Whole-Transcriptome Sequencing of Antler Tissue Reveals That circRNA2829 Regulates Chondrocyte Proliferation and Differentiation via the miR-4286-R+1/FOXO4 Axis
Abstract
:1. Introduction
2. Result
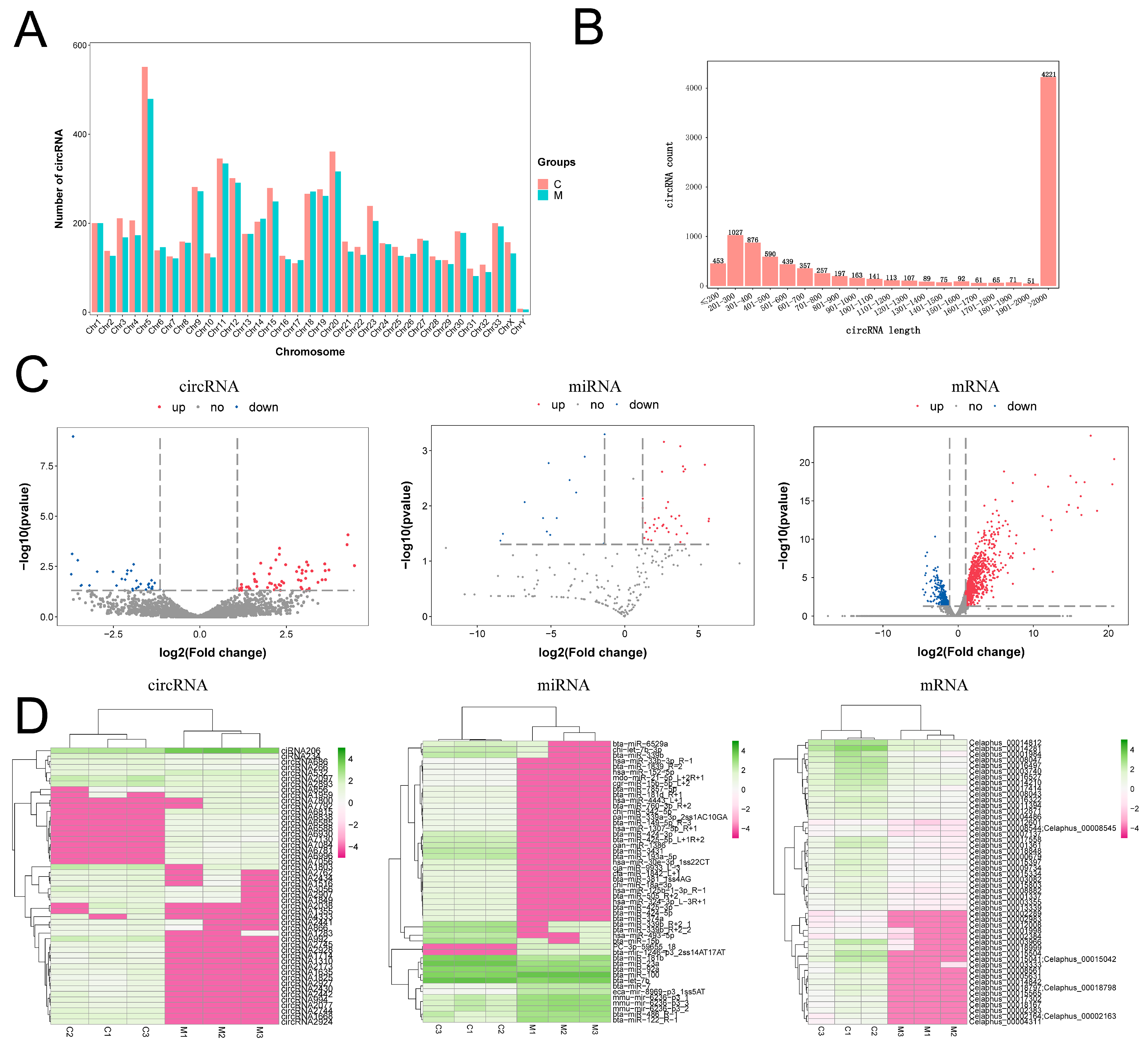
2.1. Analysis of the Differentially Expressed circRNAs, miRNAs, and mRNAs in Tip Tissue of Antlers
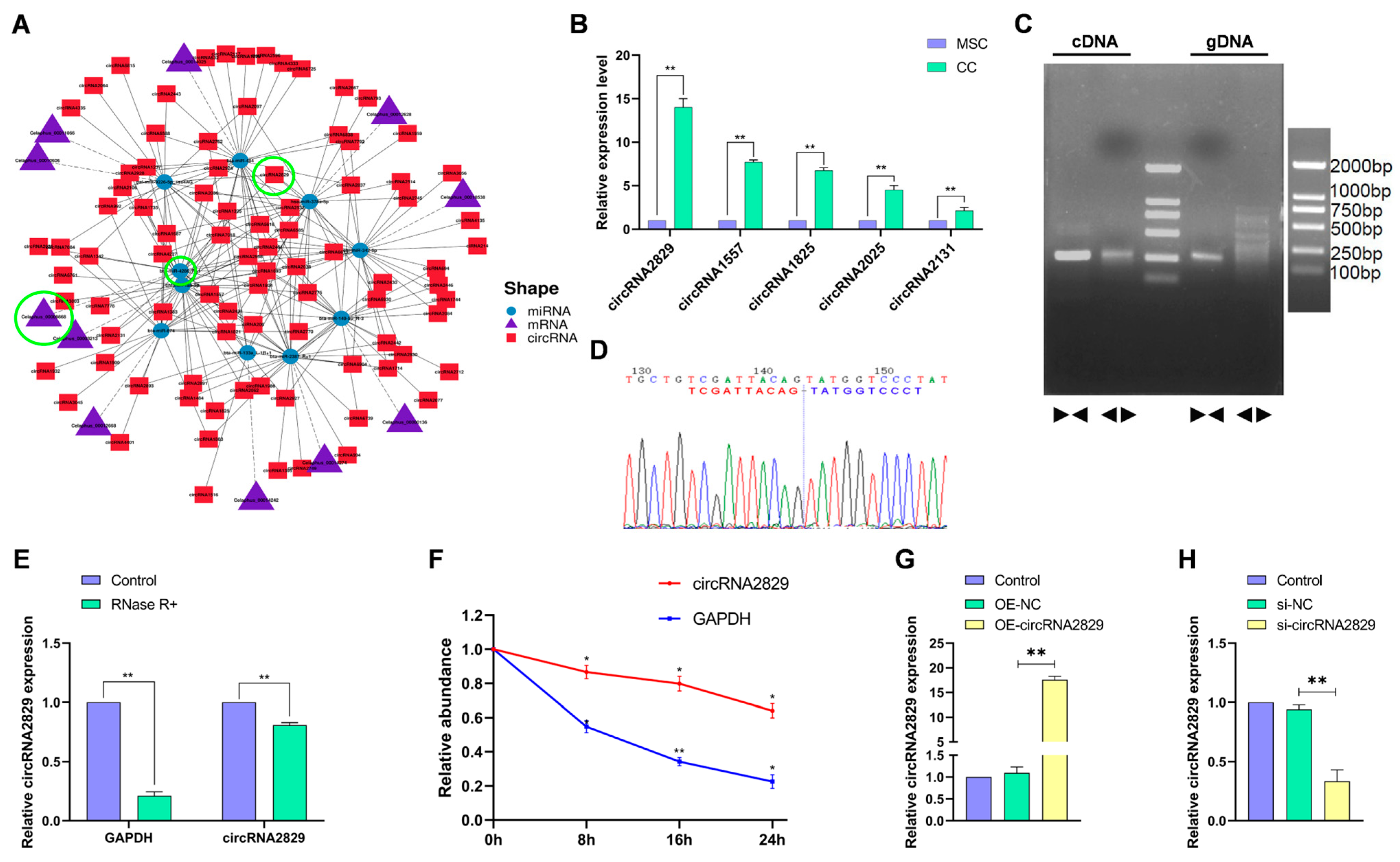
2.2. Validation of High-Throughput Sequencing Results
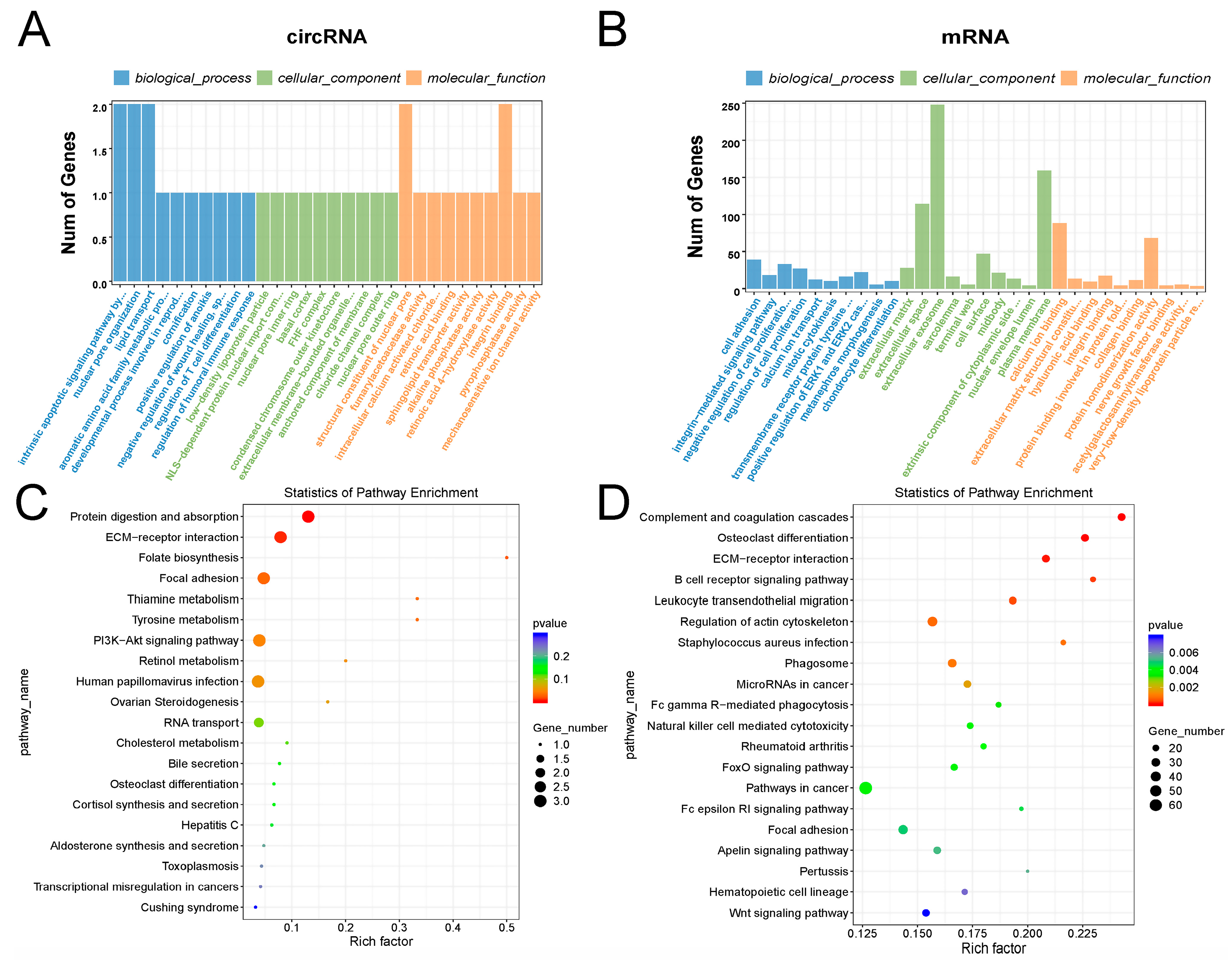
2.3. GO and KEGG Analysis of circRNAs and mRNAs
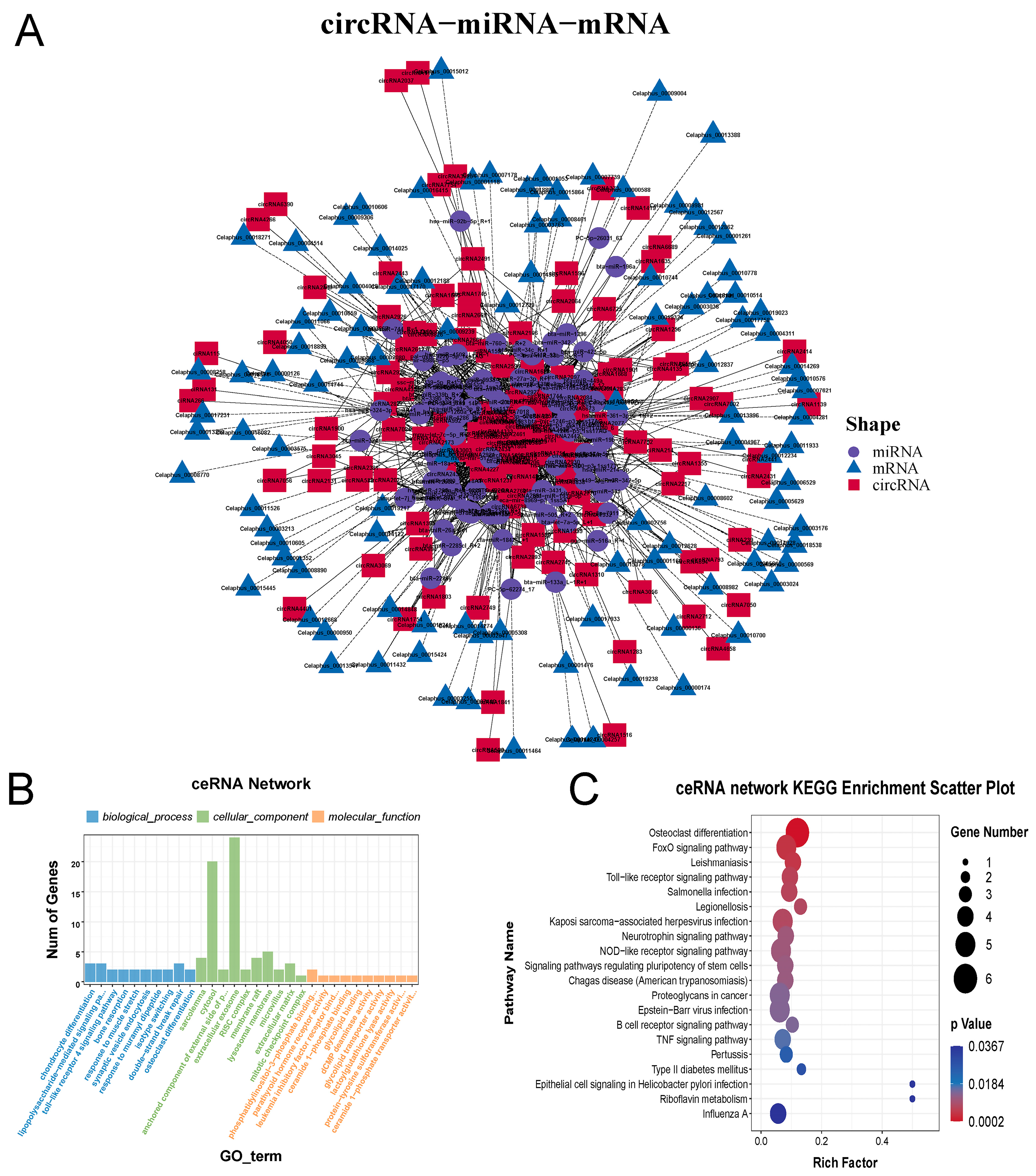
2.4. Construction and Analysis of circRNA-miRNA-mRNA ceRNA Network
2.5. CircRNA2829 Was Identified as a Circular RNA Related to Antler Regeneration and Differentiation
2.6. CircRNA2829 Promotes Chondrocyte Proliferation and Differentiation
2.7. MiR-4286-R+1 Inhibits Chondrocyte Proliferation and Differentiation
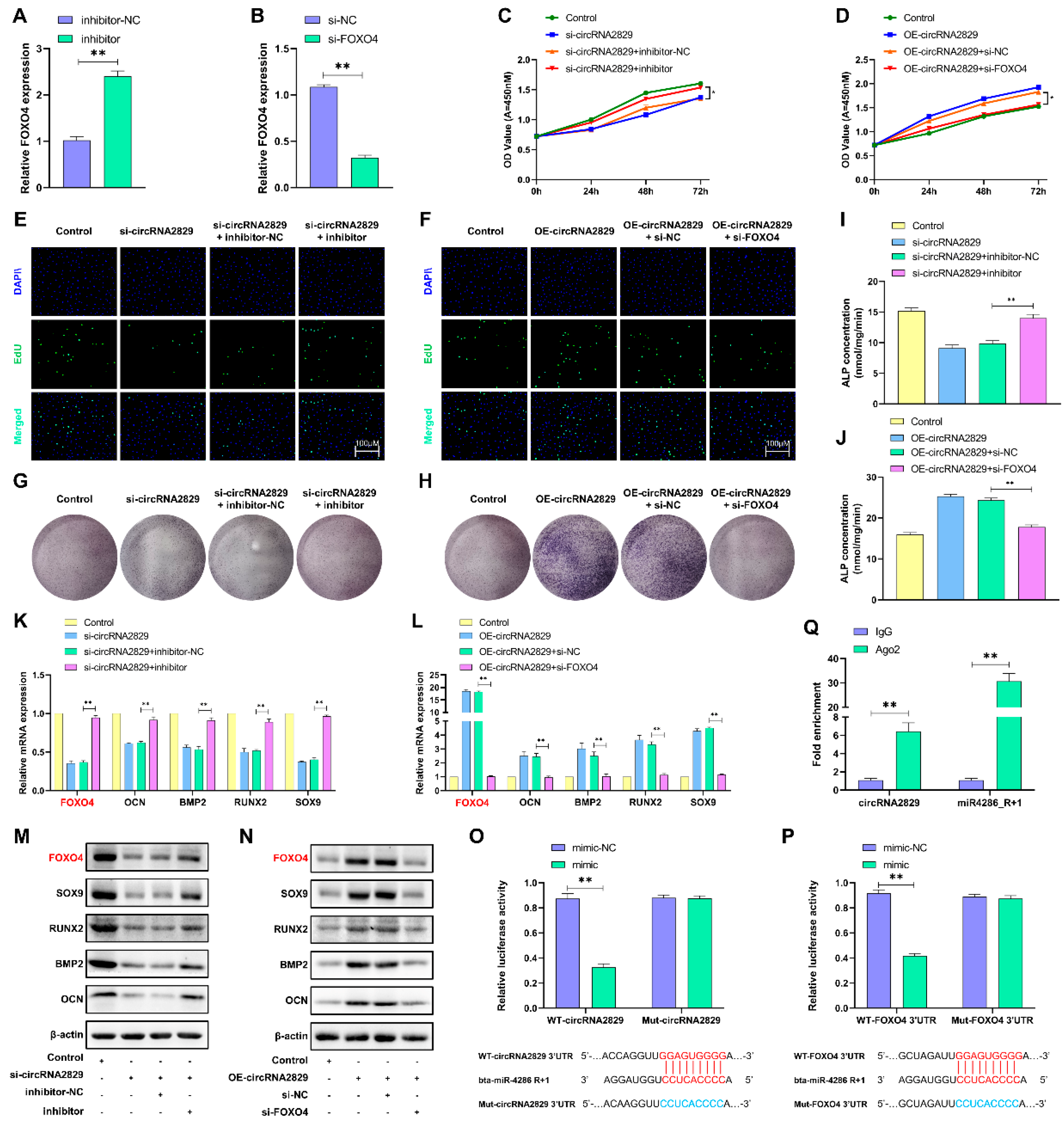
2.8. CircRNA2829 Promotes Chondrocyte Proliferation and Differentiation via the miR4286-R+1/FOXO4 Axis
3. Discussion
4. Materials and Methods
4.1. Sample Collection and Cell Culture
4.2. RNA-Seq and Sequencing Data Analysis
4.3. Real-Time Quantitative PCR
4.4. Construction of the ceRNA Networks
4.5. GO and KEGG Pathway Analysis
4.6. CircRNA Circular Structure Validation
4.7. Vector Construction and Cell Transfection
4.8. Cell Proliferation Assay
4.9. Alkaline Phosphatase (ALP) Staining and Alkaline Phosphatase Assay
4.10. Western Blot Assay
4.11. Dual-Luciferase Reporter Assay
4.12. RIP Assay
4.13. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Price, J.; Allen, S. Exploring the mechanisms regulating regeneration of deer antlers. Philos. Trans. R. Soc. London. Ser. B Biol. Sci. 2004, 359, 809–822. [Google Scholar] [CrossRef] [PubMed]
- Kierdorf, U.; Kierdorf, H.; Szuwart, T. Deer antler regeneration: Cells, concepts, and controversies. J. Morphol. 2007, 268, 726–738. [Google Scholar] [CrossRef] [PubMed]
- Barrell, G.K.; Davies, R.; Bailey, C.I. Immunocytochemical localization of oestrogen receptors in perichondrium of antlers in red deer (Cervus elaphus). Reprod. Fertil. Dev. 1999, 11, 189–192. [Google Scholar] [CrossRef] [PubMed]
- Allen, S.P.; Maden, M.; Price, J.S. A role for retinoic acid in regulating the regeneration of deer antlers. Dev. Biol. 2002, 251, 409–423. [Google Scholar] [CrossRef] [Green Version]
- Sun, Z.; Ma, Y.; Chen, F.; Wang, S.; Chen, B.; Shi, J. miR-133b and miR-199b knockdown attenuate TGF-β1-induced epithelial to mesenchymal transition and renal fibrosis by targeting SIRT1 in diabetic nephropathy. Eur. J. Pharmacol. 2018, 837, 96–104. [Google Scholar] [CrossRef]
- Wan, P.; Su, W.; Zhang, Y.; Li, Z.; Deng, C.; Li, J.; Jiang, N.; Huang, S.; Long, E.; Zhuo, Y. LncRNA H19 initiates microglial pyroptosis and neuronal death in retinal ischemia/reperfusion injury. Cell Death Differ. 2020, 27, 176–191. [Google Scholar] [CrossRef] [Green Version]
- Zheng, S.; Zhang, X.; Odame, E.; Xu, X.; Chen, Y.; Ye, J.; Zhou, H.; Dai, D.; Kyei, B.; Zhan, S.; et al. CircRNA-Protein Interactions in Muscle Development and Diseases. Int. J. Mol. Sci. 2021, 22, 3262. [Google Scholar] [CrossRef]
- Xiong, J.; Zhang, H.; Wang, Y.; Cheng, Y.; Luo, J.; Chen, T.; Xi, Q.; Sun, J.; Zhang, Y. Rno_circ_0001004 Acts as a miR-709 Molecular Sponge to Regulate the Growth Hormone Synthesis and Cell Proliferation. Int. J. Mol. Sci. 2022, 23, 1413. [Google Scholar] [CrossRef]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef]
- Kulcheski, F.R.; Christoff, A.P.; Margis, R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J. Biotechnol. 2016, 238, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Panda, A.C. Circular RNAs Act as miRNA Sponges. Adv. Exp. Med. Biol. 2018, 1087, 67–79. [Google Scholar] [CrossRef] [PubMed]
- Sen, R.; Ghosal, S.; Das, S.; Balti, S.; Chakrabarti, J. Competing endogenous RNA: The key to posttranscriptional regulation. Sci. World J. 2014, 2014, 896206. [Google Scholar] [CrossRef] [PubMed]
- Feleke, M.; Bennett, S.; Chen, J.; Hu, X.; Williams, D.; Xu, J. New physiological insights into the phenomena of deer antler: A unique model for skeletal tissue regeneration. J. Orthop. Transl. 2021, 27, 57–66. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.Y.; Li, Y.J.; Jiang, R.F.; Li, Y.T.; Feng, J.; Hu, W. Effects and mechanism of lncRNA-27785.1 that regulates TGF-β1 of Sika deer on antler cell proliferation. J. Cell. Physiol. 2021, 236, 5742–5756. [Google Scholar] [CrossRef]
- Chen, D.Y.; Yang, M.; Sun, Z.T.; Song, M.M.; Yao, H.B.; Long, G.H.; Hu, W. Notch4 affects the proliferation and differentiation of deer antler chondrocytes through the Smad3/lncRNA27785.1 axis. Cell. Signal. 2022, 98, 110429. [Google Scholar] [CrossRef]
- Yan, Y.; Chen, D.; Han, X.; Liu, M.; Hu, W. MiRNA-19a and miRNA-19b regulate proliferation of antler cells by targeting TGFBR2. Mammal Res. 2020, 65, 339–348. [Google Scholar] [CrossRef]
- Jiang, Q.; Liu, C.; Li, C.P.; Xu, S.S.; Yao, M.D.; Ge, H.M.; Sun, Y.N.; Li, X.M.; Zhang, S.J.; Shan, K.; et al. Circular RNA-ZNF532 regulates diabetes-induced retinal pericyte degeneration and vascular dysfunction. J. Clin. Investig. 2020, 130, 3833–3847. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, J.; Ma, J.; Sun, T.; Zhou, Q.; Wang, W.; Wang, G.; Wu, P.; Wang, H.; Jiang, L.; et al. Exosomal circRNAs: Biogenesis, effect and application in human diseases. Mol. Cancer 2019, 18, 116. [Google Scholar] [CrossRef] [Green Version]
- Xue, C.; Li, G.; Lu, J.; Li, L. Crosstalk between circRNAs and the PI3K/AKT signaling pathway in cancer progression. Signal. Transduct. Target. Ther. 2021, 6, 400. [Google Scholar] [CrossRef]
- Bjørklund, G.; Svanberg, E.; Dadar, M.; Card, D.J.; Chirumbolo, S.; Harrington, D.J.; Aaseth, J. The Role of Matrix Gla Protein (MGP) in Vascular Calcification. Curr. Med. Chem. 2020, 27, 1647–1660. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wei, R.; Wang, M.; Ma, L.; Zhang, Z.; Chen, L.; Guo, Q.; Guo, S.; Zhu, S.; Zhang, S.; et al. MGP Promotes Colon Cancer Proliferation by Activating the NF-κB Pathway through Upregulation of the Calcium Signaling Pathway. Mol. Ther. Oncolytics 2020, 17, 371–383. [Google Scholar] [CrossRef]
- Cui, C.; Bi, R.; Liu, W.; Guan, S.; Li, P.; Song, D.; Xu, R.; Zheng, L.; Yuan, Q.; Zhou, X.; et al. Role of PTH1R Signaling in Prx1(+) Mesenchymal Progenitors during Eruption. J. Dent. Res. 2020, 99, 1296–1305. [Google Scholar] [CrossRef] [PubMed]
- Amano, K.; Densmore, M.; Fan, Y.; Lanske, B. Ihh and PTH1R signaling in limb mesenchyme is required for proper segmentation and subsequent formation and growth of digit bones. Bone 2016, 83, 256–266. [Google Scholar] [CrossRef]
- Wu, P.P.; Hu, W.L.; Yin, C.C.; Fei, J.W. FOXO4 maintains senescence in human umbilical cord mesenchymal stem cells by repressing apoptosis. Sheng Li Xue Bao [Acta Physiol. Sin.] 2020, 72, 426–432. [Google Scholar]
- Ren, X.; Shan, W.H.; Wei, L.L.; Gong, C.C.; Pei, D.S. ACP5: Its Structure, Distribution, Regulation and Novel Functions. Anti-Cancer Agents Med. Chem. 2018, 18, 1082–1090. [Google Scholar] [CrossRef]
- Suttamanatwong, S.; Franceschi, R.T.; Carlson, A.E.; Gopalakrishnan, R. Regulation of matrix Gla protein by parathyroid hormone in MC3T3-E1 osteoblast-like cells involves protein kinase A and extracellular signal-regulated kinase pathways. J. Cell. Biochem. 2007, 102, 496–505. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Yu, J.; Wang, Q.; Zhang, L.; Chen, X.; Cao, Y.; Zhao, J.; Xu, Y.; Jiang, D.; Wang, Y.; et al. Tartrate-Resistant Acid Phosphatase 5/ACP5 Interacts with p53 to Control the Expression of SMAD3 in Lung Adenocarcinoma. Mol. Ther. Oncolytics 2020, 16, 272–288. [Google Scholar] [CrossRef] [Green Version]
- Lång, P.; Patlaka, C.; Andersson, G. Tartrate-resistant acid phosphatase type 5/ACP5 promotes cell cycle entry of 3T3-L1 preadipocytes by increasing IGF-1/Akt signaling. FEBS Lett. 2021, 595, 2616–2627. [Google Scholar] [CrossRef]
- Gong, C.; Ai, J.; Fan, Y.; Gao, J.; Liu, W.; Feng, Q.; Liao, W.; Wu, L. NCAPG Promotes The Proliferation of Hepatocellular Carcinoma through PI3K/AKT Signaling. Onco. Targets Ther. 2019, 12, 8537–8552. [Google Scholar] [CrossRef] [Green Version]
- Baar, M.P.; Brandt, R.M.C.; Putavet, D.A.; Klein, J.D.D.; Derks, K.W.J.; Bourgeois, B.R.M.; Stryeck, S.; Rijksen, Y.; van Willigenburg, H.; Feijtel, D.A.; et al. Targeted Apoptosis of Senescent Cells Restores Tissue Homeostasis in Response to Chemotoxicity and Aging. Cell 2017, 169, 132–147.e116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, U.; Pal, D.; Prasad, R. Alkaline phosphatase: An overview. Indian J. Clin. Biochem. 2014, 29, 269–278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schirle, N.T.; Sheu-Gruttadauria, J.; MacRae, I.J. Structural basis for microRNA targeting. Science 2014, 346, 608–613. [Google Scholar] [CrossRef]
- Almeida, M. Unraveling the role of FoxOs in bone–insights from mouse models. Bone 2011, 49, 319–327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, X.; Su, P.; Yin, C.; Lin, X.; Wang, X.; Gao, Y.; Patil, S.; War, A.R.; Qadir, A.; Tian, Y.; et al. The Roles of FoxO Transcription Factors in Regulation of Bone Cells Function. Int. J. Mol. Sci. 2020, 21, 692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, D.; Gong, Y.; Xu, L.; Zhou, M.; Li, J.; Song, J. Bidirectional regulation of osteogenic differentiation by the FOXO subfamily of Forkhead transcription factors in mammalian MSCs. Cell Prolif. 2019, 52, e12540. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, C.; Clark, D.E.; Lord, E.A.; Stanton, J.A.; Suttie, J.M. Sampling technique to discriminate the different tissue layers of growing antler tips for gene discovery. Anat. Rec. 2002, 268, 125–130. [Google Scholar] [CrossRef]
- Kechin, A.; Boyarskikh, U.; Kel, A.; Filipenko, M. cutPrimers: A New Tool for Accurate Cutting of Primers from Reads of Targeted Next Generation Sequencing. J. Comput. Biol. 2017, 24, 1138–1143. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [Green Version]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frazee, A.C.; Pertea, G.; Jaffe, A.E.; Langmead, B.; Salzberg, S.L.; Leek, J.T. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 2015, 33, 243–246. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kong, L.; Zhang, Y.; Ye, Z.-Q.; Liu, X.-Q.; Zhao, S.-Q.; Wei, L.; Gao, G. CPC: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35, W345–W349. [Google Scholar] [CrossRef]
- Sun, L.; Luo, H.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Liu, Y.; Chen, R.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.; Salzberg, S.L. TopHat-Fusion: An algorithm for discovery of novel fusion transcripts. Genome Biol. 2011, 12, R72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.-O.; Dong, R.; Zhang, Y.; Zhang, J.-L.; Luo, Z.; Zhang, J.; Chen, L.-L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.O.; Wang, H.B.; Zhang, Y.; Lu, X.; Chen, L.L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef] [Green Version]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [Green Version]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, H.; Jiang, R.; Chen, D.; Li, Y.; Song, M.; Sun, Z.; Long, G.; Wu, L.; Hu, W. Whole-Transcriptome Sequencing of Antler Tissue Reveals That circRNA2829 Regulates Chondrocyte Proliferation and Differentiation via the miR-4286-R+1/FOXO4 Axis. Int. J. Mol. Sci. 2023, 24, 7204. https://doi.org/10.3390/ijms24087204
Yao H, Jiang R, Chen D, Li Y, Song M, Sun Z, Long G, Wu L, Hu W. Whole-Transcriptome Sequencing of Antler Tissue Reveals That circRNA2829 Regulates Chondrocyte Proliferation and Differentiation via the miR-4286-R+1/FOXO4 Axis. International Journal of Molecular Sciences. 2023; 24(8):7204. https://doi.org/10.3390/ijms24087204
Chicago/Turabian StyleYao, Haibo, Renfeng Jiang, Danyang Chen, Yanjun Li, Mengmeng Song, Zitong Sun, Guohui Long, Lei Wu, and Wei Hu. 2023. "Whole-Transcriptome Sequencing of Antler Tissue Reveals That circRNA2829 Regulates Chondrocyte Proliferation and Differentiation via the miR-4286-R+1/FOXO4 Axis" International Journal of Molecular Sciences 24, no. 8: 7204. https://doi.org/10.3390/ijms24087204




