Relationship between XPA, XPB/ERCC3, XPF/ERCC4, and XPG/ERCC5 Polymorphisms and the Susceptibility to Head and Neck Carcinoma: A Systematic Review, Meta-Analysis, and Trial Sequential Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Identification of Articles
2.3. Selection Criteria
2.4. Data Summary
2.5. Quality Evaluation
2.6. Statistical Analyses
3. Results
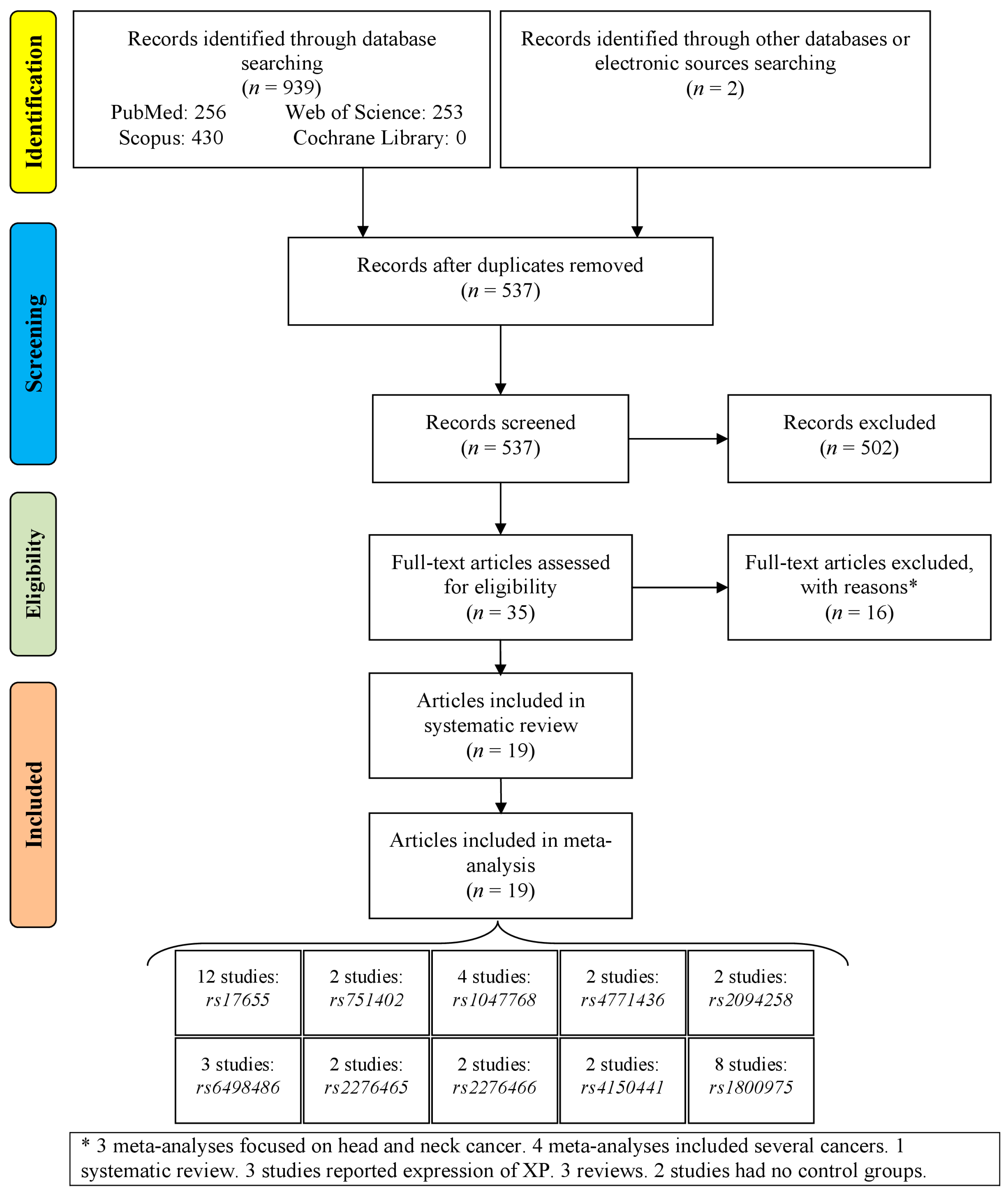
3.1. Study Selection
3.2. Characteristics of the Articles
3.3. Pooled Analysis
3.4. Subgroup Analysis
3.5. Meta-Regression Analysis
3.6. Sensitivity Analysis
3.7. TSA
3.8. Publication Bias
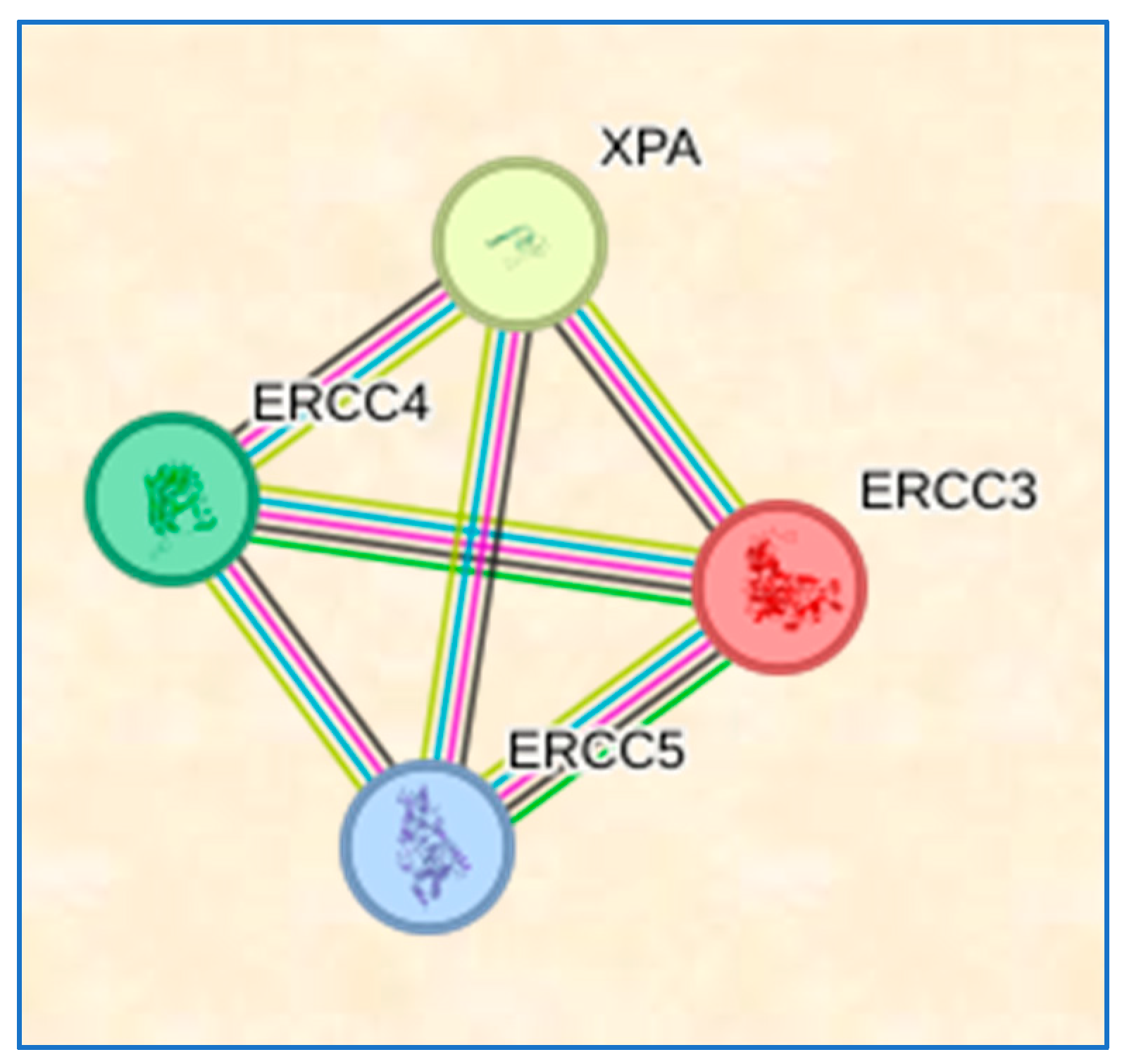
3.9. STRING Results
4. Discussion
5. Conclusions
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Barsouk, A.; Aluru, J.S.; Rawla, P.; Saginala, K.; Barsouk, A. Epidemiology, Risk Factors, and Prevention of Head and Neck Squamous Cell Carcinoma. Med. Sci. 2023, 11, 42. [Google Scholar] [CrossRef]
- Dhull, A.K.; Atri, R.; Dhankhar, R.; Chauhan, A.K.; Kaushal, V. Major risk factors in head and neck cancer: A retrospective analysis of 12-year experiences. World J. Oncol. 2018, 9, 80. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.-H.; Xirasagar, S.; Cheng, Y.-F.; Chen, C.-S.; Chang, W.-P.; Lin, H.-C. Trends in the incidence of head and neck cancer: A nationwide population-based study. Oral Oncol. 2023, 140, 106391. [Google Scholar] [CrossRef] [PubMed]
- Machiels, J.-P.; Leemans, C.R.; Golusinski, W.; Grau, C.; Licitra, L.; Gregoire, V. Squamous cell carcinoma of the oral cavity, larynx, oropharynx and hypopharynx: EHNS–ESMO–ESTRO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2020, 31, 1462–1475. [Google Scholar] [CrossRef] [PubMed]
- Lacko, M.; Braakhuis, B.J.; Sturgis, E.M.; Boedeker, C.C.; Suárez, C.; Rinaldo, A.; Ferlito, A.; Takes, R.P. Genetic susceptibility to head and neck squamous cell carcinoma. Int. J. Radiat. Oncol. Biol. Phys. 2014, 89, 38–48. [Google Scholar] [CrossRef]
- Mohammadi, H.; Momeni Roochi, M.; Rezaei, F.; Garajei, A.; Heidar, H.; Ghaderi, B.; Sadeghi, M. Association between the CYP1A1 MspI polymorphism and risk of head and neck cancer: A meta-analysis. Sci. Rep. 2022, 12, 1527. [Google Scholar] [CrossRef]
- Mohammadi, H.; Roochi, M.M.; Sadeghi, M.; Garajei, A.; Heidar, H.; Ghaderi, B.; Tadakamadla, J.; Meybodi, A.A.; Dallband, M.; Mostafavi, S.; et al. Association of N-acetyltransferases 1 and 2 Polymorphisms with Susceptibility to Head and Neck Cancers-A Meta-Analysis, Meta-Regression, and Trial Sequential Analysis. Medicina 2021, 57, 1095. [Google Scholar] [CrossRef]
- Mozaffari, H.R.; Rostamnia, M.; Sharifi, R.; Safaei, M.; Zavattaro, E.; Tadakamadla, S.K.; Imani, M.M.; Sadeghi, M.; Golshah, A.; Moradpoor, H.; et al. A PRISMA-compliant meta-analysis on association between X-ray repair cross complementing (XRCC1, XRCC2, and XRCC3) polymorphisms and oral cancer susceptibility. Gene 2021, 781, 145524. [Google Scholar] [CrossRef]
- Rezaei, F.; Mohammadi, H.; Heydari, M.; Sadeghi, M.; Mozaffari, H.R.; Khavid, A.; Godiny, M.; Brand, S.; Dürsteler, K.M.; Beatrix Brühl, A.; et al. Association between IL-8 (-251T/A) and IL-6 (-174G/C) Polymorphisms and Oral Cancer Susceptibility: A Systematic Review and Meta-Analysis. Medicina 2021, 57, 405. [Google Scholar] [CrossRef]
- Lee, T.-H.; Kang, T.-H. DNA oxidation and excision repair pathways. Int. J. Mol. Sci. 2019, 20, 6092. [Google Scholar] [CrossRef]
- Huang, R.; Zhou, P.-K. DNA damage repair: Historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct. Target. Ther. 2021, 6, 254. [Google Scholar] [CrossRef] [PubMed]
- Kokic, G.; Chernev, A.; Tegunov, D.; Dienemann, C.; Urlaub, H.; Cramer, P. Structural basis of TFIIH activation for nucleotide excision repair. Nat. Commun. 2019, 10, 2885. [Google Scholar] [CrossRef] [PubMed]
- Okamura, K.; Toyoda, M.; Hata, K.; Nakabayashi, K.; Umezawa, A. Whole-exome sequencing of fibroblast and its iPS cell lines derived from a patient diagnosed with xeroderma pigmentosum. Genom. Data 2015, 6, 4–6. [Google Scholar] [CrossRef]
- Zebian, A.; Shaito, A.; Mazurier, F.; Rezvani, H.R.; Zibara, K. XPC beyond nucleotide excision repair and skin cancers. Mutat. Res. Rev. Mutat. Res. 2019, 782, 108286. [Google Scholar] [CrossRef] [PubMed]
- Bootsma, D.; Hoeijmakers, J.H. The genetic basis of xeroderma pigmentosum. Ann. Genet. 1991, 34, 143–150. [Google Scholar]
- Nagaria, P.; Robert, C.; Rassool, F.V. DNA double-strand break response in stem cells: Mechanisms to maintain genomic integrity. Biochim. Biophys. Acta (BBA)-Gen. Subj. 2013, 1830, 2345–2353. [Google Scholar] [CrossRef]
- Kumar, N.; Moreno, N.C.; Feltes, B.C.; Menck, C.F.; Houten, B.V. Cooperation and interplay between base and nucleotide excision repair pathways: From DNA lesions to proteins. Genet. Mol. Biol. 2020, 43, e20190104. [Google Scholar] [CrossRef] [PubMed]
- Tse, D.; Zhai, R.; Zhou, W.; Heist, R.S.; Asomaning, K.; Su, L.; Lynch, T.J.; Wain, J.C.; Christiani, D.C.; Liu, G. Polymorphisms of the NER pathway genes, ERCC1 and XPD are associated with esophageal adenocarcinoma risk. Cancer Causes Control 2008, 19, 1077–1083. [Google Scholar] [CrossRef]
- Machado, C.R.; Vieira-da-Rocha, J.P.; Mendes, I.C.; Rajão, M.A.; Marcello, L.; Bitar, M.; Drummond, M.G.; Grynberg, P.; Oliveira, D.A.; Marques, C. Nucleotide excision repair in T rypanosoma brucei: Specialization of transcription-coupled repair due to multigenic transcription. Mol. Microbiol. 2014, 92, 756–776. [Google Scholar] [CrossRef]
- McCullough, L.E.; Santella, R.M.; Cleveland, R.J.; Millikan, R.C.; Olshan, A.F.; North, K.E.; Bradshaw, P.T.; Eng, S.M.; Terry, M.B.; Shen, J. Polymorphisms in DNA repair genes, recreational physical activity and breast cancer risk. Int. J. Cancer 2014, 134, 654–663. [Google Scholar] [CrossRef]
- Jiang, H.-Y.; Zeng, Y.; Xu, W.-D.; Liu, C.; Wang, Y.-J.; Wang, Y.-D. Genetic association between the XPG Asp1104His polymorphism and head and neck cancer susceptibility: Evidence based on a meta-analysis. Asian Pac. J. Cancer Prev. 2015, 16, 3645–3651. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Ling, H.; Lu, Y.; Wu, X.; Cai, M.; Su, B.; Zou, Y. Meta-analysis of the relationship between excision repair cross-complementing Group 5 rs17655 gene polymorphism and head and neck cancer susceptibility. J. Cancer Res. Ther. 2018, 14, S1041–S1047. [Google Scholar] [PubMed]
- Wu, L.; Gao, X.; Ye, D.; Ding, Y.; Yang, X.; Liu, W. Association of the XPA A23G polymorphism with the risk of head and neck carcinomas: Evidence from 5491 subjects. Mol. Clin. Oncol. 2015, 3, 649–654. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xu, J.; Yang, X.; Wu, Y.; Cheng, B.; Chen, D.; Bai, B. Association of single nucleotide polymorphisms of nucleotide excision repair genes with laryngeal cancer risk and interaction with cigarette smoking and alcohol drinking. Tumor Biol. 2014, 35, 4659–4665. [Google Scholar] [CrossRef]
- Avci, H.; Iplik, E.S.; Aydemir, L.; Acar, S.; Kiyak, E.; Ünür, M.; Cakmakoglu, B. Are XPD and XPG gene variants related to the mechanism of oral squamous cell carcinoma? Cell. Mol. Biol. 2018, 64, 94–99. [Google Scholar] [CrossRef]
- Hall, J.; Hashibe, M.; Boffetta, P.; Gaborieau, V.; Moullan, N.; Chabrier, A.; Zaridze, D.; Shangina, O.; Szeszenia-Dabrowska, N.; Mates, D. The association of sequence variants in DNA repair and cell cycle genes with cancers of the upper aerodigestive tract. Carcinogenesis 2007, 28, 665–671. [Google Scholar] [CrossRef]
- Lu, B.; Li, J.; Gao, Q.; Yu, W.; Yang, Q.; Li, X. Laryngeal cancer risk and common single nucleotide polymorphisms in nucleotide excision repair pathway genes ERCC1, ERCC2, ERCC3, ERCC4, ERCC5 and XPA. Gene 2014, 542, 64–68. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. Int. J. Surg. 2021, 88, 105906. [Google Scholar] [CrossRef]
- Peterson, J.; Welch, V.; Losos, M.; Tugwell, P. The Newcastle-Ottawa scale (NOS) for assessing the quality of nonrandomised studies in meta-analyses. Ott. Ott. Hosp. Res. Inst. 2011, 2, 1–12. [Google Scholar]
- DerSimonian, R.; Laird, N. Meta-analysis in clinical trials revisited. Contemp. Clin. Trials 2015, 45, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Mantel, N.; Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 1959, 22, 719–748. [Google Scholar]
- Imberger, G.; Thorlund, K.; Gluud, C.; Wetterslev, J. False-positive findings in Cochrane meta-analyses with and without application of trial sequential analysis: An empirical review. BMJ Open 2016, 6, e011890. [Google Scholar] [CrossRef]
- Wetterslev, J.; Jakobsen, J.C.; Gluud, C. Trial sequential analysis in systematic reviews with meta-analysis. BMC Med. Res. Methodol. 2017, 17, 39. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, R.; Ramroth, H.; Becher, H.; Dietz, A.; Schmezer, P.; Popanda, O. Laryngeal cancer risk associated with smoking and alcohol consumption is modified by genetic polymorphisms in ERCC5, ERCC6 and RAD23B but not by polymorphisms in five other nucleotide excision repair genes. Int. J. Cancer 2009, 125, 1431–1439. [Google Scholar] [CrossRef]
- An, J.; Liu, Z.; Hu, Z.; Li, G.; Wang, L.-E.; Sturgis, E.M.; El-Naggar, A.K.; Spitz, M.R.; Wei, Q. Potentially functional single nucleotide polymorphisms in the core nucleotide excision repair genes and risk of squamous cell carcinoma of the head and neck. Cancer Epidemiol. Biomark. Prev. 2007, 16, 1633–1638. [Google Scholar] [CrossRef]
- Bau, D.; Tsai, M.; Huang, C.; Lee, C.; Tseng, H.; Lo, Y.; Tsai, Y.; Tsai, F. Relationship between polymorphisms of nucleotide excision repair genes and oral cancer risk in Taiwan: Evidence for modification of smoking habit. Chin. J. Physiol. 2007, 50, 294. [Google Scholar] [PubMed]
- Cui, Y.; Morgenstern, H.; Greenland, S.; Tashkin, D.P.; Mao, J.; Cao, W.; Cozen, W.; Mack, T.M.; Zhang, Z.F. Polymorphism of Xeroderma Pigmentosum group G and the risk of lung cancer and squamous cell carcinomas of the oropharynx, larynx and esophagus. Int. J. Cancer 2006, 118, 714–720. [Google Scholar] [CrossRef] [PubMed]
- Jelonek, K.; Gdowicz-Kłosok, A.; Pietrowska, M.; Borkowska, M.; Korfanty, J.; Rzeszowska-Wolny, J.; Widłak, P. Association between single-nucleotide polymorphisms of selected genes involved in the response to DNA damage and risk of colon, head and neck, and breast cancers in a Polish population. J. Appl. Genet. 2010, 51, 343–352. [Google Scholar] [CrossRef]
- Ma, H.; Yu, H.; Liu, Z.; Wang, L.-E.; Sturgis, E.M.; Wei, Q. Polymorphisms of XPG/ERCC5 and risk of squamous cell carcinoma of the head and neck. Pharmacogenet. Genom. 2012, 22, 50. [Google Scholar] [CrossRef]
- Nigam, K.; Yadav, S.K.; Samadi, F.M.; Bhatt, M.L.; Gupta, S.; Sanyal, S. Risk modulation of oral pre cancer and cancer with polymorphisms in XPD and XPG genes in North Indian population. Asian Pac. J. Cancer Prev. APJCP 2019, 20, 2397. [Google Scholar] [CrossRef]
- Sugimura, T.; Kumimoto, H.; Tohnai, I.; Fukui, T.; Matsuo, K.; Tsurusako, S.; Mitsudo, K.; Ueda, M.; Tajima, K.; Ishizaki, K. Gene–environment interaction involved in oral carcinogenesis: Molecular epidemiological study for metabolic and DNA repair gene polymorphisms. J. Oral Pathol. Med. 2006, 35, 11–18. [Google Scholar] [CrossRef]
- Sun, Y.; Tan, L.; Li, H.; Qin, X.; Liu, J. Association of NER pathway gene polymorphisms with susceptibility to laryngeal cancer in a Chinese population. Int. J. Clin. Exp. Pathol. 2015, 8, 11615. [Google Scholar]
- Wen, S.X.; Tang, P.Z.; Zhang, X.M.; Zhao, D.; Guo, Y.L.; Tan, W.; Lin, D.X. Association between genetic polymorphism in xeroderma pigmentosum G gene and risks of laryngeal and hypopharyngeal carcinomas. Zhongguo Yi Xue Ke Xue Yuan Xue Bao. Acta Acad. Med. Sin. 2006, 28, 703–706. [Google Scholar]
- Xue, M.; Qiu-xu, W.; Mo-ye, C.; Gang, Q.; Yu-peng, W. DNA repair gene polymorphisms in ERCC4 rs6498486 and ERCC5 rs751402 and risk of salivary gland tumors. Shanghai J. Stomatol. 2013, 22, 438. [Google Scholar]
- Yuan, H.; Li, H.; Ma, H.; Niu, Y.; Wu, Y.; Zhang, S.; Hu, Z.; Shen, H.; Chen, N. Genetic polymorphisms in key DNA repair genes and risk of head and neck cancer in a Chinese population. Exp. Ther. Med. 2012, 3, 719–724. [Google Scholar] [CrossRef]
- Zavras, A.I.; Yoon, A.J.; Chen, M.-K.; Lin, C.-W.; Yang, S.-F. Association between polymorphisms of DNA repair gene ERCC5 and oral squamous cell carcinoma. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2012, 114, 624–629. [Google Scholar] [CrossRef]
- Zhu, Y.; Guo, L.; Wang, S.; Yu, Q.; Lu, J. Association of Smoking and XPG, CYP1A1, OGG1, ERCC5, ERCC1, MMP2, and MMP9 Gene Polymorphisms with the early detection and occurrence of Laryngeal Squamous Carcinoma. J. Cancer 2018, 9, 968. [Google Scholar] [CrossRef]
- Yu, H.; Liu, Z.; Huang, Y.J.; Yin, M.; Wang, L.E.; Wei, Q. Association between single nucleotide polymorphisms in ERCC4 and risk of squamous cell carcinoma of the head and neck. PLoS ONE 2012, 7, e41853. [Google Scholar] [CrossRef]
- Kamileri, I.; Karakasilioti, I.; Garinis, G.A. Nucleotide excision repair: New tricks with old bricks. Trends Genet. 2012, 28, 566–573. [Google Scholar] [CrossRef]
- Wood, R.D.; Mitchell, M.; Lindahl, T. Human DNA repair genes, 2005. Mutat. Res./Fundam. Mol. Mech. Mutagen. 2005, 577, 275–283. [Google Scholar] [CrossRef]
- Petruseva, I.; Evdokimov, A.; Lavrik, O. Molecular mechanism of global genome nucleotide excision repair. Acta Nat. 2014, 6, 23–34. [Google Scholar] [CrossRef]
- Liu, J.; He, C.; Xing, C.; Yuan, Y. Nucleotide excision repair related gene polymorphisms and genetic susceptibility, chemotherapeutic sensitivity and prognosis of gastric cancer. Mutat. Res./Fundam. Mol. Mech. Mutagen. 2014, 765, 11–21. [Google Scholar] [CrossRef]
- Xue, M.-H.; Li, G.-Y.; Wu, X.-J.; Zhang, C.-X.; Zhang, C.-F.; Zhu, K.-X. Genetic variability of genes in NER pathway influences the treatment outcome of gastric cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 5563. [Google Scholar]
- Shuck, S.C.; Short, E.A.; Turchi, J.J. Eukaryotic nucleotide excision repair: From understanding mechanisms to influencing biology. Cell Res. 2008, 18, 64–72. [Google Scholar] [CrossRef]
- d’Errico, M.; Parlanti, E.; Teson, M.; de Jesus, B.M.B.; Degan, P.; Calcagnile, A.; Jaruga, P.; Bjørås, M.; Crescenzi, M.; Pedrini, A.M. New functions of XPC in the protection of human skin cells from oxidative damage. EMBO J. 2006, 25, 4305–4315. [Google Scholar] [CrossRef]
- Niedernhofer, L.J.; Odijk, H.; Budzowska, M.; Van Drunen, E.; Maas, A.; Theil, A.F.; De Wit, J.; Jaspers, N.; Beverloo, H.B.; Hoeijmakers, J.H. The structure-specific endonuclease Ercc1-Xpf is required to resolve DNA interstrand cross-link-induced double-strand breaks. Mol. Cell. Biol. 2004, 24, 5776–5787. [Google Scholar] [CrossRef]
- Nagai, A.; Saijo, M.; Kuraoka, I.; Matsuda, T.; Kodo, N.; Nakatsu, Y.; Mimaki, T.; Mino, M.; Biggerstaff, M.; Wood, R.D. Enhancement of damage-specific DNA binding of XPA by interaction with the ERCC1 DNA repair protein. Biochem. Biophys. Res. Commun. 1995, 211, 960–966. [Google Scholar] [CrossRef]
- Nocentini, S.; Coin, F.; Saijo, M.; Tanaka, K.; Egly, J.-M. DNA damage recognition by XPA protein promotes efficient recruitment of transcription factor II H. J. Biol. Chem. 1997, 272, 22991–22994. [Google Scholar] [CrossRef]
- Patrick, S.M.; Turchi, J.J. Xeroderma pigmentosum complementation group A protein (XPA) modulates RPA-DNA interactions via enhanced complex stability and inhibition of strand separation activity. J. Biol. Chem. 2002, 277, 16096–16101. [Google Scholar] [CrossRef]
- Tudek, B.; Winczura, A.; Janik, J.; Siomek, A.; Foksinski, M.; Oliński, R. Involvement of oxidatively damaged DNA and repair in cancer development and aging. Am. J. Transl. Res. 2010, 2, 254. [Google Scholar]
- Liu, J.; Zhang, Z.; Cao, X.-L.; Lei, D.-P.; Wang, Z.-Q.; Jin, T.; Pan, X.-L. XPA A23G polymorphism and susceptibility to cancer: A meta-analysis. Mol. Biol. Rep. 2012, 39, 6791–6799. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Liu, X.; Tang, L.-L.; Long, J.-T.; Zhu, J.; Hua, R.-X.; Li, J. XPG gene polymorphisms and cancer susceptibility: Evidence from 47 studies. Oncotarget 2017, 8, 37263. [Google Scholar] [CrossRef] [PubMed]
- Rodrigo, J.P.; Suárez, C.; González, M.V.; Lazo, P.S.; Ramos, S.; Coto, E.; Alvarez, I.; García, L.A.; Martínez, J.A. Variability of genetic alterations in different sites of head and neck cancer. Laryngoscope 2001, 111, 1297–1301. [Google Scholar] [CrossRef]
| The Study, Publication Year | Country | Ethnicity | Number of Cases/Controls | Control Source | Polymorphism: HWE p-Value in Controls | Genotyping Method | Tumor Site | Quality Score |
|---|---|---|---|---|---|---|---|---|
| Abbasi, 2009 [34] | Germany | Caucasian | 248/647 | PB | rs17655: <0.0001 rs1047768: 0.7616 rs1800975: 0.7373 | PCR-RFLP | LC | 8 |
| An, 2007 [35] | USA | Caucasian | 829/854 | HB | rs1800975: 0.0010 rs17655: 0.4245 | PCR | HNC | 8 |
| Avci, 2018 [25] | Turkey | Caucasian | 111/148 | PB | rs17655: 0.1716 | PCR | OC | 9 |
| Bau, 2007 [36] | Taiwan | Asian | 154/105 | PB | rs1800975: 0.8954 | PCR | OC | 8 |
| Cui, 2006 [37] | USA | Mixed | 443/911 | PB | rs17655: <0.0001 | PCR | NPC | 9 |
| Hall, 2007 [26] | France | Caucasian | 597/770 | HB | rs1800975: 0.2248 | PCR | HNC | 8 |
| Jelonek, 2010 [38] | Poland | Caucasians | 66/113 | PB | rs1800975: 0.0551 | PCR | HNC | 6 |
| Li, 2014 [24] | China | Asian | 211/210 | HB | rs17655: 0.0003 rs1047768: 0.3326 rs2276465: 0.0023 rs2276466: 0.0970 rs6498486: 0.1517 rs4150441: 0.0034 rs1800975: <0.0001 | PCR | LC | 7 |
| Lu, 2014 [27] | China | Asian | 176/176 | HB | rs17655: 0.0007 rs2276465: 0.0071 rs6498486: 0.1479 rs4150441: 0.0127 rs1800975: <0.0001 | PCR | LC | 7 |
| Ma, 2012 [39] | USA | Mixed | 1059/1059 | PB | rs17655: 0.1749 rs1047768: 0.6694 rs2094258: 0.0920 rs4771436: 0.9424 | PCR-RFLP | HNC | 9 |
| Nigam, 2019 [40] | India | Asian | 67/288 | PB | rs17655: 0.7511 | PCR-RFLP | OC | 8 |
| Sugimura, 2006 [41] | Japan | Asian | 122/241 | HB | rs17655: <0.0001 rs1800975: 0.0496 | PCR | OC | 7 |
| Sun, 2015 [42] | China | Asian | 271/271 | HB | rs2094258: 0.8255 | PCR-RFLP | LC | 8 |
| Wen, 2006 [43] | China | Asian | 175/525 | HB | rs17655: 0.0026 | PCR-RFLP | NPC | 9 |
| Xue, 2013 [44] | China | Asian | 142/275 | HB | rs751402: 0.3033 rs6498486: 0.4273 | PCR-RFLP | OC | 8 |
| Yu, 2012 [48] | USA | Mixed | 1040/1046 | HB | rs2276466: 0.0636 | PCR-RFLP | HNC | 8 |
| Yuan, 2012 [45] | China | Asian | 394/884 | HB | rs17655: <0.0001 | PCR | HNC | 8 |
| Zavras, 2012 [46] | Taiwan | Asian | 239/336 | HB | rs751402: 0.3984 | TaqMan and PCR | OC | 6 |
| Zhu, 2018 [47] | China | Asian | 199/190 | HB | rs17655: 0.6655 rs1047768: 0.3839 rs4771436: 0.5694 | PCR | LC | 8 |
| Polymorphism (N) | Genetic Model | OR | 95%CI | Z-Value | p-Value | I2 | Pheterogeneity | |
|---|---|---|---|---|---|---|---|---|
| Min. | Max. | |||||||
| rs17655 (12) | C vs. G | 0.95 | 0.86 | 1.05 | 1.03 | 0.30 | 56% | 0.009 |
| CC vs. GG | 0.86 | 0.75 | 1.00 | 1.98 | 0.05 | 34% | 0.12 | |
| GC vs. GG | 1.26 | 0.94 | 1.71 | 1.53 | 0.13 | 85% | <0.00001 | |
| CC + GC vs. GG | 1.47 | 0.98 | 2.19 | 1.88 | 0.06 | 93% | <0.00001 | |
| CC vs. GG + GC | 0.89 | 0.81 | 0.99 | 2.14 | 0.03 | 31% | 0.14 | |
| rs751402 (2) | T vs. C | 1.28 | 1.05 | 1.57 | 2.38 | 0.02 | 0% | 0.55 |
| TT vs. CC | 1.74 | 1.10 | 2.74 | 2.39 | 0.02 | 0% | 0.75 | |
| CT vs. CC | 0.65 | 0.48 | 0.89 | 2.65 | 0.008 | 5% | 0.31 | |
| TT + CT vs. CC | 2.22 | 1.04 | 4.74 | 2.07 | 0.04 | 87% | 0.005 | |
| TT vs. CC + CT | 2.48 | 0.78 | 7.93 | 1.53 | 0.12 | 85% | 0.01 | |
| rs1047768 (4) | T vs. C | 0.92 | 0.74 | 1.13 | 0.81 | 0.42 | 72% | 0.01 |
| TT vs. CC | 0.91 | 0.64 | 1.31 | 0.50 | 0.62 | 60% | 0.06 | |
| CT vs. CC | 1.05 | 0.90 | 1.22 | 0.66 | 0.51 | 0% | 0.96 | |
| TT + CT vs. CC | 1.06 | 0.92 | 1.22 | 0.74 | 0.46 | 0% | 0.96 | |
| TT vs. CC + CT | 1.03 | 0.88 | 1.21 | 0.37 | 0.71 | 0% | 0.99 | |
| rs4771436 (2) | G vs. T | 1.02 | 0.89 | 1.16 | 0.25 | 0.81 | 0% | 0.68 |
| GG vs. TT | 1.03 | 0.73 | 1.44 | 0.15 | 0.88 | 0% | 0.61 | |
| TG vs. TT | 1.02 | 0.86 | 1.20 | 0.23 | 0.81 | 0% | 0.92 | |
| GG + TG vs. TT | 3.08 | 0.33 | 28.49 | 0.99 | 0.32 | 98% | <0.00001 | |
| GG vs. TT + TG | 1.02 | 0.73 | 1.42 | 0.11 | 0.91 | 0% | 0.62 | |
| rs2094258 (2) | A vs. G | 1.05 | 0.92 | 1.20 | 0.71 | 0.48 | 16% | 0.28 |
| AA vs. GG | 1.09 | 0.75 | 1.57 | 0.44 | 0.66 | 33% | 0.22 | |
| GA vs. GG | 1.06 | 0.89 | 1.25 | 0.65 | 0.51 | 0% | 0.64 | |
| AA + GA vs. GG | 1.06 | 0.9 | 1.24 | 0.69 | 0.49 | 0% | 0.42 | |
| AA vs. GG + GA | 1.06 | 0.74 | 1.53 | 0.33 | 0.74 | 22% | 0.26 | |
| rs6498486 (3) | C vs. A | 1.16 | 0.96 | 1.41 | 1.56 | 0.12 | 0% | 0.93 |
| CC vs. AA | 1.36 | 0.89 | 2.09 | 1.41 | 0.16 | 0% | 0.96 | |
| AC vs. AA | 1.13 | 0.87 | 1.46 | 0.92 | 0.36 | 0% | 0.99 | |
| CC + AC vs. AA | 1.17 | 0.92 | 1.50 | 1.28 | 0.20 | 0% | 0.97 | |
| CC vs. AA + AC | 1.29 | 0.86 | 1.95 | 1.22 | 0.22 | 0% | 0.96 | |
| rs2276465 (2) | C vs. A | 1.00 | 0.67 | 1.47 | 0.02 | 0.98 | 71% | 0.07 |
| CC vs. AA | 1.39 | 0.94 | 2.06 | 1.64 | 0.10 | 0% | 0.97 | |
| AC vs. AA | 1.16 | 0.85 | 1.59 | 0.93 | 0.35 | 0% | 0.96 | |
| CC vs. AC + AA | 0.74 | 0.28 | 1.96 | 0.60 | 0.55 | 92% | 0.0003 | |
| CC vs. AA + AC | 0.93 | 0.48 | 1.80 | 0.22 | 0.83 | 69% | 0.07 | |
| rs2276466 (2) | G vs. C | 0.96 | 0.85 | 1.09 | 0.58 | 0.56 | 0% | 0.39 |
| GG vs. CC | 0.30 | 0.02 | 4.54 | 0.87 | 0.39 | 98% | <0.00001 | |
| CG vs. CC | 1.08 | 0.92 | 1.28 | 0.94 | 0.35 | 0% | 0.87 | |
| GG + CG vs. CC | 1.03 | 0.88 | 1.20 | 0.32 | 0.75 | 0% | 0.77 | |
| GG vs. CC + CG | 0.84 | 0.50 | 1.39 | 0.69 | 0.49 | 50% | 0.16 | |
| rs4150441 (2) | G vs. A | 1.09 | 0.88 | 1.34 | 0.80 | 0.43 | 0% | 0.85 |
| GG vs. AA | 1.20 | 0.80 | 1.80 | 0.90 | 0.37 | 0% | 0.87 | |
| AG vs. AA | 1.12 | 0.82 | 1.53 | 0.70 | 0.48 | 0% | 0.93 | |
| GG + AG vs. AA | 1.26 | 0.95 | 1.67 | 1.59 | 0.11 | 0% | 0.47 | |
| GG vs. AA + AG | 1.08 | 0.74 | 1.57 | 0.38 | 0.70 | 0% | 0.88 | |
| rs1800975 (8) | A vs. G | 0.78 | 0.49 | 1.23 | 1.07 | 0.28 | 96% | <0.00001 |
| AA vs. GG | 0.91 | 0.77 | 1.07 | 1.11 | 0.27 | 17% | 0.29 | |
| GA vs. GG | 1.00 | 0.82 | 1.23 | 0.02 | 0.99 | 55% | 0.03 | |
| AA + GA vs. GG | 0.94 | 0.85 | 1.06 | 1.01 | 0.31 | 43% | 0.09 | |
| AA vs. GG + GA | 0.66 | 0.35 | 1.26 | 1.26 | 0.21 | 94% | <0.00001 | |
| Polymorphism (N) | Subgroup (N) | Variables | Allelic | Homozygous | Heterozygous | Dominant | Recessive |
|---|---|---|---|---|---|---|---|
| rs17655 (12) | Ethnicity | ||||||
| Asian (7) | OR (95%CI) | 0.87 (0.79, 0.97) | 0.81 (0.67, 0.97) | 1.05 (0.67, 1.65) | 1.22 (0.66, 2.66) | 0.86 (0.68, 1.09) | |
| p-value | 0.009 | 0.03 | 0.83 | 0.52 | 0.22 | ||
| I2 | 67% | 50% | 85% | 93% | 52% | ||
| Caucasian (2) | OR (95%CI) | 1.01 (0.82, 1.24) | 0.68 (0.37, 1.26) | 1.98 (0.97, 4.06) | 1.47 (0.98, 2.19) | 0.64 (0.35, 1.17) | |
| p-value | 0.23 | 0.22 | 0.06 | 0.06 | 0.15 | ||
| I2 | 0% | 0% | 81% | 93% | 0% | ||
| Mixed (3) | OR (95%CI) | 1.05 (0.96, 1.15) | 1.01 (0.79, 1.28) | 1.39 (0.85, 2.28) | 1.76 (0.70, 4.42) | 0.93 (0.81, 1.07) | |
| p-value | 0.30 | 0.95 | 0.19 | 0.23 | 0.31 | ||
| I2 | 19% | 0% | 82% | 96% | 0% | ||
| Sample size | |||||||
| ≥400 (6) | OR (95%CI) | 0.99 (0.92, 1.06) | 0.89 (0.75, 1.05) | 1.36 (0.90, 2.60) | 1.74 (1.03, 2.93) | 0.91 (0.82, 1.02) | |
| p-value | 0.82 | 0.15 | 0.15 | 0.04 | 0.11 | ||
| I2 | 53% | 28% | 90% | 95% | 36% | ||
| <400 (6) | OR (95%CI) | 0.89 (0.77, 1.03) | 0.80 (0.60, 1.07) | 1.12 (0.74, 1.69) | 1.14 (0.60, 2.18) | 0.82 (0.64, 1.04) | |
| p-value | 0.11 | 0.13 | 0.60 | 0.69 | 0.10 | ||
| I2 | 61% | 50% | 65% | 88% | 32% | ||
| Control source | |||||||
| HB (7) | OR (95%CI) | 0.90 (0.77, 1.05) | 0.86 (0.64, 1.15) | 1.13 (0.73, 1.73) | 0.32 (0.74, 2.36) | 0.90 (0.73, 1.10) | |
| p-value | 0.18 | 0.31 | 0.59 | <0.0001 | 0.29 | ||
| I2 | 67% | 60% | 84% | 93% | 56% | ||
| PB (5) | OR (95%CI) | 1.03 (0.93, 1.13) | 0.86 (0.67, 1.10) | 1.47 (0.89, 2.43) | 1.69 (0.87, 3.28) | 0.83 (0.69, 0.99) | |
| p-value | 0.56 | 0.22 | 0.13 | 0.12 | 0.04 | ||
| I2 | 1% | 0% | 89% | 95% | 0% | ||
| Cancer subtype | |||||||
| OC (3) | OR (95%CI) | 1.01 (0.82, 1.25) | 0.82 (0.59, 1.15) | 1.37 (0.71, 2.66) | 1.62 (0.65, 4.07) | 1.02 (0.71, 1.48) | |
| p-value | 0.91 | 0.25 | 0.35 | 0.30 | 0.90 | ||
| I2 | 10% | 0% | 75% | 89% | 0% | ||
| LC (4) | OR (95%CI) | 0.80 (0.62, 1.04) | 0.60 (0.45, 0.76) | 1.11 (0.54, 2.29) | 0.90 (0.43, 1.89) | 0.64 (0.51, 0.82) | |
| p-value | 0.10 | 0.0004 | 0.77 | 0.78 | 0.0003 | ||
| I2 | 73% | 20% | 90% | 93% | 0% | ||
| NPC (2) | OR (95%CI) | 0.99 (0.94, 1.04) | 0.98 (0.71, 1.35) | 1.21 (0.35, 4.19) | 2.67 (1.08, 6.63) | 0.93 (0.77, 1.12) | |
| p-value | 0.68 | 0.90 | 0.77 | 0.03 | 0.43 | ||
| I2 | 84% | 0% | 94% | 92% | 3% | ||
| rs1800975 (8) | Ethnicity | ||||||
| Asian (4) | OR (95%CI) | 0.99 (0.85, 1.15) | 0.97 (0.73, 1.29) | 1.22 (0.94, 2.95) | 1.08 (0.86, 1.35) | 0.89 (0.69, 1.14) | |
| p-value | 0.88 | 0.86 | 0.13 | 0.53 | 0.35 | ||
| I2 | 33% | 24% | 0% | 0% | 48% | ||
| Caucasian (3) | OR (95%CI) | 0.53 (0.18, 1.59) | 0.83 (0.52, 1.32) | 0.80 (0.58, 1.10) | 0.81 (0.69, 0.96) | 0.26 (0.21, 0.33) | |
| p-value | <0.0001 | 0.43 | 0.17 | 0.02 | <0.0001 | ||
| I2 | 98% | 52% | 60% | 66% | 96% | ||
| Mixed (1) | OR (95%CI) | 0.99 (0.86, 1.14) | 0.91 (0.68, 1.22) | 1.10 (0.90, 1.35) | 1.05 (0.87, 1.27) | 0.87 (0.66, 1.14) | |
| p-value | 0.87 | 0.53 | 0.36 | 0.62 | 0.31 | ||
| I2 | - | - | - | - | - | ||
| Sample size | |||||||
| ≥400 (4) | OR (95%CI) | 0.72 (0.33, 1.57) | 0.95 (0.79, 1.15) | 0.97 (0.76, 1.24) | 0.97 (0.78, 1.21) | 0.63 (0.22, 1.82) | |
| p-value | 0.41 | 0.62 | 0.81 | 0.81 | 0.39 | ||
| I2 | 98% | 0% | 98% | 64% | 97% | ||
| <400 (4) | OR (95%CI) | 0.87 (0.74, 1.03) | 0.79 (0.57, 1.10) | 1.08 (0.81, 1.44) | 0.90 (0.67, 1.22) | 0.76 (0.57, 1.01) | |
| p-value | 0.12 | 0.17 | 0.59 | 0.50 | 0.06 | ||
| I2 | 44% | 45% | 48% | 23% | 43% | ||
| Control source | |||||||
| HB (5) | OR (95%CI) | 0.73 (0.38, 1.43) | 0.91 (0.76, 1.09) | 1.04 (0.79, 1.38) | 0.96 (0.79, 1.17) | 0.44 (0.37, 0.52) | |
| p-value | 0.36 | 0.30 | 0.75 | 0.67 | <0.0001 | ||
| I2 | 98% | 11% | 69% | 51% | 95% | ||
| PB (3) | OR (95%CI) | 0.91 (0.68, 1.20) | 0.92 (0.63, 1.35) | 0.97 (0.75, 1.26) | 0.89 (0.59, 1.35) | 0.83 (0.64, 1.08) | |
| p-value | 0.50 | 0.68 | 0.83 | 0.59 | 0.17 | ||
| I2 | 54% | 49% | 32% | 51% | 31% | ||
| Cancer subtype | |||||||
| OC (2) | OR (95%CI) | 084 (0.67, 1.06) | 0.70 (0.44, 1.13) | 1.41 (0.93, 2.15) | 0.96(0.66, 1.41) | 0.66 (0.45, 0.96) | |
| p-value | 0.14 | 0.15 | 0.11 | 0.84 | 0.03 | ||
| I2 | 4% | 8% | 0% | 0% | 36% | ||
| LC (3) | OR (95%CI) | 1.09 (0.94, 1.27) | 1.16 (0.87, 1.55) | 1.09 (0.87, 1.36) | 1.11 (0.90, 1.36) | 1.12 (0.85, 1.46) | |
| p-value | 0.24 | 0.31 | 0.47 | 0.32 | 0.42 | ||
| I2 | 0% | 0% | 0% | 0% | 0% |
| Polymorphism (N) | Variable | Model | Coefficient | Standard Error | 95% Lower | 95% Upper | Z-Value | p-Value |
|---|---|---|---|---|---|---|---|---|
| rs17655 (12) | Publication year | C vs. G | −0.0005 | 0.0003 | −0.0011 | 0.0001 | −1.57 | 0.1171 |
| CC vs. GG | −0.0005 | 0.0006 | −0.0016 | 0.0006 | −0.93 | 0.3518 | ||
| GC vs. GG | 0.0003 | 0.0010 | −0.0017 | 0.0023 | 0.28 | 0.7816 | ||
| CC + GC vs. GG | −0.0007 | 0.0014 | −0.0033 | 0.0020 | −0.50 | 0.6152 | ||
| CC vs. GG + GC | −0.0005 | 0.0004 | −0.0014 | 0.0003 | −1.18 | 0.2369 | ||
| Sample size | C vs. G | 0.0001 | 0.0001 | −0.0001 | 0.0002 | 0.66 | 0.5112 | |
| CC vs. GG | 0.0001 | 0.0002 | −0.0002 | 0.0005 | 0.66 | 0.5084 | ||
| GC vs. GG | 0.0002 | 0.0003 | −0.0004 | 0.0009 | 0.72 | 0.4708 | ||
| CC + GC vs. GG | 0.0002 | 0.0004 | −0.0007 | 0.0011 | 0.40 | 0.6867 | ||
| CC vs. GG + GC | < 0.0001 | 0.0001 | −0.0003 | 0.0003 | 0.22 | 0.8232 | ||
| Quality score | C vs. G | 0.1013 | 0.0790 | −0.0535 | 0.2561 | 1.28 | 0.1995 | |
| CC vs. GG | 0.0961 | 0.1522 | −0.2022 | 0.3944 | 0.63 | 0.5278 | ||
| GC vs. GG | −0.0671 | 0.2663 | −0.5890 | 0.4548 | −0.25 | 0.8010 | ||
| CC + GC vs. GG | 0.1956 | 0.3539 | −0.4971 | 0.8902 | 0.56 | 0.5786 | ||
| CC vs. GG + GC | 0.1062 | 0.1169 | −0.1229 | 0.3352 | 0.91 | 0.3636 | ||
| rs1800975 (8) | Publication year | A vs. G | −0.0006 | 0.0018 | −0.0014 | 0.0030 | −0.32 | 0.7519 |
| AA vs. GG | −0.0011 | 0.0010 | −0.0031 | 0.0010 | −1.04 | 0.2948 | ||
| GA vs. GG | −0.0007 | 0.0009 | −0.0025 | 0.0012 | −0.71 | 0.4793 | ||
| AA + GA vs. GG | −0.0008 | 0.0008 | −0.0024 | 0.0007 | −1.03 | 0.3045 | ||
| AA vs. GG + GA | −0.0010 | 0.0025 | −0.0059 | 0.0039 | −0.40 | 0.6859 | ||
| Sample size | A vs. G | −0.0005 | 0.0007 | −0.0018 | 0.0009 | −0.67 | 0.5021 | |
| AA vs. GG | −0.0002 | 0.0003 | −0.0008 | 0.0004 | −0.70 | 0.4845 | ||
| GA vs. GG | −0.0003 | 0.0003 | −0.0009 | 0.0004 | −0.83 | 0.4070 | ||
| AA + GA vs. GG | −0.0002 | 0.0003 | −0.0007 | 0.0003 | −0.78 | 0.4328 | ||
| AA vs. GG + GA | −0.0006 | 0.0009 | −0.0024 | 0.0012 | −0.66 | 0.5120 | ||
| Quality score | A vs. G | 0.1647 | 0.5359 | −0.8830 | 1.2178 | 0.31 | 0.7548 | |
| AA vs. GG | 0.2972 | 0.2988 | −0.2883 | 0.8828 | 0.99 | 0.3198 | ||
| GA vs. GG | 0.2057 | 0.2717 | −0.3268 | 0.7381 | 0.76 | 0.4490 | ||
| AA + GA vs. GG | 0.2352 | 0.2339 | −0.2232 | 0.6936 | 1.01 | 0.3145 | ||
| AA vs. GG + GA | 0.2765 | 0.7316 | −1.1574 | 1.7104 | 0.38 | 0.7055 |
| Polymorphism (Number of Studies without a Deviation) | Genetic Model | OR | 95%CI | Z-Value | p-Value | I2 | Pheterogeneity | |
|---|---|---|---|---|---|---|---|---|
| Min. | Max. | |||||||
| rs17655 (5) | C vs. G | 0.99 | 0.90 | 1.09 | 0.25 | 0.81 | 0% | 0.85 |
| CC vs. GG | 0.95 | 0.75 | 1.21 | 0.42 | 0.68 | 0% | 0.69 | |
| GC vs. GG | 1.03 | 0.89 | 1.20 | 0.42 | 0.68 | 0% | 0.59 | |
| CC + GC vs. GG | 1.00 | 0.87 | 1.15 | 0.01 | 0.99 | 30% | 0.22 | |
| CC vs. GG + GC | 0.96 | 0.82 | 1.12 | 0.54 | 0.59 | 0% | 0.89 | |
| rs1800975 (4) | A vs. G | 0.62 | 0.26 | 1.49 | 1.08 | 0.28 | 98% | <0.00001 |
| AA vs. GG | 0.87 | 0.68 | 1.12 | 1.08 | 0.28 | 29% | 0.24 | |
| GA vs. GG | 0.84 | 0.63 | 1.12 | 1.20 | 0.23 | 51% | 0.11 | |
| AA + GA vs. GG | 0.84 | 0.63 | 1.11 | 1.22 | 0.22 | 55% | 0.08 | |
| AA vs. GG + GA | 0.49 | 0.15 | 1.63 | 1.16 | 0.25 | 96% | <0.00001 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Imani, M.M.; Basamtabar, M.; Akbari, S.; Sadeghi, E.; Sadeghi, M. Relationship between XPA, XPB/ERCC3, XPF/ERCC4, and XPG/ERCC5 Polymorphisms and the Susceptibility to Head and Neck Carcinoma: A Systematic Review, Meta-Analysis, and Trial Sequential Analysis. Medicina 2024, 60, 478. https://doi.org/10.3390/medicina60030478
Imani MM, Basamtabar M, Akbari S, Sadeghi E, Sadeghi M. Relationship between XPA, XPB/ERCC3, XPF/ERCC4, and XPG/ERCC5 Polymorphisms and the Susceptibility to Head and Neck Carcinoma: A Systematic Review, Meta-Analysis, and Trial Sequential Analysis. Medicina. 2024; 60(3):478. https://doi.org/10.3390/medicina60030478
Chicago/Turabian StyleImani, Mohammad Moslem, Masoumeh Basamtabar, Sattar Akbari, Edris Sadeghi, and Masoud Sadeghi. 2024. "Relationship between XPA, XPB/ERCC3, XPF/ERCC4, and XPG/ERCC5 Polymorphisms and the Susceptibility to Head and Neck Carcinoma: A Systematic Review, Meta-Analysis, and Trial Sequential Analysis" Medicina 60, no. 3: 478. https://doi.org/10.3390/medicina60030478





