Y-Box-Binding Proteins Have a Dual Impact on Cellular Translation
Abstract
:1. Introduction
2. Results
2.1. YB-1 and YB-3 Expressions Partially Restore Translation in HEK293TΔYB-1ΔYB-3 Cells
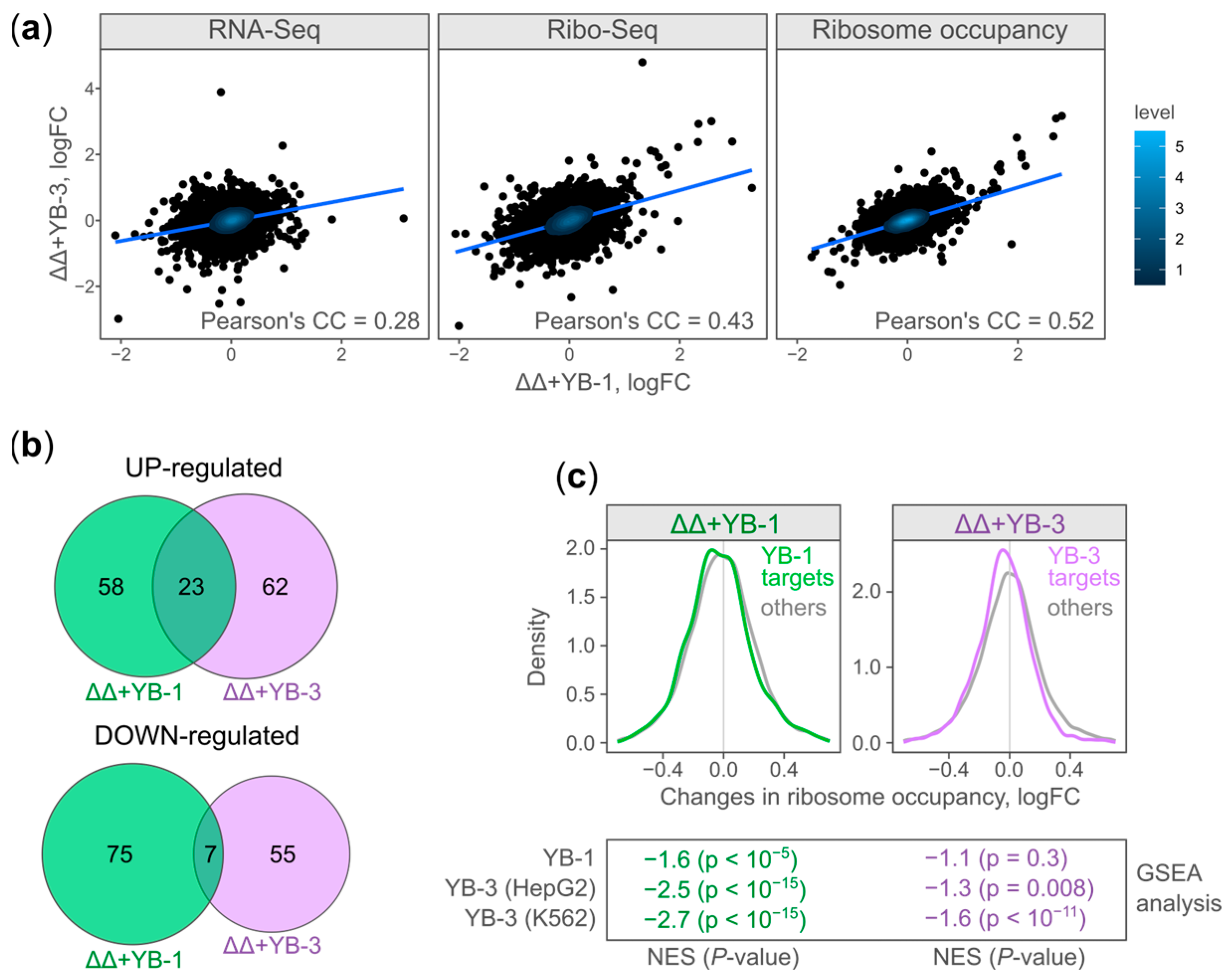
2.2. Translational Changes Correlate in YB-1- and YB-3-Expressing Cells
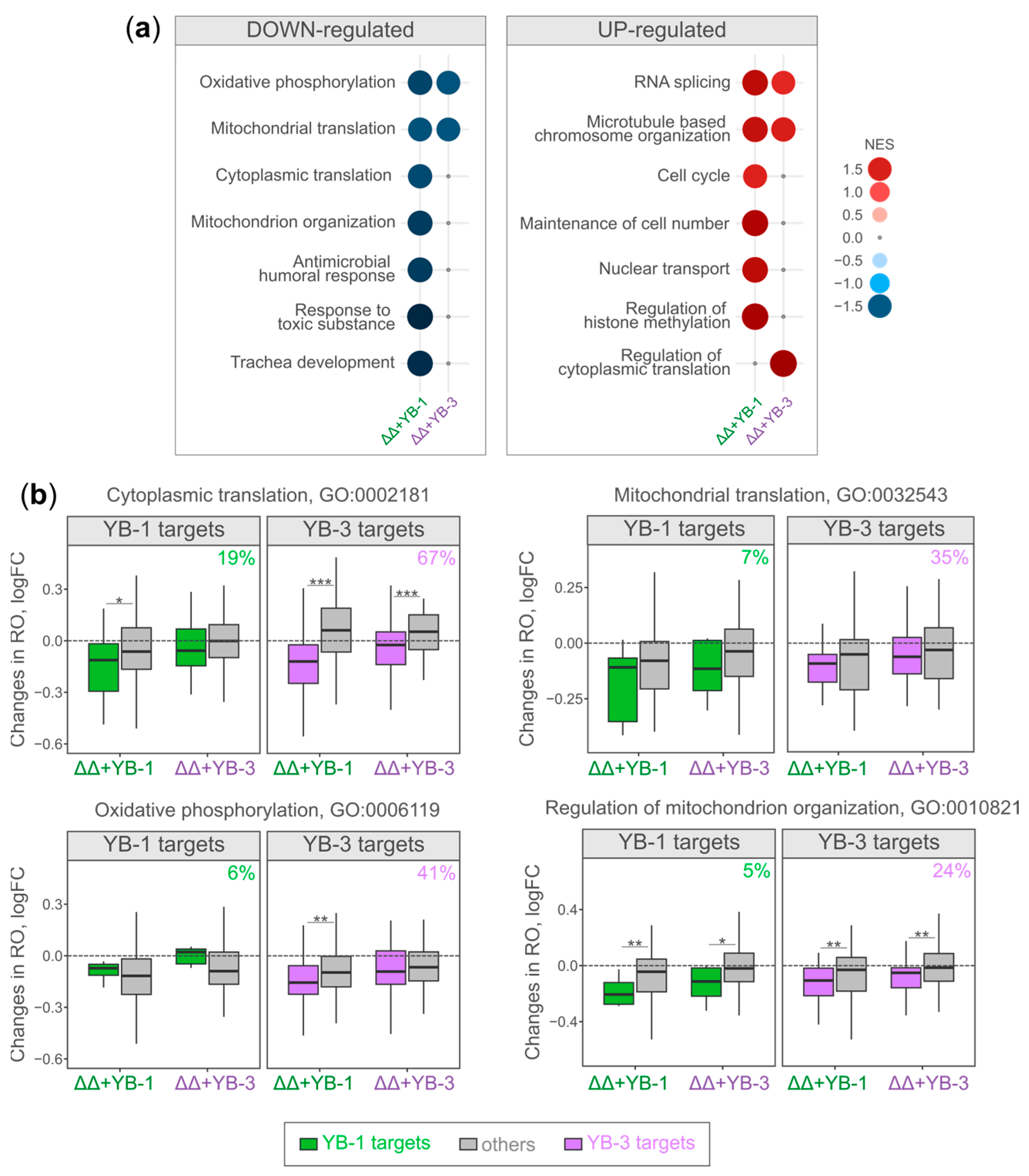
2.3. Translationally Altered Genes in YB-1- and YB-3-Expressing Cells Participate in Similar Biological Processes
3. Discussion
4. Materials and Methods
4.1. Plasmids
4.2. Cell Cultures
4.3. Western Blot and Antibodies
4.4. Sucrose Gradient
4.5. Ribo-Seq and RNA-Seq
4.6. PAR-CLIP
4.7. High-Throughput Sequencing and Data Processing
4.8. PAR-CLIP and eCLIP Data Processing
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mordovkina, D.; Lyabin, D.N.; Smolin, E.A.; Sogorina, E.M.; Ovchinnikov, L.P.; Eliseeva, I. Y-Box Binding Proteins in MRNP Assembly, Translation, and Stability Control. Biomolecules 2020, 10, 591. [Google Scholar] [CrossRef] [PubMed]
- Kleene, K.C. Y-Box Proteins Combine Versatile Cold Shock Domains and Arginine-Rich Motifs (ARMs) for Pleiotropic Functions in RNA Biology. Biochem. J. 2018, 475, 2769–2784. [Google Scholar] [CrossRef]
- Eliseeva, I.A.; Kim, E.R.; Guryanov, S.G.; Ovchinnikov, L.P.; Lyabin, D.N. Y-Box-Binding Protein 1 (YB-1) and Its Functions. Biochemistry 2011, 76, 1402–1433. [Google Scholar] [CrossRef]
- Lindquist, J.A.; Mertens, P.R. Cold Shock Proteins: From Cellular Mechanisms to Pathophysiology and Disease. Cell Commun. Signal. 2018, 16, 63. [Google Scholar] [CrossRef] [PubMed]
- Yin, Q.; Zheng, M.; Luo, Q.; Jiang, D.; Zhang, H.; Chen, C. YB-1 as an Oncoprotein: Functions, Regulation, Post-Translational Modifications, and Targeted Therapy. Cells 2022, 11, 1217. [Google Scholar] [CrossRef]
- Rybalkina, E.Y.; Moiseeva, N.I. Role of YB-1 Protein in Inflammation. Biochem. 2022, 87, S94–S102. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.-J.; Zhu, H.; Mu, S.-R.; Wei, W.-J.; Yuan, X.; Wang, M.; Liu, Y.; Hui, J.; Huang, Y. Crystal Structure of a Y-Box Binding Protein 1 (YB-1)-RNA Complex Reveals Key Features and Residues Interacting with RNA. J. Biol. Chem. 2019, 294, 10998–11010. [Google Scholar] [CrossRef]
- Philippe, B.; Matsumoto, K.; Wolffe, A.P. Sequence-Specific RNA Recognition by the Xenopus Y-Box Proteins. J. Biol. Chem. 1995, 270, 28297–28303. [Google Scholar] [CrossRef] [PubMed]
- Eliseeva, I.A.; Sogorina, E.M.; Smolin, E.A.; Kulakovskiy, I.V.; Lyabin, D.N. Diverse Regulation of YB-1 and YB-3 Abundance in Mammals. Biochemistry 2022, 87, S48–S70. [Google Scholar] [CrossRef]
- Lyabin, D.N.; Eliseeva, I.A.; Smolin, E.A.; Doronin, A.N.; Budkina, K.S.; Kulakovskiy, I.V.; Ovchinnikov, L.P. YB-3 Substitutes YB-1 in Global mRNA Binding. RNA Biol. 2020, 17, 487–499. [Google Scholar] [CrossRef]
- Lyabin, D.N.; Smolin, E.A.; Budkina, K.S.; Eliseeva, I.A.; Ovchinnikov, L.P. Towards the Mechanism(s) of YB-3 Synthesis Regulation by YB-1. RNA Biol. 2021, 18, 1630–1641. [Google Scholar] [CrossRef]
- Van Nostrand, E.L.; Pratt, G.A.; Shishkin, A.A.; Gelboin-Burkhart, C.; Fang, M.Y.; Sundararaman, B.; Blue, S.M.; Nguyen, T.B.; Surka, C.; Elkins, K.; et al. Robust Transcriptome-Wide Discovery of RNA-Binding Protein Binding Sites with Enhanced CLIP (ECLIP). Nat. Methods 2016, 13, 508–514. [Google Scholar] [CrossRef]
- Minich, W.B.; Ovchinnikov, L.P. Role of Cytoplasmic MRNP Proteins in Translation. Biochimie 1992, 74, 477–483. [Google Scholar] [CrossRef]
- Ro, S.-H.; Fay, J.; Cyuzuzo, C.I.; Jang, Y.; Lee, N.; Song, H.-S.; Harris, E.N. SESTRINs: Emerging Dynamic Stress-Sensors in Metabolic and Environmental Health. Front. Cell Dev. Biol. 2020, 8, 603421. [Google Scholar] [CrossRef]
- Budanov, A.V.; Karin, M. P53 Target Genes Sestrin1 and Sestrin2 Connect Genotoxic Stress and MTOR Signaling. Cell 2008, 134, 451–460. [Google Scholar] [CrossRef]
- Pause, A.; Belsham, G.J.; Gingras, A.-C.; Donzé, O.; Lin, T.-A.; Lawrence, J.C.; Sonenberg, N. Insulin-Dependent Stimulation of Protein Synthesis by Phosphorylation of a Regulator of 5′-Cap Function. Nature 1994, 371, 762–767. [Google Scholar] [CrossRef] [PubMed]
- Merrick, W.C. EIF4F: A Retrospective. J. Biol. Chem. 2015, 290, 24091–24099. [Google Scholar] [CrossRef]
- Ji, Z.; Song, R.; Huang, H.; Regev, A.; Struhl, K. Transcriptome-Scale RNase-Footprinting of RNA-Protein Complexes. Nat. Biotechnol. 2016, 34, 410–413. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Liu, Y.; Gerloff, D.; Rohde, C.; Pauli, C.; Köhn, M.; Misiak, D.; Oellerich, T.; Schwartz, S.; Schmidt, L.-H.; et al. NOP10 Predicts Lung Cancer Prognosis and Its Associated Small Nucleolar RNAs Drive Proliferation and Migration. Oncogene 2021, 40, 909–921. [Google Scholar] [CrossRef]
- Sloan, K.E.; Leisegang, M.S.; Doebele, C.; Ramírez, A.S.; Simm, S.; Safferthal, C.; Kretschmer, J.; Schorge, T.; Markoutsa, S.; Haag, S.; et al. The Association of Late-Acting SnoRNPs with Human Pre-Ribosomal Complexes Requires the RNA Helicase DDX21. Nucleic Acids Res. 2015, 43, 553–564. [Google Scholar] [CrossRef] [PubMed]
- Burke, M.F.; Logan, M.K.; Hebert, M.D. Identification of Additional Regulatory RNPs That Impact RRNA and U6 SnRNA Methylation. Biol. Open 2018, 7, bio036095. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.H.; Books, J.T.; Ley, T.J. YB-1 Is Important for Late-Stage Embryonic Development, Optimal Cellular Stress Responses, and the Prevention of Premature Senescence. Mol. Cell. Biol. 2005, 25, 4625–4637. [Google Scholar] [CrossRef]
- Lu, Z.H.; Books, J.T.; Ley, T.J. Cold Shock Domain Family Members YB-1 and MSY4 Share Essential Functions during Murine Embryogenesis. Mol. Cell. Biol. 2006, 26, 8410–8417. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome Engineering Using the CRISPR-Cas9 System. Nat. Protoc. 2013, 8, 2281–2308. [Google Scholar] [CrossRef] [PubMed]
- Gerashchenko, M.V.; Gladyshev, V.N. Translation Inhibitors Cause Abnormalities in Ribosome Profiling Experiments. Nucleic Acids Res. 2014, 42, e134. [Google Scholar] [CrossRef] [PubMed]
- McGlincy, N.J.; Ingolia, N.T. Transcriptome-Wide Measurement of Translation by Ribosome Profiling. Methods 2017, 126, 112–129. [Google Scholar] [CrossRef]
- Danan, C.; Manickavel, S.; Hafner, M. PAR-CLIP: A Method for Transcriptome-Wide Identification of RNA Binding Protein Interaction Sites. In Post-Transcriptional Gene Regulation; Humana Press: New York, NY, USA, 2016; pp. 153–173. [Google Scholar]
- Lyabin, D.N.; Eliseeva, I.A.; Ovchinnikov, L.P. YB-1 Synthesis Is Regulated by MTOR Signaling Pathway. PLoS ONE 2012, 7, e52527. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet J. 2011, 17, 10. [Google Scholar] [CrossRef]
- Shen, W.; Le, S.; Li, Y.; Hu, F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS ONE 2016, 11, e0163962. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Dunn, J.G.; Weissman, J.S. Plastid: Nucleotide-Resolution Analysis of next-Generation Sequencing and Genomics Data. BMC Genom. 2016, 17, 958. [Google Scholar] [CrossRef]
- Zhang, Y.; Parmigiani, G.; Johnson, W.E. ComBat-Seq: Batch Effect Adjustment for RNA-Seq Count Data. NAR Genom. Bioinform. 2020, 2, lqaa078. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Korotkevich, G.; Sukhov, V.; Budin, N.; Shpak, B.; Artyomov, M.N.; Sergushichev, A. Fast Gene Set Enrichment Analysis. bioRxiv 2021, 060012. [Google Scholar] [CrossRef]
- Liberzon, A.; Subramanian, A.; Pinchback, R.; Thorvaldsdóttir, H.; Tamayo, P.; Mesirov, J.P. Molecular Signatures Database (MSigDB) 3.0. Bioinformatics 2011, 27, 1739–1740. [Google Scholar] [CrossRef]
- Gu, Z.; Hübschmann, D. SimplifyEnrichment: A Bioconductor Package for Clustering and Visualizing Functional Enrichment Results. Genom. Proteom. Bioinform. 2023, 21, 190–202. [Google Scholar] [CrossRef] [PubMed]
- Sievers, C.; Schlumpf, T.; Sawarkar, R.; Comoglio, F.; Paro, R. Mixture Models and Wavelet Transforms Reveal High Confidence RNA-Protein Interaction Sites in MOV10 PAR-CLIP Data. Nucleic Acids Res. 2012, 40, e160. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and Memory-Efficient Alignment of Short DNA Sequences to the Human Genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Comoglio, F.; Sievers, C.; Paro, R. Sensitive and Highly Resolved Identification of RNA-Protein Interaction Sites in PAR-CLIP Data. BMC Bioinform. 2015, 16, 32. [Google Scholar] [CrossRef]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. Gigascience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: A Flexible Suite of Utilities for Comparing Genomic Features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, S.; Kulakovskiy, I.V.; Severin, J.; Koido, M.; Alam, T.; Abugessaisa, I.; Buyan, A.; Chang, H.Y.; Dostie, J.; Itoh, M.; et al. Systematic Identification of Cis-Interacting LncRNAs and Their Targets. bioRxiv 2021. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef] [PubMed]
- Github. Available online: https://github.com/YeoLab/eclip/tree/master/bin (accessed on 10 January 2020).
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Eliseeva, I.A.; Buyan, A.I.; Smolin, E.A.; Kaliadzenka, K.S.; Popov, S.; Kulakovskiy, I.V.; Lyabin, D.N. Y-Box-Binding Proteins Have a Dual Impact on Cellular Translation. Int. J. Mol. Sci. 2024, 25, 1736. https://doi.org/10.3390/ijms25031736
Eliseeva IA, Buyan AI, Smolin EA, Kaliadzenka KS, Popov S, Kulakovskiy IV, Lyabin DN. Y-Box-Binding Proteins Have a Dual Impact on Cellular Translation. International Journal of Molecular Sciences. 2024; 25(3):1736. https://doi.org/10.3390/ijms25031736
Chicago/Turabian StyleEliseeva, Irina A., Andrey I. Buyan, Egor A. Smolin, Karina S. Kaliadzenka, Sergey Popov, Ivan V. Kulakovskiy, and Dmitry N. Lyabin. 2024. "Y-Box-Binding Proteins Have a Dual Impact on Cellular Translation" International Journal of Molecular Sciences 25, no. 3: 1736. https://doi.org/10.3390/ijms25031736




