Abstract
The WUSCHEL-related homeobox (WOX) transcription factor plays a vital role in stem cell maintenance and organ morphogenesis, which are essential processes for plant growth and development. Dendrobium chrysotoxum, D. huoshanense, and D. nobile are valued for their ornamental and medicinal properties. However, the specific functions of the WOX gene family in Dendrobium species are not well understood. In our study, a total of 30 WOX genes were present in the genomes of the three Dendrobium species (nine DchWOXs, 11 DhuWOXs, and ten DnoWOXs). These 30 WOXs were clustered into ancient clades, intermediate clades, and WUS/modern clades. All 30 WOXs contained a conserved homeodomain, and the conserved motifs and gene structures were similar among WOXs belonging to the same branch. D. chrysotoxum and D. huoshanense had one pair of fragment duplication genes and one pair of tandem duplication genes, respectively; D. nobile had two pairs of fragment duplication genes. The cis-acting regulatory elements (CREs) in the WOX promoter region were mainly enriched in the light response, stress response, and plant growth and development regulation. The expression pattern and RT-qPCR analysis revealed that the WOXs were involved in regulating the floral organ development of D. chrysotoxum. Among them, the high expression of DchWOX3 suggests that it might be involved in controlling lip development, whereas DchWOX5 might be involved in controlling ovary development. In conclusion, this work lays the groundwork for an in-depth investigation into the functions of WOX genes and their regulatory role in Dendrobium species’ floral organ development.
1. Introduction
The homeobox transcription factors (HB TFs) are key regulators of plant and animal cell fates and differentiation, and homeobox genes were first discovered in Drosophila [1,2]. Meanwhile, more homeobox members continue to be found in other eukaryotes. The WUSCHEL (WUS) gene is the prototypic member of the plant-specific WUS homeobox (WOX) protein family, one of several HB TF families [3]. A total of 14 homologs of AtWUS were searched in the Arabidopsis genome, and these genes were named WOXs [4]. The WOX TFs contain a short stretch of amino acids that folds into a DNA-binding domain (called a homeodomain), which forms helix–loop–helix–turn–helix structures in space [5].
In plants, the WOX genes are extensively distributed. According to the evolutionary origins among genes, the members of the WOX gene family in plants can be clustered into three clades: the ancient clade, the intermediate clade, and the WUS/modern clade. All plant species (from algae to angiosperms) contain varying amounts of WOX genes belonging to the ancient clade; the intermediate clade is found in pteridophytes, gymnosperms, and angiosperms; and the WUS/modern clade is exclusively found in angiosperms [6]. In the WOX gene family of Arabidopsis, there are three members (AtWOX10, AtWOX13–14) in the ancient clade, four members (AtWOX8–9, AtWOX11–12) in the intermediate clade, and eight members (AtWUS, AtWOX1–7) in the WUS/modern clade.
The WOX gene family is involved in plant growth and development, as well as in the stress response. WOX family members belonging to different clades fulfill different biological functions in the development of plant flowers, floral meristems, roots, and other organs. The WOX genes of the ancient clade participate in the regulation of plant roots and flower development. AtWOX13 is expressed in floral meristem tissues, inflorescences, and young flower buds and is particularly highly expressed in developing carpels. WOX13 promotes replum development by negatively regulating the JAG/FIL genes [7]. AtWOX14 is found only in Brassicaceae, where it is expressed early in lateral root formation and specific to the development of anthers [8]. The intermediate clade mainly affects embryo patterning and root organogenesis. WOX8 and WOX9 are homologous genes that play vital roles in embryo and inflorescence development and are species-specific in their functions [2,9,10,11]. The genes WOX11 and WOX12, which are homologous, participate in the process of de novo root organogenesis in Arabidopsis [12]. The WUS clade mainly affects the development of the floral meristem and leaf and stem cell maintenance. For example, Arabidopsis WUS genes can maintain stem cell homeostasis at all developmental stages in the shoot apical meristem (SAM) [13,14]. Meanwhile, WUS genes are also able to act as activators to regulate the size of the floral meristem tissue [15]. WOX1 and WOX3 redundantly regulate abaxial–adaxial growth in the leaf and floral meristem [16,17]. AtWOX2 is required to initiate the embryogenic shoot meristem stem cell program in Arabidopsis [18]. WOX5 is critical for stem cell maintenance in the root apical meristem (RAM) [19,20]. In addition, the WOX gene family plays an important role in the response to environmental stresses, such as salt, cold, and drought. For example, GhWOX4 positively regulates drought tolerance in cotton; PagWOX11/12a positively regulates the salt tolerance of poplar [21,22].
The WOX genes act as transcription factors to activate or repress the expression of other genes on the one hand, as described above for the role played by WOXs in plants. On the other hand, the upstream part of the WOX coding region contains abundant CREs to receive the action of other regulatory factors. For example, maize ZMSP10/14/26 regulates the expression of the ZmWOX3A gene in coat precursor cells by directly binding to its promoter [23]. In summary, the combination of cis- and trans-acting factors exerts a regulatory effect on gene expression, while playing an indispensable role in plant growth, development, and evolution [24].
Orchidaceae, one of the largest angiosperm groups, contains over 750 genera and 28,000 species [25,26]. It is widely distributed, with the exception of the North and South Poles and extremely arid desert areas, and has the greatest distribution in the tropics. Orchids are highly evolved taxa within angiosperms and are one of the most studied taxa in biological research [27]. Dendrobium is the second-largest genus in the orchid family and is a typical epiphyte [28]. Most Dendrobium species have valuable medicinal stems, while their flowers and leaves have excellent ornamental value. In recent years, the completion of the whole-genome sequencing of D. catenatum [29], D. chrysotoxum [30], D. huoshanense [31], and D. nobile [32], etc., has provided valuable information revealing the genetic and molecular mechanisms of the formation of important traits in Dendrobium. The regulatory function of the WOX genes in model plants such as Arabidopsis has been relatively comprehensively researched. However, there is little knowledge about how the WOX genes affect the growth and development of Dendrobium species.
In our study, we identified the WOX gene family in three Dendrobium species (D. chrysotoxum, D. huoshanense, and D. nobile), and systematically analyzed their basic traits, including their chromosomal localization, phylogenetics, motif compositions, gene structures, collinearity, and CREs. Meanwhile, the expression pattern of WOXs in the D. chrysotoxum flower parts was analyzed. This project aimed to preliminarily elucidate the evolutionary and potential biological roles of the WOX gene family in Dendrobium species, and to provide new insights into the study of the molecular regulatory mechanisms of the WOX genes in D. chrysotoxum flower development.
2. Results
2.1. Identification and Physicochemical Properties of the WOX Gene Family
The WOX genes in three Dendrobium species were screened by BLAST and HMMER. The result showed that ten, nine, and 11 WOXs were identified in the genomes of D. chrysotoxum, D. huoshanense, and D. nobile, respectively. According to the order distribution of the chromosomes, these WOXs were named DchWOX1–10, DhuWOX1–9, and DnoWOX1–11.
To characterize the WOX genes of the Dendrobium species in more detail, we predicted the physicochemical properties of 30 WOX proteins using ExPASy. The results are as follows (Table 1). The number of amino acids (AA) varied from 110 aa (DchWOX1) to 328 aa (DchWOX4), and the molecular weight (Mw) ranged from 12.84 kDa (DchWOX1) to 35.30 kDa (DnoWOX4). Among the 30 WOXs, 12 were basic proteins with an isoelectric point (pI) higher than 8.00; the remaining 17, with a pI ranging from 5.26 (DhuWOX8) to 7.79 (DnoWOX4), were neutral or weakly acidic proteins. Additionally, the grand average of hydrophilic (GRAVY) values of all WOX proteins were less than zero, suggesting their strong hydrophilicity. The instability indexes (II) of all WOX members exceeded 40, implying that these proteins are unstable [33]. All WOX proteins were found to be in the nucleus according to subcellular location predictions, indicating that they might also function there, like most TFs.

Table 1.
Characteristics of the WOXs from three Dendrobium species.
2.2. Chromosomal Localization of WOXs
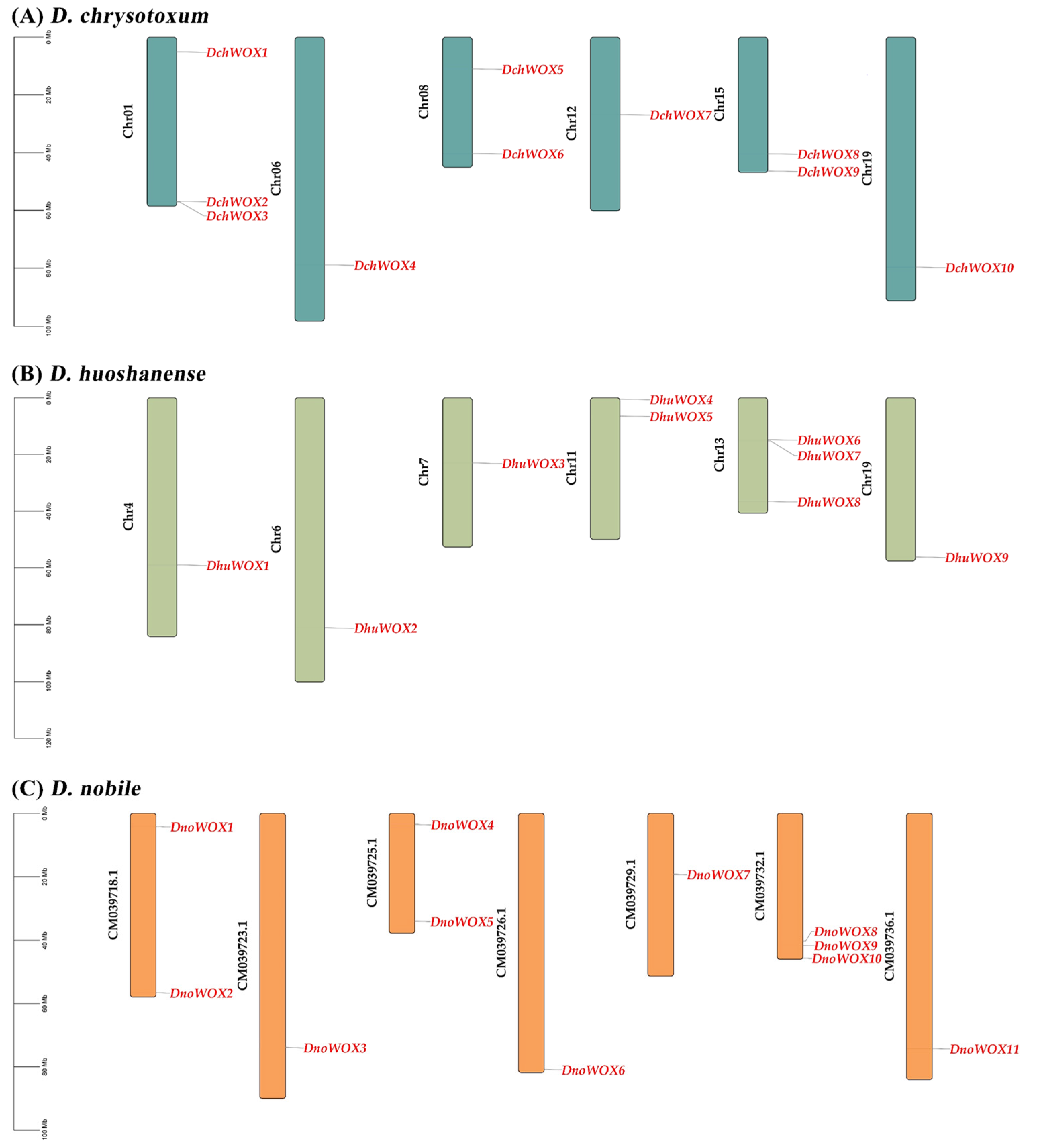
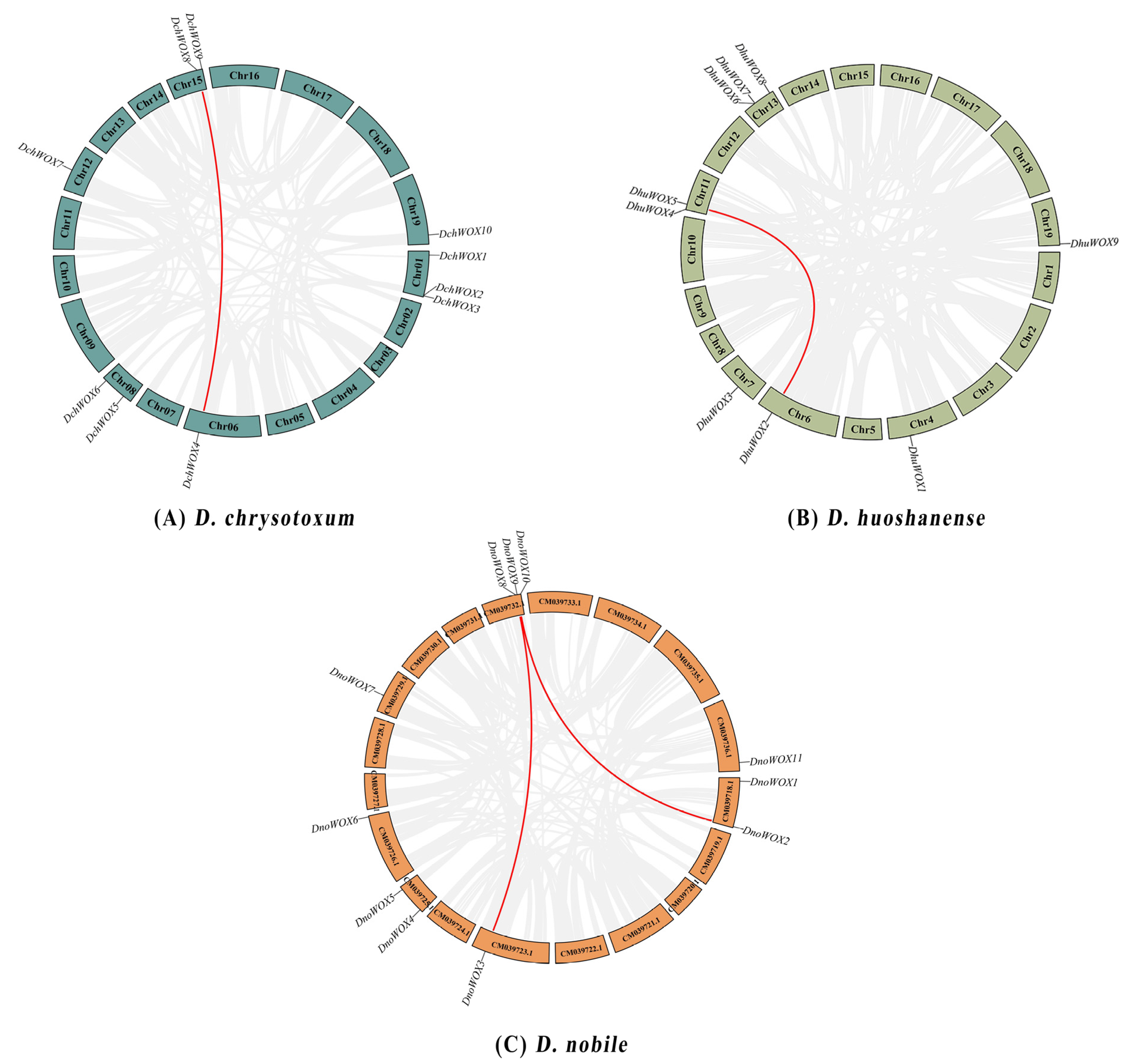
As illustrated in Figure 1A, ten DchWOXs were unevenly present on six chromosomes of D. chrysotoxum (Chr01, 06, 08, 12, 15, and 19) (Figure 1A). Nine DhuWOXs were distributed on six chromosomes, Chr4, 6, 7, 11, 13, and 19, of D. huoshanense (Figure 1B). The results of the chromosome mapping for D. nobile showed that 11 DnoWOXs were distributed across seven chromosomes (Figure 1C). In addition, we observed a pair of tandem repeat genes in D. chrysotoxum (DchWOX2 and DchWOX3) and D. huoshanense (DhuWOX6 and DhuWOX7).
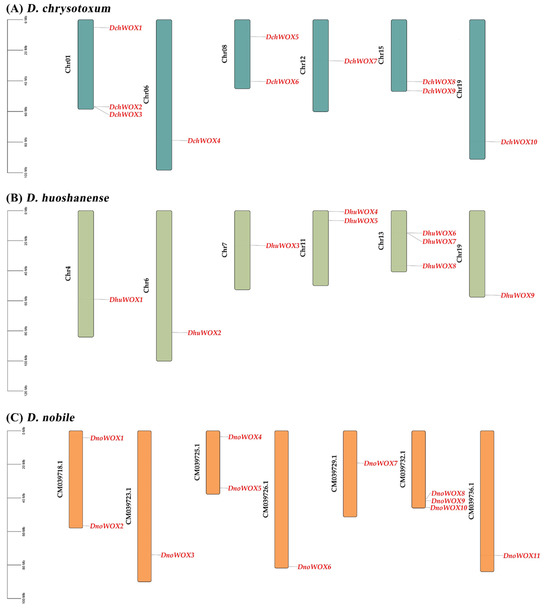
Figure 1.
Chromosome distribution of WOX genes. (A) D. chrysotoxum. (B) D. huoshanense. (C) D. nobile. Black labels are chromosome names and red labels are gene names.
2.3. Phylogenetic Analysis of WOXs
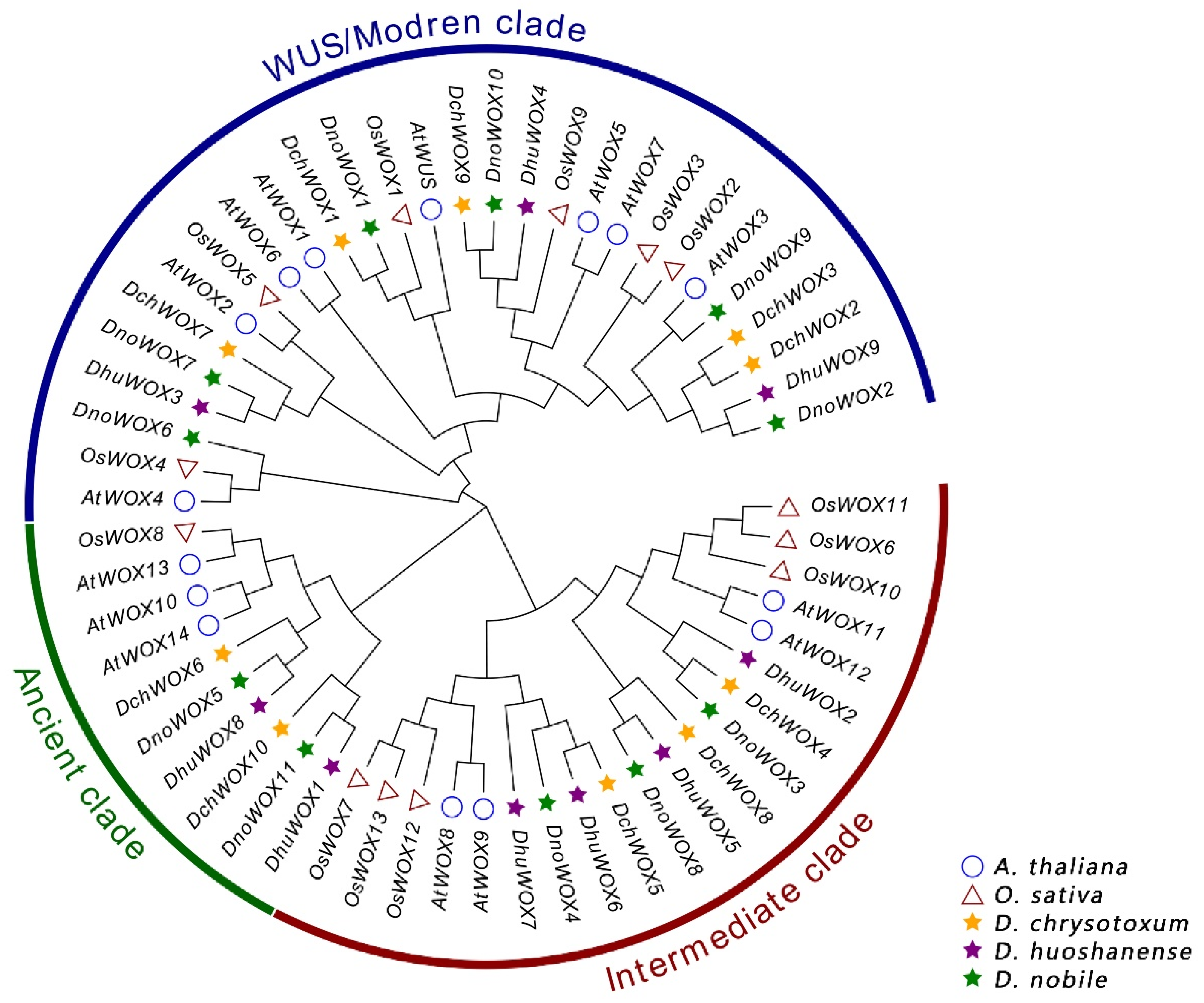
We created a phylogenetic tree of the WOX genes to analyze the evolution of the WOX genes in the Dendrobium species (Figure 2). The evolutionary tree included 30 WOXs from three Dendrobium species, 15 AtWOXs from A. thaliana, and 13 OsWOXs from O. sativa. All WOX protein sequences have been collected with Table S1. According to the classification of the WOXs’ evolutionary relationships in A. thaliana, the 30 WOXs in the Dendrobium species can be similarly clustered into the ancient clade (six WOX genes), the intermediate clade (ten WOX genes), and the WUS/modern clade (14 WOX genes). The WUS/modern clade has the largest number of WOX genes, while the ancient clade has the fewest.
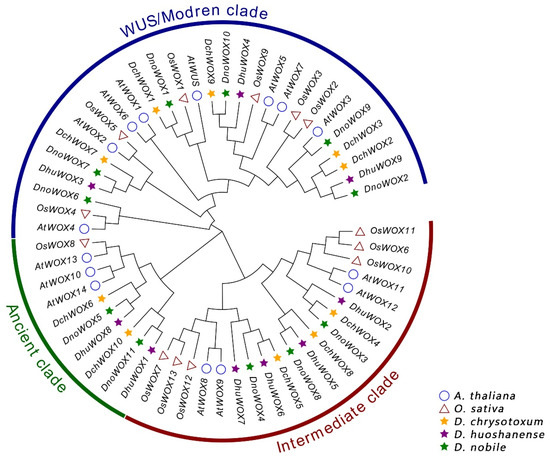
Figure 2.
Phylogenetic tree of WOXs in A. thaliana, O. sativa, and three Dendrobium species.
2.4. Gene Structure and Conserved Motifs of WOXs
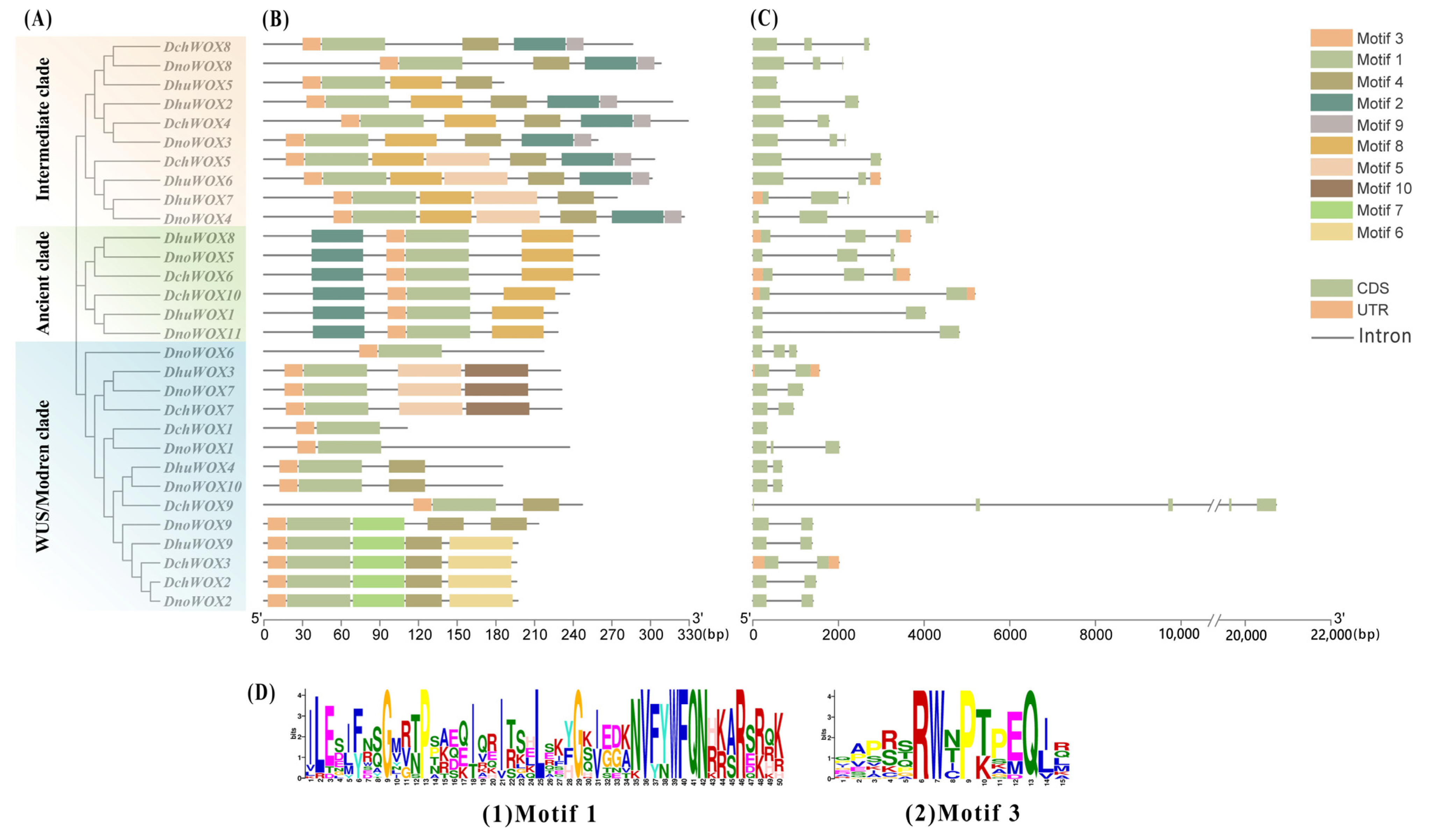
The conserved motifs of the 30 WOXs in the three Dendrobium species were evaluated through the online prediction website MEME (Figure 3B). The results demonstrated that, whereas the motif structures varied by clade, WOXs within the same clade had comparable motif structures. Ten conserved motifs were detected in the 30 WOXs. Table S2 has listed all motif sequences. All WOXs contain motif 1 and motif 3 simultaneously; motif 6, motif 7, and motif 10 are exclusive to the WUS/modern clade; and motif 9 is found only in the intermediate clade. The distinct roles of various WOXs may be conferred by the particular distributions of various structures.
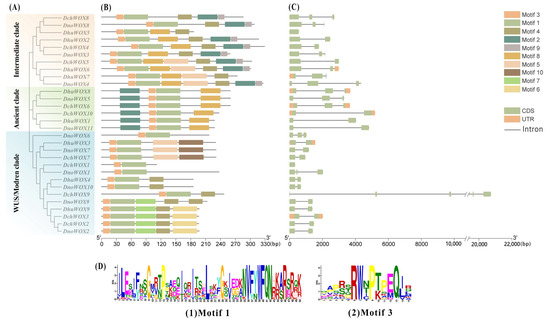
Figure 3.
Phylogenetic tree, motifs, and structures of WOXs in three Dendrobium species. (A) Phylogenetic tree of 30 WOXs. (B) Conserved motifs of 30 WOX proteins. (C) Intron and exon structures of 30 WOX genes. (D) Sequence logos of motif 1 and motif 3.
We visualized the number and distribution of the WOXs’ introns and exons to further reveal the gene structures of the WOXs in the three Dendrobium species (Figure 3C). Most Dendrobium WOXs contain 1–2 introns. Notably, three introns were detected in DnoWOX4 and four introns were detected in DchWOX9, while DchWOX1 and DhuWOX5 had no introns. The gene structures of WOX members belonging to the same clade are similar. In particular, in the ancient clade, the phylogenetic tree divides six genes into two structurally similar subclades. DhuWOX8, DnoWOX5, and DchWOX6 are clustered as subclades with two introns, while DchWOX10, DhuWOX1, and DnoWOX11 are clustered as a subclade with one intron.
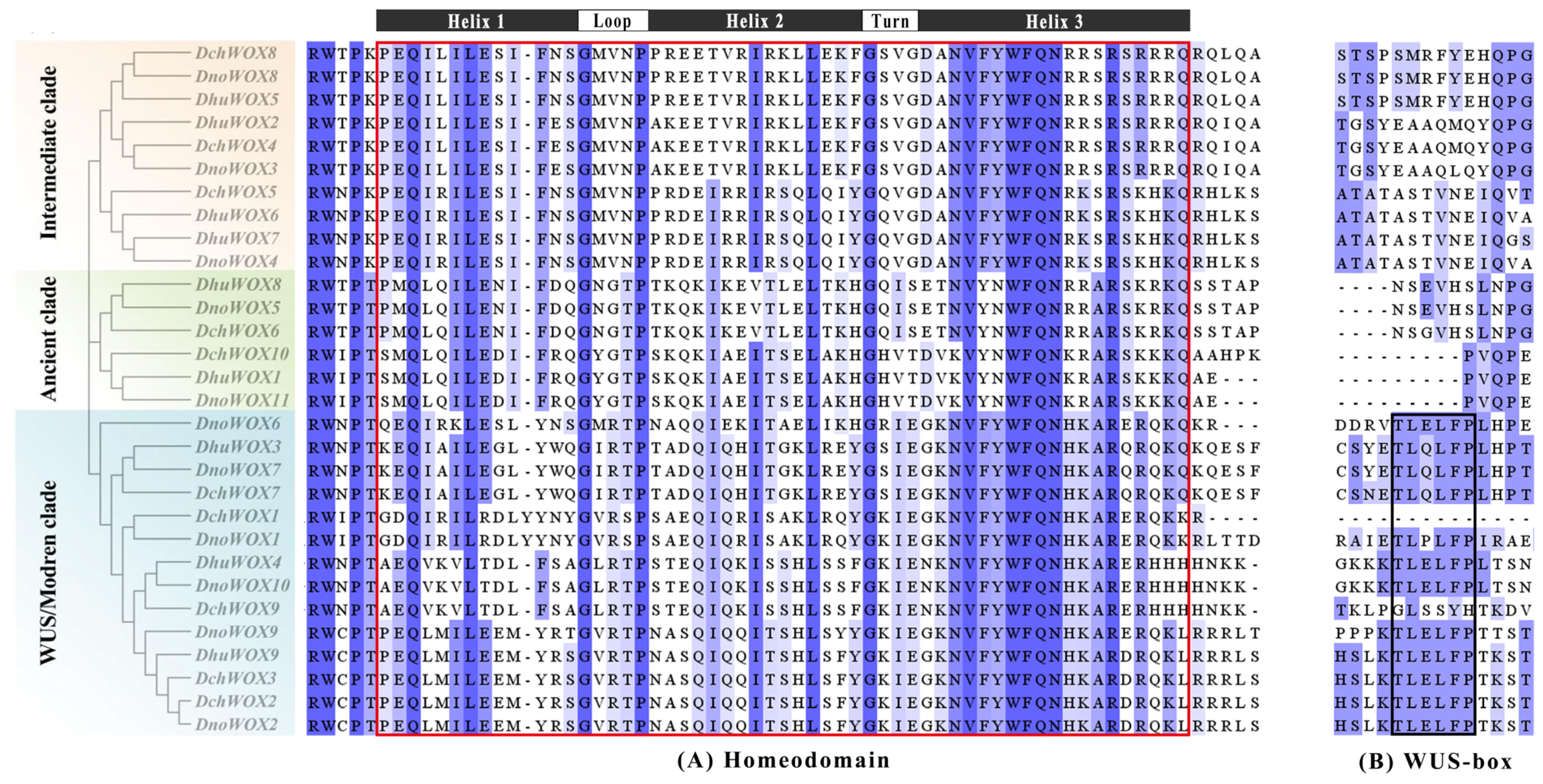
Multiple sequence pairs of the 30 WOXs showed that all WOXs contained a helix–turn–helix–loop–helix region unique to the homeodomain (Figure 4A). Twelve WOXs in the WUS/modern clade contain the WUS-box (TL-LFP-) (Figure 4B).
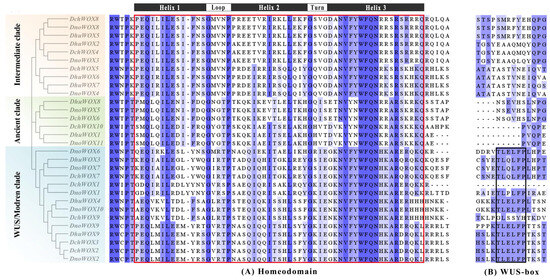
Figure 4.
Multiple sequence alignment results of the WOX gene family in three Dendrobium species. (A) Homeodomain. (B) WUS-box. The red box indicates the homeodomain and the black box indicates the WUS-box domain.
2.5. Synteny Analysis and Ka/Ks Value of WOX Gene Family
The D. chrysotoxum genome contains a pair of segmental duplication genes, DchWOX4 and DchWOX8 on Chr06 and Chr15 (Figure 5A). Similarly, the D. huoshanense genome contains one pair of fragment duplication genes, DhuWOX2 and DhuWOX5 on Chr6 and Chr11 (Figure 5B). Two pairs of segmental duplicates were found in the D. nobile genome, DnoWOX2 and DnoWOX9 on CM039718.1 and CM039732.1, and DnoWOX3 and DnoWOX8 on CM039723.1 and CM039732.1, respectively (Figure 5C). Furthermore, the Ka/Ks ratios of these four gene pairs were all less than 0.5, ranging from 0.13 to 0.2 (Table 2).
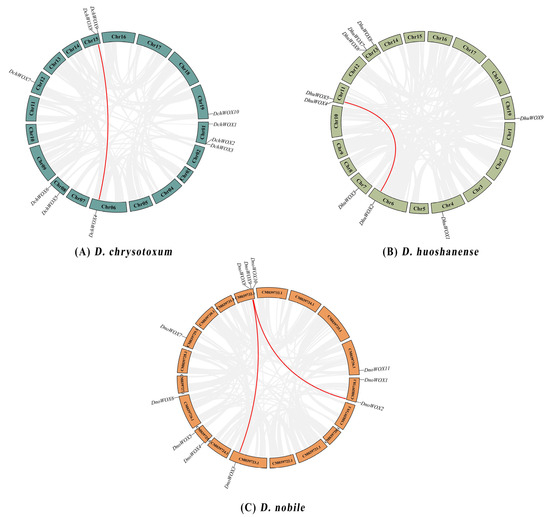
Figure 5.
Synteny analysis of the WOX gene family. (A) D. chrysotoxum. (B) D. huoshanense. (C) D. nobile. The red lines refer to segmental duplicate gene pairs.

Table 2.
The Ka/Ks of the WOX gene family in the three Dendrobium species.
2.6. Cis-Acting Elements Analysis of WOXs
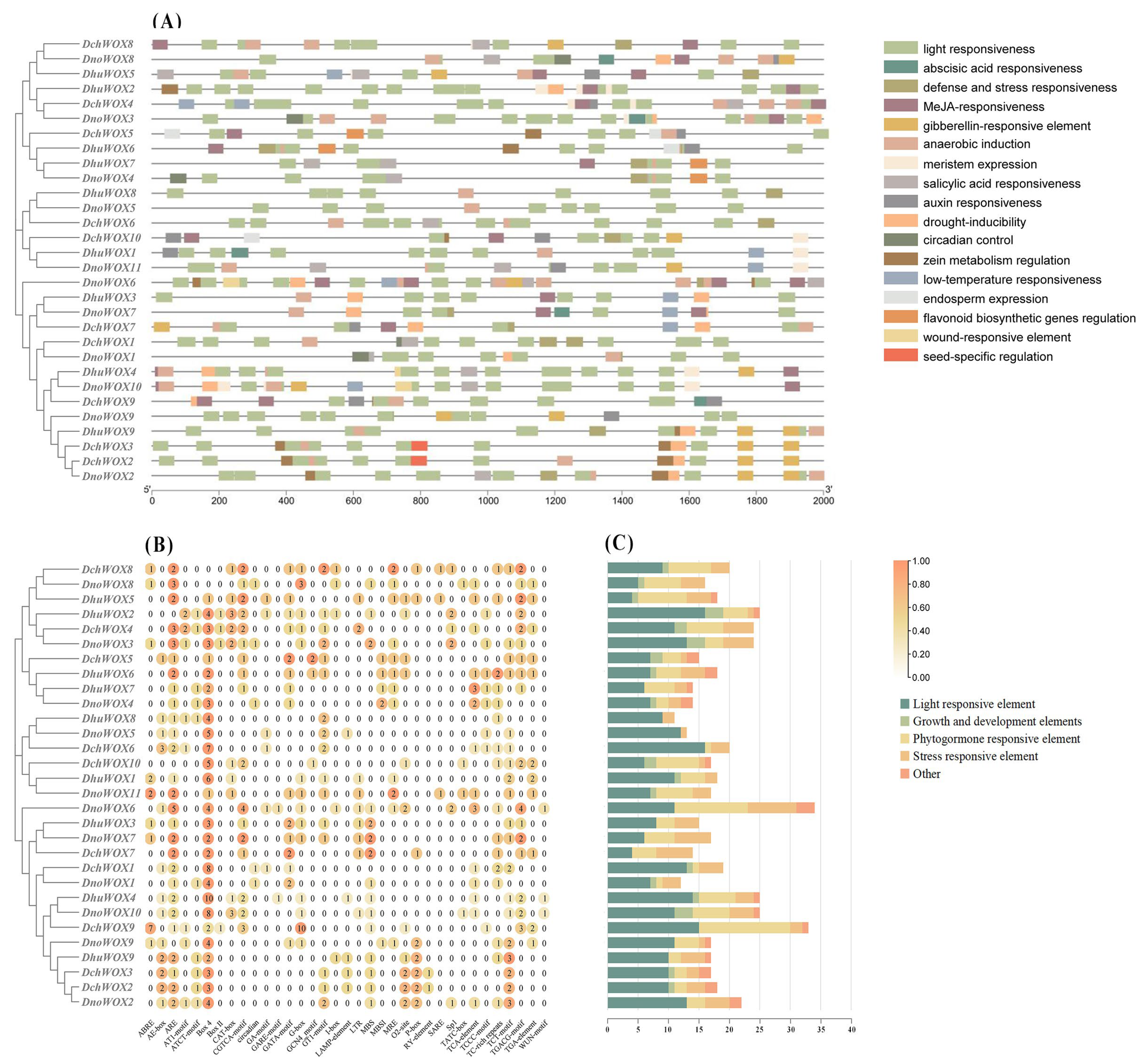
We extracted 2000 bp upstream of the CDS of the 30 WOX genes to identify the CREs to predict the potential regulatory functions of the WOX genes in the Dendrobium species. In total, 569 CREs belonging to 35 types and 17 response functions were found in the three Dendrobium species (Figure 6 and Table S3).
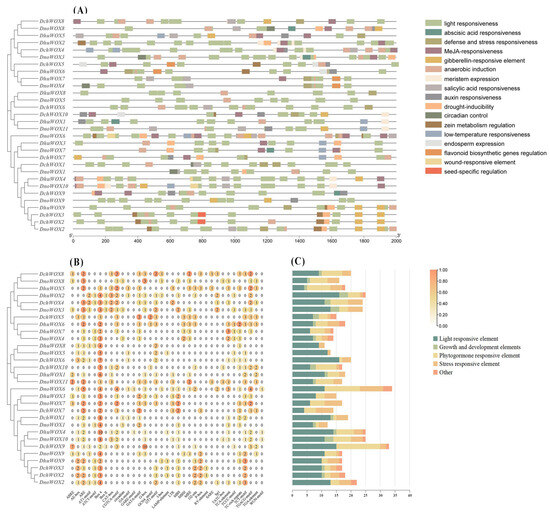
Figure 6.
The CREs in the promoter regions of 30 WOX genes. (A) Distribution of the WOX CREs; (B) the number of CREs; (C) statistics on the number of different categories of CREs. The types and numbers of CREs are listed in Table S3.
We classified the retrieved CREs into four categories: growth and development elements, phytohormone responsiveness, stress repressiveness, and light responsiveness. The growth and development element category includes endosperm expression, circadian control, and meristem expression. Interestingly, among them, the frequency of meristem expression is the largest. Five types of CRE exist within the category of phytohormone responsiveness. This includes abscisic acid (ABA), methyl jasmonate (MeJA), auxin, gibberellin, and salicylic acid responsiveness. The stress repressiveness category had four types of CRE, including defense and stress responsiveness, anaerobic induction, drought, and low-temperature stress, with anaerobic induction being the most frequent. In addition, light responsiveness accounts for almost half of all CREs (269/569), and there is a large frequency of light responsiveness in each WOX gene.
As shown in Figure 6C, DchWOX9 in D. chrysotoxum has the largest number (33 CREs) of elements, DhuWOX2 and DhuWOX4 in D. huoshanense have the most (25 CREs), and DnoWOX6 in D. nobile has the most (34 CREs).
2.7. Expression Pattern Analysis of WOX Gene Family in D. chrysotoxum
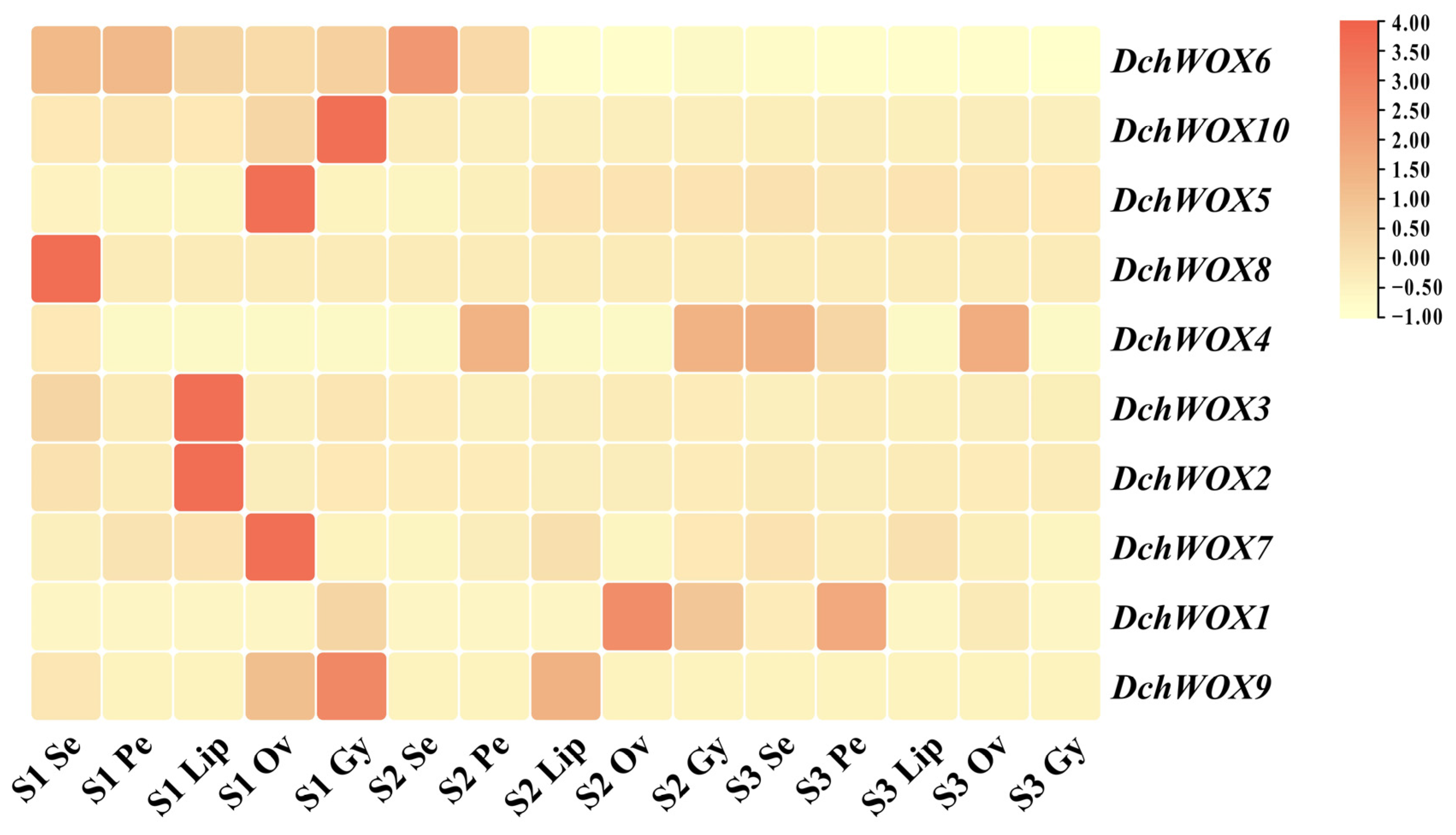
We performed expression analyses based on transcriptome data from different flower parts in the three developmental periods of D. chrysotoxum (Figure 7). In the transcriptome heatmap, DchWOX6 and DchWOX10 of the ancient clade were expressed at significantly higher levels in S1. However, DchWOX6 was similarly expressed in all five floral parts, whereas DchWOX10 exhibited high expression only in the gynostemium. Of the three members of the intermediate clade, DchWOX5 had higher expression in the ovary of S1, DchWOX8 in the sepal of S1, and DchWOX4 had lower expression in S1 than S2 and S3. Among the five WOXs of the WUS/modern clade, DchWOX2 and DchWOX3 displayed similar expression levels throughout flower development, and they had higher expression amounts in the lip of S1; DchWOX7 had higher expression in the ovary of S1; DchWOX1 was highly expressed in the ovary of S2; and DchWOX9 gynostemium expression was highest in S1.
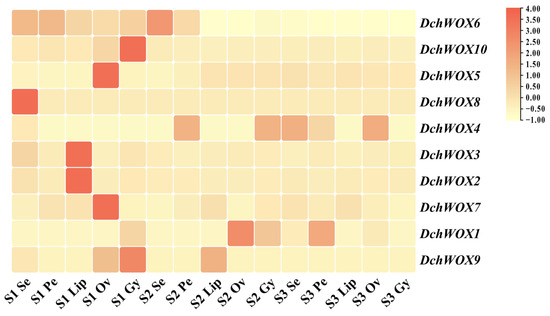
Figure 7.
The expression levels of ten DchWOXs in D. chrysotoxum at five floral parts and three developmental periods. S1: unpigmented bud stage; S2: pigmented bud stage; S3: early flowering stage; Se: sepal; Pe: petal; Lip: lip; Ov: ovary; Gy: gynostemium. Table S4 lists the FPKM values for the WOXs in D. chrysotoxum.
2.8. RT-qPCR Analysis of WOX Genes in D. chrysotoxum
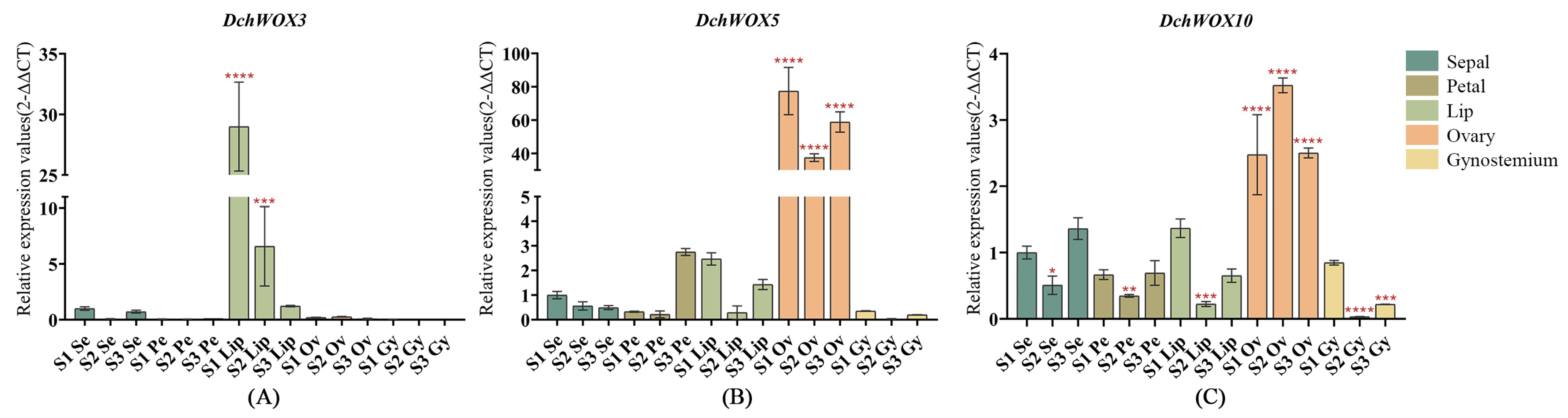
We selected DchWOX3, DchWOX5, and DchWOX10 from different clades for RT-qPCR experiments to further elucidate the expression patterns of the WOXs during the development of different flower parts in D. chrysotoxum (Figure 8). As shown, the DchWOX3 RT-qPCR results are in general agreement with the transcriptome data, i.e., DchWOX3 showed very low expression in other parts of the flower, while it was significantly expressed in the S1 lip, and its expression was gradually downregulated during flower development (Figure 8A). DchWOX5 was consistently expressed in the ovary during the three periods, suggesting that DchWOX5 is involved in regulating ovary development (Figure 8B). The transcriptome expression heatmap showed that DchWOX10 was significantly expressed during S1 in the gynostemium. However, the RT-qPCR results indicated a trend of increasing followed by decreasing expression of DchWOX10 (Figure 8C). These differences may have resulted from imperfect correlations between the samples used for transcriptome sequencing and the samples used for RT-qPCR.
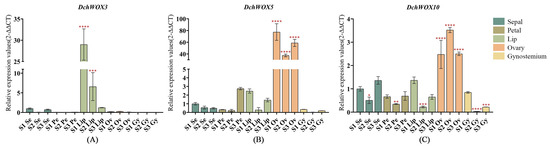
Figure 8.
RT-qPCR verified the expression of DchWOXs in the developing flower organs of D. chrysotoxum. (A) DchWOX3; (B) DchWOX5; (C) DchWOX10. The Y-axis represents the relative expression values (2−∆∆CT). The error bars indicate three RT-qPCR biological replicates. p-values in the significance test are indicated by red asterisks (* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001). Table S5 shows the primer sequences of the DchWOXs and reference gene.
3. Discussion
A class of TFs unique to plants, the WOX family is involved in critical regulatory functions in important developmental programs like organ morphogenesis and stem cell maintenance [3,34]. In our study, 30 WOXs were identified from the genomes of three species: D. chrysotoxum contained nine DchWOXs, D. huoshanense contained 11 DhuWOXs, and D. nobile contained 10 DnoWOXs. The three Dendrobium species had similar numbers of WOXs to D. catenatum (14) [35], Phalaenopsis equestris (10) [36], A. thaliana (15), O. sativa (13), Sorghum bicolor (11) [37], Vitis vinifera (12) [38], tobacco (9) [39], and Triticum aestivum (14) [40], but differed from those of dicotyledonous plants such as Brassica napus (58) [41], soybean (33) [42], and Gossypium hirsutum (40) [43]. Variations in the size of a species’ genome or processes like gene and genome replication may be the cause of the variations in the number of WOXs between species [44].
All three Dendrobium species in this study had undergone at least two whole-genome duplication (WGD) events [30,31,32]. According to the chromosome distribution map (Figure 1), both D. chrysotoxum and D. huoshanense harbored a single pair of tandem repeat genes. The synteny analysis showed that both D. chrysotoxum and D. huoshanense had a single pair of genes with segmental duplication, and D. nobile had two pairs of genes with segmental duplication (Figure 5). It is probably because of these duplication events that the WOX genes differed in number and distribution among the three species. In addition, the Ka/Ks ratios of the four WOX gene pairs detected in this study were all less than one, revealing that these WOXs underwent strong purifying selection during evolution (Table 2) [45]. This enables them to remain highly conserved in evolving Dendrobium species, maintaining the specific biological functions of WOX proteins [46].
The phylogenetic analysis of the 30 WOXs from the Dendrobium species compared with those from Arabidopsis and O. sativa showed that the distribution of the WOXs in Dendrobium species is conserved (Figure 2). Like most plants, such as O. sativa, Picea abies, and Eriobotrya japonica, the WUS/modern clade had the highest number of WOX genes and the ancient clade had the lowest number of WOX genes among the three Dendrobium species [36,47,48]. The loss of certain WOX genes occurred in the three Dendrobium species, except for WOX1/6/7/8/14, which is unique to dicotyledons. For example, D. chrysotoxum and D. huoshanense both lost WOX4, and only D. nobile retained the homologous gene for AtWOX4 (DnoWOX6). Strikingly, DchWOX1 and DnoWOX1 were well clustered into a subclade with AtWUS and OsWOX1 (AtWUS homologous gene). AtWUS was shown to be the prototype of the Arabidopsis WOX gene family, so we speculate that DchWOX1 is the prototype of the WOXs of D. chrysotoxum, and DnoWOX1 is the prototype of the WOXs of D. nobile [3]. However, similar to D. catenatum, the prototype gene was absent in D. huoshanense [35]. We hypothesize that, during evolution, there may have been functional redundancy among WOX family members to compensate for the functions performed by the missing genes, or some species-specific WOXs may have arisen [3,39,49].
Supported by the conserved motifs and intron patterns, the highly conserved gene structure guarantees the conserved function of each clade or subclade. WOX genes in the same subfamily tend to have similar numbers of introns and exons, and they also share similarities in gene structure (Figure 3) [50,51]. The gene of the ancient clade has a more conserved gene structure than the other two clades’ genes, consistent with the WOX genes of Arabidopsis, Poplar, and Sorghum [37]. The conserved ancient clade is present in all plants, and we hypothesize that strict conservation ensures that these WOX proteins perform indispensable functions in plant evolution [6]. The concatenated motif 3 and motif 1 (Figure 3A) correspond to the homeodomain sequence shown in Figure 4 and are present in all 30 WOX proteins. The homeodomain exhibits a helix–turn–helix–loop–helix structure, which ensures that it can differentiate between sequence-specific targets with precise spatiotemporal organization (Figure 4A) [52]. Similar to most plants, such as Arabidopsis, rice, and maize, only WUS/modern clade members contained the WUS-box (TL-LFP-) (Figure 4B) [4,39]. In summary, the sequence and structure conservation of the WOX gene family members maintains their functional integrity across species.
Transcriptional regulation occurs mainly through the promoter and its associated CREs to activate or repress gene expression [53]. The WOX gene family is extensively involved in regulating the development of various plant organs and contributes to abiotic stress and phytohormone signaling. The promoter regions of these 30 WOXs were rich in light-responsive elements, suggesting that WOXs play an essential role in regulating the light response (Figure 6) [54]. The meristem expression is the most frequent of the growth and developmental components. DchWOX4, DhuWOX2, and DnoWOX3 had two, three, and three meristem expression elements, respectively (Figure 6B), and these three genes shared a branch with AtWOX11 and AtWOX12. Since AtWOX11 and AtWOX12 participate in new root organ development in Arabidopsis [12], DchWOX4, DhuWOX2, and DnoWOX3 are speculated to regulate the differentiation of roots in the three Dendrobium species, respectively. The stress-repressive CREs in the promoter region of the WOXs mainly include anaerobic induction, drought inducibility, and low-temperature responsiveness, which implies that the WOX genes are essential for plants to respond to abiotic stresses. This has been verified in Arabidopsis and O. sativa; for example, the rab21 promoter drives OsWOX13 overexpression in O. sativa, thereby improving its drought tolerance [55]. Elements associated with the plant hormone response in the promoter region of the WOXs are ABA, IAA, SA, GA, and MeJA. Many studies have revealed that the WOX is affected by IAA, ABA, and GA during plant growth and development [22,56]. The MeJA response element is the most abundant phytohormone response element. It is involved in plant defense responses and also regulates plant growth and development. [57,58]. To summarize, the WOX gene family in Dendrobium species has a vital function in plant growth, development, and the stress response by mediating phytohormone regulation.
The transcriptome analysis of Arabidopsis, O. sativa, Fragaria vesca, and Nelumbo nucifera indicated that NnWOX14 was significantly expressed in the carpel of N. nucifera; FvWOX9 and FvWOX9a in F. vesca showed significant expression in the process of development; and WOX family members are expressed in the flowers of both Arabidopsis and O. sativa [37,59,60]. All of these observations suggest the significance of the WOX gene in the formation of floral organs in plants. Therefore, we combined transcriptomic data from the flowers of D. chrysotoxum with RT-qPCR experiments to identify the important regulatory role of the WOX genes in D. chrysotoxum flower development (Figure 7 and Figure 8). According to the results of DchWOX3 being significantly expressed in the S1 lip (Figure 8A), along with the development process of gradually reducing the amount of its expression, we speculate that DchWOX3 may participate in regulating lip growth. It has been found that PeWOX9A, PeWOX9B, and DcWOX9 are highly expressed in the gynoecium in D. catenatum and P. equestris, respectively, and the overexpression of DcWOX9 in Arabidopsis resulted in staminate and pistil sterility [36]. In our study, DchWOX5 was significantly expressed in the ovary during the three periods (Figure 8B). PeWOX9A, PeWOX9B, DcWOX9, and DchWOX5 both share a clade with AtWOX9, indicating their specific roles in regulating gynoecium and ovary development, which need to be further verified.
4. Materials and Methods
4.1. Data Sources
The genome files of D. chrysotoxum (accession number: PRJNA664445) and D. nobile (accession number: PRJNA725550) were retrieved from the NCBI (https://www.ncbi.nlm.nih.gov/, accessed on 25 September 2023). The genome file of D. huoshanense (accession number: CNA0014590) was retrieved from the China Nucleotide Sequence Archive (CNSA, https://ftp.cngb.org/, accessed on 25 September 2023). A total of 15 A. thaliana WOX protein sequences were retrieved from TAIR (http://www.arabidopsis.org, accessed on 25 September 2023). The 13 WOX protein sequences of O. sativa were retrieved from PlantTFDB (http://planttfdb.gao-lab.org/, accessed on 25 September 2023).
4.2. Identification and Physicochemical Properties of WOXs
Candidate WOX genes were searched in the genomes of three Dendrobium species in the two-way BLAST tool of the TBtools v2.003 software, using the 15 WOXs of A. thaliana as probes, respectively [61,62]. Meanwhile, using the Simple HMM Search tool of TBtools, the Hidden Markov Model (HMM) file of the homeodomain (PF00046) from the Pfam database (http://pfam.xfam.org/search, accessed on 27 September 2023) was utilized to further identify WOX family members in the three Dendrobium species. Candidate WOXs identified by BLAST and HMM were uploaded to NCBI CD-Search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi, accessed on 27 September 2023) for structural analysis, and only genes with conserved typical homeodomains of WOXs were retained.
All finalized sequences of the 30 WOX proteins were uploaded to the online software ExPASy (https://www.expasy.org/, accessed on 27 September 2023) for physicochemical property analysis, to obtain the amino acids (aa), molecular weight (MW), isoelectric point (pI), instability index (II), aliphatic index (AI), and grand average of hydropathicity (GRAVY) of all WOX proteins [63]. Then, the subcellular localization prediction of the WOX family members was performed by the online program Cell-PLoc 2.0 (http://www.csbio.sjtu.edu.cn/bioinf/Cell-PLoc-2/, accessed on 27 September 2023).
4.3. Chromosomal Localization
Based on the annotation data of the three Dendrobium genomes, chromosomal localization maps of the WOX genes were produced using Gene Location Visualize from the GTF/GFF program of TBtools.
4.4. Phylogenetic Analysis of WOX Gene Family
The protein sequences of 15 AtWOXs, 13 OsWOXs, ten DchWOXs, nine DhuWOXs, and 11 DnoWOXs were uploaded into the MEGA11 software, and then these 58 WOX protein sequences were used to achieve sequence alignment using the Clustal W function (default parameters), and the phylogenetic trees of the five species were constructed using the maximum likelihood (1000 bootstrap replication) [64,65]. The editing and beautification of the phylogenetic tree was performed by Evolview 3.0. (http://www.evolgenius.info/evolview/#/treeview, accessed on 8 October 2023) [66].
4.5. Protein Conservative Domain and Gene Structure Analysis
The prediction of conserved structural domains for the 30 WOXs in Dendrobium was accomplished by utilizing the CDD program from NCBI (https://www.ncbi.nlm.nih.gov/cdd, accessed on 10 October 2023). The identification of conserved motifs for the 30 WOXs was performed by the MEME online program (https://meme-suite.org/meme/tools/meme, accessed on 10 October 2023) [67]. Gene Structure View in TBtools was employed to map the phylogenetic trees, conserved motifs, and gene structures in combination. The WOX protein sequence alignment was performed by Clustal W of MEGA 11 and then beautified by jalview (Version: 2.11.3.2).
4.6. Synteny Analysis of WOX Gene Family
The identification of intra-species duplicate genes in the three Dendrobium species was performed using the One Step MCScanx function of TBtools [68]. In Advance Circos of TBtools, the duplication patterns of the three Dendrobium species were visualized. Then, the calculation of the Ka, Ks, and Ka/Ks values for the gene pairs was accomplished by the Simple Ka/Ks Calculator in TBtools.
4.7. Cis-Acting Regulatory Element Analysis
First, Gtf/Gff3 Sequence Extract and Fasta Extract of TBtools were used to extract 2000 bp upstream of the 30 WOX genes. Second, to complete the prediction of the CREs, the acquired sequences were submitted to the online website PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/, accessed on 13 October 2023). Finally, the distribution of the acquired CREs was visualized using the Basic Biosequence View module of TBtools, while the categories and number of CREs were counted and plotted in Excel 2016 [69].
4.8. Expression Pattern and RT-qPCR Analysis
D. chrysotoxum plant materials were taken from the Forest Orchid Garden of Fujian Agriculture and Forestry University for transcriptome sequencing and RT-qPCR, including five flower parts (sepal, petal, lip, ovary, and gynostemium) in three periods (unpigmented bud stage, pigmented bud stage, and early flowering stage).
The transcriptome sequencing and library construction of the five flower parts from the three periods of D. chrysotoxum development were performed by BGI Genomics Co., Ltd. (Shenzhen, China). RESM v1.2.8 was used for transcript quantification and to calculate the FPKM value for each sample. Based on the FPKM value, heatmaps of gene expression are created in the HeatMap program of TBtools.
Further validation of the expression patterns of the three WOX genes was achieved by RT-qPCR experiments. The FastPure Plant Total RNA Isolation Kit (for polysaccharide- and polyphenol-rich tissues) (Vazyme Biotech Co., Ltd., Nanjing, China) was used to extract total RNA from D. chrysotoxum samples. The Hifair® AdvanceFast One-Step RT-gDNA Digestion SuperMix for qPCR (YEASEN, Shanghai, China) was used to generate the cDNA for the quantitative PCR. Based on the transcription data, DchActin (Maker75111) was selected as the reference gene. The WOX gene sequences were submitted to the Primer Premier 5 software to design specific PCR primers (Table S5). The TSINGKE ArtiCanATM SYBR qPCR Mix was used for the RT-qPCR analysis on the Bio-Rad/CFX Connect Real-Time PCR Detection System (Bio-Rad Laboratories, Hercules, CA, USA). Three biological replicates were carried out for all experiments. Finally, the relative expression of the three WOX genes was calculated using the 2−∆∆CT method with S1 Se as the reference. The data were visualized using GraphPad Prism 7.0.
5. Conclusions
In this study, 10, 11, and 9 WOX genes were identified in the genomes of D. chrysotoxum, D. huoshanense, and D. nobile, respectively, and chromosomal localization, phylogeny, gene structure, and motif composition analyses were performed. In addition, based on the transcriptome and RT-qPCR experiments, we analyzed the expression patterns of the DchWOXs in five floral parts of D. chrysotoxum at three developmental periods. In conclusion, our results provide useful information for the in-depth exploration of the biological roles of the WOX gene family, as well as floral developmental studies in Dendrobium species.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/ijms25105352/s1.
Author Contributions
X.L. wrote the manuscript. X.H. and M.Z. performed the experiments. Q.Z. and Y.H. participated in the plant sample collection and data analysis. X.Z. revised the manuscript. B.C. and Z.L. created and planned the entire project and coordinated all collaborators. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Forestry Peak Discipline Construction Project of Fujian Agriculture and Forestry University (72202200205).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The genome sequences and annotation files of D. chrysotoxum (accession number: PRJNA664445) and D. nobile (accession number: PRJNA725550) were downloaded from the National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/, accessed on 25 September 2023) genome database. The genome file of D. huoshanense (accession number: CNA0014590) was downloaded from the China Nucleotide Sequence Archive (CNSA, https://ftp.cngb.org/, accessed on 25 September 2023).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Gehring, W.J.; Affolter, M.; Bürglin, T. Homeodomain proteins. Annu. Rev. Biochem. 1994, 63, 487. [Google Scholar] [CrossRef] [PubMed]
- Breuninger, H.; Rikirsch, E.; Hermann, M.; Ueda, M.; Laux, T. Differential Expression of WOX Genes Mediates Apical-Basal Axis Formation in the Arabidopsis Embryo. Dev. Cell 2008, 14, 867–876. [Google Scholar] [CrossRef] [PubMed]
- Enrico, C.; Christophe, T.; Michiel, V. The role of WOX genes in flower development. Ann. Bot. 2014, 114, 1545–1553. [Google Scholar]
- Haecker, A.; Groß-Hardt, R.; Geiges, B.; Sarkar, A.; Breuninger, H.; Herrmann, M.; Laux, T. Expression dynamics of WOX genes mark cell fate decisions during early embryonic patterning in Arabidopsis thaliana. Development 2004, 131, 657–668. [Google Scholar] [CrossRef] [PubMed]
- Van der Graaff, E.; Laux, T.; Rensing, S.A. The WUS homeobox-containing (WOX) protein family. Genome Biol. 2009, 10, 248. [Google Scholar] [CrossRef] [PubMed]
- Lian, G.; Ding, Z.; Wang, Q.; Zhang, D.; Xu, J. Origins and Evolution of WUSCHEL-Related Homeobox Protein Family in Plant Kingdom. Sci. World J. 2014, 2014, 534140. [Google Scholar] [CrossRef] [PubMed]
- Romera-Branchat, M.; Ripoll, J.J.; Yanofsky, M.F.; Pelaz, S. The WOX13 homeobox gene promotes replum formation in the Arabidopsis thaliana fruit. Plant J. 2012, 73, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Deveaux, Y.; Toffano-Nioche, C.; Claisse, G.; Thareau, V.; Morin, H.; Laufs, P.; Moreau, H.; Kreis, M.; Lecharny, A. Genes of the most conserved WOX clade in plants affect root and flower development in Arabidopsis. BMC Evol. Biol. 2008, 8, 291. [Google Scholar] [CrossRef] [PubMed]
- Weigel, D.; Lippman, Z.B.; Cohen, O.; Alvarez, J.P.; Abu-Abied, M.; Pekker, I.; Paran, I.; Eshed, Y.; Zamir, D. The Making of a Compound Inflorescence in Tomato and Related Nightshades. PLoS Biol. 2008, 6, e288. [Google Scholar]
- Wang, W.; Li, G.; Zhao, J.; Chu, H.; Lin, W.; Zhang, D.; Wang, Z.; Liang, W. DWARF TILLER1, a WUSCHEL-Related Homeobox Transcription Factor, Is Required for Tiller Growth in Rice. PLoS Genet. 2014, 10, e1004154. [Google Scholar] [CrossRef]
- Hendelman, A.; Zebell, S.; Rodriguez-Leal, D.; Dukler, N.; Lippman, Z.B. Conserved pleiotropy of an ancient plant homeobox gene uncovered by cis-regulatory dissection. Cell 2021, 184 (Suppl. S1), 1724–1739. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sheng, L.; Xu, Y.; Li, J.; Yang, Z.; Huang, H.; Xu, L. WOX11 and 12 Are Involved in the First-Step Cell Fate Transition during de Novo Root Organogenesis in Arabidopsis. Plant Cell 2014, 26, 1081–1093. [Google Scholar] [CrossRef] [PubMed]
- Mayer, K.F.X.; Schoof, H.; Haecker, A.; Lenhard, M.; Jürgens, G.; Laux, T. Role of WUSCHEL in Regulating Stem Cell Fate in the Arabidopsis Shoot Meristem. Cell 1998, 95, 805–815. [Google Scholar] [CrossRef] [PubMed]
- Jha, P.; Ochatt, S.J.; Kumar, V. WUSCHEL: A master regulator in plant growth signaling. Plant Cell Rep. 2020, 39, 431–444. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, M.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis WUSCHEL Is a Bifunctional Transcription Factor That Acts as a Repressor in Stem Cell Regulation and as an Activator in Floral Patterning. Plant Cell 2009, 21, 3493–3505. [Google Scholar] [CrossRef] [PubMed]
- Nakata, M.; Matsumoto, N.; Tsugeki, R.; Rikirsch, E.; Okada, K. Roles of the middle domain-specific WUSCHEL-RELATED HOMEOBOX genes in early development of leaves in Arabidopsis. Plant Cell 2012, 24, 519–535. [Google Scholar] [CrossRef]
- Honda, E.; Yew, C.L.; Yoshikawa, T.; Sato, Y.; Hibara, K.I.; Itoh, J.I. LEAF LATERAL SYMMETRY1, a Member of the WUSCHEL-RELATED HOMEOBOX3 Gene Family, Regulates Lateral Organ Development Differentially from Other Paralogs, NARROW LEAF2 and NARROW LEAF3 in Rice. Plant Cell Physiol. 2018, 59, 376–391. [Google Scholar] [CrossRef] [PubMed]
- Batool, H.S.; François, T.J.; Juliane, R.; Kurt, Z.; Andrea, R. Constitutive Overexpression of a Conifer WOX2 Homolog Affects Somatic Embryo Development in Pinus pinaster and Promotes Somatic Embryogenesis and Organogenesis in Arabidopsis Seedlings. Front. Plant Sci. 2022, 13, 838421. [Google Scholar]
- Sarkar, A.K.; Luijten, M.; Miyashima, S.; Lenhard, M.; Hashimoto, T.; Nakajima, K.; Scheres, B.; Heidstra, R.; Laux, T. Conserved factors regulate signalling in Arabidopsis thaliana shoot and root stem cell organizers. Nature 2007, 446, 811–814. [Google Scholar] [CrossRef]
- Lopez-Moya, F.; Escudero, N.; Zavala-Gonzalez, E.A.; Esteve-Bruna, D.; Blázquez, M.A.; Alabadí, D.; Lopez-Llorca, L.V. Induction of auxin biosynthesis and WOX5 repression mediate changes in root development in Arabidopsis exposed to chitosan. Sci. Rep. 2017, 7, 16813. [Google Scholar] [CrossRef]
- Wang, L.Q.; Wen, S.S.; Wang, R.; Wang, C.; Gao, B.; Lu, M.Z. PagWOX11/12a activates PagCYP736A12 gene that facilitates salt tolerance in poplar. Plant Biotechnol. J. 2021, 19, 2249–2260. [Google Scholar] [CrossRef]
- Sajjad, M.; Wei, X.; Liu, L.; Li, F.; Ge, X. Transcriptome Analysis Revealed GhWOX4 Intercedes Myriad Regulatory Pathways to Modulate Drought Tolerance and Vascular Growth in Cotton. Int. J. Mol. Sci. 2021, 22, 898. [Google Scholar] [CrossRef] [PubMed]
- Kong, D.; Pan, X.; Jing, Y.; Zhao, Y.; Duan, Y.; Yang, J.; Wang, B.; Liu, Y.; Shen, R.; Cao, Y.; et al. ZmSPL10/14/26 are required for epidermal hair cell fate specification on maize leaf. New Phytol. 2021, 230, 1533–1549. [Google Scholar] [CrossRef] [PubMed]
- Schmitz, R.J.; Grotewold, E.; Stam, M. Cis-regulatory sequences in plants: Their importance, discovery, and future challenges. Plant Cell 2022, 34, 718–741. [Google Scholar] [CrossRef]
- Chase, M.W.; Cameron, K.M.; Freudenstein, J.V.; Pridgeon, A.M.; Salazar, G.; van den Berg, C.; Schuiteman, A. An updated classification of Orchidaceae. Bot. J. Linn. Soc. 2015, 177, 151–174. [Google Scholar] [CrossRef]
- Christenhusz, M.J.M.; Byng, J.W. The number of known plants species in the world and its annual increase. Phytotaxa 2016, 261, 201–217. [Google Scholar] [CrossRef]
- Liu, H.; Luo, Y.-B. Protecting Orchids in Nature Reserves: Research and Restoration Needs. Bot. Rev. 2010, 76, 137–139. [Google Scholar] [CrossRef]
- Pridgeon, A.M.; Cribb, P.; Chase, M.W.; Rasmussen, F.N. Genera Orchidacearum Volume 6: Epidendroideae (Part 3); OUP: Oxford, UK, 2014. [Google Scholar]
- Niu, Z.; Zhu, F.; Fan, Y.; Li, C.; Zhang, B.; Zhu, S.; Hou, Z.; Wang, M.; Yang, J.; Xue, Q.; et al. The chromosome-level reference genome assembly for Dendrobium officinale and its utility of functional genomics research and molecular breeding study. Acta Pharm. Sin. B 2021, 11, 2080–2092. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, G.-Q.; Zhang, D.; Liu, X.-D.; Xu, X.-Y.; Sun, W.-H.; Yu, X.; Zhu, X.; Wang, Z.-W.; Zhao, X.; et al. Chromosome-scale assembly of the Dendrobium chrysotoxum genome enhances the understanding of orchid evolution. Hortic. Res. 2021, 8, 183. [Google Scholar] [CrossRef]
- Bangxing, H.; Yi, J.; Jun, D.; Tao, Z.; Fangli, G.; Qun, Z.; Fucheng, Z.; Xiangwen, S.; Hui, D.; Peipei, W.; et al. A chromosome-level genome assembly of Dendrobium huoshanense using long reads and Hi-C data. Genome Biol. Evol. 2020, 12, 2486–2490. [Google Scholar]
- Xu, Q.; Niu, S.-C.; Li, K.-L.; Zheng, P.-J.; Zhang, X.-J.; Jia, Y.; Liu, Y.; Niu, Y.-X.; Yu, L.-H.; Chen, D.-F.; et al. Chromosome-Scale Assembly of the Dendrobium nobile Genome Provides Insights Into the Molecular Mechanism of the Biosynthesis of the Medicinal Active Ingredient of Dendrobium. Front. Genet. 2022, 13, 844622. [Google Scholar] [CrossRef]
- Gasteiger, E. Protein identification and analysis tools on the ExPASy server. In The Proteomics Protocols Handbook; Humana Press: Totowa, NJ, USA, 2005. [Google Scholar]
- Lin, H.; Niu, L.; McHale, N.A.; Ohme-Takagi, M.; Mysore, K.S.; Tadege, M. Evolutionarily conserved repressive activity of WOX proteins mediates leaf blade outgrowth and floral organ development in plants. Proc. Natl. Acad. Sci. USA 2013, 110, 366–371. [Google Scholar] [CrossRef]
- Li, H.; Li, C.; Wang, Y.; Qin, X.; Meng, L.; Sun, X. Genome-Wide Analysis of the WOX Transcription Factor Genes in Dendrobium catenatum Lindl. Genes 2022, 13, 1481. [Google Scholar] [CrossRef] [PubMed]
- Ramkumar, T.R.; Kanchan, M.; Upadhyay, S.K.; Sembi, J.K. Identification and characterization of WUSCHEL-related homeobox (WOX) gene family in economically important orchid species Phalaenopsis equestris and Dendrobium catenatum. Plant Gene 2018, 14, 37–45. [Google Scholar] [CrossRef]
- Zhang, X.; Zong, J.; Liu, J.; Yin, J.; Zhang, D. Genome-Wide Analysis of WOX Gene Family in Rice, Sorghum, Maize, Arabidopsis and Poplar. J. Integr. Plant Biol. 2010, 52, 1016–1026. [Google Scholar] [CrossRef] [PubMed]
- Gambino, G.; Minuto, M.; Boccacci, P.; Perrone, I.; Vallania, R.; Gribaudo, I. Characterization of expression dynamics of WOX homeodomain transcription factors during somatic embryogenesis in Vitis vinifera. J. Exp. Bot. 2011, 62, 1089. [Google Scholar] [CrossRef] [PubMed]
- Xuemei, Z.; Yingying, G.; Peng, Z.; Meng-Xiang, S. Comparative Analysis of WUSCHEL-Related Homeobox Genes Revealed Their Parent-of-Origin and Cell Type-Specific Expression Pattern During Early Embryogenesis in Tobacco. Front. Plant Sci. 2018, 9, 311. [Google Scholar]
- Shi, L.; Wang, K.; Du, L.; Song, Y.; Li, H.; Ye, X. Genome-Wide Identification and Expression Profiling Analysis of WOX Family Protein-Encoded Genes in Triticeae Species. Multidiscip. Digit. Publ. Inst. 2021, 22, 9325. [Google Scholar] [CrossRef]
- Wang, M.-M.; Liu, M.-M.; Ran, F.; Guo, P.-C.; Ke, Y.-Z.; Wu, Y.-W.; Wen, J.; Li, P.-F.; Li, J.-N.; Du, H. Global Analysis of WOX Transcription Factor Gene Family in Brassica napus Reveals Their Stress- and Hormone-Responsive Patterns. Int. J. Mol. Sci. 2018, 19, 3470. [Google Scholar] [CrossRef]
- Hao, Q.; Zhang, L.; Yang, Y.; Shan, Z.; Zhou, X.A. Genome-Wide Analysis of the WOX Gene Family and Function Exploration of GmWOX18 in Soybean. Plants 2019, 8, 215. [Google Scholar] [CrossRef]
- Ruibin, S.; Xue, Z.; Dan, M.; Chuanliang, L. Identification and Evolutionary Analysis of Cotton (Gossypium hirsutum) WOX Family Genes and Their Potential Function in Somatic Embryogenesis. Int. J. Mol. Sci. 2023, 24, 11077. [Google Scholar] [CrossRef]
- Flagel, L.E.; Wendel, J.F. Gene duplication and evolutionary novelty in plants. New Phytol. 2009, 183, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Gao, X.; Wu, Y.; Wei, C.; Li, M.H.; Liu, D.K.; Liu, Z.J. Identification and Analysis of PEPC Gene Family Reveals Functional Diversification in Orchidaceae and the Regulation of Bacterial-TypePEPC. Int. J. Mol. Sci. 2024, 25, 2055. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Guan, C.; Chen, Y.; Shi, Y.; Long, O.; Lin, H.; Zhang, K.; Zhou, M. Evolutionary analysis of MADS-box genes in buckwheat species and functional study of FdMADS28 in flavonoid metabolism. Plant Physiol. Biochem. 2024, 210, 108637. [Google Scholar] [CrossRef]
- Hedman, H.; Zhu, T.; von Arnold, S.; Sohlberg, J.J. Analysis of the WUSCHEL-RELATED HOMEOBOX gene family in the conifer picea abiesreveals extensive conservation as well as dynamic patterns. BMC Plant Biol. 2013, 13, 89. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Yang, M.; Liu, X.; Xia, Y.; Hu, R.; Xia, Q.; Jing, D.; Guo, Q. Genome-wide analysis of the WOX gene family and the role of EjWUSa in regulating flowering in loquat (Eriobotrya japonica). Front. Plant Sci. 2022, 13, 1024515. [Google Scholar] [CrossRef]
- Feng, C.; Zou, S.; Gao, P.; Wang, Z. In silico identification, characterization expression profile of WUSCHEL-Related Homeobox (WOX) gene family in two species of kiwifruit. Peer J. 2021, 9, e12348. [Google Scholar] [CrossRef]
- Muhammad Tajo, S.; Pan, Z.; He, S.; Chen, B.; Km, Y.; Mahmood, T.; Bello Sadau, S.; Shahid Iqbal, M.; Gereziher, T.; Suleiman Abubakar, U.; et al. Characterization of WOX genes revealed drought tolerance, callus induction, and tissue regeneration in Gossypium hirsutum. Front. Genet. 2022, 13, 928055. [Google Scholar] [CrossRef]
- Zhang, Z.A.; Liu, M.Y.; Ren, S.N.; Liu, X.; Gao, Y.H.; Zhu, C.Y.; Niu, H.Q.; Chen, B.W.; Liu, C.; Yin, W.; et al. Identification of WUSCHEL-related homeobox gene and truncated small peptides in transformation efficiency improvement in Eucalyptus. BMC Plant Biol. 2023, 23, 604. [Google Scholar] [CrossRef]
- Li, M.; Wang, R.; Liu, Z.; Wu, X.; Wang, J. Genome-wide identification and analysis of the WUSCHEL-related homeobox (WOX) gene family in allotetraploid Brassica napus reveals changes in WOX genes during polyploidization. BMC Genom. 2019, 20, 317. [Google Scholar] [CrossRef]
- Hernandez-Garcia, C.M.; Finer, J.J. Identification and validation of promoters and cis-acting regulatory elements. Plant Sci. 2014, 217–218, 109–119. [Google Scholar] [CrossRef]
- Guo, L.; Lu, S.; Liu, T.; Nai, G.; Ren, J.; Gou, H.; Chen, B.; Mao, J. Genome-Wide Identification and Abiotic Stress Response Analysis of PP2C Gene Family in Woodland and Pineapple Strawberries. Int. J. Mol. Sci. 2023, 24, 4049. [Google Scholar] [CrossRef] [PubMed]
- Pham-Thi, M.-T.; Sug, K.J.; Songhwa, C.; Mi, J.K.; Gang-Seob, L.; Dong-Eun, K.; Jong-Joo, C.; Ik, S.S.; Hie, N.B.; Yeon-Ki, K. A WUSCHEL Homeobox Transcription Factor, OsWOX13, Enhances Drought Tolerance and Triggers Early Flowering in Rice. Mol. Cells 2018, 41, 781–798. [Google Scholar]
- Tao, Z.; Ruonan, L.; Jialing, X.; Lang, Y.; Rongchen, W.; Yunde, Z. The YUCCA-Auxin-WOX11 Module Controls Crown Root Development in Rice. Front. Plant Sci. 2018, 9, 523. [Google Scholar]
- Ji, J.; Shimizu, R.; Sinha, N.; Scanlon, M.J. Analyses of WOX4 transgenics provide further evidence for the evolution of theWOXgene family during the regulation of diverse stem cell functions. Plant Signal. Behav. 2014, 5, 916–920. [Google Scholar] [CrossRef] [PubMed]
- Fiume, E.; Fletcher, J.C. Regulation of Arabidopsis Embryo and Endosperm Development by the Polypeptide Signaling Molecule CLE8. Plant Cell 2012, 24, 1000–1012. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.Z.; Huang, J.; Lin, Z.C.; Wang, F.; Yang, S.M.; Jiang, X.; Ahmad, S.; Zhou, Y.Z.; Lan, S.; Liu, Z.J.; et al. Genome-Wide Analysis of WUSCHEL-Related Homeobox Gene Family in Sacred Lotus (Nelumbo nucifera). Int. J. Mol. Sci. 2023, 24, 14216. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Zhao, X.; Miao, Y.; Wang, D.; Zhang, Z.; Liu, Y. Genome-Wide Identification and Expression Profile Analysis of the WUSCHEL-Related Homeobox (WOX) Genes in Woodland Strawberry (Fragaria vesca). Horticulturae 2022, 8, 1043. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Xia, R. TBtools-II: A One for All, All for One Bioinformatics Platform for Biological Big-data Mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Duvaud, S.; Gabella, C.; Lisacek, F.; Stockinger, H.; Ioannidis, V.; Durinx, C. Expasy, the Swiss Bioinformatics Resource Portal, as designed by its users. Nucleic Acids Res. 2021, 49, W216–W227. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2015, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Riccucci, E.; Vanni, C.; Vangelisti, A.; Fambrini, M.; Giordani, T.; Cavallini, A.; Mascagni, F.; Pugliesi, C. Genome-Wide Analysis of WOX Multigene Family in Sunflower (Helianthus annuus L.). Int. J. Mol. Sci. 2023, 24, 3352. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, B.; Gao, S.; Lercher, M.J.; Hu, S.; Chen, W.-H. Evolview v3: A webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 2019, 47, W270–W275. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.h.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Lescot, M. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).