Deciphering of SOX9 Functions in Pancreatic Cancer Cells
Abstract
1. Introduction
2. Results
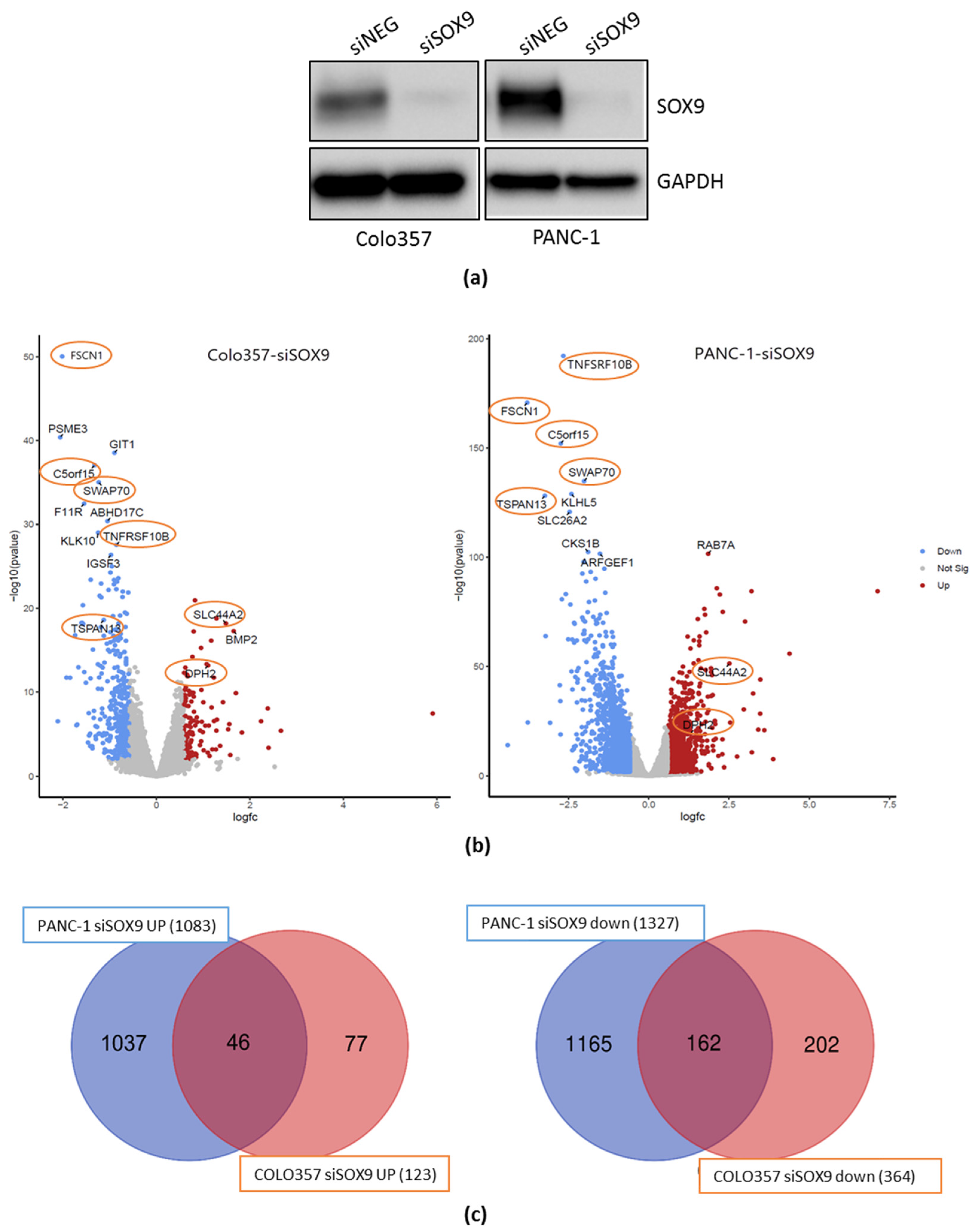
2.1. Generation and Evaluation of cDNA Libraries
2.2. The Transcriptomic Alterations of PAAD Cells upon Knockdown of the SOX9 Gene
2.2.1. Gene Ontology
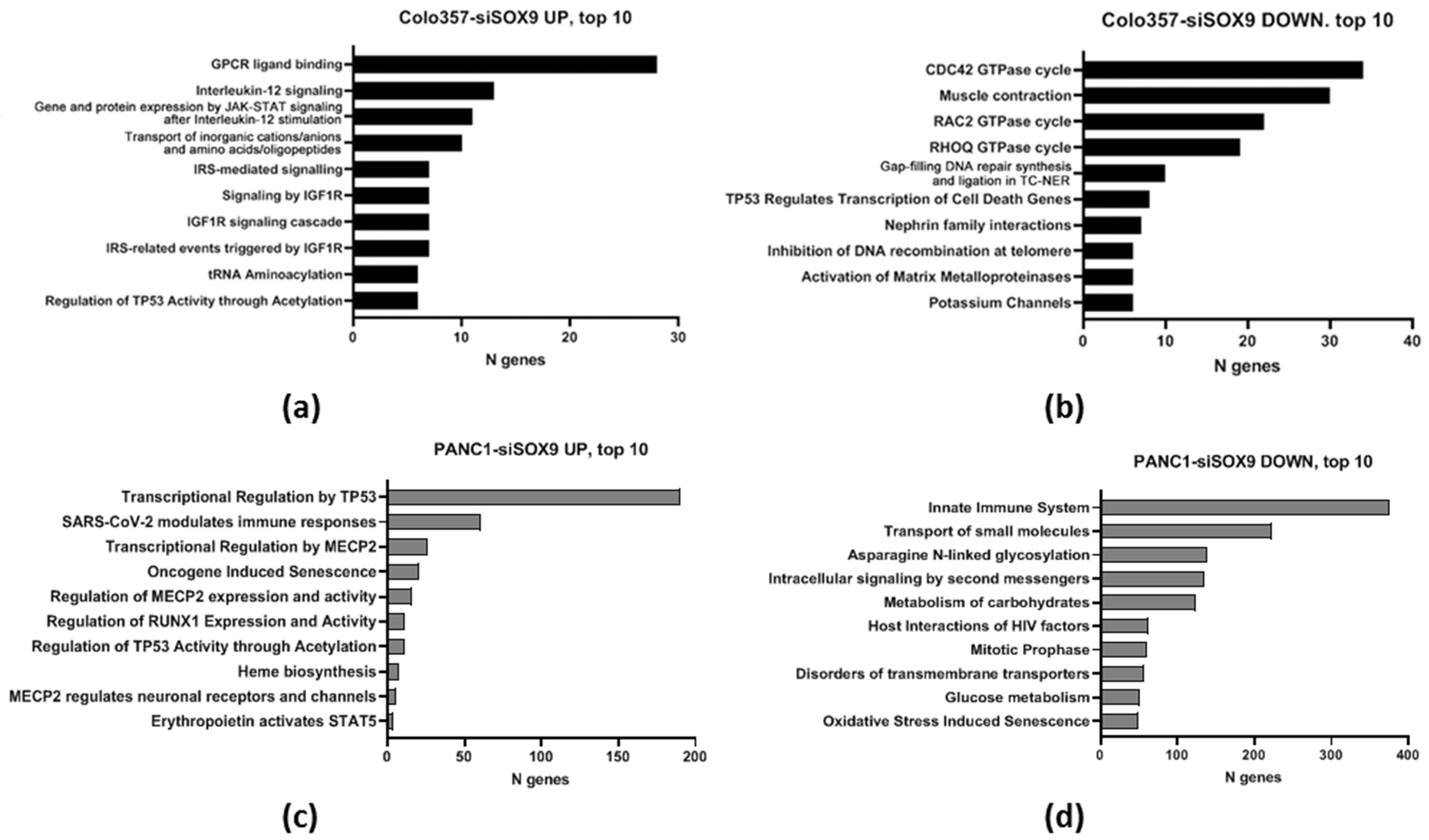
2.2.2. Reactome Gene Representation Analysis
2.2.3. Reactome Gene Expression Analysis
| Pathway | COLO357–siSOX9 | PANC-1–siSOX9 | ||
|---|---|---|---|---|
| Av. FC 2 | Number of Genes | Av. FC | Number of Genes | |
| Up-regulated pathways | ||||
| Regulation of TP53 activity through acetylation (R-HSA-6804758) | 5.09 | 6 | 19.63 | 11 |
| Erythropoietin activates phospholipase C gamma (PLCG) (R-HSA-9027277) | 28.57 | 1 | 12.49 | 4 |
| Erythropoietin activates STAT5 (R-HSA-9027283) | 28.57 | 1 | 5.31 | 4 |
| Organic anion transporters (R-HSA-428643) | 15.01 | 1 | 14.81 | 4 |
| Defective SLC17A5 causes Salla disease (SD) and ISSD (R-HSA-5619035) | 15.01 | 1 | 46.81 | 1 |
| Down-regulated pathways | ||||
| MAPK6/MAPK4 signaling (R-HSA-5687128) | −17.95 | 15 | −8.011 | 35 |
| Opposite regulated pathways | ||||
| Erythropoietin activates phosphoinositide-3-kinase (PI3K) (R-HSA-9027276) | 4.75 (up) | 3 | −0.12 (down) | 7 |
| Tandem pore domain potassium channels (R-HSA-1296346) | −16.0 (down) | 1 | 21.05 (up) | 1 |
| Tandem of pore domain in a weak inwardly rectifying K+ channels (TWIK) (R-HSA-1299308) | −16.0 (down) | 1 | 21.05 (up) | 1 |
| Phase 4—resting membrane potential (R-HSA-5576886) | −16.66 (down) | 3 | 12.03 (up) | 2 |
| TP53 regulates transcription of cell death genes (R-HSA-5633008) | −0.35 (down) | 8 | 4.59 (up) | 24 |
| Alternative complement activation (R-HSA-173736) | −39.74 (down) | 1 | N.i. 3 | N.i. |
| Activation of C3 and C5 (R-HSA-174577) | −39.74 (down) | 1 | N.i. | N.i. |
2.2.4. GSEA Expression Analysis
| Hallmark Name | SIZE | ES | NES | NOM p-Val | FDR q-Val | FWER p-Val |
|---|---|---|---|---|---|---|
| COLO357–siSOX9 up-regulated Hallmark gene sets | ||||||
| MYC_TARGETS_V1 2 | 36 | 0.39 | 2.28 | 0 | 0.0031 | 0.002 |
| MYC_TARGETS_V2 2 | 36 | 0.43 | 2.22 | 0 | 0.0022 | 0.003 |
| UV_RESPONSE_DN 2 | 25 | 0.38 | 1.91 | 0.0071 | 0.018 | 0.03 |
| COLO357–siSOX9 down-regulated Hallmark gene sets | ||||||
| ESTROGEN_RESPONSE_LATE 2 | 57 | −0.42 | −2.03 | 0 | 0.015 | 0.019 |
| PANC-1–siSOX9 up-regulated Hallmark gene sets | ||||||
| KRAS_SIGNALING_DN 2 | 30 | 0.40 | 1.78 | 0.071 | 0.016 | 0.035 |
| PANC-1–siSOX9 down-regulated Hallmark gene sets | ||||||
| E2F_TARGETS 2 | 118 | −0.42 | −2.06 | 0 | 0.042 | 0.04 |
| (KRAS_SIGNALING_UP) 2 | 54 | (−0.43) | (−1.82) | 0 | (0.030) | (0.053) |
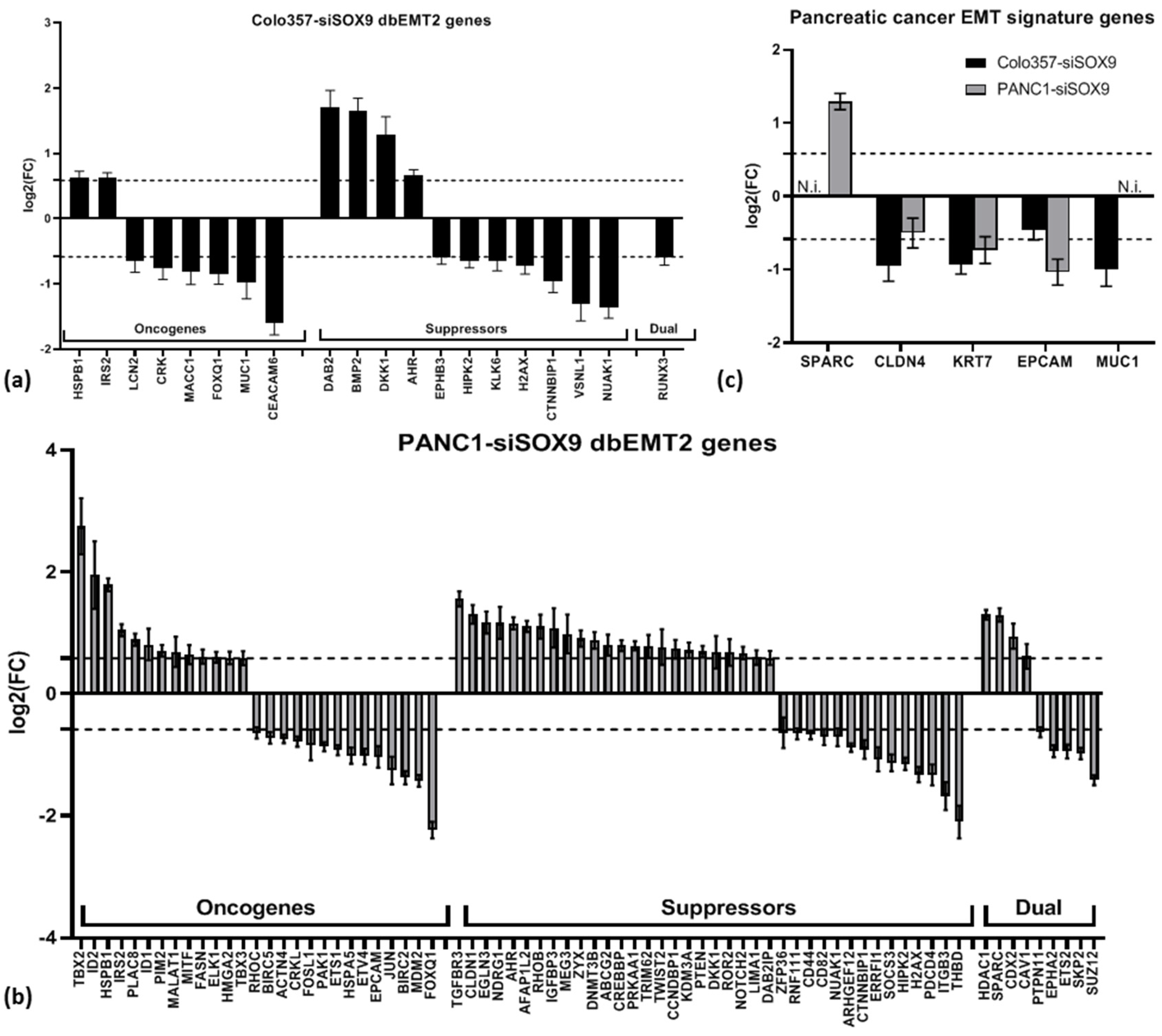
2.2.5. Oncogenes, Tumor Suppressors, EMT Genes
2.2.6. P53 Gene-Related Alterations
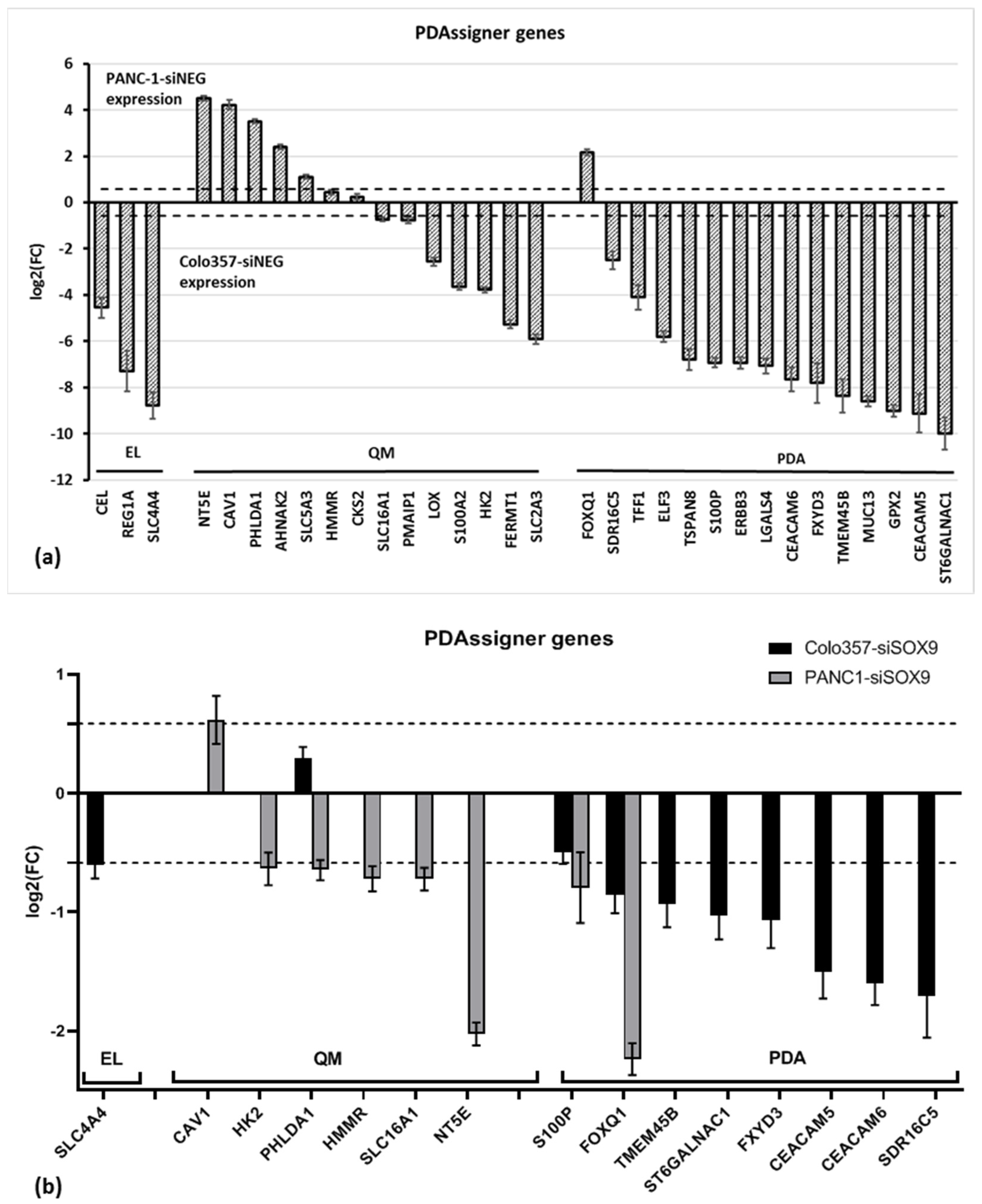
2.2.7. PDAssigner
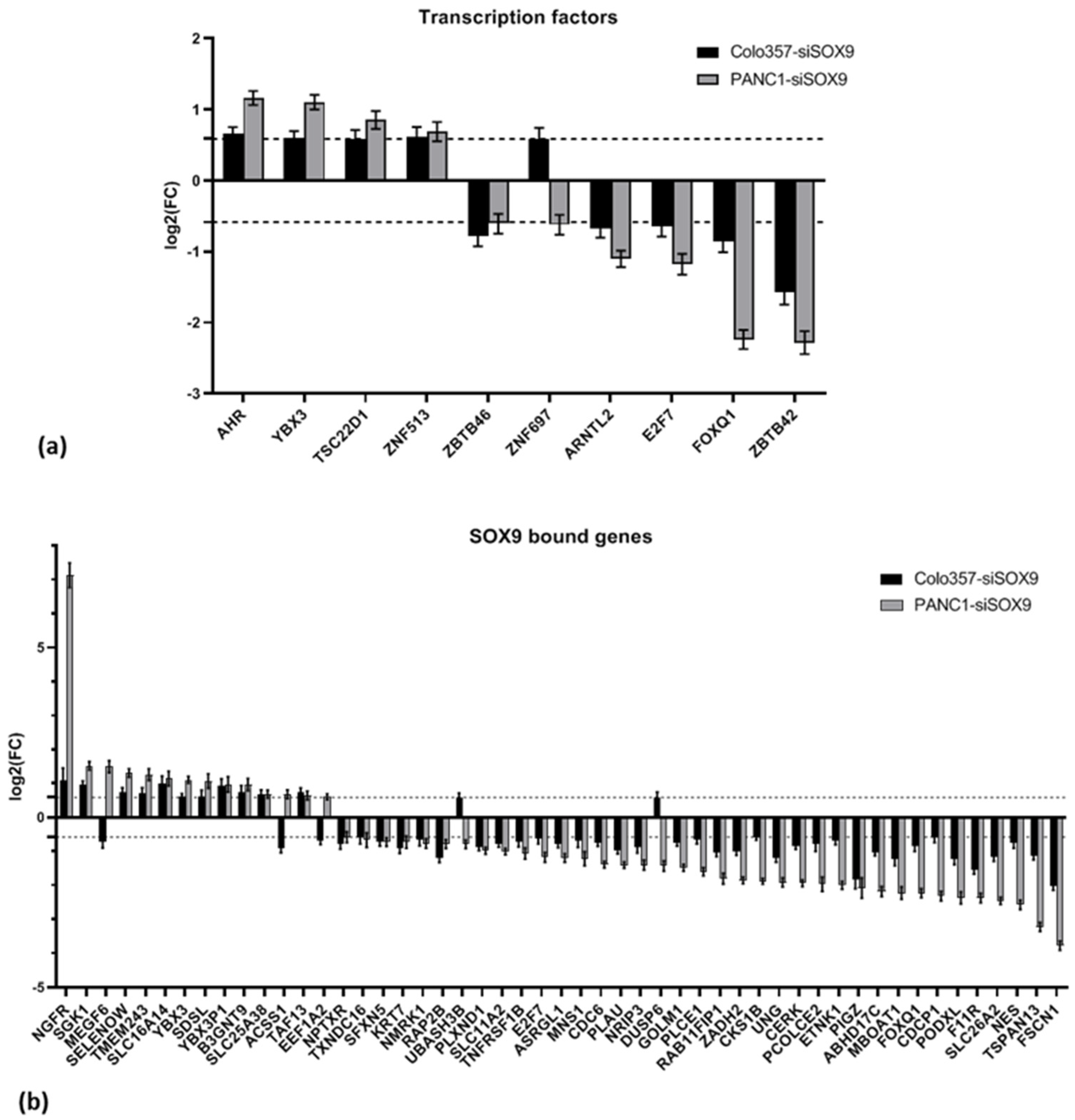
2.2.8. Transcription Factors
2.2.9. Genes Bound to SOX9 by ChIP
2.3. Analysis of Pancreatic Cancer Crucial and Prognostic Gene Sets
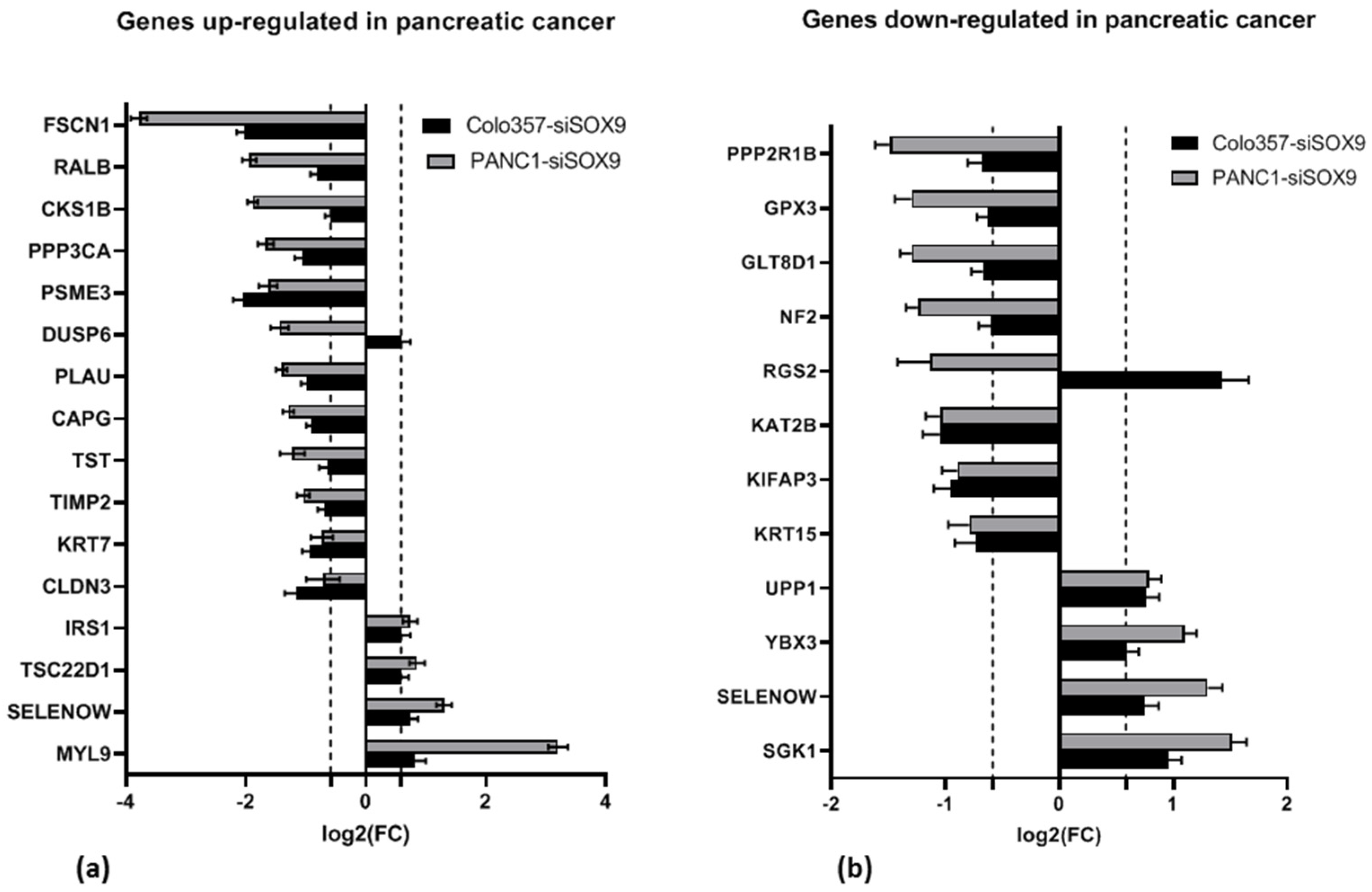
2.3.1. Differentially Expressed Genes in Pancreatic Cancer
2.3.2. Prognostic Gene Signatures
| Pancreatic Cancer Gene Signature | Reference | Number of Genes in a Signature | Genes Affected Under SOX9 Knock-Down | |
|---|---|---|---|---|
| COLO357–siSOX9 | PANC-1–siSOX9 | |||
| Gene signature of metastasis | [44] | 17 | No genes * | COL1A1, EIF4E2 |
| Gene signature for metastasis and survival | [45] | 6 | No genes * | No genes * |
| Patients outcome gene expression signature | [46] | 36 | No genes * | ADM, ITGBL1, CDK2AP1, ARRB1, SEMA3A, PHLDA1, BLM, KIF14, PXN |
| Survival gene expression signature | [47] | 19 | No genes * | AVPI1, CLIP4, RGS20 |
| Genetic predictors for survival under chemotherapy | [48] | 23 | CDRL2, RFNLA | CDRL2 |
| Gene expression signature for prognosis | [49] | 5 | E2F7 | E2F7 |
| KRAS-associated metabolic genes for prognosis | [50] | 6 | GPX3 | GPX3 |
| An EMT-related gene signature for predicting response to adjuvant chemotherapy | [51] | 8 | ITGB6 | DLX2 |
| Prognostic gene signature | [52] | 18 | CD9, UBASH3B, FYN | GNA15, UBASH3B, SERPINE1 |
| Metabolism-related genes as targets for immunotherapy | [53] | 24 | SLC1A3 | IGFBP6, HK2, SLC2A1, ALDH1A3 |
| MYC-based signature for prognosis and chemoresistance | [54] | 7 | ITGB6 | No genes * |

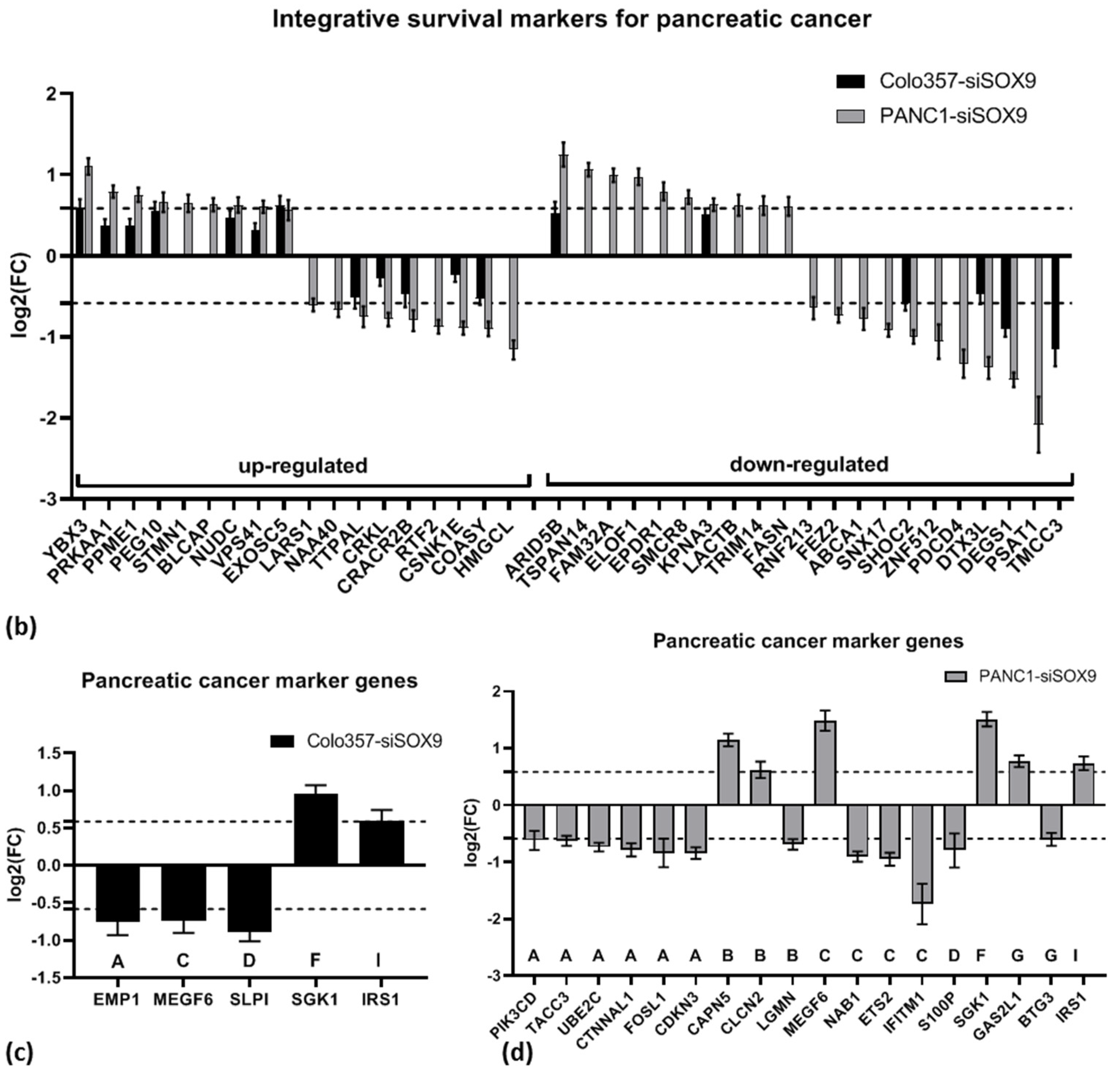
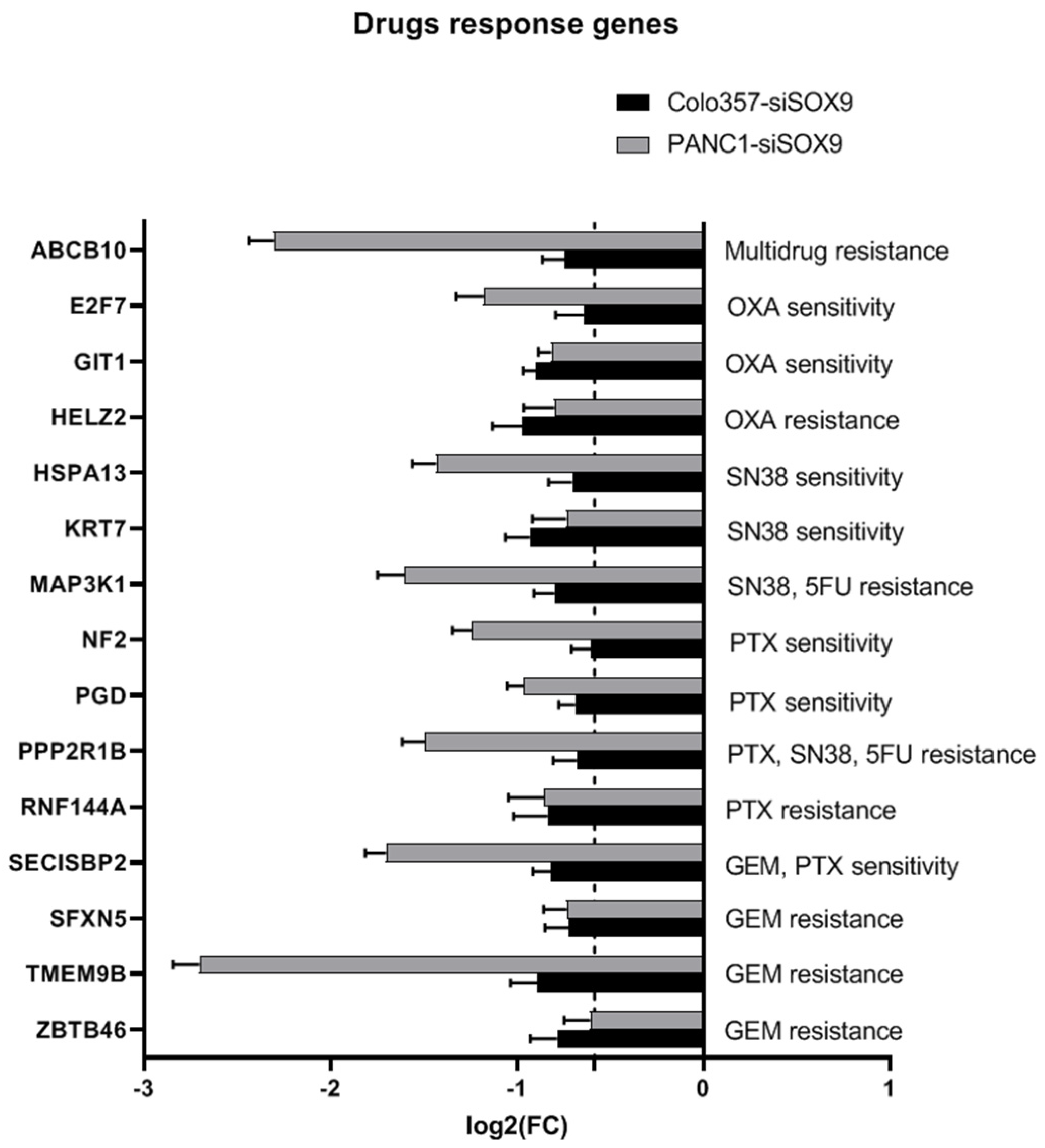
2.3.3. Drug Resistance Genes
3. Discussion
3.1. Primary Analysis of Transcriptomes
3.2. Pathway and Hallmarks Analysis: Proliferation and Cell Cycle
3.3. Oncogenes and Tumor Suppressor Genes
3.4. PDAssigner
3.5. Transcription Factors
3.6. Genes Bound to SOX9 by ChIP
3.7. Differentially Expressing Genes
3.8. Prognostic Gene Signatures
3.9. Drug Resistance Genes
4. Materials and Methods
4.1. Materials and Cell Cultures
4.2. Knockdown of SOX9 via RNA Interference
4.3. Western Blot Analysis
4.4. RNA-Sequencing (RNA-Seq)
4.5. Genes and Pathways Annotation
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ChIP | Chromatin immunoprecipitation |
| EMT | Epithelial–mesenchymal transition |
| ERK | Extracellular signal-regulated kinase |
| FC | Fold change |
| GO | Gene Ontology |
| GSEA | Gene set enrichment analysis |
| HMG | High mobility group |
| MAPK | Mitogen-activated protein kinase |
| NF-κB | Nuclear factor kappa B |
| PAAD | Pancreatic adenocarcinoma |
| PDA | Pancreatic ductal adenocarcinoma |
| QM-PDA | Quasi-mesenchymal pancreatic adenocarcinoma |
| RNA-Seq | RNA-sequencing |
| SHH | Sonic Hedgehog |
| SiRNA | Small interfering RNA |
| siNEG | Neutral siRNA (negative control) |
| siSOX9 | SOX9-specific siRNA |
| TF | Transcription factor |
| TGF-β | Transforming growth factor |
| TPM | Transcripts per million |
| bp | Base pairs |
References
- Grimm, D.; Bauer, J.; Wise, P.; Krüger, M.; Simonsen, U.; Wehland, M.; Infanger, M.; Corydon, T.J. The role of sox family members in solid tumours and metastasis. Semin. Cancer Biol. 2020, 67, 122–153. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.H.; Jankowski, A.; Cheah, K.S.; Prabhakar, S.; Jauch, R. Soxe transcription factors form selective dimers on non-compact DNA motifs through multifaceted interactions between dimerization and high-mobility group domains. Sci. Rep. 2015, 5, 10398. [Google Scholar] [CrossRef] [PubMed]
- Seymour, P.A. Sox9: A master regulator of the pancreatic program. Rev. Diabet. Stud. 2014, 11, 51–83. [Google Scholar] [CrossRef]
- Ming, Z.; Vining, B.; Bagheri-Fam, S.; Harley, V. Sox9 in organogenesis: Shared and unique transcriptional functions. Cell. Mol. Life Sci. 2022, 79, 522. [Google Scholar] [CrossRef] [PubMed]
- Jasim, S.A.; Farhan, S.H.; Ahmad, I.; Hjazi, A.; Kumar, A.; Jawad, M.A.; Pramanik, A.; Altalbawy, M.A.F.; Alsaadi, S.B.; Abosaoda, M.K. A cutting-edge investigation of the multifaceted role of sox family genes in cancer pathogenesis through the modulation of various signaling pathways. Funct. Integr. Genom. 2025, 25, 6. [Google Scholar] [CrossRef]
- Wang, J.; Wan, X.; Le, Q. Cross-regulation between sox9 and the canonical wnt signalling pathway in stem cells. Front. Mol. Biosci. 2023, 10, 1250530. [Google Scholar] [CrossRef]
- Nigam, M.; Mishra, A.P.; Deb, V.K.; Dimri, D.B.; Tiwari, V.; Bungau, S.G.; Bungau, A.F.; Radu, A.F. Evaluation of the association of chronic inflammation and cancer: Insights and implications. Biomed. Pharmacother. 2023, 164, 115015. [Google Scholar] [CrossRef]
- Bi, Q.; Wu, J.Y.; Qiu, X.M.; Zhang, J.D.; Sun, Z.J.; Wang, W. Tumor-associated inflammation: The tumor-promoting immunity in the early stages of tumorigenesis. J. Immunol. Res. 2022, 2022, 3128933. [Google Scholar] [CrossRef]
- Kapoor, G.; Prakash, S.; Jaiswal, V.; Singh, A.K. Chronic inflammation and cancer: Key pathways and targeted therapies. Cancer Investig. 2025, 43, 1–23. [Google Scholar] [CrossRef]
- He, C.; Zhao, C.; Liu, S.; Zhong, Y.; Liu, L.; Cai, D. Down-regulation of mir-138-5p protects chondrocytes atdc5 and chon-001 from il-1 beta-induced inflammation via up-regulating sox9. Curr. Pharm. Des. 2020, 25, 4613–4621. [Google Scholar] [CrossRef]
- Scharf, G.M.; Kilian, K.; Cordero, J.; Wang, Y.; Grund, A.; Hofmann, M.; Froese, N.; Wang, X.; Kispert, A.; Kist, R.; et al. Inactivation of sox9 in fibroblasts reduces cardiac fibrosis and inflammation. JCI Insight 2019, 5, e126721. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.Y.; Lian, P.; Zheng, P.S. Sox9, a potential tumor suppressor in cervical cancer, transactivates p21waf1/cip1 and suppresses cervical tumor growth. Oncotarget 2015, 6, 20711–20722. [Google Scholar] [CrossRef] [PubMed]
- Saegusa, M.; Hashimura, M.; Suzuki, E.; Yoshida, T.; Kuwata, T. Transcriptional up-regulation of sox9 by nf-κb in endometrial carcinoma cells, modulating cell proliferation through alteration in the p14arf/p53/p21waf1 pathway. Am. J. Pathol. 2012, 181, 684–692. [Google Scholar] [CrossRef]
- Shakhova, O.; Cheng, P.; Mishra, P.J.; Zingg, D.; Schaefer, S.M.; Debbache, J.; Hausel, J.; Matter, C.; Guo, T.; Davis, S.; et al. Antagonistic cross-regulation between sox9 and sox10 controls an anti-tumorigenic program in melanoma. PLoS Genet. 2015, 11, e1004877. [Google Scholar] [CrossRef] [PubMed]
- Prevostel, C.; Rammah-Bouazza, C.; Trauchessec, H.; Canterel-Thouennon, L.; Busson, M.; Ychou, M.; Blache, P. Sox9 is an atypical intestinal tumor suppressor controlling the oncogenic wnt/ss-catenin signaling. Oncotarget 2016, 7, 82228–82243. [Google Scholar] [CrossRef]
- Kopantzev, E.; Kondratyeva, L.; Kopantseva, M.; Kashkin, K.; Gnatenko, D.; Grigorieva, E.; Alekseenko, I.; Safina, D.; Chernov, I. Sox9 protein in pancreatic cancer regulates multiple cellular networks in a cell-specific manner. Biomedicines 2022, 10, 1466. [Google Scholar] [CrossRef]
- Panda, M.; Tripathi, S.K.; Biswal, B.K. Sox9: An emerging driving factor from cancer progression to drug resistance. Biochim. Biophys. Acta Rev. Cancer 2021, 1875, 188517. [Google Scholar] [CrossRef]
- Higashihara, T.; Yoshitomi, H.; Nakata, Y.; Kagawa, S.; Takano, S.; Shimizu, H.; Kato, A.; Furukawa, K.; Ohtsuka, M.; Miyazaki, M. Sex determining region y box 9 induces chemoresistance in pancreatic cancer cells by induction of putative cancer stem cell characteristics and its high expression predicts poor prognosis. Pancreas 2017, 46, 1296–1304. [Google Scholar] [CrossRef]
- Tripathi, S.K.; Sahoo, R.K.; Biswal, B.K. Sox9 as an emerging target for anticancer drugs and a prognostic biomarker for cancer drug resistance. Drug Discov. Today 2022, 27, 2541–2550. [Google Scholar] [CrossRef]
- Carrasco-Garcia, E.; Lopez, L.; Moncho-Amor, V.; Carazo, F.; Aldaz, P.; Collado, M.; Bell, D.; Gaafar, A.; Karamitopoulou, E.; Tzankov, A.; et al. Sox9 triggers different epithelial to mesenchymal transition states to promote pancreatic cancer progression. Cancers 2022, 14, 916. [Google Scholar] [CrossRef]
- Lieber, M.; Mazzetta, J.; Nelson-Rees, W.; Kaplan, M.; Todaro, G. Establishment of a continuous tumor-cell line (panc-1) from a human carcinoma of the exocrine pancreas. Int. J. Cancer 1975, 15, 741–747. [Google Scholar] [CrossRef] [PubMed]
- Morgan, R.T.; Woods, L.K.; Moore, G.E.; Quinn, L.A.; McGavran, L.; Gordon, S.G. Human cell line (colo 357) of metastatic pancreatic adenocarcinoma. Int. J. Cancer 1980, 25, 591–598. [Google Scholar] [CrossRef] [PubMed]
- Sipos, B.; Moser, S.; Kalthoff, H.; Torok, V.; Lohr, M.; Kloppel, G. A comprehensive characterization of pancreatic ductal carcinoma cell lines: Towards the establishment of an in vitro research platform. Virchows Arch. 2003, 442, 444–452. [Google Scholar] [CrossRef]
- Collisson, E.A.; Sadanandam, A.; Olson, P.; Gibb, W.J.; Truitt, M.; Gu, S.; Cooc, J.; Weinkle, J.; Kim, G.E.; Jakkula, L.; et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat. Med. 2011, 17, 500–503. [Google Scholar] [CrossRef]
- Moore, P.S.; Sipos, B.; Orlandini, S.; Sorio, C.; Real, F.X.; Lemoine, N.R.; Gress, T.; Bassi, C.; Kloppel, G.; Kalthoff, H.; et al. Genetic profile of 22 pancreatic carcinoma cell lines. Analysis of k-ras, p53, p16 and dpc4/smad4. Virchows Arch. 2001, 439, 798–802. [Google Scholar] [CrossRef]
- Sirivatanauksorn, V.; Sirivatanauksorn, Y.; Gorman, P.A.; Davidson, J.M.; Sheer, D.; Moore, P.S.; Scarpa, A.; Edwards, P.A.; Lemoine, N.R. Non-random chromosomal rearrangements in pancreatic cancer cell lines identified by spectral karyotyping. Int. J. Cancer 2001, 91, 350–358. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdottir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The molecular signatures database (msigdb) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Grutzmann, R.; Boriss, H.; Ammerpohl, O.; Luttges, J.; Kalthoff, H.; Schackert, H.K.; Kloppel, G.; Saeger, H.D.; Pilarsky, C. Meta-analysis of microarray data on pancreatic cancer defines a set of commonly dysregulated genes. Oncogene 2005, 24, 5079–5088. [Google Scholar] [CrossRef]
- Wang, X.; Yao, S.; Luo, G.; Zhou, Y.; Fang, Q. Downregulation of rps14 inhibits the proliferation and metastasis of estrogen receptor-positive breast cancer cells. Anti-Cancer Drugs 2021, 32, 1019–1028. [Google Scholar] [CrossRef]
- Hu, S.; Cai, J.; Fang, H.; Chen, Z.; Zhang, J.; Cai, R. Rps14 promotes the development and progression of glioma via p53 signaling pathway. Exp. Cell Res. 2023, 423, 113451. [Google Scholar] [CrossRef]
- Zhou, X.; Hao, Q.; Liao, J.M.; Liao, P.; Lu, H. Ribosomal protein s14 negatively regulates c-myc activity. J. Biol. Chem. 2013, 288, 21793–21801. [Google Scholar] [CrossRef]
- Xu, Y.; Xu, D. Repair pathway choice for double-strand breaks. Essays Biochem. 2020, 64, 765–777. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Liu, Y.; Zheng, C.; Qu, H. Dbemt 2.0: An updated database for epithelial-mesenchymal transition genes with experimentally verified information and precalculated regulation information for cancer metastasis. J. Genet. Genom. 2019, 46, 595–597. [Google Scholar] [CrossRef] [PubMed]
- Aiello, N.M.; Maddipati, R.; Norgard, R.J.; Balli, D.; Li, J.; Yuan, S.; Yamazoe, T.; Black, T.; Sahmoud, A.; Furth, E.E.; et al. Emt subtype influences epithelial plasticity and mode of cell migration. Dev. Cell 2018, 45, 681–695.e4. [Google Scholar] [CrossRef]
- Lambert, S.A.; Jolma, A.; Campitelli, L.F.; Das, P.K.; Yin, Y.; Albu, M.; Chen, X.; Taipale, J.; Hughes, T.R.; Weirauch, M.T. The human transcription factors. Cell 2018, 172, 650–665. [Google Scholar] [CrossRef] [PubMed]
- Shih, H.P.; Seymour, P.A.; Patel, N.A.; Xie, R.; Wang, A.; Liu, P.P.; Yeo, G.W.; Magnuson, M.A.; Sander, M. A gene regulatory network cooperatively controlled by pdx1 and sox9 governs lineage allocation of foregut progenitor cells. Cell Rep. 2015, 13, 326–336. [Google Scholar] [CrossRef]
- Missiaglia, E.; Blaveri, E.; Terris, B.; Wang, Y.H.; Costello, E.; Neoptolemos, J.P.; Crnogorac-Jurcevic, T.; Lemoine, N.R. Analysis of gene expression in cancer cell lines identifies candidate markers for pancreatic tumorigenesis and metastasis. Int. J. Cancer 2004, 112, 100–112. [Google Scholar] [CrossRef]
- Terris, B.; Blaveri, E.; Crnogorac-Jurcevic, T.; Jones, M.; Missiaglia, E.; Ruszniewski, P.; Sauvanet, A.; Lemoine, N.R. Characterization of gene expression profiles in intraductal papillary-mucinous tumors of the pancreas. Am. J. Pathol. 2002, 160, 1745–1754. [Google Scholar] [CrossRef]
- Logsdon, C.D.; Simeone, D.M.; Binkley, C.; Arumugam, T.; Greenson, J.K.; Giordano, T.J.; Misek, D.E.; Kuick, R.; Hanash, S. Molecular profiling of pancreatic adenocarcinoma and chronic pancreatitis identifies multiple genes differentially regulated in pancreatic cancer. Cancer Res. 2003, 63, 2649–2657. [Google Scholar]
- Han, H.; Bearss, D.J.; Browne, L.W.; Calaluce, R.; Nagle, R.B.; Von Hoff, D.D. Identification of differentially expressed genes in pancreatic cancer cells using cdna microarray. Cancer Res. 2002, 62, 2890–2896. [Google Scholar] [PubMed]
- Ryu, B.; Jones, J.; Blades, N.J.; Parmigiani, G.; Hollingsworth, M.A.; Hruban, R.H.; Kern, S.E. Relationships and differentially expressed genes among pancreatic cancers examined by large-scale serial analysis of gene expression. Cancer Res. 2002, 62, 819–826. [Google Scholar]
- Tan, Z.J.; Hu, X.G.; Cao, G.S.; Tang, Y. Analysis of gene expression profile of pancreatic carcinoma using cdna microarray. World J. Gastroenterol. 2003, 9, 818–823. [Google Scholar] [CrossRef]
- Ramaswamy, S.; Ross, K.N.; Lander, E.S.; Golub, T.R. A molecular signature of metastasis in primary solid tumors. Nat. Genet. 2003, 33, 49–54. [Google Scholar] [CrossRef]
- Stratford, J.K.; Bentrem, D.J.; Anderson, J.M.; Fan, C.; Volmar, K.A.; Marron, J.S.; Routh, E.D.; Caskey, L.S.; Samuel, J.C.; Der, C.J.; et al. A six-gene signature predicts survival of patients with localized pancreatic ductal adenocarcinoma. PLoS Med. 2010, 7, e1000307. [Google Scholar] [CrossRef]
- Haider, S.; Wang, J.; Nagano, A.; Desai, A.; Arumugam, P.; Dumartin, L.; Fitzgibbon, J.; Hagemann, T.; Marshall, J.F.; Kocher, H.M.; et al. A multi-gene signature predicts outcome in patients with pancreatic ductal adenocarcinoma. Genome Med. 2014, 6, 105. [Google Scholar] [CrossRef] [PubMed]
- Kirby, M.K.; Ramaker, R.C.; Gertz, J.; Davis, N.S.; Johnston, B.E.; Oliver, P.G.; Sexton, K.C.; Greeno, E.W.; Christein, J.D.; Heslin, M.J.; et al. Rna sequencing of pancreatic adenocarcinoma tumors yields novel expression patterns associated with long-term survival and reveals a role for angptl4. Mol. Oncol. 2016, 10, 1169–1182. [Google Scholar] [CrossRef] [PubMed]
- Bianconi, D.; Heller, G.; Spies, D.; Herac, M.; Gleiss, A.; Liebmann-Reindl, S.; Unseld, M.; Kieler, M.; Scheithauer, W.; Streubel, B.; et al. Biochemical and genetic predictors of overall survival in patients with metastatic pancreatic cancer treated with capecitabine and nab-paclitaxel. Sci. Rep. 2017, 7, 4851. [Google Scholar] [CrossRef]
- Kim, J.; Jo, Y.H.; Jang, M.; Nguyen, N.N.Y.; Yun, H.R.; Ko, S.H.; Shin, Y.; Lee, J.S.; Kang, I.; Ha, J.; et al. Pac-5 gene expression signature for predicting prognosis of patients with pancreatic adenocarcinoma. Cancers 2019, 11, 1749. [Google Scholar] [CrossRef]
- Ma, Z.; Li, Z.; Ma, Z.; Zhou, Z.; Zhuang, H.; Liu, C.; Huang, B.; Zou, Y.; Zheng, Z.; Yang, L.; et al. Development of a kras-associated metabolic risk model for prognostic prediction in pancreatic cancer. BioMed Res. Int. 2021, 2021, 9949272. [Google Scholar] [CrossRef]
- Feng, Z.; Li, K.; Lou, J.; Wu, Y.; Peng, C. An emt-related gene signature for predicting response to adjuvant chemotherapy in pancreatic ductal adenocarcinoma. Front. Cell Dev. Biol. 2021, 9, 665161. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.S.; Jiang, M.; Wang, Y.J.; Xu, S.F.; Jiang, F.Y.; Han, Y.T.; Liu, Z.W.; Yu, H. Platelet-related gene risk score: A predictor for pancreatic cancer microenvironmental signature, chemosensitivity and prognosis. Am. J. Cancer Res. 2023, 13, 6113–6124. [Google Scholar] [PubMed]
- Choe, S.; Kwak, W.; Kim, E.; Shin, S.; Shin, M.; Koh, H.J.; Yoon, H. Promising therapeutic approach in pancreatic cancer: Metabolism-related genes. Front. Biosci. 2024, 29, 137. [Google Scholar] [CrossRef]
- Dong, B.; Zhang, Y.; Gao, H.; Liu, J.; Li, J. Machine learning developed a myc expression feature-based signature for predicting prognosis and chemoresistance in pancreatic adenocarcinoma. Biochem. Genet. 2024, 62, 4191–4214. [Google Scholar] [CrossRef]
- Donahue, T.R.; Tran, L.M.; Hill, R.; Li, Y.; Kovochich, A.; Calvopina, J.H.; Patel, S.G.; Wu, N.; Hindoyan, A.; Farrell, J.J.; et al. Integrative survival-based molecular profiling of human pancreatic cancer. Clin. Cancer Res. 2012, 18, 1352–1363. [Google Scholar] [CrossRef]
- Tiriac, H.; Belleau, P.; Engle, D.D.; Plenker, D.; Deschenes, A.; Somerville, T.D.D.; Froeling, F.E.M.; Burkhart, R.A.; Denroche, R.E.; Jang, G.H.; et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Discov. 2018, 8, 1112–1129. [Google Scholar] [CrossRef] [PubMed]
- Seth, S.; Li, C.Y.; Ho, I.L.; Corti, D.; Loponte, S.; Sapio, L.; Del Poggetto, E.; Yen, E.Y.; Robinson, F.S.; Peoples, M.; et al. Pre-existing functional heterogeneity of tumorigenic compartment as the origin of chemoresistance in pancreatic tumors. Cell Rep. 2019, 26, 1518–1532.e9. [Google Scholar] [CrossRef]
- Hou, Z.; Lin, J.; Ma, Y.; Fang, H.; Wu, Y.; Chen, Z.; Lin, X.; Lu, F.; Wen, S.; Yu, X.; et al. Single-cell rna sequencing revealed subclonal heterogeneity and gene signatures of gemcitabine sensitivity in pancreatic cancer. Front. Pharmacol. 2023, 14, 1193791. [Google Scholar] [CrossRef]
- Wei, X.; Zhou, X.; Zhao, Y.; He, Y.; Weng, Z.; Xu, C. A 14-gene gemcitabine resistance gene signature is significantly associated with the prognosis of pancreatic cancer patients. Sci. Rep. 2021, 11, 6087. [Google Scholar] [CrossRef]
- Marques, A.V.L.; Ruginsk, B.E.; Prado, L.O.; de Lima, D.E.; Daniel, I.W.; Moure, V.R.; Valdameri, G. The association of abc proteins with multidrug resistance in cancer. Biochim. Biophys. Acta Mol. Cell Res. 2025, 1872, 119878. [Google Scholar] [CrossRef]
- Cheng, Z.; Liang, X.; Zhang, C.; Wang, R.; Wei, T.; Ning, B.; Poreba, E.; Li, L.; Wang, H.; Ding, J. Sox9-transactived long non-coding rna neat1 promotes the self-renewal of liver cancer stem cells through pka/hippo signaling. Signal Transduct. Target. Ther. 2021, 6, 87. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, V.B.; Costa-Silva, P.F.; Araújo-Silva, H.; Miranda, G.; Correia-Pinto, J.; Moura, R.S. Molecular insights of hippo signaling in the chick developing lung. Biochim. Biophys. Acta Gene Regul. Mech. 2023, 1866, 194904. [Google Scholar] [CrossRef] [PubMed]
- Balachandran, S.; Narendran, A. The developmental origins of cancer: A review of the genes expressed in embryonic cells with implications for tumorigenesis. Genes 2023, 14, 604. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chen, X.; Zhu, L.; Lao, Z.; Zhou, T.; Zang, L.; Ge, W.; Jiang, M.; Xu, J.; Cao, Y.; et al. Sox9 is a critical regulator of tspan8-mediated metastasis in pancreatic cancer. Oncogene 2021, 40, 4884–4893. [Google Scholar] [CrossRef]
- Das, K.K.; Heeg, S.; Pitarresi, J.R.; Reichert, M.; Bakir, B.; Takano, S.; Kopp, J.L.; Wahl-Feuerstein, A.; Hicks, P.; Sander, M.; et al. Etv5 regulates ductal morphogenesis with sox9 and is critical for regeneration from pancreatitis. Dev. Dyn. 2018, 247, 854–866. [Google Scholar] [CrossRef]
- Kopp, J.L.; von Figura, G.; Mayes, E.; Liu, F.F.; Dubois, C.L.; Morris, J.P.t.; Pan, F.C.; Akiyama, H.; Wright, C.V.; Jensen, K.; et al. Identification of sox9-dependent acinar-to-ductal reprogramming as the principal mechanism for initiation of pancreatic ductal adenocarcinoma. Cancer Cell 2012, 22, 737–750. [Google Scholar] [CrossRef]
- Zhou, H.; Qin, Y.; Ji, S.; Ling, J.; Fu, J.; Zhuang, Z.; Fan, X.; Song, L.; Yu, X.; Chiao, P.J. Sox9 activity is induced by oncogenic kras to affect mdc1 and mcms expression in pancreatic cancer. Oncogene 2018, 37, 912–923. [Google Scholar] [CrossRef]
- Hong, Y.; Chen, W.; Du, X.; Ning, H.; Chen, H.; Shi, R.; Lin, S.; Xu, R.; Zhu, J.; Wu, S.; et al. Upregulation of sex-determining region y-box 9 (sox9) promotes cell proliferation and tumorigenicity in esophageal squamous cell carcinoma. Oncotarget 2015, 6, 31241–31254. [Google Scholar] [CrossRef]
- Boykevisch, S.; Zhao, C.; Sondermann, H.; Philippidou, P.; Halegoua, S.; Kuriyan, J.; Bar-Sagi, D. Regulation of ras signaling dynamics by sos-mediated positive feedback. Curr. Biol. 2006, 16, 2173–2179. [Google Scholar] [CrossRef]
- Jeng, H.H.; Taylor, L.J.; Bar-Sagi, D. Sos-mediated cross-activation of wild-type ras by oncogenic ras is essential for tumorigenesis. Nat. Commun. 2012, 3, 1168. [Google Scholar] [CrossRef]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G.; et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Yamaguchi, K. Epigenetic modifications and reprogramming in paternal pronucleus: Sperm, preimplantation embryo, and beyond. Cell. Mol. Life Sci. 2017, 74, 1957–1967. [Google Scholar] [CrossRef]
- Yang, P.; Wu, W.; Macfarlan, T.S. Maternal histone variants and their chaperones promote paternal genome activation and boost somatic cell reprogramming. BioEssays 2015, 37, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Sykes, S.M.; Mellert, H.S.; Holbert, M.A.; Li, K.; Marmorstein, R.; Lane, W.S.; McMahon, S.B. Acetylation of the p53 DNA-binding domain regulates apoptosis induction. Mol. Cell 2006, 24, 841–851. [Google Scholar] [CrossRef]
- Ren, H.Y.; Komatsu, N.; Shimizu, R.; Okada, K.; Miura, Y. Erythropoietin induces tyrosine phosphorylation and activation of phospholipase c-gamma 1 in a human erythropoietin-dependent cell line. J. Biol. Chem. 1994, 269, 19633–19638. [Google Scholar] [CrossRef] [PubMed]
- Oda, A.; Sawada, K.; Druker, B.J.; Ozaki, K.; Takano, H.; Koizumi, K.; Fukada, Y.; Handa, M.; Koike, T.; Ikeda, Y. Erythropoietin induces tyrosine phosphorylation of jak2, stat5a, and stat5b in primary cultured human erythroid precursors. Blood 1998, 92, 443–451. [Google Scholar] [CrossRef] [PubMed]
- Aldaz, P.; Otaegi-Ugartemendia, M.; Saenz-Antonanzas, A.; Garcia-Puga, M.; Moreno-Valladares, M.; Flores, J.M.; Gerovska, D.; Arauzo-Bravo, M.J.; Sampron, N.; Matheu, A.; et al. Sox9 promotes tumor progression through the axis bmi1-p21(cip). Sci. Rep. 2020, 10, 357. [Google Scholar] [CrossRef]
- Bogucka, K.; Marini, F.; Rosigkeit, S.; Schloeder, J.; Jonuleit, H.; David, K.; Schlackow, M.; Rajalingam, K. Erk3/mapk6 is required for kras-mediated nsclc tumorigenesis. Cancer Gene Ther. 2021, 28, 359–374. [Google Scholar] [CrossRef]
- Arrigo, A.P.; Gibert, B. Hspb1, hspb5 and hspb4 in human cancers: Potent oncogenic role of some of their client proteins. Cancers 2014, 6, 333–365. [Google Scholar] [CrossRef]
- Schafer, C.; Seeliger, H.; Bader, D.C.; Assmann, G.; Buchner, D.; Guo, Y.; Ziesch, A.; Palagyi, A.; Ochs, S.; Laubender, R.P.; et al. Heat shock protein 27 as a prognostic and predictive biomarker in pancreatic ductal adenocarcinoma. J. Cell. Mol. Med. 2012, 16, 1776–1791. [Google Scholar] [CrossRef]
- Day, E.; Poulogiannis, G.; McCaughan, F.; Mulholland, S.; Arends, M.J.; Ibrahim, A.E.; Dear, P.H. Irs2 is a candidate driver oncogene on 13q34 in colorectal cancer. Int. J. Exp. Pathol. 2013, 94, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Zheng, C.; Chen, S.; He, Z.; Hua, H.; Sun, C.; Yu, C. Foxq1 promotes pancreatic cancer cell proliferation, tumor stemness, invasion and metastasis through regulation of ldha-mediated aerobic glycolysis. Cell Death Dis. 2023, 14, 699. [Google Scholar] [CrossRef]
- Bannoura, S.F.; Khan, H.Y.; Azmi, A.S. Kras g12d targeted therapies for pancreatic cancer: Has the fortress been conquered? Front. Oncol. 2022, 12, 1013902. [Google Scholar] [CrossRef] [PubMed]
- Edelman, H.E.; McClymont, S.A.; Tucker, T.R.; Pineda, S.; Beer, R.L.; McCallion, A.S.; Parsons, M.J. Sox9 modulates cancer biomarker and cilia genes in pancreatic cancer. Hum. Mol. Genet. 2021, 30, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Ala, M. Target c-myc to treat pancreatic cancer. Cancer Biol. Ther. 2022, 23, 34–50. [Google Scholar] [CrossRef]
- Han, L.; Huo, Y.; Huang, L.; Zheng, Y.; Yu, X.; Zhang, N.; Yang, M. Genome-wide functional integration identified maz-controlled rps14 dysregulation in hepatocellular carcinoma. Arch. Toxicol. 2024, 98, 985–997. [Google Scholar] [CrossRef]
- Vivanco, I.; Palaskas, N.; Tran, C.; Finn, S.P.; Getz, G.; Kennedy, N.J.; Jiao, J.; Rose, J.; Xie, W.; Loda, M.; et al. Identification of the jnk signaling pathway as a functional target of the tumor suppressor pten. Cancer Cell 2007, 11, 555–569. [Google Scholar] [CrossRef]
- Bracken, A.P.; Dietrich, N.; Pasini, D.; Hansen, K.H.; Helin, K. Genome-wide mapping of polycomb target genes unravels their roles in cell fate transitions. Genes Dev. 2006, 20, 1123–1136. [Google Scholar] [CrossRef]
- Mozaffari, N.L.; Pagliarulo, F.; Sartori, A.A. Human ctip: A ‘double agent’ in DNA repair and tumorigenesis. Semin. Cell Dev. Biol. 2021, 113, 47–56. [Google Scholar] [CrossRef]
- Furuta, S.; Wang, J.M.; Wei, S.; Jeng, Y.M.; Jiang, X.; Gu, B.; Chen, P.L.; Lee, E.Y.; Lee, W.H. Removal of brca1/ctip/zbrk1 repressor complex on ang1 promoter leads to accelerated mammary tumor growth contributed by prominent vasculature. Cancer Cell 2006, 10, 13–24. [Google Scholar] [CrossRef]
- Yu, Y.; Chen, L.; Zhao, G.; Li, H.; Guo, Q.; Zhu, S.; Li, P.; Min, L.; Zhang, S. Rbbp8/ctip suppresses p21 expression by interacting with ctbp and brca1 in gastric cancer. Oncogene 2020, 39, 1273–1289. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.; Yan, Y.; Huang, Q.; Zheng, D. MDMX in cancer: A partner of p53 and a p53-independent effector. Biol. Targets Ther. 2024, 18, 61–78. [Google Scholar] [CrossRef]
- Huang, Y.; Che, X.; Wang, P.W.; Qu, X. P53/mdm2 signaling pathway in aging, senescence and tumorigenesis. Semin. Cancer Biol. 2024, 101, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Farhadi, E.; Khomeijani-Farahani, M.; Nikbakhsh, R.; Azizan, A.; Soltani, S.; Barekati, H.; Mahmoudi, M. The potential role of circular rnas in regulating p53 in different types of cancers. Pathol. Res. Pract. 2024, 261, 155488. [Google Scholar] [CrossRef]
- Zheltukhin, A.O.; Chumakov, P.M. Constitutive and induced functions of the p53 gene. Biochemistry 2010, 75, 1692–1721. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Z.; Huang, L.; Wu, C.; Zhang, B. Mir-29b suppresses proliferation and mobility by targeting sox12 and dnmt3b in pancreatic cancer. Anti-Cancer Drugs 2019, 30, 281–288. [Google Scholar] [CrossRef]
- Ge, J.N.; Yan, D.; Ge, C.L.; Wei, M.J. Lncrna c9orf139 can regulate the growth of pancreatic cancer by mediating the mir-663a/sox12 axis. World J. Gastrointest. Oncol. 2020, 12, 1272–1287. [Google Scholar] [CrossRef]
- Yu, D.; Zhao, Z.; Wang, L.; Qiao, S.; Yang, Z.; Wen, Q.; Zhu, G. Sox21-as1 activated by stat6 promotes pancreatic cancer progression via up-regulation of sox21. J. Transl. Med. 2022, 20, 511. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Liu, W.; Zhang, H.M.; Xie, G.Y.; Miao, Y.R.; Xia, M.; Guo, A.Y. Htftarget: A comprehensive database for regulations of human transcription factors and their targets. Genom. Proteom. Bioinform. 2020, 18, 120–128. [Google Scholar] [CrossRef]
- Li, P.; Hu, Y.; Yi, J.; Li, J.; Yang, J.; Wang, J. Identification of potential biomarkers to differentially diagnose solid pseudopapillary tumors and pancreatic malignancies via a gene regulatory network. J. Transl. Med. 2015, 13, 361. [Google Scholar] [CrossRef]
- Shroff, S.; Rashid, A.; Wang, H.; Katz, M.H.; Abbruzzese, J.L.; Fleming, J.B.; Wang, H. Sox9: A useful marker for pancreatic ductal lineage of pancreatic neoplasms. Hum. Pathol. 2014, 45, 456–463. [Google Scholar] [CrossRef] [PubMed]
- Christgen, M.; Schniewind, B.; Jueschke, A.; Ungefroren, H.; Kalthoff, H. Gemcitabine-mediated apoptosis is associated with increased cd95 surface expression but is not inhibited by dn-fadd in colo357 pancreatic cancer cells. Cancer Lett. 2005, 227, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Schniewind, B.; Christgen, M.; Kurdow, R.; Haye, S.; Kremer, B.; Kalthoff, H.; Ungefroren, H. Resistance of pancreatic cancer to gemcitabine treatment is dependent on mitochondria-mediated apoptosis. Int. J. Cancer 2004, 109, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Martinez, M.; Fendley, G.A.; Saxberg, A.D.; Zoghbi, M.E. Stimulation of the human mitochondrial transporter abcb10 by zinc-mesoporphrin. PLoS ONE 2020, 15, e0238754. [Google Scholar] [CrossRef]
- Seguin, A.; Takahashi-Makise, N.; Yien, Y.Y.; Huston, N.C.; Whitman, J.C.; Musso, G.; Wallace, J.A.; Bradley, T.; Bergonia, H.A.; Kafina, M.D.; et al. Reductions in the mitochondrial abc transporter abcb10 affect the transcriptional profile of heme biosynthesis genes. J. Biol. Chem. 2017, 292, 16284–16299. [Google Scholar] [CrossRef]
- Oiso, S.; Takayama, Y.; Nakazaki, R.; Matsunaga, N.; Motooka, C.; Yamamura, A.; Ikeda, R.; Nakamura, K.; Takeda, Y.; Kariyazono, H. Factors involved in the cisplatin resistance of kcp-4 human epidermoid carcinoma cells. Oncol. Rep. 2014, 31, 719–726. [Google Scholar] [CrossRef]
- Yang, W.; Gong, P.; Yang, Y.; Yang, C.; Yang, B.; Ren, L. Circ-abcb10 contributes to paclitaxel resistance in breast cancer through let-7a-5p/dusp7 axis. Cancer Manag. Res. 2020, 12, 2327–2337. [Google Scholar] [CrossRef]
- Wu, Z.; Gong, Q.; Yu, Y.; Zhu, J.; Li, W. Knockdown of circ-abcb10 promotes sensitivity of lung cancer cells to cisplatin via mir-556-3p/ak4 axis. BMC Pulm. Med. 2020, 20, 10. [Google Scholar] [CrossRef]
- Skinner, K.T.; Palkar, A.M.; Hong, A.L. Genetics of abcb1 in cancer. Cancers 2023, 15, 4236. [Google Scholar] [CrossRef]
- Kopantzev, E.P.; Kopantseva, M.R.; Grankina, E.V.; Mikaelyan, A.; Egorov, V.I.; Sverdlov, E.D. Activation of igf/igf-ir signaling pathway fails to induce epithelial-mesenchymal transition in pancreatic cancer cells. Pancreatology 2019, 19, 390–396. [Google Scholar] [CrossRef]
- Ewels, P.; Magnusson, M.; Lundin, S.; Kaller, M. Multiqc: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef] [PubMed]
- Galaxy, C. The galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2022 update. Nucleic Acids Res. 2022, 50, W345–W351. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. Hisat: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. Featurecounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed]
- Batut, B.; Hiltemann, S.; Bagnacani, A.; Baker, D.; Bhardwaj, V.; Blank, C.; Bretaudeau, A.; Brillet-Gueguen, L.; Cech, M.; Chilton, J.; et al. Community-driven data analysis training for biology. Cell Syst. 2018, 6, 752–758.e1. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for rna-seq data with deseq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The gene ontology consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Gene Ontology, C.; Aleksander, S.A.; Balhoff, J.; Carbon, S.; Cherry, J.M.; Drabkin, H.J.; Ebert, D.; Feuermann, M.; Gaudet, P.; Harris, N.L.; et al. The gene ontology knowledgebase in 2023. Genetics 2023, 224, iyad031. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using david bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Kolberg, L.; Raudvere, U.; Kuzmin, I.; Adler, P.; Vilo, J.; Peterson, H. G:Profiler—Interoperable web service for functional enrichment analysis and gene identifier mapping (2023 update). Nucleic Acids Res. 2023, 51, W207–W212. [Google Scholar] [CrossRef]
- Koopmans, F.; van Nierop, P.; Andres-Alonso, M.; Byrnes, A.; Cijsouw, T.; Coba, M.P.; Cornelisse, L.N.; Farrell, R.J.; Goldschmidt, H.L.; Howrigan, D.P.; et al. Syngo: An evidence-based, expert-curated knowledge base for the synapse. Neuron 2019, 103, 217–234.e4. [Google Scholar] [CrossRef] [PubMed]
- Seal, R.L.; Braschi, B.; Gray, K.; Jones, T.E.M.; Tweedie, S.; Haim-Vilmovsky, L.; A Bruford, E. Genenames.Org: The hgnc resources in 2023. Nucleic Acids Res. 2023, 51, D1003–D1009. [Google Scholar] [CrossRef] [PubMed]
- Babicki, S.; Arndt, D.; Marcu, A.; Liang, Y.; Grant, J.R.; Maciejewski, A.; Wishart, D.S. Heatmapper: Web-enabled heat mapping for all. Nucleic Acids Res. 2016, 44, W147–W153. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.D.; Ebert, D.; Muruganujan, A.; Mushayahama, T.; Albou, L.; Mi, H. Panther: Making genome-scale phylogenetics accessible to all. Protein Sci. 2022, 31, 8–22. [Google Scholar] [CrossRef]
- Gillespie, M.; Jassal, B.; Stephan, R.; Milacic, M.; Rothfels, K.; Senff-Ribeiro, A.; Griss, J.; Sevilla, C.; Matthews, L.; Gong, C.; et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 2022, 50, D687–D692. [Google Scholar] [CrossRef]
- Kondratyeva, L.; Kashkin, K.; Kopantzev, E.; Chernov, I. Effects of SOX9 Downregulation on the Transcriptomic Landscape of Pancreatic Ductal Adenocarcinoma Cells. Available online: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE287340 (accessed on 17 January 2025).
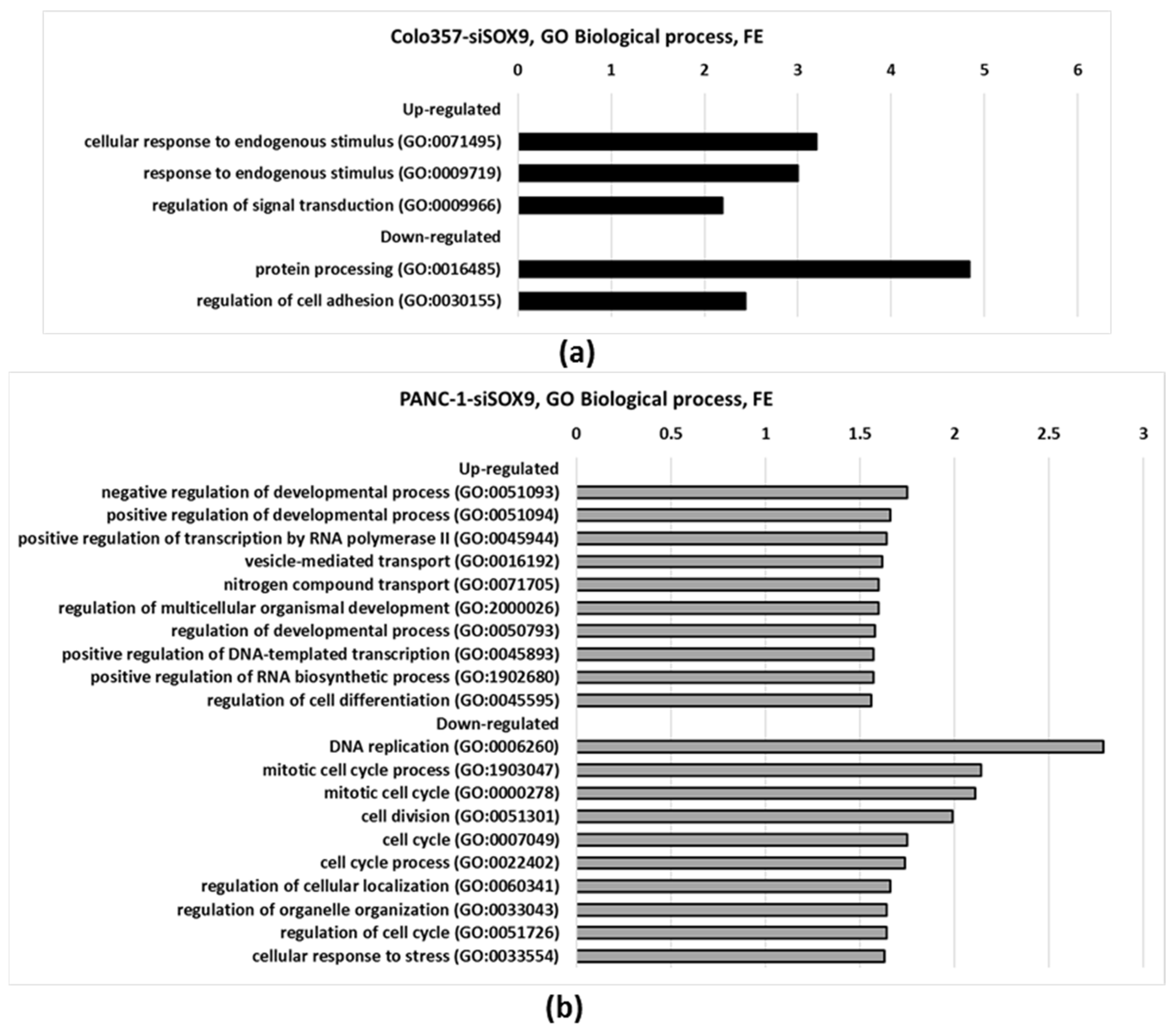
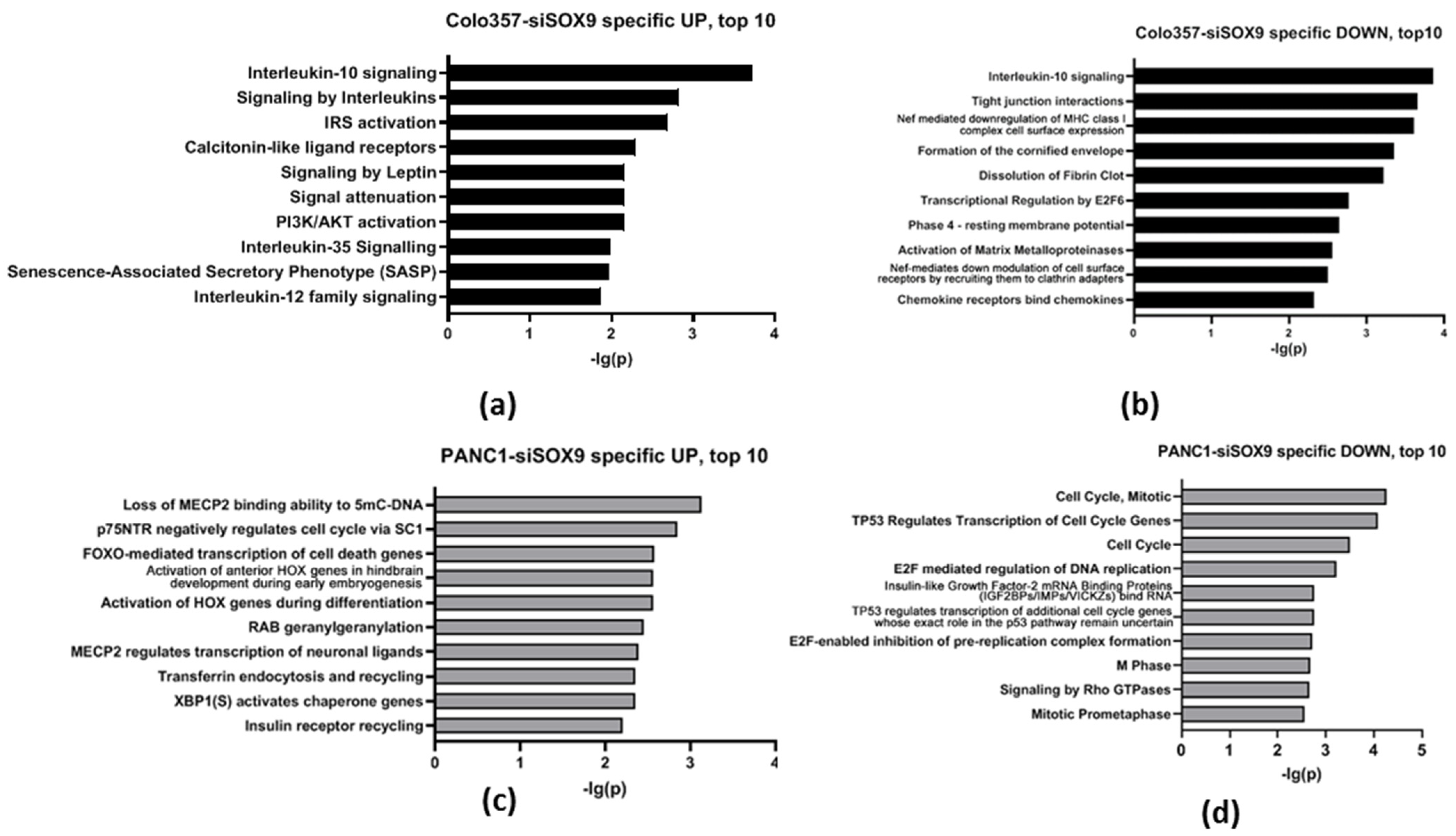
| Gene Ontology | Up-Regulated Genes | Down-Regulated Genes |
|---|---|---|
| Biological process | Regulation of signal transduction (GO:0009966) | N.i. 1 |
| Molecular function | N.i. | Protein binding (GO:0005515) |
| Cellular components | N.i. | Endomembrane system (GO:0012505) |
| Panther pathway | Gonadotropin-releasing hormone receptor pathway (P06664) | cytoplasm (GO:0005737). |
| Pathway | COLO357–siSOX9 Genes | PANC-1–siSOX9 Genes |
|---|---|---|
| Common pathways with up-regulated genes | ||
| R-HSA-112412 SOS-mediated signaling | IRS1, IRS2 | IRS1, IRS2, SOS1 |
| R-HSA-2892245 POU5F1 (OCT4), SOX2, NANOG repress genes related to differentiation | TSC22D1, DKK1 | CDX2, DKK1, EOMES, TSC22D1 |
| R-HSA-9821993 replacement of protamines by nucleosomes in the male pronucleus | H2BC12, MYL9 | H2BC8, H2BC12, H2BC21, HDAC1, H3-3A, MYL9, THBS1, TNRC6A |
| R-HSA-171306 packaging of telomere ends R-HSA-73728 RNA polymerase I promoter opening R-HSA-5334118 DNA methylation | H2BC12 | DNMT3B, H2BC12, H2BC8, H2BC21, H3-3A |
| Common pathways with down-regulated genes | ||
| R-HSA-69206 G1/S transition R-HSA-453279 mitotic G1 phase and G1/S transition | CDC6, CKS1B, PSMB10, PSME3, PPP2R1B, RBBP4, RRM2 | CABLES1, CCN1, CCNB1, CDC6, CDK1, CDK6, CKS1B, MCM8, ORC1, ORC6, POLA2, POLE2, PPP2R1B, PRIM1, PRIM2, PSMA4, PSMD1, PSME3, RBBP4, RBL2, SKP2 |
| R-HSA-111464 SMAC(DIABLO)-mediated dissociation of IAP:caspase complexes R-HSA-111463 SMAC (DIABLO) binds to IAPs R-HSA-111469 SMAC, XIAP-regulated apoptotic response R-HSA-111459 activation of caspases through apoptosome-mediated cleavage | APAF1, CASP7 | CASP7, DCP2, XIAP |
| Pathway | COLO357–siSOX9 Genes (Up-Regulated) | PANC-1–siSOX9 Genes (Down-Regulated) |
|---|---|---|
| R-HSA-2559582 senescence-associated secretory phenotype (SASP) | H2BC12, IL6 | CCN1, CDC26, CDC27, CDK6, CXCL8, FOS, H2AC11, H2AJ, H2AX, H2BC11, JUN, RPS6KA3, UBE2C |
| R-HSA-2559583 cellular senescence | H2BC12, IL6, MAP2K6 | AGO4, CBX6, CBX8, CCN1, CDC26, CDC27, CDK6, CXCL8, ETS1, ETS2, FOS, H2AC11, H2AJ, H2AX, H2BC11, HMGA1, JUN, MAPK14, MAPK9, MDM2, MSN, POT1, RBBP4, RPS6KA3, SUZ12, TNRC6B, UBE2C |
| Pathway | COLO357–siSOX9 Genes (Up-Regulated) | COLO357-siSOX9 Genes (Down–Regulated) |
|---|---|---|
| R-HSA-6783783 interleukin-10 signaling | CCL20, IL12A, IL6 | CXCL1, CXCL2, CXCL3, IL1A, IL1R2, TNFRSF1B |
| R-HSA-449147 signaling by interleukins | CCL20, DUSP6, FYN, IL12A, IL6, IRS1, IRS2, LCP1, MAP2K6, ZEB1 | CRK, CXCL1, CXCL2, CXCL3, FSCN1, IL1A, IL1R2, IL1RAP, LCN2, LGALS9, MUC1, PITPNA, PPP2R1B, PSMB10, PSME3, SERPINB2, TNFRSF1B |
| Hallmark Name | SIZE | ES | NES | NOM p-Val | FDR q-Val | FWER p-Val |
|---|---|---|---|---|---|---|
| COLO357–siSOX9 up-regulated gene sets | ||||||
| MYC_UP.V1_UP | 30 | 0.42 | 2.24 | 0.0040 | 0.013 | 0.02 |
| COLO357–siSOX9 down-regulated gene sets | ||||||
| LEF1_UP.V1_DN | 41 | −0.55 | −2.45 | 0 | 0 | 0 |
| RPS14_DN.V1_UP | 30 | −0.54 | −2.23 | 0 | 0.0050 | 0.009 |
| PANC-1–siSOX9 up-regulated gene sets | ||||||
| PTEN_DN.V1_UP | 28 | 0.49 | 2.11 | 0.0035 | 0.013 | 0.012 |
| TGFB_UP.V1_DN | 65 | 0.37 | 2.03 | 0 | 0.014 | 0.026 |
| PANC-1–siSOX9 down-regulated gene sets | ||||||
| PRC2_EZH2_UP.V1_UP | 62 | −0.53 | −2.36 | 0 | 0 | 0 |
| CTIP_DN.V1_DN | 25 | −0.60 | −2.14 | 0 | 0.0024 | 0.005 |
| Process | COLO357–siSOX9 | PANC-1–siSOX9 |
|---|---|---|
| R-HSA-6804754 regulation of TP53 expression (up) | N.i. 2 | Supplementary Tables S6 and S16 |
| R-HSA-6804116 TP53 regulates transcription of genes involved in G1 cell cycle arrest (down) | Supplementary Tables S16 and S17 | Supplementary Tables S6, S16 and S17 |
| R-HSA-6791312 TP53 regulates transcription of cell cycle genes (down) | Supplementary Tables S16 and S17 | Supplementary Tables S6, S16 and S17 |
| R-HSA-6804115 TP53 regulates transcription of additional cell cycle genes whose exact role in the p53 pathway remain uncertain (down) | N.i. | Supplementary Table S6 |
| R-HSA-6803211 TP53 regulates transcription of death receptors and ligands (down) | Supplementary Table S6 | N.s. 3 |
| R-HSA-5633008 TP53 regulates transcription of cell death genes | Supplementary Tables S8 and S9 (Down) | N.s. |
| R-HSA-6804758 regulation of TP53 activity through acetylation (up) | Supplementary Tables S8 and S9 | Supplementary Tables S8 and S10 |
| R-HSA-3700989 transcriptional regulation by TP53 (up) | N.i. | Supplementary Tables S10 and S16 |
| R-HSA-6804114 TP53 regulates transcription of genes involved in G2 cell cycle arrest (up) | N.s. | Supplementary Table S16 |
| R-HSA-6804759 regulation of TP53 activity through association with co-factors (up) | N.s. | Supplementary Table S16 |
| R-HSA-5633007 regulation of TP53 activity (up) | Supplementary Table S17 | Supplementary Table S17 |
| R-HSA-6804757 regulation of TP53 degradation (up) | Supplementary Table S17 | Supplementary Table S17 |
| R-HSA-6806003 regulation of TP53 expression and degradation (up) | Supplementary Table S17 | Supplementary Table S17 |
| Transcription Factor | Pathways 1 |
|---|---|
| siSOX9 up-regulated TF’s | |
| AHR | Aryl hydrocarbon receptor signaling (R-HSA-8937144); Endogenous sterols (R-HSA-211976); Xenobiotics (R-HSA-211981) |
| TSC22D1 | POU5F1 (OCT4), SOX2, NANOG repress genes related to differentiation (R-HSA-2892245); Transcriptional regulation of pluripotent stem cells (R-HSA-452723) |
| siSOX9 down-regulated TF’s | |
| ARNTL2 | BMAL1:CLOCK,NPAS2 activates circadian gene expression (R-HSA-1368108) |
| E2F7 | TP53 regulates transcription of genes involved in G1 cell cycle arrest (R-HSA-6804116) TP53 regulates transcription of cell cycle genes (R-HSA-6791312) Transcriptional regulation by TP53 (R-HSA-3700989) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kashkin, K.; Kondratyeva, L.; Kopantzev, E.; Abramov, I.; Zhukova, L.; Chernov, I. Deciphering of SOX9 Functions in Pancreatic Cancer Cells. Int. J. Mol. Sci. 2025, 26, 2652. https://doi.org/10.3390/ijms26062652
Kashkin K, Kondratyeva L, Kopantzev E, Abramov I, Zhukova L, Chernov I. Deciphering of SOX9 Functions in Pancreatic Cancer Cells. International Journal of Molecular Sciences. 2025; 26(6):2652. https://doi.org/10.3390/ijms26062652
Chicago/Turabian StyleKashkin, Kirill, Liya Kondratyeva, Eugene Kopantzev, Ivan Abramov, Lyudmila Zhukova, and Igor Chernov. 2025. "Deciphering of SOX9 Functions in Pancreatic Cancer Cells" International Journal of Molecular Sciences 26, no. 6: 2652. https://doi.org/10.3390/ijms26062652
APA StyleKashkin, K., Kondratyeva, L., Kopantzev, E., Abramov, I., Zhukova, L., & Chernov, I. (2025). Deciphering of SOX9 Functions in Pancreatic Cancer Cells. International Journal of Molecular Sciences, 26(6), 2652. https://doi.org/10.3390/ijms26062652






