Advanced High-Content Phenotypic Screening to Identify Drugs That Ameliorate the Inhibition of Skeletal Muscle Cell Differentiation Induced by Cancer Cachexia Serum
Abstract
1. Introduction
2. Results
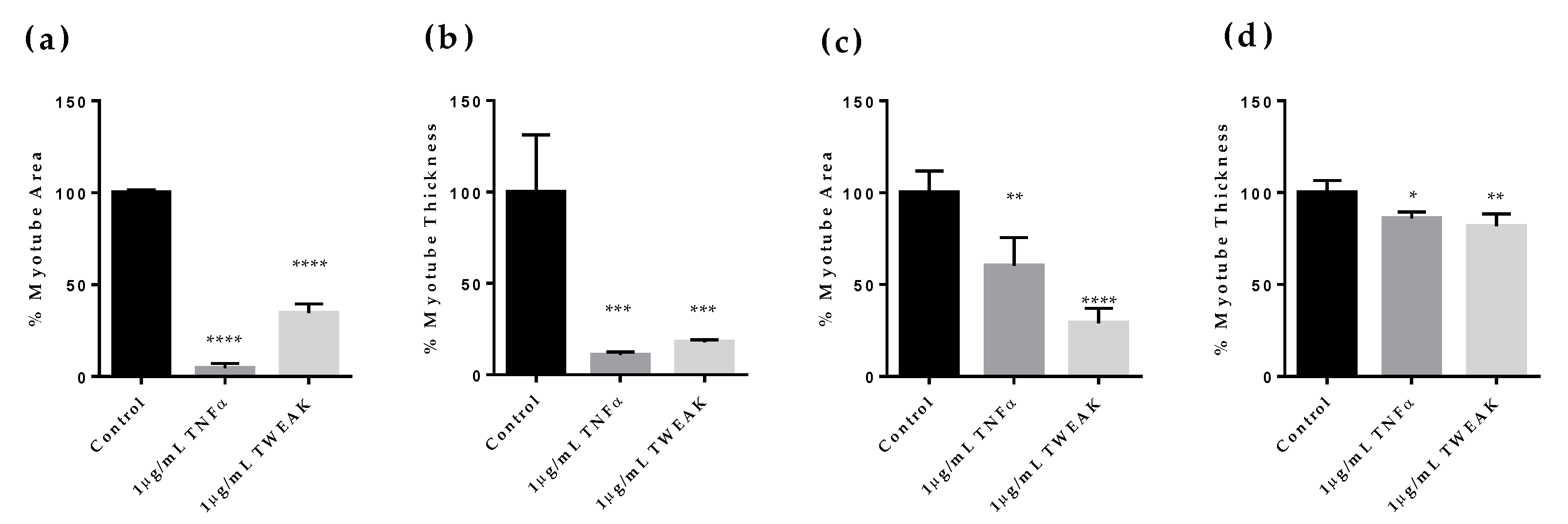
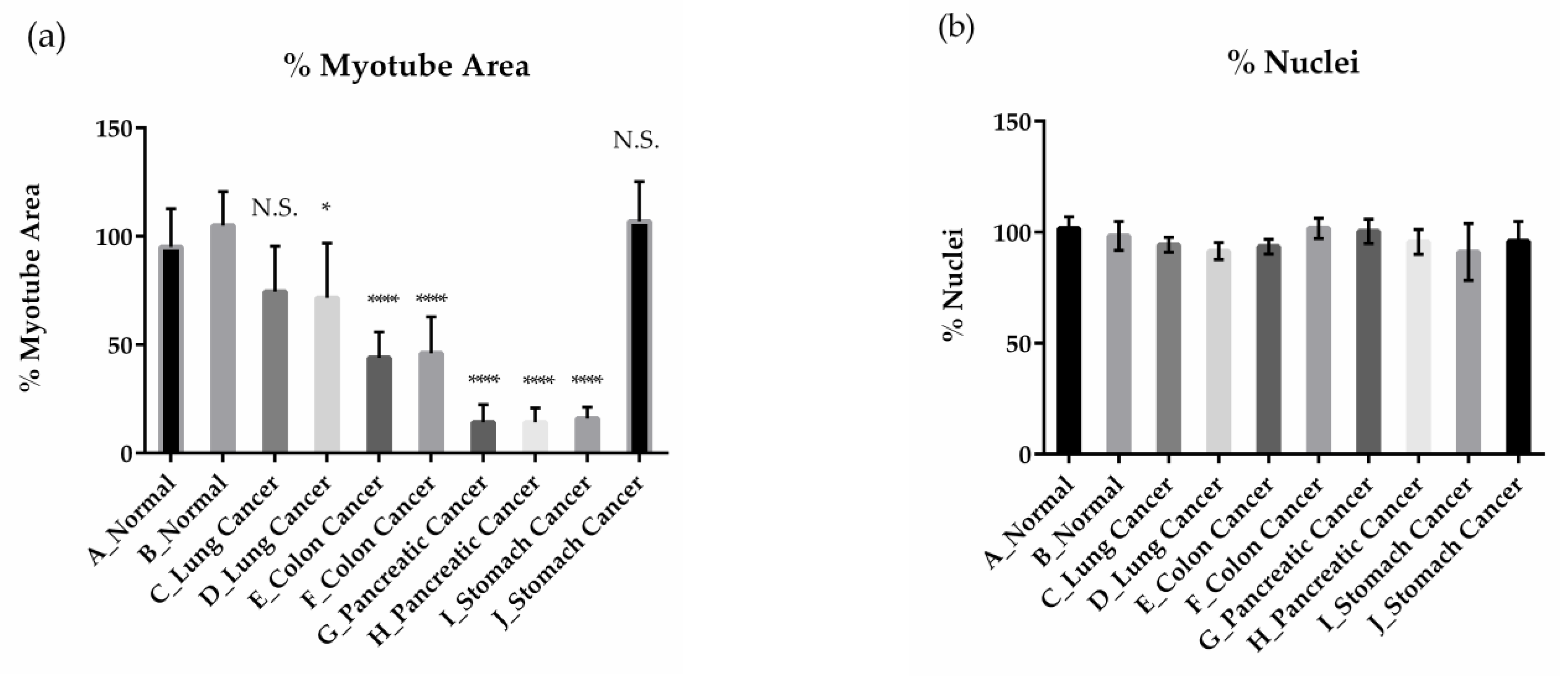
2.1. Effects of Cancer Patient Serum on Human Skeletal Muscle Differentiation and Atrophy In Vitro
2.2. Establishment of a High-Content Phenotypic Screening System Using Cancer Patient Serum for Drug Screening
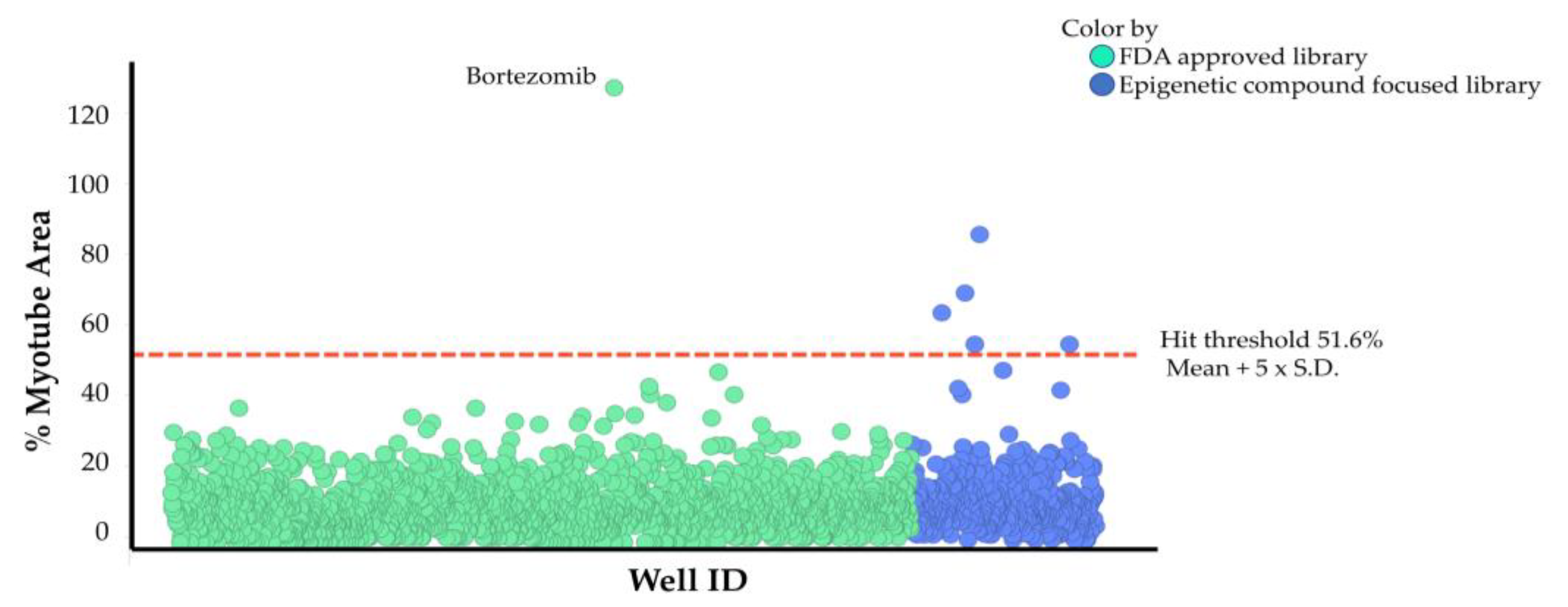
2.3. Drug Screening Using High-Content Phenotypic Screening with Cancer Patient Sera
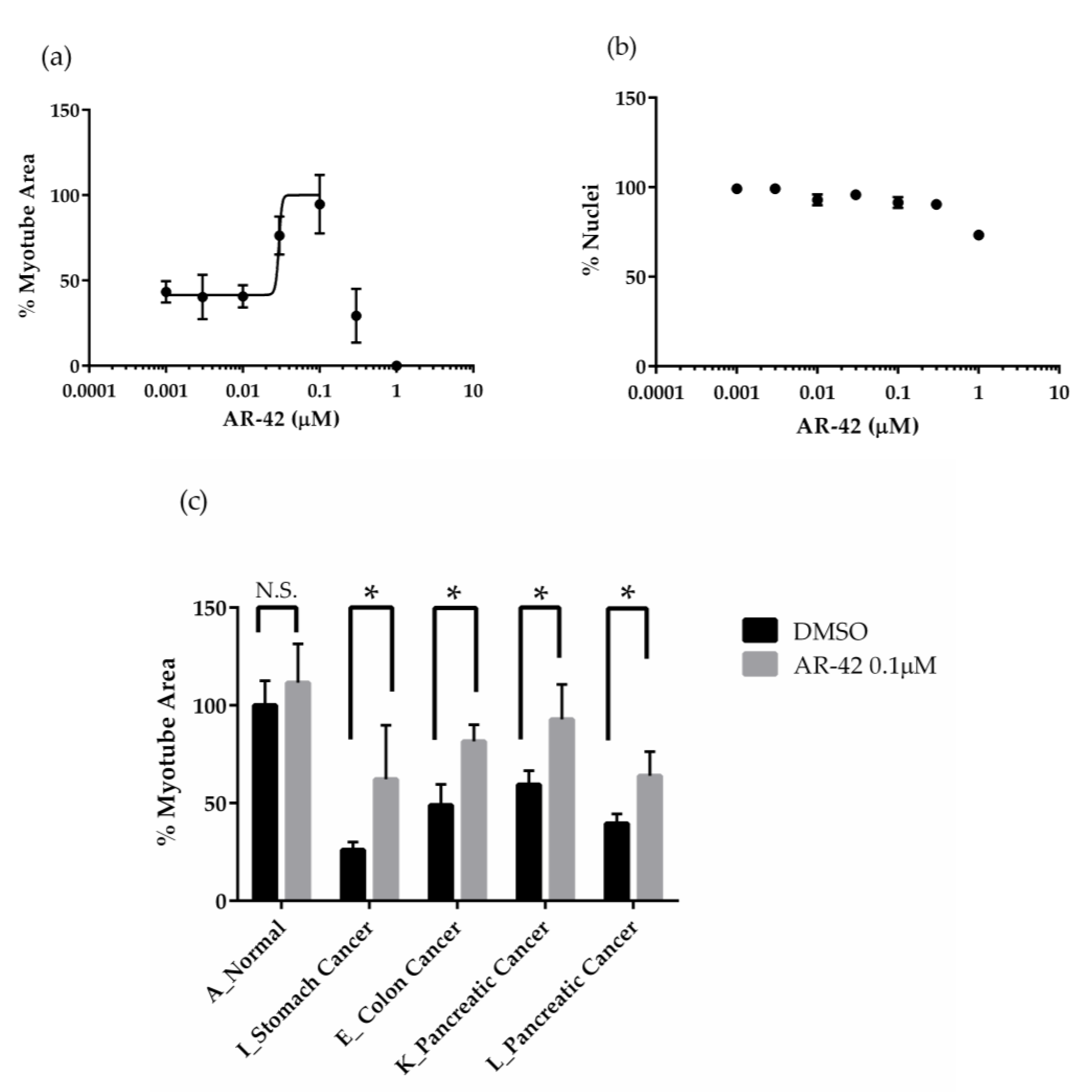
2.4. The Effects of AR-42 on Muscle Differentiation Inhibition by Cancer Patient Sera
3. Discussion
4. Materials and Methods
4.1. Human Cell Culture and Evaluation of Skeletal Muscle Differentiation and Atrophy
4.2. High Content, Phenotypic Screening
4.3. Immunostaining and Fluorescence Analysis
4.4. Quantitative Real Time PCR
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fearon, K.; Strasser, F.; Anker, S.D.; Bosaeus, I.; Bruera, E.; Fainsinger, R.L.; Jatoi, A.; Loprinzi, C.; MacDonald, N.; Mantovani, G.; et al. Definition and Classification of Cancer Cachexia: An International Consensus. Lancet Oncol. 2011, 12, 489–495. [Google Scholar] [CrossRef] [PubMed]
- Argilés, J.M.; Stemmler, B.; López-Soriano, F.J.; Busquets, S. Inter-Tissue Communication in Cancer Cachexia. Nat. Rev. Endocrinol. 2019, 15, 9–20. [Google Scholar] [CrossRef]
- Lim, S.; Brown, J.L.; Washington, T.A.; Greene, N.P. Development and Progression of Cancer Cachexia: Perspectives from Bench to Bedside. Sports Med. Health Sci. 2020, 2, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Baracos, V.E.; Martin, L.; Korc, M.; Guttridge, D.C.; Fearon, K.C.H. Cancer-Associated Cachexia. Nat. Rev. Dis. Primers 2018, 4, 17105. [Google Scholar] [CrossRef]
- Mueller, T.C.; Bachmann, J.; Prokopchuk, O.; Friess, H.; Martignoni, M.E. Molecular Pathways Leading to Loss of Skeletal Muscle Mass in Cancer Cachexia—Can Findings from Animal Models Be Translated to Humans? BMC Cancer 2016, 16, 75. [Google Scholar] [CrossRef]
- Jatoi, A.; Dakhil, S.R.; Nguyen, P.L.; Sloan, J.A.; Kugler, J.W.; Rowland, K.M.; Soori, G.S.; Wender, D.B.; Fitch, T.R.; Novotny, P.J.; et al. A Placebo-controlled Double Blind Trial of Etanercept for the Cancer Anorexia/Weight Loss Syndrome: Results from N00C1 from the North Central Cancer Treatment Group. Cancer 2007, 110, 1396–1403. [Google Scholar] [CrossRef]
- Jatoi, A.; Ritter, H.L.; Dueck, A.; Nguyen, P.L.; Nikcevich, D.A.; Luyun, R.F.; Mattar, B.I.; Loprinzi, C.L. A Placebo-Controlled, Double-Blind Trial of Infliximab for Cancer-Associated Weight Loss in Elderly and/or Poor Performance Non-Small Cell Lung Cancer Patients (N01C9). Lung Cancer 2010, 68, 234–239. [Google Scholar] [CrossRef]
- Kim-Muller, J.Y.; Song, L.; LaCarubba Paulhus, B.; Pashos, E.; Li, X.; Rinaldi, A.; Joaquim, S.; Stansfield, J.C.; Zhang, J.; Robertson, A.; et al. GDF15 Neutralization Restores Muscle Function and Physical Performance in a Mouse Model of Cancer Cachexia. Cell Rep. 2023, 42, 111947. [Google Scholar] [CrossRef]
- Tseng, Y.-C.; Kulp, S.K.; Lai, I.-L.; Hsu, E.-C.; He, W.A.; Frankhouser, D.E.; Yan, P.S.; Mo, X.; Bloomston, M.; Lesinski, G.B.; et al. Preclinical Investigation of the Novel Histone Deacetylase Inhibitor AR-42 in the Treatment of Cancer-Induced Cachexia. J. Natl. Cancer Inst. 2015, 107, djv274. [Google Scholar] [CrossRef]
- Martin, A.; Gallot, Y.S.; Freyssenet, D. Molecular Mechanisms of Cancer Cachexia-related Loss of Skeletal Muscle Mass: Data Analysis from Preclinical and Clinical Studies. J. Cachexia Sarcopenia Muscle 2023, 14, 1150–1167. [Google Scholar] [CrossRef]
- Mirzoev, T.M. Skeletal Muscle Recovery from Disuse Atrophy: Protein Turnover Signaling and Strategies for Accelerating Muscle Regrowth. Int. J. Mol. Sci. 2020, 21, 7940. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, J.A.; Pothuraju, R.; Jain, M.; Batra, S.K.; Nasser, M.W. Advances in Cancer Cachexia: Intersection between Affected Organs, Mediators, and Pharmacological Interventions. Biochim. Biophys. Acta (BBA)—Rev. Cancer 2020, 1873, 188359. [Google Scholar] [CrossRef] [PubMed]
- Malla, J.; Zahra, A.; Venugopal, S.; Selvamani, T.Y.; Shoukrie, S.I.; Selvaraj, R.; Dhanoa, R.K.; Hamouda, R.K.; Mostafa, J. What Role Do Inflammatory Cytokines Play in Cancer Cachexia? Cureus 2022, 14, e26798. [Google Scholar] [CrossRef]
- Karalaki, M.; Fili, S.; Philippou, A.; Koutsilieris, M. Muscle Regeneration: Cellular and Molecular Events. In Vivo 2009, 23, 779–796. [Google Scholar] [PubMed]
- Fields, D.P.; Roberts, B.M.; Simon, A.K.; Judge, A.R.; Fuller, D.D.; Mitchell, G.S. Cancer Cachexia Impairs Neural Respiratory Drive in Hypoxia but Not Hypercapnia. J. Cachexia Sarcopenia Muscle 2019, 10, 63–72. [Google Scholar] [CrossRef]
- Inaba, S.; Hinohara, A.; Tachibana, M.; Tsujikawa, K.; Fukada, S. Muscle Regeneration Is Disrupted by Cancer Cachexia without Loss of Muscle Stem Cell Potential. PLoS ONE 2018, 13, e0205467. [Google Scholar] [CrossRef]
- Guttridge, D.C.; Mayo, M.W.; Madrid, L.V.; Wang, C.-Y.; Baldwin, A.S., Jr. NF-κB-Induced Loss of MyoD Messenger RNA: Possible Role in Muscle Decay and Cachexia. Science 2000, 289, 2363–2366. [Google Scholar] [CrossRef]
- Fabio, P.; Domiziana, C.; Alessandro, F.; Gabriella, B.; Francesco, M.B.; Paola, C. Muscle Wasting and Impaired Myogenesis in Tumor Bearing Mice Are Prevented by ERK Inhibition. PLoS ONE 2010, 5, e1360413. [Google Scholar] [CrossRef]
- Panguluri, S.K.; Bhatnagar, S.; Kumar, A.; McCarthy, J.J.; Srivastava, A.K.; Cooper, N.G.; Lundy, R.F.; Kumar, A. Genomic Profiling of Messenger RNAs and MicroRNAs Reveals Potential Mechanisms of TWEAK-Induced Skeletal Muscle Wasting in Mice. PLoS ONE 2010, 5, e8760. [Google Scholar] [CrossRef]
- Tajrishi, M.M.; Sato, S.; Shin, J.; Zheng, T.S.; Burkly, L.C.; Kumar, A. The TWEAK–Fn14 Dyad Is Involved in Age-Associated Pathological Changes in Skeletal Muscle. Biochem. Biophys. Res. Commun. 2014, 446, 1219–1224. [Google Scholar] [CrossRef]
- Martin, A.; Freyssenet, D. Phenotypic Features of Cancer Cachexia-related Loss of Skeletal Muscle Mass and Function: Lessons from Human and Animal Studies. J. Cachexia Sarcopenia Muscle 2021, 12, 252–273. [Google Scholar] [CrossRef] [PubMed]
- Guigni, B.A.; Van Der Velden, J.; Kinsey, C.M.; Carson, J.A.; Toth, M.J. Effects of Conditioned Media from Murine Lung Cancer Cells and Human Tumor Cells on Cultured Myotubes. Am. J. Physiol.-Endocrinol. Metab. 2020, 318, E22–E32. [Google Scholar] [CrossRef] [PubMed]
- Menconi, M.; Gonnella, P.; Petkova, V.; Lecker, S.; Hasselgren, P. Dexamethasone and Corticosterone Induce Similar, but Not Identical, Muscle Wasting Responses in Cultured L6 and C2C12 Myotubes. J. Cell. Biochem. 2008, 105, 353–364. [Google Scholar] [CrossRef] [PubMed]
- Barreto, R.; Waning, D.L.; Gao, H.; Liu, Y.; Zimmers, T.A.; Bonetto, A. Chemotherapy-Related Cachexia Is Associated with Mitochondrial Depletion and the Activation of ERK1/2 and P38 MAPKs. Oncotarget 2016, 7, 43442–43460. [Google Scholar] [CrossRef]
- Ubachs, J.; Van De Worp, W.R.P.H.; Vaes, R.D.W.; Pasmans, K.; Langen, R.C.; Meex, R.C.R.; Van Bijnen, A.A.J.H.M.; Lambrechts, S.; Van Gorp, T.; Kruitwagen, R.F.P.M.; et al. Ovarian Cancer Ascites Induces Skeletal Muscle Wasting in Vitro and Reflects Sarcopenia in Patients. J. Cachexia Sarcopenia Muscle 2022, 13, 311–324. [Google Scholar] [CrossRef]
- Owens, J.; Moreira, K.; Bain, G. Characterization of Primary Human Skeletal Muscle Cells from Multiple Commercial Sources. In Vitro Cell. Dev.Biol.-Anim. 2013, 49, 695–705. [Google Scholar] [CrossRef]
- Miller, S.C.; Ito, H.; Blau, H.M.; Torti, F.M. Tumor Necrosis Factor Inhibits Human Myogenesis In Vitro. Mol. Cell. Biol. 1988, 8, 2295–2301. [Google Scholar] [CrossRef]
- Dogra, C.; Changotra, H.; Mohan, S.; Kumar, A. Tumor Necrosis Factor-like Weak Inducer of Apoptosis Inhibits Skeletal Myogenesis through Sustained Activation of Nuclear Factor-κB and Degradation of MyoD Protein. J. Biol. Chem. 2006, 281, 10327–10336. [Google Scholar] [CrossRef]
- Dogra, C.; Changoua, H.; Wedhas, N.; Qin, X.; Wergedal, J.E.; Kumar, A. TNF-related Weak Inducer of Apoptosis (TWEAK) Is a Potent Skeletal Muscle-wasting Cytokine. FASEB J. 2007, 21, 1857–1869. [Google Scholar] [CrossRef]
- Bar, H.; Zweifach, A. Z’ Does Not Need to Be > 0.5. SLAS Discov. 2020, 25, 1000–1008. [Google Scholar] [CrossRef]
- Novotny-Diermayr, V.; Sangthongpitag, K.; Hu, C.Y.; Wu, X.; Sausgruber, N.; Yeo, P.; Greicius, G.; Pettersson, S.; Liang, A.L.; Loh, Y.K.; et al. SB939, a Novel Potent and Orally Active Histone Deacetylase Inhibitor with High Tumor Exposure and Efficacy in Mouse Models of Colorectal Cancer. Mol. Cancer Ther. 2010, 9, 642–652. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.C.S.; Chan, A.H.Y.; Ganesan, A. Thirty Years of HDAC Inhibitors: 2020 Insight and Hindsight. J. Med. Chem. 2020, 63, 12460–12484. [Google Scholar] [CrossRef] [PubMed]
- Hassig, C.A.; Symons, K.T.; Guo, X.; Nguyen, P.-M.; Annable, T.; Wash, P.L.; Payne, J.E.; Jenkins, D.A.; Bonnefous, C.; Trotter, C.; et al. KD5170, a Novel Mercaptoketone-Based Histone Deacetylase Inhibitor That Exhibits Broad Spectrum Antitumor Activity in Vitro and in Vivo. Mol. Cancer Ther. 2008, 7, 1054–1065. [Google Scholar] [CrossRef]
- Kozikowski, A.P.; Tapadar, S.; Luchini, D.N.; Kim, K.H.; Billadeau, D.D. Use of the Nitrile Oxide Cycloaddition (NOC) Reaction for Molecular Probe Generation: A New Class of Enzyme Selective Histone Deacetylase Inhibitors (HDACIs) Showing Picomolar Activity at HDAC6. J. Med. Chem. 2008, 51, 4370–4373. [Google Scholar] [CrossRef]
- Chou, C.J.; Herman, D.; Gottesfeld, J.M. Pimelic Diphenylamide 106 Is a Slow, Tight-Binding Inhibitor of Class I Histone Deacetylases. J. Biol. Chem. 2008, 283, 35402–35409. [Google Scholar] [CrossRef]
- Balasubramanian, S.; Ramos, J.; Luo, W.; Sirisawad, M.; Verner, E.; Buggy, J.J. A Novel Histone Deacetylase 8 (HDAC8)-Specific Inhibitor PCI-34051 Induces Apoptosis in T-Cell Lymphomas. Leukemia 2008, 22, 1026–1034. [Google Scholar] [CrossRef]
- Tseng, Y.-C.; Liva, S.G.; Dauki, A.M.; Sovic, M.; Henderson, S.E.; Kuo, Y.-C.; Benedict, J.A.; Kulp, S.K.; Campbell, M.; Bekaii-Saab, T.; et al. Overcoming Resistance to Anabolic Selective Androgen Receptor Modulator (SARM) Therapy in Experimental Cancer Cachexia with Histone Deacetylase Inhibitor AR-42. bioRxiv 2017. [Google Scholar] [CrossRef]
- Salmi-Smail, C.; Fabre, A.; Dequiedt, F.; Restouin, A.; Castellano, R.; Garbit, S.; Roche, P.; Morelli, X.; Brunel, J.M.; Collette, Y. Modified Cap Group Suberoylanilide Hydroxamic Acid Histone Deacetylase Inhibitor Derivatives Reveal Improved Selective Antileukemic Activity. J. Med. Chem. 2010, 53, 3038–3047. [Google Scholar] [CrossRef]
- Abdelmoez, A.M.; Sardón Puig, L.; Smith, J.A.B.; Gabriel, B.M.; Savikj, M.; Dollet, L.; Chibalin, A.V.; Krook, A.; Zierath, J.R.; Pillon, N.J. Comparative Profiling of Skeletal Muscle Models Reveals Heterogeneity of Transcriptome and Metabolism. Am. J. Physiol.-Cell Physiol. 2020, 318, C615–C626. [Google Scholar] [CrossRef]
- Kadakia, K.C.; Hamilton-Reeves, J.M.; Baracos, V.E. Current Therapeutic Targets in Cancer Cachexia: A Pathophysiologic Approach. Am. Soc. Clin. Oncol. Educ. Book 2023, 43, e389942. [Google Scholar] [CrossRef]
- Neshan, M.; Tsilimigras, D.I.; Han, X.; Zhu, H.; Pawlik, T.M. Molecular Mechanisms of Cachexia: A Review. Cells 2024, 13, 252. [Google Scholar] [CrossRef] [PubMed]
- Cao, Z.; Zhao, K.; Jose, I.; Hoogenraad, N.J.; Osellame, L.D. Biomarkers for Cancer Cachexia: A Mini Review. Int. J. Mol. Sci. 2021, 22, 4501. [Google Scholar] [CrossRef] [PubMed]
- Arneson, P.C.; Doles, J.D. Impaired Muscle Regeneration in Cancer-Associated Cachexia. Trends Cancer 2019, 5, 579–582. [Google Scholar] [CrossRef] [PubMed]
- He, W.A.; Berardi, E.; Cardillo, V.M.; Acharyya, S.; Aulino, P.; Thomas-Ahner, J.; Wang, J.; Bloomston, M.; Muscarella, P.; Nau, P.; et al. NF-kB-Mediated Pax7 Dysregulation in the Muscle Microenvironment Promotes Cancer Cachexia. J. Clin. Investig. 2013, 123, 4821–4835. [Google Scholar] [CrossRef]
- Nishima, S.; Kashiwada, T.; Saito, Y.; Yuge, S.; Ishii, T.; Matsuda, K.; Kamio, K.; Seike, M.; Fukuhara, S.; Gemma, A. Bortezomib Induces Rho-Dependent Hyperpermeability of Endothelial Cells Synergistically with Inflammatory Mediators. BMC Pulm. Med. 2024, 24, 617. [Google Scholar] [CrossRef]
- Wei, L.; Zhou, W.; Croissant, J.D.; Johansen, F.-E.; Prywes, R.; Balasubramanyam, A.; Schwartz, R.J. RhoA Signaling via Serum Response Factor Plays an Obligatory Role in Myogenic Differentiation. J. Biol. Chem. 1998, 273, 30287–30294. [Google Scholar] [CrossRef]
- Nevi, L.; Pöllänen, N.; Penna, F.; Caretti, G. Targeting Epigenetic Regulators with HDAC and BET Inhibitors to Modulate Muscle Wasting. Int. J. Mol. Sci. 2023, 24, 16404. [Google Scholar] [CrossRef]
- Licandro, S.A.; Crippa, L.; Pomarico, R.; Perego, R.; Fossati, G.; Leoni, F.; Steinkühler, C. The Pan HDAC Inhibitor Givinostat Improves Muscle Function and Histological Parameters in Two Duchenne Muscular Dystrophy Murine Models Expressing Different Haplotypes of the LTBP4 Gene. Skelet. Muscle 2021, 11, 19. [Google Scholar] [CrossRef]
- Mercuri, E.; Vilchez, J.J.; Boespflug-Tanguy, O.; Zaidman, C.M.; Mah, J.K.; Goemans, N.; Müller-Felber, W.; Niks, E.H.; Schara-Schmidt, U.; Bertini, E.; et al. Safety and Efficacy of Givinostat in Boys with Duchenne Muscular Dystrophy (EPIDYS): A Multicentre, Randomised, Double-Blind, Placebo-Controlled, Phase 3 Trial. Lancet Neurol. 2024, 23, 393–403. [Google Scholar] [CrossRef]
- Iezzi, S.; Cossu, G.; Nervi, C.; Sartorelli, V.; Puri, P.L. Stage-Specific Modulation of Skeletal Myogenesis by Inhibitors of Nuclear Deacetylases. Proc. Natl. Acad. Sci. USA 2002, 99, 7757–7762. [Google Scholar] [CrossRef]
- Black, B.L.; Olson, E.N. Transcriptional control of muscle development by Myocyte Enhancer FACTOR-2 (MEF2) Proteins. Annu. Rev. Cell Dev. Biol. 1998, 14, 167–196. [Google Scholar] [CrossRef] [PubMed]
- Miska, E.A. HDAC4 Deacetylase Associates with and Represses the MEF2 Transcription Factor. EMBO J. 1999, 18, 5099–5107. [Google Scholar] [CrossRef] [PubMed]
- Ratti, F.; Ramond, F.; Moncollin, V.; Simonet, T.; Milan, G.; Méjat, A.; Thomas, J.-L.; Streichenberger, N.; Gilquin, B.; Matthias, P.; et al. Histone Deacetylase 6 Is a FoxO Transcription Factor-Dependent Effector in Skeletal Muscle Atrophy. J. Biol. Chem. 2015, 290, 4215–4224. [Google Scholar] [CrossRef] [PubMed]
- Ying Wang, J.; Peruzzi, F.; Lassak, A.; Del Valle, L.; Radhakrishnan, S.; Rappaport, J.; Khalili, K.; Amini, S.; Reiss, K. Neuroprotective Effects of IGF-I against TNFα-Induced Neuronal Damage in HIV-Associated Dementia. Virology 2003, 305, 66–76. [Google Scholar] [CrossRef][Green Version]
- Witowski, J.; Tayama, H.; Książek, K.; Wanic-Kossowska, M.; Bender, T.O.; Jörres, A. Human Peritoneal Fibroblasts Are a Potent Source of Neutrophil-Targeting Cytokines: A Key Role of IL-1β Stimulation. Lab. Investig. 2009, 89, 414–424. [Google Scholar] [CrossRef]



| ¥ | Patient Diagnosis | Grade | TNM Classification |
|---|---|---|---|
| A | – * | – * | – * |
| B | – * | – * | – * |
| C | Lung Cancer | III | T3, N0, M0 |
| D | Lung Cancer | IIIA | T2, N2, M0 |
| E | Colon Cancer | III | T3, Nx, M0 |
| F | Colon Cancer | III | T3, N0, M0 |
| G | Pancreatic Cancer | IV | Unknown |
| H | Pancreatic Cancer | IV | Unknown |
| I | Stomach Cancer | IV | T3, Nx, M1 |
| J | Stomach Cancer | Unknown | Unknown |
| K | Pancreatic Cancer | III | T4, N0, M0 |
| L | Pancreatic Cancer | III | T4, N0, M0 |
| M | Pancreatic Cancer | IV | Unknown |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nakane, A.; Nakagawa, H.; Nagata, H. Advanced High-Content Phenotypic Screening to Identify Drugs That Ameliorate the Inhibition of Skeletal Muscle Cell Differentiation Induced by Cancer Cachexia Serum. Pharmaceuticals 2025, 18, 445. https://doi.org/10.3390/ph18040445
Nakane A, Nakagawa H, Nagata H. Advanced High-Content Phenotypic Screening to Identify Drugs That Ameliorate the Inhibition of Skeletal Muscle Cell Differentiation Induced by Cancer Cachexia Serum. Pharmaceuticals. 2025; 18(4):445. https://doi.org/10.3390/ph18040445
Chicago/Turabian StyleNakane, Atsushi, Hiroyuki Nakagawa, and Hidetaka Nagata. 2025. "Advanced High-Content Phenotypic Screening to Identify Drugs That Ameliorate the Inhibition of Skeletal Muscle Cell Differentiation Induced by Cancer Cachexia Serum" Pharmaceuticals 18, no. 4: 445. https://doi.org/10.3390/ph18040445
APA StyleNakane, A., Nakagawa, H., & Nagata, H. (2025). Advanced High-Content Phenotypic Screening to Identify Drugs That Ameliorate the Inhibition of Skeletal Muscle Cell Differentiation Induced by Cancer Cachexia Serum. Pharmaceuticals, 18(4), 445. https://doi.org/10.3390/ph18040445






