ABCG Transporters in the Adaptation of Rice to Salt Stresses
Abstract
:1. Introduction
2. Results
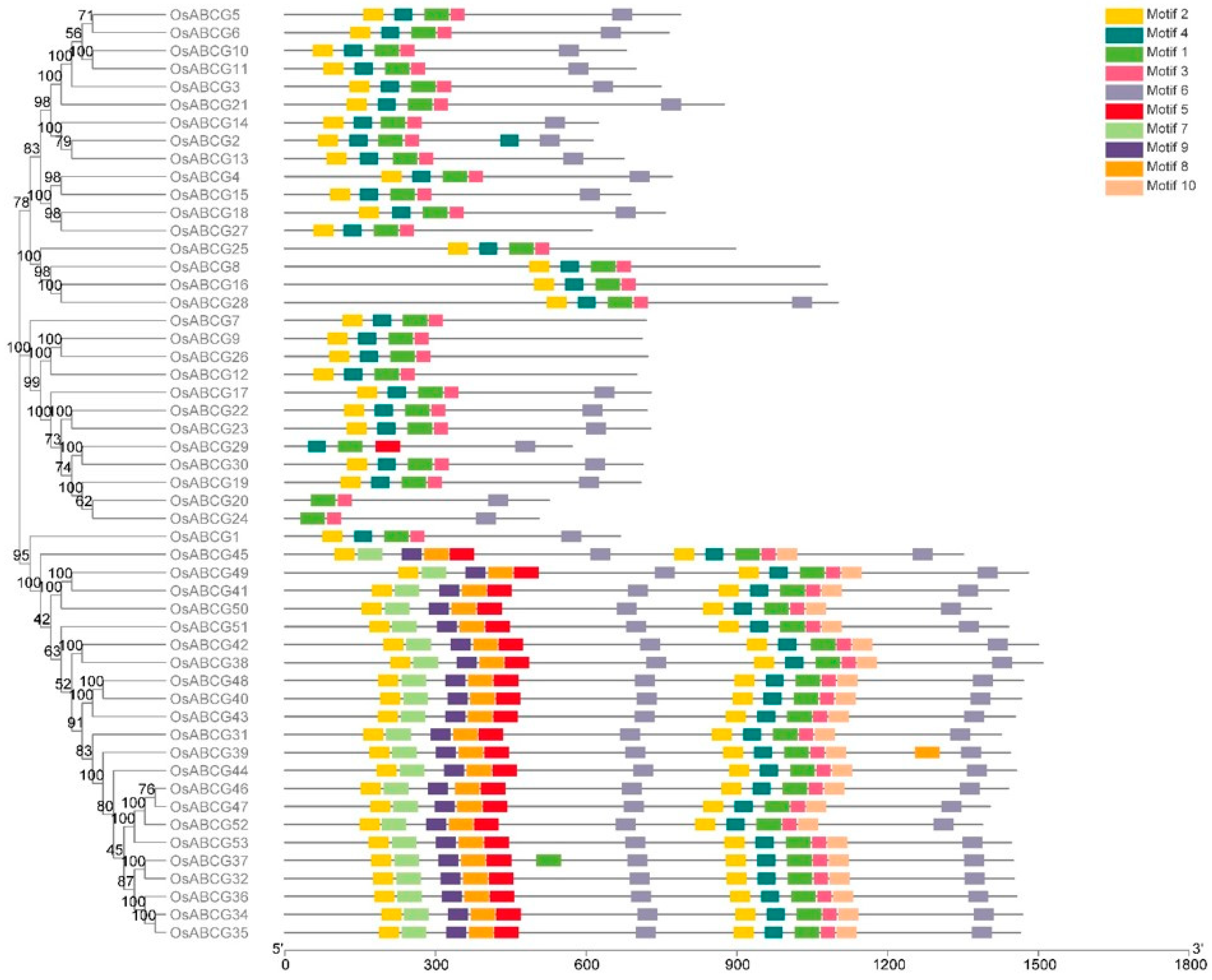
2.1. Identification of Rice OsABCGs, Phylogenetic and Evolutionary Analysis
2.2. Gene Structure Analysis of ABCGs
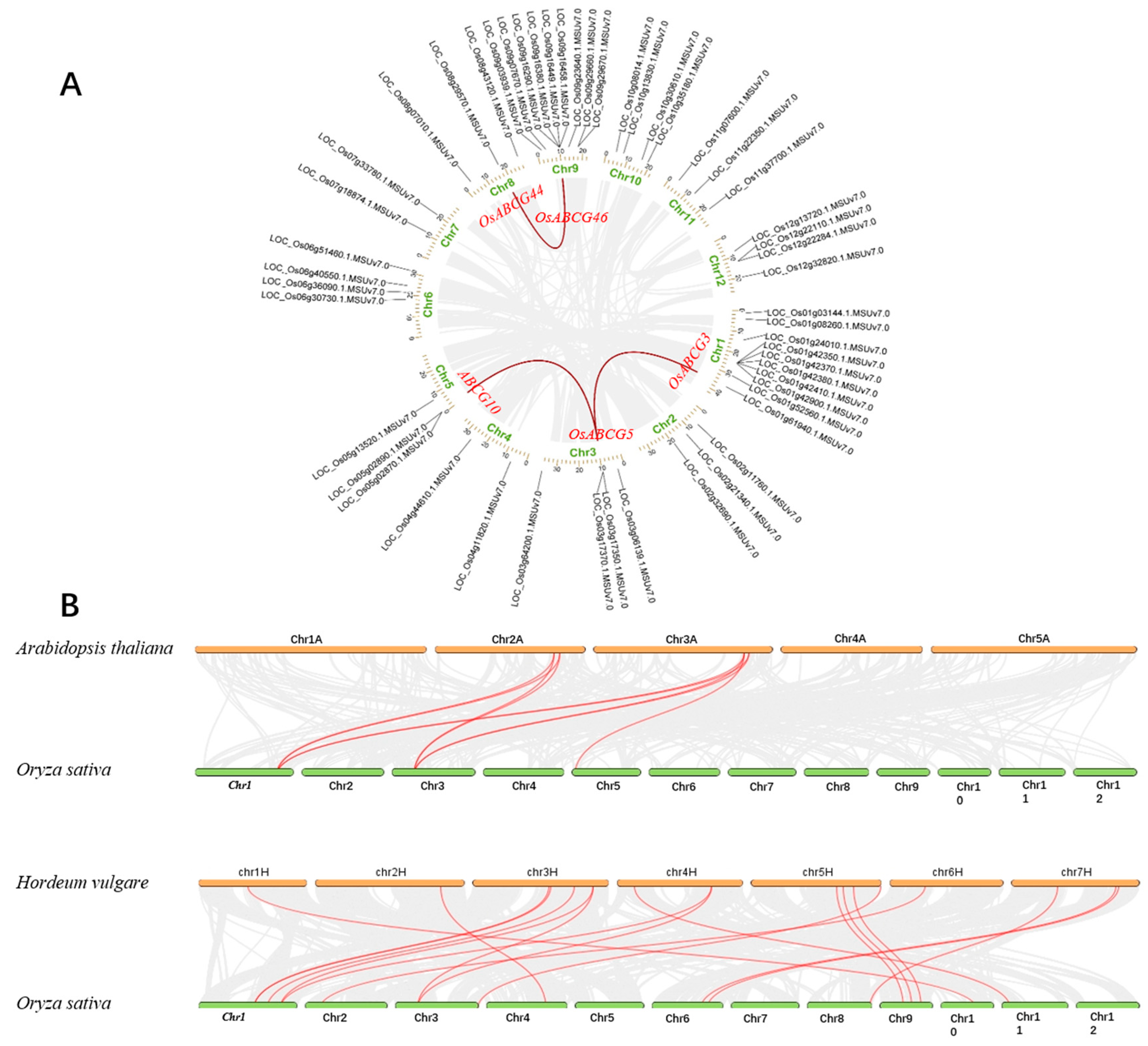
2.3. Chromosomal Location and Gene Duplication Analysis of OsABCGs
2.4. Analysis of ABCGs Promoter
2.5. Expression Pattern Analysis of Rice ABCG Family Members under Salt Stress
2.6. Perform Salt Tolerance Screening in Yeast
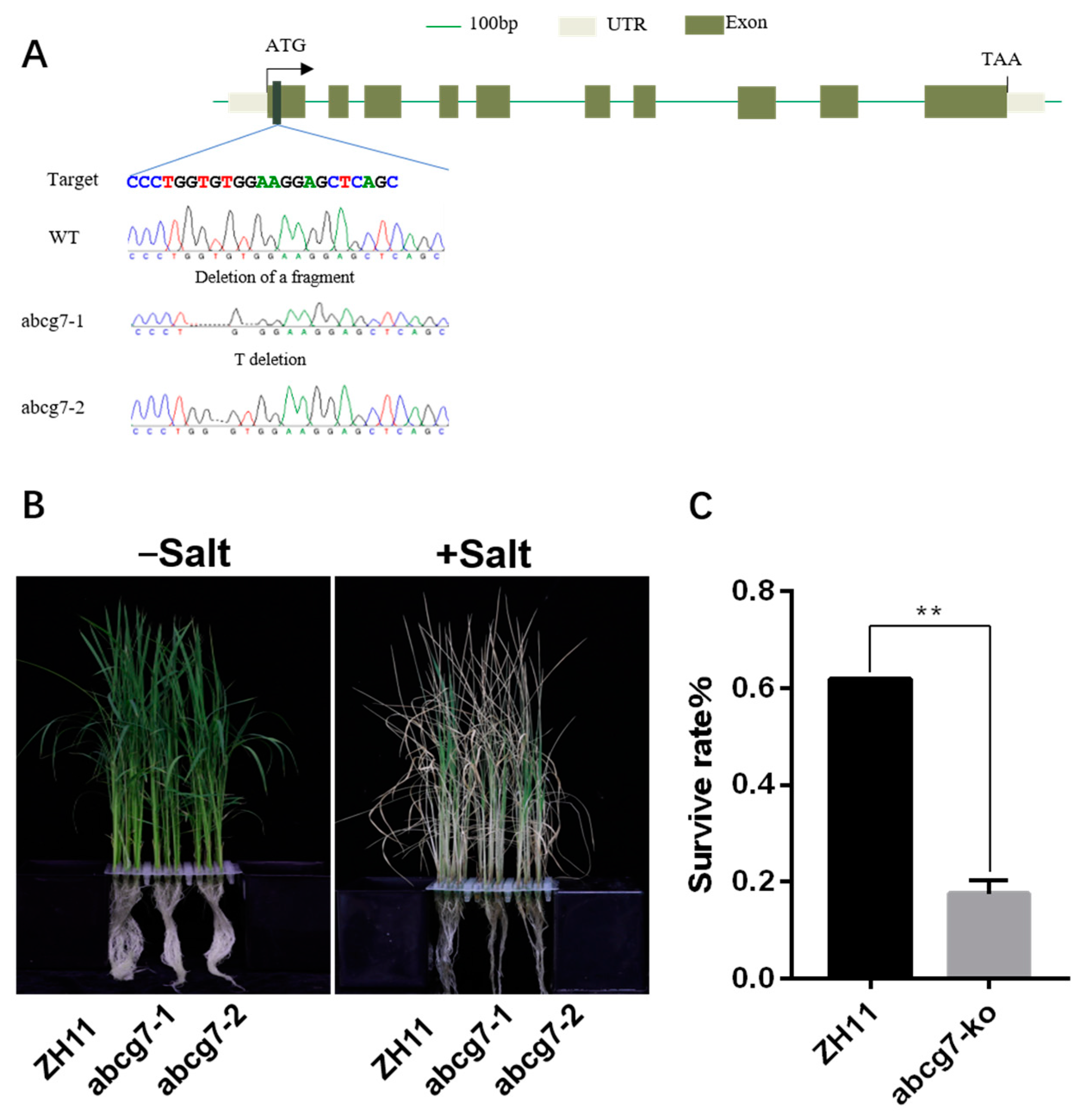
2.7. Mutations of OsABCG7 Exhibits a Salt-Sensitive Phenotype
3. Discussion
4. Materials and Methods
4.1. Identification of Rice OsABCGs Phylogenetic and Evolutionary Analysis
4.2. Analysis of Conserved Protein Motifs of OsABCGs
4.3. Analysis of Chromosomal Location and Gene Duplication
4.4. Analysis of Cis-Elements of OsABCG Genes
4.5. RNA Isolation and Quantitative Real-Time PCR Analysis under Salt Stress
4.6. Transport Assays in Yeast
4.7. Plant Materials and Treatments
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Raghuvanshi, R.; Srivastava, A.K.; Verulkar, S.; Suprasanna, P. Unlocking Allelic Diversity for Sustainable Development of Salinity Stress Tolerance in Rice. Curr. Genom. 2021, 22, 393–403. [Google Scholar] [CrossRef] [PubMed]
- Pardo, J.M. Biotechnology of Water and Salinity Stress Tolerance. Curr. Opin. Biotechnol. 2010, 21, 185–196. [Google Scholar] [CrossRef]
- Zhu, J.K. Regulation of Ion Homeostasis under Salt Stress. Curr. Opin. Plant Biol. 2003, 6, 441–445. [Google Scholar] [CrossRef]
- Chen, T.; Shabala, S.; Niu, Y.; Chen, Z.-H.; Shabala, L.; Meinke, H.; Venkataraman, G.; Pareek, A.; Xu, J.; Zhou, M. Molecular Mechanisms of Salinity Tolerance in Rice. Crop J. 2021, 9, 506–520. [Google Scholar] [CrossRef]
- Parida, A.K.; Das, A.B. Salt Tolerance and Salinity Effects on Plants: A Review. Ecotoxicol. Environ. Saf. 2005, 60, 324–349. [Google Scholar] [CrossRef]
- Wen, J.; Zhang, W. Research Progress on Gene Mapping and Cloning for Salt Tolerance and Variety Improvement for Salt Tolerance by Molecular Marker-Assisted Selection in Rice. Chin. J. Rice Sci. 2017, 31, 111–123. [Google Scholar] [CrossRef]
- Yao, D.; Wu, J.; Hu, Z.; Bai, B.; Zhuang, W.; Li, J.; Deng, Q. Physiological Mechanism and Breeding Strat-Egy of Rice Saline-Alkaline Tolerance. Hybrid. Rice 2019, 4, 1–7. [Google Scholar] [CrossRef]
- Wang, C.; Zhang, Y.; Zhao, L.; Chen, T.; Zhao, Q.; Yao, S.; Zhou, L.; Zhao, C.; Liang, W.; Sun, M.; et al. Research Status, Problems and Suggestions on Salt-Alkali Tolerant Rice. China Rice 2019, 25, 1–6. [Google Scholar]
- Qin, H.; Li, Y.; Huang, R. Advances and Challenges in the Breeding of Salt-Tolerant Rice. Int. J. Mol. Sci. 2020, 21, 8385. [Google Scholar] [CrossRef]
- Wang, G.; Ni, G.; Feng, G.; Burrill, M.; Li, J.; Zhang, J.; Zhang, F. Saline-Alkali Soil Reclamation and Utilization in China: Progress and Prospects. Front. Agric. Sci. Eng. 2024, 11, 216–228. [Google Scholar] [CrossRef]
- Zhang, B.; Liu, Z.; Zhou, R.; Cheng, P.; Li, H.; Wang, Z.; Liu, Y.; Li, M.; Zhao, Z.; Hu, Z.; et al. Genome-wide analysis of soybean DnaJA-family genes and functional characterization of GmDnaJA6 responses to saline and alkaline stress. Crop J. 2023, 11, 1230–1241. [Google Scholar] [CrossRef]
- Lu, M.; Chen, Z.; Dang, Y.; Li, J.; Wang, J.; Zheng, H.; Li, S.; Wang, X.; Du, X.; Sui, N. Identification of the MYB Gene Family in Sorghum Bicolor and Functional Analysis of SbMYBAS1 in Response to Salt Stress. Plant Mol. Biol. 2023, 113, 249–264. [Google Scholar] [CrossRef]
- Kang, J.; Park, J.; Choi, H.; Burla, B.; Kretzschmar, T.; Lee, Y.; Martinoia, E.; Angus, M.; Philip, A.R.; Lacey, S.; et al. Plant ABC Transporters. Arab. Book 2011, 9, e0153. [Google Scholar] [CrossRef] [PubMed]
- Verrier, P.J.; Bird, D.; Burla, B.; Dassa, E.; Forestier, C.; Geisler, M.; Klein, M.; Kolukisaoglu, Ü.; Lee, Y.; Martinoia, E. Plant ABC Proteins—A Unified Nomenclature and Updated Inventory. Trends Plant Sci. 2008, 13, 151–159. [Google Scholar] [CrossRef]
- Matsuda, S.; Funabiki, A.; Furukawa, K.; Komori, N.; Koike, M.; Tokuji, Y.; Takamure, I.; Kato, K. Genome–Wide Analysis and Expression Profiling of Half–Size ABC Protein Subgroup G in Rice in Response to Abiotic Stress and Phytohormone Treatments. Mol. Genet. Genom. 2012, 287, 819–835. [Google Scholar] [CrossRef]
- Niu, L.; Li, H.; Song, Z.; Dong, B.; Cao, H.; Liu, T.; Du, T.; Yang, W.; Amin, R.; Wang, L.; et al. The Functional Analysis of ABCG Transporters in the Adaptation of Pigeon Pea (Cajanus Cajan) to Abiotic Stresses. PeerJ 2021, 9, e10688. [Google Scholar] [CrossRef] [PubMed]
- Jasinski, M.; Ducos, E.; Martinoia, E.; Boutry, M. The ATP-Binding Cassette Transporters: Structure, Function, and Gene Family Comparison between Rice and Arabidopsis. Plant Physiol. 2003, 131, 1169–1177. [Google Scholar] [CrossRef]
- Wu, S.; Fang, C.; Li, Z.; Wang, Y.; Pan, S.; Wu, Y.; An, X.; Long, Y.; Wan, X. ATP-Binding Cassette G Transporters and Their Multiple Roles Especially for Male Fertility in Arabidopsis, Rice and Maize. Int. J. Mol. Sci. 2022, 23, 9304. [Google Scholar] [CrossRef]
- Gupta, B.B.; Selter, L.L.; Baranwal, V.K.; Arora, D.; Mishra, S.K.; Sirohi, P.; Poonia, A.K.; Chaudhary, R.; Kumar, R.; Krattinger, S.G.; et al. Updated Inventory, Evolutionary and Expression Analyses of G (PDR) Type ABC Transporter Genes of Rice. Plant Physiol. Biochem. 2019, 142, 429–439. [Google Scholar] [CrossRef]
- Zhang, Y.; Kilambi, H.V.; Liu, J.; Bar, H.; Lazary, S.; Egbaria, A.; Ripper, D.; Charrier, L.; Belew, Z.M.; Wulff, N.; et al. ABA Homeostasis and Long-Distance Translocation Are Redundantly Regulated by ABCG ABA Importers. Sci. Adv. 2021, 7, eabf6069. [Google Scholar] [CrossRef]
- Ashraf, M.A.; Akihiro, T.; Ito, K.; Kumagai, S.; Sugita, R.; Tanoi, K.; Rahman, A. ATP Binding Cassette Proteins ABCG37 and ABCG33 Function as Potassium-Independent Cesium Uptake Carriers in Arabidopsis Roots. Mol. Plant 2021, 14, 664–678. [Google Scholar] [CrossRef]
- Li, H.; Li, C.; Sun, D.; Yang, Z.M. OsPDR20 Is an ABCG Metal Transporter Regulating Cadmium Accumulation in Rice. J. Environ. Sci. 2024, 136, 21–34. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Deng, X.; Qian, J.; Liu, T.; Ju, M.; Li, J.; Yang, Q.; Zhu, X.; Li, W.; Liu, C.; et al. Arabidopsis ABCG14 forms a homodimeric transporter for multiple cytokinins and mediates long-distance transport of isopentenyladenine-type cytokinins. Plant Commun. 2023, 4, 100468. [Google Scholar] [CrossRef] [PubMed]
- Nawkar, G.M.; Legris, M.; Goyal, A.; Schmid-sieger, E.; Fleury, J.; Mucciolo, A.; Bellis, D.D.; Trevisan, M.; Schueler, A.; Fankhauser, C. Air Channels Create a Directional Light Signal to Regulate Hypocotyl Phototropism. Science 2023, 382, 935–940. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, Y.; Zhang, D.; Liang, J. Endomembrane-Biased Dimerization of ABCG16 and ABCG25 Transporters Determines Their Substrate Selectivity in ABA-Regulated Plant Growth and Stress Responses. Mol. Plant 2024, 17, 478–495. [Google Scholar] [CrossRef]
- Moons, A. Transcriptional Profiling of the PDR Gene Family in Rice Roots in Response to Plant Growth Regulators, Redox Perturbations and Weak Organic Acid Stresses. Planta 2008, 229, 53–71. [Google Scholar] [CrossRef]
- Guo, Z.; Yuan, X.; Li, L.; Zeng, M.; Yang, J.; Tang, H.; Duan, C. Genome-Wide Analysis of the ATP-Binding Cassette (ABC) Transporter Family in Zea Mays, L. and Its Response to Heavy Metal Stresses. Int. J. Mol. Sci. 2022, 23, 2109. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.-U.; Song, W.-Y.; Hong, D.; Ko, D.; Yamaoka, Y.; Jang, S.; Yim, S.; Lee, E.; Khare, D.; Kim, K.; et al. Plant ABC Transporters Enable Many Unique Aspects of a Terrestrial Plant’s Lifestyle. Mol. Plant 2016, 9, 338–355. [Google Scholar] [CrossRef]
- Zhang, Z.; Tong, T.; Fang, Y.; Zheng, J.; Zhang, X.; Niu, C.; Li, J.; Zhang, X.; Xue, D. Genome-Wide Identification of Barley ABC Genes and Their Expression in Response to Abiotic Stress Treatment. Plants 2020, 9, 1281. [Google Scholar] [CrossRef]
- Taylor, J.S.; Raes, J. Duplication and Divergence: The Evolution of New Genes and Old Ideas. Annu. Rev. Genet. 2004, 38, 615–643. [Google Scholar] [CrossRef]
- Koonin, E.V. Orthologs, Paralogs, and Evolutionary Genomics. Annu. Rev. Genet. 2024, 39, 309–338. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; Tester, M. Mechanisms of Salinity Tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef]
- Li, J.; Han, G.; Sun, C.; Sui, N. Research advances of MYB transcription factors in plant stress resistance and breeding. Plant Signal Behav. 2019, 14, 1613131. [Google Scholar] [CrossRef]
- Fan, Y.; Zhang, Y.; Rui, C.; Xu, N.; Zhang, H.; Wang, J.; Malik, W.; Han, M.; Zhao, L.; Lu, X.; et al. Zinc Finger Transcription Factor ZAT Family Genes Confer Multi-Tolerances in Gossypium hirsutum L. J. Cotton Res. 2021, 4, 24. [Google Scholar] [CrossRef]
- Hussain, S.; Zhang, R.; Liu, S.; Li, R.; Wang, Y.; Chen, Y.; Hou, H.; Dai, Q. Methyl Jasmonate Alleviates the Deleterious Effects of Salinity Stress by Augmenting Antioxidant Enzyme Activity and Ion Homeostasis in Rice (Oryza sativa L.). Agronomy 2022, 12, 2343. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA 11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Zhao, W.; Zheng, S.; Ling, H.Q. An efficient regeneration system and Agrobacterium-mediated transformation of Chinese upland rice cultivar Handao297. Plant Cell Tiss. Organ. Cult. 2011, 106, 475–483. [Google Scholar] [CrossRef]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, D.; Hu, Y.; Tang, L.; Du, Y.; Mao, R.; Sheng, X.; Liu, H.; Liu, X.; Zhao, B.; Lei, D. ABCG Transporters in the Adaptation of Rice to Salt Stresses. Int. J. Mol. Sci. 2024, 25, 10724. https://doi.org/10.3390/ijms251910724
Zhang D, Hu Y, Tang L, Du Y, Mao R, Sheng X, Liu H, Liu X, Zhao B, Lei D. ABCG Transporters in the Adaptation of Rice to Salt Stresses. International Journal of Molecular Sciences. 2024; 25(19):10724. https://doi.org/10.3390/ijms251910724
Chicago/Turabian StyleZhang, Dan, Yuanyi Hu, Li Tang, Yaxi Du, Ruihua Mao, Xiabing Sheng, Huimin Liu, Xiaolin Liu, Bingran Zhao, and Dongyang Lei. 2024. "ABCG Transporters in the Adaptation of Rice to Salt Stresses" International Journal of Molecular Sciences 25, no. 19: 10724. https://doi.org/10.3390/ijms251910724




