In a previous study, we identified 45 phenotypically distinct, traditionally named landraces across five Amazon tributaries in northeastern Perú [
17]. We found that growers identified landraces using a range of traits and named them accordingly. We also observed distinctions among landraces even when they were planted in the same chacra. These observations suggested that genetic polymorphism is present, including divergence among yuca types. Our analyses of polymorphism at STR loci in this study confirmed the presence of genetic variation in yuca in the upper Amazon, and revealed previously unrecognized relationships between diversity, relatedness, traditional nomenclature, and geography.
4.1. Genetic Diversity
STR genotypes in our sample revealed extensive variation. Polymorphism was present at all 13 of the loci we examined, which combined to form 43 multilocus genotypes (MLGs) in the 43 landraces we collected. Thus, every landrace was unique. This pattern is similar to patterns observed in prior studies, which have consistently reported genetic distinctions between landraces. For instance, in an investigation of 596 wild, bitter, and sweet yuca landraces in the Brazilian Amazon, Alves-Pereira et al. [
43] identified 351 MLGs defined by 14 STRs. Thus, the majority (64%) were unique, although many (36%) were identical to at least one other type. Diversity in yuca has also been reported on smaller spatial scales. In an isolated Wayāpi village in French Guiana, Duputié et al. [
18] genotyped 10 STRs in 436 specimens, from 61 named landraces and found 383 MLGs. This pattern was noteworthy because it revealed that the average Wayāpi landrace harbored more than 6 MLGs. Thus, genetic diversity is found within named, Amazonian yuca landraces. Our sampling strategy, which focused on identifying different landraces, did not allow us to investigate the issue in our own sample.
Studies beyond the Americas have found genetic diversity and landrace differences as well. In an investigation of 11 villages in Uganda, Kizito et al. [
46] genotyped 288 specimens representing 93 yuca varieties, and encountered patterns similar to those reported by Duputié et al. [
18]. Different MLGs occurred within yuca varieties, and some MLGs were shared among them. However, on average, varieties were significantly differentiated. This was noteworthy because yuca was introduced to Africa from South America just 500 years ago, which might be expected to result in founder effects reducing diversity. Nonetheless, the two continents exhibited congruent patterns. Kizito et al.’s findings were reinforced by an analysis of eight loci in 522 bitter and sweet landraces from South America, Africa, and Oceania by Bradbury et al. [
44]. Bradbury et al. found 188 MLGs, including 86 MLGs across 203 landraces in Africa. Thus, many landraces had identical MLGs. In sum, while studies of STRs in yuca performed to date are not fully comparable due to their differing sampling schemes, they uniformly characterize yuca as a genetically diverse crop that harbors polymorphism throughout its range, including diversity both within and between landraces.
We found additional evidence of differentiation between landraces in patterns of heterozygosity and the distribution of variation within and across landraces and rivers. First, our tests for Hardy–Weinberg equilibrium revealed a significant deficit of heterozygosity at every locus (
p < 0.001 at 12 of 13 loci and
p < 0.012 for the thirteenth) (
Table 2). Departures from expectation in Hardy–Weinberg tests indicate that genotype frequencies are inconsistent with panmixia, with up- and downward departures from expectation indicating nonrandom mating. Reduced heterozygosity is a classic signature of population subdivision, with the subpopulations in our study being individual landraces [
40]. It occurs because genetic drift proceeds more rapidly within subpopulations than across the population as a whole, resulting in disproportionate losses of diversity in them. Second, our AMOVA tests revealed that the fraction of overall genetic variance accounted for by between-landrace differences (38.84%) was significantly greater than expected given overall diversity in the sample. This is predicted under subdivision because STR allele frequencies in different subpopulations diverge due to genetic drift.
Our field observations of chacras and growers’ cultivation techniques were consistent with our inferences about landrace differentiation based on STRs, suggesting that landraces represent subdivided populations. We found that landraces shown to us by growers were always phenotypically distinct and were readily distinguishable (
Figure 7). Phenotypic differences were evident even among landraces planted in the same chacra and thus could not be due to environmental effects. We further noted that growers are motivated to maintain separation for practical reasons. Different landraces are often preferred for different purposes, such as roasting, making casabe (yuca bread), brewing yuca beer (masato), and production of fariña (yuca meal). Admixture between landraces, if permitted, has the potential to produce plants with unpredictable phenotypes, reducing their usefulness. Growers’ tradition of propagation through cloning, rather than through cultivation of sexually produced seedings, which might be hybrids, ensures that barriers to admixture are high. Thus, observed patterns of phenotypic variation, genetic diversity, and cultivation practices all support the notion that landraces are distinct entities, and they are maintained by strong human influences.
Evidence that landraces are distinct raises questions about their origins. Our observations suggest that diversity in yuca is generated by sexual reproduction. We noted that although growers claimed to rely strictly on clonal reproduction rather than through propagation of seedlings, which might be undesirable hybrids, mature chacras usually contained some plants with flowers and fruits. Further, nearly all of the chacras we observed were interplanted, with two or more landraces growing side by side. Thus, sexual reproduction was occurring, and likely included hybridization between landraces. Seedlings produced this way were not fostered by growers. Their maturation lagged behind the main crop, so they would require extra attention, and they were treated as weeds. However, we speculate that seedlings are occasionally adopted when they show desirable traits, and may come into permanent cultivation. We also noted that abandoned chacras persist ferally, and sexual reproduction in them was likely unrestricted. This raises the possibility that growers augment or even fully establish new chacras with sexually produced feral plants.
In addition to revealing distinctions between landraces, genotypes in our sample revealed an unexpected pattern: an excess of within-landrace heterozygosity. Although heterozygosity across our sample as a whole was lower than expected, which was consistent with population subdivision, heterozygosity within landraces was higher than expected given the variation they did contain. Specifically, the proportion of loci that were heterozygous (Hsobs) was significantly greater (p < 0.001) than the proportion expected under random mating (Hsexp) in 39 of 43 of our sampled landraces. Moreover, the mean observed value was nearly twice the expectation. In addition, the results of the AMOVA indicated that the fraction of variance accounted for by within-landrace diversity, 58.11%, was the largest single contributor to overall genetic variance, and was highly significant statistically (p < 0.001). This suggested that heterozygous plants are, through some mechanism, favored in yuca chacras.
Excess heterozygosity in within landraces is consistent with the long-held hypothesis that heterozygotes have fitness advantages [
48,
49]. In the case of yuca, these could arise as the result of natural selection imposed by the environment, artificial selection imposed by growers, or both. Evidence of heterozygote advantage has been reported in genes mediating a range of bioprocesses, from fat storage to pesticide resistance [
49]. A striking number participate in vertebrate immunity, a pattern attributed to heterozygote advantages in detecting pathogens [
49]. Our STR data do not pinpoint specific genes influencing seedling or landrace survivorship. However, they do suggest that such genes are present because the STRs are unlikely to be under selective pressure themselves. Landraces’ high heterozygosity must be due to breeding regimes favoring hybridization. Given past findings, an evident possibility is that heterozygosity in genes encoding yuca’s immune system provides advantages in resisting pathogens. Another possibility is that heterozygotes exhibit phenotypes preferred by growers, such as enhanced productivity, and are thus more likely to persist as the result of human imposed pressure—artificial selection.
4.2. Relationships between Landraces
In a previous study, we found substantial phenotypic variation in our sample, even among landraces collected from the same chacra. However, we found little evidence of landrace clustering in principal components analyses, which can reflect the presence of divergent lineages. Nonetheless, we could not rule out the presence of population structure with respect to genetics, because it can occur even when phenotypic differences are not evident. This phenomenon, cryptic population structure, is common and well documented in natural populations [
50].
In contrast to phenotypes, genotypes in our sample did show evidence of phylogenetic structure. However, it was weak. In the neighbor joining tree, three clades supported by 100% of bootstrap replicates were present, but the branches separating them were short (
Figure 4). Thus, the differences between the three clades were consistent, but small. In addition, while the presence of additional subclades was supported with bootstrap values of 51% to 88%, the subclades always contained two lineages, as opposed to defining more inclusive groups. The tree topology also suggested that eight landraces originating from the Orosa river may form a subclade, but it was not well supported statistically (<50% of bootstrap replicates). The results of our principal components analysis were consistent with these findings (
Figure 6). They revealed that the majority of landraces formed a single tight cluster, rather than separate clusters as would be expected if clades are highly differentiated. In addition, several landraces from the Orosa river showed evidence of divergence from the main cluster, suggesting that they may be separate, but they were dispersed and did not form a distinct group.
A key observation in our study was that landraces have traditional names, which often reflect their phenotypes. Moreover, some traditional names are common across our study region. For example, we observed chacras planted with amarilla at multiple locations. Señorita was similarly common. This suggested that landrace name might reflect genetic relatedness, with landraces having the same name being more closely related. If so, it would imply that landraces are not just maintained locally, they are maintained across broader geographic ranges. However, we found little evidence that landraces’ names reflected their relatedness. Across the 28 specimens with names occurring more than once in our sample, 27 were most closely related to a landrace with a different name (
Table 7). For instance, of the six amarilla specimens we collected, the closest relative of each was a landrace with a different name. Our first amarilla exemplar was most closely related to a landrace named piririca, the second was most closely related to arpón, and so on. Across the entirety of our sample, the sole exception to the pattern was a specimen named motelillo collected on the Itaya river, which was most similar to a specimen named motelillo from the Pintuyacu river, more than 200 km away via river and 60 km direct distance. Thus, in our sample, landrace name was a poor indicator of genetic similarity. The explanation for this finding is not clear. Resolving it will require an investigation of growers’ naming conventions, such as whether they incorporate phenotypic information, whether they change over time, and whether names are retained when landraces are transported from location to location.
4.3. Geography and Population Structure
The geographical distribution of populations is frequently a factor shaping their genetic relationships [
51]. For instance, populations often exhibit isolation by distance, in which populations near each other are more similar genetically than are populations far apart. Populations can exhibit phylogeographic structure as well, with geographic structure shaping not just genetic distance but also phylogenetic relationships. An underlying factor in both of these phenomena is migration. Isolation by distance and phylogeographic structure can only emerge if migration rates between populations are sufficiently low to allow differentiation to occur. This raises questions about the relationships between genetic variation and geography in our yuca sample. On the one hand, the presence of distinct landraces and the geographic distance between chacras, which exceeded 150 km direct distance and 325 km river distance in some cases, suggested that regional differentiation was likely. On the other, the region’s human residents travel regularly, which may result in yuca migration. Residents often transport yuca from place to place in the process of moving their households or trading, for instance (
Figure 8). If sufficiently common, this might have prevented the evolution of regional differences.
The associations we found between genetic distance, physical distance, and river of origin supported the presence of population structure with respect to geography. However, the structuring was weak (
Figure 5). The presence of isolation by distance was strongly supported by Mantel tests, which revealed highly significant (
p < 0.001) associations between genetic distance and both measures of geographic distance. These had slopes of
β = 5.14 × 10
−4 and
β = 2.31 × 10
−4 for direct and river distance, respectively, which translated to 10% increases in genetic distance over ~190 km and ~425 km. However, their correlation coefficients (Mantel’s
r) were low, 0.12 and 0.20. Thus, while they were significantly associated, geographic and genetic distance were poor indicators of one another. The weakness of population structure with respect to geography was also reflected in the results of the AMOVA. In that analysis, differences between rivers were the lowest contributor to overall variance (3.05%). Differences between landraces on the same river (38.84%) and within landraces (58.11%) were higher. These patterns suggested that the yuca population in the region is not panmictic. However, the barriers to regional admixture are low. This situation can emerge through a variety of processes. For instance, it can occur following recent, rapid population dispersals, when subpopulations have had little time to differentiate. It can also occur when migration rates between subpopulations are high but not unlimited. Our data do not indicate which factor is responsible. Yuca’s domestication within the last several thousand years suggests that recent, rapid dispersal did occur. Further, the long-term frequency of human travel by both foot and canoe suggests that regional migration rates may have been high. The increasing accessibility of motorboats suggests that recent migration rates may be particularly high today. These possibilities are not mutually exclusive. Thus, we conclude that migration rates have been high, but we have little information about the relative importance of factors affecting it.
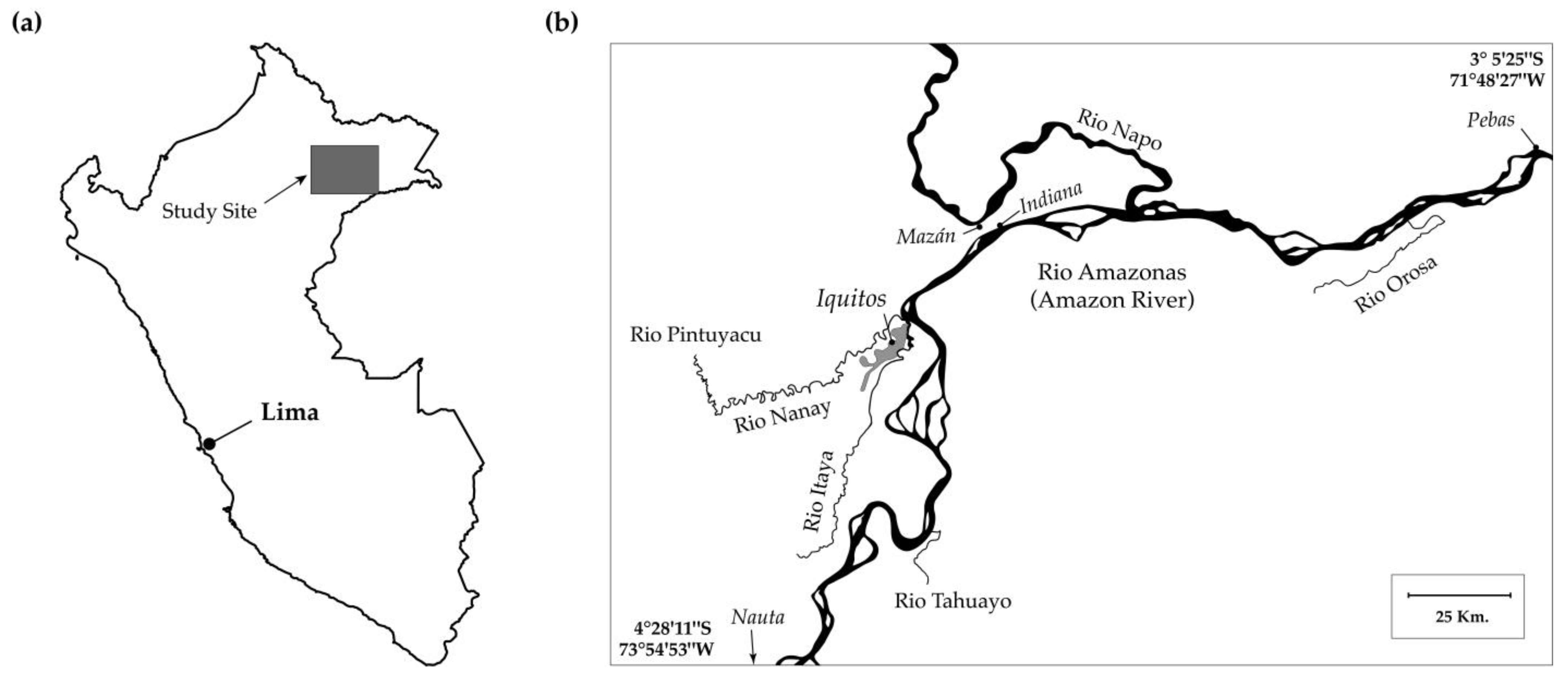
The results of the DAPC shed further light on the relationship between geography and population structure. It supported the results of the Mantel and AMOVA tests, suggesting that that spatial structure is present but weak. In the DAPC, the clusters representing the Pintuyacu and Nanay rivers overlapped extensively. This is consistent with the Pintuyacu being a tributary of the Nanay, such that they are expected to harbor related yuca landraces. Similarly, the cluster representing the Orosa river was distant from the clusters representing the other rivers, which is consistent with its location, more than 100 km away by river (
Figure 1). However, patterns of clustering were not always consistent with geography. For instance, the Itaya river cluster overlapped extensively with that of the Tahuayo, but it did not overlap with the Nanay. This is inconsistent with geography because the Nanay and Itaya enter the main channel of the Amazon within 5 km of each other, and thus would be expected to share more variation than either does with the Tahuayo, which is more than 55 km away by river.
While they did not have significance values attached to them, the pairwise genetic distances we observed between rivers recapitulated the results of the DAPC, suggesting that geography and population structure are related, but weak. Some differences between rivers were consistent with their geography. For instance, the genetic distance between the Nanay and Pintuyacu rivers was ~0.0 for all three measures (GST, G′ST, and D), which is consistent with the Pintuyacu being a direct tributary of the Nanay. This was reflected in the overlap between the Nanay and Pintuyacu in the DAPC. However, other distances were inconsistent with their geography. For instance, GST and G′ST between the Itaya and the Tahuayo were lower than between the Itaya and the Nanay, despite the Itaya and Nanay’s proximity (<5 km) relative to the Tahuayo, 55 km away.
Our observations of human travel likely explain the weak relationships we found between genetic variation and geography at our study site. We found that people in our study region, like people throughout Amazonía, often travel. The use of canoes is ubiquitous among populations living near rivers and other water bodies, allowing travel with heavy loads such as passengers, personal belongings and goods, and harvests. River travel certainly occurs, including the transport of yuca landraces for personal use, sharing with family, and occasionally exchange with strangers. The clonal propagation of yuca makes this straightforward because it requires only stem cuttings. We regularly observed travelers carrying yuca cuttings to establish new chacras (
Figure 8). Travel overland is also common. While rainforest terrain is stereotypically viewed as impenetrable, its inhabitants are familiar with regional topography and can travel for days on established trails. Overland travel does not allow the transport of the heavy loads possible with canoes. However, overland distances between locations can be substantially shorter than travel by river, and can be covered more quickly. In addition, yuca cuttings are light (usually stem pieces roughly 5 cm in diameter and 1.5 m long) and easily carried. Therefore, we hypothesize that both river and overland travel contribute to yuca’s migration, reducing the effects of geography on genetic differentiation, but not eliminating it. Taken together, these findings suggest that while husbandry practices maintain the distinctness of landraces, human travel obscures their geographic relationships.