The Future of DNA Barcoding: Reflections from Early Career Researchers
Abstract
:1. Introduction
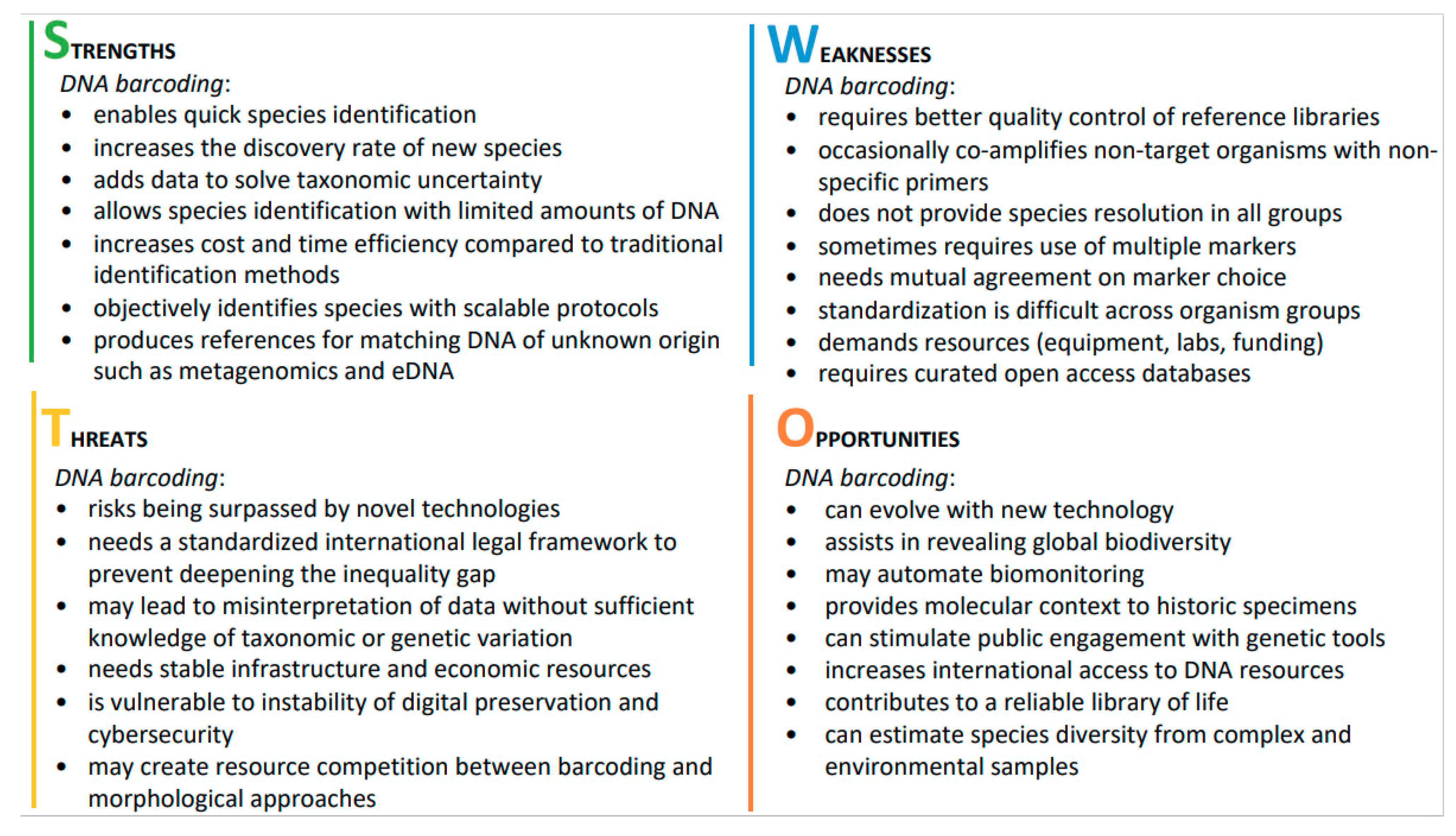
2. SWOT Analysis and Early Career Opinions
2.1. DNA Barcoding Offers Efficient, Affordable, and High-Throughput Solutions
2.2. DNA Barcoding will Survive and Thrive with Accessible and Curated Reference Libraries
2.3. DNA Barcoding Enhances Biodiversity Discovery and Monitoring
2.4. DNA Barcoding Methodology Is the Foundation for Automation and Accelerated Biodiversity Assessments
2.5. DNA Barcoding for Everyone, Everywhere
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; deWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. Ser. B 2003, 270, 313–321. [Google Scholar] [CrossRef] [Green Version]
- Bush, A.; Monk, A.W.; Compson, G.Z.; Peters, D.L.; Porter, M.T.; Shokralla, S.; Wright, G.T.M.; Hajibabaei, M.; Baird, J.D. DNA metabarcoding reveals metacommunity dynamics in a threatened boreal wetland wilderness. Proc. Natl. Acad. Sci. USA 2020, 117, 8539–8545. [Google Scholar] [CrossRef] [Green Version]
- Pennisi, E. DNA barcodes jump-start search for new species. Science 2019, 364, 920–921. [Google Scholar] [CrossRef]
- DeSalle, R.; Goldstein, P. Review and interpretation of trends in DNA barcoding. Front. Ecol. Evol. 2019, 7, 302. [Google Scholar] [CrossRef] [Green Version]
- Taberlet, P.; Bonin, A.; Zinger, L.; Coissac, E. Environmental DNA: For Biodiversity Research and Monitoring; Oxford University Press: Oxford, UK, 2018. [Google Scholar]
- Galimberti, A.; Casiraghi, M.; Bruni, I.; Guzzetti, L.; Cortis, P.; Berterame, N.M.; Labra, M. From DNA barcoding to personalized nutrition: The evolution of food traceability. Curr. Opin. Food Sci. 2019, 28, 41–48. [Google Scholar] [CrossRef]
- De Boer, H.J.; Ichim, M.C.; Newmaster, S. DNA barcoding and pharmacovigilance of herbal medicines. Drug Saf. 2015, 38, 611–620. [Google Scholar] [CrossRef] [PubMed]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar] [CrossRef] [Green Version]
- Hobern, D. BIOSCAN: DNA barcoding to accelerate taxonomy and biogeography for conservation and sustainability. Genome 2021, 64, 161–164. [Google Scholar] [CrossRef] [PubMed]
- Hobern, D.; Hebert, P.D.N. BIOSCAN-revealing eukaryote diversity, dynamics, and interactions. Biodivers. Inf. Sci. Stand. 2019, 3, e37333. [Google Scholar] [CrossRef] [Green Version]
- Hebert, P.D.N.; Braukmann, T.W.; Prosser, S.W.; Ratnasingham, S.; DeWaard, J.R.; Ivanova, N.V.; Janzen, H.D.; Hallwachs, W.; Sones, E.J.; Zakharov, E. A Sequel to Sanger: Amplicon sequencing that scales. BMC Genom. 2018, 19, 219. [Google Scholar] [CrossRef] [Green Version]
- Kress, W.J. Plant DNA barcodes: Applications today and in the future. J. Syst. Evol. 2017, 55, 291–307. [Google Scholar] [CrossRef] [Green Version]
- Govender, A.; Willows-Munro, S. The utility of DNA barcoding as a tool to assess the success of ecological restoration using Hemiptera as a biological indicator. Restor. Ecol. 2019, 27, 1409–1419. [Google Scholar] [CrossRef]
- Taberlet, P.; Coissac, E.; Pompanon, F.; Brochmann, C.; Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 2012, 21, 2045–2050. [Google Scholar] [CrossRef]
- Beng, K.C.; Corlett, R.T. Applications of environmental DNA (eDNA) in ecology and conservation: Opportunities, challenges and prospects. Biodivers. Conserv. 2020, 29, 2089–2121. [Google Scholar] [CrossRef]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, N.R.; Anokhin, P.A.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef]
- Berry, O.; Bulman, C.; Bunce, M.; Coghlan, M.; Murray, D.C.; Ward, R.D. Comparison of morphological and DNA metabarcoding analyses of diets in exploited marine fishes. Mar. Ecol. Prog. Ser. 2015, 540, 167–181. [Google Scholar] [CrossRef] [Green Version]
- Deiner, K.; Bik, H.M.; Machler, E.; Seymour, M.; Lacoursiere-Roussel, A.; Altermatt, F.; Creer, S.; Bista, I.; Lodge, M.D.; de Vere, N.; et al. Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Mol. Ecol. 2017, 26, 5872–5895. [Google Scholar] [CrossRef] [PubMed]
- Alsos, I.G.; Lammers, Y.; Yoccoz, N.G.; Jørgensen, T.; Sjögren, P.; Gielly, L.; Edwards, M.E. Plant DNA metabarcoding of lake sediments: How does it represent the contemporary vegetation. PLoS ONE 2018, 13, e0195403. [Google Scholar] [CrossRef]
- Creer, S.; Deiner, K.; Frey, S.; Porazinska, D.; Taberlet, P.; Thomas, W.K.; Potter, C.; Bik, H.M. The ecologist’s field guide to sequenceh-based identification of biodiversity. Methods Ecol. Evol. 2016, 7, 1008–1018. [Google Scholar] [CrossRef]
- Srivathsan, A.; Lee, L.; Katoh, K.; Hartop, E.; Kutty, S.N.; Wong, J.; Yeo, D.; Meier, R. MinION barcodes: Biodiversity discovery and identification by everyone, for everyone. bioRxiv 2021, 434692. [Google Scholar] [CrossRef]
- Janzen, D.H.; Hallwachs, W.; Pereira, G.; Blanco, R.; Masis, A.; Chavarria, M.M.; Chavarria, F.; Guadamuz, A.; Araya, M.; Smith, M.A.; et al. Using DNA-barcoded Malaise trap samples to measure impact of a geothermal energy project on the biodiversity of a Costa Rican old-growth rainforest. Genome 2020, 63, 407–436. [Google Scholar] [CrossRef]
- Joly, S.; Davies, T.J.; Archambault, A.; Bruneau, A.; Derry, A.; Kembel, S.W.; Peres-Neto, P.; Vamosi, J.; Wheeler, T.A. Ecology in the age of DNA barcoding: The resource, the promise and the challenges ahead. Mol. Ecol. Resour. 2014, 14, 221–232. [Google Scholar] [CrossRef] [PubMed]
- Obringer, R.; Rachunok, B.; Maia-Silva, D.; Arbabzadeh, M.; Nateghi, R.; Madani, K. The overlooked environmental footprint of increasing Internet use. Resour. Conserv. Recycl. 2021, 167, 105389. [Google Scholar] [CrossRef]
- Hollingsworth, P.M.; Graham, S.W.; Little, D.P. Choosing and using a plant DNA barcode. PLoS ONE 2011, 6, e19254. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, Y.; Henry, R.J.; Rossetto, M.; Wang, Y.; Chen, S. Plant DNA barcoding: From gene to genome. Biol. Rev. 2015, 90, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.-F.; Ma, H.; Ci, X.-Q.; Li, L.; Song, Y.; Liu, B.; Li, H.W.; Wang, S.L.; Qu, X.J.; Hu, J.L.; et al. Can plastid genome sequencing be used for species identification in Lauraceae? Bot. J. Linn. Soc. 2021, boab018. [Google Scholar] [CrossRef]
- Hollingsworth, P.M.; Li, D.-Z. van der Bank, M. Twyford, A.D. Telling plant species apart with DNA: From barcodes to genomes. Philos. Trans. R. Soc. B 2016, 371, 20150338. [Google Scholar]
- Fontes, J.T.; Vieira, P.E.; Ekrem, T.; Soares, P.; Costa, F.O. BAGS: An automated Barcode, Audit & Grade System for DNA barcode reference libraries. Mol. Ecol. Resour. 2021, 21, 573–583. [Google Scholar] [PubMed]
- Siddall, M.E.; Fontanella, F.M.; Watson, S.C.; Kvist, S.; Erséus, C. Barcoding bamboozled by bacteria: Convergence to metazoan mitochondrial primer targets by marine microbes. Syst. Biol. 2009, 58, 445–451. [Google Scholar] [CrossRef] [Green Version]
- Ratnasingham, S.; Hebert, P.D.N. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE 2013, 8, e66213. [Google Scholar] [CrossRef] [Green Version]
- Weigand, H.; Beermann, A.J.; Čiampor, F.; Costa, F.O.; Csabai, Z.; Duarte, S.; Geiger M., F.; Grabowski, M.; Rimet, F.; Rulik, B.; et al. DNA barcode reference libraries for the monitoring of aquatic biota in Europe: Gap-analysis and recommendations for future work. Sci. Total Environ. 2019, 678, 499–524. [Google Scholar] [CrossRef] [PubMed]
- Weigand, A.M.; Jochum, A.; Pfenninger, M.; Steinke, D.; Klussmann-Kolb, A. A new approach to an old conundrum—DNA barcoding sheds new light on phenotypic plasticity and morphological stasis in microsnails (Gastropoda, Pulmonata, Carychiidae). Mol. Ecol. Resour. 2011, 11, 255–265. [Google Scholar] [CrossRef]
- Pentinsaari, M.; Ratnasingham, S.; Miller, S.E.; Hebert, P.D.N. BOLD and GenBank revisited—Do identification errors arise in the lab or in the sequence libraries? PLoS ONE 2020, 15, e0231814. [Google Scholar] [CrossRef] [PubMed]
- Rimet, F.; Aylagas, E.; Borja, A.; Bouchez, A.; Canino, A.; Chauvin, C.; Chonova, T.; Ciampor, F., Jr.; Costa, F.О.; Ferrari, B.J.D.; et al. Metadata standards and practical guidelines for specimen and DNA curation when building barcode reference libraries for aquatic life. Metabarcoding Metagenom. 2021, 5, e58056. [Google Scholar]
- Porter, T.M.; Hajibabaei, M. Over 2.5 million COI sequences in GenBank and growing. PLoS ONE 2018, 13, e0200177. [Google Scholar] [CrossRef] [Green Version]
- Krachunov, M.; Nisheva, M.; Vassilev, D. Machine learning models for error detection in metagenomics and polyploid sequencing data. Information 2019, 10, 110. [Google Scholar] [CrossRef] [Green Version]
- Sharkey, M.J.; Janzen, D.H.; Hallwachs, W.; Chapman, E.G.; Smith, M.A.; Dapkey, T.; Brown, A.; Ratnasingham, S.; Naik, S.; Manjunath, R.; et al. Minimalist revision and description of 403 new species in 11 subfamilies of Costa Rican braconid parasitoid wasps, including host records for 219 species. ZooKeys 2021, 1013, 1–666. [Google Scholar] [CrossRef]
- Montes-Ortiz, L.; Elías-Gutiérrez, M. Water mite diversity (Acariformes: Prostigmata: Parasitengonina: Hydrachnidiae) from karst ecosystems in southern of Mexico: A barcoding approach. Diversity 2020, 12, 329. [Google Scholar] [CrossRef]
- Buenaventura, E.; Valverde-Castro, C.; Wolff, M.; Triana-Chavez, O.; Gómez-Palacio, A. DNA barcoding for identifying synanthropic flesh flies (Diptera, Sarcophagidae) of Colombia. Acta Tropica 2018, 182, 291–297. [Google Scholar] [CrossRef]
- Ćelepirović, N.; Novak Agbaba, S.; Karija Vlahović, M. DNA Barcoding of Fungi in the Forest Ecosystem of the Psunj and Papuk Mountains in Croatia. South-East Eur. For. 2020, 11, 145–152. [Google Scholar] [CrossRef]
- IPBES. Summary for Policymakers of the Global Assessment Report on Biodiversity and Ecosystem Services of the Intergovernmental; IPBES Secretariat: Bonn, Germany, 2019. [Google Scholar]
- Madden, M.J.L.; Young, R.G.; Brown, J.W.; Miller, S.E.; Frewin, A.J.; Hanner, R.H. Using DNA barcoding to improve invasive pest identification at U.S. ports-of-entry. PLoS ONE 2019, 14, e0222291. [Google Scholar] [CrossRef] [PubMed]
- Raclariu, A.C.; Heinrich, M.; Ichim, M.C.; de Boer, H. Benefits and limitations of DNA barcoding and metabarcoding in herbal product authentication. Phytochem. Anal. 2018, 29, 123–128. [Google Scholar] [CrossRef]
- Solano, J.; Anabalón, L.; Figueroa, A.; Gangitano, D. ITS barcoding using high resolution melting analysis of Cannabis sativa drug seizures in Chile: A forensic application. Forensic Sci. Int. 2020, 316, 110550. [Google Scholar] [CrossRef] [PubMed]
- Kuzmina, M.L.; Braukmann, T.W.A.; Zakharov, E.V. Finding the pond through the weeds: eDNA reveals underestimated diversity of pondweeds. Appl. Plant. Sci. 2018, 6, e01155. [Google Scholar] [CrossRef] [PubMed]
- Kress, J.W.; Mazet, J.A.K.; Hebert, P.D.N. Intercepting pandemics through genomics. Proc. Natl. Acad. Sci. USA 2020, 117, 13852–13855. [Google Scholar] [CrossRef] [PubMed]
- Vieira, P.E.; Desiderato, A.; Holdich, D.M.; Soares, P.; Creer, S.; Carvalho, G.R.; Costa, F.O.; Queiroga, H. Deep segregation in the open ocean: Macaronesia as an evolutionary hotspot for low dispersal marine invertebrates. Mol. Ecol. 2019, 28, 1784–1800. [Google Scholar] [CrossRef] [PubMed]
- Hupało, K.; Teixeira, M.A.L.; Rewicz, T.; Sezgin, M.; Iannilli, V.; Karaman, G.S.; Grabowski, M.; Costa, F.O. Persistence of phylogeographic footprints helps to understand cryptic diversity detected in two marine amphipods widespread in the Mediterranean basin. Mol. Phylogen. Evol. 2019, 132, 53–66. [Google Scholar] [CrossRef]
- Sigsgaard, E.E.; Nielsen, I.B.; Carl, H.; Krag, M.A.; Knudsen, S.W.; Xing, Y.; Holm-Hansen, H.T.; Møller, P.R.; Thomsen, P.F. Seawater environmental DNA reflects seasonality of a coastal fish community. Mar. Biol. 2017, 164, 128. [Google Scholar] [CrossRef]
- Parsons, K.M.; Everett, M.; Dahlheim, M.; Park, L. Water, water everywhere: Environmental DNA can unlock population structure in elusive marine species. R. Soc. Open Sci. 2018, 5, 180537. [Google Scholar] [CrossRef] [Green Version]
- Bohmann, K.; Gopalakrishnan, S.; Nielsen, M.; Nielsen, L.D.S.B.; Jones, G.; Streicker, D.G.; Gilbert, M.T.P. Using DNA metabarcoding for simultaneous inference of common vampire bat diet and population structure. Mol. Ecol. Resour. 2018, 18, 1050–1063. [Google Scholar] [CrossRef] [Green Version]
- Zizka, V.M.; Geiger, M.F.; Leese, F. DNA metabarcoding of stream invertebrates reveals spatio-temporal variation but consistent status class assessments in a natural and urban river. Ecol. Indic. 2020, 115, 106383. [Google Scholar] [CrossRef]
- Sigsgaard, E.E.; Jensen, M.R.; Winkelmann, I.E.; Møller, P.R.; Hansen, M.M.; Thomsen, P.F. Population-level inferences from environmental DNA—current status and future perspectives. Evol. Appl. 2020, 13, 245–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stransky, C.; Baumann, H.; Fevolden, S.E.; Harbitz, A.; Høie, H.; Nedreaas, K.H.; Salberg, A.-B.; Skarstein, T.H. Separation of Norwegian coastal cod and Northeast Arctic cod by outer otolith shape analysis. Fish. Res. 2008, 90, 26–35. [Google Scholar] [CrossRef]
- Zobel, M.; Davison, J.; Edwards, M.E.; Brochmann, C.; Coissac, E.; Taberlet, P.; Willerslev, E.; Moora, M. Ancient environmental DNA reveals shifts in dominant mutualisms during the late Quaternary. Nat. Commun. 2018, 9, 139. [Google Scholar] [CrossRef] [Green Version]
- Alsos, I.G.; Lavergne, S.; Merkel, M.K.F.; Boleda, M.; Lammers, Y.; Alberti, A.; Pouchon, C.; Denoeud, F.; Pitelkova, I.; Pușcaș, M. The treasure vault can be opened: Large-scale genome skimming works well using herbarium and silica gel dried material. Plants 2020, 9, 432. [Google Scholar] [CrossRef] [Green Version]
- Lobo, J.; Costa, P.M.; Teixeira, M.A.; Ferreira, M.S.; Costa, M.H.; Costa, F.O. Enhanced primers for amplification of DNA barcodes from a broad range of marine metazoans. BMC Ecol. 2013, 13, 34. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leite, L.A.R. Mitochondrial pseudogenes in insect DNA barcoding: Differing points of view on the same issue. Biota Neotrop. 2012, 12, 301–308. [Google Scholar] [CrossRef] [Green Version]
- Miya, M.; Sato, Y.; Fukunaga, T.; Sado, T.; Poulsen, J.Y.; Sato, K.; Minamoto, T.; Yamamoto, S.; Yamanaka, H.; Araki, H.; et al. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: Detection of more than 230 subtropical marine species. R. Soc. Open Sci. 2015, 2, 150088. [Google Scholar] [CrossRef] [Green Version]
- Taberlet, P.; Coissac, E.; Pompanon, F.; Gielly, L.; Miquel, C.; Valentini, A.; Vermat, T.; Corthier, G.; Brochmann, C.; Willerslev, E. Power and limitations of the chloroplast trnL (UAA) intron for plant DNA barcoding. Nucleic Acids Res. 2007, 35, e14. [Google Scholar] [CrossRef] [Green Version]
- Doyle, J.; Uthicke, S. Sensitive environmental DNA detection via lateral flow assay (dipstick)-A case study on corallivorous crown-of-thorns sea star (Acanthaster cf. solaris) detection. Environ. DNA 2021, 3, 323–342. [Google Scholar] [CrossRef]
- Naaum, A.M.; Cusa, M.; Singh, M.; Bleicher, Z.; Elliott, C.; Goodhead, I.B.; Hanner, R.H.; Helyar, S.J.; Mariani, S.; Rice, J.E.; et al. Validation of FASTFISH-ID: A new commercial platform for rapid fish species authentication via universal closed-tube barcoding. Food Res. Int. 2021, 141, 110035. [Google Scholar] [CrossRef] [PubMed]
- Kobori, H.; Dickinson, J.L.; Washitani, I.; Sakurai, R.; Amano, T.; Komatsu, N.; Kitamura, W.; Takagawa, S.; Koyama, K.; Ogawara, T.; et al. Citizen science: A new approach to advance ecology, education, and conservation. Ecol. Res. 2016, 31, 1–19. [Google Scholar] [CrossRef] [Green Version]
- Steinke, D.; Breton, V.; Berzitis, E.; Hebert, P.D.N. The School Malaise Trap Program: Coupling educational outreach with scientific discovery. PLoS Biol. 2017, 15, e2001829. [Google Scholar] [CrossRef]
- Watanabe, M.E. The Nagoya Protocol: The conundrum of defining digital sequence information. Bioscience 2019, 69, 480. [Google Scholar] [CrossRef] [Green Version]
- Bond, M.R.; Scott, D. Digital biopiracy and the (dis)assembling of the Nagoya Protocol. Geoforum 2020, 117, 24–32. [Google Scholar] [CrossRef]
- Houssen, W.; Sara, R.; Jaspars, M. Digital Sequence Information on Genetic Resources: Concept, Scope and Current Use; Convention on Biological Conservation CBD/DSI/AHTEG: Montreal, QC, Canada, 2020. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grant, D.M.; Brodnicke, O.B.; Evankow, A.M.; Ferreira, A.O.; Fontes, J.T.; Hansen, A.K.; Jensen, M.R.; Kalaycı, T.E.; Leeper, A.; Patil, S.K.; et al. The Future of DNA Barcoding: Reflections from Early Career Researchers. Diversity 2021, 13, 313. https://doi.org/10.3390/d13070313
Grant DM, Brodnicke OB, Evankow AM, Ferreira AO, Fontes JT, Hansen AK, Jensen MR, Kalaycı TE, Leeper A, Patil SK, et al. The Future of DNA Barcoding: Reflections from Early Career Researchers. Diversity. 2021; 13(7):313. https://doi.org/10.3390/d13070313
Chicago/Turabian StyleGrant, Danielle M., Ole Bjørn Brodnicke, Ann M. Evankow, André O. Ferreira, João T. Fontes, Aslak Kappel Hansen, Mads Reinholdt Jensen, Tuğba Ergül Kalaycı, Alexandra Leeper, Shalaka Kiran Patil, and et al. 2021. "The Future of DNA Barcoding: Reflections from Early Career Researchers" Diversity 13, no. 7: 313. https://doi.org/10.3390/d13070313
APA StyleGrant, D. M., Brodnicke, O. B., Evankow, A. M., Ferreira, A. O., Fontes, J. T., Hansen, A. K., Jensen, M. R., Kalaycı, T. E., Leeper, A., Patil, S. K., Prati, S., Reunamo, A., Roberts, A. J., Shigdel, R., Tyukosova, V., Bendiksby, M., Blaalid, R., Costa, F. O., Hollingsworth, P. M., ... Ekrem, T. (2021). The Future of DNA Barcoding: Reflections from Early Career Researchers. Diversity, 13(7), 313. https://doi.org/10.3390/d13070313









