Peripheral Blood CD8+ T-Lymphocyte Immune Response in Benign and Subpopulations of Breast Cancer Patients
Abstract
1. Introduction
2. Results
2.1. Proteomic Analysis Using Orbitrap Exploris™ 480 MS Coupled to an UHPLC
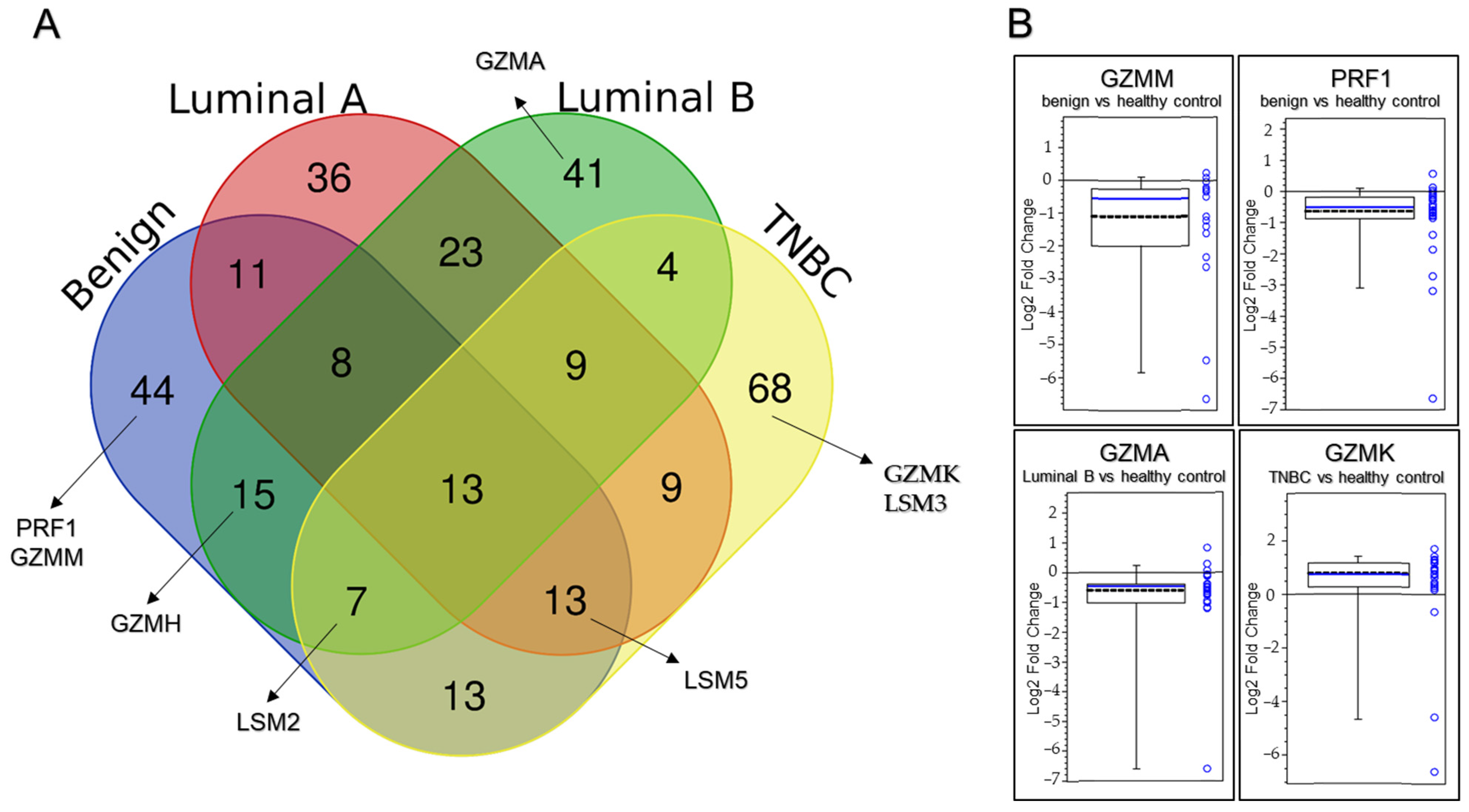
2.2. Label-Free Quantitative Proteomic Analysis of CD8+ T Lymphocytes in the Peripheral Blood Samples
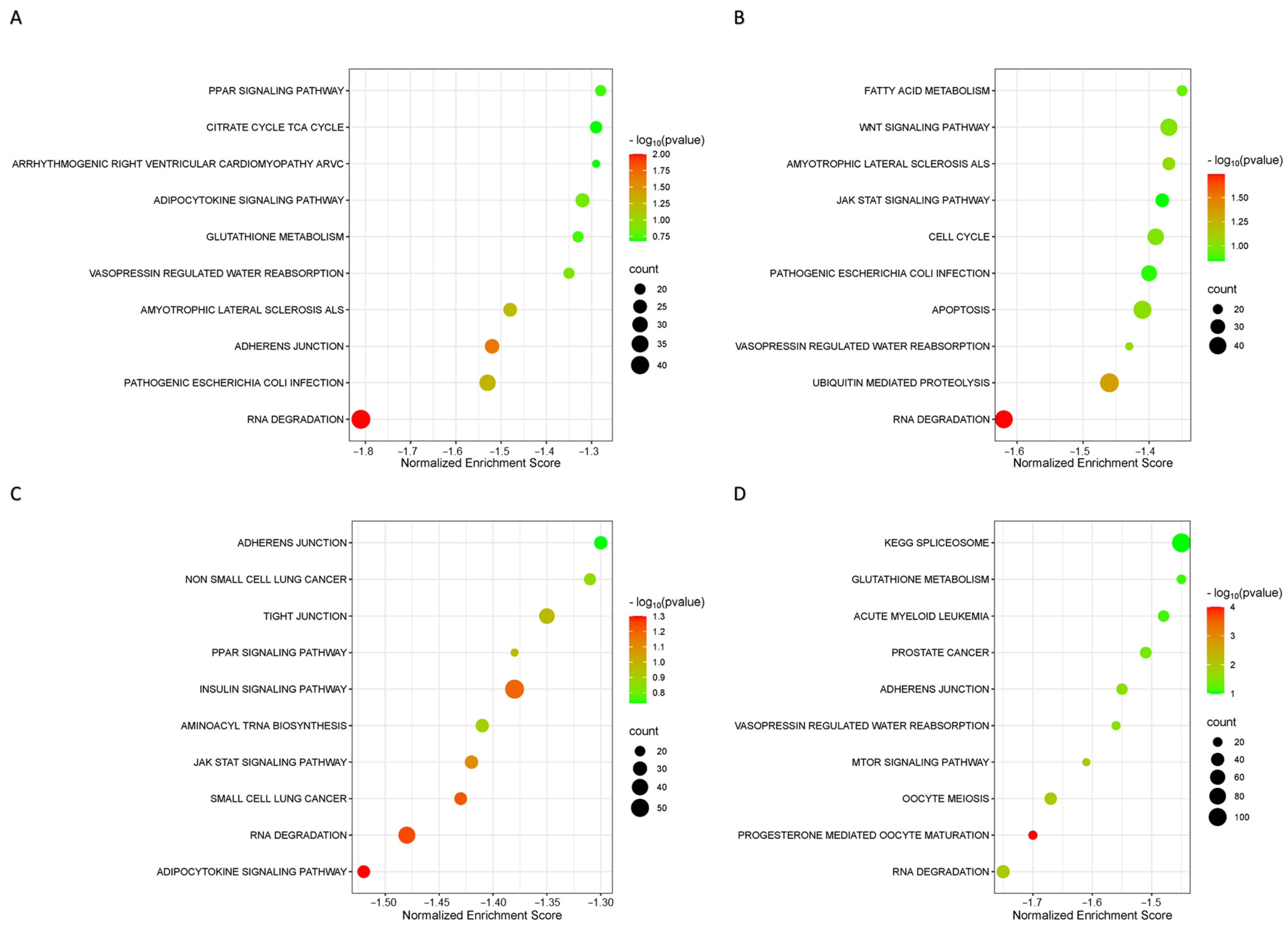
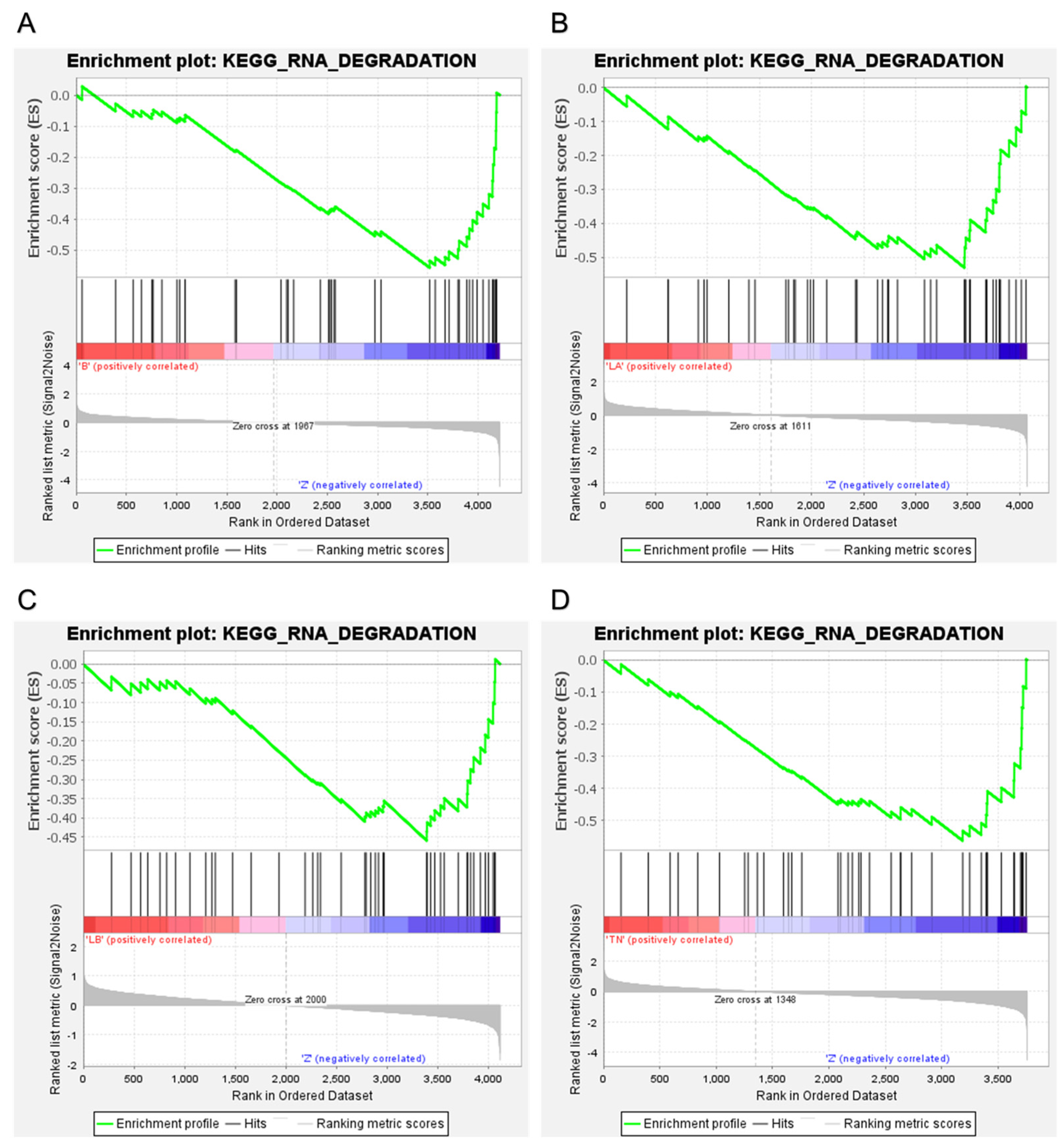
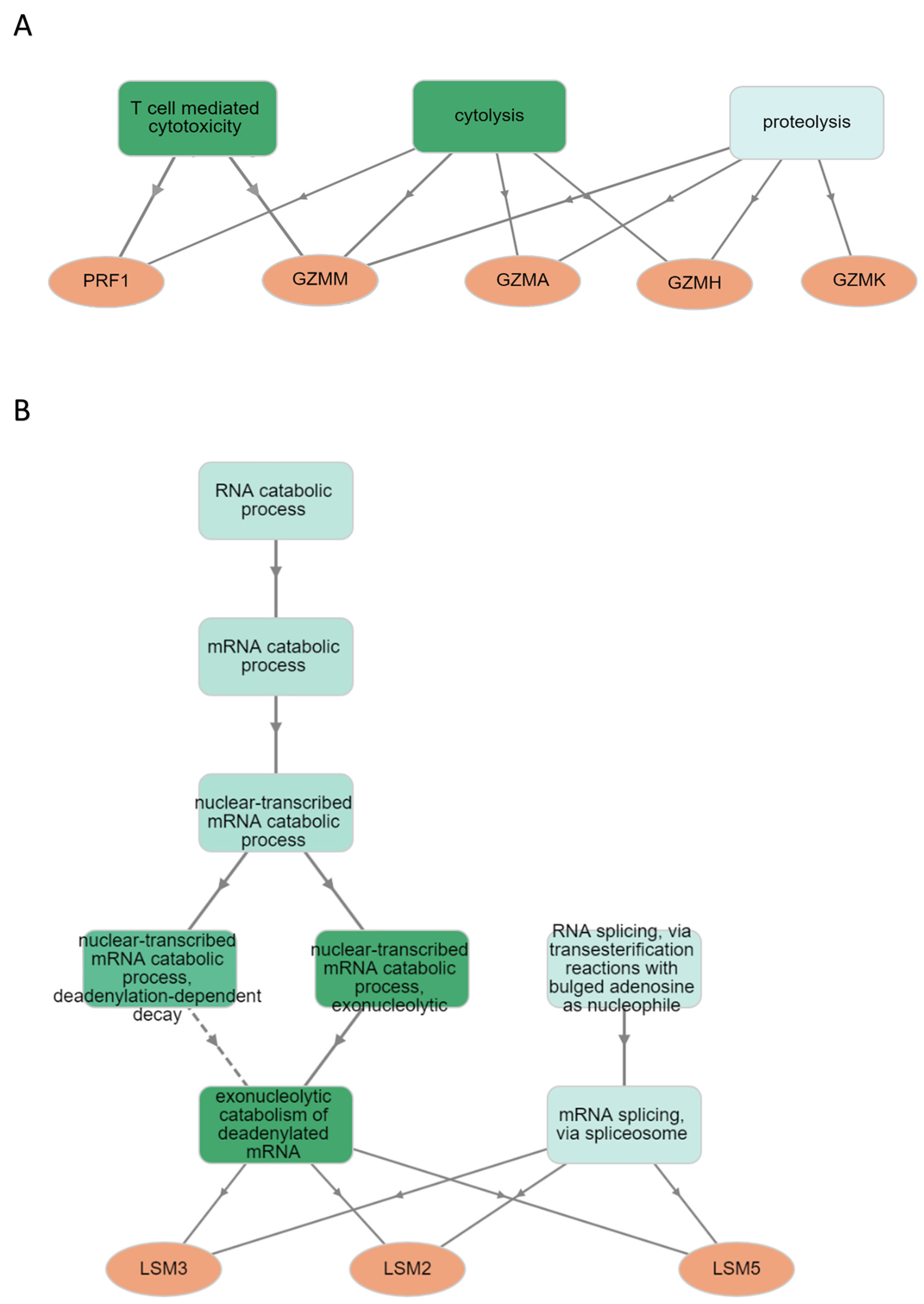
2.3. Orbitrap-MS Analysis and Pathway Enrichment Analysis of the Differential Blood Proteins in CD8+ T Lymphocytes
3. Discussion
4. Materials and Methods
4.1. Patient’s Selection
4.2. Immunohistochemistry
4.3. Blood Sampling
4.4. Isolation of CD8+ T-Lymphocytes
4.5. In Solution Digestion
4.6. Proteomics Analysis: Nano-UHPLC and ESI-MS
4.7. Identification and Label-Free Protein Quantification
4.8. Gene Set Enrichment Analysis
4.9. Bioinformatics Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Waks, A.G.; Winer, E.P. Breast Cancer Treatment: A Review. JAMA 2019, 321, 288. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, F.; Harbeck, N.; Barrios, C.H.; Bergh, J.; Cortés, J.; El Saghir, N.; Francis, P.A.; Hudis, C.A.; Ohno, S.; Partridge, A.H.; et al. Research Needs in Breast Cancer. Ann. Oncol. 2017, 28, 208–217. [Google Scholar] [CrossRef] [PubMed]
- DeMichele, A.; Yee, D.; Esserman, L. Mechanisms of Resistance to Neoadjuvant Chemotherapy in Breast Cancer. N. Engl. J. Med. 2017, 377, 2287–2289. [Google Scholar] [CrossRef] [PubMed]
- McCoach, C.E.; Bivona, T.G. Engineering Multidimensional Evolutionary Forces to Combat Cancer. Cancer Discov. 2019, 9, 587–604. [Google Scholar] [CrossRef] [PubMed]
- Ganapathy-Kanniappan, S.; Geschwind, J.-F.H. Tumor Glycolysis as a Target for Cancer Therapy: Progress and Prospects. Mol. Cancer 2013, 12, 152. [Google Scholar] [CrossRef] [PubMed]
- Gu-Trantien, C.; Loi, S.; Garaud, S.; Equeter, C.; Libin, M.; de Wind, A.; Ravoet, M.; Le Buanec, H.; Sibille, C.; Manfouo-Foutsop, G.; et al. CD4+ Follicular Helper T Cell Infiltration Predicts Breast Cancer Survival. J. Clin. Investig. 2013, 123, 2873–2892. [Google Scholar] [CrossRef] [PubMed]
- Westermann, J.; Pabst, R. Lymphocyte Subsets in the Blood: A Diagnostic Window on the Lymphoid System? Immunol. Today 1990, 11, 406–410. [Google Scholar] [CrossRef] [PubMed]
- Yerinde, C.; Siegmund, B.; Glauben, R.; Weidinger, C. Metabolic Control of Epigenetics and Its Role in CD8+ T Cell Differentiation and Function. Front. Immunol. 2019, 10, 2718. [Google Scholar] [CrossRef] [PubMed]
- Almeida, L.; Lochner, M.; Berod, L.; Sparwasser, T. Metabolic Pathways in T Cell Activation and Lineage Differentiation. Semin. Immunol. 2016, 28, 514–524. [Google Scholar] [CrossRef]
- Roncati, L.; Barbolini, G.; Piacentini, F.; Piscioli, F.; Pusiol, T.; Maiorana, A. Prognostic Factors for Breast Cancer: An Immunomorphological Update. Pathol. Oncol. Res. 2015, 22, 449–452. [Google Scholar] [CrossRef] [PubMed]
- Poschke, I.; De Boniface, J.; Mao, Y.; Kiessling, R. Tumor-induced Changes in the Phenotype of Blood-derived and Tumor-associated T Cells of Early Stage Breast Cancer Patients. Int. J. Cancer 2012, 131, 1611–1620. [Google Scholar] [CrossRef] [PubMed]
- Hay, Z.L.Z.; Slansky, J.E. Granzymes: The Molecular Executors of Immune-Mediated Cytotoxicity. Int. J. Mol. Sci. 2022, 23, 1833. [Google Scholar] [CrossRef] [PubMed]
- Bots, M.; Medema, J.P. Granzymes at a Glance. J. Cell Sci. 2006, 119, 5011–5014. [Google Scholar] [CrossRef] [PubMed]
- Jenne, D.E.; Tschopp, J. Granzymes, a Family of Serine Proteases Released from Granules of Cytolytic T Lymphocytes upon T Cell Receptor Stimulation. Immunol. Rev. 1988, 103, 53–71. [Google Scholar] [CrossRef] [PubMed]
- Dotiwala, F.; Mulik, S.; Polidoro, R.B.; Ansara, J.A.; Burleigh, B.A.; Walch, M.; Gazzinelli, R.T.; Lieberman, J. Killer Lymphocytes Use Granulysin, Perforin and Granzymes to Kill Intracellular Parasites. Nat. Med. 2016, 22, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Law, R.H.P.; Lukoyanova, N.; Voskoboinik, I.; Caradoc-Davies, T.T.; Baran, K.; Dunstone, M.A.; D’Angelo, M.E.; Orlova, E.V.; Coulibaly, F.; Verschoor, S.; et al. The Structural Basis for Membrane Binding and Pore Formation by Lymphocyte Perforin. Nature 2010, 468, 447–451. [Google Scholar] [CrossRef] [PubMed]
- Odake, S.; Kam, C.M.; Narasimhan, L.; Poe, M.; Blake, J.T.; Krahenbuhl, O.; Tschopp, J.; Powers, J.C. Human and Murine Cytotoxic T Lymphocyte Serine Proteases: Subsite Mapping with Peptide Thioester Substrates and Inhibition of Enzyme Activity and Cytolysis by Isocoumarins. Biochemistry 1991, 30, 2217–2227. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, D.; Lieberman, J. Death by a Thousand Cuts: Granzyme Pathways of Programmed Cell Death. Annu. Rev. Immunol. 2008, 26, 389–420. [Google Scholar] [CrossRef]
- Choi, J.O.; Ham, J.H.; Hwang, S.S. RNA Metabolism in T Lymphocytes. Immune Netw. 2022, 22, e39. [Google Scholar] [CrossRef]
- Spathas, N.; Goussia, A.C.; Koliou, G.-A.; Gogas, H.; Zagouri, F.; Batistatou, A.; Charchanti, A.V.; Papoudou-Bai, A.; Bobos, M.; Chrisafi, S.; et al. Association between CD8+ Tumor Infiltrating Lymphocytes and the Clinical Outcome of Patients with Operable Breast Cancer Treated with Adjuvant Dose-Dense Chemotherapy—A 10 Year Follow-Up Report of a Hellenic Cooperative Oncology Group Observational Study. Cancers 2022, 14, 5635. [Google Scholar] [CrossRef]
- Valenza, C.; Taurelli Salimbeni, B.; Santoro, C.; Trapani, D.; Antonarelli, G.; Curigliano, G. Tumor Infiltrating Lymphocytes across Breast Cancer Subtypes: Current Issues for Biomarker Assessment. Cancers 2023, 15, 767. [Google Scholar] [CrossRef]
- Chang, C.-M.; Wu, C.-L.; Lu, Y.-T. Cancer-Associated Immune Deficiency: A Form of Accelerated Immunosenescence? In Topics in Cancer Survivorship; InTech: London, UK, 2012. [Google Scholar]
- Li, M.; Xu, J.; Jiang, C.; Zhang, J.; Sun, T. Predictive and Prognostic Role of Peripheral Blood T-Cell Subsets in Triple-Negative Breast Cancer. Front. Oncol. 2022, 12, 842705. [Google Scholar] [CrossRef]
- Papaioannou, E.; Sakellakis, M.; Melachrinou, M.; Tzoracoleftherakis, E.; Kalofonos, H.; Kourea, E. A Standardized Evaluation Method for FOXP3+ Tregs and CD8+ T-cells in Breast Carcinoma: Association With Breast Carcinoma Subtypes, Stage and Prognosis. Anticancer Res. 2019, 39, 1217–1232. [Google Scholar] [CrossRef] [PubMed]
- Song, I.H.; Heo, S.-H.; Bang, W.S.; Park, H.S.; Park, I.A.; Kim, Y.-A.; Park, S.Y.; Roh, J.; Gong, G.; Lee, H.J. Predictive Value of Tertiary Lymphoid Structures Assessed by High Endothelial Venule Counts in the Neoadjuvant Setting of Triple-Negative Breast Cancer. Cancer Res. Treat. 2017, 49, 399–407. [Google Scholar] [CrossRef] [PubMed]
- Berke, G. The Binding and Lysis of Target Cells by Cytotoxic Lymphocytes: Molecular and Cellular Aspects. Annu. Rev. Immunol. 1994, 12, 735–773. [Google Scholar] [CrossRef]
- Catalfamo, M.; Henkart, P.A. Perforin and the Granule Exocytosis Cytotoxicity Pathway. Curr. Opin. Immunol. 2003, 15, 522–527. [Google Scholar] [CrossRef]
- Voskoboinik, I.; Dunstone, M.A.; Baran, K.; Whisstock, J.C.; Trapani, J.A. Perforin: Structure, Function, and Role in Human Immunopathology. Immunol. Rev. 2010, 235, 35–54. [Google Scholar] [CrossRef]
- Hombach, A.; Köhler, H.; Rappl, G.; Abken, H. Human CD4+ T Cells Lyse Target Cells via Granzyme/Perforin upon Circumvention of MHC Class II Restriction by an Antibody-Like Immunoreceptor. J. Immunol. 2006, 177, 5668–5675. [Google Scholar] [CrossRef]
- Andrade, F.; Roy, S.; Nicholson, D.; Thornberry, N.; Rosen, A.; Casciola-Rosen, L. Granzyme B Directly and Efficiently Cleaves Several Downstream Caspase Substrates: Implications for CTL-Induced Apoptosis. Immunity 1998, 8, 451–460. [Google Scholar] [CrossRef]
- Zhou, Z.; He, H.; Wang, K.; Shi, X.; Wang, Y.; Su, Y.; Wang, Y.; Li, D.; Liu, W.; Zhang, Y.; et al. Granzyme A from Cytotoxic Lymphocytes Cleaves GSDMB to Trigger Pyroptosis in Target Cells. Science 2020, 368, eaaz7548. [Google Scholar] [CrossRef] [PubMed]
- Martinvalet, D.; Zhu, P.; Lieberman, J. Granzyme A Induces Caspase-Independent Mitochondrial Damage, a Required First Step for Apoptosis. Immunity 2005, 22, 355–370. [Google Scholar] [CrossRef] [PubMed]
- Tahbaz, B.; Amirrasouli, H.; Kazerouni, F.; Rahimipour, A.; Mohammadzade, J. Comparative Study of Serum Levels of Granzyme H & Estrogen In Patients Suffering from Breast Cancer. Arch. Adv. Biosci. 2013, 4, 26–28. [Google Scholar]
- Razvi, N.Z.; Khurshid, R.; Nagra, S.A. To Study the Significance of Apoptotic Enzyme Granzyme H in Breast Cancer Patients. J. Ayub Med. Coll. Abbottabad 2008, 20, 84–86. [Google Scholar] [PubMed]
- Mahrus, S.; Craik, C.S. Selective Chemical Functional Probes of Granzymes A and B Reveal Granzyme B Is a Major Effector of Natural Killer Cell-Mediated Lysis of Target Cells. Chem. Biol. 2005, 12, 567–577. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Kam, C.M.; Powers, J.C.; Aebersold, R.; Greenberg, A.H. Purification of Three Cytotoxic Lymphocyte Granule Serine Proteases That Induce Apoptosis through Distinct Substrate and Target Cell Interactions. J. Exp. Med. 1992, 176, 1521–1529. [Google Scholar] [CrossRef]
- Zhao, T.; Zhang, H.; Guo, Y.; Zhang, Q.; Hua, G.; Lu, H.; Hou, Q.; Liu, H.; Fan, Z. Granzyme K Cleaves the Nucleosome Assembly Protein SET to Induce Single-Stranded DNA Nicks of Target Cells. Cell Death Differ. 2006, 14, 489–499. [Google Scholar] [CrossRef]
- Zhao, T.; Zhang, H.; Guo, Y.; Fan, Z. Granzyme K Directly Processes Bid to Release Cytochrome c and Endonuclease G Leading to Mitochondria-Dependent Cell Death. J. Biol. Chem. 2007, 282, 12104–12111. [Google Scholar] [CrossRef]
- Mogilenko, D.A.; Shpynov, O.; Andhey, P.S.; Arthur, L.; Swain, A.; Esaulova, E.; Brioschi, S.; Shchukina, I.; Kerndl, M.; Bambouskova, M.; et al. Comprehensive Profiling of an Aging Immune System Reveals Clonal GZMK+ CD8+ T Cells as Conserved Hallmark of Inflammaging. Immunity 2021, 54, 99–115.e12. [Google Scholar] [CrossRef]
- de Poot, S.A.H.; Westgeest, M.; Hostetter, D.R.; van Damme, P.; Plasman, K.; Demeyer, K.; Broekhuizen, R.; Gevaert, K.; Craik, C.S.; Bovenschen, N. Human and Mouse Granzyme M Display Divergent and Species-Specific Substrate Specificities. Biochem. J. 2011, 437, 431–442. [Google Scholar] [CrossRef]
- Wang, S.; Xia, P.; Shi, L.; Fan, Z. FADD Cleavage by NK Cell Granzyme M Enhances Its Self-Association to Facilitate Procaspase-8 Recruitment for Auto-Processing Leading to Caspase Cascade. Cell Death Differ. 2012, 19, 605–615. [Google Scholar] [CrossRef]
- Kelly, J.M.; Waterhouse, N.J.; Cretney, E.; Browne, K.A.; Ellis, S.; Trapani, J.A.; Smyth, M.J. Granzyme M Mediates a Novel Form of Perforin-Dependent Cell Death. J. Biol. Chem. 2004, 279, 22236–22242. [Google Scholar] [CrossRef] [PubMed]
- Łabno, A.; Tomecki, R.; Dziembowski, A. Cytoplasmic RNA Decay Pathways–Enzymes and Mechanisms. Biochim. Biophys. Acta-Mol. Cell Res. 2016, 1863, 3125–3147. [Google Scholar] [CrossRef] [PubMed]
- Schoenberg, D.R.; Maquat, L.E. Regulation of Cytoplasmic MRNA Decay. Nat. Rev. Genet. 2012, 13, 246–259. [Google Scholar] [CrossRef]
- Akiyama, T.; Suzuki, T.; Yamamoto, T. RNA Decay Machinery Safeguards Immune Cell Development and Immunological Responses. Trends Immunol. 2021, 42, 447–460. [Google Scholar] [CrossRef]
- Wolf, T.; Jin, W.; Zoppi, G.; Vogel, I.A.; Akhmedov, M.; Bleck, C.K.E.; Beltraminelli, T.; Rieckmann, J.C.; Ramirez, N.J.; Benevento, M.; et al. Dynamics in Protein Translation Sustaining T Cell Preparedness. Nat. Immunol. 2020, 21, 927–937. [Google Scholar] [CrossRef]
- Wilusz, C.J.; Wilusz, J. Lsm Proteins and Hfq: Life at the 3′ End. RNA Biol. 2013, 10, 592–601. [Google Scholar] [CrossRef]
- Goldhirsch, A.; Winer, E.P.; Coates, A.S.; Gelber, R.D.; Piccart-Gebhart, M.; Thürlimann, B.; Senn, H.-J.; Panel Members. Personalizing the treatment of women with early breast cancer: Highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann. Oncol. 2013, 24, 2206–2223. [Google Scholar] [CrossRef]
- Giuliano, A.E.; Edge, S.B.; Hortobagyi, G.N. Eighth edition of the AJCC cancer staging manual: Breast cancer. Ann. Surg. Oncol. 2018, 25, 1783–1785. [Google Scholar] [CrossRef]


| Characteristic | Benign | Luminal A (n) | Luminal B (n) | TNBC (n) |
|---|---|---|---|---|
| Mean (IQR) age | 53 (39–73) | 69 (57–76) | 61 (46–81) | 72 (57–83) |
| Site, Left/Right | Left (8) | Left (3), Right (5) | Left (3), Right (5) | Left (2), Right(1) |
| Histologic type | Fibrosclerosis (4) Xantogranuloma(1) Ductus ectasia (1) Osseal metaplasis (1) Intracanalicular adenoma (1) | Invasive ductal carcinoma (8) | Invasive ductal carcinoma (8) | Invasive ductal carcinoma (3) |
| Histological grade | - | G1 (7), G2 (1) | G1 (3), G2 (5) | G3 (3) |
| Pathological tumour size | - | pT1 (5), pT2 (3) | pT1 (5), pT2 (3) | pT1 (1), pT2 (2) |
| Pathological node status | - | pN0 (7), pN1 (1) | pN0 (4), pN1 (1), pN2 (3) | pN0 (1), pN2 (2) |
| Lymph node ratio | - | ≤20 (8) | ≤20 (5), 0.21–0.65 (3) | ≤20 (2), 0.21–0.65 (1) |
| Prognostic stage | - | IA (4), IIA (4) | IA (4), IIB (1), IIIA (3) | IA (1), IIIA (2) |
| Metastasis | - | Axillary lymph node metastasis (1) | Axillary lymph node metastasis (4) | Axillary lymph node metastasis (2) |
| Proliferation index (Ki-67) | - | <14% (8) | ≥14% (8) | ≥14% (3) |
| ER expression | - | 90–100% (8) | 30–95% (8) | - |
| PR expression | - | 60–100% (8) | 20–90% (8) | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lenárt, M.; Bober, P.; Marcin, M.; Tkáčiková, S.; Kacírová, M.; Alexovič, M.; Tóth, D.; Madárová, N.; Radoňak, J.; Urdzík, P.; et al. Peripheral Blood CD8+ T-Lymphocyte Immune Response in Benign and Subpopulations of Breast Cancer Patients. Int. J. Mol. Sci. 2024, 25, 6423. https://doi.org/10.3390/ijms25126423
Lenárt M, Bober P, Marcin M, Tkáčiková S, Kacírová M, Alexovič M, Tóth D, Madárová N, Radoňak J, Urdzík P, et al. Peripheral Blood CD8+ T-Lymphocyte Immune Response in Benign and Subpopulations of Breast Cancer Patients. International Journal of Molecular Sciences. 2024; 25(12):6423. https://doi.org/10.3390/ijms25126423
Chicago/Turabian StyleLenárt, Marek, Peter Bober, Miroslav Marcin, Soňa Tkáčiková, Mária Kacírová, Michal Alexovič, Dávid Tóth, Natália Madárová, Jozef Radoňak, Peter Urdzík, and et al. 2024. "Peripheral Blood CD8+ T-Lymphocyte Immune Response in Benign and Subpopulations of Breast Cancer Patients" International Journal of Molecular Sciences 25, no. 12: 6423. https://doi.org/10.3390/ijms25126423
APA StyleLenárt, M., Bober, P., Marcin, M., Tkáčiková, S., Kacírová, M., Alexovič, M., Tóth, D., Madárová, N., Radoňak, J., Urdzík, P., Fedačko, J., & Sabo, J. (2024). Peripheral Blood CD8+ T-Lymphocyte Immune Response in Benign and Subpopulations of Breast Cancer Patients. International Journal of Molecular Sciences, 25(12), 6423. https://doi.org/10.3390/ijms25126423






