Diarrheagenic Escherichia coli in Stool Specimens Collected from Patients Attending Primary Healthcare Facilities in Ethiopia: Whole-Genome Sequencing-Based Molecular Characterization
Abstract
:1. Introduction
2. Results
2.1. Whole-Genome Sequencing of DEC Isolates
2.2. Serotypes, Phylogenetic Groups, and Multilocus Sequence Typing of DEC Isolates
2.3. Virulence-Associated Genes of DEC Isolates
2.4. Antimicrobial Resistance Genes in DEC Isolates
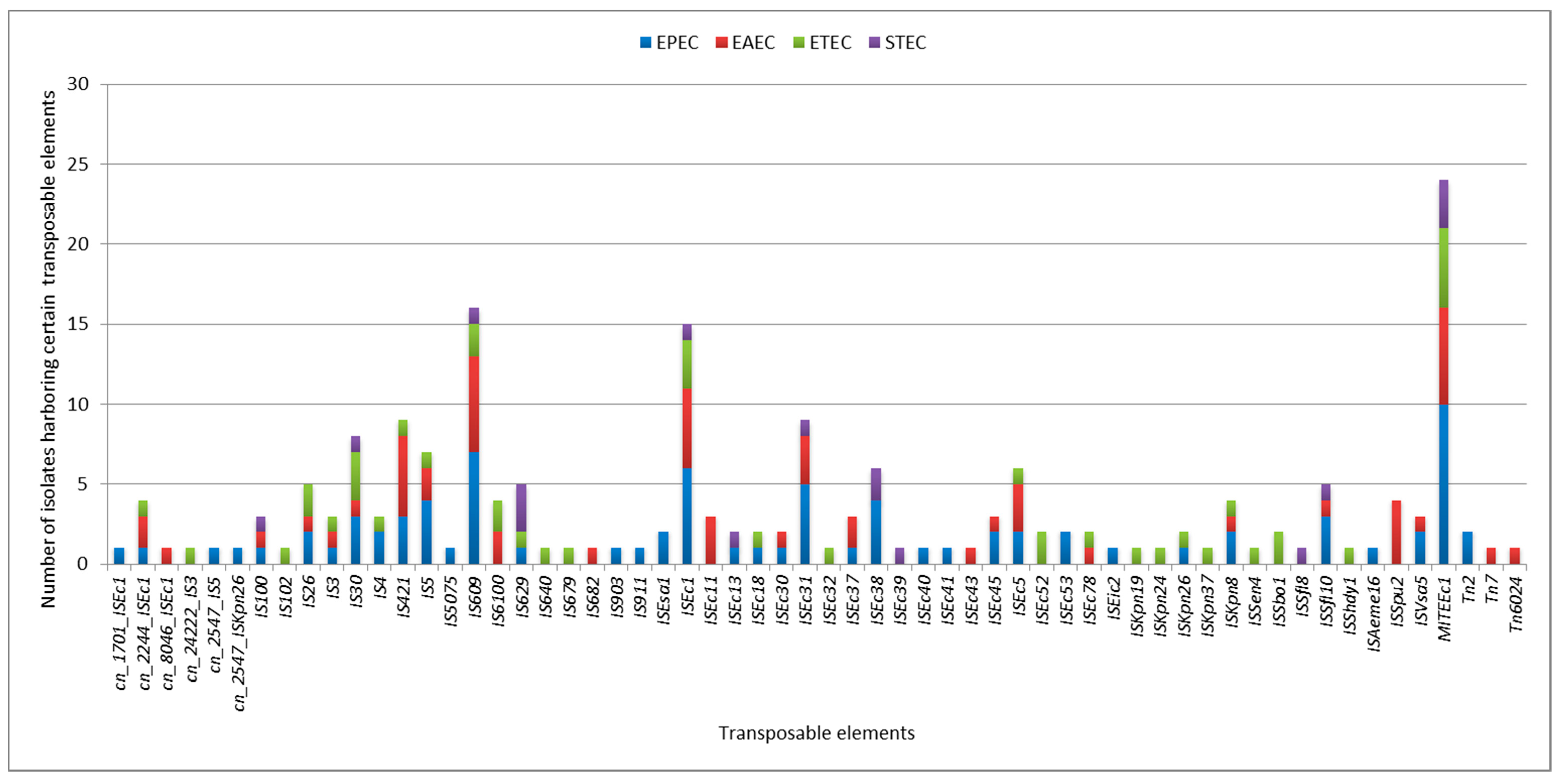
2.5. Plasmids and Other Mobile Genetic Elements
3. Discussion
4. Materials and Methods
4.1. Bacterial Isolates
4.2. DNA Extraction and PCR Detection of E. coli Pathotypes Genes
4.3. Whole-Genome Sequencing, Raw Data Pre-Processing, De Novo Assembly and Quality Control
4.4. Phylogenetic Groups, In Silico Multilocus Sequence Typing, and In Silico Serotyping
4.5. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Basavaraju, M.; Gunashree, B.S. Escherichia coli: An Overview of Main Characteristics. In Escherichia coli—Old and New Insights; Erjavec, M.S., Ed.; IntechOpen: London, UK, 2022; pp. 1–16. [Google Scholar]
- Galindo-Méndez, M. Antimicrobial Resistance in Escherichia coli. In E. coli Infections—Importance of Early Diagnosis and Efficient Treatment; IntechOpen: London, UK, 2020; pp. 1–13. [Google Scholar]
- World Health Organization. Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report 2022; World Health Organization: Geneva, Switzerland, 2022; Available online: https://www.who.int/publications/i/item/9789240062702 (accessed on 15 July 2024).
- Poirel, L.; Madec, J.-Y.; Lupo, A.; Schink, A.-K.; Skieffer, N.; Nordmann, P.; Schwarz, S. Antimicrobial Resistance in Escherichia coli. Microbiol. Spectr. 2018, 6, ARBA-0026-2017. [Google Scholar] [CrossRef] [PubMed]
- Asmare, Z.; Erkihun, M.; Abebe, W.; Tamrat, E. Antimicrobial resistance and ESBL production in uropathogenic Escherichia coli: A systematic review and meta-analysis in Ethiopia. JAC-Antimicrobial Resist. 2024, 6, dlae068. [Google Scholar] [CrossRef] [PubMed]
- Wolde, A.; Deneke, Y.; Sisay, T.; Mathewos, M. Molecular Characterization and Antimicrobial Resistance of Pathogenic Escherichia coli Strains in Children from Wolaita Sodo, Southern Ethiopia. J. Trop. Med. 2022, 2022, 9166209. [Google Scholar] [CrossRef] [PubMed]
- Gebresilasie, Y.M.; Tullu, K.D.; Yeshanew, A.G. Resistance pattern and maternal knowledge, attitude and practices of suspected Diarrheagenic Escherichia coli among children under 5 years of age in Addis Ababa, Ethiopia: Cross sectional study. Antimicrob. Resist. Infect. Control 2018, 7, 110. [Google Scholar] [CrossRef]
- Bedane, T.D.; Megersa, B.; Abunna, F.; Waktole, H.; Woldemariyam, F.T.; Tekle, M.; Shimelis, E.; Gutema, F.D. Occurrence, molecular characterization, and antimicrobial susceptibility of sorbitol non-fermenting Escherichia coli in lake water, fish and humans in central Oromia, Ethiopia. Sci. Rep. 2024, 14, 12461. [Google Scholar] [CrossRef]
- Makvana, S.; Krilov, L.R. Escherichia coli Infections. Pediatr. Rev. 2015, 36, 167–171. [Google Scholar] [CrossRef]
- Lawal, O.U.; Parreira, V.R.; Goodridge, L. The Biology and the Evolutionary Dynamics of Diarrheagenic Escherichia coli Pathotypes. In Escherichia coli—Old and New Insights; Erjavec, M.S., Ed.; IntechOpen: London, UK, 2022; pp. 1–38. [Google Scholar]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004, 2, 123–140. [Google Scholar] [CrossRef]
- Robins-Browne, R.M.; Holt, K.E.; Ingle, D.J.; Hocking, D.M.; Yang, J.; Tauschek, M. Are Escherichia coli pathotypes still relevant in the era of whole-genome sequencing? Front. Cell. Infect. Microbiol. 2016, 6, 141. [Google Scholar] [CrossRef]
- Gomes, T.A.T.; Elias, W.P.; Scaletsky, I.C.A.; Guth, B.E.C.; Rodrigues, J.F.; Piazza, R.M.F.; Ferreira, L.C.S.; Martinez, M.B. Diarrheagenic Escherichia coli. Brazilian J. Microbiol. 2016, 47, 3–30. [Google Scholar] [CrossRef]
- Servin, A.L. Pathogenesis of human diffusely adhering Escherichia coli expressing Afa/Dr adhesins (Afa/Dr DAEC): Current insights and future challenges. Clin. Microbiol. Rev. 2014, 27, 823–869. [Google Scholar] [CrossRef]
- Ramya, R.P.; Roy, S.; Thamizhmani, R.; Sugunan, A.P. Diarrheagenic Escherichia coli infections among the children of Andaman Islands with special reference to pathotype distribution and clinical profile. J. Epidemiol. Glob. Health 2017, 7, 305–308. [Google Scholar] [CrossRef] [PubMed]
- Spano, L.C.; da Cunha, K.F.; Monfardini, M.V.; de Cássia Bergamaschi Fonseca, R.; Scaletsky, I.C.A. High prevalence of diarrheagenic Escherichia coli carrying toxin-encoding genes isolated from children and adults in southeastern Brazil. BMC Infect. Dis. 2017, 17, 773. [Google Scholar] [CrossRef] [PubMed]
- Kalule, J.B.; Bester, L.A.; Banda, D.L.; Abera, D.F.; Chikuse, F.; Tesema, S.K.; Ebenezer, F.-N. Molecular Epidemiology of Diarrhoeagenic Escherichia coli in Africa: A Systematic Review and Meta-Analysis. medRxiv (Cold Spring Harb. Lab.) 2023, 1–40. [Google Scholar] [CrossRef]
- Nataro, J.P.; Kaper, J.B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 1998, 11, 142–201. [Google Scholar] [CrossRef] [PubMed]
- Canizalez-Roman, A.; Flores-Villaseñor, H.M.; Gonzalez-Nuñez, E.; Velazquez-Roman, J.; Vidal, J.E.; Muro-Amador, S.; Alapizco-Castro, G.; Alberto Díaz-Quiñonez, J.; León-Sicairos, N. Surveillance of diarrheagenic Escherichia coli strains isolated from diarrhea cases from children, adults and elderly at Northwest of Mexico. Front. Microbiol. 2016, 7, 1924. [Google Scholar] [CrossRef]
- Zelelie, T.Z.; Eguale, T.; Yitayew, B.; Abeje, D.; Alemu, A.; Seman, A.; Jass, J.; Mihret, A.; Abebe, T. Molecular epidemiology and antimicrobial susceptibility of diarrheagenic Escherichia coli isolated from children under age five with and without diarrhea in Central Ethiopia. PLoS ONE 2023, 18, e0288517. [Google Scholar] [CrossRef]
- Addisu, M.; Id, B.; Demlie, T.B.; Chekole, W.S. Molecular identification of diarrheagenic Escherichia coli pathotypes and their antibiotic resistance patterns among diarrheic children and in contact calves in Bahir Dar city. PLoS ONE 2022, 17, e0275229. [Google Scholar] [CrossRef]
- Abey, S.L.; Teka, M.; Bitew, A.B.; Molla, W.; Ejo, M.; Dagnaw, G.G.; Adugna, T.; Nigatu, S.; Mengistu, B.A.; Kinde, M.Z.; et al. Detection and antibiogram profile of diarrheagenic Escherichia coli isolated from two abattoir settings in northwest Ethiopia: A one health perspective. One Health Outlook 2024, 6, 8. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. 2013, 5, 58–65. [Google Scholar] [CrossRef]
- Uelze, L.; Grützke, J.; Borowiak, M.; Hammerl, J.A.; Juraschek, K.; Deneke, C.; Tausch, S.H.; Malorny, B. Typing methods based on whole genome sequencing data. One Health Outlook 2020, 2, 3. [Google Scholar] [CrossRef]
- Catoiu, E.A.; Phaneuf, P.; Monk, J.; Palsson, B.O. Whole-genome sequences from wild-type and laboratory-evolved strains define the alleleome and establish its hallmarks. Proc. Natl. Acad. Sci. USA 2023, 120, e2218835120. [Google Scholar] [CrossRef] [PubMed]
- Croxen, M.A.; Law, R.J.; Scholz, R.; Keeney, K.M.; Wlodarska, M.; Finlay, B.B. Recent advances in understanding enteric pathogenic Escherichia coli. Clin. Microbiol. Rev. 2013, 26, 822–880. [Google Scholar] [CrossRef] [PubMed]
- Belina, D.; Gobena, T.; Kebede, A.; Chimdessa, M.; Hailu, Y.; Hald, T. Occurrence of Diarrheagenic Pathogens and Their Coinfection Profiles in Diarrheic under Five Children and Tracked Human Contacts in Urban and Rural Settings of Eastern Ethiopia. Microbiol. Insights 2023, 16, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Ifeanyi, C.I.C.; Ikeneche, N.F.; Bassey, B.E.; Al-Gallas, N.; Aissa, R.B.; Boudabous, A. Diarrheagenic Escherichia coli pathotypes isolated from children with diarrhea in the federal capital territory abuja, Nigeria. J. Infect. Dev. Ctries. 2015, 9, 165–174. [Google Scholar] [CrossRef]
- Konaté, A.; Dembélé, R.; Kagambèga, A.; Soulama, I.; Kaboré, W.A.D.; Sampo, E.; Cissé, H.; Sanou, A.; Serme, S.; Zongo, S.; et al. Molecular characterization of diarrheagenic Escherichia coli in children less than 5 years of age with diarrhea in Ouagadougou, Burkina Faso. Eur. J. Microbiol. Immunol. 2017, 7, 220–228. [Google Scholar] [CrossRef]
- Esimogu, O.K.; Adamu, A.M.; Nafarnda, W.D. Prevalence of Pathotypes of Diarrheagenic Escherichia coli among Diarrhea Patients in Kwali, Federal Capital Territory, Nigeria. J. Epidemiol. Soc. Niger. 2023, 6, 13–23. [Google Scholar] [CrossRef]
- Al-Gallas, N.; Bahri, O.; Bouratbeen, A.; Haasen, A.B.; Aissa, R.B. Etiology of acute diarrhea in children and adults in Tunis, Tunisia, with emphasis on diarrheagenic Escherichia coli: Prevalence, phenotyping, and molecular epidemiology. Am. J. Trop. Med. Hyg. 2007, 77, 571–582. [Google Scholar] [CrossRef]
- Saka, H.K.; Dabo, N.T.; Muhammad, B.; García-Soto, S.; Ugarte-Ruiz, M.; Alvarez, J. Diarrheagenic Escherichia coli Pathotypes From Children Younger Than 5 Years in Kano State, Nigeria. Front. Public Health 2019, 7, 348. [Google Scholar] [CrossRef]
- Mabika, R.M.; Liabagui, S.L.O.; Moundounga, H.K.; Mounioko, F.; Souza, A.; Yala, J.F. Molecular Prevalence and Epidemiological Characteristics of Diarrheagenic E. coli in Children under 5 Years Old in the City of Koula-Moutou, East-Central Gabon. Open J. Med. Microbiol. 2021, 11, 157–175. [Google Scholar] [CrossRef]
- Yeda, R.; Makalliwa, G.; Apondi, E.; Sati, B.; Riziki, L.; Ouma, C.; Anguko, E.; Opot, B.; Okoth, R.; Koech, E.J.; et al. Comparative prevalence of diarrheagenic Escherichia coli between children below five years with close contact with food animals in Kisumu County, Kenya. Pan Afr. Med. J. 2024, 47, 25. [Google Scholar] [CrossRef]
- Manhique-coutinho, L.; Chiani, P.; Michelacci, V.; Taviani, E.; Fernando, A.; Bauhofer, L.; Chissaque, A.; Cossa-moiane, I.; Sambo, J.; Chilaúle, J.; et al. Molecular characterization of diarrheagenic Escherichia coli isolates from children with diarrhea: A cross-sectional study in four provinces of Mozambique. Int. J. Infect. Dis. 2022, 121, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Stoppe, N.d.C.; Silva, J.S.; Carlos, C.; Sato, M.I.Z.; Saraiva, A.M.; Ottoboni, L.M.M.; Torres, T.T. Worldwide phylogenetic group patterns of Escherichia coli from commensal human and wastewater treatment plant isolates. Front. Microbiol. 2017, 8, 2512. [Google Scholar] [CrossRef] [PubMed]
- Olowe, O.A.; Adefioye, O.J.; Ajayeoba, T.A.; Schiebel, J.; Weinreich, J.; Ali, A.; Burdukiewicz, M.; Rödiger, S.; Schierack, P. Phylogenetic grouping and biofilm formation of multidrug resistant Escherichia coli isolates from humans, animals and food products in South-West Nigeria. Sci. Afr. 2019, 6, e00158. [Google Scholar] [CrossRef]
- Adjhi, D.; Nazaire, K.; Etienne, D.; Marcellin, D.; Mireille, D. Virulence, serotype and phylogenetic groups of diarrhoeagenic Escherichia coli isolated during digestive infections in Abidjan, Cte dIvoire. Afr. J. Biotechnol. 2014, 13, 998–1008. [Google Scholar] [CrossRef]
- El-baz, R.; Said, H.S.; Abdelmegeed, E.S.; Barwa, R. Characterization of virulence determinants and phylogenetic background of multiple and extensively drug resistant Escherichia coli isolated from different clinical sources in Egypt. Appl. Microbiol. Biotechnol. 2022, 106, 1279–1298. [Google Scholar] [CrossRef]
- Khairy, R.M.; Mohamed, E.S.; Ghany, H.M.A.; Abdelrahim, S.S. Phylogenic classification and virulence genes profiles of uropathogenic E. coli and diarrhegenic E. coli strains isolated from community acquired infections. PLoS ONE 2019, 14, e0222441. [Google Scholar] [CrossRef]
- Escobar-Páramo, P.; Le Menac’h, A.; Le Gall, T.; Amorin, C.; Gouriou, S.; Picard, B.; Skurnik, D.; Denamur, E. Identification of forces shaping the commensal Escherichia coli genetic structure by comparing animal and human isolates. Environ. Microbiol. 2006, 8, 1975–1984. [Google Scholar] [CrossRef]
- Javadi, M.; Bouzari, S.; Oloomi, M. Horizontal Gene Transfer and the Diversity of Escherichia coli. In Escherichia coli—Recent Advances on Physiology, Pathogenesis and Biotechnological Applications; Samie, A., Ed.; IntechOpen: London, UK, 2017; pp. 1–17. [Google Scholar]
- Tegha, G.; Ciccone, E.J.; Krysiak, R.; Kaphatika, J.; Chikaonda, T.; Ndhlovu, I.; van Duin, D.; Hoffman, I.; Juliano, J.J.; Wang, J. Genomic epidemiology of Escherichia coli isolates from a tertiary referral center in lilongwe, Malawi. Microb. Genom. 2021, 7, 000490. [Google Scholar] [CrossRef]
- Okeke, I.N.; Wallace-Gadsden, F.; Simons, H.R.; Matthews, N.; Labar, A.S.; Hwang, J.; Wain, J. Multi-locus sequence typing of enteroaggregative Escherichia coli isolates from nigerian children uncovers multiple lineages. PLoS ONE 2010, 5, e14093. [Google Scholar] [CrossRef]
- Riley, L.W. Distinguishing Pathovars from Nonpathovars: Escherichia coli. Microbiol. Spectr. 2020, 8, 1–23. [Google Scholar] [CrossRef]
- El-Baky, R.M.A.; Ibrahim, R.A.; Mohamed, D.S.; Ahmed, E.F.; Hashem, Z.S. Prevalence of virulence genes and their association with antimicrobial resistance among pathogenic E. coli isolated from Egyptian patients with different clinical infections. Infect. Drug Resist. 2020, 13, 1221–1236. [Google Scholar] [CrossRef] [PubMed]
- Dahwash, S.M.; Raheema, R.H.; Albahadili, M.A.; Maslat, A.H. Distribution of Phylogenetics and Virulence Genes of Uropathogenic Escherichia coli Among Urinary Tract Infection in Pregnant Women. Biochem. Cell. Arch. 2021, 21, 449–456. [Google Scholar]
- Tarchouna, M.; Ferjani, A.; Ben-Selma, W.; Boukadida, J. Distribution of uropathogenic virulence genes in Escherichia coli isolated from patients with urinary tract infection. Int. J. Infect. Dis. 2013, 17, e450–e453. [Google Scholar] [CrossRef] [PubMed]
- Mbanga, J.; Mudzana, R. Virulence Factors and Antibiotic Resistance Patterns of Uropathogenic Escherichia coli. Afr. J. Agric. Res. 2014, 8, 3678–3686. [Google Scholar]
- Adwan, G.; Adwan, K.; Bourinee, H. Molecular characterization of some new E. coli strains theoretically responsible for both intestinal and extraintestinal infections. Int. J. Med. Res. Health Sci. 2016, 5, 158–163. [Google Scholar]
- Nielsen, K.L.; Stegger, M.; Kiil, K.; Godfrey, P.A.; Feldgarden, M.; Lilje, B.; Andersen, P.S.; Frimodt-Møller, N. Whole-genome comparison of urinary pathogenic Escherichia coli and faecal isolates of UTI patients and healthy controls. Int. J. Med. Microbiol. 2018, 307, 497–507. [Google Scholar] [CrossRef]
- Santos, A.C.d.M.; Santos, F.F.; Silva, R.M.; Gomes, T.A.T. Diversity of Hybrid- and Hetero-Pathogenic Escherichia coli and Their Potential Implication in More Severe Diseases. Front. Cell. Infect. Microbiol. 2020, 10, 339. [Google Scholar] [CrossRef]
- McCoy, C.S.; Mannion, A.J.; Feng, Y.; Madden, C.M.; Artim, S.C.; Au, G.G.; Dolan, M.; Haupt, J.L.; Burns, M.A.; Sheh, A.; et al. Cytotoxic Escherichia coli strains encoding colibactin, cytotoxic necrotizing factor, and cytolethal distending toxin colonize laboratory common marmosets (Callithrix jacchus). Sci. Rep. 2021, 11, 2309. [Google Scholar] [CrossRef]
- Valat, C.; Haenni, M.; Arnaout, Y.; Drapeau, A.; Hirchaud, E.; Touzain, F.; Boyer, T.; Delannoy, S.; Vorimore, F.; Fach, P.; et al. F74 plasmids are major vectors of virulence genes in bovine NTEC2. Lett. Appl. Microbiol. 2022, 75, 355–362. [Google Scholar] [CrossRef]
- Pérès, S.Y.; Marchès, O.; Daigle, F.; Nougayrède, J.P.; Hérault, F.; Tasca, C.; De Rycke, J.; Oswald, E. A new cytolethal distending toxin (CDT) from Escherichia coli producing CNF2 blocks HeLa cell division in G2/M phase. Mol. Microbiol. 1997, 24, 1095–1107. [Google Scholar] [CrossRef]
- Brunder, W.; Schmidt, H.; Karch, H. EspP, a novel extracellular serine protease of enterohaemorrhagic Escherichia coli O157:H7 cleaves human coagulation factor V. Mol. Microbiol. 1997, 24, 767–778. [Google Scholar] [CrossRef] [PubMed]
- Nasrollahian, S.; Graham, J.P.; Halaji, M. A review of the mechanisms that confer antibiotic resistance in pathotypes of E. coli. Front. Cell. Infect. Microbiol. 2024, 14, 1387497. [Google Scholar] [CrossRef] [PubMed]
- Zenebe, T.; Eguale, T.; Desalegn, Z.; Beshah, D.; Gebre-Selassie, S.; Mihret, A.; Abebe, T. Distribution of ß-Lactamase Genes Among Multidrug-Resistant and Extended-Spectrum ß-Lactamase-Producing Diarrheagenic Escherichia coli from Under-Five Children in Ethiopia. Infect. Drug Resist. 2023, 16, 7041–7054. [Google Scholar] [CrossRef] [PubMed]
- Dela, H.; Egyir, B.; Majekodunmi, A.O.; Behene, E.; Yeboah, C.; Ackah, D.; Bongo, R.N.A.; Bonfoh, B.; Zinsstag, J.; Bimi, L.; et al. Diarrhoeagenic E. coli occurrence and antimicrobial resistance of Extended Spectrum Beta-Lactamases isolated from diarrhoea patients attending health facilities in Accra, Ghana. PLoS ONE 2022, 17, e0268991. [Google Scholar] [CrossRef]
- Ramatla, T.; Tawana, M.; Lekota, K.E.; Thekisoe, O. Antimicrobial resistance genes of Escherichia coli, a bacterium of “One Health” importance in South Africa: Systematic review and meta-analysis. AIMS Microbiol. 2023, 9, 75–89. [Google Scholar] [CrossRef]
- Evans, D.R.; Griffith, M.P.; Sundermann, A.J.; Shutt, K.A.; Saul, M.I.; Mustapha, M.M.; Marsh, J.W.; Cooper, V.S.; Harrison, L.H.; Van Tyne, D. Systematic detection of horizontal gene transfer across genera among multidrug-resistant bacteria in a single hospital. eLife 2020, 9, e53886. [Google Scholar] [CrossRef]
- Smith, J.L.; Fratamico, P.M.; Gunther, N.W. Shiga toxin-producing Escherichia coli. Adv. Appl. Microbiol. 2014, 86, 145–197. [Google Scholar] [CrossRef]
- Bielaszewska, M.; Prager, R.; Köck, R.; Mellmann, A.; Zhang, W.; Tschäpe, H.; Tarr, P.I.; Karch, H. Shiga toxin gene loss and transfer in vitro and in vivo during enterohemorrhagic Escherichia coli O26 infection in humans. Appl. Environ. Microbiol. 2007, 73, 3144–3150. [Google Scholar] [CrossRef]
- Rahal, E.A.; Fadlallah, S.M.; Nassar, F.J.; Kazzi, N. Approaches to treatment of emerging Shiga toxin-producing Escherichia coli infections highlighting the O104:H4 serotype. Front. Cell. Infect. Microbiol. 2015, 5, 24. [Google Scholar] [CrossRef]
- Pokharel, P.; Dhakal, S.; Dozois, C.M. The diversity of Escherichia coli pathotypes and vaccination strategies against this versatile bacterial pathogen. Microorganisms 2023, 11, 344. [Google Scholar] [CrossRef]
- World Health Organization & Food and Agriculture Organization of the United Nations. Shiga Toxin-Producing Escherichia coli (STEC) and Food: Attribution, Characterization, and Monitoring; World Health Organization: Geneva, Switzerland, 2018. [Google Scholar]
- Liu, Y.; Thaker, H.; Wang, C.; Xu, Z.; Dong, M. Diagnosis and Treatment for Shiga Toxin-Producing Escherichia coli Associated Hemolytic Uremic Syndrome. Toxins 2023, 15, 10. [Google Scholar] [CrossRef] [PubMed]
- Wolde, D.; Eguale, T.; Alemayehu, H.; Medhin, G.; Haile, A.F.; Pirs, M.; Smrdel, K.S.; Avberšek, J.; Kušar, D.; Kišek, T.C.; et al. Antimicrobial Susceptibility and Characterization of Extended-Spectrum β -Lactamase-Producing Escherichia coli Isolated from Stools of Primary Healthcare Patients in Ethiopia. Antibiotics 2024, 13, 93. [Google Scholar] [CrossRef] [PubMed]
- Seegene. AllplexTM GI-Bacteria (II) Assay; Seegene Inc.: Seoul, Republic of Korea, 2022; Volume 1. [Google Scholar]
- Mainda, G.; Lupolova, N.; Sikakwa, L.; Richardson, E.; Bessell, P.R.; Malama, S.K.; Kwenda, G.; Stevens, M.P.; Bronsvoort, B.M.D.; Muma, J.B.; et al. Whole Genome Sequence Analysis Reveals Lower Diversity and Frequency of Acquired Antimicrobial Resistance (AMR) Genes in E. coli from Dairy Herds Compared with Human Isolates from the Same Region of Central Zambia. Front. Microbiol. 2019, 10, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Illumina. Illumina DNA Prep Reference Guide; Illumina, Inc.: San Diego, CA, USA, 2020. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Babraham Bioinformatics. FastQC A Quality Control Tool for High Throughput Sequence Data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 11 October 2023).
- Prjibelski, A.; Antipov, D.; Meleshko, D.; Lapidus, A.; Korobeynikov, A. Using SPAdes De Novo Assembler. Curr. Protoc. Bioinform. 2020, 70, e102. [Google Scholar] [CrossRef]
- Mikheenko, A.; Prjibelski, A.; Saveliev, V.; Antipov, D.; Gurevich, A. Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 2018, 34, i142–i150. [Google Scholar] [CrossRef]
- Beghain, J.; Bridier-Nahmias, A.; Le Nagard, H.; Denamur, E.; Clermont, O. ClermonTyping: An easy-to-use and accurate in silico method for Escherichia genus strain phylotyping. Microb. Genom. 2018, 4, e000192. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Microbiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef]
- Joensen, K.G.; Tetzschner, A.M.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and easy in silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.H.K.; Bortolaia, V.; Tansirichaiya, S.; Aarestrup, F.M.; Roberts, A.P.; Petersen, T.N. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J. Antimicrob. Chemother. 2021, 76, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Han, S.; Saha, S.; Oler, E.; Peters, H.; Grant, J.R.; Stothard, P.; Gautam, V. PHASTEST: Faster than PHASTER, better than PHAST. Nucleic Acids Res. 2023, 51, W443–W450. [Google Scholar] [CrossRef] [PubMed]


| Characteristics | Number of DEC Isolates (%) | |||
|---|---|---|---|---|
| EPEC (n = 10) | EAEC (n = 6) | ETEC (n = 5) | STEC (n = 3) | |
| Study site | ||||
| Addis Ababa (n = 12) | 4 (16.7) | 3 (12.5) | 2 (8.3) | 3 (12.5) |
| Hossana (n = 12) | 6 (25) | 3 (12.5) | 3 (12.5) | 0 |
| Sex | ||||
| Male (n = 10) | 6 (25.0) | 3 (12.5) | 0 | 1 (4.2) |
| Female (n = 14) | 4 (16.7) | 3 (12.5) | 5 (20.8) | 2 (8.3) |
| Participants | ||||
| Diarrheic (n = 14) | 7 (29.2) | 3 (12.5) | 4 (16.7) | 0 |
| Non-diarrheic (n = 10) | 3 (12.5) | 3 (12.5) | 1 (4.2) | 3 (12.5) |
| Age group | ||||
| 0–4 (n = 3) | 2 (8.3) | 0 | 1 (4.2) | 0 |
| 5–9 (n = 2) | 1 (4.2) | 1 (8.3) | 0 | 0 |
| 10–14 (n = 3) | 2 (8.3) | 0 | 0 | 1 (4.2) |
| 15–19 (n = 2) | 0 | 1 (4.2) | 1 (4.2) | 0 |
| 20–45 (n = 11) | 5 (20.8) | 3 (12.5) | 2 (8.3) | 1 (4.2) |
| 46–65 (n = 3) | 0 | 1 (4.2) | 1 (4.2) | 1 (4.2) |
| Pathotype as Determined by WGS | Phylogroup | ST | Serotype | Pathotype Specific Virulence Genes |
|---|---|---|---|---|
| EPEC | A | 10 | O90:H40 | eae-g02-θ |
| A | 10 | O90:H40 | eae-g02-θ | |
| A | 382 | H5 | eae-e08-π | |
| B1 | 40 | O21:H21 | eae-b01a-β | |
| B1 | 517 | O111:H19 | eae-e02-ε | |
| B1 | 517 | H19 | eae-e02-ε | |
| B1 | 616 | H21 | eae-e06-η | |
| B1 | 4038 | O93:H28 | eae-e06-η | |
| B2 | 2346 | O142:H34 | eae-a01-α | |
| E | 8508 | O136:H49 | eae-g02-θ | |
| EAEC | A | 10 | O65:H12 | aggR |
| A | 34 | O99:H33 | aggR | |
| B1 | 295 | O171:H5 | aggR | |
| B1 | 678 | O104:H4 | aggR | |
| B1 | 1136 | O59:H19 | aggR | |
| B1 | 1136 | O167:H19 | aggR | |
| ETEC | A | 10 | O159:H21 | estah-STa3 |
| A | 1564 | H21 | estah-STa3 | |
| B1 | 155 | O58:H51 | eltIAB-8 | |
| B1 | 314 | O112ab:H21 | eltIAB-11 | |
| B1 | 314 | O112ab:H21 | eltIAB-11 | |
| STEC | B1 | 101 | O117:H12 | stx1 (stx1a) |
| B1 | 155 | H25 | stx2 (stx2c) | |
| B1 | 737 | O81:H21 | stx1 (stx1c) |
| Virulence-Associated Genes and Their Encoded Proteins/Functions | Prevalence in Each Pathotypes N (%), N = 24 | |||||
|---|---|---|---|---|---|---|
| EPEC (n = 10) | EAEC (n = 6) | ETEC (n = 5) | STEC (n = 3) | |||
| Adhesins | aap | Dispersin, antiaggregation protein | 0 | 6 (25) | 0 | 0 |
| afaA | Transcriptional regulator | 1 (4.2) | 0 | 0 | 0 | |
| afaC | Outer membrane usher protein | 1 (4.2) | 0 | 0 | 0 | |
| afaD | Afimbrial adhesion | 1 (4.2) | 3 (12.5) | 0 | 0 | |
| agg3A | AAF/III major fimbrial subunit | 0 | 3 (12.5) | 0 | 0 | |
| agg3B | AAF/III minor adhesin | 0 | 3 (12.5) | 0 | 0 | |
| agg3C | Usher, AAF/III assembly unit | 0 | 3 (12.5) | 0 | 0 | |
| agg3D | Chaperone, AAF/III assembly unit | 0 | 3 (12.5) | 0 | 0 | |
| agg4A | AAF/IV major fimbrial subunit | 0 | 2 (8.3) | 0 | 0 | |
| agg4C | Usher, AAF/IV assembly unit | 0 | 2 (8.3) | 0 | 0 | |
| aggA | AAF/I major fimbrial subunit | 0 | 1 (4.2) | 0 | 0 | |
| aggC | Usher, AAF/I assembly unit | 0 | 1 (4.2) | 0 | 0 | |
| aggD | Chaperone, AAF/I assembly unit | 0 | 1 (4.2) | 0 | 0 | |
| aggR | AraC transcriptional activator | 0 | 6 (25) | 0 | 0 | |
| cif | Type III secreted effector | 4 (16.7) | 0 | 0 | 0 | |
| csgA | Curli major subunit CsgA | 9 (37.5) | 6 (25) | 5 (20.8) | 2 (8.3) | |
| cswR | CS12 Transcriptional activator | 0 | 0 | 2 (8.3) | 0 | |
| eaeA | Intimin | 10 (41.7) | 0 | 0 | 0 | |
| efa1 | EHEC factor for adherence; lymphostatin Efa1-LifA | 1 (4.2) | 0 | 0 | 0 | |
| etpB | EtpB Nonfimbrial adhesin/TPS transporter | 0 | 0 | 1 (4.2) | 0 | |
| faeC | F4 (K88) Minor fimbrial subunit | 0 | 0 | 0 | 1 (4.2) | |
| fdeC | Intimin-like adhesin FdeC | 5 (20.8) | 2 (8.3) | 3 (12.5) | 3 (12.5) | |
| fimH | Type 1 fimbriae | 8 (33.3) | 5 (20.8) | 4 (16.7) | 3 (12.5) | |
| hra | Heat-resistant agglutinin | 0 | 2 (8.3) | 1 (4.2) | 0 | |
| iha | Adherence protein | 0 | 2 (8.3) | 0 | 3 (12.5) | |
| lpfA | Long polar fimbriae | 4 (16.7) | 4 (16.7) | 3 (12.5) | 3 (12.5) | |
| nlpI | Lipoprotein NlpI precursor | 10 (41.7) | 6 (25) | 5 (20.8) | 3 (12.5) | |
| perA | EPEC adherence factor | 1 (4.2) | 0 | 0 | 0 | |
| tir | Translocated intimin receptor protein | 9 (37.5) | 0 | 0 | 0 | |
| yehA | Outer membrane lipoprotein, YHD fimbrial cluster | 6 (25) | 3 (12.5) | 4 (16.7) | 2 (8.3) | |
| yehB | Usher, YHD fimbrial cluster | 8 (33.3) | 6 (25) | 5 (20.8) | 3 (12.5) | |
| yehC | Chaperone, YHD fimbrial cluster | 8 (33.3) | 6 (25) | 5 (20.8) | 3 (12.5) | |
| yehD | Major pilin subunit, YHD fimbrial cluster | 8 (33.3) | 5 (20.8) | 5 (20.8) | 3 (12.5) | |
| yfcV | Fimbrial protein | 1 (4.2) | 0 | 0 | 0 | |
| Protectins | aaiC | Type VI secretion protein | 0 | 3 (12.5) | 0 | 0 |
| aar | AggR-activated regulator | 0 | 4 (16.7) | 0 | 0 | |
| aatA | Dispersin transporter protein | 0 | 6 (25) | 0 | 0 | |
| aamR:FN554766 | Not defined | 0 | 1 (4.2) | 0 | 0 | |
| anr | AraC negative regulator | 3 (12.5) | 5 (20.8) | 1 (4.2) | 0 | |
| aslA | Arylsulfatase-like protein | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| capU | Hexosyltransferase homolog | 2 (8.3) | 3 (12.5) | 1 (4.2) | 0 | |
| cba | Colicin B | 0 | 0 | 0 | 1 (4.2) | |
| cea | Colicin E1 | 0 | 1 (4.2) | 0 | 0 | |
| cia | Colicin Ia | 1 (4.2) | 0 | 0 | 1 (4.2) | |
| cib | Colicin Ib | 0 | 0 | 2 (8.3) | 0 | |
| cma | Colicin M | 0 | 0 | 0 | 1 (4.2) | |
| colE2 | Colicin E2 | 0 | 0 | 0 | 1 (4.2) | |
| colE5 | Colicin E5 lysis protein Lys | 0 | 2 (8.3) | 0 | 0 | |
| cvaC | Microcin C | 1 (4.2) | 0 | 0 | 0 | |
| eilA | Salmonella HilA homolog | 1 (4.2) | 0 | 0 | 0 | |
| espA | Type III secretions system | 10 (41.7) | 0 | 0 | 0 | |
| espB | Secreted protein B | 1 (4.2) | 0 | 0 | 0 | |
| espC | Escherichia coli enterotoxin EspC | 1 (4.2) | 0 | 0 | 0 | |
| espF | EPEC secreted protein F; Type III secretion system | 4 (16.7) | 0 | 0 | 0 | |
| espI | Serine protease autotransporters of Enterobacteriaceae (SPATE) EspI | 0 | 1 (4.2) | 0 | 1 (4.2) | |
| espJ | Prophage-encoded type III secretion system effector | 5 (20.8) | 0 | 0 | 0 | |
| espP | Extracellular serine protease plasmid-encoded | 0 | 0 | 0 | 1 (4.2) | |
| espY2 | Non-LEE-encoded type III secreted effector | 1 (4.2) | 0 | 0 | 0 | |
| etpC | EtpC Glycotransferase | 2 (8.3) | 0 | 0 | 0 | |
| etpD | Type II secretion protein EtpD | 1 (4.2) | 0 | 1 (4.2) | 0 | |
| etsC | Putative type I secretion outer membrane protein | 1 (4.2) | 0 | 1 (4.2) | 0 | |
| gad | Glutamate decarboxylase | 3 (12.5) | 2 (8.3) | 2 (8.3) | 0 | |
| hha | Hemolysin expression modulator Hha (previous rmoA) | 5 (20.8) | 1 (4.2) | 0 | 0 | |
| iss | Increased serum survival | 4 (16.7) | 2 (8.3) | 1 (4.2) | 2 (8.3) | |
| kpsE | Capsule polysaccharide export inner-membrane protein | 0 | 0 | 0 | 1 (4.2) | |
| mchB | Precursor of microcin H47 | 0 | 1 (4.2) | 0 | 1 (4.2) | |
| mchC | MchC protein | 0 | 1 (4.2) | 0 | 1 (4.2) | |
| mchF | ABC transporter protein MchF | 1 (4.2) | 1 (4.2) | 0 | 1 (4.2) | |
| mcmA | Microcin M | 0 | 1 (4.2) | 0 | 1 (4.2) | |
| neuC | Polysialic acid capsule biosynthesis protein | 0 | 2 (8.3) | 0 | 0 | |
| nleB | Non-LEE encoded effector B | 6 (25) | 0 | 0 | 0 | |
| ompT | Outer membrane protease (protein protease 7) | 3 (12.5) | 0 | 1 (4.2) | 2 (8.3) | |
| ORF3 | Isoprenoid Biosynthesis | 0 | 6 (25) | 0 | 0 | |
| ORF4 | Putative isopentenyl-diphosphate delta-isomerase | 0 | 6 (25) | 0 | 0 | |
| shiA | Homologue of the Shigella flexneri SHI-2 pathogenicity island gene shiA | 0 | 2 (8.3) | 1 (4.2) | 0 | |
| shiB | Homologue of the Shigella flexneri SHI-2 pathogenicity island gene shiB | 1 (4.2) | 3 (12.5) | 0 | 1 (4.2) | |
| tia | Tia invasion determinant | 0 | 0 | 0 | 1 (4.2) | |
| traJ | Protein TraJ (Positive regulator of conjugal transfer operon) | 4 (16.7) | 3 (12.5) | 3 (12.5) | 0 | |
| traT | Outer membrane protein complement resistance | 3 (12.5) | 3 (12.5) | 3 (12.5) | 3 (12.5) | |
| tsh | Serine protease autotransporter of Enterobacteriaceae (SPATE)-Immunoglobulin A1 protease- Temperature-sensitive hemagglutinin | 1 (4.2) | 0 | 0 | 0 | |
| Iron uptake | aalF | CS23 Minor structural subunit | 0 | 0 | 0 | 1 (4.2) |
| chuA | Outer membrane hemin receptor | 2 (8.3) | 0 | 0 | 0 | |
| fyuA | Yersiniabactin siderophore receptor | 0 | 3 (12.5) | 0 | 1 (4.2) | |
| ireA | Iron-regulated outer membrane protein IreA | 0 | 2 (8.3) | 0 | 0 | |
| iroN | Salmochelin siderophore receptor protein | 1 (4.2) | 0 | 0 | 0 | |
| irp2 | Yersiniabactin non-ribosomal peptide synthetase | 0 | 2 (8.3) | 0 | 1 (4.2) | |
| iucC | Aerobactin synthetase | 1 (4.2) | 4 (16.7) | 0 | 1 (4.2) | |
| iutA | Ferric aerobactin receptor | 1 (4.2) | 4 (16.7) | 0 | 1 (4.2) | |
| sitA | Iron transport protein | 1 (4.2) | 2 (8.3) | 1 (4.2) | 0 | |
| terC | Tellurium ion resistance protein | 9 (37.5) | 6 (25) | 5 (20.8) | 3 (12.5) | |
| Toxins | astA | Heat-stable enterotoxin EAST-1 | 1 (4.2) | 4 (16.7) | 4 (16.7) | 0 |
| cdt-IIIB | Cytolethal distending toxin III subunit B | 0 | 0 | 0 | 1 (4.2) | |
| cnf2 | Cytotoxic necrotizing factor 2 | 0 | 0 | 0 | 1 (4.2) | |
| ehxA | Enterohaemolysin | 0 | 0 | 0 | 1 (4.2) | |
| eltIAB-8 | Heat-labile enterotoxin LTIh-8 | 0 | 0 | 3 (12.5) | 0 | |
| eltIAB-11 | Heat-labile enterotoxin LTIh-11 | |||||
| estah-STa3 | Heat-stable enterotoxin STa3 human variant | 0 | 0 | 2 (8.3) | 0 | |
| hlyA | Hemolysin A | 1 (4.2) | 0 | 0 | 0 | |
| hlyE | Avian E. coli haemolysin | 8 (33.3) | 6 (25) | 4 (16.7) | 3 (12.5) | |
| hlyF | Hemolysin F | 1 (4.2) | 0 | 1 (4.2) | 0 | |
| pet | Autotransporter enterotoxin | 1 (4.2) | 0 | 0 | 0 | |
| pic | Serine protease autotransporter of Enterobacteriaceae (SPATE) Pic | 0 | 2 (8.3) | 0 | 0 | |
| sat | Serine protease autotransporter of Enterobacteriaceae (SPATE) Sat | 2 (8.3) | 2 (8.3) | 0 | 0 | |
| senB | Plasmid-encoded enterotoxin | 2 (8.3) | 0 | 0 | 0 | |
| sigA | Serine protease autotransporter of Enterobacteriaceae (SPATE) Shigella IgA-like protease homologue | 0 | 3 (12.5) | 0 | 0 | |
| stx1 | Shiga toxin 1 | 0 | 0 | 0 | 2 (8.3) | |
| stx2 | Shiga toxin 2 | 0 | 0 | 0 | 1 (4.2) | |
| subA | Subtilase toxin subunit | 0 | 0 | 0 | 1 (4.2) | |
| toxB | Toxin B | 1 (4.2) | 0 | 0 | 0 | |
| vat | Serine protease autotransporter of Enterobacteriaceae (SPATE) Vat | 0 | 0 | 0 | 1 (4.2) | |
| Virulence-Associated Genes and Their Encoded Proteins/Functions | Prevalence in Each Phylogenetic Group N (%), N = 24 | |||||
|---|---|---|---|---|---|---|
| A (n = 7) | B1 (n = 15) | B2 (n = 1) | E (n = 1) | |||
| Adhesins | aap | Dispersin, antiaggregation protein | 2 (8.3) | 4 (16.7) | 0 | 0 |
| afaA | Transcriptional regulator | 1 (4.2) | 0 | 0 | 0 | |
| afaC | Outer membrane usher protein | 1 (4.2) | 0 | 0 | 0 | |
| afaD | Afimbrial adhesion | 2 (8.3) | 2 (8.3) | 0 | 0 | |
| agg3A | AAF/III major fimbrial subunit | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| agg3B | AAF/III minor adhesin | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| agg3C | Usher, AAF/III assembly unit | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| agg3D | Chaperone, AAF/III assembly unit | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| agg4A | AAF/IV major fimbrial subunit | 0 | 2 (8.3) | 0 | 0 | |
| agg4C | Usher, AAF/IV assembly unit | 0 | 2 (8.3) | 0 | 0 | |
| aggA | AAF/I major fimbrial subunit | 1 (4.2) | 0 | 0 | 0 | |
| aggC | Usher, AAF/I assembly unit | 1 (4.2) | 0 | 0 | 0 | |
| aggD | Chaperone, AAF/I assembly unit | 1 (4.2) | 0 | 0 | 0 | |
| aggR | AraC transcriptional activator | 2 (8.3) | 4 (16.7) | 0 | 0 | |
| cif | Type III secreted effector | 2 (8.3) | 1 (4.2) | 0 | 1 (4.2) | |
| csgA | Curli major subunit CsgA | 7 (29.2) | 15 (62.5) | 1 (4.2) | 1 (4.2) | |
| cswR | CS12 Transcriptional activator | 0 | 2 (8.3) | 0 | 0 | |
| eaeA | Intimin | 3 (12.5) | 4 (16.7) | 1 (4.2) | 1 (4.2) | |
| efa1 | EHEC factor for adherence; lymphostatin Efa1-LifA | 0 | 1 (4.2) | 0 | 0 | |
| etpB | EtpB nonfimbrial adhesin/TPS transporter | 1 (4.2) | 0 | 0 | 0 | |
| faeC | F4 (K88) Minor fimbrial subunit | 0 | 1 (4.2) | 0 | 0 | |
| fdeC | Intimin-like adhesin FdeC | 0 | 12 (50) | 0 | 1 (4.2) | |
| fimH | Type 1 fimbriae | 6 (25) | 13 (54.2) | 0 | 1 (4.2) | |
| hra | Heat-resistant agglutinin | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| iha | Adherence protein | 0 | 5 (20.8) | 0 | 0 | |
| lpfA | Long polar fimbriae | 0 | 14 (58.3) | 0 | 0 | |
| nlpI | Lipoprotein NlpI precursor | 6 (25) | 12 (50) | 1 (4.2) | 1 (4.2) | |
| perA | EPEC adherence factor | 0 | 0 | 1 (4.2) | 0 | |
| tir | Translocated intimin receptor protein | 3 (12.5) | 5 (20.8) | 0 | 1 (4.2) | |
| yehA | Outer membrane lipoprotein, YHD fimbrial cluster | 7 (29.2) | 7 (29.2) | 1 (4.2) | 0 | |
| yehB | Usher, YHD fimbrial cluster | 5 (20.8) | 15 (62.5) | 1 (4.2) | 1 (4.2) | |
| yehC | Chaperone, YHD fimbrial cluster | 5 (20.8) | 15 (62.5) | 1 (4.2) | 1 (4.2) | |
| yehD | Major pilin subunit, YHD fimbrial cluster | 5 (20.8) | 14 (58.3) | 1 (4.2) | 1 (4.2) | |
| yfcV | Fimbrial protein | 0 | 0 | 1 (4.2) | 1 (4.2) | |
| Protectins | aaiC | Type VI secretion protein | 1 (4.2) | 1 (4.2) | 0 | 0 |
| aar | AggR-activated regulator | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| aatA | Dispersin transporter protein | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| aamR:FN554766 | Not defined | 0 | 1 (4.2) | 0 | 0 | |
| anr | AraC negative regulator | 4 (16.7) | 2 (8.3) | 0 | 0 | |
| aslA | Arylsulfatase-like protein | 1 (4.2) | 0 | 1 (4.2) | 0 | |
| capU | Hexosyltransferase homolog | 1 (4.2) | 3 (12.5) | 0 | 0 | |
| cba | Colicin B | 0 | 1 (4.2) | 0 | 0 | |
| cea | Colicin E1 | 1 (4.2) | 0 | 0 | 0 | |
| cia | Colicin Ia | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| cib | Colicin Ib | 2 (8.3) | 0 | 0 | 0 | |
| cma | Colicin M | 0 | 1 (4.2) | 0 | 0 | |
| colE2 | Colicin E2 | 0 | 1 (4.2) | 0 | 0 | |
| colE5 | Colicin E5 lysis protein Lys | 0 | 2 (8.3) | 0 | 0 | |
| cvaC | Microcin C | 1 (4.2) | 0 | 0 | 0 | |
| eilA | Salmonella HilA homolog | 0 | 0 | 0 | 1 (4.2) | |
| espA | Type III secretions system | 3 (12.5) | 5 (20.8) | 1 (4.2) | 1 (4.2) | |
| espB | Secreted protein B | 0 | 1 (4.2) | 0 | 0 | |
| espC | Escherichia coli enterotoxin EspC | 0 | 0 | 1 (4.2) | 0 | |
| espF | EPEC secreted protein F; Type III secretion system | 1 (4.2) | 3 (12.5) | 0 | 0 | |
| espI | Serine protease autotransporter of Enterobacteriaceae (SPATE) EspI | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| espJ | Prophage-encoded type III secretion system effector | 0 | 3 (12.5) | 1 (4.2) | 1 (4.2) | |
| espP | Extracellular serine protease plasmid-encoded | 0 | 1 (4.2) | 0 | 0 | |
| espY2 | Non-LEE-encoded type III secreted effector | 0 | 0 | 0 | 1 (4.2) | |
| etpC | EtpC Glycotransferase | 1 (4.2) | 0 | 0 | 0 | |
| etpD | Type II secretion protein EtpD | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| etsC | Putative type I secretion outer membrane protein | 1 (4.2) | 0 | 0 | 0 | |
| gad | Glutamate decarboxylase | 2 (8.3) | 4 (16.7) | 0 | 0 | |
| hha | Hemolysin expression modulator Hha (previous rmoA) | 1 (4.2) | 3 (12.5) | 0 | 1 (4.2) | |
| iss | Increased serum survival | 3 (12.5) | 5 (20.8) | 0 | 0 | |
| kpsE | Capsule polysaccharide export inner-membrane protein | 0 | 1 (4.2) | 0 | 0 | |
| mchB | Precursor of microcin H47 | 0 | 2 (8.3) | 0 | 0 | |
| mchC | MchC protein | 0 | 2 (8.3) | 0 | 0 | |
| mchF | ABC transporter protein MchF | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| mcmA | Microcin M | 0 | 2 (8.3) | 0 | 0 | |
| neuC | Polysialic acid capsule biosynthesis protein | 0 | 1 (4.2) | 0 | 0 | |
| nleB | Non-LEE encoded effector B | 3 (12.5) | 2 (8.3) | 0 | 1 (4.2) | |
| ompT | Outer membrane protease (protein protease 7) | 1 (4.2) | 3 (12.5) | 1 (4.2) | 0 | |
| ORF3 | Isoprenoid Biosynthesis | 2 (8.3) | 4 (16.7) | 0 | 0 | |
| ORF4 | Putative isopentenyl-diphosphate delta-isomerase | 2 (8.3) | 4 (16.7) | 0 | ||
| shiA | Homologue of the Shigella flexneri SHI-2 pathogenicity island gene shiA | 2 (8.3) | 1 (4.2) | 0 | 0 | |
| shiB | Homologue of the Shigella flexneri SHI-2 pathogenicity island gene shiB | 1 (4.2) | 4 (16.7) | 0 | 0 | |
| tia | Tia invasion determinant | 0 | 1 (4.2) | 0 | 0 | |
| traJ | Protein TraJ (Positive regulator of conjugal transfer operon) | 4 (16.7) | 3 (12.5) | 1 (4.2) | 0 | |
| traT | Outer membrane protein complement resistance | 3 (12.5) | 6 (25) | 0 | 0 | |
| tsh | Serine protease autotransporter of Enterobacteriaceae (SPATE)-Immunoglobulin A1 protease- Temperature-sensitive hemagglutinin | 1 (4.2) | 0 | 0 | 0 | |
| Iron uptake | aalF | CS23 Minor structural subunit | 0 | 1 (4.2) | 0 | 0 |
| chuA | Outer membrane hemin receptor | 0 | 0 | 1 (4.2) | 1 (4.2) | |
| fyuA | Yersiniabactin siderophore receptor | 2 (8.3) | 2 (8.3) | 0 | 0 | |
| ireA | Iron-regulated outer membrane protein IreA | 0 | 1 (4.2) | 0 | 0 | |
| iroN | Salmochelin siderophore receptor protein | 1 (4.2) | 0 | 0 | 0 | |
| irp2 | Yersiniabactin non-ribosomal peptide synthetase | 1 (4.2) | 2 (8.3) | 0 | 0 | |
| iucC | Aerobactin synthetase | 1 (4.2) | 4 (16.7) | 0 | 0 | |
| iutA | Ferric aerobactin receptor | 1 (4.2) | 4 (16.7) | 0 | 0 | |
| sitA | Iron transport protein | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| terC | Tellurium ion resistance protein | 7 (29.2) | 15 (62.5) | 1 (4.2) | 1 (4.2) | |
| Toxins | astA | Heat-stable enterotoxin EAST-1 | 3 (12.5) | 4 (16.7) | 0 | 0 |
| cdt-IIIB | Cytolethal distending toxin III subunit B | 0 | 1 (4.2) | 0 | 0 | |
| cnf2 | Cytotoxic necrotizing factor 2 | 0 | 1 (4.2) | 0 | 0 | |
| ehxA | Enterohaemolysin | 0 | 1 (4.2) | 0 | 0 | |
| eltIAB-8 | Heat-labile enterotoxin LTIh-8 | 0 | 1 (4.2) | 0 | 0 | |
| eltIAB-11 | Heat-labile enterotoxin LTIh-11 | 0 | 2 (8.3) | 0 | 0 | |
| estah-STa3 | Heat-stable enterotoxin STa3 human variant | 2 (8.3) | 0 | 0 | 0 | |
| hlyA | Hemolysin A | 0 | 1 (4.2) | 0 | 0 | |
| hlyE | Avian E. coli haemolysin | 6 (25) | 15 (62.5) | 0 | 1 (4.2) | |
| hlyF | Hemolysin F | 1 (4.2) | 0 | 0 | 0 | |
| pet | Autotransporter enterotoxin | 0 | 1 (4.2) | 0 | 0 | |
| pic | Serine protease autotransporter of Enterobacteriaceae (SPATE) Pic | 1 (4.2) | 1 (4.2) | 0 | 0 | |
| sat | Serine protease autotransporter of Enterobacteriaceae (SPATE) Sat | 1 (4.2) | 3 (12.5) | 0 | 0 | |
| senB | Plasmid-encoded enterotoxin | 0 | 2 (8.3) | 0 | 0 | |
| sigA | Serine protease autotransporter of Enterobacteriaceae (SPATE), Shigella IgA-like protease homologue | 0 | 2 (8.3) | 0 | 0 | |
| stx1 | Shiga toxin 1 | 0 | 2 (8.3) | 0 | 0 | |
| stx2 | Shiga toxin 2 | 0 | 1 (4.2) | 0 | 0 | |
| subA | Subtilase toxin subunit | 0 | 1 (4.2) | 0 | 0 | |
| toxB | Toxin B | 0 | 0 | 1 (4.2) | 0 | |
| vat | Serine protease autotransporter of Enterobacteriaceae (SPATE) Vat | 0 | 1 (4.2) | 0 | 0 | |
| Antimicrobial Class | Antimicrobial Resistance Gene (ARG) | Prevalence N (%) | Conferring Resistance to | Found in DEC Pathotype |
|---|---|---|---|---|
| β-lactams | blaTEM-1B | 12 (50) | amoxicillin, ampicillin, piperacillin, ticarcillin, cephalothin | EPEC, EAEC, ETEC |
| blaTEM-122 | 1 (4.2) | amoxicillin, amoxicillin + clavulanic acid, ampicillin, ampicillin + clavulanic acid, piperacillin, piperacillin + tazobactam, ticarcillin, ticarcillin + clavulanic acid | ETEC | |
| blaTEM-163 | 1 (4.2) | amoxicillin, amoxicillin + clavulanic acid, ampicillin, ampicillin + clavulanic acid, piperacillin, piperacillin + tazobactam, ticarcillin, ticarcillin + clavulanic acid | EPEC | |
| blaCTX-M-3 | 1 (4.2) | amoxicillin, ampicillin, cefepime, cefotaxime, ceftazidime, piperacillin, aztreonam, ticarcillin, ceftriaxone | EPEC | |
| blaCTX-M-15 | 2 (8.3) | amoxicillin, ampicillin, cefepime, cefotaxime, ceftazidime, piperacillin, aztreonam, ticarcillin, ceftriaxone | ETEC | |
| Sulfonamides | sul1 | 2 (8.3) | sulfamethoxazole | EPEC |
| sul2 | 8 (33.3) | sulfamethoxazole | EPEC, EAEC, ETEC | |
| Aminoglycosides | aph(6)-Id | 6 (25) | streptomycin | EPEC, ETEC |
| aph(3″)-Ib | 6 (25) | streptomycin | EPEC, ETEC | |
| aac(3)-IIa | 1 (4.2) | gentamicin, tobramycin | EPEC | |
| aadA1 | 2 (8.3) | streptomycin, spectinomycin | EAEC, ETEC | |
| aadA5 | 1 (4.2) | streptomycin, spectinomycin | EPEC | |
| aadA24 | 1 (4.2) | streptomycin, spectinomycin | EPEC | |
| Tetracyclines | tet(A) | 4 (16.7) | tetracycline, doxycycline | EPEC |
| tet(B) | 3 (12.5) | tetracycline, doxycycline, minocycline | EPEC, ETEC | |
| Quinolones | qnrS1 | 2 (8.3) | ciprofloxacin | ETEC |
| Trimethoprim | dfrA1 | 2 (8.3) | trimethoprim | EAEC, ETEC |
| dfrA7 | 1 (4.2) | trimethoprim | EPEC | |
| dfrA8 | 3 (12.5) | trimethoprim | EPEC, EAEC, ETEC | |
| dfrA14 | 1 (4.2) | trimethoprim | EPEC | |
| dfrA15 | 1 (4.2) | trimethoprim | ETEC | |
| dfrA17 | 1 (4.2) | trimethoprim | EPEC | |
| Macrolide/lincosamide/streptogramin (MLS) group | mph(A) | 6 (25) | erythromycin, azithromycin, spiramycin, telithromycin | EPEC, EAEC, ETEC |
| erm(B) | 1 (4.2) | lincomycin, clindamycin, erythromycin, quinupristin, pristinamycin IA, virginiamycin S | EAEC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wolde, D.; Eguale, T.; Medhin, G.; Haile, A.F.; Alemayehu, H.; Mihret, A.; Pirs, M.; Strašek Smrdel, K.; Avberšek, J.; Kušar, D.; et al. Diarrheagenic Escherichia coli in Stool Specimens Collected from Patients Attending Primary Healthcare Facilities in Ethiopia: Whole-Genome Sequencing-Based Molecular Characterization. Int. J. Mol. Sci. 2024, 25, 10251. https://doi.org/10.3390/ijms251910251
Wolde D, Eguale T, Medhin G, Haile AF, Alemayehu H, Mihret A, Pirs M, Strašek Smrdel K, Avberšek J, Kušar D, et al. Diarrheagenic Escherichia coli in Stool Specimens Collected from Patients Attending Primary Healthcare Facilities in Ethiopia: Whole-Genome Sequencing-Based Molecular Characterization. International Journal of Molecular Sciences. 2024; 25(19):10251. https://doi.org/10.3390/ijms251910251
Chicago/Turabian StyleWolde, Deneke, Tadesse Eguale, Girmay Medhin, Aklilu Feleke Haile, Haile Alemayehu, Adane Mihret, Mateja Pirs, Katja Strašek Smrdel, Jana Avberšek, Darja Kušar, and et al. 2024. "Diarrheagenic Escherichia coli in Stool Specimens Collected from Patients Attending Primary Healthcare Facilities in Ethiopia: Whole-Genome Sequencing-Based Molecular Characterization" International Journal of Molecular Sciences 25, no. 19: 10251. https://doi.org/10.3390/ijms251910251






