Abstract
Taxonomic studies of bambusicolous fungi in China and Thailand have resulted in the collection of three fascinating saprobic coelomycetes strains. Morphology coupled with combined gene analysis of ITS, LSU, TUB2, and TEF1-α DNA sequence data showed that they belong to the genus Apiospora, family Apiosporaceae. A new species from Thailand, Apiospora mukdahanensis, and new records of A. locuta-pollinis from China are herein described. In addition, based on both morphological data coupled with phylogenetics and nomenclatural analyses, A. mori is proposed as a new combination. Maximum likelihood, maximum parsimony and Bayesian analyses were performed to clarify the phylogenetic affinities of the species obtained in this study. Newly obtained strains are compared with morphologically- and phylogenetically-related taxa. The comprehensive descriptions, illustrations, and updated phylogeny are provided and discussed for intra-and intergeneric relationships within Apiospora species.
1. Introduction
Apiospora is a large genus in the family Apiosporaceae (Amphisphaeriales, Sordariomycetes, Ascomycota) [1,2], which is ecologically diverse and distributed worldwide [2,3,4,5]. Most species have been identified as saprobes and endophytes of a range of plant hosts, mainly occurring on the family Poaceae [2,3,4,5,6,7,8,9,10,11,12]. In addition, some species have been reported as plant pathogens. For instance, A. kogelbergensis causes the blight disease of Schizostachyum [13], A. sacchari causes the damping-off of durum wheat (Triticum durum) [14], and A. xenocordella causes fruit blight on pistachio (Pistacia vera) [15]. Apiospora shows a cosmopolitan distribution in diverse substrates, including air [4,16], soil [4,16,17,18], freshwater [19], marine environments [20,21,22,23,24,25], lichens [26], insect guts [27], and human tissues [3,28,29,30]. Interestingly, some species (e.g., A. arundinis, A. sacchari) have been reported as a source of useful bioactive compounds, such as antifungal agents and enzymes [21,22,31], possessing great potential for their commercial applications in the pharmaceutical industries.
Apiospora is classified by asexual morph characteristics that produce basauxic conidiophores and unicellular globose to obovoid conidia, usually rounded in face view and lenticular in side view, with a longitudinal germ slit [2,7,10,11,32]. The sexual morph is characterized as having multi-locular perithecial stromata, clavate to broadly cylindrical asci and hyaline ascospores surrounded by a thick gelatinous sheath [2,7,8,11]. Apiospora was previously known as the sexual morph of the genus Arthrinium [33,34]. According to the International Code of Nomenclature for Algae, Fungi, and Plants (ICN) [35], Apiospora was a synonym of Arthrinium due to the early introduction of Arthinium and is more commonly used in the literature [3]. Crous and Groenewald [3] and Wang et al. [4] provided the upgraded phylogenetic trees of Arthrinium species (=Apiospora) using combined ITS, TEF1-α, and TUB2 sequence data with additional strains (collected from various hosts, substrates, and locations) and indicated that Arthrinium seems to be a species complex which needs further taxonomic revision and epitypification. Multi-gene phylogeny of ITS, LSU, TEF1-α, and TUB2 sequences conducted by Pintos et al. [5] revealed that Arthrinium caricicola, the type species, and other species of Arthrinium mostly found in Carex sp. formed independent lineages unrelated to other species of Arthrinium, and reported that Apiospora occurred on other hosts. However, the taxonomic placement of both genera was uncertain until Pintos and Alvarado [2] resolved this issue and presented Arthrinium and Apiospora as well-supported distinct clades suggesting they are separate genera.
The morphological identification of Apiospora species is challenging because most species share similar morphological characteristics (e.g., conidia). In addition, their morphological features can vary depending on incubation periods and different substrates [3]. Thus, morphological characteristics integrated with molecular phylogeny have been widely accepted to distinguish Apiospora species [3,4,5,6,7,8,9,12,17,36,37,38,39]. Presently, 117 epithets are recognized in Apiospora [40], comprising 76 Arthrinium species, which were synonymized under Apiospora [2,12,25]. The taxonomic position of other taxa, which lack sequencing information and comprehensive morphological descriptions, remain uncertain and require further study. In this study, we isolated apiospora-like taxa from bamboo in China and Thailand. The morphological characteristics and molecular analyses of ITS, LSU, TUB2, and TEF1-α were applied to determine a new species, Apiospora mukdahanensis, and one new record of A. locuta-pollinis. Furthermore, Arthrinium mori is also transferred to Apiospora as a new combination on the basis of phylogenetic evidence. The host association, geographical distribution, and species diversity of Apiospora are also discussed.
2. Materials and Methods
2.1. Sample Collection, Fungal Isolation and Morphological Examination
Dead and decaying bamboo specimens were collected during a series of field trips conducted in China and Thailand from the year 2019–2021. The specimens were packed in zip-lock plastic bags prior for further study. Fungal colonies on the host substrate were observed using a stereo microscope (Nikon SMZ800N, Tokyo, Japan). The micromorphological characteristics were documented and photographed with a compound microscope (Nikon Eclipse Ni U, Tokyo, Japan) equipped with Nikon DS-Ri2 camera. The measurements of fungal structures (i.e., conidiomata, conidiophore mother cells, conidiophores, conidiogenous cells, and conidia) were made using the Tarosoft (R) Image Frame Work program. Images used for figures were combined and edited using Adobe Photoshop CS6 Extended version 10.0 software (Adobe Systems, San Jose, CA, USA). Single-spore isolation was conducted to isolate the fungus as detailed in Senanayake et al. [41]. The germinating conidia were inoculated on potato dextrose agar (PDA) and incubated at 28 °C for two weeks. Culture characteristics were observed and described after four weeks. Axenic cultures were kept in 2 mL sterilized screw-cap tubes containing PDA for short-term storage and duplicated in 10% glycerol for long-term storage. The type specimen and living culture of Apiospora mukdahanensis were deposited in the herbarium of Mae Fah Luang University (MFLU) and the Mae Fah Luang University Culture Collection (MFLUCC), respectively. The specimens of A. locuta-pollinis and cultures were deposited in the Herbarium of Cryptogams Kunming Institute of Botany Academia Sinica (KUN-HKAS), and the Kunming Institute of Botany Culture Collection, Kunming, China (KUNCC), respectively. Index Fungorum numbers and Faces of Fungi numbers were obtained for the new taxa as detailed in Index Fungorum [42] and Jayasiri et al. [43], respectively.
2.2. DNA Extraction, PCR Amplification and Sequencing
The total genomic DNA was extracted from mature mycelium grown on PDA at 28 °C for two weeks using a Biospin Fungus Genomic DNA Extraction Kit (BioFlux®, Hangzhou, China). Polymerase chain reaction (PCR) amplification was applied to amplify DNA fragments with three phylogenetic markers, including the internal transcribed spacers region of ribosomal DNA (ITS; ITS1-5.8s-ITS2) using primers ITS5 and ITS4 [44]; the partial 28S large subunit nuclear ribosomal DNA (LSU) using primers LR0R and LR5 [45]; and the translation elongation factor 1-α (TEF1-α) using primers EF1-728F and EF-2 [46,47]. The PCR reaction was carried out in the final volume of 25 μL, containing 2 μL template DNA (50 ng/μL), 12.5 μL of PCR Master Mix (0.5 mM of each primer, 50 U Taq DNA polymerase 400 mM of each dNTP, and 3 mM MgCl2), 1 μL of each forward and reward primer and 8.5 μL of the sterilized double-distilled water (ddH2O). The PCR thermal cycling programs for ITS, LSU and TEF1-α were adjusted by an initial denaturation at 94 °C for 3 min, followed by 40 cycles of denaturation at 94 °C for 45 s, annealing at 56 °C for 50 s, extension at 72 °C for 1 min and a final extension step at 72 °C for 10 min. The PCR products were processed for purification and sequencing by TsingKe Biological Technology, Kunming City, Yunnan Province, China. Newly generated sequences in this study were deposited in Genbank (Table 1).

Table 1.
List of the taxa used in phylogenetic reconstruction and their corresponding GenBank numbers.
2.3. Phylogenetic Analyses
The sequences generated by this study were supplemented with the related taxa resulting from the nucleotide blast search in GenBank (www.ncbi.nlm.nih.gov/blast/, accessed on 1 September 2022) and recent publications [2,12,25,39,48,49] (Table 1). The matrix of consensus sequences was aligned using MAFFT v. 7.475 on the web portal (http://mafft.cbrc.jp/alignment/server/index.html) [50] and then the ambiguous sites were manually trimmed using BioEdit 7.1.3.0 [51]. Phylogenetic trees based on the concatenated ITS, LSU, TUB2, and TEF1-α sequence data (analysis 1) and ITS, LSU, and TEF1-α sequence data (analysis 2) were inferred to clarify the phylogenetic relationships of Apiospora species using maximum likelihood (ML) and Bayesian inference (BI) analyses. In order to clarify the phylogenetic placements of new strains and related strains in A. locuta-pollinis/marii clade, ML, maximum parsimony (MP), and BI were analyzed based on the concatenated ITS, LSU, TUB2, and TEF1-α sequence data (analysis 3). Phylogenetic trees of these combined gene datasets were further compared to check the congruence of the tree topologies.
ML analyses were implemented using RAxML-HPC2 (v.8.2.12) on the CIPRES web portal (http://www.phylo.org/portal2/) [52]. The GTRGAMMAI model of nucleotide substitution with 1000 rapid bootstrap replicates was used. BI analyses were performed using MrBayes v.3.2.7a via the CIPRES web portal (http://www.phylo.org/portal2/). The optimal substitution model of nucleotide evolution was determined using MrModeltest v. 2.3 [53]. In the first and second analyses, GTR + I + G was chosen as the best-fit model for the ITS, LSU, and TEF1-α datasets, and HKY + I + G for the TUB2 dataset. For the third analysis, the best-fit model for the ITS, LSU, TUB2, and TEF1-α datasets were as follows: SYM + I, GTR, GTR + G, and K80 + I. Ten million Markov chain Monte Carlo sampling (MCMC) generations were run with six simultaneous Markov chains to calculate Bayesian posterior probabilities [54,55,56]. Trees were sampled every 100th generation. When the average standard deviation of split frequencies was constantly below 0.01, the runs were automatically stopped and the first 25% of the generated trees were discarded. The remaining trees were used to evaluate the posterior probabilities (PP) of the majority rule consensus tree. MP analyses were conducted with the heuristic search option, as implemented in PAUP v. 4.0b10 [57]. Clade stability was determined using a bootstrap analysis with 1000 replicates, random sequence additions with maxtrees set to 1000 [58]. The MP tree was described for descriptive tree statistics including Tree Length (TL), Consistency Index (CI), Retention Index (RI), Relative Consistency Index (RC), and Homoplasy Index (HI) under different optimality criteria. Phylogenetic trees were viewed in FigTree v1.4.0 [59] and formatted using Adobe Photoshop CS6 software (Adobe Systems, San Jose, CA, USA).
2.4. Host and Geographical Distribution of Apiospora Species
To investigate geographical distribution and host-substratum diversity of the Apiospora species, the data were summarized based on the USDA fungal database (https://nt.ars-grin.gov/fungaldatabases/, accessed on 1 September 2022), academic literature, and this study).
3. Results
3.1. Phylogenetic Analyses
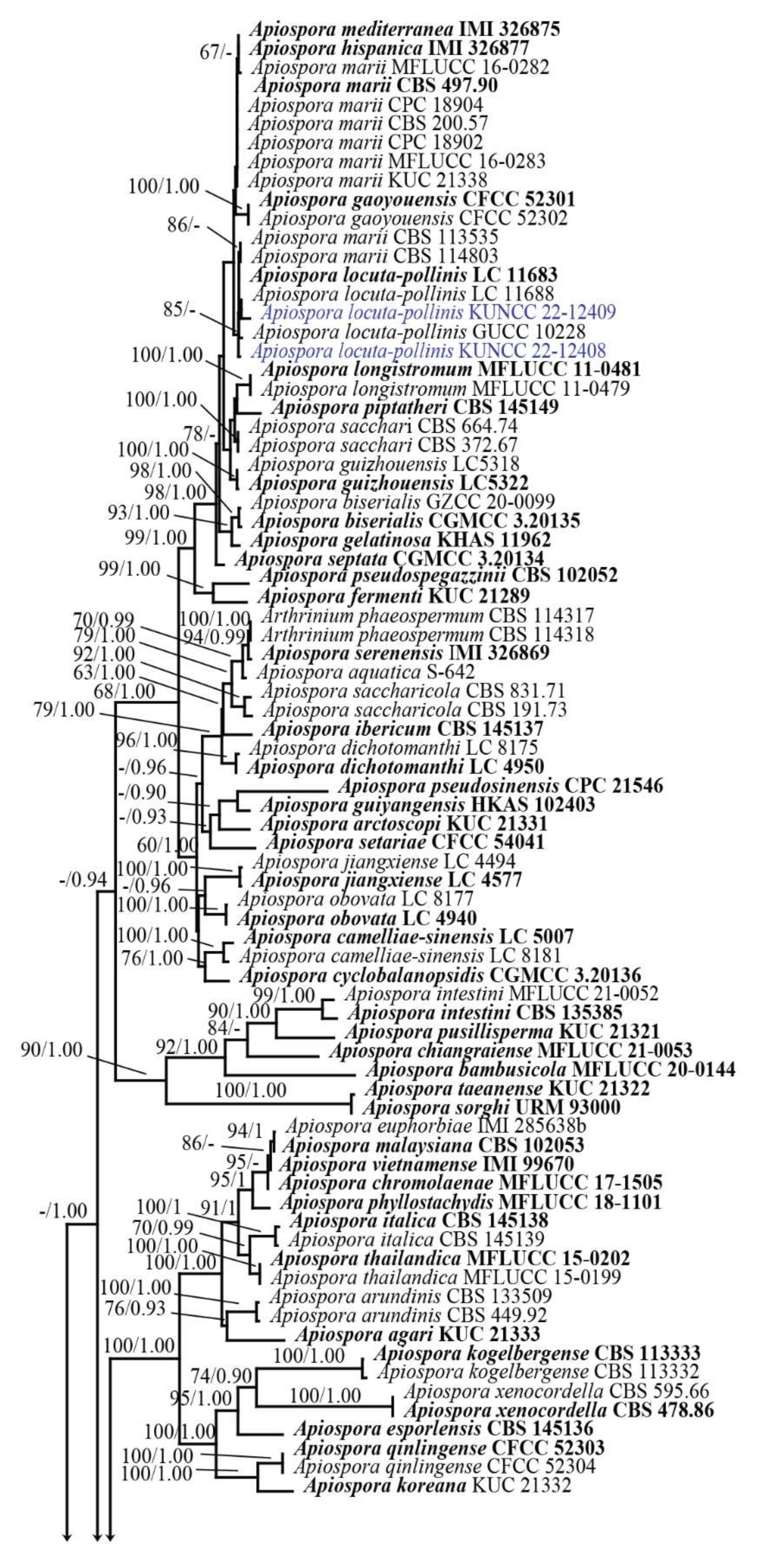
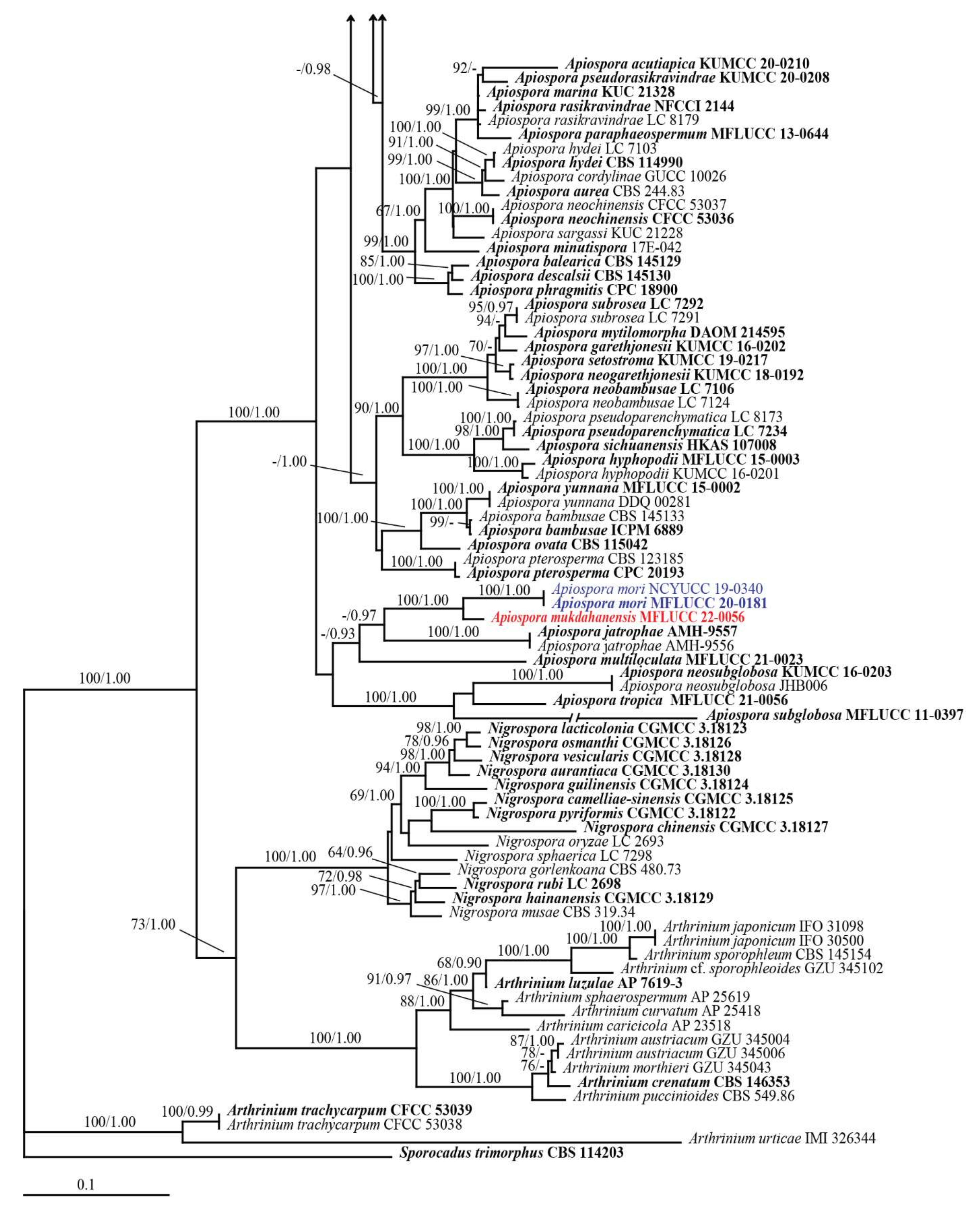
Analysis 1: The combination of ITS, LSU, TUB2, and TEF1-α sequence dataset comprised 156 taxa of Apiospora, and other related taxa in the family Apiosporaceae. Sporocadus trimorphus (CBS 114203) was selected as the outgroup taxon (Figure 1). The final alignment consisted of 3264 total characters, including gaps (ITS: 1–641 bp, LSU: 642–1524 bp, TUB2: 1525–2366 bp, TEF1-α: 2367–3264 bp). The RAxML analysis of the integrated dataset yielded a best scoring tree with a final ML optimization likelihood value of −39,292.041642. The aligned sequence matrix contained 1874 distinct alignment patterns, with 38.58% undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.237643, C = 0.255831, G = 0.254014, T = 0.252512; substitution rates AC = 1.214378, AG = 2.729624, AT = 1.181789, CG = 1.027511, CT = 4.357413, and GT = 1.000000; gamma distribution shape parameter α = 0.294728. Bayesian analysis resulted in the average standard deviation of split frequencies as 0.009934.
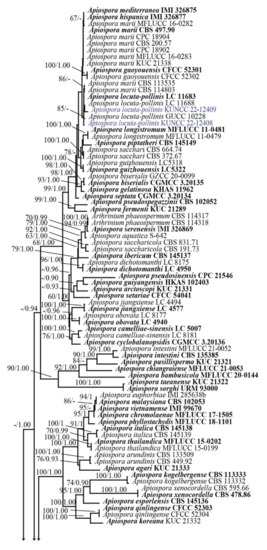
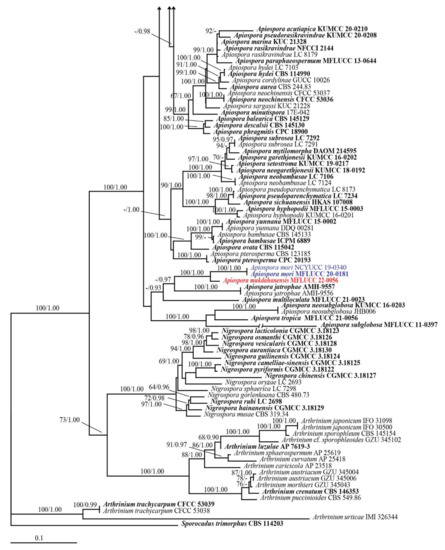
Figure 1.
Phylogenetic tree retrieved from RAxML analyses of a combined ITS, LSU, TUB2, and TEF1-α data sequence of Apiospora, and other related taxa in the family Apiosporaceae. Bootstrap support values for ML equal or greater than 60% and Bayesian posterior probabilities greater than 0.90 are indicated at the nodes as ML/PP. The ex-type strains are in bold. The new species are in red and new record and new combination species are in blue. The tree is rooted to Sporocadus trimorphus (CBS 114203).
Analysis 2: The combination of ITS, LSU, and TEF1-α sequence dataset comprised 156 taxa of Apiospora, and other related taxa in Apiosporaceae. Sporocadus trimorphus (CBS 114203) were selected as the outgroup taxon (Figure S1). The final alignment consisted of 2422 total characters, including gaps (ITS: 1–641 bp, LSU: 642–1524 bp, TEF1-α: 1525–2422 bp). The RAxML analysis of the integrated dataset yielded a best scoring tree with a final ML optimization likelihood value of −24,247.334486. The aligned sequence matrix contained 1182 distinct alignment patterns, with 37.22% undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.245462, C = 0.240858, G = 0.258736, T = 0.254944; substitution rates AC = 1.136736, AG = 2.122936, AT = 1.134884, CG = 1.008342, CT = 4.360445, and GT = 1.000000; gamma distribution shape parameter α = 0.245115. Bayesian analysis resulted in the average standard deviation of split frequencies as 0.009872.
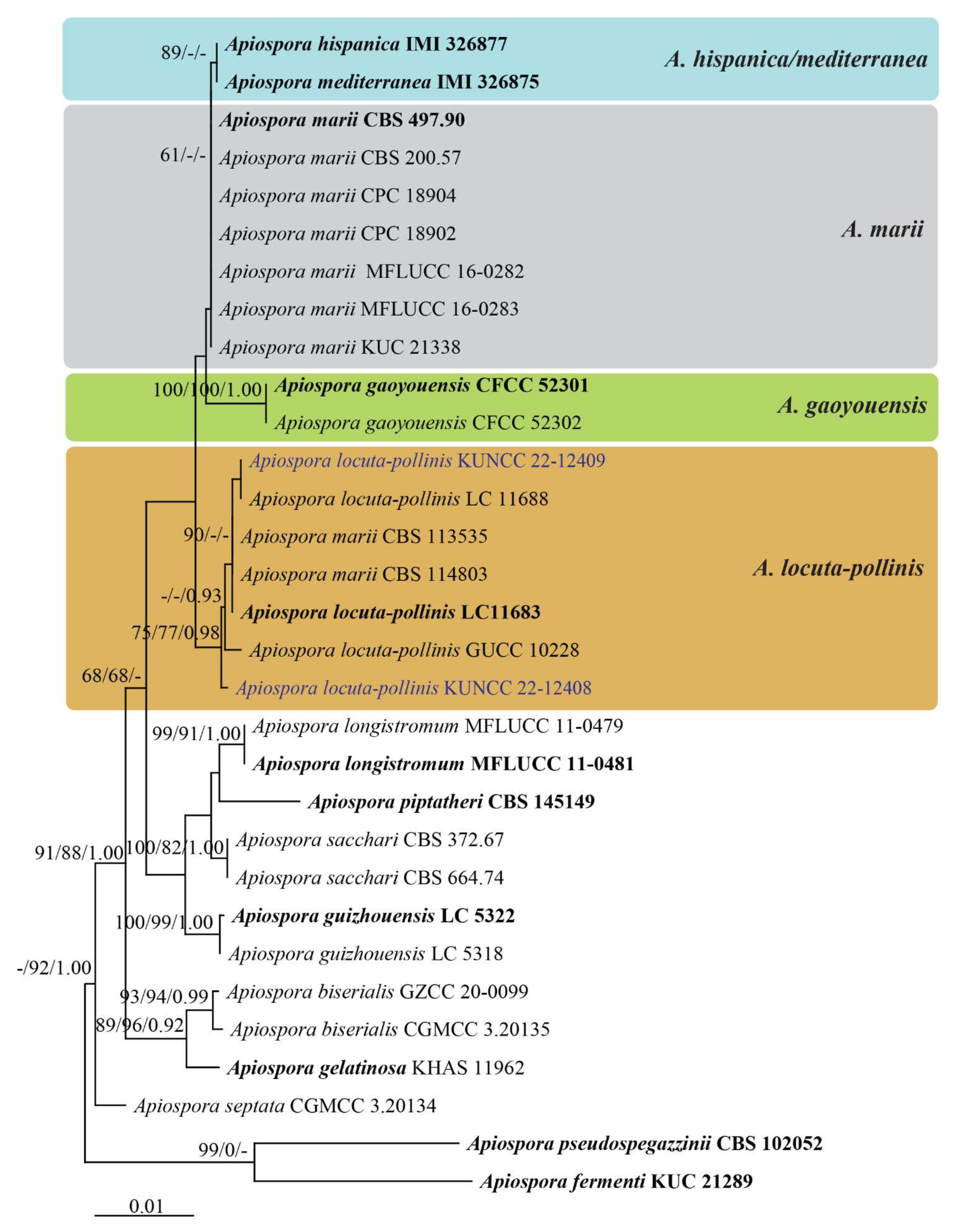
Analysis 3: The combination of ITS, LSU, TUB2, and TEF1-α sequence dataset comprised 31 taxa in Apiospora locuta-pollinis/marii clade. Apiospora fermenti KUC 2189 and A. pseudospegazzinii CBS 102052 were selected as the outgroup taxa (Figure 2). The final alignment consisted of 2735 total characters, including gaps (ITS: 1–626 bp, LSU: 627–1470 bp, TUB2: 1471–2296 bp, TEF1-α: 2297–2735 bp). The RAxML analysis of the integrated dataset yielded a best scoring tree with a final ML optimization likelihood value of −5544.668718. The aligned sequence matrix contained 363 distinct alignment patterns, with 25.58% undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.237367, C = 0.256117, G = 0.247967, T = 0.258550; substitution rates AC = 1.390352, AG = 3.604609, AT = 1.633776, CG = 0.564937, CT = 4.605733, and GT = 1.000000; gamma distribution shape parameter α = 0.020000. The maximum parsimonious dataset consisted of 2744 characters of which 2523 were constant, 119 parsimony-informative and 102 parsimony-uninformative. The parsimony analysis of the data matrix resulted in a single most parsimonious tree (TL = 266, CI = 0.883, RI = 0.914, RC = 0.808, HI = 0.117). Bayesian analysis resulted in the average standard deviation of split frequencies as 0.009680.
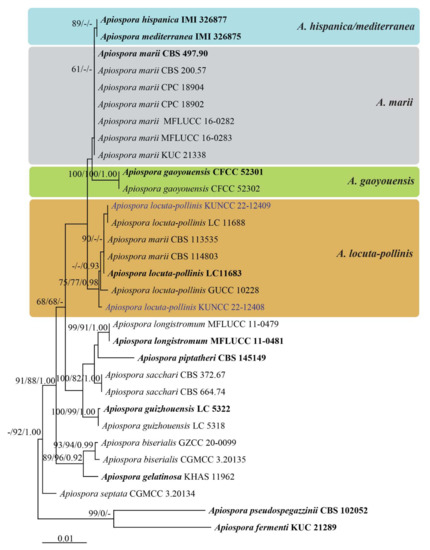
Figure 2.
Phylogenetic tree retrieved from RAxML analyses of a combined ITS, LSU, TUB2, and TEF1-α data sequence of taxa in Apiospora locuta-pollinis/marii clade. Bootstrap support values for ML and MP equal or greater than 60% and Bayesian posterior probabilities greater than 0.90 are indicated at the nodes as ML/MP/PP. The ex-type strains are in bold. The new record strains are in blue. The tree is rooted to Apiospora fermenti KUC 2189 and A. pseudospegazzinii CBS 102052.
Phylogenetic analyses inferred from ML and BI analyses were not significantly different and showed congruent topologies. The overall tree topologies of the concatenated ITS-LSU-TUB2-TEF1-α sequence dataset (Figure 1) were also congruent with the tree topologies of a concatenated ITS-LSU-TEF1-α sequence matrix (Figure S1). However, the first analysis (Figure 1) revealed higher statistical support than that from the second analysis (Figure S1). Therefore, the phylogenetic results of the concatenated ITS-LSU-TUB2-TEF1-α sequence matrix was selected for discussion in this study. Phylogenetic results showed that the new species, Apiospora mukdahanensis formed a well-resolved clade sister to A. mori with significant support (100% ML/1.00 PP, Figure 1). The new strains, KUNCC 22-12408 and KUNCC 22-12409 clustered in the same clade with A. locuta-pollinis including the ex-type strain (LC 11683) with 85% ML support (Figure 1).
The multigene phylogeny based on the ITS-LSU-TUB2-TEF1-α sequence data of the species in the first clade (Apiospora locuta-pollinis/marii clade) revealed a similar result between ML, MP, and BI analyses. The results indicated that A. locuta-pollinis formed a well-supported clade with 75% ML/77% MP/0.98 PP support including the new strain KUNCC 22-12408 which formed a separated branch basal to other A. locuta-pollinis strains and the strain KUNCC 22-12409 shared the same branch length with A. locuta-pollinis (LC 11688) (Figure 2), whereas the type of A. hispanica, A. mediterranea, and A. marii were not well separated and clustered together with low support (Figure 2). Apiospora gaoyouensis formed an independent clade basal to A. marii with significant support (100% ML/100% MP/1.00 PP, Figure 2). The two strains of A. marii (CBS 113535 and CBS 114803) grouped within A. locuta-pollinis clade (Figure 2).
3.2. Taxonomy
3.2.1. Apiospora mori (Tennakoon, C.H. Kuo and K.D. Hyde) Monkai and Phookamsak, comb. nov.
Index Fungorum number: IF559913; Facesoffungi number: FoF 12871
Basionym: Arthrinium mori Tennakoon, C.H. Kuo and K.D. Hyde, Fungal diversity 108: 215 (2021).
Notes: Arthrinium mori was introduced by Tennakoon et al. [60] from dead leaves of Morus australis in Taiwan. Tennakoon et al. [60] noted that Ar. mori forms a well-supported branch sister to Ar. jatrophae with 86% ML/1.00 PP support, but differs from Ar. jatrophae in having smaller conidia (4.5–5.5 × 4–5 vs. 6.5–9.7 × 3.2–6.5 μm) [6]. In our phylogenetic analyses, Ar. mori constitutes an independent clade sister to Apiospora mukdahanensis with high support (100% ML/1.00 PP, Figure 1). Based on the phylogenetic analysis, 55 Arthrinium species were proposed as new combinations of Apiospora, but they did not include Ar. mori [2]. Thus, we transferred Ar. mori under the new combination A. mori, on the basis of morphological and molecular data.
3.2.2. Apiospora mukdahanensis Monkai and Phookamsak, sp. nov.
Index Fungorum number: IF559912; Facesoffungi number: FoF 12853; Figure 3.
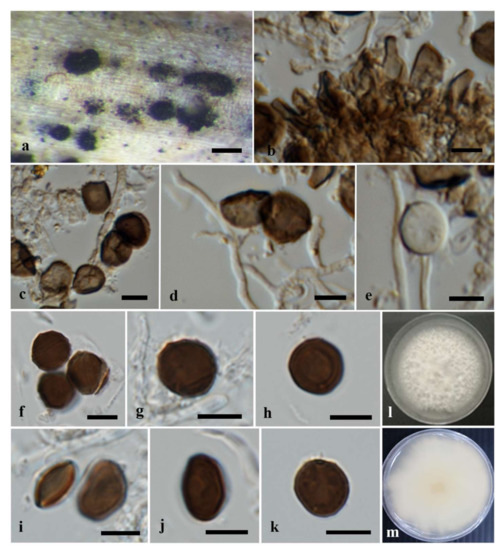
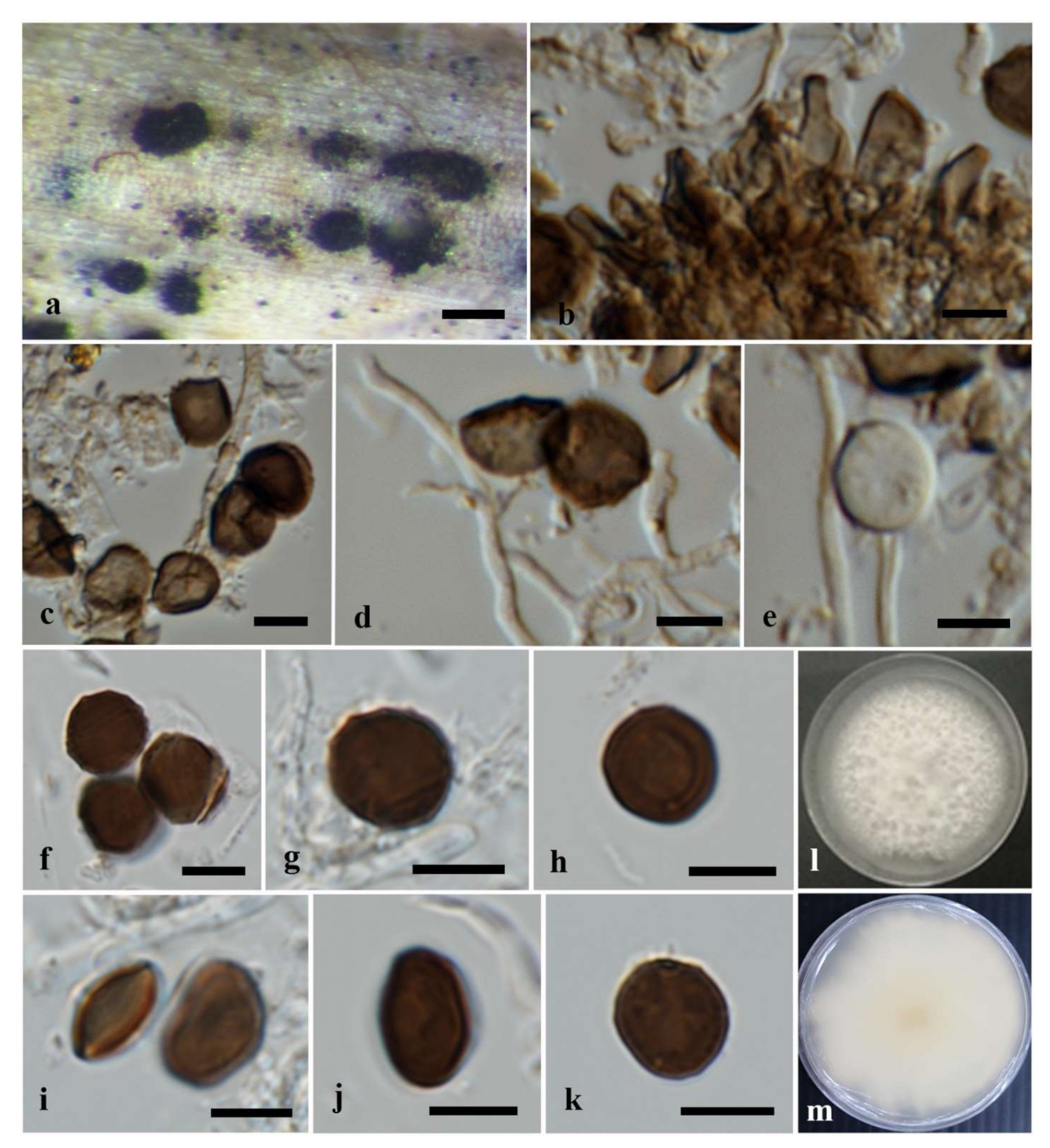
Figure 3.
Apiospora mukdahanensis (MFLU 22-0104, holotype). (a) Conidiomata on host substrate. (b) Conidiophore mother cells. (c–e) Conidiophores, conidiogenous cells and conidia. (f–k) Conidia. (l,m) Colonies on PDA ((l) from above, (m) from reverse). Scale bars: (a) = 200 μm, (b–k) = 5 μm.
Etymology: Referring to the locality, Mukdahan Province, Thailand, where the holotype was collected.
Holotype: MFLU 22-0104
Saprobic on dead bamboo leaves. Sexual morph: Undetermined. Asexual morph: Conidiomata sporodochia, rounded to ovoid, pulvinate, dark brown to black, 150–400 μm diam. Mycelium basal, consists of smooth, brown to dark brown, branched septate hyphae. Conidiophore mother cells arising from the mycelial mat, obpyriform to lageniform, dark brown, smooth, (6–)7–8.5(–10.5) × (2.5–)3.5–4(–5.5) μm (x = 8.5 × 4 μm, n = 12). Conidiophores basauxic, arising from conidiophore mother cells, cylindrical, pale brown, septate, with brown transversal septa, straight or flexuous. Conidiogenous cells polyblastic, smooth, hyaline to pale brown, ampulliform, cylindrical or lageniform, (6–)8.5–14(–20) × 1.5–2(–3) μm (x = 11 × 2 μm, n = 15). Conidia brown to dark brown, smooth to slightly roughened, thick-walled, globose to subglobose, or irregularly round in face view, lenticular in side view (6–)–7.5(–8) × (5–)–6.5(–7) μm (x = 7 × 6 μm, n = 30), with a pale equatorial slit, and a central scar.
Culture characteristics: Colonies on PDA reached at 9 cm diam. in 2 weeks at 28 °C, flat, fluffy, spreading, with abundant aerial mycelium, irregular margin, white to cream, in reverse, white and pale yellowish in the center.
Material examined: THAILAND, Mukdahan Province, Tambon Phang Daeng, Amphoe Dong Luang, on dead bamboo leaves, 24 July 2019, E. Yasanthika, E2B-4 (MFLU 22-0104, holotype), ex-type living culture, MFLUCC 22-0056.
Notes: The nucleotide BLAST search of ITS region showed that Apiospora mukdahanensis (MFLUCC 22-0056) has high similarity with Arthrinium sp. strain 4–13 (99.19%), Arthrinium sp. strain NF34_TK10 and A. mori strain MFLU 18-2514 (98.47%). The nucleotide BLAST search of LSU region showed that A. mukdahanensis (MFLUCC 22-0056) has high similarity with Apiospora sp. strain NF34_TK10 (99.03%), Apiospora sp. strain JYZ-2021a (98.95%) and A. mori MFLU 18-2514 (98.83%). The nucleotide BLAST search of TEF1-α region showed that A. mukdahanensis (MFLUCC 22-0056) has high similarity with A. phragmitis strain MFLUCC 18-0099 (95.96%), A. locuta-pollinis strain LC 11689, LC 11688 (95.92%), and Arthrinium sp. strain MFLU 18-2333 (95.64%).
Based on phylogenetic analysis, Apiospora mukdahanensis formed an independent lineage sister to A. mori with 100% ML and 1.00 PP support (Figure 1). Morphologically, Apiospora mukdahanensis is distinct from A. mori in having larger conidia (6–8.1 × 5.1–6.9 vs. 4.5–5.5 × 4–5 μm) with slightly roughened wall, whereas A. mori has smooth-walled conidia [60]. The new species resembles A. jatrophae in having overlapping size range of conidia (6–8.1 × 5.1–6.9 vs. 6.5–9.7 × 3.2–6.5 μm) [6]. However, A. jatrophae differs from A. mukdahanensis in having thick multi-septate conidiophores with brown transverse septa and two types of conidia including smooth-walled, lenticular conidia and anomalous conidia [6]. The base-pair comparison of ITS gene region showed 4.1% base pair differences (21/508 bp) between A. mukdahanensis and A. mori (MFLUCC 20-0181 and NCYUCC 19-0340) and 9.6% base pair differences (48/498 bp) between A. mukdahanensis and A. jatrophae (AMH-9556, AMH-9557). However, the TEF1-α sequence data are not available for A. mori and A. jatrophae to compare with our new species.
3.2.3. Apiospora locuta-pollinis (F. Liu and L. Cai) Pintos and P. Alvarado, Fungal Systematics and Evolution 7: 206 (2021)
Index Fungorum number: IF837763; Facesoffungi number: FoF 05221, Figure 4.
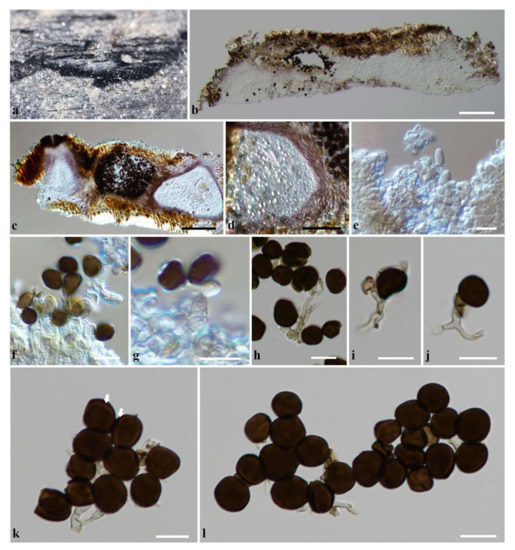
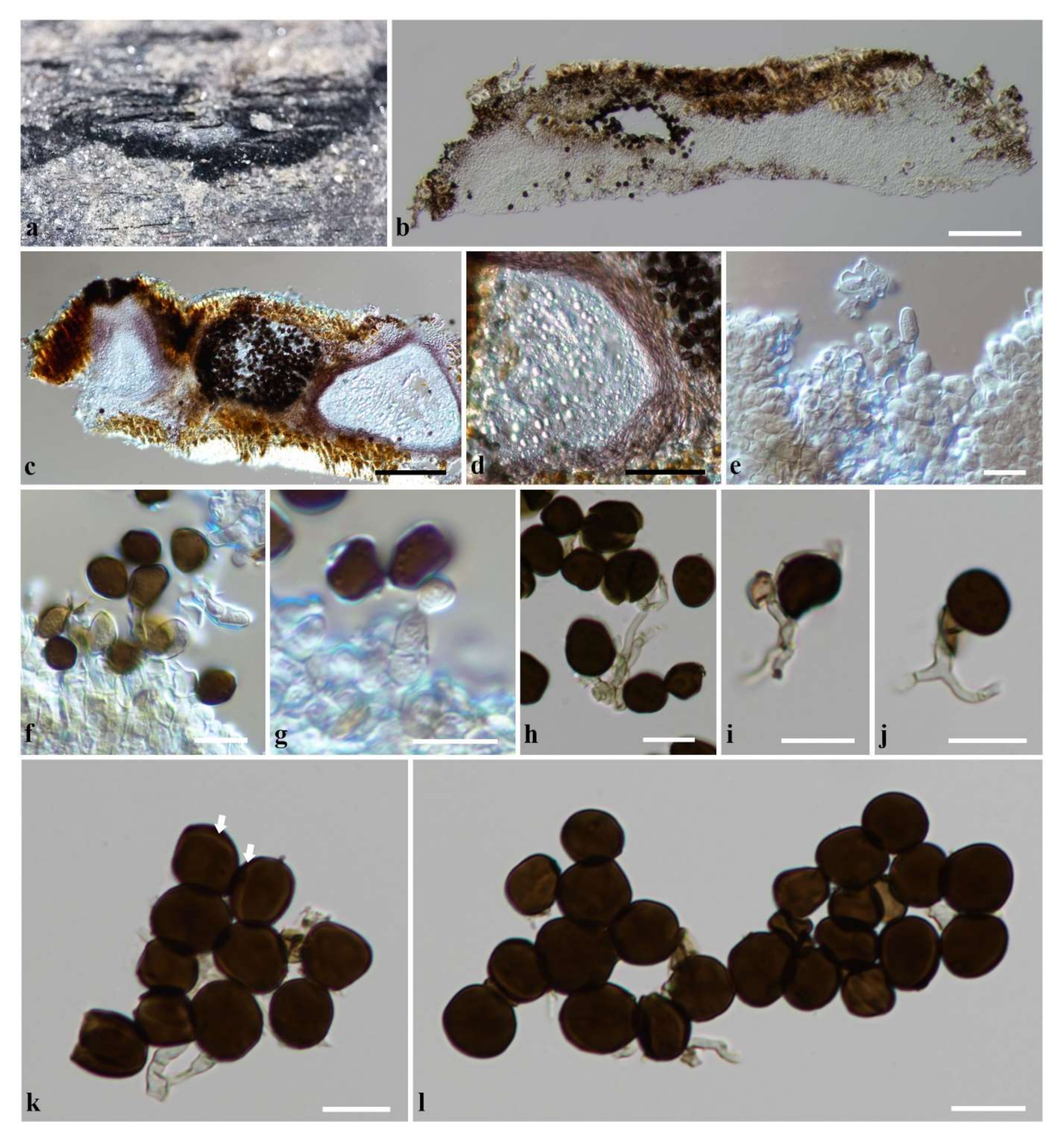
Figure 4.
Apiospora locuta-pollinis (KUN-HKAS 124566). (a) Conidiomata on host substrate. (b,c) Section through the stromata. (d) Section through pycnidial wall. (e) Conidiophore mother cells. (f,g) Conidiogenous cells giving rise conidia. (h–j) Conidiophores (sterile cells attached to the conidiophore in (i,j)). (k,l) Conidia (equatorial slit is indicated by arrows in (k)). Scale bars: (b,c) = 100 µm, (d) = 50 µm, (e–l) = 10 µm.
Basionym: Arthrinium locuta-pollinis F. Liu and L. Cai, Mycosphere 9(6): 1094 (2018).
Saprobic on decaying stems of bamboo. Sexual morph: Immature state found associated with asexual morph on host. Asexual morph: Conidiomata associated with the sexual morph, pycnidial, raised, stromatic at base, 150–230 µm high, 450–830 µm diam., covered by black conidial masses on host surface, laying parallel to the longitudinal axis of the stem, clustered, solitary to gregarious, subepidermal to erumpent, globose to subconical, or hemispherical, uni- to multi-loculate; individual pycnidium 200–260 µm diam., 150–210 µm high, glabrous, ostiolate, opening by longitudinal splitting of the epidermis. Pycnidial wall (15–)20–40(–50) µm wide, of unequal thickness, slightly thick at sides towards apex, thinner at base, composed of several cell layers of reddish-brown to dark brown pseudoparenchymatous cells of textura angularis to textura prismatica, with outer layers intermixed with the host’s tissues. Conidiophore mother cells (3.5–)4.5–9(–12) × 3.5–5.5 µm (x = 7.8 × 4.8 µm, n = 30), arising in dense packs from hyaline to light brown, irregularly angled palisade-like cells (4–9 × (2.8–)3–4.5 µm) in the stroma, subhyaline to pale brown, ampulliform to cylindrical, tapering towards rounded apex with small granules. Conidiophores (4–)5–10(–20) × (1.5–)2–3.5 µm (x = 8.9 × 2.2 µm, n = 30), basauxic, arising from conidiophore mother cells, conspicuous, short, septate, branched, smooth, pale brown, flexuous. Conidiogenous cells (3.5–)4.5–6.5(–9) × 2.5–4.5 µm (x = 6 × 3.5 µm, n = 30), polyblastic, straight or flexuous, cylindrical to subcylindrical or ampulliform, hyaline to light brown, smooth or with small granules, moderately brown, denticulate, sometimes flattened. Conidia (8–)9–13 × (7–)9–12 µm (x = 10.5 × 10 µm, n = 50), globose to obovoid, or ellipsoidal, dark brown, smooth-walled, with a paler equatorial slit, sometimes small piece of the denticle remains attached to the base of the conidium, (1–)2–5(–10) × 1.5–3 µm ( = 3.8 × 1.9 µm, n = 20). Sterile cells are brown, leaf-like, attached to the conidiophore.
Material examined: CHINA, Yunnan Province, Honghe Autonomous Prefecture, Honghe County, Honghe Hani Rice Terraces, on decaying stem of bamboo, 26 January 2021, R. Phookamsak, bn01 (KUN-HKAS 124566), living culture = KUNCC 22-12408; Honghe Hani Rice Terraces, on dead stem of bamboo, 26 January 2021, R. Phookamsak, bn11 (KUN-HKAS 124567), living culture = KUNCC 22-12409.
Notes: The nucleotide BLAST search of ITS region showed that Apiospora locuta-pollinis (KUNCC 22-12408 and KUNCC 22-12409) has high similarity with A. marii strain CBS 497.90, isolate A4, CPC 18904, CPC 18902, CBS 200.57, CBS 113535, Arthrinium sp. strain MFLUCC 16-0282, Fungal sp. BMP3011 (100%). The nucleotide BLAST search of LSU region showed that A. locuta-pollinis (KUNCC 22-12408 and KUNCC 22-12409) has high similarity with A. guizhouensis strain LC5318, A. locuta-pollinis isolate SICAUCC 22-0037, A. sacchari strain CBS 664.74, CBS 372.67, CBS 301.49, CBS 212.30, A. marii strain CBS 113535, A. biserialis isolate CS19-17 and A. guizhouensis strain KUMCC 20-0206 (100%). The nucleotide BLAST search of TEF1-α region showed that A. locuta-pollinis (KUNCC 22-12408) has high similarity with A. marii isolate GUCC 10214 (100%) and A. locuta-pollinis (KUNCC 22-12409) has high similarity with A. locuta-pollinis strain LC 11683, A. marii culture-collection CBS 113535, CBS 114803, GUCC 10254, GUCC 10227, A. locuta-pollinis strain LC 11688, LC 11689 (100%).
In our multigene phylogeny, two new strains (KUNCC 22-12408 and KUNCC 22-12409) shared a close relationship with Apiospora locuta-pollinis based on ML, MP, and BI analyses (Figure 1 and Figure 2). The base-pair comparison of ITS gene regions indicated that strain KUNCC 22-12408 is identical to strain KUNCC 22-12409 and other A. locuta-pollinis strains, except for a 1 bp difference with GUCC 10228. In the base-pair comparison of TEF1-α gene regions, strain KUNCC 22-12409 had no base pair differences with all the A. locuta-pollinis strains and strain KUNCC 22-12408 had a 2 bp difference (0.5%) with KUNCC 22-12409 and other strains of A. locuta-pollinis.
Morphological characteristics of our new strains (KUNCC 22-12408 and KUNCC 22-12409) were compared with the type strain of Apiospora locuta-pollinis (LC 11683) (Table 2). Both new strains are similar to the type strain of A. locuta-pollinis in having globose to subglobose conidia with hyaline equatorial rim, however they have larger conidia (10.5 × 10 and 9.6 × 8 vs. 7.1 × 6.4 μm) (Table 2). The conidiogenous cells of strain KUNCC 22-12409 are more similar to the type strain (LC 11683) in being subglobose to ampulliform to doliiform, but with smaller size (3.2 × 2.2 vs. 4.9 × 3.8 μm) (Table 2)., whereas the strain KUNCC 22-12408 has cylindrical to subcylindrical or ampulliform conidiogenous cells with larger size compared to other strains (6 × 3.5 µm) (Table 2). The morphological description of A. locuta-pollinis was based on cultures and its conidiophores were reduced to conidiogenous cells [61]. Thus, the comparisons of conidiomata and conidiophores characteristics between these strains were not possible. In addition, we found some morphological differences between two new strains including strain KUNCC 22-12408 which had longer conidiophore mother cells and conidiogenous cells, but shorter conidiophores compared to strain KUNCC 22-12409 (Table 2).

Table 2.
Morphological comparison among Apiospora locuta-pollinis strains.
Apiospora locuta-pollinis was previously isolated from hive-stored pollen of Brassica campestris in Hubei province, China [61], whereas the new strains KUNCC 22-12408 were isolated from decaying bamboo and KUNCC 22-12409 was isolated from dead bamboo in Yunnan province, China. Although there were some morphological variations among the new strains and the type strain of A. locuta-pollinis, the multigene phylogeny and DNA sequence comparisons (ITS and TEF1-α gene regions) did not provide the necessary support to delineate them as a distinct species. Therefore, we treated these strains as new records of A. locuta-pollinis. It is possible that the strain KUNCC 22-12408 is a new species due to the significant morphological and phylogenetic differences, which caused it to form a separated branch to other A. locuta-pollinis strains. Further taxonomic studies are needed to resolve their conspecific status.
3.3. Host and Geographical Distribution of Apiospora Species
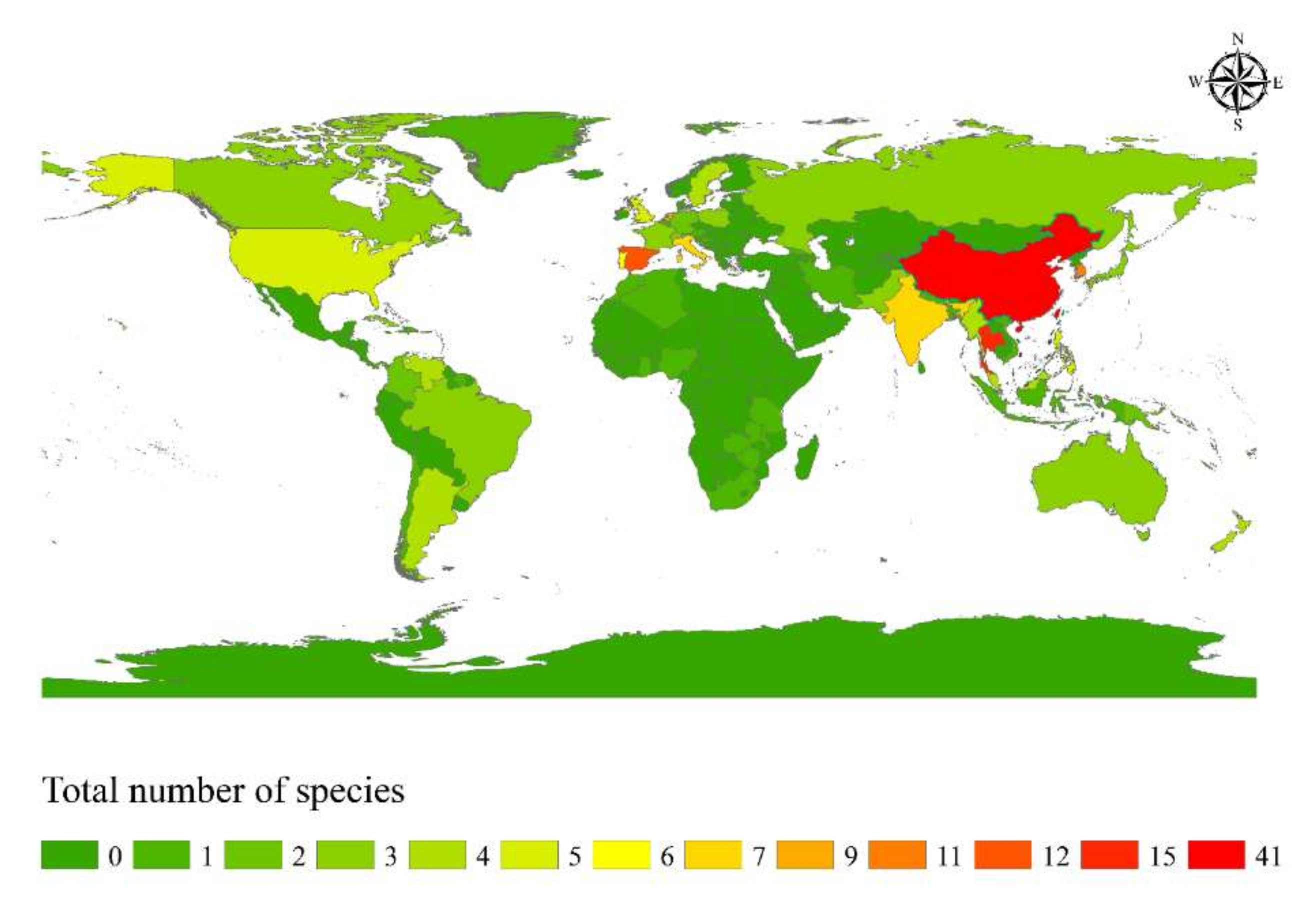
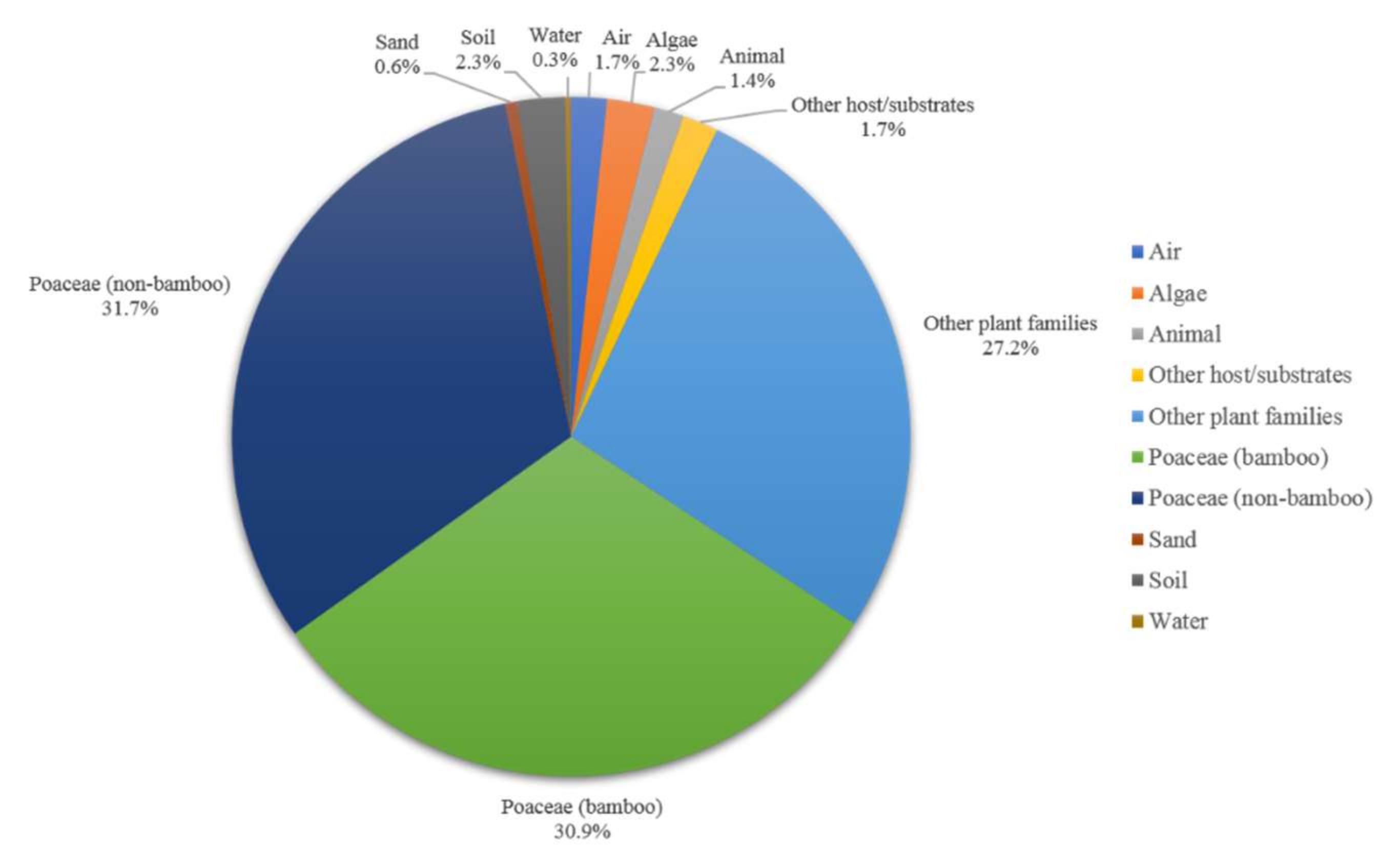
Based on species distribution data, Apiospora is widely distributed in temperate, subtropical, and tropical areas, including Africa, America, Asia, Australia, and Europe (Figure 5). The highest species diversity of Apiospora was found in Asia (e.g., China, India, Japan, Thailand) (Figure 5). However, the data reflect areas in which there have been reports of Apiospora species and may thus reflect hotspots of research, and not just species hotspots. Areas shown to be devoid of Apiospora species may just be areas that require further study. The host-substratum diversity of Apiospora species (Figure 6) showed that most species have been found exclusively associated with Poaceae (63%), including bamboo (31%) and non-bamboo (32%). The most common bamboo genera associated with Apiospora are Bambusa, Phyllostachys, and Arundinaria and the most common non-bamboo genera are Saccharum, Phragmites, and Arundo.
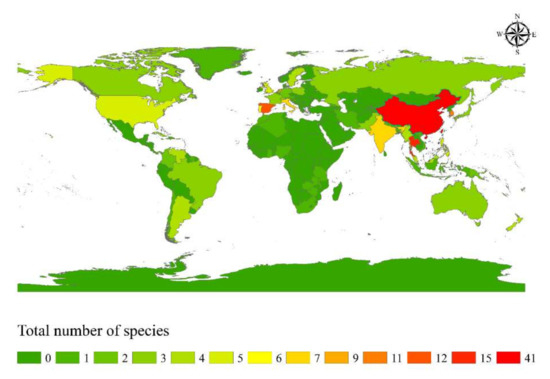
Figure 5.
Species diversity hotspot countries of Apiospora.
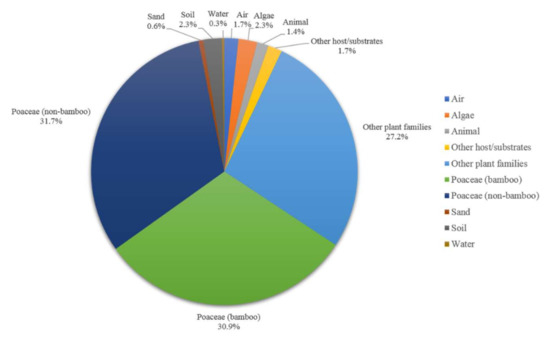
Figure 6.
Host-substratum diversity of Apiospora species.
4. Discussion
Our study provides better insight into interspecific and intraspecific variation in Apiospora, particularly in the A. locuta-pollinis/marii clade. Phylogenetic analyses of four gene markers (ITS, LSU, TUB2, and TEF1-α) revealed the distinct relationships between A. marii, A. locuta-pollinis, and A. gaoyouensis (Figure 1 and Figure 2). Morphologically, they have similar conidia characteristics (globose to elongate ellipsoid in surface, lenticular in side view) with an overlapping size range: 7.2–7.5 µm diam. in Apiospora marii, 5.5–9 µm diam. in A. locuta-pollinis, and 5–8 µm diam. in A. gaoyouensis (Table 3).

Table 3.
Morphological comparison among Apiospora species in the A. locuta-pollinis/marii clade.
In the Apiospora marii clade, A. hispanica and A. mediterranea are not well-resolved (Figure 1 and Figure 2). Morphologically, A. marii and A. hispanica have overlapping sizes of conidia (7.2–7.5 × 6.1–6.5 vs. 7.5–8.5 × 6.2–7.6 µm) (Table 3), whereas A. mediterranea has larger conidia (9–9.5 × 7.5–9 µm) (Table 3). The base-pair difference of ITS and TUB2 sequence data indicated that they are consistent. However, the LSU sequence data of A. hispanica and A. mediterranea are in short length (320 base pairs) and their TEF1-α sequence data were lacking. The morphological reexamination and molecular data of the type specimens of A. hispanica and A. mediterranea are required to confirm a putative synonymy.
In the Apiospora locuta-pollinis clade, two strains of A. marii CBS 113535 and CBS 114803 clustered together (Figure 2). Pintos et al. [5] and Garin et al. [62] also reported that A. marii CBS 114803 had a well-supported lineage distant from the A. marii clade. Tian et al. [12] synonymized A. pseudomarii GUCC 10228 as A. locuta-pollinis based on morphology and phylogeny. Crous and Groenewald [3] provided the sequence data of these strains in which CBS 113535 was isolated from oats in Sweden and CBS 114803 was isolated from the culm of Arundinaria hindsi in Hongkong. Thus, we treated these strains as A. locuta-pollinis based on the phylogenetic evidence.
TUB2 and TEF1-α gene regions are essential phylogenetic markers for accurate identification of Apiospora species [3,4,5,9,38]. Most of the recent studies have used multigene phylogenetic analyses of integrated ITS, LSU, TUB2, and TEF1-α sequence data for Apiospora species delineation [2,5,10,11,12,24,25,38]. However, our phylogenetic analysis based on the integrated ITS, LSU, and TEF1-α sequence data also provided the necessary resolution to distinguish species of Apiospora (Figure S1). In addition, the TEF1-α gene region could support the species delineation between A. marii and A. locuta-pollinis. As they have a 10 bp difference (2.3%) in the TEF1-α gene region, whereas no difference was found in TUB2 gene region. It seems that the TUB2 gene region is uninformative for the separation of species in the A. locuta-pollinis/marii clade. Nevertheless, with the lack of TUB2 sequences in our new strains, the TEF1-α gene region was not enough to resolve their placements within A. locuta-pollinis lineage. We suggest that TUB2 gene and other protein-coding genes, such as RPB2, should be included for further phylogenetic analysis to confirm their actual identity and placements.
Our study also revealed significant morphological variation among Apiospora locuta-pollinis strains. This result was also observed for other Apiospora species. For instance, A. yunnana, introduced by Dai et al. [8], based on a sexual morph on bamboo culms, had larger conidia in cultures (15.5–26.5 μm diam.), compared to the strains CFCC 52311, CFCC 5231 which were isolated and described directly from bamboo culms (10–16 μm diam.) [9]. Apiospora pseudoparenchymatica, introduced by Wang et al. [4], and was isolated from living leaves of bamboo and described by its asexual morph. Feng et al. [11] found a new record of A. pseudoparenchymatica from decaying bamboo culms and noted the significant difference in the characteristics of conidiophores and conidiogenous cells of the new strain, GZCC 20–0117 (on WA) compared to the type specimen, LC 8173 (on PDA and MEA) [4]. The significant variation in morphology might be due to the differences in substrates (natural substrates or cultures), growth conditions, hosts, and habitats. This observation makes our finding all the more valuable, we found both new strains from the same host substrate and habitat. The only difference is strain KUNCC 22-12408, which was from the decaying state and strain KUNCC 22-12409 was from the dead state of bamboo. Therefore, our study highlighted the great impact of environmental factors on morphological variation.
Geographically, Asia was found to be home to greatest diversity of Apiospora (Figure 5) and is likely the result of the rich diversity of bamboo genera/species, especially those in China [63]. More extensive sampling of these hosts will surely result in the discovery of additional new species. Although the host preference of Apiospora is the family Poaceae, there remains a number of species which have been found on other plant families (Figure 6), such as A. euphorbiae (on Euphorbiaceae, Lauraceae, Pinaceae, Zingiberaceae) [64], and A. jiangxiensis (on Lauraceae, Primulaceae, Theaceae) [4]., On the contrary, some species seem to be host-specific, such as A. pterosperma which has only been found on Cyperaceae (Lepidosperma and Machaerina) [3], and A. rhododendri has only been reported from Ericaceae (Rhododendron) [64]. Some species are ubiquitous, with a diverse range of hosts and habitats (e.g., A. arundinis, A. marii, A. rasikravindrae, A. serenensis) [2,3,4,5,8,12,64]. It is noteworthy that the asexual morph is more frequently discovered from different ecological habitats, and it is possible they are the cause of certain plant diseases [2,62]. For example, A. arundinis which has been isolated from soil, water, and numerous plant hosts [4,64], is also reported as causing some plant diseases, such as brown culm streak of Phyllostachys praecox [65] and the leaf spot of rosemary (Salvia rosmarinus) [66]. Apiospora marii has been isolated from air, sand, and different plant hosts [3,5], also reported as causing die-back of olives (Olea europaea) [62].
5. Conclusions
Based on the rate of discovery, their diverse host ranges and different life-styles, the species number of Apiospora is likely to continue to increase in the future [39,48]. There are still a number of regions that have remained unstudied in terms of Apiospora, which will likely further add to the current list of species within this genus. A comprehensive survey of these unknown regions along with a polyphasic taxonomic study of Apiospora is necessary, especially focusing on Poaceae species, will enable a better understanding of host relationships and the ecological significance of this group of fungi.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/d14110918/s1, Figure S1: Phylogenetic tree retrieved from RAxML analyses of a combined ITS, LSU and TEF1-α sequence data of Apiospora, and other related taxa in the family Apiosporaceae. Bootstrap support values for ML equal or greater than 60% and Bayesian posterior probabilities greater than 0.90 are indicated at the nodes as ML/PP. The ex-type strains are in bold. The new species are in red and new record and new combination species are in blue. The tree is rooted to Sporocadus trimorphus (CBS 114203).
Author Contributions
Conceptualization, J.M. and R.P.; data curation, J.M.; formal analysis, J.M., D.S.T. and S.X.; funding acquisition, J.X. and S.L.; investigation, J.M., R.P., S.X. and Q.L.; methodology, J.M. and R.P.; project administration, J.M. and R.P.; resources, J.M., R.P., S.X. and Q.L.; supervision, J.X., J.K. and S.L.; validation, J.M., R.P., D.J.B. and S.L.; writing—original draft, J.M. and R.P.; writing—review and editing, J.M., R.P., D.S.T., D.J.B., P.E.M. and J.K. All authors have read and agreed to the published version of the manuscript.
Funding
The research was supported by Post–Doctoral Fellowship 2022 for Chiang Mai University (Grant Nos. R000030592 and R000031743). This research study is also supported by Yunnan Provincial Science and Technology Department, Key Project (Grant No. 202101AS070045), the Project “Conservation of characteristic woody germplasm resources and construction of woody circular agriculture in the dry and hot valley of Honghe River” (Grant No. 202003AD150004) and NSFC-CGIAR Project “Characterization of roots and their associated rhizosphere microbes in agroforestry systems: ecological restoration in high-phosphorus environment” (Grant No. 31861143002).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Data can be found within the manuscript.
Acknowledgments
J.M. would like to thank the Post–Doctoral Fellowship 2022 from Chiang Mai University, Thailand (Grant No. R000030592). R.P. would like to thank the Post–Doctoral Fellowship 2022 from Chiang Mai University, Thailand (Grant No. R000031743). J.M. is grateful to Erandi Yasanthika and Areerat Manowong for their assistance during this research. Shaun Pennycook from Landcare Research, Auckland, New Zealand, is thanked for advising on the taxon name. We also acknowledge the Biology Experimental Center, Germplasm Bank of Wild Species, Kunming Institute of Botany, Chinese Academy of Sciences for providing the molecular laboratory facilities.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Hyde, K.D.; Norphanphoun, C.; Maharachchikumbura, S.S.N.; Bhat, D.J.; Jones, E.B.G.; Bundhun, D.; Chen, Y.J.; Bao, D.F.; Boonmee, S.; Calabon, M.S.; et al. Refined families of Sordariomycetes. Mycosphere 2020, 11, 305–1059. [Google Scholar] [CrossRef]
- Pintos, Á.; Alvarado, P. Phylogenetic delimitation of Apiospora and Arthrinium. Fungal Syst. Evol. 2021, 7, 197–221. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Groenewald, J.Z. A phylogenetic re-evaluation of Arthrinium. IMA Fungus 2013, 4, 133–154. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Tan, X.M.; Liu, F.; Cai, L. Eight new Arthrinium species from China. MycoKeys 2018, 1, 1–24. [Google Scholar] [CrossRef]
- Pintos, Á.; Alvarado, P.; Planas, J.; Jarling, R. Six new species of Arthrinium from Europe and notes about A. caricicola and other species found in Carex spp. hosts. MycoKeys 2019, 49, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, R.; Kulkarni, G.; Sonawane, M.S.; Shouche, Y.S. A new endophytic species of Arthrinium (Apiosporaceae) from Jatropha podagrica. Mycoscience 2014, 55, 118–123. [Google Scholar] [CrossRef]
- Dai, D.Q.; Jiang, H.B.; Tang, L.Z.; Bhat, D.J. Two new species of Arthrinium (Apiosporaceae, Xylariales) associated with bamboo from Yunnan, China. Mycosphere 2016, 7, 1332–1345. [Google Scholar] [CrossRef]
- Dai, D.Q.; Phookamsak, R.; Wijayawardene, N.N.; Li, W.J.; Bhat, D.J.; Xu, J.C.; Taylor, J.E.; Hyde, K.D.; Chukeatirote, E. Bambusicolous fungi. Fungal Divers. 2017, 82, 1–105. [Google Scholar] [CrossRef]
- Jiang, N.; Li, J.; Tian, C.M. Arthrinium species associated with bamboo and reed plants in China. Fungal Syst. Evol. 2018, 2, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Senanayake, I.C.; Bhat, J.D.; Cheewangkoon, R.; Xie, N. Bambusicolous Arthrinium Species in Guangdong Province, China. Front. Microbiol. 2020, 11, 602773. [Google Scholar] [CrossRef]
- Feng, Y.; Liu, J.K.J.; Lin, C.G.; Chen, Y.Y.; Xiang, M.M.; Liu, Z.Y. Additions to the genus Arthrinium (Apiosporaceae) from bamboos in China. Front. Microbiol. 2021, 12, 661281. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.G.; Karunarathna, S.C.; Mapook, A.; Promputtha, I.; Xu, J.C.; Bao, D.F.; Tibpromma, S. One new species and two new host records of Apiospora from bamboo and maize in Northern Thailand with thirteen new combinations. Life 2021, 11, 1071. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.; Luo, F.; Zhang, H.; Fang, X.; Zhu, T.; Li, S. First report of Arthrinium kogelbergense causing blight disease of Bambusa intermedia in Sichuan Province, China. Plant Dis. 2021, 105, 214. [Google Scholar] [CrossRef] [PubMed]
- Mavragani, D.C.; Abdellatif, L.; McConkey, B.; Hamel, C.; Vujanovic, V. First report of damping-off of durum wheat caused by Arthrinium sacchari in the semi-arid Saskatchewan fields. Plant Dis. 2007, 91, 469. [Google Scholar] [CrossRef]
- Aiello, D.; Gulisano, S.; Gusella, G.; Polizzi, G.; Guarnaccia, V. First report of fruit blight caused by Arthrinium xenocordella on Pistacia vera in Italy. Plant Dis. 2018, 102, 1853. [Google Scholar] [CrossRef]
- Zhang, Z.F.; Liu, F.; Zhou, X.; Liu, X.Z.; Liu, S.J.; Cai, L. Culturable mycobiota from Karst caves in China, with descriptions of 20 new species. Persoonia 2017, 39, 1–31. [Google Scholar] [CrossRef]
- Singh, S.M.; Yadav, L.S.; Singh, P.N.; Hepat, R.; Sharma, R.; Singh, S.K. Arthrinium rasikravindrii sp. nov. from Svalbard, Norway. Mycotaxon 2013, 122, 449–460. [Google Scholar] [CrossRef]
- Das, K.; Lee, S.Y.; Choi, H.W.; Eom, A.H.; Cho, Y.J.; Jung, H.Y. Taxonomy of Arthrinium minutisporum sp. nov., Pezicula neosporulosa, and Acrocalymma pterocarpi: New records from soil in Korea. Mycobiology 2020, 48, 450–463. [Google Scholar] [CrossRef]
- Luo, Z.L.; Hyde, K.D.; Liu, J.K.; Maharachchikumbura, S.S.N.; Jeewon, R.; Bao, D.F.; Bhat, D.J.; Lin, C.G.; Li, W.L.; Yang, J.; et al. Freshwater Sordariomycetes. Fungal Divers. 2019, 99, 451–660. [Google Scholar] [CrossRef] [Green Version]
- Suryanarayanan, T.S. Fungal endosymbionts of seaweeds. In Biology of Marine Fungi; Raghukumar, C., Ed.; Springer: Dordrecht, The Netherlands, 2012; pp. 53–70. [Google Scholar]
- Hong, J.-H.; Jang, S.; Heo, Y.M.; Min, M.; Lee, H.; Lee, Y.; Lee, H.; Kim, J.J. Investigation of marine-derived fungal diversity and their exploitable biological activities. Mar. Drugs 2015, 13, 4137–4155. [Google Scholar] [CrossRef]
- Heo, Y.M.; Kim, K.; Ryu, S.M.; Kwon, S.L.; Park, M.Y.; Kang, J.E.; Hong, J.H.; Lim, Y.W.; Kim, C.; Kim, B.S.; et al. Diversity and ecology of marine Algicolous Arthrinium species as a source of bioactive natural products. Mar. Drugs 2018, 16, 508. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, M.S.; Oh, S.-Y.; Lee, S.; Eimes, J.A.; Lim, Y.W. Fungal diversity and enzyme activity associated with sailfin sandfish egg masses in Korea. Fungal Ecol. 2018, 34, 1–9. [Google Scholar] [CrossRef]
- Kwon, S.L.; Park, M.S.; Jang, S.; Lee, Y.M.; Heo, Y.M.; Hong, J.H.; Lee, H.; Jang, Y.; Park, J.H.; Kim, C.; et al. The genus Arthrinium (Ascomycota, Sordariomycetes, Apiosporaceae) from marine habitats from Korea, with eight new species. IMA Fungus 2021, 12, 13. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.L.; Cho, M.; Lee, Y.M.; Kim, C.; Lee, S.M.; Ahn, B.J.; Lee, H.; Kim, J.J. Two unrecorded Apiospora species isolated from marine substrates in Korea with eight new combinations (A. piptatheri and A. rasikravindrae). Mycobiology 2022, 50, 46–54. [Google Scholar] [CrossRef]
- He, Y.; Zhang, Z. Diversity of organism in the Usnea longissima lichen. Afr. J. Microbiol. Res. 2012, 6, 4797–4804. [Google Scholar]
- Crous, P.W.; Wingfield, M.J.; Le Roux, J.J.; Richardson, D.M.; Strasberg, D.; Shivas, R.G.; Alvarado, P.; Edwards, J.; Moreno, G.; Sharma, R. Fungal Planet description sheets: 371–399. Pers. Mol. Phylogeny Evol. Fungi 2015, 35, 264. [Google Scholar] [CrossRef]
- Rai, M.K. Mycosis in man due to Arthrinium phaeospermum var. indicum. First case report. Mycoses 1989, 32, 472–475. [Google Scholar]
- Zhao, Y.M.; Deng, C.R.; Chen, X. Arthrinium phaeospermum causing dermatomycosis, a new record of China. Acta Mycol. Sin. 1990, 9, 232–235. [Google Scholar]
- De Hoog, G.S.; Guarro, J.; Gené, J.; Figueras, M.J. Atlas of Clinical Fungi, 2nd ed.; CBS: Utrecht, The Netherlands, 2000; 1126p. [Google Scholar]
- Shrestha, P.; Ibáñez, A.B.; Bauer, S.; Glassman, S.I.; Szaro, T.M.; Bruns, T.D.; Taylor, J.W. Fungi isolated from Miscanthus and sugarcane: Biomass conversion, fungal enzymes, and hydrolysis of plant cell wall polymers. Biotechnol. Biofuels 2015, 8, 1. [Google Scholar] [CrossRef] [Green Version]
- Kunze, G. Zehn neue Pilzgattungen. Mykol. Hefte 1817, 1, 1–18. [Google Scholar]
- Ellis, M.B. Dematiaceous Hyphomycetes: IV. Mycol. Pap. 1963, 29, 1–33. [Google Scholar]
- Seifert, K.; Morgan-Jones, G.; Gams, W.; Kendrick, B. The Genera of Hyphomycetes. [CBS Biodiversity Series 9]; CBSKNAW Fungal Biodiversity Centre: Utrecht, The Netherlands, 2011; pp. 1–1997. [Google Scholar]
- McNeill, J.; Barrie, F.R.; Buck, W.R.; Demoulin, V.; Greuter, W.; Hawksworths, D.L.; Herendeen, P.S.; Knapp, S.; Marhold, K.; Prado, J.; et al. International code of nomenclature for algae, fungi and plants (Melbourne Code) adopted by the Eighteenth International Botanical Congress Melbourne, Australia, July 2011. Regnum Veg. 2012, 154, 1–140. [Google Scholar]
- Senanayake, I.C.; Maharachchikumbura, S.S.N.; Hyde, K.D.; Bhat, J.D.; Jones, E.B.G.; McKenzie, E.H.C.; Dai, D.Q.; Daranagama, D.A.; Dayarathne, M.C.; Goonasekara, I.D.; et al. Towards unraveling relationships in Xylariomycetidae (Sordariomycetes). Fungal Divers. 2015, 73, 73–144. [Google Scholar] [CrossRef]
- Réblová, M.; Miller, A.N.; Rossman, A.Y.; Seifert, K.; Crous, P.; Hawksworth, D.; Adel-Wahab, M.A.; Cannon, P.F.; Daranagama, D.A.; De Beer, Z.W.; et al. Recommendations for competing sexual-asexually typified generic names in Sordariomycetes (except Diaporthales, Hypocreales, and Magnaporthales). IMA Fungus 2016, 7, 131–153. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, H.B.; Hyde, K.D.; Doilom, M.; Karunarathna, S.C.; Xu, J.C.; Phookamsak, R. Arthrinium setostromum (Apiosporaceae, Xylariales), a novel species associated with dead bamboo from Yunnan, China. Asian J. Mycol. 2019, 2, 254–268. [Google Scholar] [CrossRef]
- Phukhamsakda, C.; Nilsson, R.H.; Bhunjun, C.S.; de Farias, A.R.G.; Sun, Y.R.; Wijesinghe, S.N.; Raza, M.; Bao, D.-F.; Lu, L.; Tibpromma, S.; et al. The numbers of fungi: Contributions from traditional taxonomic studies and challenges of metabarcoding. Fungal Divers. 2022, 114, 327–386. [Google Scholar] [CrossRef]
- Species Fungorum. Available online: http://www.speciesfungorum.org (accessed on 1 September 2022).
- Senanayake, I.; Rathnayaka, A.; Marasinghe, D.; Calabon, M.; Gentekaki, E.; Lee, H.; Hurdeal, V.; Pem, D.; Dissanayake, L.; Wijesinghe, S.; et al. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. Mycosphere 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Index Fungorum. Available online: http://www.indexfungorum.org (accessed on 1 September 2022).
- Jayasiri, S.C.; Hyde, K.D.; Ariyawansa, H.A.; Bhat, J.; Buyck, B.; Cai, L.; Dai, Y.C.; Abd-Elsalam, K.A.; Ertz, D.; Hidayat, I.; et al. The Faces of Fungi database: Fungal names linked with morphology, phylogeny and human impacts. Fungal Divers. 2015, 74, 3–18. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: Cambridge, MA, USA, 1990; Volume 18, p. 7. [Google Scholar]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef] [Green Version]
- O’Donnell, K.; Kistler, H.C.; Cigelnik, E.; Ploetz, R.C. Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proc. Natl. Acad. Sci. USA 1998, 95, 2044–2049. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Bhunjun, C.S.; Niskanen, T.; Suwannarach, N.; Wannathes, N.; Chen, Y.J.; McKenzie, E.H.; Maharachchikumbura, S.S.N.; Buyck, B.; Zhao, C.-L.; Fan, Y.-G.; et al. The numbers of fungi: Are the most speciose genera truly diverse? Fungal Divers. 2022, 114, 387–462. [Google Scholar] [CrossRef]
- Samarakoon, M.C.; Hyde, K.D.; Maharachchikumbura, S.S.N.; Stadler, M.; Jones, E.B.G.; Promputtha, I.; Suwannarach, N.; Camporesi, E.; Bulgakov, T.S.; Liu, J.K. Taxonomy, phylogeny, molecular dating and ancestral state reconstruction of Xylariomycetidae (Sordariomycetes). Fungal Divers. 2022, 112, 1–88. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hall, T. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In Proceedings of the Nucleic Acids Symposium Series, London, UK, 2–6 September 1999; pp. 95–98. [Google Scholar]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES science gateway for inference of large phylogenetic trees. In Proceedings of the 2010 Gateway Computing Environments Workshop (GCE), New Orleans, LA, USA, 14 November 2010; IEEE: New Orleans, LA, USA, 2010; pp. 1–8. [Google Scholar]
- Nylander, J.A. MrModeltest 2. Program Distributed by the Author; Department of Systematic Zoology, Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhaxybayeva, O.; Gogarten, J.P. Bootstrap, Bayesian probability and maximum likelihood mapping: Exploring new tools for comparative genome analyses. BMC Genom. 2002, 3, 4. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Hoehna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [Green Version]
- Swofford, D.L. PAUP* Phylogenetic Analysis Using Parsimony * (and Other Methods); Version 4.0.; Sinauer Associates: Sunderland, UK, 2002. [Google Scholar]
- Hillis, D.M.; Bull, J.J. An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst. Biol. 1993, 42, 182–192. [Google Scholar] [CrossRef]
- Rambaut, A.; Drummond, A.J. FigTree: Tree Figure Drawing Tool; Institute of Evolutionary Biology, University of Edinburgh: Edinburgh, Scotland, 2012. [Google Scholar]
- Tennakoon, D.S.; Kuo, C.H.; Maharachchikumbura, S.S.N.; Thambugala, K.M.; Gentekaki, E.; Phillips, A.J.L.; Bhat, D.J.; Wanasinghe, D.N.; de Silva, N.I.; Promputtha, I.; et al. Taxonomic and phylogenetic contributions to Celtis formosana, Ficus ampelas, F. septica, Macaranga tanarius and Morus australis leaf litter inhabiting microfungi. Fungal Divers. 2021, 108, 1–215. [Google Scholar] [CrossRef]
- Zhao, Y.Z.; Zhang, Z.F.; Cai, L.; Peng, W.J.; Liu, F. Four new filamentous fungal species from newly-collected and hive-stored bee pollen. Mycosphere 2018, 9, 1089–1116. [Google Scholar] [CrossRef]
- Gerin, D.; Nigro, F.; Faretra, F.; Pollastro, S. Identification of Arthrinium marii as causal agent of olive tree dieback in Apulia (southern Italy). Plant Dis. 2020, 104, 694–701. [Google Scholar] [CrossRef] [PubMed]
- Lobovikov, M.; Paudel, S.; Ball, L.; Piazza, M.; Guardia, M.; Ren, H.; Russo, L.; Wu, J.Q. World Bamboo Resources: A Thematic Study Prepared in the Framework of the Global Forest Resources Assessment 2005; Food and Agriculture Organization of The United Nations: Rome, Italy, 2007; Volume 18. [Google Scholar]
- Farr, D.F.; Rossman, A.Y. Fungal Databases, U.S. National Fungus Collections, ARS, USDA. Available online: https://nt.ars-grin.gov/fungaldatabases/ (accessed on 1 September 2022).
- Chen, K.; Wu, X.Q.; Huang, M.X.; Han, Y.Y. First report of brown culm streak of Phyllostachys praecox caused by Arthrinium arundinis in Nanjing, China. Plant Dis. 2014, 98, 1274. [Google Scholar] [CrossRef]
- Bagherabadi, S.; Zafari, D.; Anvar, F.G. First report of leaf spot caused by Arthrinium arundinis on rosemary in Iran. J. Plant Pathol. 2014, 96, 4–126. [Google Scholar]
- Larrondo, J.V.; Calvo, M.A. Two new species of Arthrinium from Spain. Mycologia 1990, 82, 396–398. [Google Scholar] [CrossRef]
- Larrondo, J.V.; Calvo, M.A. New contributions to the study of the genus Arthrinium. Mycologia 1992, 84, 475–478. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).