Chemical Space Exploration and Machine Learning-Based Screening of PDE7A Inhibitors
Abstract
1. Introduction
2. Results and Discussion
2.1. Chemical Information Analysis
2.2. Murcko Scaffold Analysis
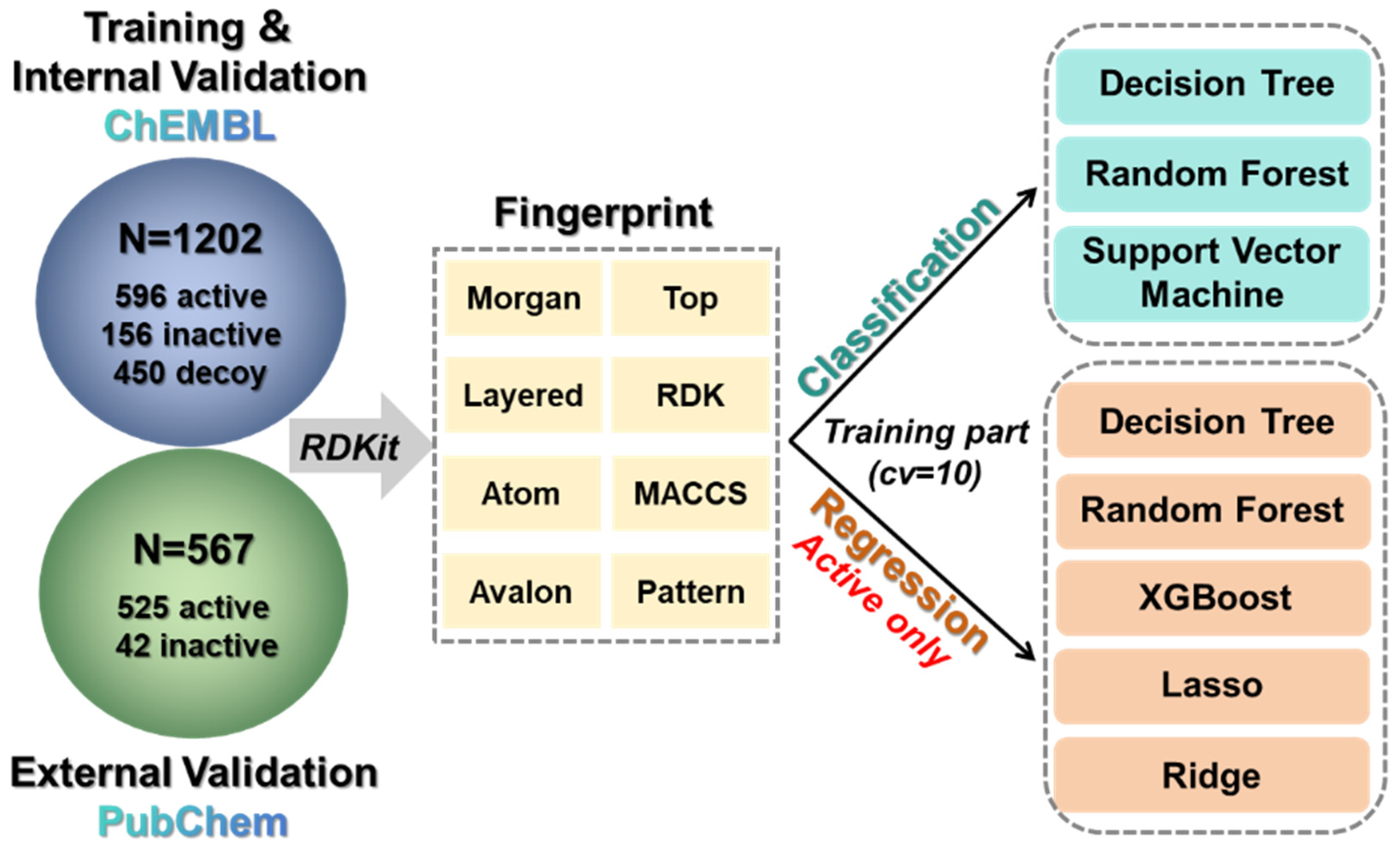
2.3. Development and Characterization of Machine Learning Models
2.4. Interpretable Machine Learning Analysis
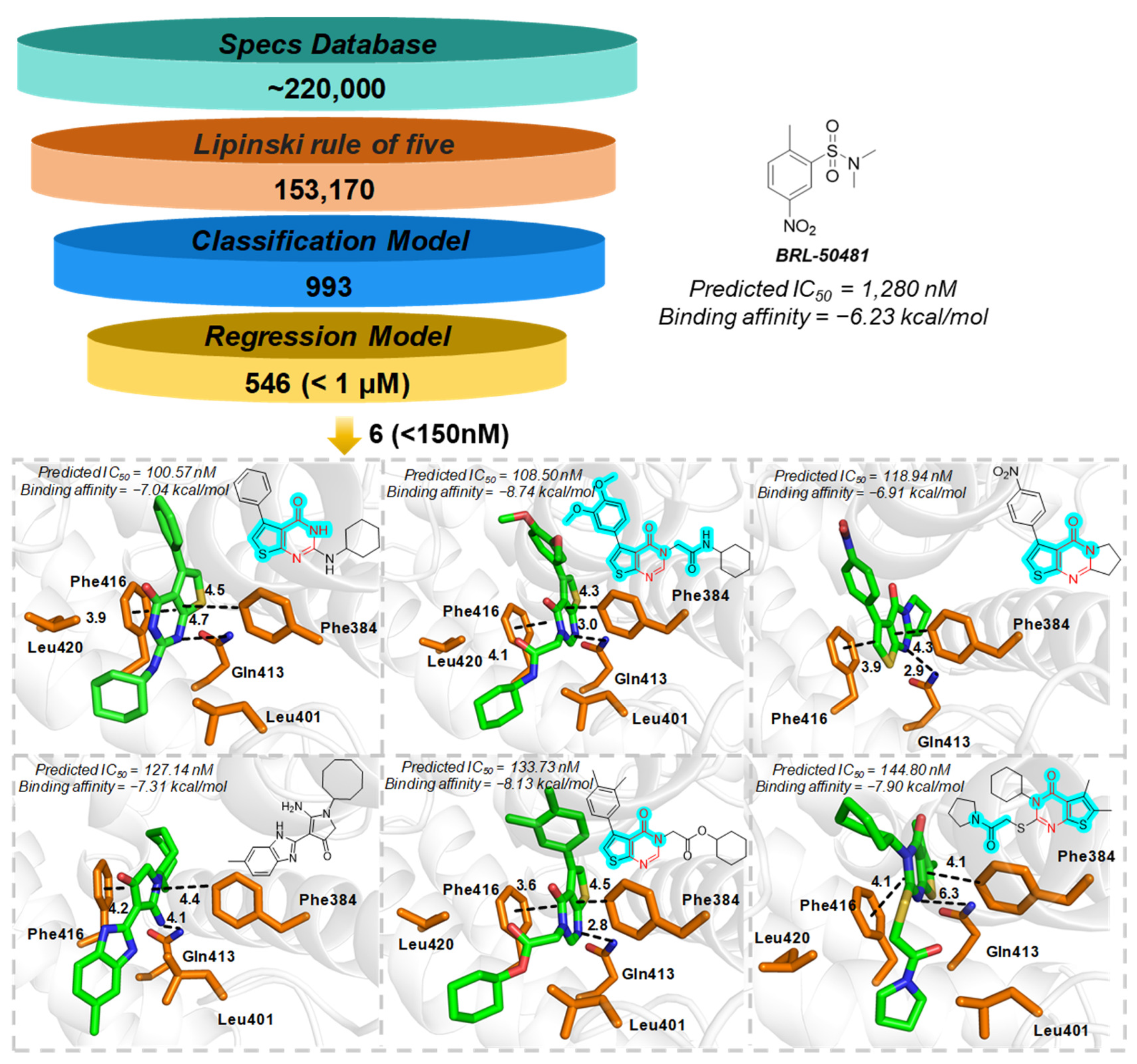
2.5. PDE7A Inhibitor Screening
3. Materials and Methods
3.1. Data Preparation
3.2. Molecular Features and Fingerprint Calculation
3.3. Machine Learning Model Construction
3.4. Model Evaluation
3.5. Feature Importance Analysis
3.6. Molecule Docking
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bender, A.T.; Beavo, J.A. Cyclic nucleotide phosphodiesterases: Molecular regulation to clinical use. Pharmacol. Rev. 2006, 58, 488–520. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.T. Targeting phosphodiesterases (PDEs) for treatment of CNS diseases. Curr. Pharm. Des. 2015, 21, 271–273. [Google Scholar] [CrossRef] [PubMed]
- Hsien Lai, S.; Zervoudakis, G.; Chou, J.; Gurney, M.E.; Quesnelle, K.M. PDE4 subtypes in cancer. Oncogene 2020, 39, 3791–3802. [Google Scholar] [CrossRef] [PubMed]
- Peng, T.; Gong, J.; Jin, Y.; Zhou, Y.; Tong, R.; Wei, X.; Bai, L.; Shi, J. Inhibitors of phosphodiesterase as cancer therapeutics. Eur. J. Med. Chem. 2018, 150, 742–756. [Google Scholar] [CrossRef]
- Manni, S.; Mauban, J.H.; Ward, C.W.; Bond, M. Phosphorylation of the cAMP-dependent protein kinase (PKA) regulatory subunit modulates PKA-AKAP interaction, substrate phosphorylation, and calcium signaling in cardiac cells. J. Biol. Chem. 2008, 283, 24145–24154. [Google Scholar] [CrossRef]
- Li, H.; Zuo, J.; Tang, W. Phosphodiesterase-4 inhibitors for the treatment of inflammatory diseases. Front. Pharmacol. 2018, 9, 1048. [Google Scholar] [CrossRef]
- Chen, L.Z.; Jiang, Z.Q.; Yang, L.F.; Fang, Y.; Lu, S.W.; Akakuru, O.U.; Huang, S.S.; Li, J.; Ma, S.Y.; Wu, A.G. HPDA/Zn as a CREB inhibitor for ultrasound imaging and stabilization of atherosclerosis plaque. Chin. J. Chem. 2023, 41, 199–206. [Google Scholar] [CrossRef]
- Kim, N.J.; Baek, J.H.; Lee, J.; Kim, H.; Song, J.K.; Chun, K.H. A PDE1 inhibitor reduces adipogenesis in mice via regulation of lipolysis and adipogenic cell signaling. Exp. Mol. Med. 2019, 51, 1–15. [Google Scholar] [CrossRef]
- Chen, L.; Staubli, S.E.; Schneider, M.P.; Kessels, A.G.; Ivic, S.; Bachmann, L.M.; Kessler, T.M. Phosphodiesterase 5 inhibitors for the treatment of erectile dysfunction: A trade-off network meta-analysis. Eur. Urol. 2015, 68, 674–680. [Google Scholar] [CrossRef]
- Zhuang, X.D.; Long, M.; Li, F.; Hu, X.; Liao, X.X.; Du, Z.M. PDE5 inhibitor sildenafil in the treatment of heart failure: A meta-analysis of randomized controlled trials. Int. J. Cardiol. 2014, 172, 581–587. [Google Scholar] [CrossRef]
- Wu, X.N.; Zhou, Q.; Huang, Y.D.; Xie, X.; Li, Z.; Wu, Y.; Luo, H.B. Structure-based discovery of orally efficient inhibitors via unique interactions with H-pocket of PDE8 for the treatment of vascular dementia. Acta Pharm. Sin. B 2022, 12, 3103–3112. [Google Scholar] [CrossRef] [PubMed]
- Beaumont, V.; Zhong, S.; Lin, H.; Xu, W.; Bradaia, A.; Steidl, E.; Gleyzes, M.; Wadel, K.; Buisson, B.; Padovan-Neto, F.E.; et al. Phosphodiesterase 10A inhibition improves cortico-basal ganglia function in huntington’s disease models. Neuron 2016, 92, 1220–1237. [Google Scholar] [CrossRef] [PubMed]
- Roser-Page, S.; Weiss, D.; Vikulina, T.; Yu, M.; Pacifici, R.; Weitzmann, M.N. Cyclic adenosine monophosphate (cAMP)-dependent phosphodiesterase inhibition promotes bone anabolism through CD8+ T cell wnt-10b production in mice. JBMR Plus 2022, 6, e10636. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, H.; Wang, W.Z.; Wang, D.; Skaggs, K.; Zhang, H.T. Phosphodiesterase 7 (PDE7): A unique drug target for central nervous system diseases. Neuropharmacology 2021, 196, 108694. [Google Scholar] [CrossRef]
- Peng, T.; Qi, B.; He, J.; Ke, H.; Shi, J. Advances in the development of phosphodiesterase-4 inhibitors. J. Med. Chem. 2020, 63, 10594–10617. [Google Scholar] [CrossRef]
- García, A.M.; Brea, J.; Morales-García, J.A.; Perez, D.I.; González, A.; Alonso-Gil, S.; Gracia-Rubio, I.; Ros-Simó, C.; Conde, S.; Cadavid, M.I.; et al. Modulation of cAMP-specific PDE without emetogenic activity: New sulfide-like PDE7 inhibitors. J. Med. Chem. 2014, 57, 8590–8607. [Google Scholar] [CrossRef]
- Castaño, T.; Wang, H.; Campillo, N.E.; Ballester, S.; González-García, C.; Hernández, J.; Pérez, C.; Cuenca, J.; Pérez-Castillo, A.; Martínez, A.; et al. Synthesis, structural analysis, and biological evaluation of thioxoquinazoline derivatives as phosphodiesterase 7 inhibitors. ChemMedChem 2009, 4, 866–876. [Google Scholar] [CrossRef]
- Redondo, M.; Brea, J.; Perez, D.I.; Soteras, I.; Val, C.; Perez, C.; Morales-García, J.A.; Alonso-Gil, S.; Paul-Fernandez, N.; Martin-Alvarez, R.; et al. Effect of phosphodiesterase 7 (PDE7) inhibitors in experimental autoimmune encephalomyelitis mice. Discovery of a new chemically diverse family of compounds. J. Med. Chem. 2012, 55, 3274–3284. [Google Scholar] [CrossRef]
- Banerjee, A.; Yadav, P.S.; Bajpai, M.; Sangana, R.R.; Gullapalli, S.; Gudi, G.S.; Gharat, L.A. Isothiazole and isoxazole fused pyrimidones as PDE7 inhibitors: SAR and pharmacokinetic evaluation. Bioorg. Med. Chem. Lett. 2012, 22, 3223–3228. [Google Scholar] [CrossRef]
- Lorthiois, E.; Bernardelli, P.; Vergne, F.; Oliveira, C.; Mafroud, A.K.; Proust, E.; Heuze, L.; Moreau, F.; Idrissi, M.; Tertre, A.; et al. Spiroquinazolinones as novel, potent, and selective PDE7 inhibitors. Part 1. Bioorg. Med. Chem. Lett. 2004, 14, 4623–4626. [Google Scholar] [CrossRef]
- Kempson, J.; Pitts, W.J.; Barbosa, J.; Guo, J.; Omotoso, O.; Watson, A.; Stebbins, K.; Starling, G.C.; Dodd, J.H.; Barrish, J.C.; et al. Fused pyrimidine based inhibitors of phosphodiesterase 7 (PDE7): Synthesis and initial structure-activity relationships. Bioorg. Med. Chem. Lett. 2005, 15, 1829–1833. [Google Scholar] [CrossRef] [PubMed]
- Gewald, R.; Rueger, C.; Grunwald, C.; Egerland, U.; Hoefgen, N. Synthesis and structure-activity relationship studies of dihydronaphthyridinediones as a novel structural class of potent and selective PDE7 inhibitors. Bioorg. Med. Chem. Lett. 2011, 21, 6652–6656. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.J.; Cieslinski, L.B.; Newton, R.; Donnelly, L.E.; Fenwick, P.S.; Nicholson, A.G.; Barnes, P.J.; Barnette, M.S.; Giembycz, M.A. Discovery of BRL 50481 [3-(N,N-dimethylsulfonamido)-4-methyl-nitrobenzene], a selective inhibitor of phosphodiesterase 7: In vitro studies in human monocytes, lung macrophages, and CD8+ T-lymphocytes. Mol. Pharmacol. 2004, 66, 1679–1689. [Google Scholar] [CrossRef]
- Shao, Y.X.; Huang, M.; Cui, W.; Feng, L.J.; Wu, Y.; Cai, Y.; Li, Z.; Zhu, X.; Liu, P.; Wan, Y.; et al. Discovery of a phosphodiesterase 9A inhibitor as a potential hypoglycemic agent. J. Med. Chem. 2014, 57, 10304–10313. [Google Scholar] [CrossRef]
- Wei, X.; Tang, X.; Liu, N.; Liu, Y.; Guan, G.; Liu, Y.; Wu, X.; Liu, Y.; Wang, J.; Dong, H.; et al. PyCoCa:A quantifying tool of carbon content in airway macrophage for assessment the internal dose of particles. Sci. Total Environ. 2022, 851, 158103. [Google Scholar] [CrossRef]
- Wei, X.; Liu, N.; Song, J.; Ren, C.; Tang, X.; Jiang, W. Effect of silica nanoparticles on cell membrane fluidity: The role of temperature and membrane composition. Sci. Total Environ. 2022, 838, 156552. [Google Scholar] [CrossRef]
- Wang, H.; Liu, Y.; Chen, Y.; Robinson, H.; Ke, H. Multiple elements jointly determine inhibitor selectivity of cyclic nucleotide phosphodiesterases 4 and 7. J. Biol. Chem. 2005, 280, 30949–30955. [Google Scholar] [CrossRef]
- Barnes, M.J.; Cooper, N.; Davenport, R.J.; Dyke, H.J.; Galleway, F.P.; Galvin, F.C.; Gowers, L.; Haughan, A.F.; Lowe, C.; Meissner, J.W.; et al. Synthesis and structure-activity relationships of guanine analogues as phosphodiesterase 7 (PDE7) inhibitors. Bioorg. Med. Chem. Lett. 2001, 11, 1081–1083. [Google Scholar] [CrossRef]
- Pitts, W.J.; Vaccaro, W.; Huynh, T.; Leftheris, K.; Roberge, J.Y.; Barbosa, J.; Guo, J.Q.; Brown, B.; Watson, A.; Donaldson, K.; et al. Identification of purine inhibitors of phosphodiesterase 7 (PDE7). Bioorg. Med. Chem. Lett. 2004, 14, 2955–2958. [Google Scholar] [CrossRef]
- Endo, Y.; Kawai, K.; Asano, T.; Amano, S.; Asanuma, Y.; Sawada, K.; Onodera, Y.; Ueo, N.; Takahashi, N.; Sonoda, Y.; et al. 2-(Isopropylamino)thieno[3,2-d]pyrimidin-4(3H)-one derivatives as selective phosphodiesterase 7 inhibitors with potent in vivo efficacy. Bioorg. Med. Chem. Lett. 2015, 25, 1910–1914. [Google Scholar] [CrossRef]
- Kawai, K.; Endo, Y.; Asano, T.; Amano, S.; Sawada, K.; Ueo, N.; Takahashi, N.; Sonoda, Y.; Nagai, M.; Kamei, N.; et al. Discovery of 2-(Cyclopentylamino)thieno[3,2-d]pyrimidin-4(3H)-one derivatives as a new series of potent phosphodiesterase 7 inhibitors. J. Med. Chem. 2014, 57, 9844–9854. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Hersey, A.; Nowotka, M.; Bento, A.P.; Chambers, J.; Mendez, D.; Mutowo, P.; Atkinson, F.; Bellis, L.J.; Cibrián-Uhalte, E.; et al. The ChEMBL database in 2017. Nucleic Acids Res. 2017, 45, D945. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xiao, J.; Suzek, T.O.; Zhang, J.; Wang, J.; Zhou, Z.; Han, L.; Karapetyan, K.; Dracheva, S.; Shoemaker, B.A.; et al. PubChem’s bioAssay database. Nucleic Acids Res. 2012, 40, D400–D412. [Google Scholar] [CrossRef] [PubMed]
- Ding, W.; Nan, Y.; Wu, J.; Han, C.; Xin, X.; Li, S.; Liu, H.; Zhang, L. Combining multi-dimensional molecular fingerprints to predict the hERG cardiotoxicity of compounds. Comput. Biol. Med. 2022, 144, 105390. [Google Scholar] [CrossRef]
- Zhou, H.; Shan, M.; Qin, L.P.; Cheng, G. Reliable prediction of cannabinoid receptor 2 ligand by machine learning based on combined fingerprints. Comput. Biol. Med. 2023, 152, 106379. [Google Scholar] [CrossRef]
- Niu, C.; Sun, X.; Hu, F.; Tang, X.; Wang, K. Molecular determinants for the chemical activation of the warmth-sensitive TRPV3 channel by the natural monoterpenoid carvacrol. J. Biol. Chem. 2022, 298, 101706. [Google Scholar] [CrossRef]
- Huang, J.X.; Zhu, B.L.; Xu, J.P.; Zhou, Z.Z. Advances in the development of phosphodiesterase 7 inhibitors. Eur. J. Med. Chem. 2023, 250, 115194. [Google Scholar] [CrossRef]
- Kalliokoski, T.; Kramer, C.; Vulpetti, A.; Gedeck, P. Comparability of mixed IC₅₀ data—A statistical analysis. PLoS ONE 2013, 8, e61007. [Google Scholar] [CrossRef]
- Feng, H.; Zhang, L.; Li, S.; Liu, L.; Yang, T.; Yang, P.; Zhao, J.; Arkin, I.T.; Liu, H. Predicting the reproductive toxicity of chemicals using ensemble learning methods and molecular fingerprints. Toxicol. Lett. 2021, 340, 4–14. [Google Scholar] [CrossRef]
- Yan, Z.; Caldwell, G.W. Metabolism profiling, and cytochrome P450 inhibition & induction in drug discovery. Curr. Top. Med. Chem. 2001, 1, 403–425. [Google Scholar] [CrossRef]
- Mysinger, M.M.; Carchia, M.; Irwin, J.J.; Shoichet, B.K. Directory of useful decoys, enhanced (DUD-E): Better ligands and decoys for better benchmarking. J. Med. Chem. 2012, 55, 6582–6594. [Google Scholar] [CrossRef] [PubMed]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef] [PubMed]
- Bemis, G.W.; Murcko, M.A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 1996, 39, 2887–2893. [Google Scholar] [CrossRef]
- Zhao, J.; Shi, X.; Wang, Z.; Xiong, S.; Lin, Y.; Wei, X.; Li, Y.; Tang, X. Hepatotoxicity assessment investigations on PFASs targeting L-FABP using binding affinity data and machine learning-based QSAR model. Ecotoxicol. Environ. Saf. 2023, 262, 115310. [Google Scholar] [CrossRef]
- Speybroeck, N. Classification and regression trees. Int. J. Public Health 2012, 57, 243–246. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Nedaie, A.; Najafi, A.A. Support vector machine with Dirichlet feature mapping. Neural Netw. 2018, 98, 87–101. [Google Scholar] [CrossRef]
- Chen, T.Q.; Guestrin, C.; Assoc Comp, M. XGBoost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (KDD), San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Tibshirani, R. The lasso method for variable selection in the Cox model. Stat. Med. 1997, 16, 385–395. [Google Scholar] [CrossRef]
- Tikhonov, A.N. Solution of incorrectly formulated problem and the regularization method. Sov. Dok. 1963, 4, 1035–1038. [Google Scholar]
- Swami, A.; Jain, R.J. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 2013, 12, 2825–2830. [Google Scholar]
- Kirk, D.; Catal, C.; Tekinerdogan, B. Precision nutrition: A systematic literature review. Comput. Biol. Med. 2021, 133, 104365. [Google Scholar] [CrossRef]
- Lundberg, S.M.; Lee, S.I. A unified approach to interpreting model predictions. In Proceedings of the 31st Annual Conference on Neural Information Processing Systems (NIPS), Long Beach, CA, USA, 4–9 December 2017. [Google Scholar]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Wang, Z.; Ma, S.; Tang, X.; Zhang, H. Chemical Space Exploration and Machine Learning-Based Screening of PDE7A Inhibitors. Pharmaceuticals 2025, 18, 444. https://doi.org/10.3390/ph18040444
Li Y, Wang Z, Ma S, Tang X, Zhang H. Chemical Space Exploration and Machine Learning-Based Screening of PDE7A Inhibitors. Pharmaceuticals. 2025; 18(4):444. https://doi.org/10.3390/ph18040444
Chicago/Turabian StyleLi, Yuze, Zhe Wang, Shengyao Ma, Xiaowen Tang, and Hanting Zhang. 2025. "Chemical Space Exploration and Machine Learning-Based Screening of PDE7A Inhibitors" Pharmaceuticals 18, no. 4: 444. https://doi.org/10.3390/ph18040444
APA StyleLi, Y., Wang, Z., Ma, S., Tang, X., & Zhang, H. (2025). Chemical Space Exploration and Machine Learning-Based Screening of PDE7A Inhibitors. Pharmaceuticals, 18(4), 444. https://doi.org/10.3390/ph18040444





