Genomic Insights into Pseudomonas protegens E1BL2 from Giant Jala Maize: A Novel Bioresource for Sustainable Agriculture and Efficient Management of Fungal Phytopathogens
Abstract
1. Introduction
2. Results
2.1. Genome Structure of P. protegens E1BL2
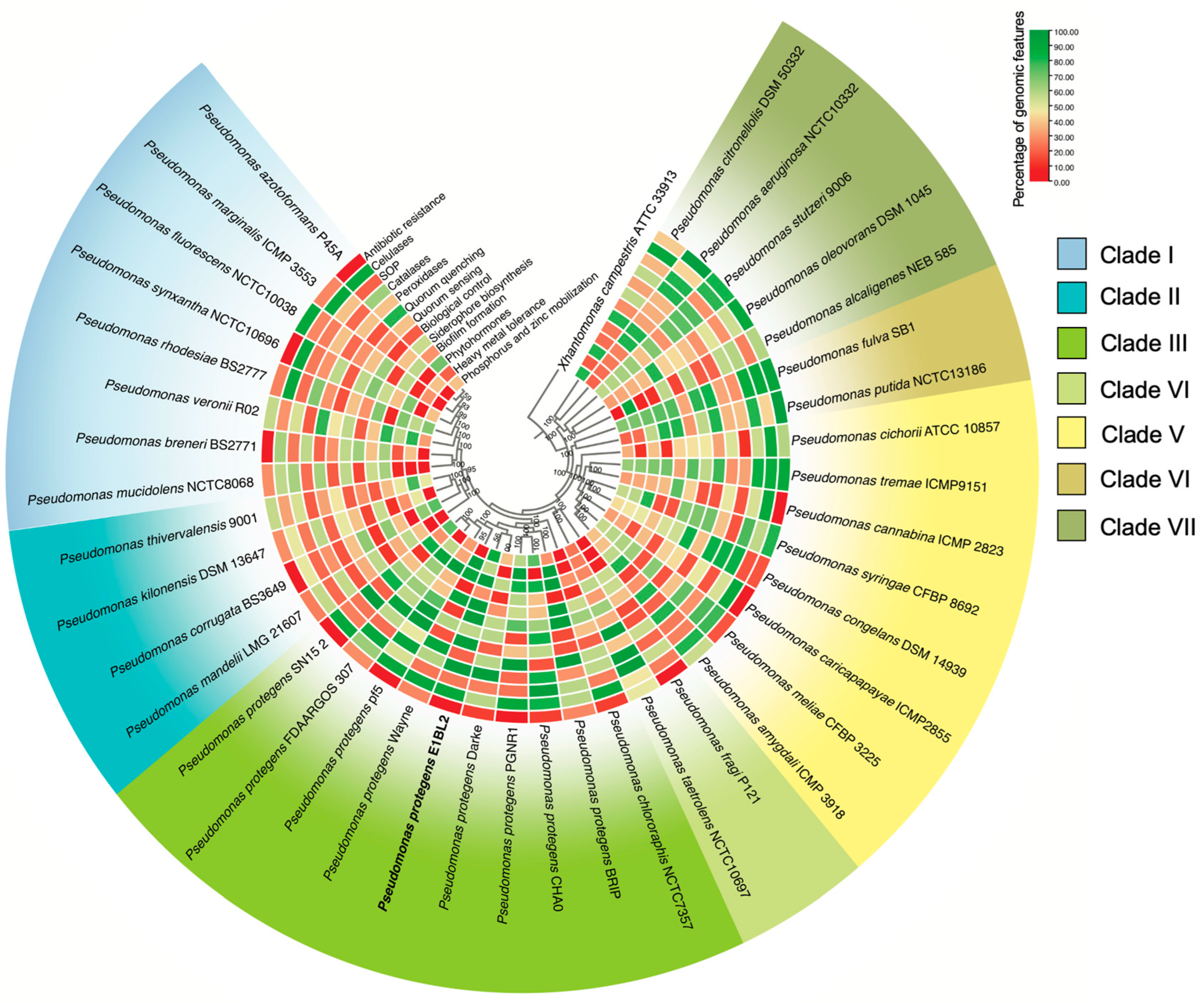
2.2. Phylogenomic and Comparative Genomics of Stain E1BL2
2.3. PGPB Characterization of P. protegens E1BL2
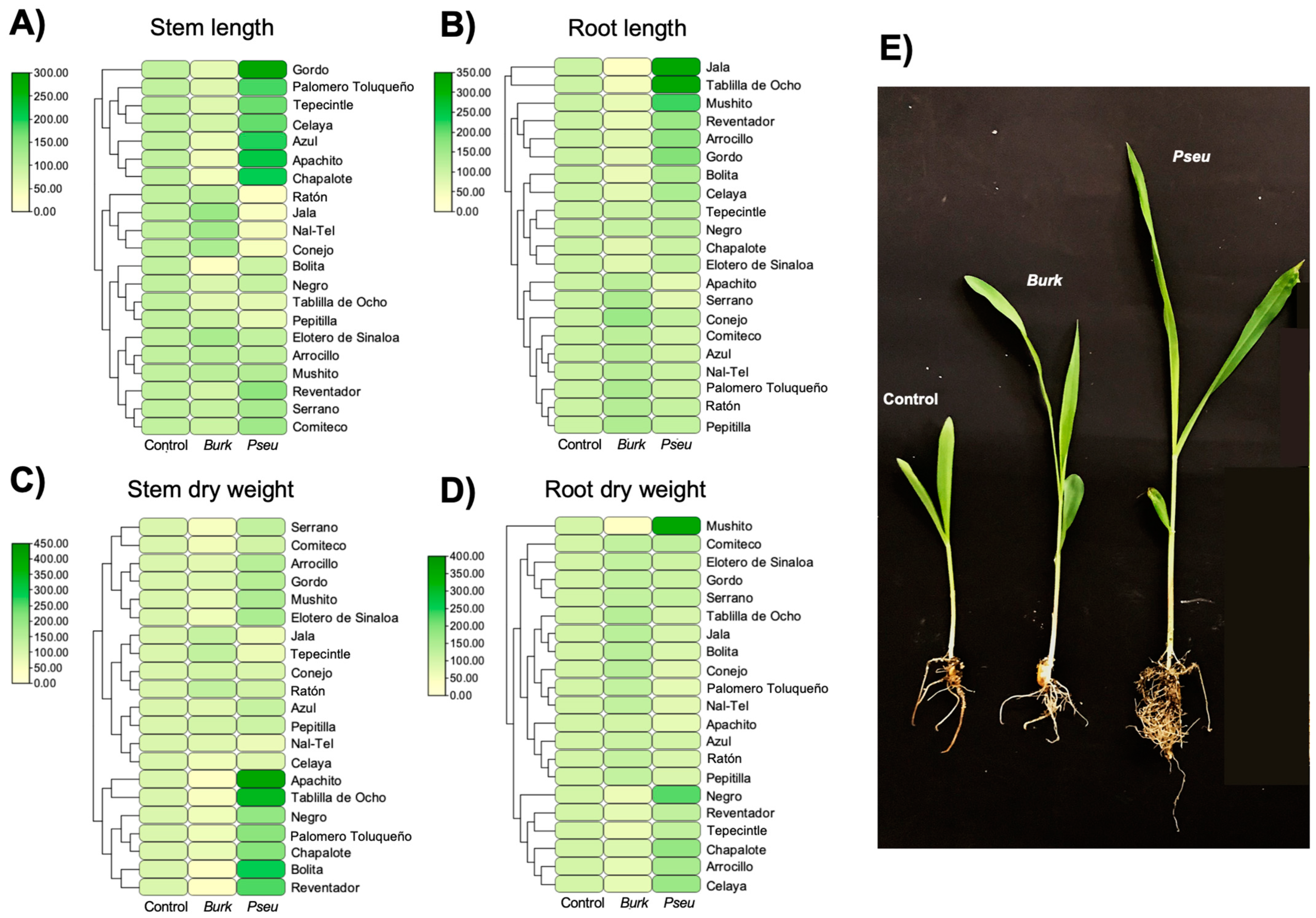
2.4. Determination of the Effect of Inoculation of Bacterial Strains in Maize Plants
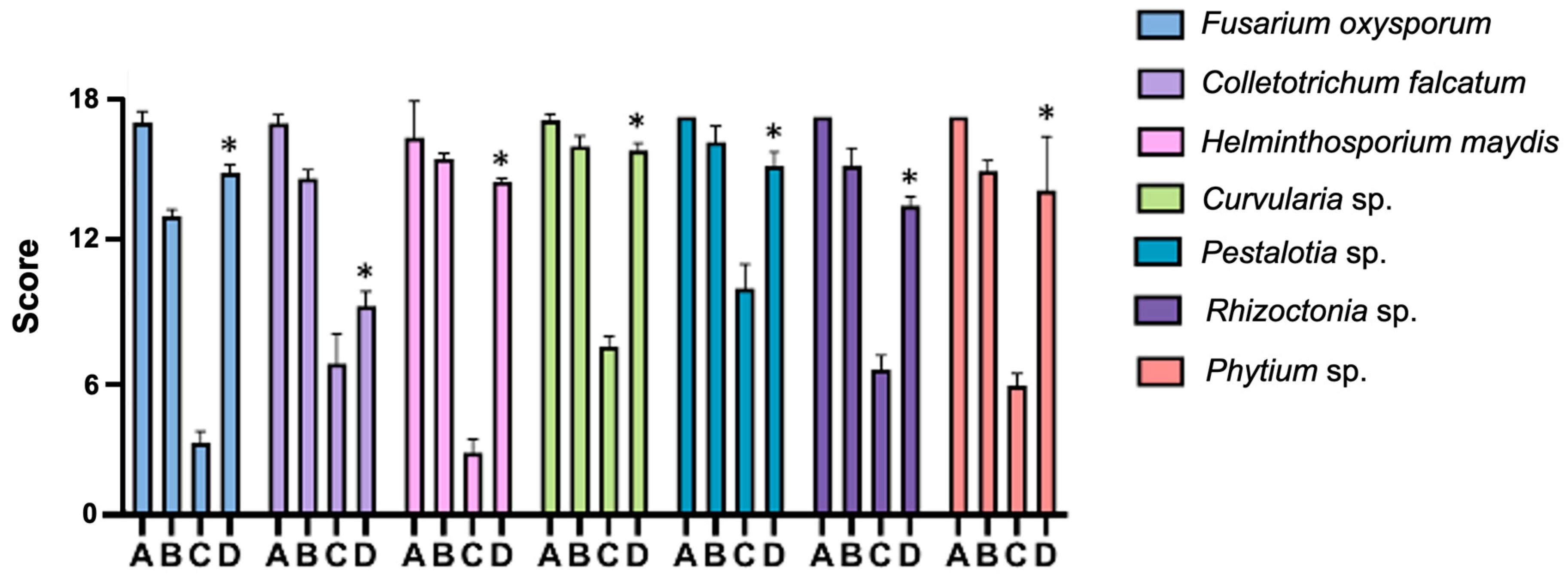
2.5. Inhibition of Fungal Growth and Germination Protection In Vitro
2.6. Maize Protection Seedling Assay in Greenhouse
2.7. Plant Field Trial
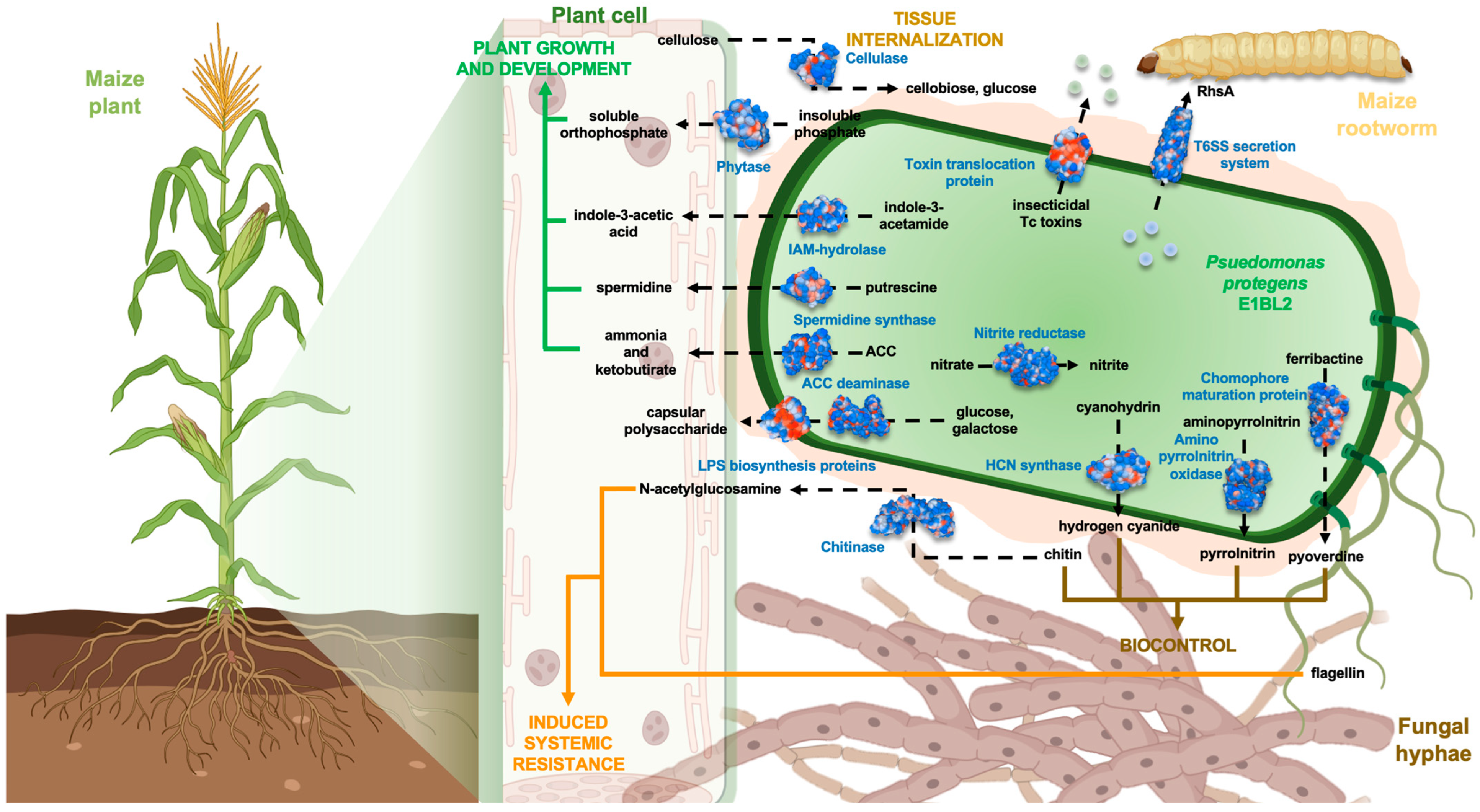
3. Discussion
4. Materials and Methods
4.1. Bacterial Strain and Growth Conditions
4.2. DNA Extraction
4.3. Whole-Genome Sequencing and Assembly and Annotation
4.4. Phylogenomic and Comparative Genome Analysis
4.5. PGPB Characterization of P. protegens E1BL2
4.6. Determination of the Effect of Inoculation of Bacterial Strains in Maize Plants
4.7. Inhibition of Fungal Growth and Germination Protection In Vitro
4.8. Maize Protection Seedling Assay in Greenhouse
4.9. Test in Planta for Maize Protection Seedling at Greenhouse Level
4.10. Plant Field Trial
4.11. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef]
- Omomowo, O.I.; Babalola, O.O. Bacterial and fungal endophytes: Tiny giants with immense beneficial potential for plant growth and sustainable agricultural productivity. Microorganisms 2019, 7, 481. [Google Scholar] [CrossRef]
- Wei, Z.; Gu, Y.; Friman, V.-P.; Kowalchuk, G.A.; Xu, Y.; Shen, Q.; Jousset, A. Initial soil microbiome composition and functioning predetermine future plant health. Sci. Adv. 2019, 5, eaaw0759. [Google Scholar] [CrossRef]
- Wu, T.; Xu, J.; Xie, W.; Yao, Z.; Yang, H.; Sun, C.; Li, X. Pseudomonas aeruginosa L10: A hydrocarbon-degrading, biosurfactant-producing, and plant-growth-promoting endophytic bacterium isolated from a reed (Phragmites australis). Front. Microbiol. 2018, 9, 1087. [Google Scholar] [CrossRef]
- Haas, D.; Défago, G. Biological control of soil-borne pathogens by fluorescent Pseudomonads. Nat. Rev. Microbiol. 2005, 3, 307–319. [Google Scholar] [CrossRef]
- Glick, B.R. Plant growth-promoting bacteria: Mechanisms and applications. Scientifica 2012, 2012, 963401. [Google Scholar] [CrossRef] [PubMed]
- Beneduzi, A.; Ambrosini, A.; Passaglia, L.M.P. Plant Growth-Promoting Rhizobacteria (PGPR): Their potential as antagonists and biocontrol agents. Genet. Mol. Biol. 2012, 35 (Suppl. S1), 1044–1051. [Google Scholar] [CrossRef] [PubMed]
- Premachandra, D.; Hudek, L.; Brau, L. Bacterial modes of action for enhancing of plant growth. J. Biotechnol. Biomater. 2016, 6, 236. [Google Scholar] [CrossRef]
- Vílchez, J.I.; Navas, A.; González-López, J.; Arcos, S.C.; Manzanera, M. Biosafety test for plant growth-promoting bacteria: Proposed environmental and human safety index (EHSI) protocol. Front. Microbiol. 2016, 6, 1514. [Google Scholar] [CrossRef]
- Rutz, D.; Frasson, D.; Sievers, M.; Blom, J.; Rezzonico, F.; Pothier, J.F.; Smits, T.H.M. Comparative genomic analysis of the biotechnological potential of the novel species Pseudomonas wadenswilerensis CCOS 864T and Pseudomonas reidholzensis CCOS 865T. Diversity 2019, 11, 204. [Google Scholar] [CrossRef]
- Preston, G.M. Plant perceptions of plant growth-promoting Pseudomonas. Phil. Trans. R. Soc. Lond. B 2004, 359, 907–918. [Google Scholar] [CrossRef] [PubMed]
- Loper, J.E.; Hassan, K.A.; Mavrodi, D.V.; Davis, E.W.; Lim, C.K.; Shaffer, B.T.; Elbourne, L.D.H.; Stockwell, V.O.; Hartney, S.L.; Breakwell, K.; et al. Comparative Genomics of Plant-Associated Pseudomonas Spp.: Insights into Diversity and Inheritance of Traits Involved in Multitrophic Interactions. PLoS Genet. 2012, 8, e1002784. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Barranquero, J.A.; Cazorla, F.M.; de Vicente, A. Pseudomonas syringae pv. syringae associated with mango trees, a particular pathogen within the “hodgepodge” of the Pseudomonas syringae Complex. Front. Plant Sci. 2019, 10, 570. [Google Scholar] [CrossRef] [PubMed]
- Li, E.; De Jonge, R.; Liu, C.; Jiang, H.; Friman, V.-P.; Pieterse, C.M.J.; Bakker, P.A.H.M.; Jousset, A. Rapid Evolution of Bacterial Mutualism in the Plant Rhizosphere. Nat. Commun. 2021, 12, 3829. [Google Scholar] [CrossRef]
- Desnoues, N.; Lin, M.; Guo, X.; Ma, L.; Carreño-Lopez, R.; Elmerich, C. Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology 2003, 149, 2251–2262. [Google Scholar] [CrossRef]
- Farhan, H.; Abdullah, B.; Hameed, A. The biological activity of bacterial vaccine of Pseudomonas putida 2 and Pseudomonas fluorescens 3 isolates to protect sesame crop (Sesamum indicum) from Fusarium fungi under field conditions. ABJNA 2010, 1, 803–811. [Google Scholar] [CrossRef]
- Mercado-Blanco, J.; Alós, E.; Rey, M.D.; Prieto, P. Pseudomonas fluorescens PICF7 displays an endophytic lifestyle in cultivated cereals and enhances yield in barley. FEMS Microbiol. Ecol. 2016, 92, fiw092. [Google Scholar] [CrossRef]
- Zhang, J.; Mavrodi, D.V.; Yang, M.; Thomashow, L.S.; Mavrodi, O.V.; Kelton, J.; Weller, D.M. Pseudomonas synxantha 2-79 transformed with pyrrolnitrin biosynthesis genes has improved biocontrol activity against soilborne pathogens of wheat and canola. Phytopathology 2020, 110, 1010–1017. [Google Scholar] [CrossRef]
- Chetverikov, S.P.; Chetverikova, D.V.; Bakaeva, M.D.; Kenjieva, A.A.; Starikov, S.N.; Sultangazin, Z.R. A promising herbicide-resistant bacterial strain of Pseudomonas protegens for stimulation of the growth of agricultural cereal grains. Appl. Biochem. Microbiol. 2021, 57, 110–116. [Google Scholar] [CrossRef]
- Hol, W.H.G.; Bezemer, T.M.; Biere, A. Getting the ecology into interactions between plants and the plant growth-promoting bacterium Pseudomonas fluorescens. Front. Plant Sci. 2013, 4, 81. [Google Scholar] [CrossRef]
- Pravisya, P.; Jayaram, K.M.; Yusuf, A. Biotic priming with Pseudomonas fluorescens induce drought stress tolerance in Abelmoschus esculentus (L.) Moench (Okra). Physiol. Mol. Biol. Plants. 2019, 25, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Çakmakçi, R.; Dönmez, F.; Aydın, A.; Şahin, F. Growth promotion of plants by plant growth-promoting rhizobacteria under greenhouse and two different field soil conditions. Soil Biol. Biochem. 2006, 38, 1482–1487. [Google Scholar] [CrossRef]
- Zhang, Q.X.; Kong, X.W.; Li, S.Y.; Chen, X.J.; Chen, X.J. Antibiotics of Pseudomonas protegens FD6 are essential for biocontrol activity. Australas. Plant Pathol. 2020, 49, 307–317. [Google Scholar] [CrossRef]
- De La Vega-Camarillo, E.; Sotelo-Aguilar, J.; Rios-Galicia, B.; Mercado-Flores, Y.; Arteaga-Garibay, R.; Villa-Tanaca, L.; Hernández-Rodríguez, C. Promotion of the Growth and Yield of Zea mays by Synthetic Microbial Communities from Jala Maize. Front. Microbiol. 2023, 14, 1167839. [Google Scholar] [CrossRef] [PubMed]
- Rios-Galicia, B.; Villagómez-Garfias, C.; De la Vega-Camarillo, E.; Guerra-Camacho, J.E.; Medina-Jaritz, N.; Arteaga-Garibay, R.I.; Villa-Tanaca, L.; Hernández-Rodríguez, C. The Mexican giant maize of Jala landrace harbour plant-growth-promoting rhizospheric and endophytic bacteria. 3 Biotech 2021, 11, 447. [Google Scholar] [CrossRef]
- Von Felten, A.; Défago, G.; Maurhofer, M. Quantification of Pseudomonas fluorescens strains F113, CHA0 and Pf153 in the rhizosphere of maize by strain-specific real-time PCR unaffected by the variability of DNA extraction efficiency. J. Microbiol. Methods 2010, 81, 108–115. [Google Scholar] [CrossRef]
- Flury, P.; Aellen, N.; Ruffner, B.; Péchy-Tarr, M.; Fataar, S.; Metla, Z.; Dominguez-Ferreras, A.; Bloemberg, G.; Frey, J.; Goesmann, A.; et al. Insect pathogenicity in plant-beneficial Pseudomonads: Phylogenetic distribution and comparative genomics. ISME J. 2016, 10, 2527–2542. [Google Scholar] [CrossRef]
- Garrido-Sanz, D.; Meier-Kolthoff, J.P.; Göker, M.; Martín, M.; Rivilla, R.; Redondo-Nieto, M. Genomic and genetic diversity within the Pseudomonas fluorescens complex. PLoS ONE 2016, 11, e0150183. [Google Scholar] [CrossRef] [PubMed]
- Peix, A.; Ramírez-Bahena, M.-H.; Velázquez, E. The current status on the taxonomy of Pseudomonas Revisited: An Update. Infec. Gen. Evol. 2018, 57, 106–116. [Google Scholar] [CrossRef]
- Maroniche, G.A.; Rubio, E.J.; Consiglio, A.; Perticari, A. Plant-associated fluorescent Pseudomonas from red lateritic soil: Beneficial characteristics and their impact on lettuce growth. J. Gen. Appl. Microbiol. 2016, 62, 248–257. [Google Scholar] [CrossRef]
- Zhao, H.; Liu, L.; Yang, L.; Gu, Q.; Li, Y.; Zhang, J.; Wu, S.; Chen, M.; Xie, X.; Wu, Q. Pseudomonas protegens FJKB0103 isolated from rhizosphere exhibits anti-methicillin-resistant Staphylococcus aureus activity. Microorganisms 2022, 10, 315. [Google Scholar] [CrossRef] [PubMed]
- Ferreiro, M.; Gallegos, M. Distinctive features of the Gac-Rsm pathway in plant-associated Pseudomonas. Environ. Microbiol. 2021, 23, 5670–5689. [Google Scholar] [CrossRef] [PubMed]
- Dimkić, I.; Janakiev, T.; Petrović, M.; Degrassi, G.; Fira, D. Plant-associated Bacillus and Pseudomonas antimicrobial activities in plant disease suppression via biological control mechanisms—A review. Physiol. Mol. Plant Pathol. 2022, 117, 101754. [Google Scholar] [CrossRef]
- Guzmán-Guzmán, P.; Santoyo, G. Action mechanisms, biodiversity, and omics approaches in biocontrol and plant growth-promoting Pseudomonas: An updated review. Biocontrol Sci. Technol. 2022, 32, 527–550. [Google Scholar] [CrossRef]
- Yi, H.-S.; Ahn, Y.-R.; Song, G.C.; Ghim, S.-Y.; Lee, S.; Lee, G.; Ryu, C.-M. Impact of a bacterial volatile 2,3-butanediol on Bacillus subtilis rhizosphere robustness. Front. Microbiol. 2016, 7, 993. [Google Scholar] [CrossRef]
- Park, J.Y.; Kang, B.R.; Ryu, C.-M.; Anderson, A.J.; Kim, Y.C. Polyamine is a critical determinant of Pseudomonas chlororaphis O6 for GacS-Dependent bacterial cell growth and biocontrol capacity: Role of polyamines in a biocontrol bacterium. Mol. Plant Pathol. 2018, 19, 1257–1266. [Google Scholar] [CrossRef]
- Burlinson, P.; Studholme, D.; Cambray-Young, J.; Heavens, D.; Rathjen, J.; Hodgkin, J.; Preston, G.M. Pseudomonas fluorescens NZI7 repels grazing by C. elegans, a natural predator. ISME J. 2013, 7, 1126–1138. [Google Scholar] [CrossRef]
- Ruiz, J.A.; Bernar, E.M.; Jung, K. Production of siderophores increases resistance to fusaric acid in Pseudomonas protegens Pf-5. PLoS ONE 2015, 10, e0117040. [Google Scholar] [CrossRef]
- Drehe, I.; Simonetti, E.; Ruiz, J.A. Contribution of the siderophores pyoverdine and enantio-pyochelin to fitness in soil of Pseudomonas protegens Pf-5. Curr. Microbiol. 2018, 75, 1560–1565. [Google Scholar] [CrossRef]
- Sexton, D.J.; Glover, R.C.; Loper, J.E.; Schuster, M. Pseudomonas protegens Pf-5 favours self-produced siderophore over free-loading in interspecies competition for iron. Environ. Microbiol. 2017, 19, 3514–3525. [Google Scholar] [CrossRef]
- Loper, J.E.; Henkels, M.D.; Rangel, L.I.; Olcott, M.H.; Walker, F.L.; Bond, K.L.; Kidarsa, T.A.; Hesse, C.N.; Sneh, B.; Stockwell, V.O.; et al. Rhizoxin analogs, orfamide A and chitinase production contribute to the toxicity of Pseudomonas protegens Strain Pf-5 to Drosophila melanogaster: Insect toxicity of Pseudomonas protegens Pf-5. Environ. Microbiol. 2016, 18, 3509–3521. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, K.; Ogiso, M.; Morohosi, T.; Seo, S. Glutamate positively regulates chitinase activity and the biocontrol efficacy of Pseudomonas protegens. Mol. Plant-Microbe Interact. 2023, 36, 323–333. [Google Scholar] [CrossRef]
- Saini, S.; Verma, A.; Kumar, A.; Prakash, A.; Sharma, S.K.; Ramesh, A.; Johri, B. Identification and characterization of antifungal metabolite producing Pseudomonas protegens strain BNJ-SS-45 isolated from rhizosphere of wheat crop (Triticum aestivum L.). Int. J. Appl. Pure Sci. Agric. 2016, 2, 69–76. [Google Scholar]
- Bellameche, F.; Jasim, M.A.; Mauch-Mani, B.; Mascher, F. Histopathological aspects of resistance in wheat to Puccinia triticina, induced by Pseudomonas protegens CHA0 and β-aminobutyric acid. Phytopathol. Mediterr. 2021, 60, 441–453. [Google Scholar] [CrossRef]
- Ashrafi, J.; Rahnama, K.; Babaeizad, V.; Ramezanpour, S.S.; Keel, C. Induction of wheat resistance to STB by the endophytic fungus Serendipita indica and Pseudomonas protegens. Iran. J. Biotech. 2021, 19, e2762. [Google Scholar] [CrossRef]
- Michavila, G.; Adler, C.; De Gregorio, P.R.; Lami, M.J.; Caram Di Santo, M.C.; Zenoff, A.M.; de Cristobal, R.E.; Vincent, P.A. Pseudomonas protegens CS1 from the lemon phyllosphere as a candidate for citrus canker biocontrol agent. Plant. Biol. J. 2017, 19, 608–617. [Google Scholar] [CrossRef] [PubMed]
- Pellicciaro, M.; Lione, G.; Giordano, L.; Gonthier, P. Biocontrol potential of Pseudomonas protegens against Heterobasidion species attacking conifers in Europe. Biol. Control 2021, 157, 104583. [Google Scholar] [CrossRef]
- Andreolli, M.; Zapparoli, G.; Angelini, E.; Lucchetta, G.; Lampis, S.; Vallini, G. Pseudomonas protegens MP12: A plant growth-promoting endophytic bacterium with broad-spectrum antifungal activity against grapevine phytopathogens. Microbiol. Res. 2019, 219, 123–131. [Google Scholar] [CrossRef]
- Meyer, G.; Bünemann, E.K.; Frossard, E.; Maurhofer, M.; Mäder, P.; Oberson, A. Gross phosphorus fluxes in a calcareous soil inoculated with Pseudomonas protegens CHA0 revealed by 33P isotopic dilution. Soil Biol. Biochem. 2017, 104, 81–94. [Google Scholar] [CrossRef]
- Singh, J.; Singh, A.V.; Upadhayay, V.K.; Khan, A.; Chandra, R. Prolific contribution of Pseudomonas protegens in Zn biofortification of wheat by modulating multifaceted physiological response under saline and non-saline conditions. World J. Microbiol. Biotechnol. 2022, 38, 227. [Google Scholar] [CrossRef]
- Bakaeva, M.; Chetverikov, S.; Timergalin, M.; Feoktistova, A.; Rameev, T.; Chetverikova, D.; Kenjieva, A.; Starikov, S.; Sharipov, D.; Hkudaygulov, G. PGP-Bacterium Pseudomonas protegens improves bread wheat growth and mitigates herbicide and drought stress. Plants 2022, 11, 3289. [Google Scholar] [CrossRef] [PubMed]
- Urias-Lugo, D.A.; Heredia, J.B.; Serna-Saldivar, S.O.; Muy-Rangel, M.D.; Valdez-Torres, J.B. Total phenolics, total anthocyanins and antioxidant capacity of native and elite blue maize hybrids (Zea mays L.). CyTA J. Food 2015, 13, 336–339. [Google Scholar] [CrossRef]
- Lacombe, A.; Wu, V.C.H.; Tyler, S.; Edwards, K. Antimicrobial action of the American cranberry constituents; phenolics, anthocyanins, and organic acids, against Escherichia coli O157:H7. Int. J. Food Microbiol. 2010, 139, 102–107. [Google Scholar] [CrossRef]
- Nalbur, B.E.; Eleren, S.Ç.; Şahin, S.; Alkan, U. Toxic effects of copper-based and synthetic organic pesticides on activated sludge. CLEAN Soil Air Water 2012, 40, 39–44. [Google Scholar] [CrossRef]
- Powers, M.J.; Sanabria-Valentín, E.; Bowers, A.A.; Shank, E.A. Inhibition of cell differentiation in Bacillus subtilis by Pseudomonas protegens. J. Bacteriol. 2015, 197, 2129–2138. [Google Scholar] [CrossRef]
- Quecine, M.C.; Kidarsa, T.A.; Goebel, N.C.; Shaffer, B.T.; Henkels, M.D.; Zabriskie, T.M.; Loper, J.E. An interspecies signaling system mediated by fusaric acid has parallel effects on antifungal metabolite production by Pseudomonas protegens strain Pf-5 and antibiosis of Fusarium spp. Appl. Environ. Microbiol. 2016, 82, 1372–1382. [Google Scholar] [CrossRef]
- Ramette, A.; Frapolli, M.; Saux, M.F.-L.; Gruffaz, C.; Meyer, J.-M.; Défago, G.; Sutra, L.; Moënne-Loccoz, Y. Pseudomonas protegens sp. nov., widespread plant-protecting bacteria producing the biocontrol compounds 2,4-diacetylphloroglucinol and pyoluteorin. System. Appl. Microbiol. 2011, 34, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Montelongo-Martínez, L.F.; Hernández-Méndez, C.; Muriel-Millan, L.F.; Hernández-Estrada, R.; Fabian-Del Olmo, M.J.; González-Valdez, A.; Soberón-Chávez, G.; Cocotl-Yañez, M. Unraveling the regulation of pyocyanin synthesis by RsmA through MvaU and RpoS in Pseudomonas aeruginosa ID4365. J. Basic Microbiol. 2023, 63, 51–63. [Google Scholar] [CrossRef]
- Ma, Z.; Geudens, N.; Kieu, N.P.; Sinnaeve, D.; Ongena, M.; Martins, J.C.; Höfte, M. Biosynthesis, chemical structure, and structure-activity relationship of orfamide lipopeptides produced by Pseudomonas protegens and related species. Front. Microbiol. 2016, 7, 382. [Google Scholar] [CrossRef]
- Rose, M.M.; Scheer, D.; Hou, Y.; Hotter, V.S.; Komor, A.J.; Aiyar, P.; Scherlach, K.; Vergara, F.; Yan, Q.; Loper, J.E.; et al. The bacterium Pseudomonas protegens antagonizes the microalga Chlamydomonas reinhardtii using a blend of toxins. Environ. Microbiol. 2021, 23, 5525–5540. [Google Scholar] [CrossRef]
- Shi, H.; Huang, X.; Wang, Z.; Guan, Y.; Zhang, X. Improvement of pyoluteorin production in Pseudomonas protegens H78 through engineering its biosynthetic and regulatory pathways. Appl. Microbiol. Biotechnol. 2019, 103, 3465–3476. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, R.L.; van Verk, M.C.; Stringlis, I.A.; Zamioudis, C.; Tommassen, J.; Pieterse, C.M.J.; Bakker, P.A.H.M. Unearthing the genomes of plant-beneficial Pseudomonas model strains WCS358, WCS374 and WCS417. BMC Genom. 2015, 16, 539. [Google Scholar] [CrossRef]
- Jousset, A.; Lara, E.; Wall, L.G.; Valverde, C. Secondary metabolites help biocontrol strain Pseudomonas fluorescens CHA0 to escape protozoan grazing. Appl. Environ. Microbiol. 2006, 72, 7083–7090. [Google Scholar] [CrossRef] [PubMed]
- Yan, Q.; Philmus, B.; Chang, J.H.; Loper, J.E. Novel mechanism of metabolic co-regulation coordinates the biosynthesis of secondary metabolites in Pseudomonas protegens. eLife 2017, 6, e22835. [Google Scholar] [CrossRef] [PubMed]
- Flury, P.; Vesga, P.; Dominguez-Ferreras, A.; Tinguely, C.; Ullrich, C.I.; Kleespies, R.G.; Keel, C.; Maurhofer, M. Persistence of root-colonizing Pseudomonas protegens in herbivorous insects throughout different developmental stages and dispersal to new host plants. ISME J. 2019, 13, 860–872. [Google Scholar] [CrossRef]
- Vesga, P.; Flury, P.; Vacheron, J.; Keel, C.; Croll, D.; Maurhofer, M. Transcriptome plasticity underlying plant root colonization and insect invasion by Pseudomonas protegens. ISME J. 2020, 14, 2766–2782. [Google Scholar] [CrossRef] [PubMed]
- Fox, A.R.; Soto, G.; Valverde, C.; Russo, D.; Lagares, A.; Zorreguieta, Á.; Alleva, K.; Pascuan, C.; Frare, R.; Mercado-Blanco, J.; et al. Major cereal crops benefit from biological nitrogen fixation when inoculated with the nitrogen-fixing bacterium Pseudomonas protegens Pf-5 X940: Robust biological nitrogen fixation in major cereal crops. Environ. Microbiol. 2016, 18, 3522–3534. [Google Scholar] [CrossRef]
- Cesa-Luna, C.; Baez, A.; Aguayo-Acosta, A.; Llano-Villarreal, R.C.; Juárez-González, V.R.; Gaytán, P.; Bustillos-Cristales, M.d.R.; Rivera-Urbalejo, A.; Muñoz-Rojas, J.; Quintero-Hernández, V. Growth inhibition of pathogenic microorganisms by Pseudomonas protegens EMM-1 and partial characterization of inhibitory substances. PLoS ONE 2020, 15, e0240545. [Google Scholar] [CrossRef]
- Fukami, J.; Nogueira, M.A.; Araujo, R.S.; Hungria, M. Accessing inoculation methods of maize and wheat with Azospirillum brasilense. AMB Express 2016, 6, 3. [Google Scholar] [CrossRef]
- Kannojia, P.; Choudhary, K.K.; Srivastava, A.K.; Singh, A.K. PGPR Bioelicitors. In PGPR Amelioration in Sustainable Agriculture; Elsevier: Amsterdam, The Netherlands, 2019; pp. 67–84. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Zamioudis, C.; Berendsen, R.L.; Weller, D.M.; Van Wees, S.C.M.; Bakker, P.A.H.M. Induced systemic resistance by beneficial microbes. Annu. Rev. Phytopathol. 2014, 52, 347–375. [Google Scholar] [CrossRef]
- Pedrosa, F.O.; Oliveira, A.L.M.; Guimarães, V.F.; Etto, R.M.; Souza, E.M.; Furmam, F.G.; Gonçalves, D.R.P.; Santos, O.J.A.P.; Gonçalves, L.S.A.; Battistus, A.G.; et al. The ammonium excreting Azospirillum brasilense strain HM053: A new alternative inoculant for maize. Plant Soil 2020, 451, 45–56. [Google Scholar] [CrossRef]
- Mercado-Flores, Y.; Cárdenas-Álvarez, I.O.; Rojas-Olvera, A.V.; Pérez-Camarillo, J.P.; Leyva-Mir, S.G.; Anducho-Reyes, M.A. Application of Bacillus subtilis in the biological control of the phytopathogenic fungus Sporisorium reilianum. Biol. Control 2014, 76, 36–40. [Google Scholar] [CrossRef]
- Benz, B.F. Archaeological evidence of teosinte domestication from Guilá Naquitz, Oaxaca. Proc. Natl. Acad. Sci. USA 2001, 98, 2104–2106. [Google Scholar] [CrossRef] [PubMed]
- Hufford, M.B.; Martínez-Meyer, E.; Gaut, B.S.; Eguiarte, L.E.; Tenaillon, M.I. Inferences from the historical distribution of wild and domesticated maize provide ecological and evolutionary insight. PLoS ONE 2012, 7, e47659. [Google Scholar] [CrossRef]
- Higdon, S.M.; Pozzo, T.; Kong, N.; Huang, B.C.; Yang, M.L.; Jeannotte, R.; Brown, C.T.; Bennett, A.B.; Weimer, B.C. Genomic characterization of a diazotrophic microbiota associated with maize aerial root mucilage. PLoS ONE 2020, 15, e0239677. [Google Scholar] [CrossRef]
- Chavéz-Díaz, I.F.; Cruz-Cárdenas, C.I.; Sandoval-Cancino, G.; Calvillo-Aguilar, F.F.; Ruíz-Ramírez, S.; Blanco-Camarillo, M.; Rojas-Anaya, E.; Ramírez-Vega, H.; Arteaga-Garibay, R.I.; Zelaya-Molina, L.X. Seedling growth promotion and potential biocontrol against phytopathogenic Fusarium by native rhizospheric Pseudomonas spp. strains from Amarillo Zamorano maize landrace. Rhizosphere 2022, 24, 100601. [Google Scholar] [CrossRef]
- Fontes-Puebla, A.A.; Bernal, J.S. Resistance and tolerance to root herbivory in maize were mediated by domestication, spread, and breeding. Front. Plant Sci. 2020, 11, 223. [Google Scholar] [CrossRef]
- Lima, A.F.; Bernal, J.; Venâncio, M.G.S.; De Souza, B.H.S.; Carvalho, G.A. Comparative tolerance levels of maize landraces and a hybrid to natural infestation of fall armyworm. Insects 2022, 13, 651. [Google Scholar] [CrossRef]
- William, S.; Feil, H.; Copeland, A. Bacterial genomic DNA isolation using CTAB. Sigma 2012, 50, 6876. [Google Scholar]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 1 May 2022).
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef]
- Blin, K.; Shaw, S.; Steinke, K.; Villebro, R.; Ziemert, N.; Lee, S.Y.; Medema, M.H.; Weber, T. AntiSMASH 5.0: Updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 2019, 47, W81–W87. [Google Scholar] [CrossRef]
- Skinnider, M.A.; Dejong, C.A.; Rees, P.N.; Johnston, C.W.; Li, H.; Webster, A.L.H.; Wyatt, M.A.; Magarvey, N.A. Genomes to natural products PRediction Informatics for Secondary Metabolomes (PRISM). Nucleic Acids Res. 2015, 43, gkv1012. [Google Scholar] [CrossRef]
- Avram, O.; Rapoport, D.; Portugez, S.; Pupko, T. M1CR0B1AL1Z3R—A User-friendly web server for the analysis of large-scale microbial genomics data. Access Microbiol. 2020, 2, 190. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Afzal, I.; Iqrar, I.; Shinwari, Z.K.; Yasmin, A. Plant growth-promoting potential of endophytic bacteria isolated from roots of wild Dodonaea viscosa L. Plant Growth Regul. 2017, 81, 399–408. [Google Scholar] [CrossRef]
- Medina-Córdova, N.; López-Aguilar, R.; Ascencio, F.; Castellanos, T.; Campa-Córdova, A.I.; Angulo, C. Biocontrol activity of the marine yeast Debaryomyces hansenii against phytopathogenic fungi and its ability to inhibit mycotoxins production in maize grain (Zea mays L.). Biol. Control 2016, 97, 70–79. [Google Scholar] [CrossRef]
- Borah, S.N.; Goswami, D.; Sarma, H.K.; Cameotra, S.S.; Deka, S. Rhamnolipid biosurfactant against Fusarium verticillioides to control stalk and ear rot disease of maize. Front. Microbiol. 2016, 7, 1505. [Google Scholar] [CrossRef]
- Díaz-Nájera, J.F.; Alvarado-Gómez, O.G.; Leyva-Mir, S.G.; Ayvar-Serna, S.; Michel-Aceves, A.C.; Vargas-Hernández, M. Identification and control of fungi causing fruits rot in pipiana pumpkin (Cucurbita argyrosperma Huber). Afr. J. Agric. Res. 2015, 10, 1150–1157. [Google Scholar]
- Cumagun, C.J.R.; Ramos, J.S.; Dimaano, A.O.; Munaut, F.; Van Hove, F. Genetic characteristics of Fusarium verticillioides from corn in the Philippines. J. Gen. Plant Pathol. 2009, 75, 405–412. [Google Scholar] [CrossRef]
- Lazarotto, M.; Muniz, M.F.B.; Poletto, T.; Dutra, C.B.; Blume, E.; Harakawa, R.; Poletto, I. First report of Pestalotiopsis Clavispora causing leaf spot of Carya illinoensis in Brazil. Plant Dis. 2012, 96, 1826. [Google Scholar] [CrossRef] [PubMed]
- Peterson, R.F.; Campbell, A.B.; Hannah, A.E. A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can. J. Res. 1948, 26, 496–500. [Google Scholar] [CrossRef]



| Gen | Protein | Genomic Location |
|---|---|---|
| bdhA | 2,3-butanediol dehydrogenase, R-alcohol forming, (R)- and (S)-acetoin-specific | 173,948–175,051 |
| acoR | Transcriptional activator of acetoin dehydrogenase | 167,454–169,322 |
| trpE | Anthranilate synthase, amidotransferase component | 553,679–554,272 |
| speE | Spermidine synthase | 163,961–166,468 |
| potA | Spermidine/putrescine import ABC transporter ATP-binding protein | 114,777–115,868 |
| ptaA | ABC transporter in pyoverdin gene cluster, periplasmic component | 32,978–33,961 |
| pchC | Pyochelin biosynthetic protein | 903,310–904,089 |
| cel3 | Cellulase | 191,379–192,467 |
| chi14 | Chitinase | 34,754–36,220 |
| hscA | Chaperon protein | 498,330–500,192 |
| htpG | Chaperon protein | 189,071–190,975 |
| nirD | Nitrite reductase | 2–253 |
| nit1 | Nitrilase | 870,762–871,667 |
| hmpA | Nitric oxide dioxygenase | 566,116–567,297 |
| flaA | Flagellin protein | 97,566–98,414 |
| flaG | Flagellar protein | 98,488–98,862 |
| rffA | Lipopolysaccharide biosynthesis protein | 94,647–95,207 |
| rfaP | Lipopolysaccharide core heptose(I) kinase | 33,553–34,359 |
| phlD | Phloroglucinol synthase | 17,378–19,240 |
| prtD | Pyocin formation protein | 538,488–539,531 |
| prnD | Aminopyrrolnitrin oxidase | 96,691–97,782 |
| hcnA | Hydrogen cyanide synthase | 265,107–265,424 |
| vgrG | Translocation effector protein | 130,901–132,085 |
| higB | Toxin HigB | 85,815–86,093 |
| aprE | Type I secretion membrane fusion protein | 595,315–596,649 |
| pslA | Pellicle/biofilm biosynthesis protein | 78,948–80,381 |
| ppkA | T6SS Serine/threonine protein kinase | 54,120–57,203 |
| pgaA | Biofilm PGA outer membrane secretin | 4930–7416 |
| pgaB | Biofilm PGA synthesis deacetylase | 2918–4915 |
| pgaC | Biofilm PGA synthesis N-glycosyltransferase | 1565–2914 |
| PGPB Traits | Phosphate Solubilization | ACCd | IAA | Metallophores | Chitinase | Protease | Cellulase | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Fe+3 | Mo+6 | Zn+2 | V+5 | ||||||||||||
| SQ | Qn | Ql | Qn | SQ | Qn | SQ | Qn | SQ | Qn | SQ | Qn | SQ | SQ | SQ | |
| (Index) | (µg/mL) | Nu | (µg/mL) | (Index) | (%) | (Index) | (%) | (Index) | (%) | (Index) | (%) | (Index) | (Index) | (Index) | |
| P. protegens E1BL2 | 1.1 ± 0.3 | 41.8 ± 1.6 | + | 4.5 ± 0.7 * | 2.0 ± 0.8 | 56 ± 3.6 | 3.0 ± 0.7 | 32 ± 3.5 | 2.1 ± 0.2 | 43 ± 4.2 | 4.4 ± 0.5 * | 60 ± 3.9 | 2.7 ± 0.9 | 1.8 ± 0.3 | 1.6 ± 0.6 * |
| B. metallica R3J3HD10 | 2.0 ± 0.2 | 81.8 ± 3.6 | - | 2.7 ± 0.9 | 3.3 ± 0.5 | 62.4 ± 4.2 | 3.4 ± 0.4 | 40.2 ± 2.1 | 3.1 ± 0.8 | 46.7 ± 3.5 | 2.0 ± 0.6 | 20.3 ± 3.4 | 2.1 ± 0.1 | 2.6 ± 0.2 | 0.6 ± 0.1 |
| Fusarium oxysporum | Colletotrichum falcatum | Helminthosporium maydis | Curvularia sp. | Pestalotia sp. | Rhizoctonia sp. | Pythium sp. | |
|---|---|---|---|---|---|---|---|
| Cupravit® (copper oxychloride) | 7 | 58.5 ± 2.3 | 77.2 ± 1.8 | 69.1 ± 0.7 | 72.7 ± 3.2 | 54.8 ± 2.7 | 69.4 ± 1.3 |
| Pseudomonas protegens E1BL2 | 63.8 ± 0.4 * | 41.0 ± 2.4 * | 90.3 ± 2.1 * | 40.2 ± 5.0 * | 73.0 ± 4.1 | 59.7 ± 6.5 | 72.7 ± 5.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De la Vega-Camarillo, E.; Sotelo-Aguilar, J.; González-Silva, A.; Hernández-García, J.A.; Mercado-Flores, Y.; Villa-Tanaca, L.; Hernández-Rodríguez, C. Genomic Insights into Pseudomonas protegens E1BL2 from Giant Jala Maize: A Novel Bioresource for Sustainable Agriculture and Efficient Management of Fungal Phytopathogens. Int. J. Mol. Sci. 2024, 25, 9508. https://doi.org/10.3390/ijms25179508
De la Vega-Camarillo E, Sotelo-Aguilar J, González-Silva A, Hernández-García JA, Mercado-Flores Y, Villa-Tanaca L, Hernández-Rodríguez C. Genomic Insights into Pseudomonas protegens E1BL2 from Giant Jala Maize: A Novel Bioresource for Sustainable Agriculture and Efficient Management of Fungal Phytopathogens. International Journal of Molecular Sciences. 2024; 25(17):9508. https://doi.org/10.3390/ijms25179508
Chicago/Turabian StyleDe la Vega-Camarillo, Esaú, Josimar Sotelo-Aguilar, Adilene González-Silva, Juan Alfredo Hernández-García, Yuridia Mercado-Flores, Lourdes Villa-Tanaca, and César Hernández-Rodríguez. 2024. "Genomic Insights into Pseudomonas protegens E1BL2 from Giant Jala Maize: A Novel Bioresource for Sustainable Agriculture and Efficient Management of Fungal Phytopathogens" International Journal of Molecular Sciences 25, no. 17: 9508. https://doi.org/10.3390/ijms25179508
APA StyleDe la Vega-Camarillo, E., Sotelo-Aguilar, J., González-Silva, A., Hernández-García, J. A., Mercado-Flores, Y., Villa-Tanaca, L., & Hernández-Rodríguez, C. (2024). Genomic Insights into Pseudomonas protegens E1BL2 from Giant Jala Maize: A Novel Bioresource for Sustainable Agriculture and Efficient Management of Fungal Phytopathogens. International Journal of Molecular Sciences, 25(17), 9508. https://doi.org/10.3390/ijms25179508







