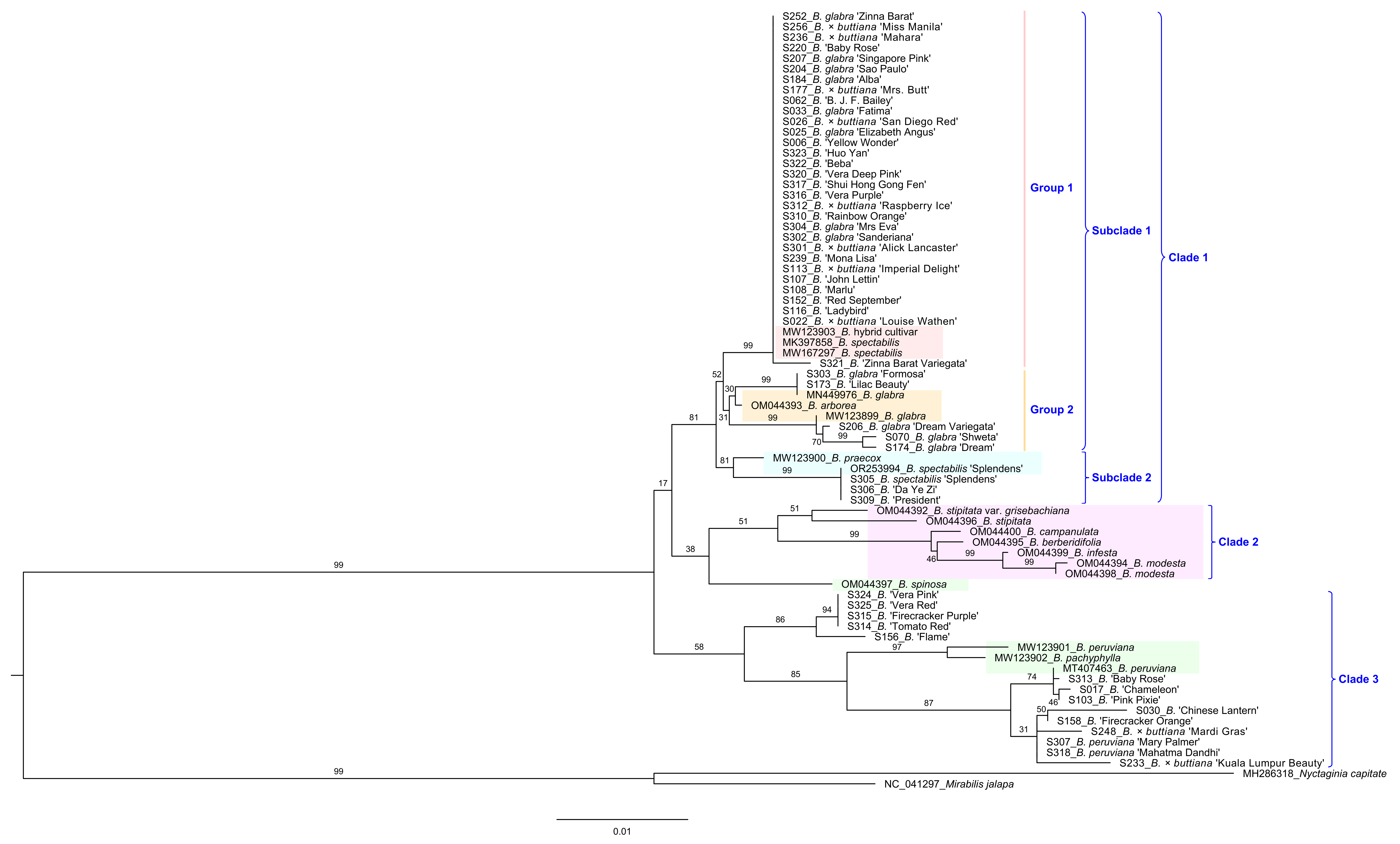
2.3. Phylogenetic Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Materials and DNA Extraction
4.2. Chloroplast Genome Sequencing, Assembly and Annotation
4.3. Genome Comparison and Variation Analysis
4.4. Phylogenetic Analysis
4.5. Primer Design and PCR Amplification
4.6. Multiple Sequence Alignment and Effectiveness of Marker Discriminatory
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Plants of the World Online. Available online: https://powo.science.kew.org/ (accessed on 28 November 2022).
- Kobayashi, K.D.; McConnell, J.; Griffis, J. Bougainvillea. In Ornamentals and Flowers; College of Tropical Agriculture and Human Resources, University of Hawaii: Honolulu, HI, USA, 2007; Volume OF-38, pp. 1–12. [Google Scholar]
- Roy, R.K.; Singh, S. Migration and domestication of Bougainvillea: A historical review. Chronica Hortic. 2016, 56, 10–15. [Google Scholar]
- Salam, P.; Bhargav, V.; Gupta, Y.C.; Nimbolkar, P.K. Evolution in Bougainvillea (Bougainvillea Commers.)—A review. J. Appl. Nat. Sci. 2017, 9, 1489–1494. [Google Scholar] [CrossRef]
- Szhualao; (Shenzhen Institute of Bougainvillea Research, Shenzhen, Guangdong, China). The International Bougainvillea Cultivars Checklist. Unpublished work. 2013. [Google Scholar]
- Zhou, Q.; Huang, K.F.; Ding, Y.L.; Guo, H.Z. Investigation and Taxonomic Identification on Introduced Ornamental Varieties in Bougainvillea in China. Acta Agric. Jiangxi 2011, 23, 53–56. [Google Scholar]
- Hammad, I. Genetic variation among Bougainvillea glabra cultivars (Nyctaginaceae) detected by RAPD markers and isozymes patterns. Res. J. Agri. Bio. Sci. 2009, 5, 63–71. [Google Scholar]
- Chatterjee, J.; Kalam, A.; Mandal, A.; Chakrabarty, D.; Datta, S.K. Use of RAPD analysis to determine genetic diversity and relationships among bougainvillea cultivars at intra and inter specific levels. Hortic. Environ. Biotechnol. 2007, 48, 43–51. [Google Scholar]
- Srivastava, R.; Shukla, S.; Soni, A.; Kumar, A. RAPD-based genetic relationships in different Bougainvillea cultivars. Crop Breed. Appl. Biotechnol. 2009, 9, 154–163. [Google Scholar] [CrossRef]
- Kumar, P.P.; Janakiram, T.; Bhatt, K.V.; Jain, R.; Prasad, K.V.; Prabhu, K.V. Molecular characterization and cultivar identification in Bougainvillea spp. using SSR markers. Indian J. Agric. Sci. 2014, 84, 1024–1030. [Google Scholar] [CrossRef]
- Nock, C.J.; Waters, D.L.; Edwards, M.A.; Bowen, S.G.; Rice, N.; Cordeiro, G.M.; Henry, R.J. Chloroplast genome sequences from total DNA for plant identification. Plant Biotechnol. J. 2011, 9, 328–333. [Google Scholar] [CrossRef] [PubMed]
- Martin, W.; Deusch, O.; Stawski, N.; Grunheit, N.; Goremykin, V. Chloroplast genome phylogenetics: Why we need independent approaches to plant molecular evolution. Trends Plant Sci. 2005, 10, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Douglas, N.A.; Manos, P.S. Molecular phylogeny of Nyctaginaceae: Taxonomy, biogeography, and characters associated with a radiation of xerophytic genera in North America. Am. J. Bot. 2007, 94, 856–872. [Google Scholar] [CrossRef]
- Ni, J.; Lee, S.Y.; Hu, X.; Wang, W.; Zhang, J.; Ruan, L.; Dai, S.; Liu, G. The complete chloroplast genome of a commercially exploited ornamental plant, Bougainvillea glabra (Caryophyllales: Nyctaginaceae). Mitochondrial DNA B Resour. 2019, 4, 3390–3391. [Google Scholar] [CrossRef] [PubMed]
- Bautista, M.A.C.; Zheng, Y.; Hu, Z.; Deng, Y.; Chen, T. Comparative Analysis of Complete Chloroplast Genome Sequences of Wild and Cultivated Bougainvillea (Nyctaginaceae). Plants 2020, 9, 1671. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Lee, S.Y.; Hu, X.; Sun, M.; Ni, J.; Wang, W.; Dai, S.; Ruan, L. Characterization of the complete chloroplast genome of ornamental plant, Bougainvillea peruviana (Nyctaginaceae). Mitochondrial DNA B Resour. 2020, 5, 3267–3268. [Google Scholar] [CrossRef] [PubMed]
- Bautista, M.A.C.; Zheng, Y.; Boufford, D.E.; Hu, Z.; Deng, Y.; Chen, T. Phylogeny and Taxonomic Synopsis of the Genus Bougainvillea (Nyctaginaceae). Plants 2022, 11, 1700. [Google Scholar] [CrossRef]
- Yan, J.; Singh, S. Migration of Bougainvillea and its Domestication: A Study. J. Greens Gardens 2019, 2, 6–12. [Google Scholar]
- Daniell, H.; Jin, S.; Zhu, X.G.; Gitzendanner, M.A.; Soltis, D.E.; Soltis, P.S. Green giant-a tiny chloroplast genome with mighty power to produce high-value proteins: History and phylogeny. Plant Biotechnol. J. 2021, 19, 430–447. [Google Scholar] [CrossRef]
- Bremer, B. A Review of Molecular Phylogenetic Studies of Rubiaceae. Ann. Mo. Bot. Gard. 2009, 96, 4–26. [Google Scholar] [CrossRef]
- Tosh, J.; Davis, A.P.; Dessein, S.; Block, P.D.; Huysmans, S.; Fay, M.F.; Smets, E.; Robbrecht, E. Phylogeny of Tricalysia (Rubiaceae) and its Relationships with Allied Genera Based on Plastid DNA Data: Resurrection of the Genus Empogona. Ann. Mo. Bot. Gard. 2009, 96, 194–213. [Google Scholar] [CrossRef]
- Block, P.D.; Rakotonasolo, F.; Ntore, S.; Razafimandimbison, S.G.; Janssens, S. Four new endemic genera of Rubiaceae (Pavetteae) from Madagascar represent multiple radiations into drylands. PhytoKeys 2018, 99, 1–66. [Google Scholar] [CrossRef]
- Cristians, S.; Bye, R.; Nieto-Sotelo, J. Molecular Markers Associated With Chemical Analysis: A Powerful Tool for Quality Control Assessment of Copalchi Medicinal Plant Complex. Front. Pharmacol. 2018, 9, 666. [Google Scholar] [CrossRef]
- Dong, W.; Liu, J.; Yu, J.; Wang, L.; Zhou, S. Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS ONE 2012, 7, e35071. [Google Scholar] [CrossRef]
- Pang, X.; Liu, C.; Shi, L.; Liu, R.; Liang, D.; Li, H.; Cherny, S.S.; Chen, S. Utility of the trnH-psbA intergenic spacer region and its combinations as plant DNA barcodes: A meta-analysis. PLoS ONE 2012, 7, e48833. [Google Scholar] [CrossRef]
- Mishra, P.; Kumar, A.; Nagireddy, A.; Mani, D.N.; Shukla, A.K.; Tiwari, R.; Sundaresan, V. DNA barcoding: An efficient tool to overcome authentication challenges in the herbal market. Plant Biotechnol. J. 2015, 14, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Shchegoleva, N.V.; Nikitina, E.V.; Juramurodov, I.J.; Zverev, A.A.; Turginov, O.T.; Jabborov, A.M.; Yusupov, Z.; Dekhkonov, D.B.; Deng, T.; Sun, H. A new species of Ranunculus (Ranunculaceae) from Western Pamir-Alay, Uzbekistan. PhytoKeys 2022, 193, 125–139. [Google Scholar] [CrossRef]
- Dong, W.P.; Xu, C.; Li, C.; Sun, J.; Zuo, Y.; Shi, S.; Cheng, T.; Guo, J.; Zhou, S. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 2015, 5, 8348. [Google Scholar] [CrossRef]
- Datta, S.K. Breeding of Bougainvillea: Past, present, and future. Nucleus 2022, 65, 239–254. [Google Scholar] [CrossRef]
- Sindhu, S.S.; Singh, B.; Saxena, N.K. Genetic improvement of bougainvillea in Indian scenario—A review. J. Ornam. Hortic. 2020, 23, 1–11. [Google Scholar] [CrossRef]
- Patel, R.K.; Jain, M. NGS QC Toolkit: A toolkit for quality control of next generation sequencing data. PLoS ONE 2012, 7, e30619. [Google Scholar] [CrossRef]
- Dierckxsens, N.; Mardulyn, P.; Smits, G. NOVOPlasty: De novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 2017, 45, e18. [Google Scholar]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Shi, L.; Zhu, Y.; Chen, H.; Zhang, J.; Lin, X.; Guan, X. CpGAVAS, an integrated web server for the annotation, visualization, analysis, and GenBank submission of completely sequenced chloroplast genome sequences. BMC Genom. 2012, 13, 715. [Google Scholar] [CrossRef] [PubMed]
- Greiner, S.; Lehwark, P.; Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019, 47, W59–W64. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [PubMed]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Amiryousefi, A.; Hyvönen, J.; Poczai, P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Available online: https://oaktrust.library.tamu.edu/handle/1969.1/104937/ (accessed on 21 March 2023).
- Kumar, P.P.; Janakiram, T.; Bhat, K.V.; Prasad, K.V.; Jain, R. Genetic divergence analysis of bougainvillea (Bougainvillea spp.) cultivars using morphological markers. Indian J. Agric. Sci. 2015, 85, 661–665. [Google Scholar] [CrossRef]
- Standard Names of Common Bougainvillea Cultivars. Bougainvillea workstation in Nursery Stock Branch of China Flower Association. Unpublished work. 2021.
- Liu, Y.M.; Ruan, L.; Zhou, H.G.; Yu, M.J. Cultivar Classification of Bougainvillea; China Forestry Publishing House: Beijing, China, 2020. [Google Scholar]
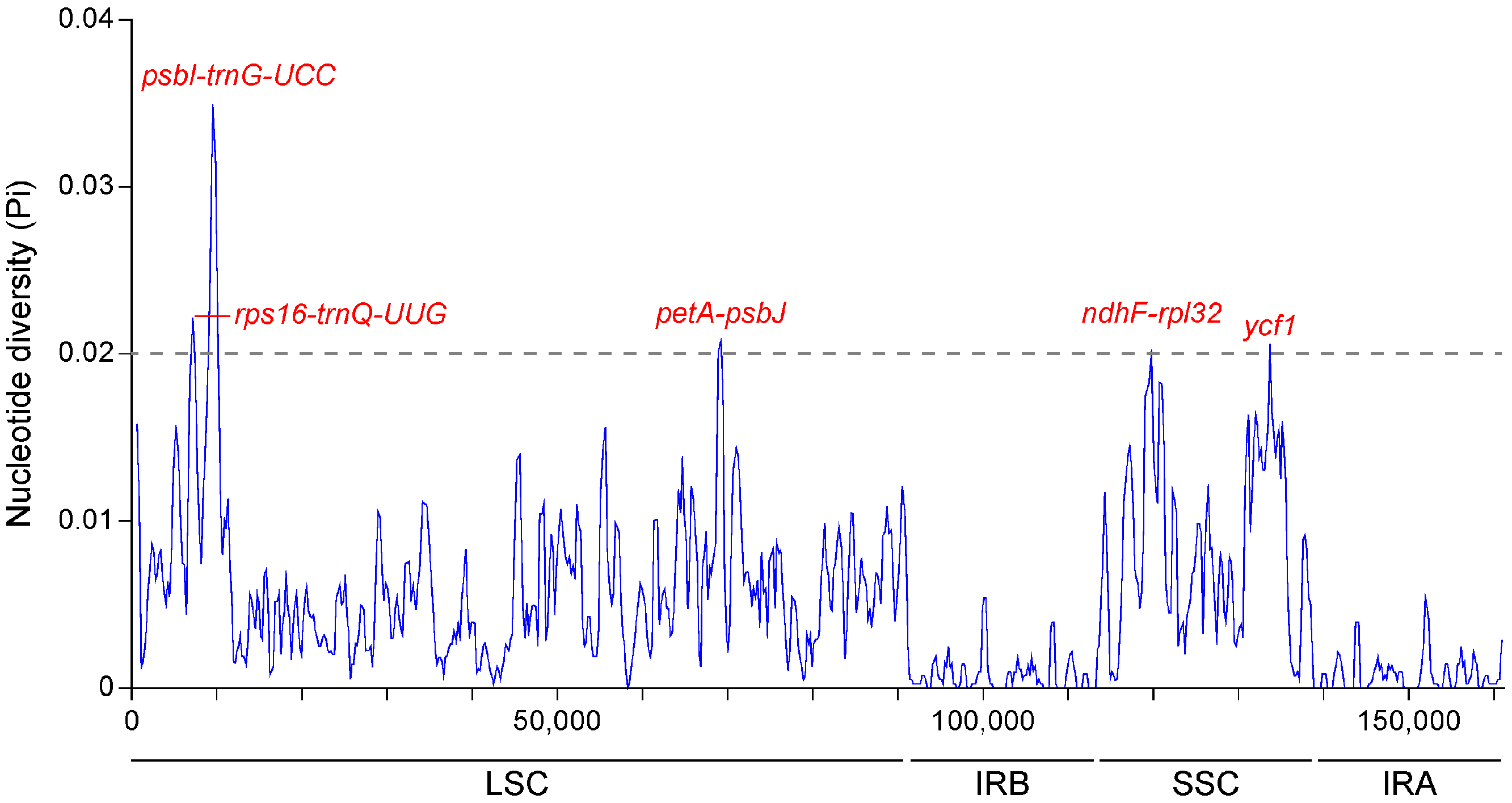
| Species | Mutational Hotspots | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| rps16-trnQ-UUG Position: 6744–7587 | psbI-trnG-UCC Position: 8957–9978 | petA-psbJ Position: 68,799–69,549 | ||||||||||
| Length | GC Content | Number of SNPs | Number of Gaps | Length | GC Content | Number of SNPs | Number of Gaps | Length | GC Content | Number of SNPs | Number of Gaps | |
| B. spectabilis ‘Splendens’ OR253994 | 723 | 23.7% | / | / | 748 | 22.3% | / | / | 628 | 28.3% | / | / |
| B. spectabilis MW167297 | 712 | 23.2% | 23 | 11 | 738 | 22.1% | 20 | 10 | 636 | 27.4% | 11 | −8 |
| B. glabra MN449976 | 701 | 24.4% | 17 | 22 | 776 | 21.4% | 12 | −28 | 635 | 28.0% | 6 | −7 |
| B. arborea OM044393 | 688 | 24.6% | 16 | 35 | 794 | 21.5% | 12 | −46 | 635 | 28.2% | 9 | −7 |
| B. praecox MW123900 | 724 | 23.6% | 16 | −1 | 692 | 23.7% | 7 | 56 | 633 | 28.3% | 6 | −5 |
| B. stipitata OM044396 | 713 | 24.4% | 26 | 10 | 798 | 21.2% | 45 | −50 | 684 | 26.0% | 20 | −56 |
| B. campanulata OM044400 | 704 | 24.6% | 29 | 19 | 745 | 22.3% | 35 | 3 | 702 | 26.4% | 21 | −74 |
| B. berberidifolia OM044395 | 710 | 24.4% | 28 | 13 | 745 | 22.1% | 34 | 3 | 672 | 27.1% | 21 | −44 |
| B. infesta OM044399 | 692 | 24.9% | 24 | 31 | 743 | 22.3% | 37 | 5 | 684 | 26.8% | 21 | −56 |
| B. modesta OM044394 | 705 | 24.4% | 28 | 18 | 751 | 22.0% | 36 | −3 | 684 | 26.8% | 21 | −56 |
| B. spinosa OM044397 | 703 | 23.8% | 20 | 20 | 752 | 21.8% | 44 | −4 | 684 | 28.4% | 19 | −56 |
| B. peruviana MT407463 | 713 | 23.6% | 30 | 10 | 716 | 22.5% | 60 | 32 | 628 | 27.7% | 18 | 0 |
| B. pachyphylla MW123902 | 710 | 23.7% | 30 | 13 | 779 | 21.4% | 62 | −31 | 628 | 27.9% | 17 | 0 |
| Total | / | / | 287 | / | / | / | 404 | / | / | / | 190 | / |
| Species | Mutational Hotspots | |||||||
|---|---|---|---|---|---|---|---|---|
| ndhF-rpl32 Position: 119,304–120,150 | ycf1 Position: 133,364–134,041 | |||||||
| Length | GC Content | Number of SNPs | Number of Gaps | Length | GC Content | Number of SNPs | Number of Gaps | |
| B. spectabilis ‘Splendens’ OR253994 | 701 | 20.4% | / | / | 636 | 22.3% | / | / |
| B. spectabilis MW167297 | 734 | 20.3% | 23 | −33 | 636 | 22.6% | 10 | 0 |
| B. glabra MN449976 | 686 | 21.1% | 22 | 15 | 636 | 22.6% | 14 | 0 |
| B. arborea OM044393 | 686 | 21.0% | 12 | 15 | 636 | 23.0% | 14 | 0 |
| B. praecox MW123900 | 672 | 20.4% | 20 | 29 | 636 | 22.2% | 7 | 0 |
| B. stipitata OM044396 | 748 | 19.7% | 23 | −47 | 657 | 21.8% | 16 | −21 |
| B. campanulata OM044400 | 764 | 19.6% | 26 | −63 | 621 | 22.2% | 19 | 15 |
| B. berberidifolia OM044395 | 746 | 19.7% | 25 | −45 | 621 | 22.5% | 14 | 15 |
| B. infesta OM044399 | 765 | 19.6% | 23 | −64 | 621 | 22.5% | 15 | 15 |
| B. modesta OM044394 | 746 | 19.7% | 23 | −45 | 621 | 22.5% | 15 | 15 |
| B. spinosa OM044397 | 737 | 20.2% | 36 | −36 | 636 | 22.5% | 12 | 0 |
| B. peruviana MT407463 | 739 | 20.3% | 30 | −38 | 642 | 22.4% | 16 | −6 |
| B. pachyphylla MW123902 | 740 | 20.3% | 34 | −39 | 642 | 22.7% | 14 | −6 |
| Total | / | / | 297 | / | / | / | 166 | / |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, X.; Lee, S.Y.; Ni, J.; Zhang, X.; Hu, X.; Zou, P.; Wang, W.; Liu, G. Comparative Analyses of Chloroplast Genome Provide Effective Molecular Markers for Species and Cultivar Identification in Bougainvillea. Int. J. Mol. Sci. 2023, 24, 15138. https://doi.org/10.3390/ijms242015138
Lin X, Lee SY, Ni J, Zhang X, Hu X, Zou P, Wang W, Liu G. Comparative Analyses of Chloroplast Genome Provide Effective Molecular Markers for Species and Cultivar Identification in Bougainvillea. International Journal of Molecular Sciences. 2023; 24(20):15138. https://doi.org/10.3390/ijms242015138
Chicago/Turabian StyleLin, Xinggu, Shiou Yih Lee, Jianzhong Ni, Xiaomin Zhang, Xing Hu, Peishan Zou, Wei Wang, and Guofeng Liu. 2023. "Comparative Analyses of Chloroplast Genome Provide Effective Molecular Markers for Species and Cultivar Identification in Bougainvillea" International Journal of Molecular Sciences 24, no. 20: 15138. https://doi.org/10.3390/ijms242015138






