Abstract
Infectious diseases have been treated using plants and their compounds for thousands of years. This knowledge has enabled modern techniques to identify specific antiviral remedies and to understand their molecular mechanism of action. Numerous active phytochemicals, such as alkaloids, terpenoids, polyphenols (phenolic acids, flavonoids, stilbenes, and lignans), coumarins, thiophenes, saponins, furyl compounds, small proteins, and peptides, are promising options for treating and preventing viral infections. It has been shown that plant-derived products can prevent or inhibit viral entry into and replication by host cells. Biotechnological advances have made it possible to engineer plants with an increased capacity for the production and accumulation of natural antiviral compounds. Plants can also be engineered to produce various types of antivirals (cytokines, antibodies, vaccines, and lectins). This study summarizes the current understanding of the antiviral activity of specific plant-derived metabolites, emphasizing their mechanisms of action and exploring the enormous potential of plants as biological factories.
1. Introduction
Emerging viral infections continue to be a significant global health concern. Phytochemicals with antiviral properties, together with advances in plant molecular farming, position plants as both sources of therapeutic compounds and platforms for producing valuable pharmaceutical proteins for viral prevention and treatment. Here, we highlight the potential of plant-based approaches in addressing current and future viral threats.
1.1. The Persisting Era of Infectious Disease
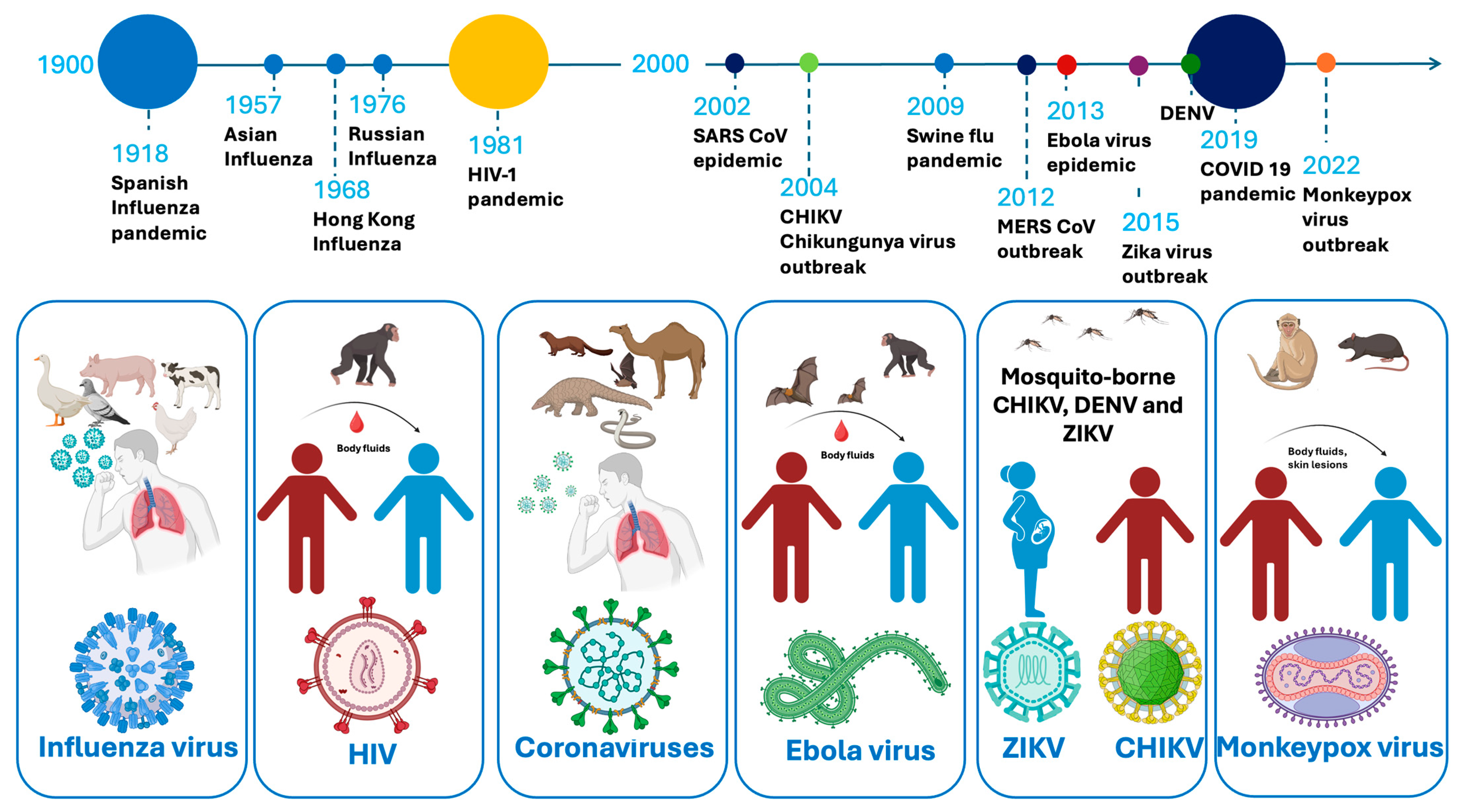
A substantial number of severe illnesses are caused by emerging and reemerging viruses, which have become increasingly prevalent due to factors such as population growth, global travel, and climate change [1,2,3]. Over the past 40 years, multiple major outbreaks have demonstrated the unpredic and evolving nature of viral epidemics. These include the 1981 human immunodeficiency virus-1 (HIV-1) pandemic, the 2002 Severe Acute Respiratory Syndrome coronavirus (SARS-CoV) outbreak, and the 2009 influenza A H1N1 (swine flu) pandemic. In 2012, the Middle East Respiratory Syndrome coronavirus (MERS-CoV) emerged, followed by the 2013 Ebola virus (EBOV) epidemic in West Africa. The 2015 Zika virus (ZIKV) outbreak in the Americas and the 2019 dengue fever virus (DENV) epidemic further underscored the global vulnerability to arboviruses. The emergence of SARS-CoV-2 in 2019 rapidly escalated into a worldwide pandemic. More recently, the 2022 Monkeypox virus (MPXV) outbreak and the 2024 resurgence of the chikungunya virus (CHIKV) in La Réunion, following its major outbreak in 2004, emphasize the ongoing challenge posed by viral epidemics (Figure 1) [4].
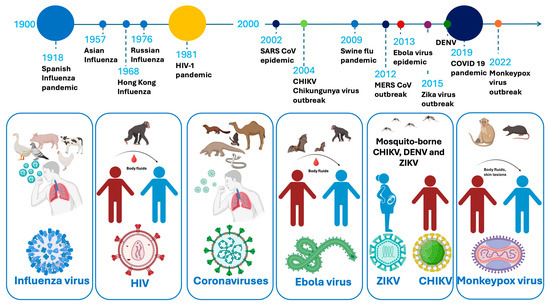
Figure 1.
A timeline of viral epidemics and pandemics from 1900 to the present, the viral host, viral vectors, and transmission methods.
However, the mortality and morbidity associated with infectious diseases have decreased over the past forty years due to the swift development of vaccines, improvements in sanitation, access to healthcare, and medical advancements [5]. New and improved biotechnologies, such as the development of mRNA vaccines and recombinant virus-like particles (VLPs) as vaccine and delivery vesicles, and the use of new expression systems (mammalian, insect, and plant cells, and whole plants), offer new approaches for rapid vaccine preparation, especially against the emergence of new viruses [6,7]. Controlling the spread of (re)emerging viruses may also depend on using robust antivirals. Traditional plant-derived remedies and plant biotechnology products can be a source of affordable new antiviral agents for anti-inflammatory and immunomodulatory therapy that can support the battle against (re)emerging viruses [8,9].
1.2. Antivirals
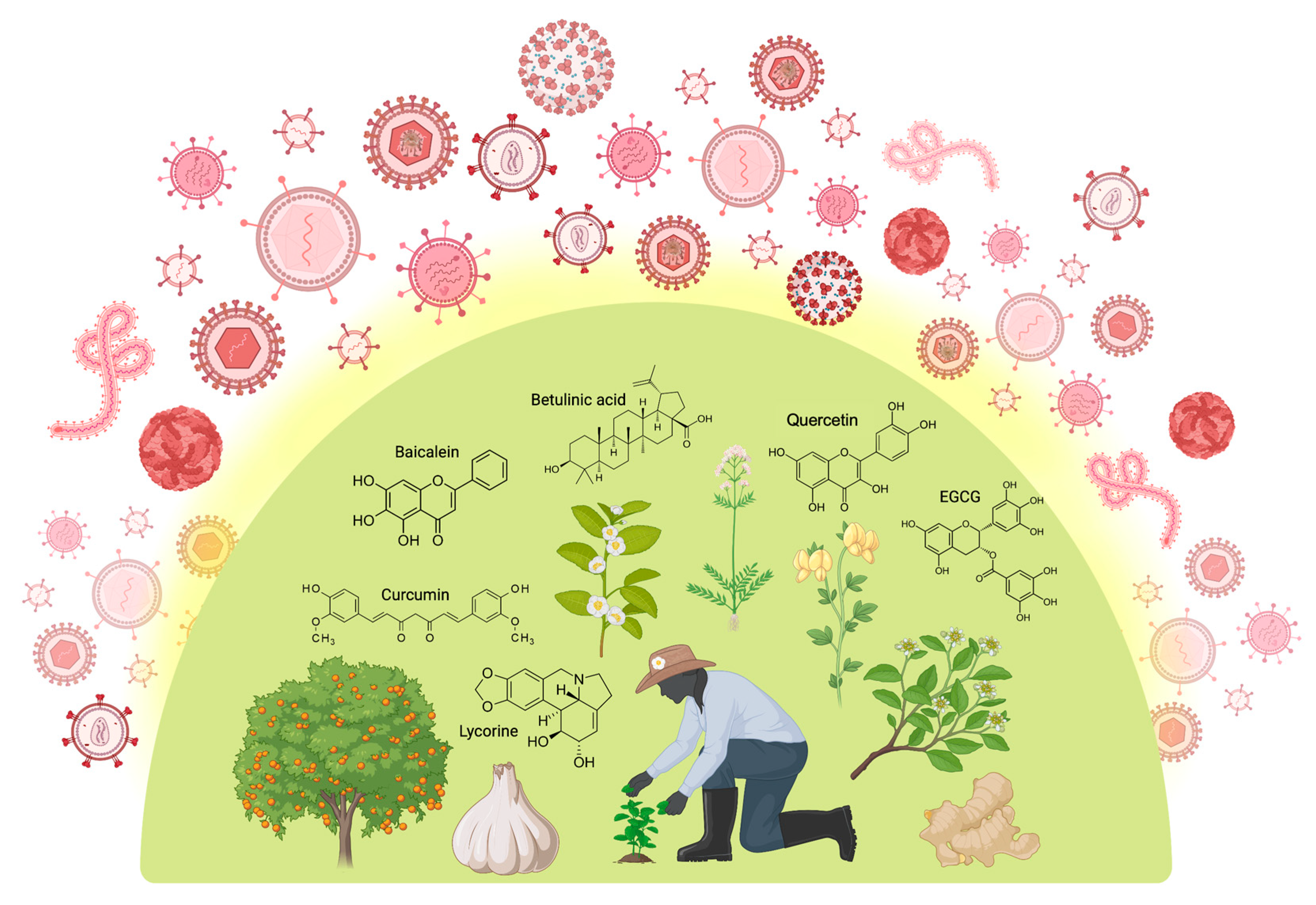
Antiviral drugs, natural and synthetic, are used to treat viral infections, with the latter synthesized like traditional medications [10]. The abundance of bioactive agents with therapeutic qualities in plants makes them suitable for use in medicine. The primary source of remedies in almost 80% of developing nations is herbal [11], and over 40% of medications today come from natural products and traditional knowledge [12]. Plant compounds such as alkaloids, flavonoids, polyphenols, and tannins have been studied for their potential antiviral properties, and their effectiveness is being evaluated continuously [13]. Figure 2 presents an overview of medicinal plants and plant-derived phytochemicals that have been used for viral protection.
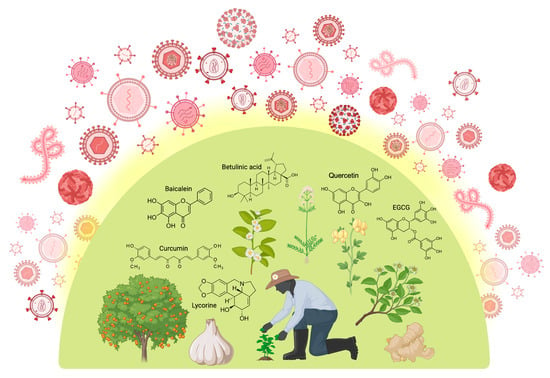
Figure 2.
Medicinal plants and plant-derived phytochemicals offer broad-spectrum protection against various stages of the viral life cycle. Polyphenols (e.g., quercetin, baicalein, curcumin, EGCG), followed by terpenoids and alkaloids such as betulinic acid and lycorine, respectively, are the most commonly occurring structural families of phytochemicals bearing pronounced antiviral properties. Created in BioRender [14].
Plant biotechnology can also advance antiviral treatments, from improving the production of natural antiviral compounds to aiding in the development of plant-based vaccines, antibodies, and therapeutic and diagnostic proteins. By harnessing the unique properties of plants and advancing genetic engineering, biotechnology has the potential to offer sustainable solutions for both human and animal health.
This review highlights the significant role of plant-derived compounds and plant biotechnology in antiviral research, focusing on recent advancements in the field. An alternative strategy for creating safe, efficient, and cost-effective antivirals and vaccines is the use of recombinant proteins and small molecules expressed in plants. By harnessing the natural capabilities of plants, we can produce therapeutic compounds, offering a sustainable and scalable approach for combating viral infections.
2. Life Cycle of (Re)Emerging Viruses—How Can Natural Products Help?
(Re)emerging viruses continue to pose significant threats to global health, with their ability to rapidly mutate, adapt to new environments, and spread across populations [15]. The increasing frequency of viral outbreaks (Figure 1) underscores the urgent need to improve the production of plant remedies, plant-derived therapeutics, and vaccines. However, identifying herbal or herbal-based remedies or developing synthetic agents that can suppress or disrupt the life cycle of viruses without causing harm to the host’s cells is a notable challenge. Furthermore, viruses utilize host cellular machinery for the synthesis and assembly of their basic components—nucleic acid (DNA or RNA), the protein coat, viral enzymes, and, occasionally, a lipid envelope—which significantly complicates the development of safe-for-the-host antivirals [11,16].
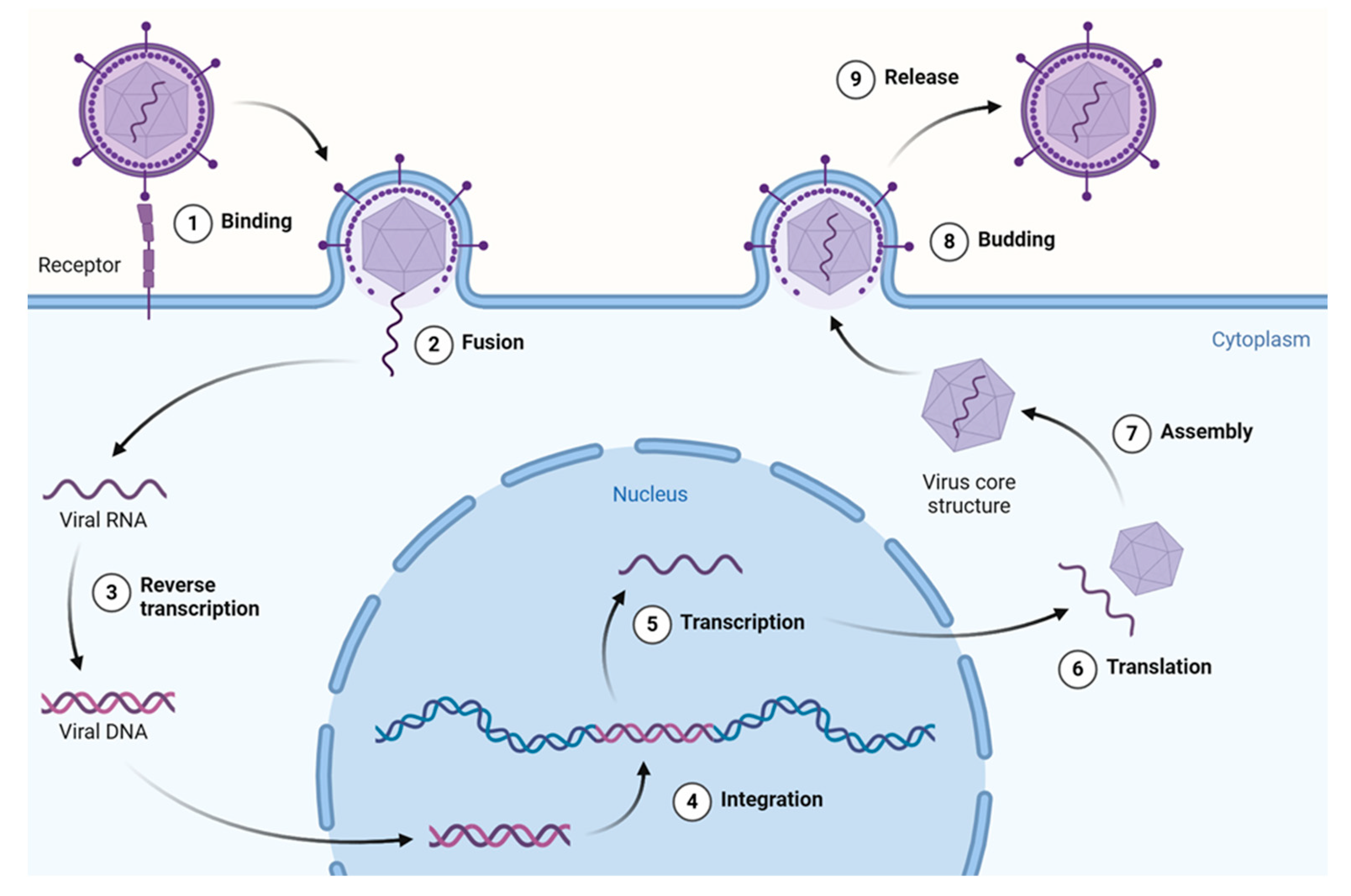
In general, the viral life cycle comprises the following events—attachment, penetration, uncoating, transcription, replication, assembly, and release—with most antivirals targeting at least one of the above steps. This cycle repeats multiple times, leading to viral spread within the host organism and the onset of potentially pathological conditions. The stages of the retrovirus life cycle are described in Figure 3.
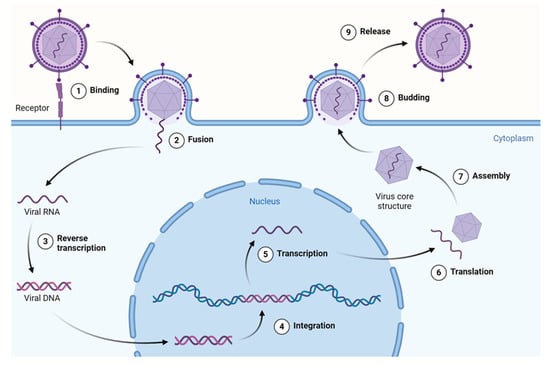
Figure 3.
Overview of the retrovirus (HIV-1) life cycle. This figure illustrates key stages from viral entry and reverse transcription to integration, replication, assembly, and budding of the mature virion. HIV-1 is shown here as a model system due to its extensively studied replication pathway and the availability of data on plant-derived compounds targeting various stages. Created using a template retrieved from Goldman-Israelow. Created in BioRender [14].
Current antiviral therapeutics exhibit their properties either by inhibiting virus attachment or entry into the cell, increasing the cell’s resistance to the virus, or deproteinizing the virus, or by producing antimetabolites that inhibit nucleic acid synthesis.
3. Plant Compounds That Target Stages of the Viral Life Cycle
A common feature of all viruses is the exploitation of host proteins for their efficient entry, replication, assembly, and exit from infected cells, all while trying to evade the host immune system. This is achieved through mechanisms of prioritizing viral protein expression and specific interactions between viral and host proteomes. Plant-derived preparations have been used to treat infectious diseases throughout human history. Phytochemicals have been shown to impact the different stages of the viral life cycle by interacting with specific viral proteins and reducing viral infectivity. Below, we describe proteins involved in coronavirus, HIV-1, influenza, dengue, Zika, Ebola, and chikungunya virus entry, replication, and viral protein synthesis and maturation. Plant-derived bioactive compounds that have been proven to exhibit antiviral properties through in vitro experiments are listed in Appendix A (Table A1 and Table A2), along with their respective specific viral protein targets and calculated IC50/EC50 values.
3.1. Phytochemicals Modulating Viral Entry, Attachment, and Fusion
Viruses must first enter their target cells through interactions between external viral structural proteins and host cell surface proteins, which trigger receptor-mediated endocytosis or direct membrane fusion.
Coronaviruses (CoVs) are a family of enveloped, positive-sense, single-stranded RNA viruses. Seven species of CoVs are pathogenic for humans, and three of them have caused pandemics of severe disease—the SARS-CoV and MERS-CoV outbreaks in 2002 and 2012, respectively, and SARS-CoV-2, which emerged in Wuhan, China, in December 2019 and caused the COVID-19 pandemic, which spread rapidly across the world due to the high transmissibility and pathogenicity of the virus [17,18,19,20]. The high pathogenicity and mortality of COVID-19 revealed the urgent need for broad-spectrum antivirals to meet the challenges of current and future coronavirus outbreaks [21].
Coronaviruses utilize the spike (S) proteins on their membrane envelopes to bind cell surface proteins such as angiotensin-converting enzyme 2 (ACE-2; SARS-CoV and SARS-CoV-2) or dipeptidyl peptidase 4 (DPP4; MERS-CoV) via the S1 subunit, while the S2 subunit mediates membrane fusion following cleavage by host surface proteases like transmembrane protease serine 2 (TMPRSS2), cathepsins, and furin [22]. Therefore, both the interaction between S proteins and their host surface protein targets and the cleavage of the S protein necessary for fusion can be targeted by antivirals.
Several molecules of plant origin have been identified by in vitro experiments as potent inhibitors of the processes of interaction and fusion of CoVs, such as cepharantine, neferine, hernandezine, and various agglutinin lectins [23,24]. Punicalin and punicalagin found in Gunnera perpensa inhibited the process of viral entry by the disruption of spike glycoprotein–host ACE2 interaction at very low concentrations—IC50 0.009 µM and 0.029 µM, respectively [25]. Similarly, quinoline carboxylic acids present in Ephedra sinica also blocked the interaction between the SARS-CoV-2 receptor-binding domain and ACE2 in a dose-dependent manner and with a low IC50 concentration [26]. The anthraquinone emodin, found in various plants, has been shown to block S protein binding to ACE2 in vitro and to limit SARS-CoV infectivity in Vero E6 cells [27]. Ohishi and coauthors tested 10 phytochemicals, and epigallocatechin gallate (the major flavonoid in green tea) exhibited the strongest inhibition of S protein–ACE2 binding, as confirmed by ELISA experiments [28]. Other phytochemicals have been proven to impair the cleavage of the S protein by host proteases and, thus, prevent virus fusion. The large polyphenol tannic acid inhibited host TMPRSS2-mediated proteolysis of the S protein with an IC50 of 2.31 µM, as well as the main viral 3CL protease, as discussed later [29]. The ubiquitous flavonol quercetin, found in many fruits and vegetables, exhibited the blockage of furin-mediated S protein cleavage [30].
The entry of human immunodeficiency virus (HIV), a retrovirus (+ssRNA), is mediated by the interaction between gp120 and gp41 envelope glycoprotein heterodimers and the CD4 T-cell surface protein, along with the engagement of the chemokine receptor CCR5 or CXCR4. The multistep interaction triggers conformational changes in gp41, which facilitate the fusion of the viral envelope with the host membrane [31]. Abrogating the binding between gp120 and CD4 or gp41-mediated membrane fusion is therefore a viable antiviral strategy.
Disrupted HIV-1 entry by the targeting of gp120, gp41, or the necessary host cell receptors has been reported for several phytochemicals. Baicalin, a flavonoid from Scutellaria baicalensis, has been widely studied in regard to its inhibitory potential, including demonstrated effects against HIV-1 entry by binding to gp120 [32].
The host cell entry of the mosquito-borne Flaviviridae (+ssRNA) family members, the dengue (DENV) and Zika (ZIKV) viruses, is cell-type-specific and is mediated through the major structural envelope (E) viral glycoprotein binding to various host factors such as surface glucosaminoglycans, the phosphatidylserine receptor TIM-1, C-type lectins such as DC-SIGN and L-SIGN, and others. These interactions trigger clathrin-mediated endocytosis, and the subsequent acidification of the endosomes causes the formation of fusogenic E glycoprotein trimers, which facilitate fusion between the viral envelope and the endosomal membranes, resulting in the release of viral RNA into the cytosol [33].
Various phytochemicals have been shown to interfere with the entry of DENV and ZIKV by targeting either the viral E protein or crucial host entry factors. For example, naringenin, a flavonoid found in citrus fruits, has been shown to inhibit DENV infection in Huh7.5 cells [34]. Similarly, baicalein demonstrated potent inhibitory effects on DENV-2 infection in human liver cells by blocking viral adsorption, and baicalin inhibited viral attachment to host cells by binding ZIKV envelope (E) protein [35,36,37]. Gossypol has been found to be an effective inhibitor of both DENV and ZIKV envelope protein region III attachment [38]. In the context of ZIKV, EGCG again emerges as a promising entry inhibitor due to its direct binding to the ZIKV envelope protein, preventing viral adsorption and internalization into host cells [39]. Curcumin has been experimentally validated to inhibit ZIKV infection by impairing viral binding to host cells and suppressing membrane fusion, through the destabilization of the viral envelope [40].
Furthermore, several studies highlight compounds capable of affecting post-entry processes linked to viral protein translation and replication. Isoquercitrin, another flavonoid, was shown to inhibit the internalization of ZIKV particles into host cells without affecting initial binding, suggesting an impact on endosomal maturation or membrane fusion steps [41]. Unlike HIV, which uses receptor-mediated conformational changes, or coronaviruses, which require proteolytic priming, flavivirus entry depends critically on low-pH-triggered E protein rearrangements, offering unique intervention points for antiviral phytochemicals.
The arthritogenic chikungunya virus (CHIKV) is a mosquito-borne alphavirus, a member of the Togaviridae family (+ssRNA). It enters target mammalian cells through receptor-mediated endocytosis via the interaction of its envelope glycoprotein with several possible cell surface receptors—matrix remodeling-associated protein 8 (MXRA8), prohibitin 1, and the TIM1 phosphatidylserine receptor—potentially also aided by binding to glucosaminoglycans such as heparan sulfate [42,43,44,45]. After the acidification of the endocytic compartment, conformational changes in the E1 viral envelope glycoprotein trigger fusion with the endosomal membrane, allowing the nucleocapsid to enter the cytosol and initiate infection.
The interaction between CHIKV E1 glycoprotein and host factors and its subsequent entry have been shown to be abrogated by some of the phytochemicals commonly mentioned in this article—epigallocatechin gallate, baicalein, and curcumin as well as the antimalarial chloroquine [40,46,47,48]. Flavaglines FL3 and FL23, along with sulfonyl amidine, also have an inhibitory effect on entry via binding to prohibitin, while the Tectona grandis (Teak) compounds 2-(butoxycarbonyl) benzoic acid (BCB), 3,7,11,15-tetramethyl-1-hexadecanol (THD), and benzene-1-carboxylic acid-2-hexadeconate (BHCD) exert their inhibition by directly binding the E1 CHIKV envelope glycoprotein [49,50].
Influenza viruses, belonging to the Orthomyxoviridae family (-ssRNA), initiate infection by utilizing the binding of their trimeric surface glycoprotein hemagglutinin (HA) to sialic acid residues on host surface glycoproteins and glycolipids. The virus is then internalized via clathrin-mediated endocytosis, and within the acidic environment of endosomes, HA undergoes a conformational change which promotes the fusion of the viral and endosomal membranes, facilitating the release of viral ribonucleoprotein complexes (vRNPs) into the cytoplasm and their subsequent transport into the nucleus for replication [51]. The additional steps of pH-dependent fusion, endosomal trafficking, and nuclear import, absent from the life cycles of the aforementioned viruses, provide further intervention points for potential antiviral strategies. Several phytochemicals have been validated to interfere with influenza virus entry, predominantly by targeting HA-mediated binding or fusion processes.
EGCG and other catechins from green tea have been shown to inactivate influenza virions by binding to HA and blocking attachment to host cells, with experimental confirmation in vivo [52]. Quercetin, a ubiquitous flavonol found in many fruits and vegetables, and its derivatives have also been reported to inhibit influenza virus infection by the same mechanism [53,54]. Moreover, some compounds target host factors that influenza viruses exploit during entry. For instance, resveratrol, a stilbenoid from Vitis vinifera (grapes), has been found to inhibit nuclear–cytoplasmic transport of viral ribonucleoproteins by modulating host PI3K/Akt signaling, thereby impairing an essential post-entry step in the influenza replication cycle [55]. Several alkaloids (harmalol, harmane, harmaline, and strychnine sulfate) had an anti-IVA adsorption effect in MDCK cells [56].
Ebola virus (EBOV), a member of the Filoviridae family (-ssRNA), initiates infection through the engagement of its heavily glycosylated surface glycoprotein (GP) with multiple host cell binding factors such as C-type lectins (e.g., DC-SIGN, L-SIGN) and phosphatidylserine receptors like TIM-1 [57]. Following internalization through macropinocytosis, EBOV traffics to late endosomes, where its GP is cleaved by endosomal proteases, notably cathepsins B and L, thus exposing the receptor-binding site necessary for interaction with the resident endosomal Niemann–Pick C1 membrane protein (NPC1). Binding to NPC1 facilitates the fusion of the viral and endosomal membranes, enabling the release of the nucleocapsid into the cytoplasm, initiating replication [58]. The reliance on host late endosomal proteases and the additional interaction with the NPC1 protein provide further points of intervention for small-molecule antivirals.
The targeting of GP-mediated binding has been demonstrated for several proanthocyanidins and flavanols from the Maesa perlarius plant, with procyanidin b2 demonstrating the strongest EBOV GP binding [59]. Additionally, quercetin in its glycosidic form has demonstrated inhibitory effects on EBOV infection by impairing viral entry, likely through interactions with GP or the disruption of endosomal trafficking, as shown in Vero E6 cell culture [60].
3.2. Phytochemicals Modulating Viral Replication, Protein Synthesis, and Maturation
Following entry into the host cell, viruses must rapidly and efficiently replicate their genomes and produce structural and non-structural proteins using host biosynthetic machinery, through interactions between viral and host proteins, allowing viruses to hijack or bypass cellular systems such as their transcriptional machinery, RNA processing pathways, and ribosomes. Viral protein production is prioritized over host proteins through various mechanisms including co-opting host RNA polymerases, employing RNA cap-snatching, generating internal ribosome entry sites (IRESs), or selectively modulating which mRNAs are translated [61]. These mechanisms not only ensure the production of essential viral components but can also serve to suppress host antiviral responses. The disruption of the interactions between viral and host proteins at this stage is also an attractive avenue for the development of antivirals.
CoV positive-sense mRNA is directly translated by host ribosomes into the pp1a and pp1ab polyproteins upon entry into the cytosol. They are then autocatalytically cleaved by the action of the viral main protease (Mpro/3CLpro, nsp5) and papain-like protease (PLpro, nsp3) within the polyprotein polypeptide chains, resulting in the generation of several non-structural proteins (NSPs), which facilitate selective translation and the formation of the replication–transcription complex (RTC), sequestered within endoplasmic reticulum-derived double-membrane vesicles. The viral RNA-dependent RNA polymerase (RdRp, nsp12) then produces full-length genomic RNA and nested subgenomic RNAs through discontinuous transcription, which yield the main structural viral proteins (S, E, M, and N). RNA helicase (nsp13) is a nucleoside triphosphatase (NTPase). Once viral proteins are synthesized, nucleocapsids are assembled in the cytoplasm and budded into the lumen of the endoplasmic reticulum (ER)–Golgi intermediate compartments. Virions are then released from the infected cell through exocytosis [62,63,64,65]. SARS-CoV-2 genome characterization provides the basis for further studies on pathogenesis and the optimization of the design of diagnostic, antiviral, and vaccination strategies [66].
Many phytochemicals have been shown in recent years to target specific proteins involved in virus replication and maturation, both in vitro and in vivo. For example, baicalin and baicalein, main flavonoids found in Scutellaria baicalensis, inhibit SARS-CoV-2 3CLPro and SARS-CoV MPro proteinases as demonstrated by isothermal titration calorimetry and enzyme binding FRET assays and in vitro assays, respectively [67,68,69]. Baicalein also inhibited SARS-CoV-helicase unwinding and ATP-ase activity [70]. The same study also examined 21 compounds of plant origin, and myricetin, quercetin, flavanone, kaempferol, and licoflavone C were found to be efficient unwinding activity inhibitors as well. Quercetin and kaempferol and their derivatives are among the most ubiquitous polyphenols in different plant parts [71]. Their aglycones and glycosides are also potent SARS-CoV 3CLPro, SARS-CoV PLPro, SARS-CoV-2 MPro, and SARS-CoV-helicase inhibitors [70,72,73,74]. Amentoflavone is a biflavonoid of apigenin and is present in a variety of plants, such as Ginkgo biloba and Hypericum perforatum [75,76]. Amentoflavone, apigenin, and its glycoside were established to inhibit SARS-CoV 3CLPro, SARS-CoV-2 RdRp, and SARS-CoV helicase in vitro [70,73,77,78]. Catechins are present in high quantities in different Camellia sinensis herbal remedies [79]. Several studies established that different catechin representatives exert inhibitory activity on proteins, responsible for CoV replication [70,80,81,82,83]. FRET analysis revealed cyanidin 3-O-galactoside to be a potent SARS-CoV-2 MPro inhibitor [82]. This compound is present in high concentrations in Vaccinium species, Aronia melanocarpa, and Sambucus ebulus [84,85,86]. A strong MERS-CoV RdRp inhibitor is lycorine, found in Lycoris radiata [87]. CoV RdRP is highly conserved among coronaviruses, and its low mutation rate makes it a promising target for drug development [88,89,90].
The HIV genome is composed of two identical single-stranded RNA molecules. The genome of the HIV provirus, also known as proviral DNA, is generated during reverse transcription by reverse transcriptase (RT) of the viral RNA genome into DNA, the degradation of the RNA, and the integration of the double-stranded HIV DNA into the human genome by HIV-1 integrase [91]. HIV-1 RT inhibition by chicoric acid and 1-methoxyoxalyl-3,5-dicaffeoylquinic acid was demonstrated by McDougall and coauthors [92]. Geraniin and corilagin have also been shown to be effective RT inhibitors in vitro [93]. In a study from 2021, anti-HIV-1 integrase and anti RT-associated RNAse activities were demonstrated in vitro for 11 compounds, isolated from Punica granatum [94]. Among them, apigenin, ellagic acid, luteolin, luteolin 7-O-glucoside, punicalins, and punicalagins were the most potent inhibitors of both enzymes. Strong inhibition of RT-associated RNA-se was detected for betulinic, oleanolic, and ursolic acids as well. A derivative of betulinic acid (3-O-(3′,3′-dimethylsuccinyl) betulinic acid) decreased RT-associated RNAse in cell culture [95].
The ZIKV genomic RNA encodes a large polyprotein that is co- and post-translationally processed into structural capsid (C), pre-membrane (prM), and envelope (E) proteins, and non-structural proteins (NS1, NS2a, NS2b, NS3, NS4A, NS4B, and NS5) [96]. The main biological role of NS5, the largest non-structural ZIKV protein, is the synthesis and capping of the viral RNA. It consists of two major functional domains—the N-terminal methyltransferase (MTase) and the C-terminal RNA-dependent RNA polymerase (RdRP) [97,98]. However, to date, effective preventive measures against the Zika disease are still lacking [99,100]. Enzyme interaction analysis established catechins, luteolin, and myricetin as possible efficient pZIKV NS2B-NS3Pro inhibitors [101]. ZIKV helicase activity is impaired by EGCG, and ZIKV RdRp activity is impaired by lycorine [102,103].
The dengue virus (DENV) genome encodes three structural proteins (C, M, and E), which form part of the mature virion, and seven non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5) [104]. NS5 is involved in mRNA capping through its methyltransferase and guanylyltransferase activities [105]. Attempts to develop anti-DENV antiviral agents appear to be very challenging due to the presence of four dengue serotypes (DENV1, DENV2, DENV3, and DENV4) that are highly prone to mutation due to the error-prone nature of their RNA polymerase [106]. Several studies focus on the few anti-DENV plant compounds that can interact with key viral proteins in vitro and in cell culture. Sotetsuflavone and apigenin are DENV RdRp inhibitors [107,108]. Myricetin is a potent DENV NS1 protease inhibitor, while bisdemethoxycurcumin and curcumin are DENV NS2B/NS3 protease inhibitors [109,110].
Chikungunya virus (CHIKV) has a ~12 kb positive-sense RNA genome with two open reading frames—the 5′-proximal ORF encodes non-structural replication proteins nsp1-4, while the 3′-ORF encodes the structural capsid, E1-3, and 6K proteins [111,112]. Upon exit from the endocytic compartment, the uncoated viral RNA is directly translated into one of the non-structural polyproteins nsp1-4, which then undergoes autoproteolysis by virtue of the nsp2 protease/helicase domain. The replication complex is then assembled and the RNA-dependent RNA polymerase nsp4 produces a negative-sense RNA template for genomic RNA synthesis and for the production of subgenomic RNA, encoding the structural polyprotein, which is then further cleaved. Other than the phytochemicals blocking its entry, several studies have found inhibitory effects of natural compounds on CHIKV replication. One study identified apigenin (IC50 = 70.8 µM), chrysin (IC50 = 126.6 µM), narigenin (IC50 = 118.4 µM), and silybin (IC50 = 92.3 µM) as potent inhibitors of CHIKV replication in BHK-21 cells, while harringtonine from Cephalotaxus harringtonia lowers negative- and positive-sense RNA and viral protein production in the same cell line [113,114]. Among other phytochemicals showing promising reductions in CHIKV viral replication or protein synthesis are berberine, prostratin, baicalein, and silymarin [48,115,116,117].
The genome of influenza A virus (IAV) is divided into eight segments—viral RNAs (vRNAs)—all present in a single IAV virion, that encode at least 16 different viral proteins, using partially overlapping open reading frames and alternative splicing [118,119]. The vRNAs are single-stranded, except for the last 13–14 nucleotides of the 5′ and 3′ termini, which are partially complementary [120]. Influenza neuraminidase (NA) cleaves the glycosidic linkage of terminal sialic acid residues on host glycoproteins. It plays a role in the release of progeny virions from infected cells [121,122]. Due to its important role in the viral life cycle, NA is considered the primary target for influenza antivirals. Several compounds have been tested for anti-IVA NA activity. Among them, gossypetin, herbacetin, kaempferol, and quercetin showed very low IC50 concentrations [123]. Other studies have revealed hispidulin, luteolin, matteucin, matteucinol, methoxymatteucin, and nepetin to also have the same activity [124,125]. 2′,4′dihydroxy-6′-methoxy-3′,5′-dimethylchalcone and myricetin-3′,5′-dimethylether-3-O-β-D-galactopyranoside exhibited inhibition against NA of four IVA subtypes in HEK293 cell culture and oxypeucedanin against two IVA subtypes [126,127]. IVA RdRp in vitro inhibition was also established for catechins [128]. Berberine, an active ingredient of Berberis sp., interfered with IVA maturation by blocking the transport of influenza A ribonucleoprotein to the cytoplasm [129]. IVA is also known to upregulate the mitogen-activated protein kinase/extracellular signal-related kinase (MAPK/ERK) pathway and hijacks this pathway for nucleolar export of the viral ribonucleoprotein for virus maturation, with berberine hampering this interaction [129].
Ebola virus (EBOV) is composed of seven genes coding at least ten proteins, among which are the following: nucleoprotein (NP), viral protein 35 (VP35), VP40, glycoprotein (GP), soluble GP (sGP), Δ-peptide, ssGP, VP30, VP24, and polymerase (L). VP35 is a component of the EBOV polymerase complex, where it serves as a bridge between the viral RNA-dependent RNA polymerase (L) and the nucleoprotein (NP) [130,131]. It is also involved in viral genome packaging and nucleocapsid formation and has been reported to possess NTPase- and helicase-like activities potentially involved in viral RNA remodeling [132,133]. Several plant components have been tested for VP35–dsRNA interactions, where cynarin and germacrane sesquiterpene 8α-(5′-hydroxyangeloyl)-salonitenolide were identified as the most potent with 90% and 60% inhibitory activity, respectively, at 100 µM [134].
4. (Re)Emerging Viruses—How Can Plant Biotechnology Help?
The use of engineered plants to produce high-quality molecules (vaccines, antibodies, cytokines, peptides, and bioactive small molecules) with pharmaceutical applications is increasing [135]. This area of biotechnology is also known as plant molecular farming (PMF) [136,137]. The advancement and optimization of PMF are due to the successful application of well-established techniques, such as plant gene engineering, increasing yields through alternative expression methods using plant viruses, genome editing biosynthetic pathways, modeling plant glycoengineering, and improving downstream processing (DSP) [138,139,140,141].
4.1. Plant-Derived Vaccines
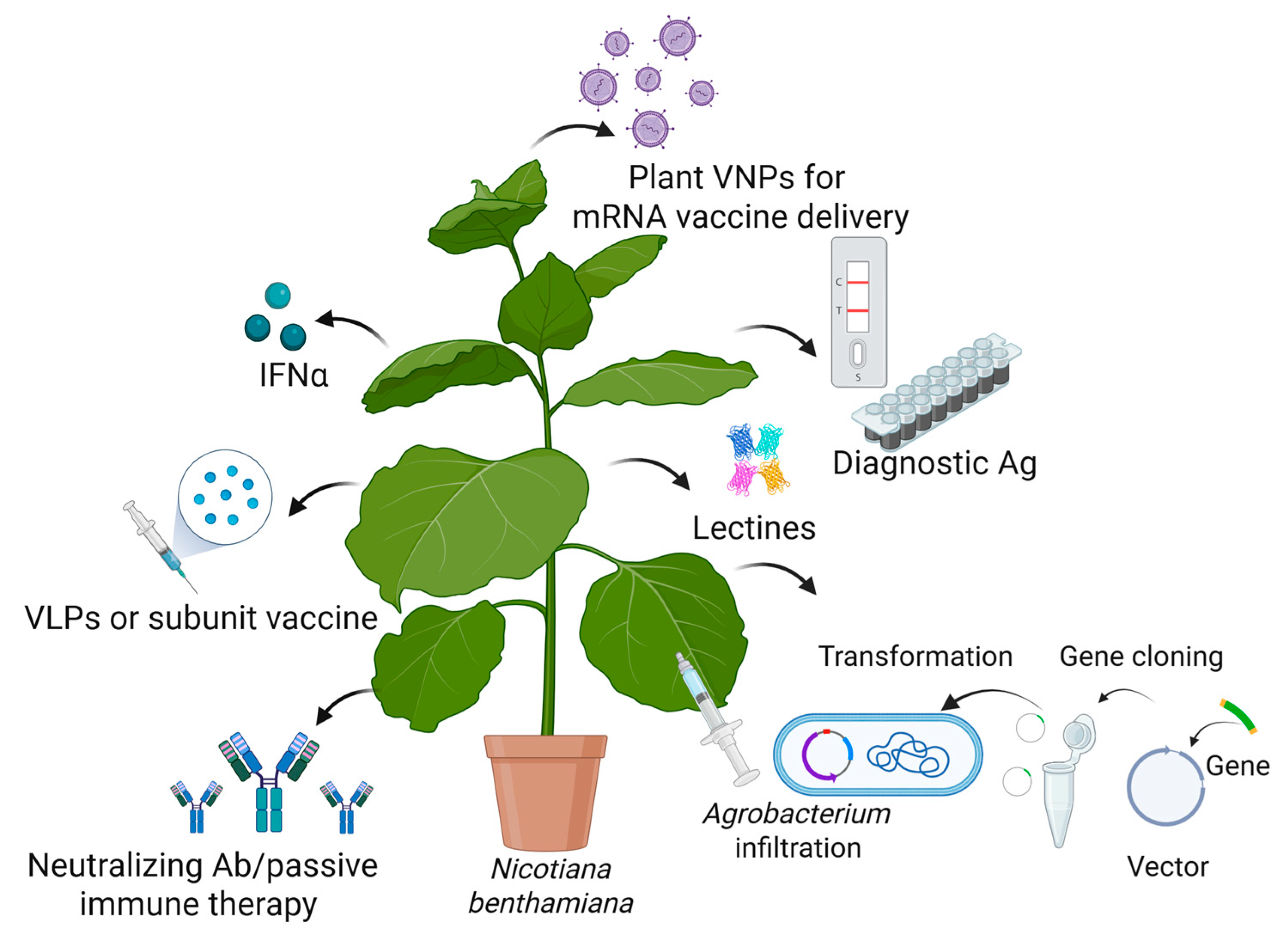
Plant biotechnology and vaccine production have emerged as a promising field that can address some of the global challenges posed by (re)emerging viruses. Plant-derived vaccines are produced using recombinant biotechnology, where the gene encoding the desired antigen is inserted into a viral vector or integrated into the plant genome [142,143]. Effective vaccinogens can then be cost-effectively manufactured in plants, providing a massive number of doses for global-scale deployment in a relatively short amount of time. PMF is based on three main technological platforms: transient expression in Nicotiana benthamiana using viral vectors, stable transgenic or transplastomic plant production, and transgenic plant cell-suspension cultures [142,143,144,145,146]. Transient expression with agroinfiltration is the process of introducing genes into plant leaves by infiltrating them with binary vectors carried by Agrobacterium tumefaciens. Figure 4 describes the use of plants as biofactories to produce recombinant proteins with antiviral applications.
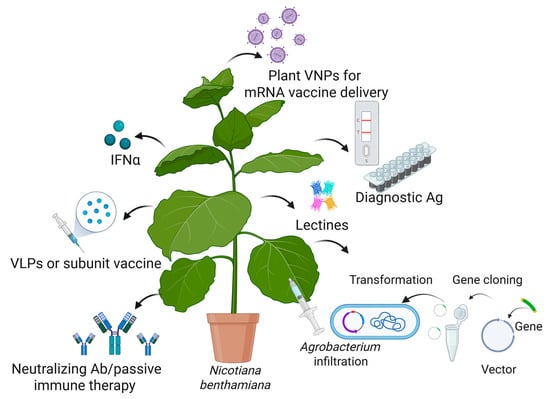
Figure 4.
Transient expression of recombinant proteins (antigens (Ag), antibodies (Ab), viral nanoparticles (VNPs), vaccines, virus-like particles (VLPs), lectins, interferon α (INFα)) in N. benthamiana and their application in the fight against emerging viruses. Created in BioRender [14].
Considering the need for vaccines and therapeutics for rapidly spreading (re)emerging viruses, transient expression systems provide the necessary time efficiency and scalability [147,148]. During the COVID-19 pandemic, several companies (Medicago Inc., Québec, QC, Canada; Kentucky BioProcessing, Owensboro, KY, USA; iBio, San Diego, CA, USA; Baiya Phytopharm, Bangkok, Thailand) used N. benthamiana to develop a vaccine against SARS-CoV-2. Their candidate vaccines passed successful phase-1 and -2 clinical trials [148,149,150]. Medicago Inc.’s vaccine successfully passed phase-3 clinical trials, and in 2022, Health Canada approved the first plant-derived SARS-CoV-2 vaccine, Covifenz [151]. Using transient expression technology, Medicago Inc. also produced and licensed a vaccine against the influenza virus. Although stable plant transformation takes longer, this approach enables large-scale and long-term recombinant protein production. The stable transformation of edible plants (maize, rice, oilseeds, lettuce, alfalfa, and tomatoes) allows the development of orally delivered vaccines, especially for veterinary applications [6,152,153,154]. To identify the most efficient plant expression system (stable, transient, or transplantomic), the viral antigens are expressed both transiently and stably. For example, HIV-1 capsid protein p24 was expressed independently and fused with the matrix subunits p17 (p17/p24) using transient and stable expression in tobacco. The Agrobacterium-mediated transient expression of p24 and p17/p24 gave the highest yield (1 mg p24/kg fresh weight). The highest intracellular accumulation of p17/p24 was observed in the chloroplast compared to the endoplasmic reticulum (ER) and cytoplasm [155]. In conclusion, the optimal production method (stable or transient) and the site for achieving sustained and robust accumulation (cytosol, chloroplast, ER, or apoplast) need to be determined empirically based on the distinct features of the chosen immunogen. The route of vaccine delivery, the capacity to create high amounts of recombinant protein expression, and the low value of downstream processing all influence the selection of technology for plant-based vaccine expression.
Plant molecular farming and plant-derived vaccines offer workable solutions addressing some of the challenges caused by outbreaks of (re)emerging viruses [156,157,158,159]. The high cost of developing and producing new vaccines is a significant obstacle for the resource-limited countries that need them the most. Developing countries heavily depend on other regions for life-saving vaccines, 99% of which are imported [160]. This stark imbalance in vaccine production can lead to unequal access and significant health disparities between developing countries and other parts of the world. Plant biotechnology companies based in Africa or Asia, producing plant-derived vaccines, therapeutics, and diagnostic reagents, may solve production and supply problems locally. Companies like Cape Biologix Technologies, South Africa, Baiya Phytopharm, Thailand, and BioApplications Inc., South Korea, are good examples of the successful implementation of local products in local markets [149,161]. Plant-derived vaccines against (re)emerging viruses developed during the past 10 years are summarized in Table 1.

Table 1.
Examples of plant-derived vaccines to prevent recently emerging viruses.
4.2. Bio-Encapsulation of mRNA Within Plant-Derived Virus-like Particles (VLPs) and Chimeric VLP Production
Virus-like particles (VLPs) that mimic a natural virion but lack the native genome have been used as vaccines or vehicles for drug and nucleic acid delivery [168]. Plant-virus-derived VLPs, such as the tobacco mosaic virus (TMV) and cowpea mosaic virus (CPMV), were used as delivery platforms for mRNA vaccine and diagnostic reagent production. A self-amplifying mRNA vaccine packaged by TMV coat proteins expressing E7 protein from human papillomavirus 16 (HPV16) elicits E7-specific IgG antibodies. The same study demonstrated that TMV VLPs are suitable for mRNA vaccine delivery in vivo. Furthermore, plant-derived VLPs are used as nanoparticles to present heterologous immunogens to the immune system, inducing potent cellular and humoral immune responses. These chimeric proteins can self-assemble due to the structural properties of the viral capsid proteins and form particles with similar morphology and structure to the native virion [169,170,171,172]. Table 2 provides the most notable examples of chimeric VLPs produced in plants.

Table 2.
Chimeric VLPs displaying foreign antigens and their immunogenic properties.
4.3. Plant-Derived Antibodies Used for Passive Immunotherapy
Passive immunotherapy is a form of immunization that uses immunoactive products, such as antibodies produced outside the patient’s body. These antibodies provide immediate protection against infections or diseases without necessitating the immune system’s involvement. This approach is particularly beneficial for immunocompromised patients or when a rapid response to a threat is not possible [183]. Antibodies represent a critical component of the vertebrate adaptive immune system. Antibodies can be produced by plants, genetically modified with the genes that encode mammalian and human immunoglobulins. Plant-derived antibodies (plantibodies) offer several advantages, including reduced production costs, the absence of animal pathogens, and scalability. Importantly, plantibodies demonstrate high stability and a long shelf life, making them suitable for application in countries with limited medical infrastructure. Model organisms such as tobacco (N. benthamiana and N. tabacum), rice, and potato are among the most commonly used plant bioreactors due to their high yield and ease of cultivation. Current approaches focus on increasing expression and shortening production time. The magnification approach, which involves the insertion of multiple provectors into Agrobacterium tumefaciens, is a recent development in monoclonal antibody (mAb) production. This method has the potential to yield functional mAbs within 14 days, making it a valuable tool for preclinical, clinical, and commercial applications [184,185,186].
In recent years, plantibodies have gained significant popularity, emerging as an alternative to conventionally used antibodies derived from mammalian cell cultures [187]. Significant parallels have been observed between plant and mammalian antibody production, particularly in terms of glycosylation. In this regard, plants typically add mannose and xylose to N-linked oligosaccharides, whereas animals add galactose and sialic acid. This variation in glycosylation has the potential to impact the immunogenicity and efficacy of plant-derived antibodies when administered to humans [188,189]. Through the use of genetic engineering, modifications have been introduced to enhance glycosylation compatibility with human physiology [190]. Antibodies containing glycans that are nearly homogeneous, possess a mammalian structure, and exhibit enhanced neutralizing capabilities have recently been generated in plants. For example, in the case of antibodies directed against the Ebola virus, the incorporation of a specific motif, such as KDEL, has been shown to direct the antibody to the endoplasmic reticulum, where glycosylation occurs. This results in the formation of N-glycans with a high mannose content, which, in turn, enhances the efficiency of antibody binding to viral particles [191]. In addition, precise control of N-glycosylation was achieved through the generation of transgenic plants (N. benthamiana) with reduced xylosyl- and fucosyltransferase (ΔXF) activity by RNA interference. The antibodies expressed in these plants contain homogeneous N-glycans (GnGn), which increases their affinity for FcγRIIIa. This study provides the first successful example of improved antiviral activity by glycoengineering, as evidenced by the production of an antibody against HIV in these plants [192].
The yield and quality of recombinant secretory IgA antibodies can be improved by the engineering of the endoplasmic reticulum (ER) in N. benthamiana [193]. Göritzer et al. expanded the ER by targeting the enzyme CTP:phosphocholine cytidylyltransferase (CCT), using CRISPR/Cas to perform site-directed mutagenesis of each of the three endogenous CCT genes in N. benthamiana. The mutant plants with a changed ER increased the yields of assembled secretory IgAs due to prolonged ER residence time and a boost in chaperone accumulation [193]. As interest in antibody production from plant systems has grown, so has their productivity. In recent years, several antibodies against some of the more serious viral diseases, such as human immunodeficiency virus, Ebola virus, West Nile virus, dengue virus, herpes simplex virus, rabies virus, and respiratory syncytial virus, have been expressed in plants. Table 3 presents a comprehensive overview of plant antibodies, their expression system, and the infectious diseases targeted.

Table 3.
Overview of plant antibodies, the system in which they are expressed, and the infectious diseases against which they are directed.
4.4. Recombinant Cytokines Produced in Plants
Cytokines are signaling proteins produced by cells to influence the immune response and help control inflammation in the human body [219]. Recombinant human cytokines have been produced in plants using stable or transient expression methods for increasing protein yield and stability [220]. Interferons, the main group of cytokines, play a central role in the regulation of immune and inflammatory responses to viral infections [221]. Interferon alfa (INF-α) is approved by the FDA in the treatment of chronic hepatitis C and B, and AIDS-related Kaposi’s sarcoma [222]. Recombinant human interferons have been successfully produced in different plant species (carrot, lettuce, potato, and rice) [222,223,224,225,226,227,228,229]. Similarly to the commercial product, a recombinant IFN demonstrated dose-dependent antivirus activity in human A549 cells [223]. These findings show that using scalable and environmentally friendly plant-based expression systems for the production of functional recombinant human proteins is feasible.
4.5. Recombinant Carbohydrate-Binding Proteins with Antiviral Activity Produced in Plants
Some native small carbohydrate-binding proteins from plants (lectins) can be used as antivirals. Lectins have the capacity to bind to and neutralize a wide variety of viruses by obstructing the glycan structures found on the surface of the virus [230]. The lectin Griffithsin (GRFT), produced from red algae, has shown broad-range antiviral efficacy against many enveloped viruses, such as HIV-1, SARS-CoV-2, MERS-CoV, HCV, Japanese encephalitis virus (JEV), herpes simplex virus-2 (HSV-2), and porcine epidemic diarrhea virus (PEDV) [231,232]. Lectins can also be obtained via genetically modified plants, thereby affording increased production and simplified purification (Table 4) [233].

Table 4.
Recombinant lectins produced in plants.
5. Engineering of Plant Biosynthetic Pathways for Overproduction of Phytochemicals
Numerous therapeutic substances naturally occur in plants. They are based on secondary metabolites (terpenoids, alkaloids, flavonoids, etc.) and can be extracted from their native producers [244]. However, many of them are produced in specific plant tissues or at certain developmental stages. This specific spatiotemporal expression pattern limits the overall yield when attempting to extract metabolites from the entire plant [9]. Plant secondary metabolites are produced via multistep biosynthetic pathways that involve a variety of enzymes, cofactors, and intermediates that have a highly specific stereochemistry. In recent years, research has increasingly focused on extensively characterizing the biosynthetic pathways of plant-derived secondary metabolites, driven by the need to produce these metabolites on a larger scale and in a more controlled and economically viable manner.
5.1. Improved Production of Phytochemicals by CRISPR-Cas9 Genome Editing
In recent years, genome editing technologies have significantly advanced our ability to manipulate plant metabolic pathways for the production of valuable bioactive compounds. CRISPR-Cas9 technology enables the targeted editing of specific genes involved in plant biosynthetic pathways. Key strategies include gene knockout, multiplex editing, and targeted mutations to optimize the biosynthetic pathways of important compounds such as alkaloids and phenols. For instance, Salvia miltiorrhiza has been genetically edited to enhance its phenolic acid content [245]. The bZIP2 gene was knocked out using CRISPR-Cas9, leading to an increased concentration of phenolic acids. In another case, Cichorium intybus (chicory), which produces sesquiterpene lactones (STLs), a group of compounds known for their antiviral and anticancer properties [246], was edited using CRISPR-Cas9 to inhibit the germacrene A synthase (CiGAS) enzyme. This led to an upregulation of phenolic acid production, enhancing the plant’s antiviral potential [247]. The applications of CRISPR/Cas9 in medicinal plants to improve the production of bioactive compounds are summarized in Table 5.

Table 5.
Modification of medicinal plants to improve the production of bioactive compounds using CRISPR/Cas9 technology for gene editing.
Prior to the advent of CRISPR/Cas technology, various non-CRISPR/Cas genetic manipulation techniques were employed to enhance the production of specialized metabolites in medicinal plants. These methods include gene silencing with RNA interference (RNAi), gene stacking (multiple genetic modifications in a single plant), and overexpression [255].
5.2. Transient Expression of Biosynthetic Enzymes
The transient expression of biosynthetic enzymes in N. benthamiana has emerged as a powerful approach for the production of complex plant-derived metabolites. This method has become a “go-to” strategy due to its speed and flexibility, allowing researchers to bypass the slower and more labor-intensive process of stable transformation [256,257]. Such successful transient expression is exemplified by the production of triterpenes, e.g., saponins, in N. benthamiana. A combinatorial approach was used to co-express 16 enzymes involved in the biosynthesis of triterpenes. This system enabled the production of a saponin molecule that could potentially be further engineered as an adjuvant for vaccines [258].
7. Conclusions
Plant-derived natural products and biotechnological innovations offer a multifaceted strategy to combat emerging viral threats. By addressing challenges in specificity, supply chain, biosynthesis, and delivery, plant-based solutions could play a transformative role in future pandemic preparedness and response. Continued interdisciplinary research is critical to unlock their full potential and ensure safety, efficacy, scalability, and sustainability. As explored in this review, an integrated approach combining genome editing, structural biology, metabolic engineering, and gene engineering can significantly enhance the development and utilization of plant-based antiviral agents. Despite the inherent challenges, such as production constraints and the complexity of biosynthetic pathways, continued innovation in these areas holds great potential for generating effective plant-derived drugs, vaccines, antibodies, and diagnostic tools. The lessons learned from the COVID-19 pandemic and technologies developed during that period underscore the importance of adaptable technologies for vaccine and antiviral production. The progress made not only provides a foundation for addressing current health crises but also serves as a strategic framework for mitigating future viral outbreaks.
Author Contributions
Conceptualization, G.Z.; writing—original draft preparation, all authors; writing—review and editing, G.Z. and G.L.L.; visualization, G.Z.; supervision, G.Z. and Y.K.-K. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Plovdiv University Research Fund MUPD25-BF-004 and by the European Regional Development Fund through Programme Research Innovation and Digitalisation for Smart Transformation, Grant agreement № BG16RFPR002-1.014-0003-C01.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| SARS-CoV | Severe Acute Respiratory Syndrome Coronavirus |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome Coronavirus 2 |
| MERS-CoV | Middle East Respiratory Syndrome Coronavirus |
| CoVs | Coronaviruses |
| H1N1 | Hemagglutinin Type 1 and Neuraminidase Type 1 (Influenza A subtype) |
| HIV-1 | Human Immunodeficiency Virus Type 1 |
| mRNA | Messenger Ribonucleic Acid |
| VLPs | Virus-Like Particles |
| IVA | Influenza Virus A |
| DENV | Dengue Virus |
| ZIKV | Zika Virus |
| EBOV | Ebola Virus |
| MPXV | Monkeypox Virus |
| RNA | Ribonucleic Acid |
| RNP | Ribonucleoprotein |
| DNA | Deoxyribonucleic Acid |
| S protein | Spike Protein |
| ACE2 | Angiotensin-Converting Enzyme 2 |
| DPP4 | Dipeptidyl Peptidase 4 |
| TMPRSS2 | Transmembrane Protease Serine 2 |
| NP | Nucleoprotein |
| VP | Viral Protein |
| GP | Glycoprotein |
| sGP | Soluble GP |
| L | Polymerase |
| NTPase | Nucleoside Triphosphatase |
| HA | Hemagglutinin |
| vRNP | Viral Ribonucleoprotein Complex |
| E protein | Envelope Protein |
| CD4 | Cluster of Differentiation 4 |
| CCR5 | C-C Chemokine Receptor Type 5 |
| CXCR4 | C-X-C Chemokine Receptor Type 4 |
| NPC1 | Niemann–Pick C1 Protein |
| E8L | A Viral Envelope Protein from MPXV |
| EGCG | Epigallocatechin Gallate |
| ASA, ASAI | Allium Sativum Lectins |
| IC50 | Half Maximal Inhibitory Concentration |
| FRET | Förster Resonance Energy Transfer |
| Mpro/3CLpro | Main Protease/3-Chymotrypsin-Like Protease (also nsp5) |
| PLpro | Papain-Like Protease (also nsp3) |
| NSPs | Non-Structural Proteins |
| RdRp | RNA-Dependent RNA Polymerase (also nsp12) |
| RTC | Replication–Transcription Complex |
| ER | Endoplasmic Reticulum |
| ERGIC | Endoplasmic Reticulum–Golgi Intermediate Compartment |
| IRES | Internal Ribosome Entry Site |
| RT | Reverse Transcriptase |
| GAGs | Glycosaminoglycans |
| DC-SIGN/L-SIGN | Dendritic Cell-Specific Intercellular Adhesion Molecule-3-Grabbing Non-Integrin/Liver/Lymph Node-Specific ICAM-3-Grabbing Integrin |
| TIM-1 | T-cell Immunoglobulin and Mucin-Domain-Containing-1 |
| PI3K/Akt | Phosphoinositide 3-Kinase/Protein Kinase B pathway |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| PMF | Plant Molecular Farming |
| DSP | Downstream Processing |
| HIV | Human Immunodeficiency Virus |
| TMV | Tobacco Mosaic Virus |
| CPMV | Cowpea Mosaic Virus |
| HPV16 | Human Papillomavirus 16 |
| AMV | Alfalfa Mosaic Virus |
| BaMV | Bamboo Mosaic Virus-Based |
| FMDV | Foot-and-Mouth Disease Virus |
| AP 205 | Bacteriophage AP 205 |
| HBV | Hepatitis B Virus |
| HEV | Hepatitis E Virus |
| BTV | Bluetongue Virus |
| AHSV | African Horse Sickness Virus |
| mAb | Monoclonal Antibody |
| KDEL | Retention Signal Sequence in Proteins |
| ΔXF | Deletion of Xylosyl- and Fucosyltransferase (enzyme activity) |
| CTP | Cytidine Triphosphate |
| CCT | CTP:phosphocholine cytidylyltransferase |
| CRISPR/Cas | Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-Associated Protein |
| IgA | Immunoglobulin A |
| INF-α | Interferon Alfa |
| FDA | Food and Drug Administration |
| GRFT | Griffithsin |
| CV-N | Cyanovirin-N |
| EC50 | Half Maximal Effective Concentration |
| STLs | Sesquiterpene Lactones |
| CiGAS | Germacrene A Synthase from Cichorium |
Appendix A

Table A1.
Plant-derived compounds affecting viral adsorption, fusion, and entry.
Table A1.
Plant-derived compounds affecting viral adsorption, fusion, and entry.
| Compound | Activity | Cell Type Tested | Target | IC50/EC50 | Plant | Reference |
|---|---|---|---|---|---|---|
| SARS-CoV-2 | ||||||
| Cepharantine | Inhibition of pre-entry, entry, and membrane fusion | Calu-3, A549, HEK293T-ACE2, Vero E6 | Blockage of host calcium channels | 0.315 µM | N/A | [23] |
| Hernandezine | Inhibition of pre-entry, entry, and membrane fusion | Calu-3, A549, HEK293T-ACE2, Vero E6 | Blockage of host calcium channels | 0.111 µM | N/A | [23] |
| Neferine | Inhibition of pre-entry, entry, and membrane fusion | Calu-3, A549, HEK293T-ACE2, Vero E6 | Blockage of host calcium channels | 0.946 µM | N/A | [23] |
| ASA | Inhibition of early viral attachment | Vero E6 | N/A | >4 µM | Allium sativum L. | [271] |
| ASA1 | Inhibition of early viral attachment | Vero E6 | N/A | >4 µM | Allium sativum L. | [271] |
| Punicalin | Inhibition of viral entry | N/A | Disruption of spike glycoprotein–host ACE2 interaction | 0.009 µM | Gunnera perpensa | [25] |
| Punicalagin | Inhibition of viral entry | N/A | Disruption of spike glycoprotein–host ACE2 interaction | 0.029 µM | Gunnera perpensa | [25] |
| Epigallocatechin gallate | Inhibition of viral entry | HEK293FT, Caco-2 | Disruption of spike glycoprotein–host ACE2 interaction | 33.9 µM | Camelia sinensis | [28] |
| 4,6-dihydroxyquinoline-2-carboxylic acid | Inhibition of viral entry | Calu-3, HEK293T-ACE2 | Disruption of spike glycoprotein–host ACE2 interaction | 0.07 µM | Ephedra sinica | [26] |
| 4-hydroxy-6-methoxyquinoline-2-carboxylic acid | Inhibition of viral entry | Calu-3, HEK293T-ACE2 | Disruption of spike glycoprotein–host ACE2 interaction | 0.15 µM | Ephedra sinica | [26] |
| 4-hydroxyquinoline-2-carboxylic acid | Inhibition of viral entry | Calu-3, HEK293T-ACE2 | Disruption of spike glycoprotein–host ACE2 interaction | 0.58 µM | Ephedra sinica | [26] |
| Dengue virus | ||||||
| Gossypol | Inhibition of viral attachment | LLC-MK2 | Envelope protein region III | 1.87 µM (DENV-1) 1.89 µM (DENV-2) 3.7 µM/(DENV-3) 2.6 µM/ (DENV-4) | Gossypium spp. | [38] |
| Baicalein | Inhibition of DENV-2 adsorption | Vero E6 | Not established | 7.14 μg/mL | Scutellaria baicalensis | [35] |
| Baicalin | Inhibition of viral adsorption | Vero E6 | Not established | 18.07 μg/mL | Scutellaria baicalensis | [37] |
| Zika virus | ||||||
| Baicalin | Inhibition of viral attachment to host cells | Vero E6 | Envelope E protein | 14 µM | Scutellaria baicalensis | [36] |
| Gossypol | Inhibition of viral attachment to host cells | Vero E6 | Envelope protein region III | 22.2 µM | Gossypium sp. | [38] |
| Curcumin | Inhibition of viral attachment to host cells | HeLa, BHK-21, Vero E6 | Envelope E protein | 1.90 µM | Curcuma longa | [40] |
| (−) Epigallocatechin gallate | Destabilization and dissolution of viral particle | Vero E6 | Viral envelope phospholipids | 21.4 µM | Camellia sinensis | [39] |
| Isoquercitrin | Inhibition of membrane-associated viral particle internalization into A549 cells | A549 | Not established | 15.5 µM | Mangifera indica | [41] |
| HIV-1 | ||||||
| Ajoene | Inhibition of adhesive interactions and fusion of leukocytes | T-lymphoblasts (H9, CEM13) | N/A | 45 µM | Allium sativum L. | [272] |
| “TFmix” (theaflavin; theaflavin-3-gallate; theaflavin-3′-gallate; theaflavin-3-3′-digallate) | Inhibition of viral attachment | H9/HIV-1IIIB cells, MT-2 | Interference with viral gp41 6-helix bundle formation | 6.25 µM | Camellia sinensis assamica | [273] |
| Cassiabrevone | Inhibition of viral attachment | U373-CD4-CXCR4 | Viral gp120-host CD4 binding | 30.96 µM | Cassia abbreviata | [274] |
| Acerosin | Inhibition of viral entry | MT-2 | Surmised blocking of virus–CD4 or CXCR4/CCR5 host cell receptor interaction | 2.7 µM | Artemisia campestris | [275] |
| Xanthomicrol | Inhibition of viral entry | MT-2 | Surmised blocking of virus–CD4 or CXCR4/CCR5 host cell receptor interaction | 17.43 µM | Artemisia campestris | [275] |
| Guibourtinidol-(4α → 8)-epiafzelechin | Inhibition of viral attachment | U373-CD4-CXCR4 | Viral gp120-host CD4 binding | 42.47 µM | Cassia abbreviata | [274] |
| Baicalin | Inhibition of viral fusion | Hos/CD4/CCR5, Hos/CD4/CXCR4 | Broad inhibition of T-cell tropic (X4) and monocyte tropic (R5) HIV-1 Env protein-mediated fusion with host CD4/CXCR4 or CD4/CCR5 | 4 µM | Scutellaria baicalensis | [32] |
| Procyanidin A (pentamer) | Inhibition of viral attachment | PBMC | Blocks binding of viral gp120 to host heparan sulfate | 7 µM | Cinnamomum cassia | [276] |
| Procyanidin A (trimer) | Inhibition of viral attachment | PBMC | Blocks binding of viral gp120 to host heparan sulfate | 7.5 µM | Cinnamomum cassia | [276] |
| Procyanidin A (pentamer) | Inhibition of viral attachment | PBMC | Blocks binding of viral gp120 to CD4 | 21.5 µM (YU2 HIV-1 envelope), 20 µM (MN HIV-1 envelope) | Cinnamomum cassia | [276] |
| Ebola virus | ||||||
| (+) Catechin | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 36.0 µM | Maesa perlarius | [59] |
| Ellagic acid | Inhibition of viral entry | A549, HeLa | Blocking Ebola glycoprotein-mediated entry | 1.4 µM (against pseudovirions in A549 cells) 10.5 µM (against EBOV in HeLa cells) | Rhodiola rosea L. | [277] |
| (−) Epicatechin | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 22.1 µM | Maesa perlarius | [59] |
| Epicatechin gallate | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 2.95 µM | Maesa perlarius | [59] |
| Epigallocatechin | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 5.53 µM | Maesa perlarius | [59] |
| Epigallocatechin-3-gallate | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 2.8 µM | Maesa perlarius | [59] |
| Gallocatechin | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 12.4 µM | Maesa perlarius | [59] |
| Procyanidin B1 | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 0.95 µM | Maesa perlarius | [59] |
| Procyanidin B2 | Inhibition of viral attachment, entry, and fusion | A549, HEK293T | Blocking Ebola glycoprotein | 0.83 µM | Maesa perlarius | [59] |
| Quercetin 3-β-O-d-glucoside | Inhibition of early stages of viral entry | Vero E6 | Not established, surmised involvement of NPC1 endosomal transporter/LDL receptors | 5.3 µM | N/A | [60] |
| Influenza A | ||||||
| Harmalol | Viricidal and antivirus adsorption effects | MDCK | Not established | 0.035 µg/mL | Peganum harmala L. | [56] |
| Harmane | Viricidal and antivirus adsorption effects | MDCK | Not established | 0.033 µg/mL | Peganum harmala L. | [56] |
| Harmaline | Viricidal and antivirus adsorption effects | MDCK | Not established | 0.056 µg/mL | Peganum harmala L. | [56] |
| Strychnine sulfate | Viricidal and antivirus adsorption effects | MDCK | Not established | 0.06 µg/mL | Peganum harmala L. | [56] |
| Pentagalloylglucose | Inhibition of viral adsorption | MDCK | Viral hemagglutinin | 2.51 µM | Phyllanthus emblica Linn | [278] |
| 5,7,3′,4′-tetra-O-methylquercetin | Blocking host cell entry and/or recognition | MDCK | Binding to H1N1 virions | 0.36 µM | Sambucus nigra L. | [54] |
| (±)-dihydromyricetin | Blocking host cell entry and/or recognition | MDCK | Binding to H1N1 virions | 8.7 µM | Sambucus nigra L. | [54] |
| Cyanidin 3-sambubioside | Inhibition of viral adsorption | MDCK | Not established | 252 µg/mL (whole-extract value) | Sambucus nigra L. | [54] |
| Quercetin | Inhibition of viral entry | MDCK | HA2 subunit of influenza hemagglutinin | 7.76 µM (A/Puerto Rico/8/34 (H1N1)), 6.23 µM (A/FM-1/47/1 (H1N1)), 2.74 µM (A/Aichi/2/68 (H3N2)) | N/A | [53] |
| Chikungunya virus | ||||||
| Epigallocatechin gallate | Inhibition of viral entry | HEK293T | Envelope glycoprotein | 12 µM | Camellia Sinensis | [47] |
| Baicalein | Inhibition of viral adsorption | Vero cells | Not established | 103.76 µM | Scutellaria baicalensis | [48] |
| Quercetagetin | Inhibition of viral adsorption | Vero cells | Not established | 25.3 µM | N/A | [48] |
| Curcumin | Affects viral glycoprotein conformation and/or membrane fluidity | HeLa | Viral envelope | 3.89 µM | Curcuma longa | [40] |
| 2-(butoxycarbonyl) benzoic acid (BCB) | Inhibition of viral entry | Vero cells | E1 CHIKV envelope glycoprotein | 2.49 µg/mL vs. Asian isolate 28.62 µg/mL vs. African isolate | Tectona grandis | [50] |
| 3,7,11,15-tetramethyl-1-hexadecanol (THD) | Inhibition of viral entry | Vero cells | E1 CHIKV envelope glycoprotein | 1.66 µg/mL vs. Asian isolate 122.4 µg/mL vs. African isolate | Tectona grandis | [50] |
| Benzene 1-carboxylic acid hexadecanoate (BHCD) | Inhibition of viral entry | Vero cells | E1 CHIKV envelope glycoprotein | 3.04 µg/mL vs. Asian isolate 76.46 µg/mL vs. African isolate | Tectona grandis | [50] |

Table A2.
Plant-derived compounds targeting specific proteins involved in viral replication and maturation.
Table A2.
Plant-derived compounds targeting specific proteins involved in viral replication and maturation.
| Compound | Target | Cell Type Tested | IC50/EC50 | Plant | Reference |
|---|---|---|---|---|---|
| Coronaviruses (SARS-CoV, SARS-CoV-2, MERS-CoV) 3CLPro | |||||
| 3′-(3-Methylbut-2-enyl)-3′,4′,7-trihydroxyflavane | MERS-CoV-3CLPro | N/A | 34.7 µM | Broussonetia papyrifera | [72] |
| 4-Hydroxyisolonchocarpin | MERS-CoV-3CLPro | N/A | 193.7 µM | Broussonetia papyrifera | [72] |
| Broussochalcone A | MERS-CoV-3CLPro | N/A | 36.2 µM | Broussonetia papyrifera | [72] |
| Broussochalcone B | MERS-CoV-3CLPro | N/A | 27.9 µM | Broussonetia papyrifera | [72] |
| Broussoflavan A | MERS-CoV-3CLPro | N/A | 125.7 µM | Broussonetia papyrifera | [72] |
| Isoliquiritigenin | MERS-CoV-3CLPro | N/A | 33.9 µM | Broussonetia papyrifera | [72] |
| Kaempferol | MERS-CoV-3CLPro | N/A | 35.3 µM | Broussonetia papyrifera | [72] |
| Kazinol A | MERS-CoV-3CLPro | N/A | 66.2 µM | Broussonetia papyrifera | [72] |
| Kazinol B | MERS-CoV-3CLPro | N/A | 31.4 µM | Broussonetia papyrifera | [72] |
| Kazinol F | MERS-CoV-3CLPro | N/A | 135.0 µM | Broussonetia papyrifera | [72] |
| Kazinol J | MERS-CoV-3CLPro | N/A | 109.2 µM | Broussonetia papyrifera | [72] |
| Papyriflavonol A | MERS-CoV-3CLPro | N/A | 64.5 µM | Broussonetia papyrifera | [72] |
| Quercetin | MERS-CoV-3CLPro | N/A | 34.8 µM | Broussonetia papyrifera | [72] |
| Quercetin-β-galactoside | MERS-CoV-3CLPro | N/A | 68.0 µM | Broussonetia papyrifera | [72] |
| 3′-(3-Methylbut-2-enyl)-3′,4′,7-trihydroxyflavane | SARS-CoV-3CLPro | N/A | 30.2 µM | Broussonetia papyrifera | [72] |
| 3-isotheaflavin-3-gallate | SARS-CoV-3CLPro | N/A | 7 µM | Camellia sinensis | [80] |
| 4-Hydroxyisolonchocarpin | SARS-CoV-3CLPro | N/A | 202.7 µM | Broussonetia papyrifera | [72] |
| Aloe emodin | SARS-CoV-3CLPro | Vero cells | 132 μM | Isatis Indigotica | [279] |
| Amentoflavone | SARS-CoV-3CLPro | N/A | 8.3 μM | Torreya nucifera | [73] |
| Apigenin | SARS-CoV-3CLPro | N/A | 280.8 μM | Torreya nucifera | [73] |
| beta-sitosterol | SARS-CoV-3CLPro | Vero cells | 115 μM | Isatis indigotica | [279] |
| Betulinic acid | SARS-CoV-3CLPro | Vero E6 | 8.2 μM | Betula pubescens | [280] |
| Broussochalcone A | SARS-CoV-3CLPro | N/A | 88.1 µM | Broussonetia papyrifera | [72] |
| Broussochalcone B | SARS-CoV-3CLPro | N/A | 57.8 µM | Broussonetia papyrifera | [72] |
| Broussoflavan A | SARS-CoV-3CLPro | N/A | 92.4 µM | Broussonetia papyrifera | [72] |
| Chalcone | SARS-CoV-3CLPro | N/A | 11.4 µM | Angelica keiskei | [72] |
| Daidzein | SARS-CoV-3CLPro | Vero cells | 105 μM | Isatis Indigotica | [279] |
| Dihydrotanshinone I | SARS-CoV-3CLPro | N/A | 14.4 µM | Salvia miltiorrhiza | [281] |
| Herbacetin | SARS-CoV-3CLPro | N/A | 33.17 µM | Rhodiola Rosea | [282] |
| Hesperetin | SARS-CoV-3CLPro | Vero cells | 60 μM | Isatis Indigotica | [279] |
| Hirsutanolol | SARS-CoV-3CLPro | N/A | 105.6 µM | Alnus japonica | [283] |
| Hirsutenone | SARS-CoV-3CLPro | N/A | 36.2 µM | Alnus japonica | [283] |
| Indigo | SARS-CoV-3CLPro | Vero cells | 300 μM | Isatis indigotica | [279] |
| Indirubin | SARS-CoV-3CLPro | Vero cells | 293 μM | Isatis indigotica | [279] |
| Isoliquiritigenin | SARS-CoV-3CLPro | N/A | 61.9 µM | Broussonetia papyrifera | [72] |
| Kaempferol | SARS-CoV-3CLPro | N/A | 116.3 µM | Broussonetia papyrifera | [72] |
| Kazinol A | SARS-CoV-3CLPro | N/A | 84.8 µM | Broussonetia papyrifera | [72] |
| Kazinol B | SARS-CoV-3CLPro | N/A | 233.3 µM | Broussonetia papyrifera | [72] |
| Kazinol F | SARS-CoV-3CLPro | N/A | 43.3 µM | Broussonetia papyrifera | [72] |
| Kazinol J | SARS-CoV-3CLPro | N/A | 64.2 µM | Broussonetia papyrifera | [72] |
| Methyl tanshinoate | SARS-CoV-3CLPro | N/A | 21.1 µM | Salvia miltiorrhiza | [281] |
| Oregonin | SARS-CoV-3CLPro | N/A | 129.5 µM | Alnus japonica | [281] |
| Papyriflavonol A | SARS-CoV-3CLPro | N/A | 103.6 µM | Broussonetia papyrifera | [72] |
| Pectolinarin | SARS-CoV-3CLPro | N/A | 37.78 µM | Cirsium heterophyllum | [282] |
| Quercetin | SARS-CoV-3CLPro | N/A | 23.8 μM | Torreya nucifera | [73] |
| Quercetin-β-galactoside | SARS-CoV-3CLPro | N/A | 128.8 µM | Broussonetia papyrifera | [72] |
| Rhoifolin | SARS-CoV-3CLPro | N/A | 27.45 µM | Citrus limon | [282] |
| Rosmariquinone | SARS-CoV-3CLPro | N/A | 21.1 µM | Salvia miltiorrhiza | [281] |
| Rubranol | SARS-CoV-3CLPro | N/A | 144.6 µM | Alnus japonica | [283] |
| Rubranoside A | SARS-CoV-3CLPro | N/A | 102.1 µM | Alnus japonica | [283] |
| Rubranoside B | SARS-CoV-3CLPro | N/A | 105.3 µM | Alnus japonica | [283] |
| Sinigrin | SARS-CoV-3CLPro | Vero cells | 121 μM | Isatis indigotica | [279] |
| Tannic acid | SARS-CoV-3CLPro | N/A | 3 µM | Rhus Coriaria | [80] |
| Tanshinone I | SARS-CoV-3CLPro | N/A | 38.7 µM | Salvia miltiorrhiza | [281] |
| Tanshinone IIA | SARS-CoV-3CLPro | N/A | 89.1 µM | Salvia miltiorrhiza | [281] |
| Tanshinone IIB | SARS-CoV-3CLPro | N/A | 24.8 µM | Salvia miltiorrhiza | [281] |
| Theaflavin | SARS-CoV-3CLPro | N/A | 56.0 µM | Camellia Sinensis | [80] |
| Theaflavin-3,3′-digallate | SARS-CoV-3CLPro | N/A | 9.5 µM | Camellia Sinensis | [80] |
| Baicalein | SARS-CoV-2-3CLPro | N/A | 0.94 μM | Scutellaria baicalensis | [67] |
| Baicalin | SARS-CoV-2-3CLPro | N/A | 6.41 μM | Scutellaria baicalensis | [67] |
| β-carotene | SARS-CoV-2-3CLPro | N/A | 17.54 μM | Vaccinium Oxycoccos | [82] |
| Cyanidin 3-O-galactoside | SARS-CoV-2-3CLPro | N/A | 9.98 μM | Vaccinium Oxycoccos | [82] |
| Epicatechin | SARS-CoV-2-3CLPro | N/A | 12.54 μM | Vaccinium Oxycoccos | [82] |
| Isoschaftoside | SARS-CoV-2-3CLPro | N/A | 30.22 μM | Camellia Sinensis | [74] |
| Kaempferol-3-O-gentiobioside | SARS-CoV-2-3CLPro | N/A | 35.89 μM | Camellia Sinensis | [74] |
| Narcissoside | SARS-CoV-2-3CLPro | N/A | 38.14 μM | Zygophyllum simplex | [74] |
| Rutin | SARS-CoV-2-3CLPro | N/A | 31.26 μM | Fagopyrum tataricum | [74] |
| Vicenin-2 | SARS-CoV-2-3CLPro | N/A | 38.86 μM | Citrus Reticulata | [74] |
| PLPro | |||||
| Kazinol F | MERS-CoV-PLpro | N/A | 39.5 µM | Broussonetia papyrifera | [72] |
| Broussochalcone A | MERS-CoV-PLpro | N/A | 42.1 µM | Broussonetia papyrifera | [72] |
| Broussoflavan A | MERS-CoV-PLpro | N/A | 49.1 µM | Broussonetia papyrifera | [72] |
| Isoliquiritigenin | MERS-CoV-PLpro | N/A | 82.2 µM | Broussonetia papyrifera | [72] |
| Kazinol A | MERS-CoV-PLpro | N/A | 88.5 µM | Broussonetia papyrifera | [72] |
| Kazinol B | MERS-CoV-PLpro | N/A | 94.9 µM | Broussonetia papyrifera | [72] |
| Kazinol J | MERS-CoV-PLpro | N/A | 55.0 µM | Broussonetia papyrifera | [72] |
| 3′-(3-methylbut-2-enyl)-3′,4′,7-trihydroxyflavane | MERS-CoV-PLpro | N/A | 48.8 µM | Broussonetia papyrifera | [72] |
| Cryptotanshinone | SARS-CoV-PLPro | N/A | 0.8 µM | Salvia miltiorrhiza | [281] |
| Diplacone | SARS-CoV-PLPro | N/A | 10.4 µM | Paulownia tomentosa | [284] |
| Broussochalcone A | SARS-CoV-PLPro | N/A | 9.2 µM | Broussonetia papyrifera | [72] |
| Broussoflavan A | SARS-CoV-PLPro | N/A | 30.4 µM | Broussonetia papyrifera | [72] |
| Chalcone | SARS-CoV-PLPro | Vero cells | 1.2 µM | Angelica keiskei | [285] |
| Curcumin | SARS-CoV-PLPro | N/A | 5.7 µM | Curcuma longa | [283] |
| Dihydrotanshinone I | SARS-CoV-PLPro | N/A | 4.9 µM | Salvia miltiorrhiza | [281] |
| 6-geranyl-4′,5,7-trihydroxy-3′,5′-dimethoxyflavanone | SARS-CoV-PLPro | N/A | 13.9 µM | Paulownia tomentosa | [284] |
| Hirsutanonol 5 | SARS-CoV-PLPro | N/A | 7.8 µM | Alnus japonica | [283] |
| Hirsutenone 2 | SARS-CoV-PLPro | N/A | 4.1 µM | Alnus japonica | [283] |
| 4-hydroxyisolonchocarpin | SARS-CoV-PLPro | N/A | 35.4 µM | Broussonetia papyrifera | [72] |
| Isoliquiritigenin | SARS-CoV-PLPro | N/A | 24.6 µM | Broussonetia papyrifera | [72] |
| Kaempferol | SARS-CoV-PLPro | N/A | 16.3 µM | Broussonetia papyrifera | [72] |
| Kazinol A | SARS-CoV-PLPro | N/A | 66.2 µM | Broussonetia papyrifera | [72] |
| Kazinol B | SARS-CoV-PLPro | N/A | 31.4 µM | Broussonetia papyrifera | [72] |
| Kazinol F | SARS-CoV-PLPro | N/A | 27.8 µM | Broussonetia papyrifera | [72] |
| Kazinol J | SARS-CoV-PLPro | N/A | 15.2 µM | Broussonetia papyrifera | [72] |
| 3′-(3-methylbut-2-enyl)-3′,4′,7-trihydroxyflavane | SARS-CoV-PLPro | N/A | 35.8 µM | Broussonetia papyrifera | [72] |
| 3′-O-methyldiplacol | SARS-CoV-PLPro | N/A | 9.5 µM | Paulownia tomentosa | [284] |
| 4′-O-methyldiplacol | SARS-CoV-PLPro | N/A | 9.2 µM | Paulownia tomentosa | [284] |
| 3′-O-methyldiplacone | SARS-CoV-PLPro | N/A | 13.2 µM | Paulownia tomentosa | [284] |
| 4′-O-methyldiplacone | SARS-CoV-PLPro | N/A | 12.7 µM | Paulownia tomentosa | [284] |
| Mimulone | SARS-CoV-PLPro | N/A | 14.4 µM | Paulownia tomentosa | [284] |
| Oregonin | SARS-CoV-PLPro | N/A | 20.1 µM | Alnus japonica | [283] |
| Tanshinone IIA | SARS-CoV-PLPro | N/A | 1.6 µM | Salvia miltiorrhiza | [281] |
| Tomentin A | SARS-CoV-PLPro | N/A | 6.2 µM | Paulownia tomentosa | [284] |
| Tomentin B | SARS-CoV-PLPro | N/A | 6.1 µM | Paulownia tomentosa | [284] |
| Tomentin C | SARS-CoV-PLPro | N/A | 11.6 µM | Paulownia tomentosa | [284] |
| Tomentin D | SARS-CoV-PLPro | N/A | 12.5 µM | Paulownia tomentosa | [284] |
| Tomentin E | SARS-CoV-PLPro | N/A | 5.0 µM | Paulownia tomentosa | [284] |
| Rubranol | SARS-CoV-PLPro | N/A | 12.3 µM | Alnus japonica | [283] |
| Rubranoside A | SARS-CoV-PLPro | N/A | 9.1 µM | Alnus japonica | [283] |
| Rubranoside B | SARS-CoV-PLPro | N/A | 8.0 µM | Alnus japonica | [283] |
| Papyriflavonol A | SARS-CoV PLPro | N/A | 3.7 µM | Broussonetia papyrifera | [72] |
| Quercetin | SARS-CoV PLPro | N/A | 8.6 µM | Broussonetia papyrifera | [72] |
| Quercetin-β-galactoside | SARS-CoV PLPro | N/A | 51.9 µM | Broussonetia papyrifera | [72] |
| Broussochalcone B | SARS-CoV-PLPro | N/A | 11.6 µM | Broussonetia papyrifera | [72] |
| RdRp | |||||
| Amentoflavone | SARS-CoV-2 RdRp | RD cells | 13.17 µM | Selaginella tamariscina | [77] |
| Baicalein | SARS-CoV-2 RdRp | Vero CCL-81 | 4.5 µM | Scutellaria baicalensis | [69] |
| Baicalin | SARS-CoV-2 RdRp | Vero CCL-81 | 9 µM | Scutellaria baicalensis | [69] |
| Corilagin | SARS-CoV-2 RdRp | Vero CCL-81 | 0.13 µM | Caesalpinia coriaria | [286] |
| Luteolin | SARS-CoV-2 RdRp | N/A | 4.6 µM | Apium Graveolens | [287] |
| Lycorine | MERS-CoV RdRp | Vero CCL-81 | 1.41 µM | Lycoris Radiata | [87] |
| Lycorine | SARS-CoV RdRp | Vero CCL-81 | 1.02 µM | Lycoris Radiata | [87] |
| Lycorine | SARS-CoV-2 RdRp | Vero CCL-81 | 0.88 µM | Lycoris Radiata | [87] |
| Quercetin | SARS-CoV-2 RdRp | N/A | 6.9 µM | Alium Cepa | [287] |
| SARS-CoV nsp13-Helicase/ATPase activity | |||||
| Myricetin | nsp13 ATPase | N/A | 2.71 µM | N/A | [78] |
| Scutellarein | nsp13 ATPase | N/A | 0.86 µM | Scutellaria baicalensis | [78] |
| Baicalein | nsp13 ATPase | N/A | 0.47 µM | Scutellaria baicalensis | [288] |
| Baicalein | nsp13 helicase | Vero E6 | 2.9 µM | Scutellaria baicalensis | [70] |
| Dihydro-myricetin | nsp13 helicase | Vero E6 | 25.6 µM | N/A | [70] |
| Diosmetin | nsp13 helicase | Vero E6 | 10.6 µM | Vicia cracca | [70] |
| Ellagic acid | nsp13 ATPase/helicase | N/A | 2.8 µM | N/A | [83] |
| Flavanone | nsp13 helicase | Vero E6 | 0.52 µM | N/A | [70] |
| Flavanone-7-O-glucoside | nsp13 helicase | Vero E6 | 2.88 µM | N/A | [70] |
| (−)-Gallocatechin gallate | nsp13 helicase | N/A | 1.34 µM | Camellia sinensis | [83] |
| Licoflavone C | nsp13 helicase | Vero E6 | 1.34 µM | Genista ephedroides | [70] |
| Kaempferol | nsp13 helicase | Vero E6 | 0.76 µM | N/A | [70] |
| Katacine | nsp13 helicase | N/A | 5.98 µM | Polygonum coriarium | [83] |
| Licoflavone C | nsp13 ATPase (in the presence of BSA, TCEP and polyrA) | Vero E6 | 18.3 µM | Genista ephedroides | [70] |
| Linoleic acid | nsp13 helicase/ATPase | N/A | 4.3 µM | N/A | [289] |
| Myricetin | nsp13 helicase | Vero E6 | 0.41 µM | N/A | [70] |
| Oleic acid | nsp13 helicase/ATPase | N/A | 14 µM | N/A | [289] |
| Gossypol | nsp13 helicase/ATPase | N/A | 1.3 µM | Gossypium spp. | [289] |
| Prunetin | nsp13 helicase | Vero E6 | 11.5 µM | Prunus emarginata | [70] |
| Punicalagin | nsp13 helicase | Vero cells, A549-ACE2 | 0.43 µM | Punica granatum | [83] |
| Quercetin | nsp13 helicase | Vero E6 | 0.53 µM | N/A | [70] |
| Rhodiosin | nsp13 helicase | N/A | 0.48 µM | Rhodiola spp. | [83] |
| Rosmanol | nsp13 helicase | N/A | 8.93 µM | Rosmarinus officinalis L. | [83] |
| Tannic acid | nsp13 helicase | N/A | 1.25 µM | N/A | [83] |
| Wogonin | nsp13 helicase | Vero E6 | 24.9 µM | Scutellaria baicalensis | [70] |
| Dengue virus | |||||
| Sotetsuflavone | NS5 RdRp | N/A | 0.16 µM | Dacrydium araucarioides | [107] |
| Apigenin | NS5 RdRp and restores STAT2 inhibition by NS5 | IFN-I competent Huh7 cells, engineered K562 cell platform | EC50 29.7 µM | N/A | [108] |
| Luteolin | NS5 RdRp and restores STAT2 inhibition by NS5 | IFN-I competent Huh7 cells, engineered K562 cell platform | EC50 9.2 µM | N/A | [108] |
| Bisdemethoxycurcumin | NS2B/NS3 protease (DENV2) | BHK-21 cells | 36.23 µM | Curcuma longa | [110] |
| Curcumin | NS2B/NS3 protease (DENV2) | BHK-21 cells | 66.0 µM | Curcuma longa | [110] |
| Myricetin | NS2B/NS3 protease | N/A | 8.46 µM | N/A | [109] |
| Influenza A | |||||
| Apigenin | Neuraminidase | MDCK | 33 µM | Rhodiola rosea roots | [123] |
| Astragalin | Neuraminidase | MDCK | 38 µM | Rhodiola rosea roots | [123] |
| Cosmosiin | Neuraminidase | MDCK | 47 µM | Rhodiola rosea roots | [123] |
| Demethoxymatteucinol | Neuraminidase | MDCK | 30 µM | Pentarhizidium orientale | [125] |
| Gossypetin | Neuraminidase | MDCK | 3 µM | Rhodiola rosea roots | [123] |
| Herbacetin | Neuraminidase | MDCK | 9 µM | Rhodiola rosea roots | [123] |
| Hispidulin | Neuraminidase | MDCK | 19.83 µM | Salvia plebeia R. Br | [124] |
| 3′-hydroxy-5′-methoxy-6,8-dimethylhuazhongilexone | Neuraminidase | MDCK | 24 µM | Pentarhizidium orientale | [125] |
| Kaempferol | Neuraminidase | MDCK | 11 µM | Rhodiola rosea roots | [123] |
| Linocinamarin | Neuraminidase | MDCK | 44 µM | Rhodiola rosea roots | [123] |
| Luteolin | Neuraminidase | MDCK | 17.96 µM | Salvia plebeia R. Br | [124] |
| Matteucin | Neuraminidase | MDCK | 24 µM | Pentarhizidium orientale | [125] |
| Matteucinol | Neuraminidase | MDCK | 25 µM | Pentarhizidium orientale | [125] |
| Methoxymatteucin | Neuraminidase | MDCK | 25 µM | Pentarhizidium orientale | [125] |
| Nicotiflorin | Neuraminidase | MDCK | 32 µM | Rhodiola rosea roots | [123] |
| Quercetin | Neuraminidase | MDCK | 2 µM | Rhodiola rosea roots | [123] |
| Rosmarinic acid methyl ester | Neuraminidase | MDCK | 16.65 µM | Salvia plebeia R. Br | [124] |
| Rutin | Neuraminidase | MDCK | 34 µM | Rhodiola rosea roots | [123] |
| Rhodiolinin | Neuraminidase | MDCK | 10 µM | Rhodiola rosea roots | [123] |
| Rhodionin | Neuraminidase | MDCK | 32 µM | Rhodiola rosea roots | [123] |
| Rhodiosin | Neuraminidase | MDCK | 57 µM | Rhodiola rosea roots | [123] |
| Nepetin | Neuraminidase | MDCK | 11.18 µM | Salvia plebeia R. Br | [124] |
| 2′,4′dihydroxy-6′-methoxy-3′,5′-dimethylchalcone | Neuraminidase | HEK293, MDCK | 8.23 µM (H1N1) 5.07 µM (H9N2) 7.02 μM (H1N1 WT) 8.84 μM (H1N1-H274Y mutation) | Cleistocalyx operculatus | [126] |
| Myricetin-3′,5′-dimethylether-3-O-β-D-galactopyranoside | Neuraminidase | HEK293, MDCK | 8.86 µM (H1N1) 6.50 µM (H9N2) 7.10 μM (H1N1 WT) 9.34 μM (H1N1-H274Y mutation) | Cleistocalyx operculatus | [126] |
| Berberine | HAE cells, blocks nuclear export of IAV ribonucleoprotein to cytoplasm | MDCK, A549, LET1, HAE | 16 µM | Berberis sp. | [129] |
| HIV-1 | |||||
| Oleanolic acid | Protease | N/A | 10 µg/mL | Xanthoceras sorbifolia | [290] |
| 3-oxotirucalla-7, 24-dien-21-oic acid | Protease | N/A | 20 µg/mL | Xanthoceras sorbifolia | [290] |
| Apigenin | Integrase | N/A | 22 µM | Punica granatum | [94] |
| Ellagic acid | Integrase | N/A | 0.075 µM | Punica granatum | [94] |
| Betulinic acid | Integrase | N/A | 96.5 µM | Punica granatum | [94] |
| Kuwanon-L | RT-associated RDDP | TZM-bl (modified HeLa) | 0.99 µM | Xanthocer assorbifolia | [291] |
| Kuwanon-L | RT-associated RNase | TZM-bl (modified HeLa) | 0.57 µM | Xanthocer assorbifolia | [291] |
| Luteolin | Integrase | N/A | 6.5 µM | Punica granatum | [94] |
| Luteolin 7-O-glucoside | Integrase | N/A | 8.5 µM | Punica granatum | [94] |
| Punicalins | Integrase | N/A | 0.09 µM | Punica granatum | [94] |
| Punicalagins | Integrase | N/A | 0.065 µM | Punica granatum | [94] |
| Corilagin | Reverse transcriptase | MT4 T-lymphoid, MAGI cells | 9.3 µM | Phyllanthus amarus | [93] |
| Norisoboldine | Reverse transcriptase | N/A | 153.7 μg/mL | Croton echinocarpus | [292] |
| L-chicoric acid | Reverse transcriptase (presence of heteropolymeric template) | MT-2 | 17 µM | Echinacea purpurea | [92] |
| Geraniin | Reverse transcriptase | MT4 T-lymphoid, MAGI cells | 1.9 µM | Phyllanthus amarus | [93] |
| 1-methoxyoxalyl-3,5-DCQA | Reverse transcriptase (presence of heteropolymeric template) | MT-2 | 7 µM | Echinacea spp. | [92] |
| Apigenin | RT-associated RNase | N/A | 16.1 µM | Punica granatum | [94] |
| Betulinic acid | RT-associated RNase | N/A | 2.0 µM | Punica granatum | [94] |
| Ellagic acid | RT-associated RNase | N/A | 1.4 µM | Punica granatum | [94] |
| Luteolin | RT-associated RNase | N/A | 3.7 µM | Punica granatum | [94] |
| Oleanolic acid | RT-associated RNase | N/A | 6.7 µM | Punica granatum | [94] |
| Punicalins | RT-associated RNase | N/A | 0.18 µM | Punica granatum | [94] |
| Punicalagins | RT-associated RNase | N/A | 0.12 µM | Punica granatum | [94] |
| Ursolic acid | RT-associated RNase | N/A | 5.7 µM | Punica granatum | [94] |
| 3-O-(3′,3′-dimethylsuccinyl) betulinic acid—Bevirimat | p25-to-p24 conversion | PBMC, MT-2 | 10.3 nM | Syzygium claviflorum | [95] |
| Zika virus | |||||
| Astragalin | NS2B-NS3Pro | N/A | 112.0 µM | Phytolacca americana | [101] |
| Epicatechin gallate | NS2B-NS3Pro | N/A | 98.0 µM | Camellia sinensis | [101] |
| Epigallocatechin gallate | NS2B-NS3Pro | N/A | 87.0 µM | Camellia sinensis | [101] |
| Gallocatechin gallate | NS2B-NS3Pro | N/A | 99.0 µM | Camellia sinensis | [101] |
| Luteolin | NS2B-NS3Pro | N/A | 53.0 µM | N/A | [101] |
| Myricetin | NS2B-NS3Pro | N/A | 22.0 µM | N/A | [101] |
| Rutin | NS2B-NS3Pro | N/A | 112.0 µM | N/A | [101] |
| (−)-epigallocatechin-3-gallate | NS3 helicase–ATPase activity | N/A | 2.95 µM | Camellia sinensis | [102] |
| Chikungunya virus | |||||
| Berberine | Host MAPK pathway | HEK293T | 4.5 µM | Anamirta cocculus | [115] |
| Harringtonine | Nsp2 protease | BHK-21 | 0.24 µM | Cephalotaxus harringtonia | [114] |
| Tomatidine | Nsp2 protease | Huh7 | 1.3 µM | Solanum dulcamara | [293] |
| Apigenin | Replicase complex | BHK-21 | 70.8 µM | N/A | [113] |
| Chrysin | Replicase complex | BHK-21 | 126.6 µM | N/A | [113] |
| Naringenin | Replicase complex | BHK-21 | 118.4 µM | N/A | [113] |
| Silybin | Replicase complex | BHK-21 | 92.3 µM | N/A | [113] |
| Withaferin A | Nsp2 protease | BHK-21 | 0.51 µM | Withania somnifera | [294] |
| Prostratin | Not established/replication machinery | Vero cells, BGM, HEL | 8 µM (Vero) 7.6 µM (BGM) 7.1 µM (HEL) | Trigonostemon howii | [116] |
| Baicalein | Replicase complex | Vero cells | 7 µM | Scutellaria baicalensis | [48] |
| Fisetin | Replicase complex | Vero cells | 29.5 µM | N/A | [48] |
| Quercetagetin | Replicase complex | Vero cells | 43.52 µM | N/A | [48] |
| Silymarin | Replicase complex | Vero cells | 16.9 µg/mL | Silybum marianum | [117] |
| Trigocherrierin A | CHIKV-induced cell death | Vero cells | 0.6 µM | Trigonostemon cherrieri | [295] |
References
- Han, J.J.; Song, H.A.; Pierson, S.L.; Shen-Gunther, J.; Xia, Q. Emerging Infectious Diseases Are Virulent Viruses—Are We Prepared? An Overview. Microorganisms 2023, 11, 2618. [Google Scholar] [CrossRef] [PubMed]
- Baker, R.E.; Mahmud, A.S.; Miller, I.F.; Rajeev, M.; Rasambainarivo, F.; Rice, B.L.; Takahashi, S.; Tatem, A.J.; Wagner, C.E.; Wang, L.-F.; et al. Infectious Disease in an Era of Global Change. Nat. Rev. Microbiol. 2022, 20, 193–205. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Takova, K.; Tonova, V.; Koynarski, T.; Lukov, L.L.; Minkov, I.; Pishmisheva, M.; Kotsev, S.; Tsachev, I.; Baymakova, M.; et al. The Re-Emergence of Hepatitis E Virus in Europe and Vaccine Development. Viruses 2023, 15, 1558. [Google Scholar] [CrossRef] [PubMed]
- Bhadoria, P.; Gupta, G.; Agarwal, A. Viral Pandemics in the Past Two Decades: An Overview. J. Fam. Med. Prim. Care 2021, 10, 2745–2750. [Google Scholar] [CrossRef] [PubMed]
- Fatima, M.; Hong, K.-J. Innovations, Challenges, and Future Prospects for Combination Vaccines Against Human Infections. Vaccines 2025, 13, 335. [Google Scholar] [CrossRef] [PubMed]
- Vo, D.-K.; Trinh, K.T.L. Molecular Farming for Immunization: Current Advances and Future Prospects in Plant-Produced Vaccines. Vaccines 2025, 13, 191. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.-W.; Zahmanova, G.; Minkov, I.; Lomonossoff, G.P. Plant-Based Expression and Characterization of SARS-CoV-2 Virus-like Particles Presenting a Native Spike Protein. Plant Biotechnol. J. 2022, 20, 1363–1372. [Google Scholar] [CrossRef] [PubMed]
- Trivedi, P.; Abbas, A.; Lehmann, C.; Rupasinghe, H.P.V. Antiviral and Anti-Inflammatory Plant-Derived Bioactive Compounds and Their Potential Use in the Treatment of COVID-19-Related Pathologies. J. Xenobiotics 2022, 12, 289–306. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, R.; Dubey, N.K.; Sharma, M.; Kharkwal, H.; Bajpai, R.; Srivastava, R. Boosting the Human Antiviral Response in Conjunction with Natural Plant Products. Front. Nat. Prod. 2025, 3, 1470639. [Google Scholar] [CrossRef]
- Riaz, M.; Khalid, R.; Afzal, M.; Anjum, F.; Fatima, H.; Zia, S.; Rasool, G.; Egbuna, C.; Mtewa, A.G.; Uche, C.Z.; et al. Phytobioactive Compounds as Therapeutic Agents for Human Diseases: A Review. Food Sci. Nutr. 2023, 11, 2500–2529. [Google Scholar] [CrossRef] [PubMed]
- Popoola, T.D.; Segun, P.A.; Ekuadzi, E.; Dickson, R.A.; Awotona, O.R.; Nahar, L.; Sarker, S.D.; Fatokun, A.A. West African Medicinal Plants and Their Constituent Compounds as Treatments for Viral Infections, Including SARS-CoV-2/COVID-19. Daru J. Fac. Pharm. Tehran Univ. Med. Sci. 2022, 30, 191–210. [Google Scholar] [CrossRef] [PubMed]
- Atampugbire, G.; Adomako, E.E.A.; Quaye, O. Medicinal Plants as Effective Antiviral Agents and Their Potential Benefits. Nat. Prod. Commun. 2024, 19, 1934578X241282923. [Google Scholar] [CrossRef]
- Thomas, E.; Stewart, L.E.; Darley, B.A.; Pham, A.M.; Esteban, I.; Panda, S.S. Plant-Based Natural Products and Extracts: Potential Source to Develop New Antiviral Drug Candidates. Molecules 2021, 26, 6197. [Google Scholar] [CrossRef] [PubMed]
- BioRender. Available online: https://app.biorender.com/user/signin (accessed on 18 June 2025).
- Wang, S.; Li, W.; Wang, Z.; Yang, W.; Li, E.; Xia, X.; Yan, F.; Chiu, S. Emerging and Reemerging Infectious Diseases: Global Trends and New Strategies for Their Prevention and Control. Signal Transduct. Target. Ther. 2024, 9, 223. [Google Scholar] [CrossRef] [PubMed]
- Gelderblom, H.R. Structure and Classification of Viruses. In Medical Microbiology; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996; ISBN 978-0-9631172-1-2. [Google Scholar]
- Kesheh, M.M.; Hosseini, P.; Soltani, S.; Zandi, M. An Overview on the Seven Pathogenic Human Coronaviruses. Rev. Med. Virol. 2022, 32, e2282. [Google Scholar] [CrossRef] [PubMed]
- Schoeman, D.; Fielding, B.C. Coronavirus Envelope Protein: Current Knowledge. Virol. J. 2019, 16, 69. [Google Scholar] [CrossRef] [PubMed]
- Munster, V.J.; Koopmans, M.; van Doremalen, N.; van Riel, D.; de Wit, E. A Novel Coronavirus Emerging in China—Key Questions for Impact Assessment. N. Engl. J. Med. 2020, 382, 692–694. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Li, K.; Huang, S.; Ivanov, K.I.; Yang, S.; Ji, Y.; Zhang, H.; Wu, W.; He, Y.; Zeng, Q.; et al. Anti-SARS-CoV-2 Prodrug ATV006 Has Broad-Spectrum Antiviral Activity against Human and Animal Coronaviruses. Acta Pharm. Sin. B 2025, 15, 2498–2510. [Google Scholar] [CrossRef] [PubMed]
- Wong, N.A.; Saier, M.H. The SARS-Coronavirus Infection Cycle: A Survey of Viral Membrane Proteins, Their Functional Interactions and Pathogenesis. Int. J. Mol. Sci. 2021, 22, 1308. [Google Scholar] [CrossRef] [PubMed]
- He, C.-L.; Huang, L.-Y.; Wang, K.; Gu, C.-J.; Hu, J.; Zhang, G.-J.; Xu, W.; Xie, Y.-H.; Tang, N.; Huang, A.-L. Identification of Bis-Benzylisoquinoline Alkaloids as SARS-CoV-2 Entry Inhibitors from a Library of Natural Products. Signal Transduct. Target. Ther. 2021, 6, 131. [Google Scholar] [CrossRef] [PubMed]
- Keyaerts, E.; Vijgen, L.; Pannecouque, C.; Van Damme, E.; Peumans, W.; Egberink, H.; Balzarini, J.; Van Ranst, M. Plant Lectins Are Potent Inhibitors of Coronaviruses by Interfering with Two Targets in the Viral Replication Cycle. Antiviral Res. 2007, 75, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Invernizzi, L.; Moyo, P.; Cassel, J.; Isaacs, F.J.; Salvino, J.M.; Montaner, L.J.; Tietjen, I.; Maharaj, V. Use of Hyphenated Analytical Techniques to Identify the Bioactive Constituents of Gunnera perpensa L., a South African Medicinal Plant, Which Potently Inhibit SARS-CoV-2 Spike Glycoprotein-Host ACE2 Binding. Anal. Bioanal. Chem. 2022, 414, 3971–3985. [Google Scholar] [CrossRef] [PubMed]
- Mei, J.; Zhou, Y.; Yang, X.; Zhang, F.; Liu, X.; Yu, B. Active Components in Ephedra Sinica Stapf Disrupt the Interaction between ACE2 and SARS-CoV-2 RBD: Potent COVID-19 Therapeutic Agents. J. Ethnopharmacol. 2021, 278, 114303. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.-Y.; Wu, S.-L.; Chen, J.-C.; Li, C.-C.; Hsiang, C.-Y. Emodin Blocks the SARS Coronavirus Spike Protein and Angiotensin-Converting Enzyme 2 Interaction. Antiviral Res. 2007, 74, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Ohishi, T.; Hishiki, T.; Baig, M.S.; Rajpoot, S.; Saqib, U.; Takasaki, T.; Hara, Y. Epigallocatechin Gallate (EGCG) Attenuates Severe Acute Respiratory Coronavirus Disease 2 (SARS-CoV-2) Infection by Blocking the Interaction of SARS-CoV-2 Spike Protein Receptor-Binding Domain to Human Angiotensin-Converting Enzyme 2. PLoS ONE 2022, 17, e0271112. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.-C.; Chen, Y.; Wang, Y.-C.; Wang, W.-J.; Yang, C.-S.; Tsai, C.-L.; Hou, M.-H.; Chen, H.-F.; Shen, Y.-C.; Hung, M.-C. Tannic Acid Suppresses SARS-CoV-2 as a Dual Inhibitor of the Viral Main Protease and the Cellular TMPRSS2 Protease. Am. J. Cancer Res. 2020, 10, 4538–4546. [Google Scholar] [PubMed]
- Roy, A.V.; Chan, M.; Banadyga, L.; He, S.; Zhu, W.; Chrétien, M.; Mbikay, M. Quercetin Inhibits SARS-CoV-2 Infection and Prevents Syncytium Formation by Cells Co-Expressing the Viral Spike Protein and Human ACE2. Virol. J. 2024, 21, 29. [Google Scholar] [CrossRef] [PubMed]
- Cb, W.; Jc, T.; Rw, D. Molecular Mechanisms of HIV Entry. Adv. Exp. Med. Biol. 2012, 726, 223–242. [Google Scholar] [CrossRef]
- Li, B.Q.; Fu, T.; Dongyan, Y.; Mikovits, J.A.; Ruscetti, F.W.; Wang, J.M. Flavonoid Baicalin Inhibits HIV-1 Infection at the Level of Viral Entry. Biochem. Biophys. Res. Commun. 2000, 276, 534–538. [Google Scholar] [CrossRef] [PubMed]
- Agrelli, A.; de Moura, R.R.; Crovella, S.; Brandão, L.A.C. ZIKA Virus Entry Mechanisms in Human Cells. Infect. Genet. Evol. J. Mol. Epidemiol. Evol. Genet. Infect. Dis. 2019, 69, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Frabasile, S.; Koishi, A.C.; Kuczera, D.; Silveira, G.F.; Verri, W.A.; Duarte Dos Santos, C.N.; Bordignon, J. Corrigendum: The Citrus Flavanone Naringenin Impairs Dengue Virus Replication in Human Cells. Sci. Rep. 2017, 7, 43976. [Google Scholar] [CrossRef] [PubMed]
- Zandi, K.; Teoh, B.-T.; Sam, S.-S.; Wong, P.-F.; Mustafa, M.R.; Abubakar, S. Novel Antiviral Activity of Baicalein against Dengue Virus. BMC Complement. Altern. Med. 2012, 12, 214. [Google Scholar] [CrossRef] [PubMed]
- Oo, A.; Teoh, B.T.; Sam, S.S.; Bakar, S.A.; Zandi, K. Baicalein and Baicalin as Zika Virus Inhibitors. Arch. Virol. 2019, 164, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, E.; Teoh, B.-T.; Sam, S.-S.; Lani, R.; Hassandarvish, P.; Chik, Z.; Yueh, A.; Abubakar, S.; Zandi, K. Baicalin, a Metabolite of Baicalein with Antiviral Activity against Dengue Virus. Sci. Rep. 2014, 4, 5452. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Tai, W.; Wang, N.; Li, X.; Jiang, S.; Debnath, A.K.; Du, L.; Chen, S. Identification of Novel Natural Products as Effective and Broad-Spectrum Anti-Zika Virus Inhibitors. Viruses 2019, 11, 1019. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, B.M.; Batista, M.N.; Braga, A.C.S.; Nogueira, M.L.; Rahal, P. The Green Tea Molecule EGCG Inhibits Zika Virus Entry. Virology 2016, 496, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Mounce, B.C.; Cesaro, T.; Carrau, L.; Vallet, T.; Vignuzzi, M. Curcumin Inhibits Zika and Chikungunya Virus Infection by Inhibiting Cell Binding. Antiviral Res. 2017, 142, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Gaudry, A.; Bos, S.; Viranaicken, W.; Roche, M.; Krejbich-Trotot, P.; Gadea, G.; Desprès, P.; El-Kalamouni, C. The Flavonoid Isoquercitrin Precludes Initiation of Zika Virus Infection in Human Cells. Int. J. Mol. Sci. 2018, 19, 1093. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Zhao, Z.; Chai, Y.; Jin, X.; Li, C.; Yuan, F.; Liu, S.; Gao, Z.; Wang, H.; Song, J.; et al. Molecular Basis of Arthritogenic Alphavirus Receptor MXRA8 Binding to Chikungunya Virus Envelope Protein. Cell 2019, 177, 1714–1724.e12. [Google Scholar] [CrossRef] [PubMed]
- Wintachai, P.; Wikan, N.; Kuadkitkan, A.; Jaimipuk, T.; Ubol, S.; Pulmanausahakul, R.; Auewarakul, P.; Kasinrerk, W.; Weng, W.-Y.; Panyasrivanit, M.; et al. Identification of Prohibitin as a Chikungunya Virus Receptor Protein. J. Med. Virol. 2012, 84, 1757–1770. [Google Scholar] [CrossRef] [PubMed]
- Moller-Tank, S.; Kondratowicz, A.S.; Davey, R.A.; Rennert, P.D.; Maury, W. Role of the Phosphatidylserine Receptor TIM-1 in Enveloped-Virus Entry. J. Virol. 2013, 87, 8327–8341. [Google Scholar] [CrossRef] [PubMed]
- Silva, L.A.; Khomandiak, S.; Ashbrook, A.W.; Weller, R.; Heise, M.T.; Morrison, T.E.; Dermody, T.S. A Single-Amino-Acid Polymorphism in Chikungunya Virus E2 Glycoprotein Influences Glycosaminoglycan Utilization. J. Virol. 2014, 88, 2385–2397. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Santhosh, S.R.; Tiwari, M.; Lakshmana Rao, P.V.; Parida, M. Assessment of in Vitro Prophylactic and Therapeutic Efficacy of Chloroquine against Chikungunya Virus in Vero Cells. J. Med. Virol. 2010, 82, 817–824. [Google Scholar] [CrossRef] [PubMed]
- Weber, C.; Sliva, K.; von Rhein, C.; Kümmerer, B.M.; Schnierle, B.S. The Green Tea Catechin, Epigallocatechin Gallate Inhibits Chikungunya Virus Infection. Antiviral Res. 2015, 113, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Lani, R.; Hassandarvish, P.; Shu, M.-H.; Phoon, W.H.; Chu, J.J.H.; Higgs, S.; Vanlandingham, D.; Abu Bakar, S.; Zandi, K. Antiviral Activity of Selected Flavonoids against Chikungunya Virus. Antiviral Res. 2016, 133, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Wintachai, P.; Thuaud, F.; Basmadjian, C.; Roytrakul, S.; Ubol, S.; Désaubry, L.; Smith, D.R. Assessment of Flavaglines as Potential Chikungunya Virus Entry Inhibitors. Microbiol. Immunol. 2015, 59, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Sangeetha, K.; Purushothaman, I.; Rajarajan, S. Spectral Characterisation, Antiviral Activities, in Silico ADMET and Molecular Docking of the Compounds Isolated from Tectona grandis to Chikungunya Virus. Biomed. Pharmacother. 2017, 87, 302–310. [Google Scholar] [CrossRef] [PubMed]
- Dou, D.; Revol, R.; Östbye, H.; Wang, H.; Daniels, R. Influenza A Virus Cell Entry, Replication, Virion Assembly and Movement. Front. Immunol. 2018, 9, 1581. [Google Scholar] [CrossRef] [PubMed]
- Song, J.-M.; Lee, K.-H.; Seong, B.-L. Antiviral Effect of Catechins in Green Tea on Influenza Virus. Antiviral Res. 2005, 68, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Li, R.; Li, X.; He, J.; Jiang, S.; Liu, S.; Yang, J. Quercetin as an Antiviral Agent Inhibits Influenza A Virus (IAV) Entry. Viruses 2015, 8, 6. [Google Scholar] [CrossRef] [PubMed]
- Roschek, B.; Fink, R.C.; McMichael, M.D.; Li, D.; Alberte, R.S. Elderberry Flavonoids Bind to and Prevent H1N1 Infection in Vitro. Phytochemistry 2009, 70, 1255–1261. [Google Scholar] [CrossRef] [PubMed]
- Palamara, A.T.; Nencioni, L.; Aquilano, K.; De Chiara, G.; Hernandez, L.; Cozzolino, F.; Ciriolo, M.R.; Garaci, E. Inhibition of Influenza A Virus Replication by Resveratrol. J. Infect. Dis. 2005, 191, 1719–1729. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, A.; Mahmoud, S.H.; Elshaier, Y.A.M.M.; Shama, N.M.A.; Nasr, N.F.; Ali, M.A.; El-Shazly, A.M.; Mostafa, I.; Mostafa, A. Antiviral Activities of Plant-Derived Indole and β-Carboline Alkaloids against Human and Avian Influenza Viruses. Sci. Rep. 2023, 13, 1612. [Google Scholar] [CrossRef] [PubMed]
- Bodmer, B.S.; Hoenen, T.; Wendt, L. Molecular Insights into the Ebola Virus Life Cycle. Nat. Microbiol. 2024, 9, 1417–1426. [Google Scholar] [CrossRef] [PubMed]
- Stewart, C.M.; Phan, A.; Bo, Y.; LeBlond, N.D.; Smith, T.K.T.; Laroche, G.; Giguère, P.M.; Fullerton, M.D.; Pelchat, M.; Kobasa, D.; et al. Ebola Virus Triggers Receptor Tyrosine Kinase-Dependent Signaling to Promote the Delivery of Viral Particles to Entry-Conducive Intracellular Compartments. PLoS Pathog. 2021, 17, e1009275. [Google Scholar] [CrossRef] [PubMed]
- Tsang, N.Y.; Li, W.-F.; Varhegyi, E.; Rong, L.; Zhang, H.-J. Ebola Entry Inhibitors Discovered from Maesa Perlarius. Int. J. Mol. Sci. 2022, 23, 2620. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Kroeker, A.; He, S.; Kozak, R.; Audet, J.; Mbikay, M.; Chrétien, M. Prophylactic Efficacy of Quercetin 3-β-O-d-Glucoside against Ebola Virus Infection. Antimicrob. Agents Chemother. 2016, 60, 5182–5188. [Google Scholar] [CrossRef] [PubMed]
- Rampersad, S.; Tennant, P. Chapter 3—Replication and Expression Strategies of Viruses. In Viruses: Molecular Biology, Host Interactions and Applications to Biotechnology; Academic Press: Cambridge, MA, USA, 2018; pp. 55–82. [Google Scholar] [CrossRef]
- Yang, S.J.; Lim, Y. Resveratrol Ameliorates Hepatic Metaflammation and Inhibits NLRP3 Inflammasome Activation. Metabolism. 2014, 63, 693–701. [Google Scholar] [CrossRef] [PubMed]
- Boopathi, S.; Poma, A.B.; Kolandaivel, P. Novel 2019 Coronavirus Structure, Mechanism of Action, Antiviral Drug Promises and Rule out against Its Treatment. J. Biomol. Struct. Dyn. 2021, 39, 3409–3418. [Google Scholar] [CrossRef] [PubMed]
- Jena, N.R. Drug Targets, Mechanisms of Drug Action, and Therapeutics against SARS-CoV-2. Chem. Phys. Impact 2021, 2, 100011. [Google Scholar] [CrossRef]
- Astuti, I.; Ysrafil. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An Overview of Viral Structure and Host Response. Diabetes Metab. Syndr. 2020, 14, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.-W.; Kok, K.-H.; Zhu, Z.; Chu, H.; To, K.K.-W.; Yuan, S.; Yuen, K.-Y. Genomic Characterization of the 2019 Novel Human-Pathogenic Coronavirus Isolated from a Patient with Atypical Pneumonia after Visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef] [PubMed]
- Su, H.; Yao, S.; Zhao, W.; Li, M.; Liu, J.; Shang, W.; Xie, H.; Ke, C.; Gao, M.; Yu, K.; et al. Discovery of Baicalin and Baicalein as Novel, Natural Product Inhibitors of SARS-CoV-2 3CL Protease in Vitro. bioRxiv 2020. [Google Scholar] [CrossRef]
- Su, H.; Yao, S.; Zhao, W.; Li, M.; Liu, J.; Shang, W.; Xie, H.; Ke, C.; Hu, H.; Gao, M.; et al. Anti-SARS-CoV-2 Activities in Vitro of Shuanghuanglian Preparations and Bioactive Ingredients. Acta Pharmacol. Sin. 2020, 41, 1167–1177. [Google Scholar] [CrossRef] [PubMed]
- Zandi, K.; Musall, K.; Oo, A.; Cao, D.; Liang, B.; Hassandarvish, P.; Lan, S.; Slack, R.L.; Kirby, K.A.; Bassit, L.; et al. Baicalein and Baicalin Inhibit SARS-CoV-2 RNA-Dependent-RNA Polymerase. Microorganisms 2021, 9, 893. [Google Scholar] [CrossRef] [PubMed]
- Corona, A.; Wycisk, K.; Talarico, C.; Manelfi, C.; Milia, J.; Cannalire, R.; Esposito, F.; Gribbon, P.; Zaliani, A.; Iaconis, D.; et al. Natural Compounds Inhibit SARS-CoV-2 Nsp13 Unwinding and ATPase Enzyme Activities. ACS Pharmacol. Transl. Sci. 2022, 5, 226–239. [Google Scholar] [CrossRef] [PubMed]
- Dabeek, W.M.; Marra, M.V. Dietary Quercetin and Kaempferol: Bioavailability and Potential Cardiovascular-Related Bioactivity in Humans. Nutrients 2019, 11, 2288. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Yuk, H.J.; Ryu, H.W.; Lim, S.H.; Kim, K.S.; Park, K.H.; Ryu, Y.B.; Lee, W.S. Evaluation of Polyphenols from Broussonetia Papyrifera as Coronavirus Protease Inhibitors. J. Enzyme Inhib. Med. Chem. 2017, 32, 504–512. [Google Scholar] [CrossRef] [PubMed]
- Ryu, Y.B.; Jeong, H.J.; Kim, J.H.; Kim, Y.M.; Park, J.-Y.; Kim, D.; Nguyen, T.T.H.; Park, S.-J.; Chang, J.S.; Park, K.H.; et al. Biflavonoids from Torreya Nucifera Displaying SARS-CoV 3CL(pro) Inhibition. Bioorg. Med. Chem. 2010, 18, 7940–7947. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Chen, Z.; Tao, Y.; Zhang, B.; Wu, X.; Yang, L.; Wang, Q.; Wang, Z. An Integrated Method for Optimized Identification of Effective Natural Inhibitors against SARS-CoV-2 3CLpro. Sci. Rep. 2021, 11, 22796. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Tang, N.; Lai, X.; Zhang, J.; Wen, W.; Li, X.; Li, A.; Wu, Y.; Liu, Z. Insights Into Amentoflavone: A Natural Multifunctional Biflavonoid. Front. Pharmacol. 2021, 12, 768708. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Tan, N.; Zeng, G.; Zhang, Y.; Jia, R. Amentoflavone and Its Derivatives as Novel Natural Inhibitors of Human Cathepsin B. Bioorg. Med. Chem. 2005, 13, 5819–5825. [Google Scholar] [CrossRef] [PubMed]
- Youn, K.W.; Lee, S.; Kim, J.H.; Park, Y.-I.; So, J.; Kim, C.; Cho, C.W.; Park, J. Amentoflavone from Selaginella Tamariscina Inhibits SARS-CoV-2 RNA-Dependent RNA Polymerase. Heliyon 2024, 10, e36568. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.-S.; Lee, J.; Lee, J.M.; Kim, Y.; Chin, Y.-W.; Jee, J.-G.; Keum, Y.-S.; Jeong, Y.-J. Identification of Myricetin and Scutellarein as Novel Chemical Inhibitors of the SARS Coronavirus Helicase, nsP13. Bioorg. Med. Chem. Lett. 2012, 22, 4049–4054. [Google Scholar] [CrossRef] [PubMed]
- Meyer, B.R.; White, H.M.; McCormack, J.D.; Niemeyer, E.D. Catechin Composition, Phenolic Content, and Antioxidant Properties of Commercially-Available Bagged, Gunpowder, and Matcha Green Teas. Plant Foods Hum. Nutr. Dordr. Neth. 2023, 78, 662–669. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-N.; Lin, C.P.C.; Huang, K.-K.; Chen, W.-C.; Hsieh, H.-P.; Liang, P.-H.; Hsu, J.T.-A. Inhibition of SARS-CoV 3C-like Protease Activity by Theaflavin-3,3′-Digallate (TF3). Evid.-Based Complement. Altern. Med. ECAM 2005, 2, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.T.H.; Woo, H.-J.; Kang, H.-K.; Nguyen, V.D.; Kim, Y.-M.; Kim, D.-W.; Ahn, S.-A.; Xia, Y.; Kim, D. Flavonoid-Mediated Inhibition of SARS Coronavirus 3C-like Protease Expressed in Pichia Pastoris. Biotechnol. Lett. 2012, 34, 831–838. [Google Scholar] [CrossRef] [PubMed]
- Pillai, U.J.; Cherian, L.; Taunk, K.; Iype, E.; Dutta, M. Identification of Antiviral Phytochemicals from Cranberry as Potential Inhibitors of SARS-CoV-2 Main Protease (Mpro). Int. J. Biol. Macromol. 2024, 261, 129655. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Peng, Y.; Yao, H.; Wang, Y.; Li, J.; Yang, Y.; Lin, Z. Punicalagin as an Allosteric NSP13 Helicase Inhibitor Potently Suppresses SARS-CoV-2 Replication in Vitro. Antiviral Res. 2022, 206, 105389. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.N.; Shipley, P.R. Determination of Anthocyanins in Cranberry Fruit and Cranberry Fruit Products by High-Performance Liquid Chromatography with Ultraviolet Detection: Single-Laboratory Validation. J. Aoac Int. 2011, 94, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Meng, X.; Tan, C.; Tong, Y.; Wan, M.; Wang, M.; Zhao, Y.; Deng, H.; Kong, Y.; Ma, Y. Composition and Antioxidant Activity of Anthocyanins from Aronia Melanocarpa Extracted Using an Ultrasonic-Microwave-Assisted Natural Deep Eutectic Solvent Extraction Method. Ultrason. Sonochem. 2022, 89, 106102. [Google Scholar] [CrossRef] [PubMed]
- Kiselova-Kaneva, Y.; Nashar, M.; Roussev, B.; Salim, A.; Hristova, M.; Olczyk, P.; Komosinska-Vassev, K.; Dincheva, I.; Badjakov, I.; Galunska, B.; et al. Sambucus Ebulus (Elderberry) Fruits Modulate Inflammation and Complement System Activity in Humans. Int. J. Mol. Sci. 2023, 24, 8714. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.-H.; Min, J.S.; Jeon, S.; Lee, J.; Kim, S.; Park, T.; Park, D.; Jang, M.S.; Park, C.M.; Song, J.H.; et al. Lycorine, a Non-Nucleoside RNA Dependent RNA Polymerase Inhibitor, as Potential Treatment for Emerging Coronavirus Infections. Phytomedicine Int. J. Phytother. Phytopharm. 2021, 86, 153440. [Google Scholar] [CrossRef] [PubMed]
- Hillen, H.S.; Kokic, G.; Farnung, L.; Dienemann, C.; Tegunov, D.; Cramer, P. Structure of Replicating SARS-CoV-2 Polymerase. Nature 2020, 584, 154–156. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Yan, L.; Huang, Y.; Liu, F.; Zhao, Y.; Cao, L.; Wang, T.; Sun, Q.; Ming, Z.; Zhang, L.; et al. Structure of the RNA-Dependent RNA Polymerase from COVID-19 Virus. Science 2020, 368, 779–782. [Google Scholar] [CrossRef] [PubMed]
- Kokic, G.; Hillen, H.S.; Tegunov, D.; Dienemann, C.; Seitz, F.; Schmitzova, J.; Farnung, L.; Siewert, A.; Höbartner, C.; Cramer, P. Mechanism of SARS-CoV-2 Polymerase Stalling by Remdesivir. Nat. Commun. 2021, 12, 279. [Google Scholar] [CrossRef] [PubMed]
- Hu, W.-S.; Hughes, S.H. HIV-1 Reverse Transcription. Cold Spring Harb. Perspect. Med. 2012, 2, a006882. [Google Scholar] [CrossRef] [PubMed]
- McDougall, B.; King, P.J.; Wu, B.W.; Hostomsky, Z.; Reinecke, M.G.; Robinson, W.E. Dicaffeoylquinic and Dicaffeoyltartaric Acids Are Selective Inhibitors of Human Immunodeficiency Virus Type 1 Integrase. Antimicrob. Agents Chemother. 1998, 42, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Notka, F.; Meier, G.R.; Wagner, R. Inhibition of Wild-Type Human Immunodeficiency Virus and Reverse Transcriptase Inhibitor-Resistant Variants by Phyllanthus Amarus. Antiviral Res. 2003, 58, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Sanna, C.; Marengo, A.; Acquadro, S.; Caredda, A.; Lai, R.; Corona, A.; Tramontano, E.; Rubiolo, P.; Esposito, F. In Vitro Anti-HIV-1 Reverse Transcriptase and Integrase Properties of Punica granatum L. Leaves, Bark, and Peel Extracts and Their Main Compounds. Plants 2021, 10, 2124. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Goila-Gaur, R.; Salzwedel, K.; Kilgore, N.R.; Reddick, M.; Matallana, C.; Castillo, A.; Zoumplis, D.; Martin, D.E.; Orenstein, J.M.; et al. PA-457: A Potent HIV Inhibitor That Disrupts Core Condensation by Targeting a Late Step in Gag Processing. Proc. Natl. Acad. Sci. USA 2003, 100, 13555–13560. [Google Scholar] [CrossRef] [PubMed]
- Laureti, M.; Paradkar, P.N.; Fazakerley, J.K.; Rodriguez-Andres, J. Superinfection Exclusion in Mosquitoes and Its Potential as an Arbovirus Control Strategy. Viruses 2020, 12, 1259. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Tan, X.-F.; Thurmond, S.; Zhang, Z.-M.; Lin, A.; Hai, R.; Song, J. The Structure of Zika Virus NS5 Reveals a Conserved Domain Conformation. Nat. Commun. 2017, 8, 14763. [Google Scholar] [CrossRef]
- Zhao, B.; Yi, G.; Du, F.; Chuang, Y.-C.; Vaughan, R.C.; Sankaran, B.; Kao, C.C.; Li, P. Structure and Function of the Zika Virus Full-Length NS5 Protein. Nat. Commun. 2017, 8, 14762. [Google Scholar] [CrossRef] [PubMed]
- Haddad, J.G.; Gadea, G.; Desprès, P.; El Kalamouni, C. Chapter 38—Medicinal Plants as Promising Source of Natural Antiviral Substances against Zika Virus. In Zika Virus Impact, Diagnosis, Control, and Models; Martin, C.R., Hollins Martin, C.J., Preedy, V.R., Rajendram, R., Eds.; Academic Press: Cambridge, MA, USA, 2021; pp. 397–407. ISBN 978-0-12-820267-8. [Google Scholar]
- Kang, C.; Keller, T.H.; Luo, D. Zika Virus Protease: An Antiviral Drug Target. Trends Microbiol. 2017, 25, 797–808. [Google Scholar] [CrossRef] [PubMed]
- Lim, H.-J.; Nguyen, T.T.H.; Kim, N.M.; Park, J.-S.; Jang, T.-S.; Kim, D. Inhibitory Effect of Flavonoids against NS2B-NS3 Protease of ZIKA Virus and Their Structure Activity Relationship. Biotechnol. Lett. 2017, 39, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D.; Sharma, N.; Aarthy, M.; Singh, S.K.; Giri, R. Mechanistic Insights into Zika Virus NS3 Helicase Inhibition by Epigallocatechin-3-Gallate. ACS Omega 2020, 5, 11217–11226. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Lao, Z.; Xu, J.; Li, Z.; Long, H.; Li, D.; Lin, L.; Liu, X.; Yu, L.; Liu, W.; et al. Antiviral Activity of Lycorine against Zika Virus in Vivo and in Vitro. Virology 2020, 546, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.; Kuhn, R.J. Structural Proteomics of Dengue Virus. Curr. Opin. Microbiol. 2008, 11, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Henderson, B.R.; Saeedi, B.J.; Campagnola, G.; Geiss, B.J. Analysis of RNA Binding by the Dengue Virus NS5 RNA Capping Enzyme. PLoS ONE 2011, 6, e25795. [Google Scholar] [CrossRef] [PubMed]
- Ng, L.-C.; Chem, Y.-K.; Koo, C.; Mudin, R.N.B.; Amin, F.M.; Lee, K.-S.; Kheong, C.C. 2013 Dengue Outbreaks in Singapore and Malaysia Caused by Different Viral Strains. Am. J. Trop. Med. Hyg. 2015, 92, 1150–1155. [Google Scholar] [CrossRef] [PubMed]
- Coulerie, P.; Nour, M.; Maciuk, A.; Eydoux, C.; Guillemot, J.-C.; Lebouvier, N.; Hnawia, E.; Leblanc, K.; Lewin, G.; Canard, B.; et al. Structure-Activity Relationship Study of Biflavonoids on the Dengue Virus Polymerase DENV-NS5 RdRp. Planta Med. 2013, 79, 1313–1318. [Google Scholar] [CrossRef] [PubMed]
- Acchioni, C.; Acchioni, M.; Mancini, F.; Amendola, A.; Marsili, G.; Tirelli, V.; Gwee, C.P.; Chan, K.W.-K.; Sandini, S.; Bisbocci, M.; et al. A Cellular Screening Platform, Stably Expressing DENV2 NS5, Defines a Novel Anti-DENV Mechanism of Action of Apigenin Based on STAT2 Activation. Virology 2023, 583, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Dang, M.; Lim, L.; Roy, A.; Song, J. Myricetin Allosterically Inhibits the Dengue NS2B-NS3 Protease by Disrupting the Active and Locking the Inactive Conformations. ACS Omega 2022, 7, 2798–2808. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, A.; Pilankatta, R.; Teramoto, T.; Sajith, A.M.; Nwulia, E.; Kulkarni, A.; Padmanabhan, R. Inhibition of Dengue Virus by Curcuminoids. Antiviral Res. 2019, 162, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Frolov, I.; Frolova, E.I. Molecular Virology of Chikungunya Virus. Curr. Top. Microbiol. Immunol. 2022, 435, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Bartholomeeusen, K.; Daniel, M.; LaBeaud, D.A.; Gasque, P.; Peeling, R.W.; Stephenson, K.E.; Ng, L.F.P.; Ariën, K.K. Chikungunya Fever. Nat. Rev. Dis. Primer 2023, 9, 17. [Google Scholar] [CrossRef] [PubMed]
- Pohjala, L.; Utt, A.; Varjak, M.; Lulla, A.; Merits, A.; Ahola, T.; Tammela, P. Inhibitors of Alphavirus Entry and Replication Identified with a Stable Chikungunya Replicon Cell Line and Virus-Based Assays. PLoS ONE 2011, 6, e28923. [Google Scholar] [CrossRef] [PubMed]
- Kaur, P.; Thiruchelvan, M.; Lee, R.C.H.; Chen, H.; Chen, K.C.; Ng, M.L.; Chu, J.J.H. Inhibition of Chikungunya Virus Replication by Harringtonine, a Novel Antiviral That Suppresses Viral Protein Expression. Antimicrob. Agents Chemother. 2013, 57, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.S.; Thaa, B.; Amrun, S.N.; Simarmata, D.; Rausalu, K.; Nyman, T.A.; Merits, A.; McInerney, G.M.; Ng, L.F.P.; Ahola, T. The Antiviral Alkaloid Berberine Reduces Chikungunya Virus-Induced Mitogen-Activated Protein Kinase Signaling. J. Virol. 2016, 90, 9743–9757. [Google Scholar] [CrossRef] [PubMed]
- Bourjot, M.; Delang, L.; Nguyen, V.H.; Neyts, J.; Guéritte, F.; Leyssen, P.; Litaudon, M. Prostratin and 12-O-Tetradecanoylphorbol 13-Acetate Are Potent and Selective Inhibitors of Chikungunya Virus Replication. J. Nat. Prod. 2012, 75, 2183–2187. [Google Scholar] [CrossRef] [PubMed]
- Lani, R.; Hassandarvish, P.; Chiam, C.W.; Moghaddam, E.; Chu, J.J.H.; Rausalu, K.; Merits, A.; Higgs, S.; Vanlandingham, D.; Abu Bakar, S.; et al. Antiviral Activity of Silymarin against Chikungunya Virus. Sci. Rep. 2015, 5, 11421. [Google Scholar] [CrossRef] [PubMed]
- Stubbs, T.M.; te Velthuis, A.J. The RNA-Dependent RNA Polymerase of the Influenza A Virus. Future Virol. 2014, 9, 863–876. [Google Scholar] [CrossRef] [PubMed]
- Dubois, J.; Terrier, O.; Rosa-Calatrava, M. Influenza Viruses and mRNA Splicing: Doing More with Less. mBio 2014, 5, e00070-14. [Google Scholar] [CrossRef] [PubMed]
- Robertson, J.S.; Naeve, C.W.; Webster, R.G.; Bootman, J.S.; Newman, R.; Schild, G.C. Alterations in the Hemagglutinin Associated with Adaptation of Influenza B Virus to Growth in Eggs. Virology 1985, 143, 166–174. [Google Scholar] [CrossRef] [PubMed]
- Grienke, U.; Schmidtke, M.; von Grafenstein, S.; Kirchmair, J.; Liedl, K.R.; Rollinger, J.M. Influenza Neuraminidase: A Druggable Target for Natural Products. Nat. Prod. Rep. 2012, 29, 11–36. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Gong, J.; Li, M.; Fang, H.; Xu, W. The Medicinal Potential of Influenza Virus Surface Proteins: Hemagglutinin and Neuraminidase. Curr. Med. Chem. 2011, 18, 1050–1066. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.J.; Ryu, Y.B.; Park, S.-J.; Kim, J.H.; Kwon, H.-J.; Kim, J.H.; Park, K.H.; Rho, M.-C.; Lee, W.S. Neuraminidase Inhibitory Activities of Flavonols Isolated from Rhodiola Rosea Roots and Their in Vitro Anti-Influenza Viral Activities. Bioorg. Med. Chem. 2009, 17, 6816–6823. [Google Scholar] [CrossRef] [PubMed]
- Bang, S.; Quy Ha, T.K.; Lee, C.; Li, W.; Oh, W.-K.; Shim, S.H. Antiviral Activities of Compounds from Aerial Parts of Salvia plebeia R. Br. J. Ethnopharmacol. 2016, 192, 398–405. [Google Scholar] [CrossRef] [PubMed]
- Huh, J.; Ha, T.K.Q.; Kang, K.B.; Kim, K.H.; Oh, W.K.; Kim, J.; Sung, S.H. C-Methylated Flavonoid Glycosides from Pentarhizidium Orientale Rhizomes and Their Inhibitory Effects on the H1N1 Influenza Virus. J. Nat. Prod. 2017, 80, 2818–2824. [Google Scholar] [CrossRef] [PubMed]
- Ha, T.K.Q.; Dao, T.T.; Nguyen, N.H.; Kim, J.; Kim, E.; Cho, T.O.; Oh, W.K. Antiviral Phenolics from the Leaves of Cleistocalyx Operculatus. Fitoterapia 2016, 110, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.W.; Ha, T.K.Q.; Cho, H.M.; An, J.-P.; Kim, S.K.; Kim, C.-S.; Kim, E.; Oh, W.K. Antiviral Activity of Furanocoumarins Isolated from Angelica Dahurica against Influenza a Viruses H1N1 and H9N2. J. Ethnopharmacol. 2020, 259, 112945. [Google Scholar] [CrossRef] [PubMed]
- Kuzuhara, T.; Iwai, Y.; Takahashi, H.; Hatakeyama, D.; Echigo, N. Green Tea Catechins Inhibit the Endonuclease Activity of Influenza A Virus RNA Polymerase. PLoS Curr. 2009, 1, RRN1052. [Google Scholar] [CrossRef] [PubMed]
- Botwina, P.; Owczarek, K.; Rajfur, Z.; Ochman, M.; Urlik, M.; Nowakowska, M.; Szczubiałka, K.; Pyrc, K. Berberine Hampers Influenza A Replication through Inhibition of MAPK/ERK Pathway. Viruses 2020, 12, 344. [Google Scholar] [CrossRef] [PubMed]
- Mühlberger, E.; Weik, M.; Volchkov, V.E.; Klenk, H.D.; Becker, S. Comparison of the Transcription and Replication Strategies of Marburg Virus and Ebola Virus by Using Artificial Replication Systems. J. Virol. 1999, 73, 2333–2342. [Google Scholar] [CrossRef] [PubMed]
- Leung, D.W.; Borek, D.; Luthra, P.; Binning, J.M.; Anantpadma, M.; Liu, G.; Harvey, I.B.; Su, Z.; Endlich-Frazier, A.; Pan, J.; et al. An Intrinsically Disordered Peptide from Ebola Virus VP35 Controls Viral RNA Synthesis by Modulating Nucleoprotein-RNA Interactions. Cell Rep. 2015, 11, 376–389. [Google Scholar] [CrossRef] [PubMed]
- Takamatsu, Y.; Kolesnikova, L.; Becker, S. Ebola Virus Proteins NP, VP35, and VP24 Are Essential and Sufficient to Mediate Nucleocapsid Transport. Proc. Natl. Acad. Sci. USA 2018, 115, 1075–1080. [Google Scholar] [CrossRef] [PubMed]
- Shu, T.; Gan, T.; Bai, P.; Wang, X.; Qian, Q.; Zhou, H.; Cheng, Q.; Qiu, Y.; Yin, L.; Zhong, J.; et al. Ebola Virus VP35 Has Novel NTPase and Helicase-like Activities. Nucleic Acids Res. 2019, 47, 5837–5851. [Google Scholar] [CrossRef] [PubMed]
- Corona, A.; Fanunza, E.; Salata, C.; Morwitzer, M.J.; Distinto, S.; Zinzula, L.; Sanna, C.; Frau, A.; Daino, G.L.; Quartu, M.; et al. Cynarin Blocks Ebola Virus Replication by Counteracting VP35 Inhibition of Interferon-Beta Production. Antiviral Res. 2022, 198, 105251. [Google Scholar] [CrossRef] [PubMed]
- Takova, K.; Koynarski, T.; Minkov, G.; Toneva, V.; Mardanova, E.; Ravin, N.; Lukov, G.L.; Zahmanova, G. Development and Optimization of an Enzyme Immunoassay to Detect Serum Antibodies against the Hepatitis E Virus in Pigs, Using Plant-Derived ORF2 Recombinant Protein. Vaccines 2021, 9, 991. [Google Scholar] [CrossRef] [PubMed]
- Rybicki, E.P.; Chikwamba, R.; Koch, M.; Rhodes, J.I.; Groenewald, J.-H. Plant-Made Therapeutics: An Emerging Platform in South Africa. Biotechnol. Adv. 2012, 30, 449–459. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Aljabali, A.A.A.; Takova, K.; Minkov, G.; Tambuwala, M.M.; Minkov, I.; Lomonossoff, G.P. Green Biologics: Harnessing the Power of Plants to Produce Pharmaceuticals. Int. J. Mol. Sci. 2023, 24, 17575. [Google Scholar] [CrossRef] [PubMed]
- Buyel, J.F.; Stöger, E.; Bortesi, L. Targeted Genome Editing of Plants and Plant Cells for Biomanufacturing. Transgenic Res. 2021, 30, 401–426. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Timko, M.P. Improving Protein Quantity and Quality-The Next Level of Plant Molecular Farming. Int. J. Mol. Sci. 2022, 23, 1326. [Google Scholar] [CrossRef] [PubMed]
- Strasser, R. Plant Glycoengineering for Designing Next-Generation Vaccines and Therapeutic Proteins. Biotechnol. Adv. 2023, 67, 108197. [Google Scholar] [CrossRef] [PubMed]
- Thuenemann, E.C.; Lenzi, P.; Love, A.J.; Taliansky, M.; Bécares, M.; Zuñiga, S.; Enjuanes, L.; Zahmanova, G.G.; Minkov, I.N.; Matić, S.; et al. The Use of Transient Expression Systems for the Rapid Production of Virus-like Particles in Plants. Curr. Pharm. Des. 2013, 19, 5564–5573. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Aljabali, A.A.; Takova, K.; Toneva, V.; Tambuwala, M.M.; Andonov, A.P.; Lukov, G.L.; Minkov, I. The Plant Viruses and Molecular Farming: How Beneficial They Might Be for Human and Animal Health? Int. J. Mol. Sci. 2023, 24, 1533. [Google Scholar] [CrossRef] [PubMed]
- Mardanova, E.S.; Blokhina, E.A.; Tsybalova, L.M.; Peyret, H.; Lomonossoff, G.P.; Ravin, N.V. Efficient Transient Expression of Recombinant Proteins in Plants by the Novel pEff Vector Based on the Genome of Potato virus X. Front. Plant Sci. 2017, 8, 247. [Google Scholar] [CrossRef] [PubMed]
- Peyret, H.; Lomonossoff, G.P. When Plant Virology Met Agrobacterium: The Rise of the Deconstructed Clones. Plant Biotechnol. J. 2015, 13, 1121–1135. [Google Scholar] [CrossRef] [PubMed]
- Sainsbury, F.; Lomonossoff, G.P. Extremely High-Level and Rapid Transient Protein Production in Plants without the Use of Viral Replication. Plant Physiol. 2008, 148, 1212–1218. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Dukiandjiev, S.; Minkov, I.; Andonov, A. Molecular farming—The production of recombinant subunit hepatitis b vaccine in tobacco plants. In Proceedings of the Balkan Scientific Conference of Biology, Plovdiv, Bulgaria, 19–21 May 2005; pp. 329–338. [Google Scholar]
- Rybicki, E.P. Plant-Based Vaccines against Viruses. Virol. J. 2014, 11, 205. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Takova, K.; Valkova, R.; Toneva, V.; Minkov, I.; Andonov, A.; Lukov, G.L. Plant-Derived Recombinant Vaccines against Zoonotic Viruses. Life 2022, 12, 156. [Google Scholar] [CrossRef] [PubMed]
- Team, B.P. Baiya Phytopharm Announces Preliminary Results of the Phase 1 Clinical Trial of Its COVID-19 Vaccine Baiya SARS-CoV-2 Vax 2. Baiya Phytopharm. 2022. Available online: https://baiyaphytopharm.com/baiyaphytopharm-announces-preliminary-results-of-the-phase-1-clinical-trial-of-its-covid-19-vaccine-baiya-sars-cov-2-vax-2/ (accessed on 15 June 2025).
- KBio Inc A Phase I/II, First-in-Human, Observer-Blinded, Randomized, Placebo-Controlled, Parallel Group Study to Evaluate the Safety and Immunogenicity of TAP-COVID-19 SARS-CoV-2 Vaccine with CpG Adjuvant in Healthy Adults Aged 18–49 and 50–85; clinicaltrials.gov, 2023. Available online: https://outbreak.info/resources/NCT04473690 (accessed on 15 June 2025).
- Medicago, GSK Announce Positive Results for Plant-Based COVID-19 Vaccine. Available online: https://www.pharmacytimes.com/view/medicago-gsk-announce-positive-results-for-plant-based-covid-19-vaccine (accessed on 15 June 2025).
- Ramessar, K.; Sabalza, M.; Capell, T.; Christou, P. Maize Plants: An Ideal Production Platform for Effective and Safe Molecular Pharming. Plant Sci. 2008, 174, 409–419. [Google Scholar] [CrossRef]
- Concha, C.; Cañas, R.; Macuer, J.; Torres, M.J.; Herrada, A.A.; Jamett, F.; Ibáñez, C. Disease Prevention: An Opportunity to Expand Edible Plant-Based Vaccines? Vaccines 2017, 5, 14. [Google Scholar] [CrossRef] [PubMed]
- Jain, A.; Saini, V.; Kohli, D.V. Edible Transgenic Plant Vaccines for Different Diseases. Curr. Pharm. Biotechnol. 2013, 14, 594–614. [Google Scholar] [CrossRef] [PubMed]
- Meyers, A.; Chakauya, E.; Shephard, E.; Tanzer, F.L.; Maclean, J.; Lynch, A.; Williamson, A.-L.; Rybicki, E.P. Expression of HIV-1 Antigens in Plants as Potential Subunit Vaccines. BMC Biotechnol. 2008, 8, 53. [Google Scholar] [CrossRef] [PubMed]
- Chung, Y.H.; Church, D.; Koellhoffer, E.C.; Osota, E.; Shukla, S.; Rybicki, E.P.; Pokorski, J.K.; Steinmetz, N.F. Integrating Plant Molecular Farming and Materials Research for Next-Generation Vaccines. Nat. Rev. Mater. 2022, 7, 372–388. [Google Scholar] [CrossRef] [PubMed]
- Peyret, H.; Steele, J.F.C.; Jung, J.-W.; Thuenemann, E.C.; Meshcheriakova, Y.; Lomonossoff, G.P. Producing Vaccines against Enveloped Viruses in Plants: Making the Impossible, Difficult. Vaccines 2021, 9, 780. [Google Scholar] [CrossRef] [PubMed]
- Benvenuto, E.; Broer, I.; D’Aoust, M.-A.; Hitzeroth, I.; Hundleby, P.; Menassa, R.; Oksman-Caldentey, K.-M.; Peyret, H.; Salgueiro, S.; Saxena, P.; et al. Plant Molecular Farming in the Wake of the Closure of Medicago Inc. Nat. Biotechnol. 2023, 41, 893–894. [Google Scholar] [CrossRef] [PubMed]
- Cirilo, T.M.; Grossi de Oliveira, A.L.; Costa Pinto, J.; Rihs, J.B.d.R.; Ruas, A.C.L.; Siqueira, W.F.; de Castro, J.C.; Sernizon Guimarães, N.; de Medeiros Brito, R.M.; Bueno, L.L.; et al. Assessing the Efficacy of Modified Plant Vaccine Antigens in Animal Immunization: A Systematic Review. Food Biosci. 2025, 66, 106178. [Google Scholar] [CrossRef]
- Mogojwe, H. Current and Planned Vaccine Manufacturing in Africa. Available online: https://www.clintonhealthaccess.org/report/current-and-planned-vaccine-manufacturing/ (accessed on 21 September 2023).
- BioApplications Inc. Available online: https://www.bioapplications.global/ (accessed on 15 June 2025).
- NCT05040789. Available online: https://covid-19.cochrane.org/studies/crs-18688748 (accessed on 10 September 2021).
- Yang, M.; Sun, H.; Lai, H.; Hurtado, J.; Chen, Q. Plant-produced Zika Virus Envelope Protein Elicits Neutralizing Immune Responses That Correlate with Protective Immunity against Zika Virus in Mice. Plant Biotechnol. J. 2017, 16, 572–580. [Google Scholar] [CrossRef] [PubMed]
- Phoolcharoen, W.; Bhoo, S.H.; Lai, H.; Ma, J.; Arntzen, C.J.; Chen, Q.; Mason, H.S. Expression of an Immunogenic Ebola Immune Complex in Nicotiana Benthamiana. Plant Biotechnol. J. 2011, 9, 807–816. [Google Scholar] [CrossRef] [PubMed]
- Ríos-Huerta, R.; Monreal-Escalante, E.; Govea-Alonso, D.O.; Angulo, C.; Rosales-Mendoza, S. Expression of an Immunogenic LTB-Based Chimeric Protein Targeting Zaire Ebolavirus Epitopes from GP1 in Plant Cells. Plant Cell Rep. 2017, 36, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Pillet, S.; Aubin, É.; Trépanier, S.; Poulin, J.-F.; Yassine-Diab, B.; ter Meulen, J.; Ward, B.J.; Landry, N. Humoral and Cell-Mediated Immune Responses to H5N1 Plant-Made Virus-like Particle Vaccine Are Differentially Impacted by Alum and GLA-SE Adjuvants in a Phase 2 Clinical Trial. Npj Vaccines 2018, 3, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Pillet, S.; Couillard, J.; Trépanier, S.; Poulin, J.-F.; Yassine-Diab, B.; Guy, B.; Ward, B.J.; Landry, N. Immunogenicity and Safety of a Quadrivalent Plant-Derived Virus like Particle Influenza Vaccine Candidate-Two Randomized Phase II Clinical Trials in 18 to 49 and ≥50 Years Old Adults. PLoS ONE 2019, 14, e0216533. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Lai, H. Plant-Derived Virus-like Particles as Vaccines. Hum. Vaccines Immunother. 2013, 9, 26–49. [Google Scholar] [CrossRef] [PubMed]
- Kheirvari, M.; Liu, H.; Tumban, E. Virus-like Particle Vaccines and Platforms for Vaccine Development. Viruses 2023, 15, 1109. [Google Scholar] [CrossRef] [PubMed]
- Mardanova, E.S.; Kotlyarov, R.Y.; Stuchinskaya, M.D.; Nikolaeva, L.I.; Zahmanova, G.; Ravin, N.V. High-Yield Production of Chimeric Hepatitis E Virus-Like Particles Bearing the M2e Influenza Epitope and Receptor Binding Domain of SARS-CoV-2 in Plants Using Viral Vectors. Int. J. Mol. Sci. 2022, 23, 15684. [Google Scholar] [CrossRef] [PubMed]
- Mardanova, E.S.; Vasyagin, E.A.; Kotova, K.G.; Zahmanova, G.G.; Ravin, N.V. Plant-Produced Chimeric Hepatitis E Virus-like Particles as Carriers for Antigen Presentation. Viruses 2024, 16, 1093. [Google Scholar] [CrossRef] [PubMed]
- Blokhina, E.A.; Mardanova, E.S.; Zykova, A.A.; Shuklina, M.A.; Stepanova, L.A.; Tsybalova, L.M.; Ravin, N.V. Chimeric Virus-like Particles of Physalis Mottle Virus as Carriers of M2e Peptides of Influenza a Virus. Viruses 2024, 16, 1802. [Google Scholar] [CrossRef] [PubMed]
- Dalsgaard, K.; Uttenthal, A.; Jones, T.D.; Xu, F.; Merryweather, A.; Hamilton, W.D.; Langeveld, J.P.; Boshuizen, R.S.; Kamstrup, S.; Lomonossoff, G.P.; et al. Plant-Derived Vaccine Protects Target Animals against a Viral Disease. Nat. Biotechnol. 1997, 15, 248–252. [Google Scholar] [CrossRef] [PubMed]
- Palmer, K.E.; Benko, A.; Doucette, S.A.; Cameron, T.I.; Foster, T.; Hanley, K.M.; McCormick, A.A.; McCulloch, M.; Pogue, G.P.; Smith, M.L.; et al. Protection of Rabbits against Cutaneous Papillomavirus Infection Using Recombinant Tobacco Mosaic Virus Containing L2 Capsid Epitopes. Vaccine 2006, 24, 5516–5525. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.M.; Chichester, J.A.; Mett, V.; Jaje, J.; Tottey, S.; Manceva, S.; Casta, L.J.; Gibbs, S.K.; Musiychuk, K.; Shamloul, M.; et al. A Plant-Produced Pfs25 VLP Malaria Vaccine Candidate Induces Persistent Transmission Blocking Antibodies against Plasmodium Falciparum in Immunized Mice. PLoS ONE 2013, 8, e79538. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Lai, H.; Sun, H.; Chen, Q. Virus-like Particles That Display Zika Virus Envelope Protein Domain III Induce Potent Neutralizing Immune Responses in Mice. Sci. Rep. 2017, 7, 7679. [Google Scholar] [CrossRef] [PubMed]
- Ravin, N.V.; Kotlyarov, R.Y.; Mardanova, E.S.; Kuprianov, V.V.; Migunov, A.I.; Stepanova, L.A.; Tsybalova, L.M.; Kiselev, O.I.; Skryabin, K.G. Plant-Produced Recombinant Influenza Vaccine Based on Virus-like HBc Particles Carrying an Extracellular Domain of M2 Protein. Biochem. Biokhimiia 2012, 77, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Mazalovska, M.; Takova, K.; Toneva, V.; Minkov, I.; Peyret, H.; Lomonossoff, G. Efficient Production of Chimeric Hepatitis B Virus-Like Particles Bearing an Epitope of Hepatitis E Virus Capsid by Transient Expression in Nicotiana Benthamiana. Life 2021, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.G.; Mazalovska, M.; Takova, K.H.; Toneva, V.T.; Minkov, I.N.; Mardanova, E.S.; Ravin, N.V.; Lomonossoff, G.P. Rapid High-Yield Transient Expression of Swine Hepatitis E ORF2 Capsid Proteins in Nicotiana Benthamiana Plants and Production of Chimeric Hepatitis E Virus-Like Particles Bearing the M2e Influenza Epitope. Plants 2019, 9, 29. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.; Mazalovska, M.; Toneva, V.; Minkov, I.; Lomonossoff, G. Production of Chimeric Virus-like Particles Bearing M2e Influenza Epitope in Nicotiana Benthamiana Plants. J. Biotechnol. 2015, 208, S109. [Google Scholar] [CrossRef]
- Ponndorf, D.; Meshcheriakova, Y.; Thuenemann, E.C.; Dobon Alonso, A.; Overman, R.; Holton, N.; Dowall, S.; Kennedy, E.; Stocks, M.; Lomonossoff, G.P.; et al. Plant-Made Dengue Virus-like Particles Produced by Co-Expression of Structural and Non-Structural Proteins Induce a Humoral Immune Response in Mice. Plant Biotechnol. J. 2021, 19, 745–756. [Google Scholar] [CrossRef] [PubMed]
- Rutkowska, D.A.; Mokoena, N.B.; Tsekoa, T.L.; Dibakwane, V.S.; O’Kennedy, M.M. Plant-Produced Chimeric Virus-like Particles—A New Generation Vaccine against African Horse Sickness. BMC Vet. Res. 2019, 15, 432. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A. Passive Antibody Administration (Immediate Immunity) as a Specific Defense against Biological Weapons. Emerg. Infect. Dis. 2002, 8, 833–841. [Google Scholar] [CrossRef] [PubMed]
- Hiatt, A.; Whaley, K.J.; Zeitlin, L. Plant-Derived Monoclonal Antibodies for Prevention and Treatment of Infectious Disease. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Hiatt, A.; Pauly, M. Monoclonal Antibodies from Plants: A New Speed Record. Proc. Natl. Acad. Sci. USA 2006, 103, 14645–14646. [Google Scholar] [CrossRef] [PubMed]
- Giritch, A.; Marillonnet, S.; Engler, C.; van Eldik, G.; Botterman, J.; Klimyuk, V.; Gleba, Y. Rapid High-Yield Expression of Full-Size IgG Antibodies in Plants Coinfected with Noncompeting Viral Vectors. Proc. Natl. Acad. Sci. USA 2006, 103, 14701–14706. [Google Scholar] [CrossRef] [PubMed]
- Diamos, A.G.; Hunter, J.G.L.; Pardhe, M.D.; Rosenthal, S.H.; Sun, H.; Foster, B.C.; DiPalma, M.P.; Chen, Q.; Mason, H.S. High Level Production of Monoclonal Antibodies Using an Optimized Plant Expression System. Front. Bioeng. Biotechnol. 2020, 7, 472. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Liu, L.; Voglmeir, J. mAbs N-Glycosylation: Implications for Biotechnology and Analytics. Carbohydr. Res. 2022, 514, 108541. [Google Scholar] [CrossRef] [PubMed]
- Ko, K.; Tekoah, Y.; Rudd, P.M.; Harvey, D.J.; Dwek, R.A.; Spitsin, S.; Hanlon, C.A.; Rupprecht, C.; Dietzschold, B.; Golovkin, M.; et al. Function and Glycosylation of Plant-Derived Antiviral Monoclonal Antibody. Proc. Natl. Acad. Sci. USA 2003, 100, 8013–8018. [Google Scholar] [CrossRef] [PubMed]
- Castilho, A.; Steinkellner, H. Glyco-Engineering in Plants to Produce Human-like N-Glycan Structures. Biotechnol. J. 2012, 7, 1088–1098. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Kim, D.-S.; Ko, K. Expression of a Large Single-Chain 13F6 Antibody with Binding Activity against Ebola Virus-Like Particles in a Plant System. Int. J. Mol. Sci. 2020, 21, 7007. [Google Scholar] [CrossRef] [PubMed]
- Strasser, R.; Castilho, A.; Stadlmann, J.; Kunert, R.; Quendler, H.; Gattinger, P.; Jez, J.; Rademacher, T.; Altmann, F.; Mach, L.; et al. Improved Virus Neutralization by Plant-Produced Anti-HIV Antibodies with a Homogeneous Β1,4-Galactosylated N-Glycan Profile*. J. Biol. Chem. 2009, 284, 20479–20485. [Google Scholar] [CrossRef] [PubMed]
- Göritzer, K.; Melnik, S.; Schwestka, J.; Arcalis, E.; Drapal, M.; Fraser, P.; Ma, J.K.-C.; Stoger, E. Enhancing Quality and Yield of Recombinant Secretory IgA Antibodies in Nicotiana Benthamiana by Endoplasmic Reticulum Engineering. Plant Biotechnol. J. 2025, 23, 1178–1189. [Google Scholar] [CrossRef] [PubMed]
- Niemer, M.; Mehofer, U.; Torres Acosta, J.A.; Verdianz, M.; Henkel, T.; Loos, A.; Strasser, R.; Maresch, D.; Rademacher, T.; Steinkellner, H.; et al. The Human Anti-HIV Antibodies 2F5, 2G12, and PG9 Differ in Their Susceptibility to Proteolytic Degradation: Down-Regulation of Endogenous Serine and Cysteine Proteinase Activities Could Improve Antibody Production in Plant-Based Expression Platforms. Biotechnol. J. 2014, 9, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.K.-C.; Drossard, J.; Lewis, D.; Altmann, F.; Boyle, J.; Christou, P.; Cole, T.; Dale, P.; van Dolleweerd, C.J.; Isitt, V.; et al. Regulatory Approval and a First-in-Human Phase I Clinical Trial of a Monoclonal Antibody Produced in Transgenic Tobacco Plants. Plant Biotechnol. J. 2015, 13, 1106–1120. [Google Scholar] [CrossRef] [PubMed]
- Lotter-Stark, H.C.T.; Rybicki, E.P.; Chikwamba, R.K. Plant Made Anti-HIV Microbicides—A Field of Opportunity. Biotechnol. Adv. 2012, 30, 1614–1626. [Google Scholar] [CrossRef] [PubMed]
- Loos, A.; Van Droogenbroeck, B.; Hillmer, S.; Grass, J.; Pabst, M.; Castilho, A.; Kunert, R.; Liang, M.; Arcalis, E.; Robinson, D.G.; et al. Expression of Antibody Fragments with a Controlled N-Glycosylation Pattern and Induction of Endoplasmic Reticulum-Derived Vesicles in Seeds of Arabidopsis. Plant Physiol. 2011, 155, 2036–2048. [Google Scholar] [CrossRef] [PubMed]
- Grandits, M.; Grünwald-Gruber, C.; Gastine, S.; Standing, J.F.; Reljic, R.; Teh, A.Y.-H.; Ma, J.K.-C. Improving the Efficacy of Plant-Made Anti-HIV Monoclonal Antibodies for Clinical Use. Front. Plant Sci. 2023, 14, 1126470. [Google Scholar] [CrossRef] [PubMed]
- Rademacher, T.; Sack, M.; Arcalis, E.; Stadlmann, J.; Balzer, S.; Altmann, F.; Quendler, H.; Stiegler, G.; Kunert, R.; Fischer, R.; et al. Recombinant Antibody 2G12 Produced in Maize Endosperm Efficiently Neutralizes HIV-1 and Contains Predominantly Single-GlcNAc N-Glycans. Plant Biotechnol. J. 2008, 6, 189–201. [Google Scholar] [CrossRef] [PubMed]
- Castilho, A.; Bohorova, N.; Grass, J.; Bohorov, O.; Zeitlin, L.; Whaley, K.; Altmann, F.; Steinkellner, H. Rapid High Yield Production of Different Glycoforms of Ebola Virus Monoclonal Antibody. PLoS ONE 2011, 6, e26040. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Davis, K.R. The Potential of Plants as a System for the Development and Production of Human Biologics. F1000Research 2016, 5, 912. [Google Scholar] [CrossRef] [PubMed]
- Lai, H.; Engle, M.; Fuchs, A.; Keller, T.; Johnson, S.; Gorlatov, S.; Diamond, M.S.; Chen, Q. Monoclonal Antibody Produced in Plants Efficiently Treats West Nile Virus Infection in Mice. Proc. Natl. Acad. Sci. USA 2010, 107, 2419–2424. [Google Scholar] [CrossRef] [PubMed]
- Jugler, C.; Joensuu, J.; Chen, Q. Hydrophobin-Protein A Fusion Protein Produced in Plants Efficiently Purified an Anti-West Nile Virus Monoclonal Antibody from Plant Extracts via Aqueous Two-Phase Separation. Int. J. Mol. Sci. 2020, 21, 2140. [Google Scholar] [CrossRef] [PubMed]
- Kapusta, J.; Modelska, A.; Figlerowicz, M.; Pniewski, T.; Letellier, M.; Lisowa, O.; Yusibov, V.; Koprowski, H.; Plucienniczak, A.; Legocki, A.B. A Plant-Derived Edible Vaccine against Hepatitis B Virus. FASEB J. 1999, 13, 1796–1799. [Google Scholar] [CrossRef] [PubMed]
- Joung, Y.H.; Park, S.H.; Moon, K.-B.; Jeon, J.-H.; Cho, H.-S.; Kim, H.-S. The Last Ten Years of Advancements in Plant-Derived Recombinant Vaccines against Hepatitis B. Int. J. Mol. Sci. 2016, 17, 1715. [Google Scholar] [CrossRef] [PubMed]
- Shanmugaraj, B.; Rattanapisit, K.; Manopwisedjaroen, S.; Thitithanyanont, A.; Phoolcharoen, W. Monoclonal Antibodies B38 and H4 Produced in Nicotiana Benthamiana Neutralize SARS-CoV-2 in Vitro. Front. Plant Sci. 2020, 11, 589995. [Google Scholar] [CrossRef] [PubMed]
- Frigerio, R.; Marusic, C.; Villani, M.E.; Lico, C.; Capodicasa, C.; Andreano, E.; Paciello, I.; Rappuoli, R.; Salzano, A.M.; Scaloni, A.; et al. Production of Two SARS-CoV-2 Neutralizing Antibodies with Different Potencies in Nicotiana Benthamiana. Front. Plant Sci. 2022, 13, 956741. [Google Scholar] [CrossRef] [PubMed]
- Jugler, C.; Sun, H.; Grill, F.; Kibler, K.; Esqueda, A.; Lai, H.; Li, Y.; Lake, D.; Chen, Q. Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component. Vaccines 2022, 10, 772. [Google Scholar] [CrossRef] [PubMed]
- Girard, L.S.; Fabis, M.J.; Bastin, M.; Courtois, D.; Pétiard, V.; Koprowski, H. Expression of a Human Anti-Rabies Virus Monoclonal Antibody in Tobacco Cell Culture. Biochem. Biophys. Res. Commun. 2006, 345, 602–607. [Google Scholar] [CrossRef] [PubMed]
- Pisuttinusart, N.; Rattanapisit, K.; Srisaowakarn, C.; Thitithanyanont, A.; Strasser, R.; Shanmugaraj, B.; Phoolcharoen, W. Neutralizing Activity of Anti-Respiratory Syncytial Virus Monoclonal Antibody Produced in Nicotiana Benthamiana. Hum. Vaccines Immunother. 2024, 20, 2327142. [Google Scholar] [CrossRef] [PubMed]
- Dobrovolny, M. GM Rice Delivers Antibodies against Deadly Rotavirus. Nature 2013. [Google Scholar] [CrossRef]
- Dong, J.-L.; Liang, B.-G.; Jin, Y.-S.; Zhang, W.-J.; Wang, T. Oral Immunization with pBsVP6-Transgenic Alfalfa Protects Mice against Rotavirus Infection. Virology 2005, 339, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Juárez, P.; Presa, S.; Espí, J.; Pineda, B.; Antón, M.T.; Moreno, V.; Buesa, J.; Granell, A.; Orzaez, D. Neutralizing Antibodies against Rotavirus Produced in Transgenically Labelled Purple Tomatoes. Plant Biotechnol. J. 2012, 10, 341–352. [Google Scholar] [CrossRef] [PubMed]
- Castillo-Esparza, J.F.; Gómez-Lim, M.A. Transient Expression in Cytoplasm and Apoplast of Rotavirus VP6 Protein Fused to Anti-DEC205 Antibody in Nicotiana Benthamiana and Nicotiana Sylvestris. Mol. Biotechnol. 2021, 63, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Krittanai, S.; Rattanapisit, K.; Bulaon, C.J.I.; Pitaksajjakul, P.; Keadsanti, S.; Ramasoota, P.; Strasser, R.; Phoolcharoen, W. Nicotiana Benthamiana as a Potential Source for Producing Anti-Dengue Virus D54 Neutralizing Therapeutic Antibody. Biotechnol. Rep. Amst. Neth. 2024, 42, e00844. [Google Scholar] [CrossRef] [PubMed]
- Dent, M.; Hurtado, J.; Paul, A.M.; Sun, H.; Lai, H.; Yang, M.; Esqueda, A.; Bai, F.; Steinkellner, H.; Chen, Q. Plant-Produced Anti-Dengue Virus Monoclonal Antibodies Exhibit Reduced Antibody-Dependent Enhancement of Infection Activity. J. Gen. Virol. 2016, 97, 3280–3290. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.M.; Paul, M.J.; Pinneh, E.; Shanmugaraj, B.; Ashall, J.; Ramalingam, S.; Hewson, R.; Diamond, M.S.; Fox, J.M.; Ma, J.K.-C. Characterisation of Chikungunya Virus Neutralising Monoclonal Antibodies Expressed in Tobacco Plants. J. Biotechnol. 2025, 406, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Sun, H.; Lai, H.; Neupane, B.; Bai, F.; Steinkellner, H.; Chen, Q. Plant-Produced Anti-Zika Virus Monoclonal Antibody Glycovariant Exhibits Abrogated Antibody-Dependent Enhancement of Infection. Vaccines 2023, 11, 755. [Google Scholar] [CrossRef] [PubMed]
- Kany, S.; Vollrath, J.T.; Relja, B. Cytokines in Inflammatory Disease. Int. J. Mol. Sci. 2019, 20, 6008. [Google Scholar] [CrossRef] [PubMed]
- Sirko, A.; Vaněk, T.; Góra-Sochacka, A.; Redkiewicz, P. Recombinant Cytokines from Plants. Int. J. Mol. Sci. 2011, 12, 3536–3552. [Google Scholar] [CrossRef] [PubMed]
- Bonjardim, C.A. Interferons (IFNs) Are Key Cytokines in Both Innate and Adaptive Antiviral Immune Responses—And Viruses Counteract IFN Action. Microbes Infect. 2005, 7, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Tayal, V.; Kalra, B.S. Cytokines and Anti-Cytokines as Therapeutics—An Update. Eur. J. Pharmacol. 2008, 579, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.-L.; Lin, Y.-L.; Lee, Y.-L.; Yang, N.-S.; Chan, M.-T. Expression of Bioactive Human Interferon-Gamma in Transgenic Rice Cell Suspension Cultures. Transgenic Res. 2004, 13, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Ohya, K.; Matsumura, T.; Ohashi, K.; Onuma, M.; Sugimoto, C. Expression of Two Subtypes of Human IFN-Alpha in Transgenic Potato Plants. J. Interferon Cytokine Res. Off. J. Int. Soc. Interferon Cytokine Res. 2001, 21, 595–602. [Google Scholar] [CrossRef] [PubMed]
- Arazi, T.; Slutsky, S.G.; Shiboleth, Y.M.; Wang, Y.; Rubinstein, M.; Barak, S.; Yang, J.; Gal-On, A. Engineering Zucchini Yellow Mosaic Potyvirus as a Non-Pathogenic Vector for Expression of Heterologous Proteins in Cucurbits. J. Biotechnol. 2001, 87, 67–82. [Google Scholar] [CrossRef] [PubMed]
- Arlen, P.A.; Falconer, R.; Cherukumilli, S.; Cole, A.; Cole, A.M.; Oishi, K.K.; Daniell, H. Field Production and Functional Evaluation of Chloroplast-Derived Interferon-Alpha2b. Plant Biotechnol. J. 2007, 5, 511–525. [Google Scholar] [CrossRef] [PubMed]
- Sawahel, W.A. The Production of Transgenic Potato Plants Expressing Human Alpha-Interferon Using Lipofectin-Mediated Transformation. Cell. Mol. Biol. Lett. 2002, 7, 19–29. [Google Scholar] [PubMed]
- Luchakivskaya, Y.; Kishchenko, O.; Gerasymenko, I.; Olevinskaya, Z.; Simonenko, Y.; Spivak, M.; Kuchuk, M. High-Level Expression of Human Interferon Alpha-2b in Transgenic Carrot (Daucus carota L.) Plants. Plant Cell Rep. 2011, 30, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chen, M.; Liu, X.W.; Zhang, H.C.; Shen, F.F.; Wang, G.P. Transient Expression of an Active Human Interferon-Beta in Lettuce. ResearchGate 2007, 112, 258–265. [Google Scholar] [CrossRef]
- Nabi-Afjadi, M.; Heydari, M.; Zalpoor, H.; Arman, I.; Sadoughi, A.; Sahami, P.; Aghazadeh, S. Lectins and Lectibodies: Potential Promising Antiviral Agents. Cell. Mol. Biol. Lett. 2022, 27, 37. [Google Scholar] [CrossRef] [PubMed]
- Lee, C. Griffithsin, a Highly Potent Broad-Spectrum Antiviral Lectin from Red Algae: From Discovery to Clinical Application. Mar. Drugs 2019, 17, 567. [Google Scholar] [CrossRef] [PubMed]
- Mazalovska, M.; Kouokam, J.C. Lectins as Promising Therapeutics for the Prevention and Treatment of HIV and Other Potential Coinfections. BioMed Res. Int. 2018, 2018, 3750646. [Google Scholar] [CrossRef] [PubMed]
- Bellande, K.; Lalo, A.; Ligat, L.; Roujol, D.; Jamet, E.; Canut, H. Recombinant N-Glycosylation Isoforms of Legume Lectins: Production and Purification from Nicotiana Benthamiana Leaves Following RuBisCO Depletion. Plant Physiol. Biochem. PPB 2020, 157, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Hoelscher, M.; Tiller, N.; Teh, A.Y.-H.; Wu, G.-Z.; Ma, J.K.-C.; Bock, R. High-Level Expression of the HIV Entry Inhibitor Griffithsin from the Plastid Genome and Retention of Biological Activity in Dried Tobacco Leaves. Plant Mol. Biol. 2018, 97, 357–370. [Google Scholar] [CrossRef] [PubMed]
- O’Keefe, B.R.; Vojdani, F.; Buffa, V.; Shattock, R.J.; Montefiori, D.C.; Bakke, J.; Mirsalis, J.; d’Andrea, A.-L.; Hume, S.D.; Bratcher, B.; et al. Scaleable Manufacture of HIV-1 Entry Inhibitor Griffithsin and Validation of Its Safety and Efficacy as a Topical Microbicide Component. Proc. Natl. Acad. Sci. USA 2009, 106, 6099–6104. [Google Scholar] [CrossRef] [PubMed]
- Vamvaka, E.; Arcalis, E.; Ramessar, K.; Evans, A.; O’Keefe, B.R.; Shattock, R.J.; Medina, V.; Stöger, E.; Christou, P.; Capell, T. Rice Endosperm Is Cost-Effective for the Production of Recombinant Griffithsin with Potent Activity against HIV. Plant Biotechnol. J. 2016, 14, 1427–1437. [Google Scholar] [CrossRef] [PubMed]
- Sexton, A.; Drake, P.M.; Mahmood, N.; Harman, S.J.; Shattock, R.J.; Ma, J.K.-C. Transgenic Plant Production of Cyanovirin-N, an HIV Microbicide. FASEB J. 2006, 20, 356–358. [Google Scholar] [CrossRef] [PubMed]
- Drake, P.M.W.; de Moraes Madeira, L.; Szeto, T.H.; Ma, J.K.-C. Transformation of Althaea officinalis L. by Agrobacterium Rhizogenes for the Production of Transgenic Roots Expressing the Anti-HIV Microbicide Cyanovirin-N. Transgenic Res. 2013, 22, 1225–1229. [Google Scholar] [CrossRef] [PubMed]
- O’Keefe, B.R.; Murad, A.M.; Vianna, G.R.; Ramessar, K.; Saucedo, C.J.; Wilson, J.; Buckheit, K.W.; da Cunha, N.B.; Araújo, A.C.G.; Lacorte, C.C.; et al. Engineering Soya Bean Seeds as a Scalable Platform to Produce cyanovirin-N, a non-ARV Microbicide against HIV. Plant Biotechnol. J. 2015, 13, 884–892. [Google Scholar] [CrossRef] [PubMed]
- Vamvaka, E.; Evans, A.; Ramessar, K.; Krumpe, L.R.H.; Shattock, R.J.; O’Keefe, B.R.; Christou, P.; Capell, T. Cyanovirin-N Produced in Rice Endosperm Offers Effective Pre-Exposure Prophylaxis against HIV-1BaL Infection in Vitro. Plant Cell Rep. 2016, 35, 1309–1319. [Google Scholar] [CrossRef] [PubMed]
- Sexton, A.; Harman, S.; Shattock, R.J.; Ma, J.K.-C. Design, Expression, and Characterization of a Multivalent, Combination HIV Microbicide. FASEB J. 2009, 23, 3590–3600. [Google Scholar] [CrossRef] [PubMed]
- Vamvaka, E.; Farré, G.; Molinos-Albert, L.M.; Evans, A.; Canela-Xandri, A.; Twyman, R.M.; Carrillo, J.; Ordóñez, R.A.; Shattock, R.J.; O’Keefe, B.R.; et al. Unexpected Synergistic HIV Neutralization by a Triple Microbicide Produced in Rice Endosperm. Proc. Natl. Acad. Sci. USA 2018, 115, E7854–E7862. [Google Scholar] [CrossRef] [PubMed]
- Seber Kasinger, L.E.; Dent, M.W.; Mahajan, G.; Hamorsky, K.T.; Matoba, N. A Novel Anti-HIV-1 Bispecific bNAb-Lectin Fusion Protein Engineered in a Plant-Based Transient Expression System. Plant Biotechnol. J. 2019, 17, 1646–1656. [Google Scholar] [CrossRef] [PubMed]
- Hilal, B.; Khan, M.M.; Fariduddin, Q. Recent Advancements in Deciphering the Therapeutic Properties of Plant Secondary Metabolites: Phenolics, Terpenes, and Alkaloids. Plant Physiol. Biochem. 2024, 211, 108674. [Google Scholar] [CrossRef] [PubMed]
- Shi, M.; Du, Z.; Hua, Q.; Kai, G. CRISPR/Cas9-Mediated Targeted Mutagenesis of bZIP2 in Salvia Miltiorrhiza Leads to Promoted Phenolic Acid Biosynthesis. Ind. Crops Prod. 2021, 167, 113560. [Google Scholar] [CrossRef]
- Janda, K.; Gutowska, I.; Geszke-Moritz, M.; Jakubczyk, K. The Common Cichory (Cichorium intybus L.) as a Source of Extracts with Health-Promoting Properties-A Review. Molecules 2021, 26, 1814. [Google Scholar] [CrossRef] [PubMed]
- Cankar, K.; Bundock, P.; Sevenier, R.; Häkkinen, S.T.; Hakkert, J.C.; Beekwilder, J.; van der Meer, I.M.; de Both, M.; Bosch, D. Inactivation of the Germacrene A Synthase Genes by CRISPR/Cas9 Eliminates the Biosynthesis of Sesquiterpene Lactones in Cichorium intybus L. Plant Biotechnol. J. 2021, 19, 2442–2453. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Yang, F.; Wu, Y.; Lu, J.; Gao, X.; Zhu, X.; Yang, J.; Liu, S.; Xiao, G.; Pan, X. Identification of Tanshinone I as Cap-Dependent Endonuclease Inhibitor with Broad-Spectrum Antiviral Effect. J. Virol. 2023, 97, e0079623. [Google Scholar] [CrossRef] [PubMed]
- Langat, R.; Chakrawarti, A.; Klatt, N.R. Cannabis Use in HIV: Impact on Inflammation, Immunity and the Microbiome. Curr. HIV/AIDS Rep. 2025, 22, 19. [Google Scholar] [CrossRef] [PubMed]
- Ross, I.A. (Ed.) Antiviral Activities of Cannabis. In Plant-Based Therapeutics, Volume 1: Cannabis sativa; Springer International Publishing: Cham, Switzerland, 2023; pp. 629–639. ISBN 978-3-031-35155-6. [Google Scholar]
- Liu, X.-F.; Zhu, J.; Ge, S.-Y.; Xia, L.-J.; Yang, H.-Y.; Qian, Y.-T.; Ren, F.-Z. Orally Administered Dendrobium Officinale and Its Polysaccharides Enhance Immune Functions in BALB/c Mice. Nat. Prod. Commun. 2011, 6, 867–870. [Google Scholar] [CrossRef] [PubMed]
- Kui, L.; Chen, H.; Zhang, W.; He, S.; Xiong, Z.; Zhang, Y.; Yan, L.; Zhong, C.; He, F.; Chen, J.; et al. Building a Genetic Manipulation Tool Box for Orchid Biology: Identification of Constitutive Promoters and Application of CRISPR/Cas9 in the Orchid, Dendrobium Officinale. Front. Plant Sci. 2016, 7, 2036. [Google Scholar] [CrossRef] [PubMed]
- Ouassaf, M.; Bourougaa, L.; Bahaz, F.; Alhatlani, B.Y. Exploring the Antiviral Potential of Artemisia Annua Through JAK-STAT Pathway Targeting: A Network Pharmacology Approach. Pharmaceuticals 2024, 17, 1539. [Google Scholar] [CrossRef] [PubMed]
- Qamar, F.; Ashrafi, K.; Singh, A.; Dash, P.K.; Abdin, M.Z. Artemisinin Production Strategies for Industrial Scale: Current Progress and Future Directions. Ind. Crops Prod. 2024, 218, 118937. [Google Scholar] [CrossRef]
- Sinha, S.; Sandhu, K.; Bisht, N.; Naliwal, T.; Saini, I.; Kaushik, P. Ascertaining the Paradigm of Secondary Metabolism Enhancement through Gene Level Modification in Therapeutic Plants. J. Young Pharm. 2019, 11, 337–343. [Google Scholar] [CrossRef]
- Grützner, R.; König, K.; Horn, C.; Engler, C.; Laub, A.; Vogt, T.; Marillonnet, S. A Transient Expression Tool Box for Anthocyanin Biosynthesis in Nicotiana Benthamiana. Plant Biotechnol. J. 2024, 22, 1238–1250. [Google Scholar] [CrossRef] [PubMed]
- Reed, J.; Osbourn, A. Engineering Terpenoid Production through Transient Expression in Nicotiana Benthamiana. Plant Cell Rep. 2018, 37, 1431–1441. [Google Scholar] [CrossRef] [PubMed]
- Reed, J.; Orme, A.; El-Demerdash, A.; Owen, C.; Martin, L.B.B.; Misra, R.C.; Kikuchi, S.; Rejzek, M.; Martin, A.C.; Harkess, A.; et al. Elucidation of the Pathway for Biosynthesis of Saponin Adjuvants from the Soapbark Tree. Science 2023, 379, 1252–1264. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Ding, T.; Chang, H.; Peng, Y.; Li, J.; Liang, X.; Ma, H.; Li, F.; Ren, M.; Wang, W. Plant-Derived Strategies to Fight against Severe Acute Respiratory Syndrome Coronavirus 2. Eur. J. Med. Chem. 2024, 264, 116000. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Tan, H.; Li, Q.; Chen, J.; Gao, S.; Wang, Y.; Chen, W.; Zhang, L. CRISPR/Cas9-Mediated Efficient Targeted Mutagenesis of RAS in Salvia Miltiorrhiza. Phytochemistry 2018, 148, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Bamba, T.; Kondo, A.; Hasunuma, T. Metabolomics-Based Development of Bioproduction Processes toward Industrial-Scale Production. Curr. Opin. Biotechnol. 2024, 85, 103057. [Google Scholar] [CrossRef] [PubMed]
- Chastang, T.; Pozzobon, V.; Taidi, B.; Courot, E.; Clément, C.; Pareau, D. Resveratrol Production by Grapevine Cells in Fed-Batch Bioreactor: Experiments and Modelling. Biochem. Eng. J. 2018, 131, 9–16. [Google Scholar] [CrossRef]
- Rebelo, B.A.; Folgado, A.; Ferreira, A.C.; Abranches, R. Production of the SARS-CoV-2 Spike Protein and Its Receptor Binding Domain in Plant Cell Suspension Cultures. Front. Plant Sci. 2022, 13, 995429. [Google Scholar] [CrossRef] [PubMed]
- Takova, K.; Tonova, V.; Minkov, I.; Mardanova, E.S.; Ravin, N.V.; Kotsev, S.; Pishmisheva, M.; Zahmanova, G. Plant-Based Antigen Production Strategy for SARS-CoV-2 Nucleoprotein and RBD and Its Application for Detection of Antibody Responses in COVID-19 Patients. Appl. Sci. 2025, 15, 786. [Google Scholar] [CrossRef]
- Tusé, D.; McNulty, M.; McDonald, K.A.; Buchman, L.W. A Review and Outlook on Expression of Animal Proteins in Plants. Front. Plant Sci. 2024, 15, 1426239. [Google Scholar] [CrossRef] [PubMed]
- Fischer, R.; Buyel, J.F. Molecular Farming—The Slope of Enlightenment. Biotechnol. Adv. 2020, 40, 107519. [Google Scholar] [CrossRef] [PubMed]
- Kaldis, A.; Uddin, M.S.; Guluarte, J.O.; Martin, C.; Alexander, T.W.; Menassa, R. Development of a Plant-Based Oral Vaccine Candidate against the Bovine Respiratory Pathogen Mannheimia Haemolytica. Front. Plant Sci. 2023, 14, 1251046. [Google Scholar] [CrossRef] [PubMed]
- Pantazica, A.-M.M.; Cucos, L.-M.; Stavaru, C.; Clarke, J.-L.; Branza-Nichita, N. Challenges and Prospects of Plant-Derived Oral Vaccines against Hepatitis B and C Viruses. Plants 2021, 10, 2037. [Google Scholar] [CrossRef] [PubMed]
- Lucero, M.S.; Chimeno Zoth, S.; Jaton, J.; Gravisaco, M.J.; Pinto, S.; Richetta, M.; Berinstein, A.; Gómez, E. Oral Immunization With Plant-Based Vaccine Induces a Protective Response Against Infectious Bursal Disease. Front. Plant Sci. 2021, 12, 741469. [Google Scholar] [CrossRef] [PubMed]
- Pomatto, M.A.C.; Gai, C.; Negro, F.; Massari, L.; Deregibus, M.C.; Grange, C.; De Rosa, F.G.; Camussi, G. Plant-Derived Extracellular Vesicles as a Delivery Platform for RNA-Based Vaccine: Feasibility Study of an Oral and Intranasal SARS-CoV-2 Vaccine. Pharmaceutics 2023, 15, 974. [Google Scholar] [CrossRef] [PubMed]
- Rouf, R.; Uddin, S.J.; Sarker, D.K.; Islam, M.T.; Ali, E.S.; Shilpi, J.A.; Nahar, L.; Tiralongo, E.; Sarker, S.D. Antiviral Potential of Garlic (Allium sativum) and Its Organosulfur Compounds: A Systematic Update of Pre-Clinical and Clinical Data. Trends Food Sci. Technol. 2020, 104, 219–234. [Google Scholar] [CrossRef] [PubMed]
- Tatarintsev, A.V.; Vrzheshch, P.V.; Schegolev, A.A.; Yershov, D.E.; Turgiev, A.S.; Varfolomeyev, S.D.; Kornilayeva, G.V.; Makarova, T.V.; Karamov, E.V. Ajoene Antagonizes Integrin-Dependent Processes in HIV-Infected T-Lymphoblasts. AIDS Lond. Engl. 1992, 6, 1215–1217. [Google Scholar]
- Yang, J.; Li, L.; Tan, S.; Jin, H.; Qiu, J.; Mao, Q.; Li, R.; Xia, C.; Jiang, Z.-H.; Jiang, S.; et al. A Natural Theaflavins Preparation Inhibits HIV-1 Infection by Targeting the Entry Step: Potential Applications for Preventing HIV-1 Infection. Fitoterapia 2012, 83, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Yang, X.-W.; Schols, D.; Mori, M.; Botta, B.; Chevigné, A.; Mulinge, M.; Steinmetz, A.; Schmit, J.-C.; Seguin-Devaux, C. Active Components from Cassia Abbreviata Prevent HIV-1 Entry by Distinct Mechanisms of Action. Int. J. Mol. Sci. 2021, 22, 5052. [Google Scholar] [CrossRef] [PubMed]
- Apaza Ticona, L.; Bermejo, P.; Guerra, J.A.; Abad, M.J.; Beltrán, M.; Martín Lázaro, R.; Alcamí, J.; Bedoya, L.M. Ethanolic Extract of Artemisia Campestris Subsp. Glutinosa (Besser) Batt. Inhibits HIV-1 Replication in Vitro through the Activity of Terpenes and Flavonoids on Viral Entry and NF-κB Pathway. J. Ethnopharmacol. 2020, 263, 113163. [Google Scholar] [CrossRef] [PubMed]
- Connell, B.J.; Chang, S.-Y.; Prakash, E.; Yousfi, R.; Mohan, V.; Posch, W.; Wilflingseder, D.; Moog, C.; Kodama, E.N.; Clayette, P.; et al. A Cinnamon-Derived Procyanidin Compound Displays Anti-HIV-1 Activity by Blocking Heparan Sulfate- and Co-Receptor- Binding Sites on Gp120 and Reverses T Cell Exhaustion via Impeding Tim-3 and PD-1 Upregulation. PLoS ONE 2016, 11, e0165386. [Google Scholar] [CrossRef] [PubMed]
- Cui, Q.; Du, R.; Anantpadma, M.; Schafer, A.; Hou, L.; Tian, J.; Davey, R.A.; Cheng, H.; Rong, L. Identification of Ellagic Acid from Plant Rhodiola rosea L. as an Anti-Ebola Virus Entry Inhibitor. Viruses 2018, 10, 152. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Xiong, S.; Xiang, Y.-F.; Guo, C.-W.; Ge, F.; Yang, C.-R.; Zhang, Y.-J.; Wang, Y.-F.; Kitazato, K. Antiviral Activity and Possible Mechanisms of Action of Pentagalloylglucose (PGG) against Influenza A Virus. Arch. Virol. 2011, 156, 1359–1369. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-W.; Tsai, F.-J.; Tsai, C.-H.; Lai, C.-C.; Wan, L.; Ho, T.-Y.; Hsieh, C.-C.; Chao, P.-D.L. Anti-SARS Coronavirus 3C-like Protease Effects of Isatis Indigotica Root and Plant-Derived Phenolic Compounds. Antiviral Res. 2005, 68, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Wen, C.-C.; Kuo, Y.-H.; Jan, J.-T.; Liang, P.-H.; Wang, S.-Y.; Liu, H.-G.; Lee, C.-K.; Chang, S.-T.; Kuo, C.-J.; Lee, S.-S.; et al. Specific Plant Terpenoids and Lignoids Possess Potent Antiviral Activities against Severe Acute Respiratory Syndrome Coronavirus. J. Med. Chem. 2007, 50, 4087–4095. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Kim, J.H.; Kim, Y.M.; Jeong, H.J.; Kim, D.W.; Park, K.H.; Kwon, H.-J.; Park, S.-J.; Lee, W.S.; Ryu, Y.B. Tanshinones as Selective and Slow-Binding Inhibitors for SARS-CoV Cysteine Proteases. Bioorg. Med. Chem. 2012, 20, 5928–5935. [Google Scholar] [CrossRef] [PubMed]
- Jo, S.; Kim, S.; Shin, D.H.; Kim, M.-S. Inhibition of SARS-CoV 3CL Protease by Flavonoids. J. Enzyme Inhib. Med. Chem. 2020, 35, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Jeong, H.J.; Kim, J.H.; Kim, Y.M.; Park, S.-J.; Kim, D.; Park, K.H.; Lee, W.S.; Ryu, Y.B. Diarylheptanoids from Alnus Japonica Inhibit Papain-like Protease of Severe Acute Respiratory Syndrome Coronavirus. Biol. Pharm. Bull. 2012, 35, 2036–2042. [Google Scholar] [CrossRef] [PubMed]
- Cho, J.K.; Curtis-Long, M.J.; Lee, K.H.; Kim, D.W.; Ryu, H.W.; Yuk, H.J.; Park, K.H. Geranylated Flavonoids Displaying SARS-CoV Papain-like Protease Inhibition from the Fruits of Paulownia Tomentosa. Bioorg. Med. Chem. 2013, 21, 3051–3057. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-Y.; Ko, J.-A.; Kim, D.W.; Kim, Y.M.; Kwon, H.-J.; Jeong, H.J.; Kim, C.Y.; Park, K.H.; Lee, W.S.; Ryu, Y.B. Chalcones Isolated from Angelica Keiskei Inhibit Cysteine Proteases of SARS-CoV. J. Enzyme Inhib. Med. Chem. 2016, 31, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Yi, D.; Lei, X.; Zhao, J.; Zhang, Y.; Cui, X.; Xiao, X.; Jiao, T.; Dong, X.; Zhao, X.; et al. Corilagin Inhibits SARS-CoV-2 Replication by Targeting Viral RNA-Dependent RNA Polymerase. Acta Pharm. Sin. B 2021, 11, 1555–1567. [Google Scholar] [CrossRef] [PubMed]
- Munafò, F.; Donati, E.; Brindani, N.; Ottonello, G.; Armirotti, A.; De Vivo, M. Quercetin and Luteolin Are Single-Digit Micromolar Inhibitors of the SARS-CoV-2 RNA-Dependent RNA Polymerase. Sci. Rep. 2022, 12, 10571. [Google Scholar] [CrossRef] [PubMed]
- Keum, Y.-S.; Lee, J.M.; Yu, M.-S.; Chin, Y.-W.; Jeong, Y.-J. Inhibition of SARS Coronavirus Helicase by Baicalein. Bull. Korean Chem. Soc. 2013, 34, 3187–3188. [Google Scholar] [CrossRef]
- Zeng, J.; Weissmann, F.; Bertolin, A.P.; Posse, V.; Canal, B.; Ulferts, R.; Wu, M.; Harvey, R.; Hussain, S.; Milligan, J.C.; et al. Identifying SARS-CoV-2 Antiviral Compounds by Screening for Small Molecule Inhibitors of Nsp13 Helicase. Biochem. J. 2021, 478, 2405–2423. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Nakamura, N.; Hattori, M.; Kakuda, H.; Qiao, J.; Yu, H. Inhibitory Effects on HIV-1 Protease of Constituents from the Wood of Xanthoceras Sorbifolia. J. Nat. Prod. 2000, 63, 238–242. [Google Scholar] [CrossRef] [PubMed]
- Martini, R.; Esposito, F.; Corona, A.; Ferrarese, R.; Ceresola, E.R.; Visconti, L.; Tintori, C.; Barbieri, A.; Calcaterra, A.; Iovine, V.; et al. Natural Product Kuwanon-L Inhibits HIV-1 Replication through Multiple Target Binding. Chembiochem Eur. J. Chem. Biol. 2017, 18, 374–377. [Google Scholar] [CrossRef] [PubMed]
- Ravanelli, N.; Santos, K.P.; Motta, L.B.; Lago, J.H.G.; Furlan, C.M. Alkaloids from Croton Echinocarpus Baill.: Anti-HIV Potential. S. Afr. J. Bot. 2016, 102, 153–156. [Google Scholar] [CrossRef]
- Troost, B.; Mulder, L.M.; Diosa-Toro, M.; van de Pol, D.; Rodenhuis-Zybert, I.A.; Smit, J.M. Tomatidine, a Natural Steroidal Alkaloid Shows Antiviral Activity towards Chikungunya Virus in Vitro. Sci. Rep. 2020, 10, 6364. [Google Scholar] [CrossRef] [PubMed]
- Sharma, K.B.; Subramani, C.; Ganesh, K.; Sharma, A.; Basu, B.; Balyan, S.; Sharma, G.; Ka, S.; Deb, A.; Srivastava, M.; et al. Withaferin A Inhibits Chikungunya Virus nsP2 Protease and Shows Antiviral Activity in the Cell Culture and Mouse Model of Virus Infection. PLoS Pathog. 2024, 20, e1012816. [Google Scholar] [CrossRef] [PubMed]
- Bourjot, M.; Leyssen, P.; Neyts, J.; Dumontet, V.; Litaudon, M. Trigocherrierin A, a Potent Inhibitor of Chikungunya Virus Replication. Molecules 2014, 19, 3617–3627. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).