Role and Regulatory Mechanism of circRNA_14820 in the Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells
Abstract
1. Introduction
2. Results
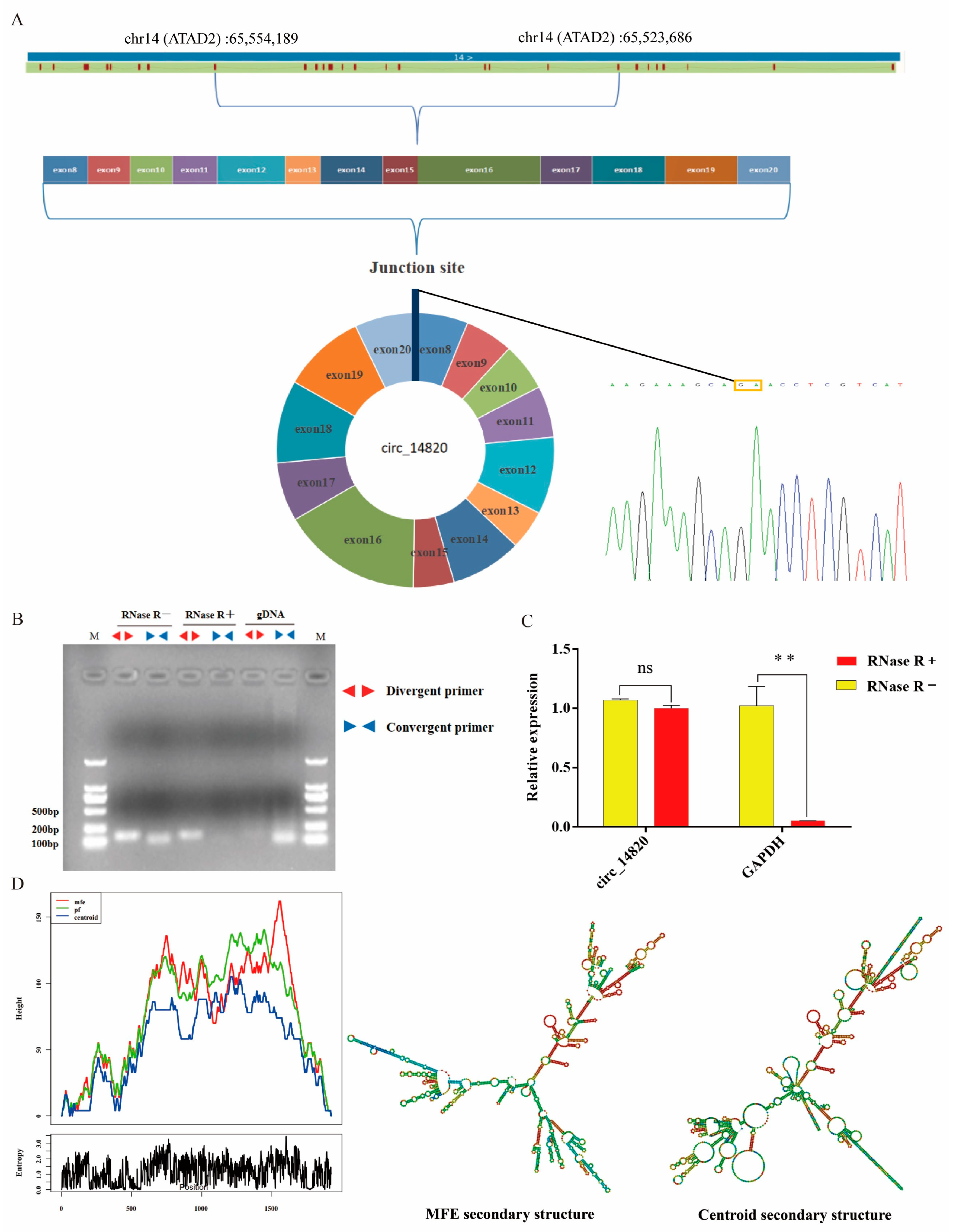
2.1. Identification of Circ_14820
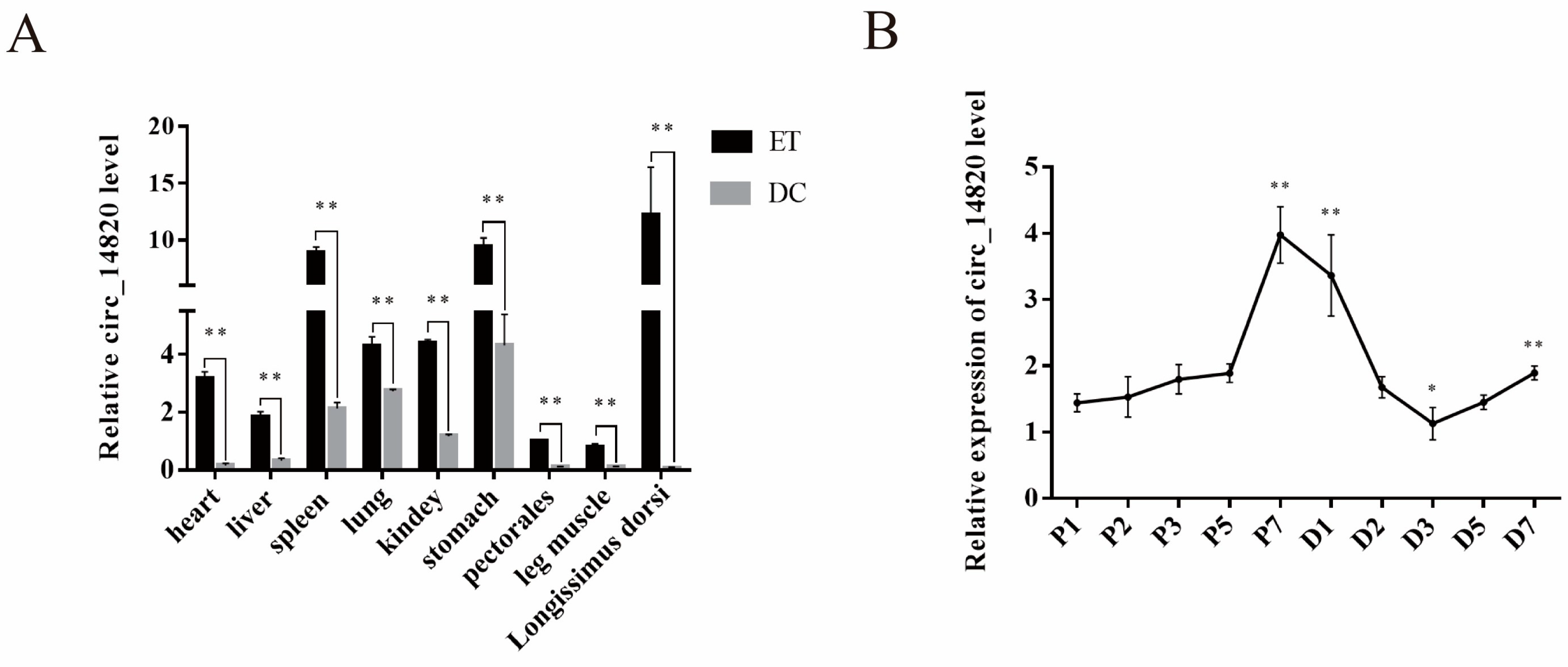
2.2. Spatiotemporal Expression Pattern of Circ_14820
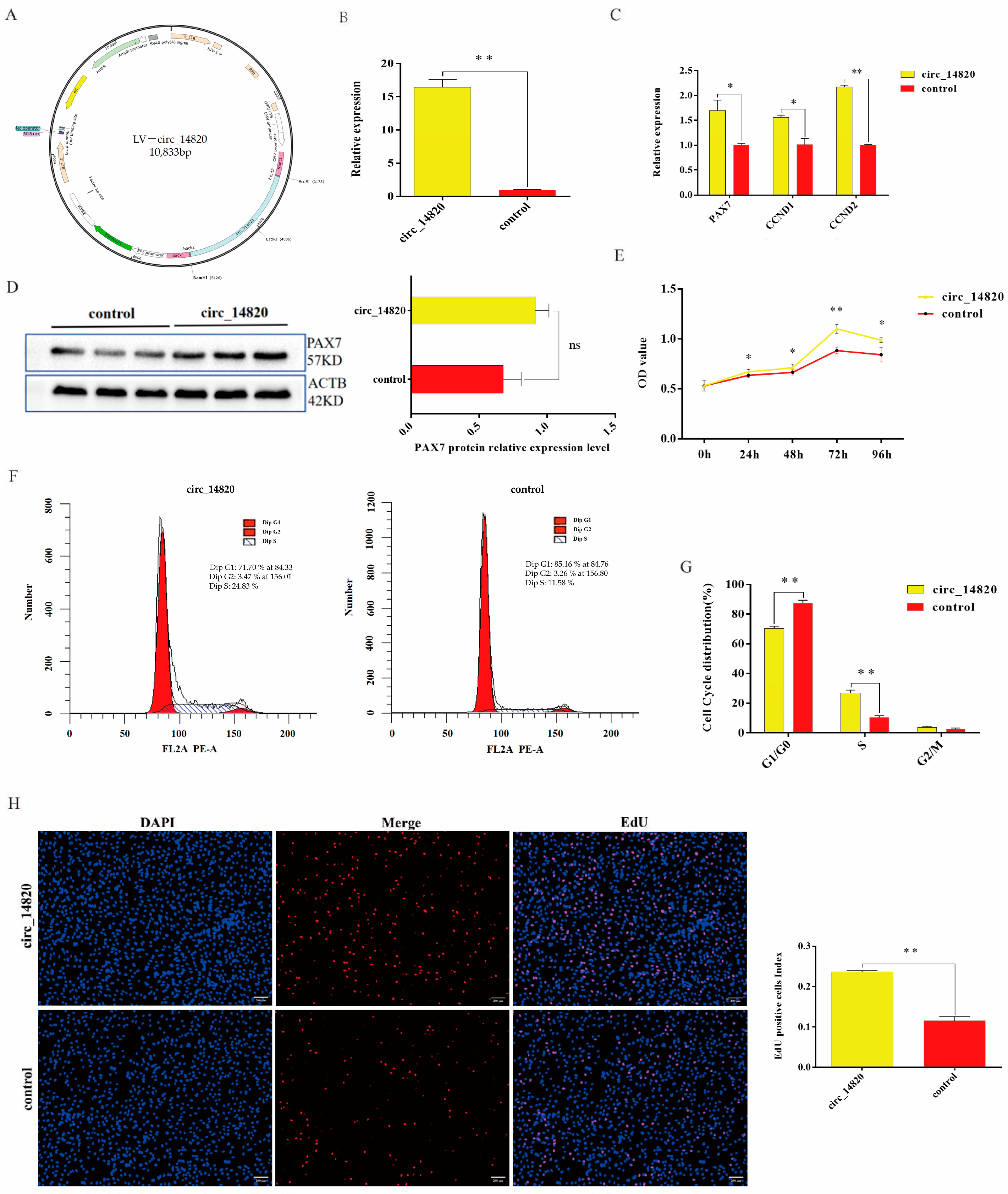
2.3. Circ_14820 Promotes Proliferation of SMSCs
2.4. Circ_14820 Inhibits Differentiation of SMSCs
2.5. Nucleoplasmic Localization of Circ_14820
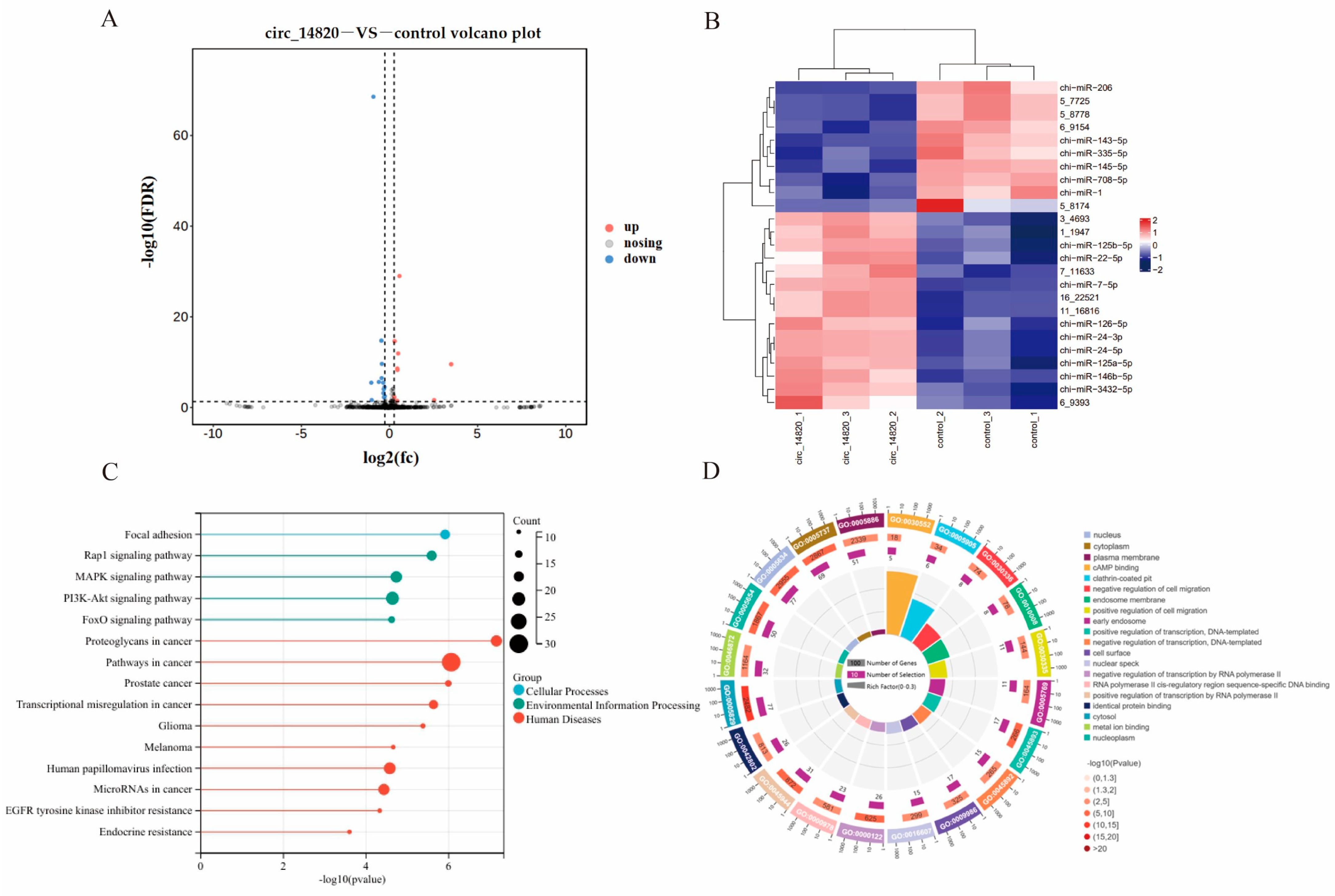
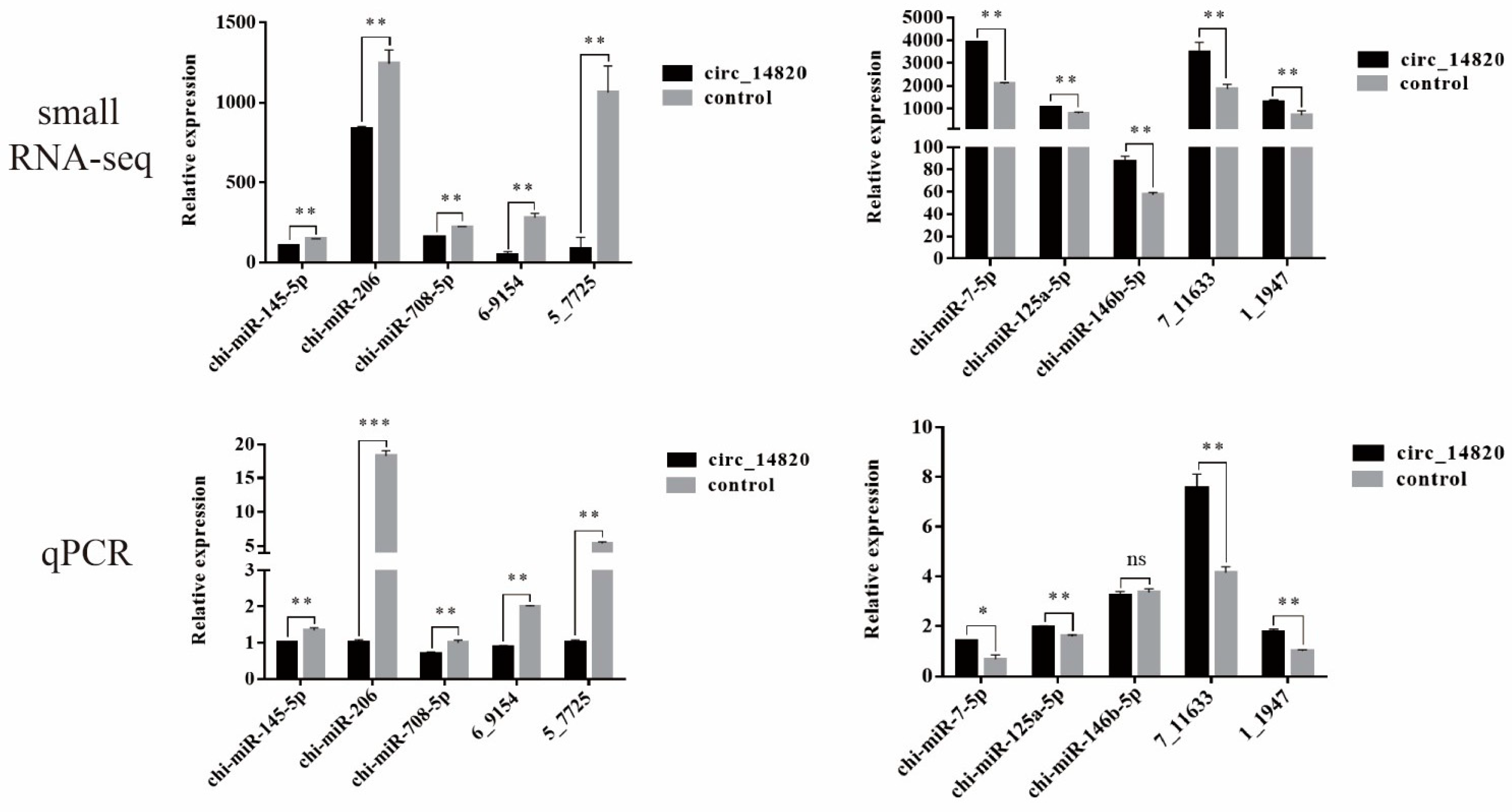
2.6. Analysis of miRNAs Regulated by Circ_14820
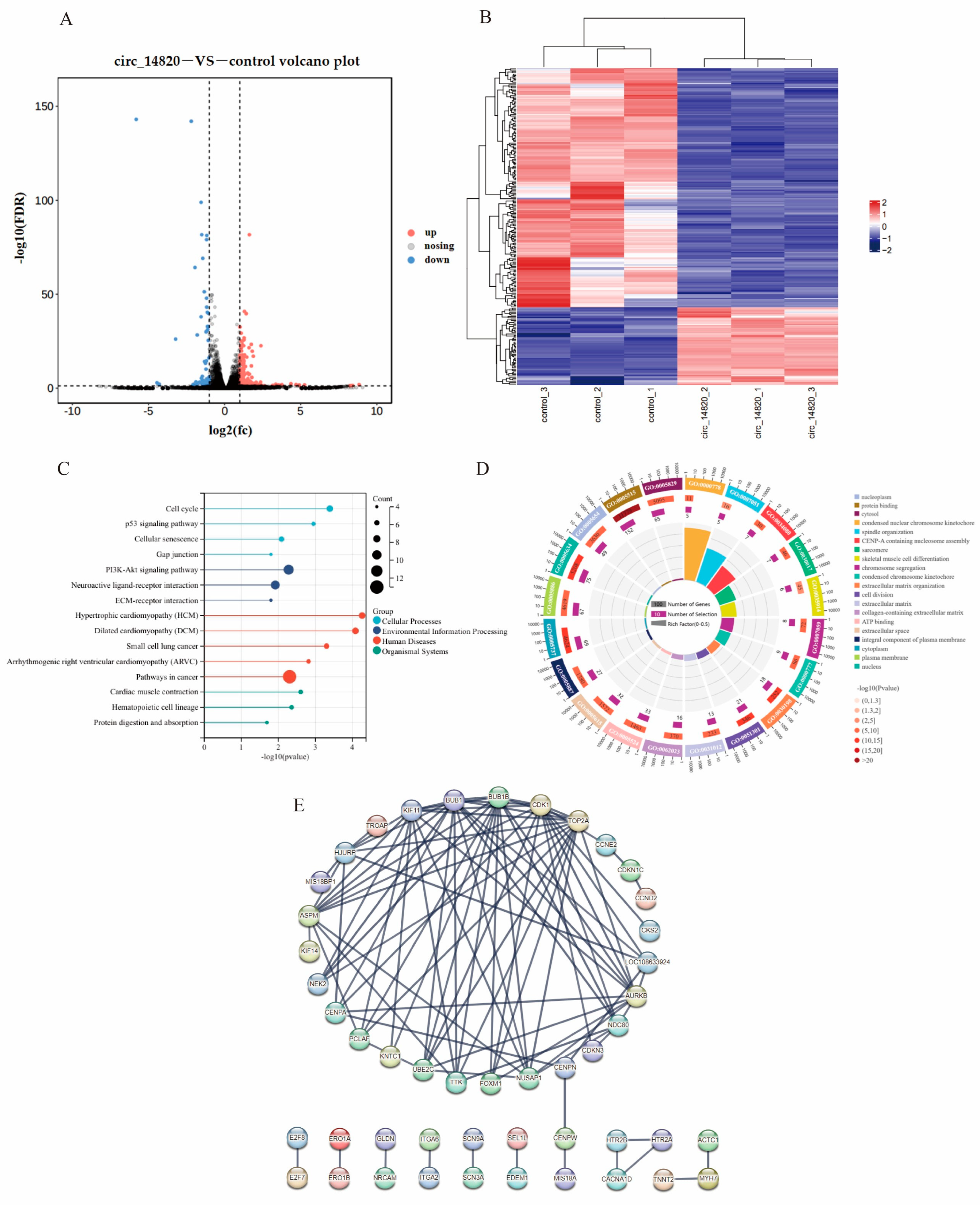
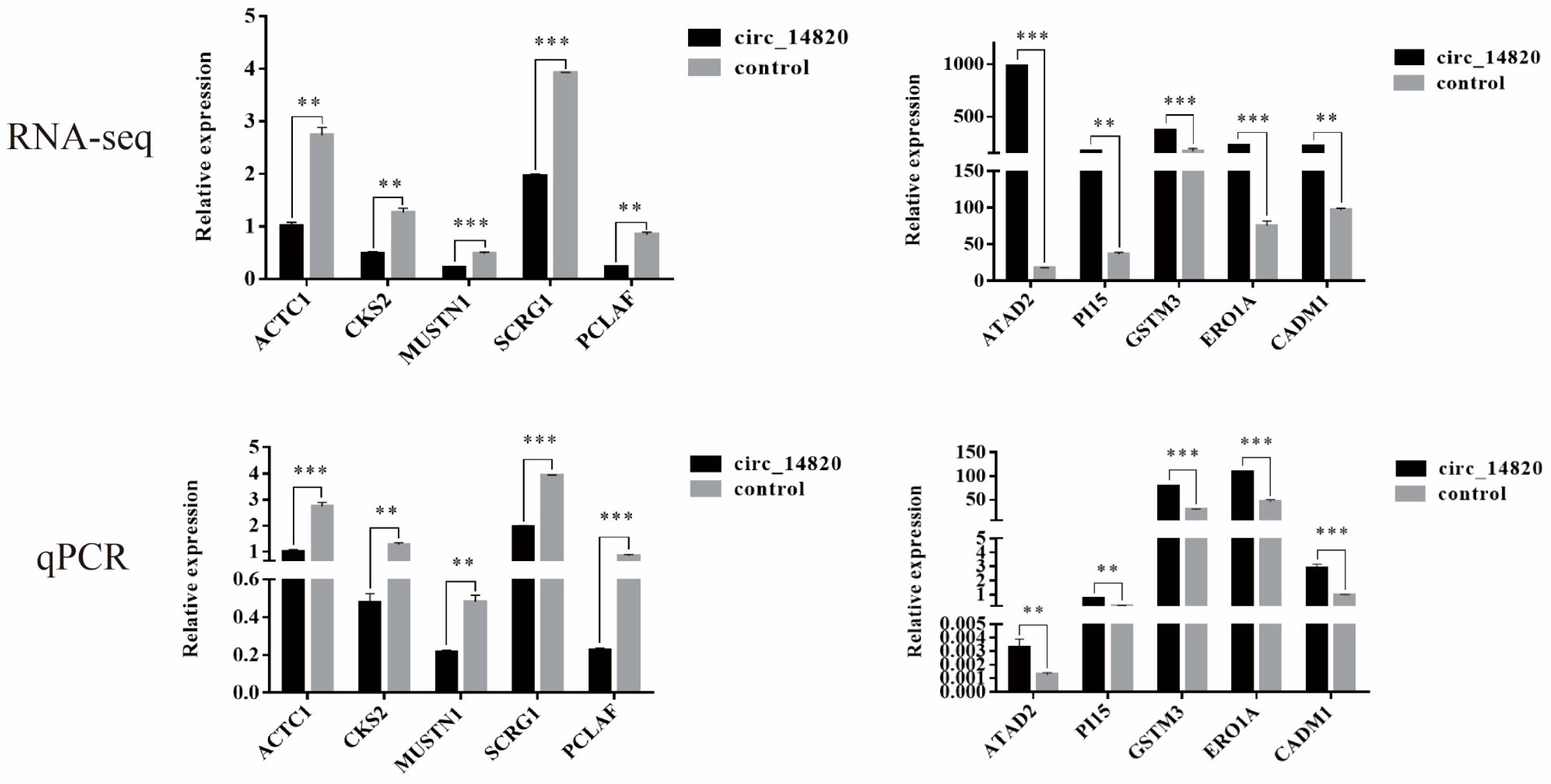
2.7. Analysis of mRNAs Regulated by Circ_14820
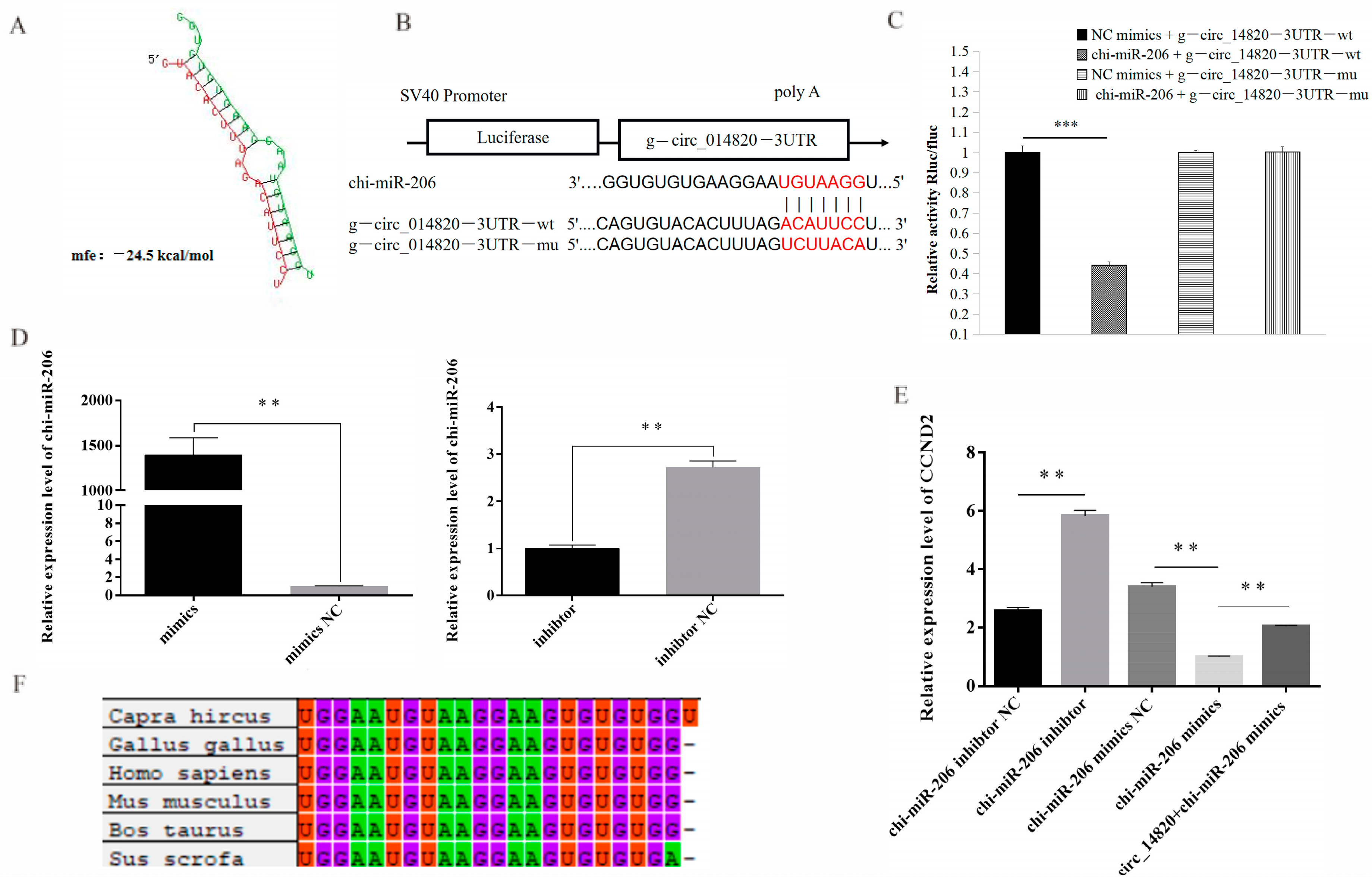
2.8. Analysis of Circ_14820-miR-206-CCND2 Regulatory Axis
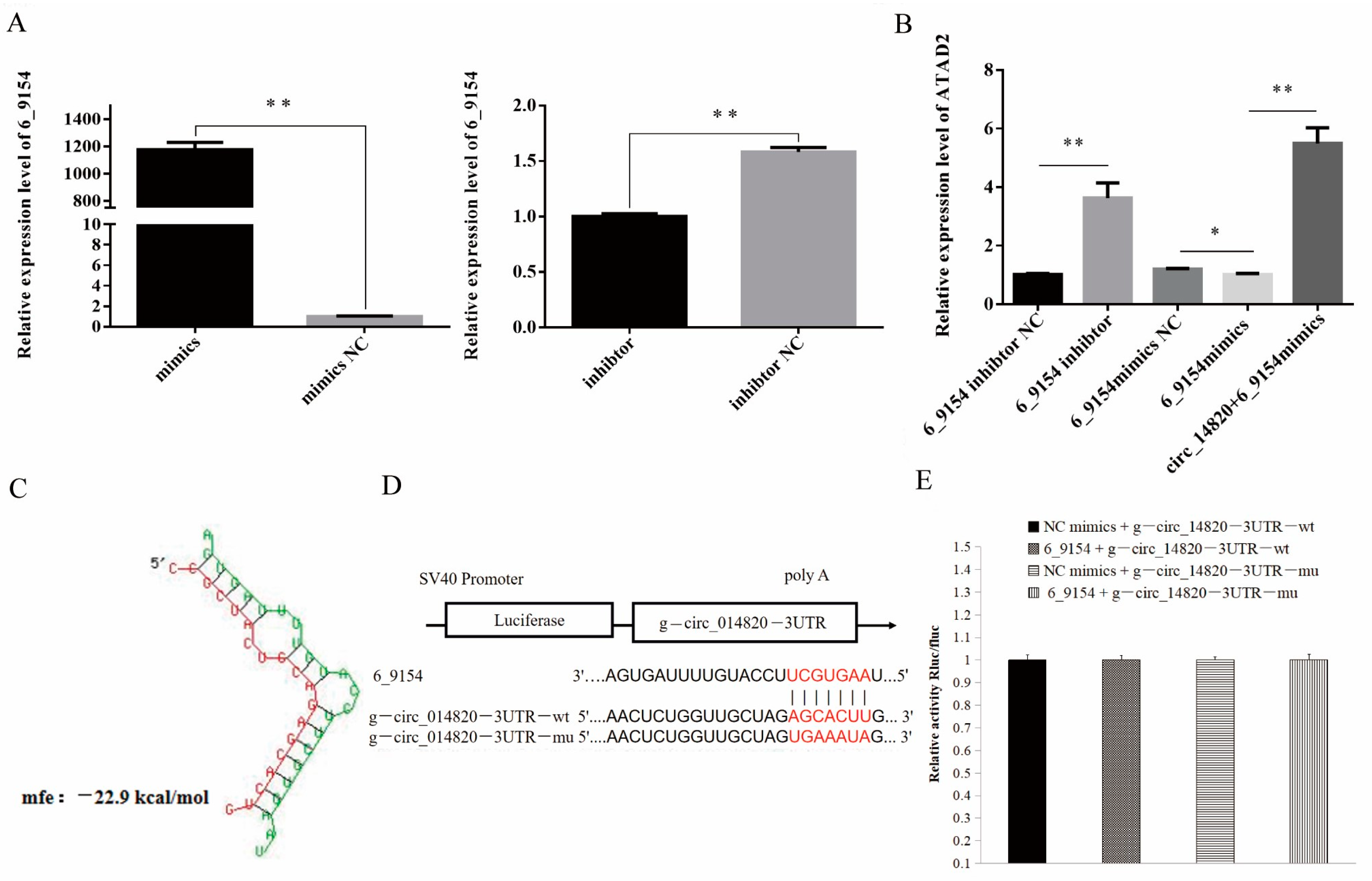
2.9. circ_14820 Regulates ATAD2 Expression through Novel miRNA 6-9154
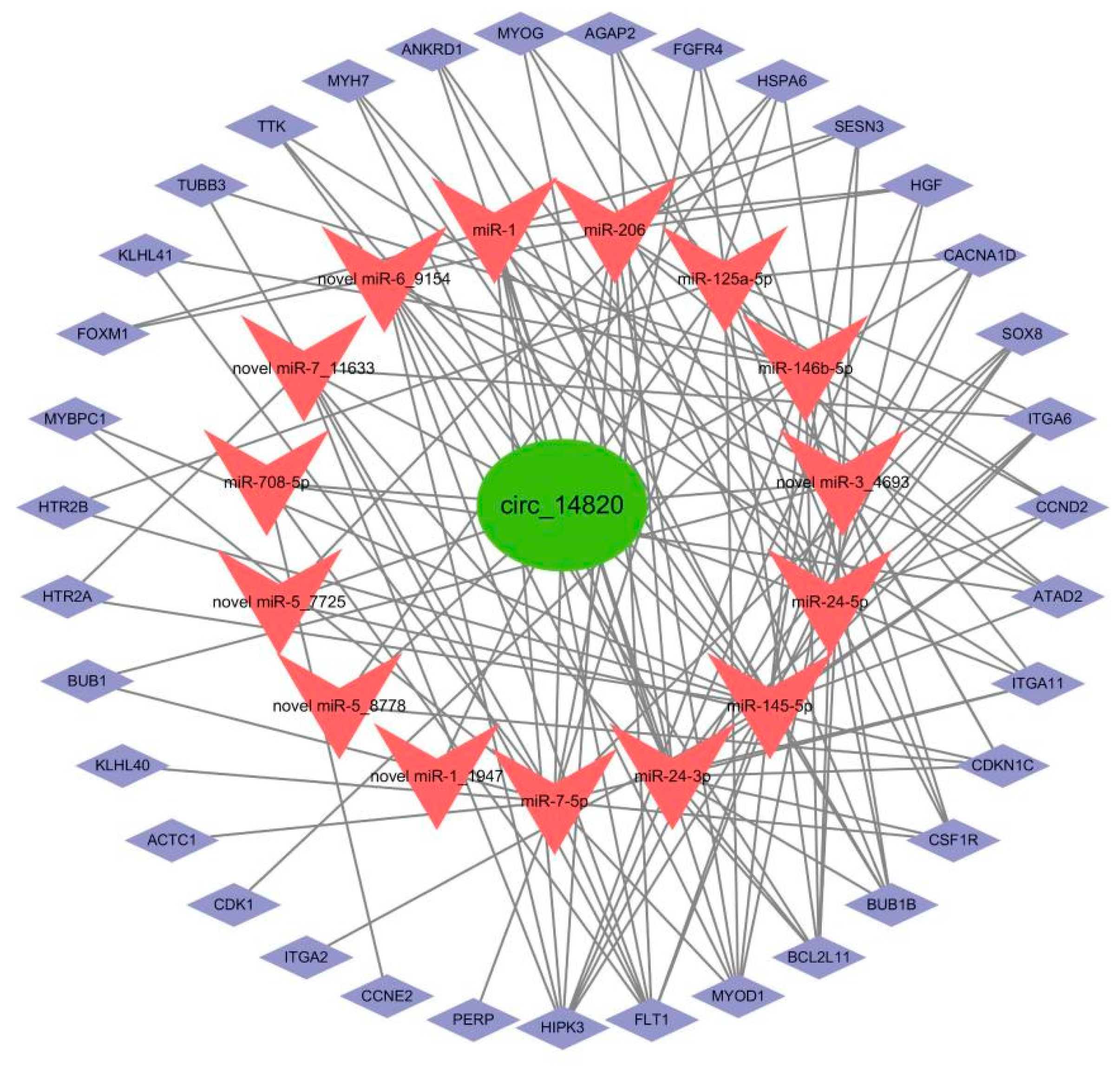
2.10. Construction of Circ_14820-miRNA-mRNA Ternary Regulatory Network
3. Discussion
4. Materials and Methods
4.1. Ethics Statement
4.2. Animal
4.3. Isolation and Identification of SMSCs
4.4. RNA Extractions and qPCR
4.5. Circ_14820 Identification
4.6. Vector Construction and Transfection
4.7. Cell Counting Kit-8 (CCK-8) and EdU Assay
4.8. MyHC Immunofluorescence Assay
4.9. Flow Cytometry Cell Cycle Analysis
4.10. Western Blotting (WB) Assay
4.11. Nucleoplasmic Separation of SMSCs
4.12. Small RNA and MRNA Sequencing Analysis
4.13. Analysis of the Luciferase Report
4.14. Bioinformatics Analysis
4.15. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Megeney, L.A.; Rudnicki, M.A. Determination versus differentiation and the MyoD family of transcription factors. Biochem. Cell Biol. 1995, 73, 723–732. [Google Scholar] [CrossRef] [PubMed]
- Nabeshima, Y.; Hanaoka, K.; Hayasaka, M.; Esumi, E.; Li, S.; Nonaka, I.; Nabeshima, Y. Myogenin gene disruption results in perinatal lethality because of severe muscle defect. Nature 1993, 364, 532–535. [Google Scholar] [CrossRef] [PubMed]
- Black, B.L.; Olson, E.N. Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins. Annu. Rev. Cell Dev. Biol. 1998, 14, 167–196. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Song, W.; Jin, C.; Huang, K.; Yu, Q.; Qi, J.; Zhang, Q.; He, Y. Pax3 and Pax7 Exhibit Distinct and Overlapping Functions in Marking Muscle Satellite Cells and Muscle Repair in a Marine Teleost, Sebastes schlegelii. Int. J. Mol. Sci. 2021, 22, 3769. [Google Scholar] [CrossRef]
- Otto, A.; Schmidt, C.; Luke, G.; Allen, S.; Valasek, P.; Muntoni, F.; Lawrence-Watt, D.; Patel, K. Canonical Wnt signalling induces satellite-cell proliferation during adult skeletal muscle regeneration. J. Cell Sci. 2008, 121, 2939–2950. [Google Scholar] [CrossRef] [PubMed]
- Mayeuf-Louchart, A.; Lagha, M.; Danckaert, A.; Rocancourt, D.; Relaix, F.; Vincent, S.D.; Buckingham, M. Notch regulation of myogenic versus endothelial fates of cells that migrate from the somite to the limb. Proc. Natl. Acad. Sci. USA 2014, 111, 8844–8849. [Google Scholar] [CrossRef] [PubMed]
- Simone, C.; Forcales, S.V.; Hill, D.A.; Imbalzano, A.N.; Latella, L.; Puri, P.L. p38 pathway targets SWI-SNF chromatin-remodeling complex to muscle-specific loci. Nature Genet. 2004, 36, 738–743. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Fu, Y.; Pang, Y.; Tong, H.; Li, S.; Yan, Y. GRP94 promotes muscle differentiation by inhibiting the PI3K/AKT/mTOR signaling pathway. J. Cell. Physiol. 2019, 234, 21211–21223. [Google Scholar] [CrossRef]
- Ling, Y.; Zheng, Q.; Zhu, L.; Xu, L.; Sui, M.; Zhang, Y.; Liu, Y.; Fang, F.; Chu, M.; Ma, Y.; et al. Trend analysis of the role of circular RNA in goat skeletal muscle development. BMC Genom. 2020, 21, 220. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef]
- Wang, X.; Li, H.; Lu, Y.; Cheng, L. Circular RNAs in Human Cancer. Front. Oncol. 2020, 10, 577118. [Google Scholar] [CrossRef]
- Dong, R.; Ma, X.K.; Chen, L.L.; Yang, L. Increased complexity of circRNA expression during species evolution. RNA Biol. 2017, 14, 1064–1074. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Yang, Y.; Niu, G.; Tang, Z.; Li, K. Genome-wide profiling of Sus scrofa circular RNAs across nine organs and three developmental stages. DNA Res. 2017, 24, 523–535. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Westholm, J.O.; Miura, P.; Olson, S.; Shenker, S.; Joseph, B.; Sanfilippo, P.; Celniker, S.E.; Graveley, B.R.; Lai, E.C. Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 2014, 9, 1966–1980. [Google Scholar] [CrossRef] [PubMed]
- Rybak-Wolf, A.; Stottmeister, C.; Glazar, P.; Jens, M.; Pino, N.; Giusti, S.; Hanan, M.; Behm, M.; Bartok, O.; Ashwal-Fluss, R.; et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell 2015, 58, 870–885. [Google Scholar] [CrossRef]
- Wei, X.; Li, H.; Yang, J.; Hao, D.; Dong, D.; Huang, Y.; Lan, X.; Plath, M.; Lei, C.; Lin, F.; et al. Circular RNA profiling reveals an abundant circLMO7 that regulates myoblasts differentiation and survival by sponging miR-378a-3p. Cell Death Dis. 2017, 8, e3153. [Google Scholar] [CrossRef]
- Li, L.; Chen, Y.; Nie, L.; Ding, X.; Zhang, X.; Zhao, W.; Xu, X.; Kyei, B.; Dai, D.; Zhan, S.; et al. MyoD-induced circular RNA CDR1as promotes myogenic differentiation of skeletal muscle satellite cells. Biochim. Biophys. Acta-Gene Regul. Mech. 2019, 1862, 807–821. [Google Scholar] [CrossRef]
- Xu, J.; Wen, Y.; Li, X.; Peng, W.; Zhang, Z.; Liu, X.; Yang, P.; Chen, N.; Lei, C.; Zhang, J.; et al. Bovine enhancer-regulated circSGCB acts as a ceRNA to regulate skeletal muscle development via enhancing KLF3 expression. Int. J. Biol. Macromol. 2024, 261, 129779. [Google Scholar] [CrossRef]
- Fu, J.; Zhang, J.; Chen, X.; Liu, Z.; Yang, X.; He, Z.; Hao, Y.; Liu, B.; Yao, D. ATPase family AAA domain-containing protein 2 (ATAD2): From an epigenetic modulator to cancer therapeutic target. Theranostics 2023, 13, 787–809. [Google Scholar] [CrossRef]
- Liu, Q.; Liu, H.; Li, L.; Dong, X.; Ru, X.; Fan, X.; Wen, T.; Liu, J. ATAD2 predicts poor outcomes in patients with ovarian cancer and is a marker of proliferation. Int. J. Oncol. 2020, 56, 219–231. [Google Scholar] [CrossRef]
- Wang, A.Q.; Lv, M.; Xu, Y.H.; Xie, P.M.; Dong, Y.Y. MiR-200b-5p inhibits proliferation of ovarian cancer cells by targeting ATAD2 and regulating PI3K/AKT signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 9860–9868. [Google Scholar] [CrossRef]
- Knudsen, E.S.; Wang, J.Y. Targeting the RB-pathway in cancer therapy. Clin. Cancer Res. 2010, 16, 1094–1099. [Google Scholar] [CrossRef]
- Liu, C.; Yang, P.; Wang, X.; Xiang, B.; E, G.; Huang, Y. Candidate circRNAs related to skeletal muscle development in Dazu black goats. Anim. Biotechnol. 2024, 35, 2286609. [Google Scholar] [CrossRef]
- Ouyang, H.; Chen, X.; Wang, Z.; Yu, J.; Jia, X.; Li, Z.; Luo, W.; Abdalla, B.A.; Jebessa, E.; Nie, Q.; et al. Circular RNAs are abundant and dynamically expressed during embryonic muscle development in chickens. DNA Res. 2018, 25, 71–86. [Google Scholar] [CrossRef]
- Yan, S.; Pei, Y.; Li, J.; Tang, Z.; Yang, Y. Recent Progress on Circular RNAs in the Development of Skeletal Muscle and Adipose Tissues of Farm Animals. Biomolecules 2023, 13, 314. [Google Scholar] [CrossRef]
- Zhou, Z.; Li, K.; Liu, J.; Zhang, H.; Fan, Y.; Chen, Y.; Han, H.; Yang, J.; Liu, Y. Expression Profile Analysis to Identify Circular RNA Expression Signatures in Muscle Development of Wu’an Goat Longissimus Dorsi Tissues. Front. Vet. Sci. 2022, 9, 833946. [Google Scholar] [CrossRef]
- Starke, S.; Jost, I.; Rossbach, O.; Schneider, T.; Schreiner, S.; Hung, L.H.; Bindereif, A. Exon circularization requires canonical splice signals. Cell Rep. 2015, 10, 103–111. [Google Scholar] [CrossRef]
- Welden, J.R.; van Doorn, J.; Nelson, P.T.; Stamm, S. The human MAPT locus generates circular RNAs. Biochim. Biophys. Acta-Mol. Basis Dis. 2018, 1864, 2753–2760. [Google Scholar] [CrossRef]
- Zhang, Z.; Fan, Y.; Deng, K.; Liang, Y.; Zhang, G.; Gao, X.; El-Samahy, M.A.; Zhang, Y.; Deng, M.; Wang, F. Circular RNA circUSP13 sponges miR-29c to promote differentiation and inhibit apoptosis of goat myoblasts by targeting IGF1. FASEB J. 2022, 36, e22097. [Google Scholar] [CrossRef]
- Fan, Y.; Zhang, Z.; Deng, K.; Kang, Z.; Guo, J.; Zhang, G.; Zhang, Y.; Wang, F. CircUBE3A promotes myoblasts proliferation and differentiation by sponging miR-28-5p to enhance expression. Int. J. Biol. Macromol. 2023, 226, 730–745. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Li, L.; Zhou, H.; Zhang, X.; Xu, X.; Dai, D.; Zhan, S.; Cao, J.; Guo, J.; Zhong, T.; et al. CircTCF4 Suppresses Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells Independent from AGO2 Binding. Int. J. Mol. Sci. 2022, 23, 12868. [Google Scholar] [CrossRef] [PubMed]
- Horak, M.; Novak, J.; Bienertova-Vasku, J. Muscle-specific microRNAs in skeletal muscle development. Dev. Biol. 2016, 410, 1–13. [Google Scholar] [CrossRef]
- Kim, H.K.; Lee, Y.S.; Sivaprasad, U.; Malhotra, A.; Dutta, A. Muscle-specific microRNA miR-206 promotes muscle differentiation. J. Cell Biol. 2006, 174, 677–687. [Google Scholar] [CrossRef]
- Xu, X.; Lu, H.; Xu, D.; Yu, Z.; Ai, N.; Wang, K.; Li, X.; He, J.; Jiang, J.; Ma, H.; et al. miR-708-5p Regulates Myoblast Proliferation and Differentiation. Vet. Sci. 2022, 9, 641. [Google Scholar] [CrossRef]
- Jin, J.; Li, F.; Fan, C.; Wu, Y.; He, C. Elevated mir-145-5p is associated with skeletal muscle dysfunction and triggers apoptotic cell death in C2C12 myotubes. J. Muscle Res. Cell Motil. 2022, 43, 135–145. [Google Scholar] [CrossRef]
- Lee, J.H.; Kim, S.W.; Han, J.S.; Shin, S.P.; Lee, S.I.; Park, T.S. Functional analyses of miRNA-146b-5p during myogenic proliferation and differentiation in chicken myoblasts. BMC Mol. Cell Biol. 2020, 21, 40. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.U.; Sass, S.; Mueller, N.S.; Krebs, S.; Bauersachs, S.; Kaiser, S.; Blum, H.; Thirion, C.; Krause, S.; Theis, F.J.; et al. Integrative Analysis of MicroRNA and mRNA Data Reveals an Orchestrated Function of MicroRNAs in Skeletal Myocyte Differentiation in Response to TNF-alpha or IGF1. PLoS ONE 2015, 10, e135284. [Google Scholar] [CrossRef]
- Yoshida, T.; Delafontaine, P. Mechanisms of IGF-1-Mediated Regulation of Skeletal Muscle Hypertrophy and Atrophy. Cells 2020, 9, 1970. [Google Scholar] [CrossRef] [PubMed]
- Sandri, M.; Sandri, C.; Gilbert, A.; Skurk, C.; Calabria, E.; Picard, A.; Walsh, K.; Schiaffino, S.; Lecker, S.H.; Goldberg, A.L. Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 2004, 117, 399–412. [Google Scholar] [CrossRef]
- Zhang, C.; Li, S.; Li, L.; Wang, R.; Luo, S.; Li, G. Stevioside Ameliorates Palmitic Acid-Induced Abnormal Glucose Uptake via the PDK4/AMPK/TBC1D1 Pathway in C2C12 Myotubes. Endocrinol. Diabetes Metab. 2024, 7, e482. [Google Scholar] [CrossRef]
- Sin, J.; Andres, A.M.; Taylor, D.J.; Weston, T.; Hiraumi, Y.; Stotland, A.; Kim, B.J.; Huang, C.; Doran, K.S.; Gottlieb, R.A. Mitophagy is required for mitochondrial biogenesis and myogenic differentiation of C2C12 myoblasts. Autophagy 2016, 12, 369–380. [Google Scholar] [CrossRef]
- Yue, B.; Wang, J.; Ru, W.; Wu, J.; Cao, X.; Yang, H.; Huang, Y.; Lan, X.; Lei, C.; Huang, B.; et al. The Circular RNA circHUWE1 Sponges the miR-29b-AKT3 Axis to Regulate Myoblast Development. Mol. Ther.-Nucl. Acids 2020, 19, 1086–1097. [Google Scholar] [CrossRef]
- Li, A.; Su, X.; Tian, Y.; Song, G.; Zan, L.; Wang, H. Effect of Actin Alpha Cardiac Muscle 1 on the Proliferation and Differentiation of Bovine Myoblasts and Preadipocytes. Animals 2021, 11, 3468. [Google Scholar] [CrossRef]
- Zhou, D.; Wang, Y.; Yang, R.; Wang, F.; Zhao, Z.; Wang, X.; Xie, L.; Tian, X.; Wang, G.; Li, B.; et al. The MyoD1 Promoted Muscle Differentiation and Generation by Activating CCND2 in Guanling Cattle. Animals 2022, 12, 2571. [Google Scholar] [CrossRef]
- Lu, Z.; Wang, D.; Wang, X.; Zou, J.; Sun, J.; Bi, Z. MiR-206 regulates the progression of osteoporosis via targeting HDAC4. Eur. J. Med. Res. 2021, 26, 8. [Google Scholar] [CrossRef]
- Li, Q.; Zhu, L.; Niu, F.; Li, Q.; Wang, C.; Yang, H.; Gao, C. Histone deacetylase HDAC4 participates in the pathological process of myocardial ischemia-reperfusion injury via MEKK1/JNK pathway by binding to miR-206. Cell Death Discov. 2021, 7, 240. [Google Scholar] [CrossRef]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Liu, Y.; Song, J.; Liu, Y.; Zhou, Z.; Wang, X. Transcription activation of circ-STAT3 induced by Gli2 promotes the progression of hepatoblastoma via acting as a sponge for miR-29a/b/c-3p to upregulate STAT3/Gli2. J. Exp. Clin. Cancer Res. 2020, 39, 101. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.N.; Liu, C.L.; Zeng, S.Q.; Liu, C.B.; Si, W.J.; Yuan, Y.; Ren, L.X.; He, Y.M.; Zhang, W.Y.; Zhang, H.Y.; et al. Identification of differentially expressed long non-coding RNAs and messenger RNAs involved with muscle development in Dazu black goats through RNA sequencing. Anim. Biotechnol. 2023, 34, 1305–1313. [Google Scholar] [CrossRef] [PubMed]
- von Maltzahn, J.; Jones, A.E.; Parks, R.J.; Rudnicki, M.A. Pax7 is critical for the normal function of satellite cells in adult skeletal muscle. Proc. Natl. Acad. Sci. USA 2013, 110, 16474–16479. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Tan, J.N.; Zhong, G.Y.; Zhong, L.; Hou, D.; Ma, S.; Wang, P.L.; Zhang, Z.H.; Lu, X.Q.; Yang, B.; et al. Hsa_circ_0020134 promotes liver metastasis of colorectal cancer through the miR-183-5p-PFN2-TGF-beta/Smad axis. Transl. Oncol. 2024, 39, 101823. [Google Scholar] [CrossRef] [PubMed]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, P.; Li, X.; Liu, C.; Han, Y.; E, G.; Huang, Y. Role and Regulatory Mechanism of circRNA_14820 in the Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells. Int. J. Mol. Sci. 2024, 25, 8900. https://doi.org/10.3390/ijms25168900
Yang P, Li X, Liu C, Han Y, E G, Huang Y. Role and Regulatory Mechanism of circRNA_14820 in the Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells. International Journal of Molecular Sciences. 2024; 25(16):8900. https://doi.org/10.3390/ijms25168900
Chicago/Turabian StyleYang, Pu, Xuelong Li, Chengli Liu, Yanguo Han, Guangxin E, and Yongfu Huang. 2024. "Role and Regulatory Mechanism of circRNA_14820 in the Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells" International Journal of Molecular Sciences 25, no. 16: 8900. https://doi.org/10.3390/ijms25168900
APA StyleYang, P., Li, X., Liu, C., Han, Y., E, G., & Huang, Y. (2024). Role and Regulatory Mechanism of circRNA_14820 in the Proliferation and Differentiation of Goat Skeletal Muscle Satellite Cells. International Journal of Molecular Sciences, 25(16), 8900. https://doi.org/10.3390/ijms25168900





