Abstract
The genus Cinnamomum exhibits a rich variety of chemotypes and is an economically important essential oil (EO)-producing plant belonging to the family Lauraceae. Here, we aimed to explore the potential differences in the terpenoid (the principal components of EOs) biosynthesis pathways of different chemotypes at the molecular level in four Cinnamomum species—C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum. Gas chromatography–mass spectrometry (GC-MS) was employed to elucidate the discrepancies in the chemical profiles and compositions of leaf EO terpenoids among the four Cinnamomum species. The results revealed significant variations in leaf EO yields. The main constituents of the leaf EOs from C. camphora var. linaloolifera and C. kanehirae were the acyclic monoterpene linalool, and those of C. longipaniculatum and C. micranthum were the monoterpene eucalyptol and the sesquiterpene β-caryophyllene, respectively. Furthermore, a comparative transcriptome analysis of the leaves from the four Cinnamomum species revealed that differentially expressed genes (DEGs) were significantly enriched in terpene-related entries. Specifically, 42 and 24 DEGs were significantly enriched to the mevalonate (MVA)/2-methylerythritol 4-phosphate (MEP) pathways and terpene synthase (TPS) activity, respectively. Most genes encoding proteins involved in the terpenoid precursor MVA and MEP pathways exhibited differential expression across the four species, which correlated with the distinct terpenoid profiles observed in their leaf EOs. Four acyclic monoterpene linalool synthase genes—Maker00024100, Maker00014813, Maker00014818, and Maker00018424—were highly expressed in C. camphora var. linaloolifera and C. kanehirae. A monoterpene eucalyptol synthesis gene, Maker00001509, was highly expressed in C. longipaniculatum, and a sesquiterpene β-stigmasterol synthesis gene, Maker00005791, was highly expressed in C. micranthum. These expression levels were subsequently validated through quantitative real-time polymerase chain reaction (qRT-PCR). In conclusion, the combined results of the GC-MS and transcriptome analyses revealed a strong correlation between the metabolite content of the EOs and gene expression. This research contributes to a better understanding of the differences in terpene accumulation in various chemotypes of Cinnamomum at the molecular and mechanistic levels, laying a solid foundation for the cultivation of an ideal Cinnamomum variety.
1. Introduction
The Cinnamomum genus comprises broad-leaved evergreen plant species belonging to the Lauraceae family; it is valued for its timber and aromatic properties and has been cultivated in China for at least 2000 years [1]. Several commercially important EOs are derived from Cinnamomum species, including C. camphora, C. kanehirae, C. longipaniculatum, and C. micranthum [2,3]. EOs are aromatic oily liquids extracted from a variety of plant materials, including leaves, roots, fruits, flowers, seeds, twigs, and bark [4]. They are commercially significant, and are primarily used in pharmaceuticals, foodstuffs, biochemicals, flavors, and fragrances [5]. The diverse chemical composition of plant EOs allows for their classification into several chemotypes, including those containing terpenes, aromatic compounds, and aliphatic compounds. For decades, numerous EO profiles and chemotypes have been identified in Cinnamomum species, with terpenoids being the primary components [6,7]. However, the principal terpenoid components of the EOs of Cinnamomum species display considerable intraspecific and interspecific variations in phytochemicals. Cinnamomum plants are classified into various chemotypes, such as linalool-type, eucalyptol-type, camphor-type, borneol-type, and nerolidol-type chemotypes [8,9,10]. Minor genetic and epigenetic alterations, which exert minimal or no influence on the plant’s morphology or anatomy, can nevertheless give rise to substantial alterations in the chemotype [11]. In recent years, efforts have been made to explore the differences in the metabolic pathways of different chemotypes, with the objective of breeding high-quality varieties with specific chemical compositions.
Previous research on the biosynthesis and metabolic regulation of plant terpenoids have determined that the formation of terpenoids occurs through the condensation of isopentenyl diphosphate (IPP) and its allylic isomer dimethylallyl diphosphate (DMAPP). IPP and DMAPP are synthesized in plastids by the 2-methylerythritol 4-phosphate pathway (MEP) and in the cytoplasm by the mevalonate (MVA) pathway [12,13,14]. Terpene synthases (TPSs) catalyze the biosynthesis of sesquiterpene from MVA pathway precursors, and diterpenes and monoterpenes from MEP pathway precursors [15,16]. The development of RNA-sequencing (RNA-Seq) has enabled comprehensive transcriptomic investigations in species lacking a reference genome [17]. The use of transcriptome sequencing has enabled the identification of genes involved in terpene biosynthesis in different chemotypes of Cinnamomum [9,18,19,20,21]. However, the lack of systematic and in-depth transcriptomic studies on terpenoids in various Cinnamomum chemotypes has hindered the effective transformation and utilization of these plants. The recent publication of several Cinnamomum whole-genome sequences provides reference data for an in-depth exploration of the molecular mechanisms underlying the formation of different Cinnamomum chemotypes at the genomic level [22,23,24,25].
In this study, we extracted the EOs from the leaves of C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum and analyzed the main terpenoid constituents in the EOs using GC-MS to determine their chemotypes. We performed a comparative analysis of reference transcriptomes from the leaves of these four Cinnamomum species, using the published whole genome of C. camphora var. linaloolifera as the reference sequence. Following functional annotation and classification, the genes involved in the MEP/MVA pathways and terpene synthesis in different Cinnamomum species were identified and their expression levels were verified.
2. Materials and Methods
2.1. Plant Material
The study utilized four species of Cinnamomum. Two species, C. kanehirae and C. micranthum, were grown in the Fujian Forestry Science and Technology Experiment Center, Nanjing, Zhangzhou City, Fujian Province, China (127°19′ E, 24°30′ N). The two other species were represented by three-year-old C. longipaniculatum and two-year-old Nan’an 1, a new variety of C. camphora var. linaloolifera, which were cultivated in the Banlin state-owned forest farm in Anxi, Fujian Province (117°57′ E, 24°55′ N). No specific permits were required for this study. The plant material was identified by Prof. Shuangquan Zou (see author list). Healthy and pest-free fresh leaves were randomly collected from four directions (east, south, west, and north) from the plants for EO extraction and identification of the major components. In addition, a portion of fresh leaves was subjected to a thorough washing with sterile water and immediately frozen in liquid nitrogen for the purposes of transcriptome sequencing and qRT-PCR. Three relevant biological replicates were generated for each sample.
2.2. Essential Oil Extraction
EOs were extracted using a modified water vapor method in our laboratory. A 100 g sample of fresh leaves was collected and placed into an oven at 105 °C for 0.5 h. After the initial drying, the sample was transferred to an incubator set at 65 °C to complete the drying process, after which, the samples were weighed (W2). The dried sample was pulverized and placed into a 500 mL volatile oil extractor with 200 mL of ultrapure water and heated for 4 h. After distillation, the EO layer, which separates and floats on the surface, was collected, dried with anhydrous sodium sulfate, and weighed (W1). The distillation process was considered complete when two consecutive measurements showed no increase in the EO collected. Finally, the EO yield was calculated using the formula W1/W2 × 100.
2.3. Gas Chromatography–Mass Spectrometry (GC-MS) Analysis
The composition of the EOs was determined by GC-MS, performed using a Shimadzu QP2020 GC-MS instrument (chromatographic column: SH-RXI-5SILMS, 30 m × 0.25 mm × 0.25 µm). The GC-MS procedure entailed maintaining the EO samples initially at 80 °C for 2 min, subsequently elevating the temperature to 160 °C at a rate of 8 °C/min, and then further increasing the temperature to 250 °C at a rate of 8 °C/min, where it was maintained for 2 min. The injection volume was 1.0 µL, and a split ratio of 20:1 was employed. The temperature of the injection port was 280 °C, the EI ion source was 230 °C, and the connection line was 200 °C. The MS scan range (m/z) was 50–650. For the analysis, the essential oils were dissolved in alcohol (30 mg/mL) and directly injected.
2.4. Transcriptome Sequencing and Assembly
High-quality RNA was extracted from healthy mature leaves of Nan’an 1 (C. camphora var. linaloolifera), C. kanehirae, C. longipaniculatum, and C. micranthum using the RNeasy Plant Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. The mass and concentration of the extracted RNA were quantified using 1% agarose gel electrophoresis and UV spectrophotometry. The quality-checked RNAs were enriched with oligo-dTs, randomly interrupted, and double-stranded cDNAs were synthesized. After the DNA was amplified by PCR with specific primers, the single-stranded DNA was circularized with a bridge primer to obtain a single-stranded circular DNA library. Afterwards, it was sequenced on the Novaseq 6000 platform (Illumina, San Diego, CA, USA). SOAPnuke v1.4.0 [26] and Trimmomatic v0.36 [27] were used to count and filter the raw data. The clean data were assembled using Bowtie2 v2.2.5 [28], with the Nan’an 1 genome as the reference genome.
2.5. Sequencing Data Analyses
RSEM v1.2.8 [29] was used to obtain the FPKM (expected number of fragments per kilobase of transcript sequence per million base pairs sequenced) values. DESeq2 [30] was used to conduct the analysis on the differentially expressed genes (DEGs). Genes with |log2(FC)| ≥ 2 and a Q value ≤ 0.05 were considered to be significantly differentially expressed. The ‘ggVolcano’ package in R was used to perform the volcano mapping, and the ‘ggVennDiagram’ and the ‘UpsetR’ packages were used to perform the Upset mapping. Based on Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG), significantly different DEGs were functionally annotated using Goseq R package. The FPKM values of candidate genes for EO synthesis in different tissues were visualized by using TBtools-II v2.101 [31].
2.6. Quantitative Real-Time PCR Evaluation
A total of 24 key genes for EO synthesis were selected for qRT-PCR analysis to verify the accuracy of the transcriptome data and the actual expression levels of the selected genes. Leaf RNA was extracted using a total plant RNA kit (Polysaccharide Polyphenol Plant Total RNA Extraction Kit, developed by Hangzhou Bo Ri Technology Co., Ltd, (Hangzhou, China) for qRT-PCR validation. Primer Premier 5.0 software was used to design the primers, and all the primer information are shown in Table A1. CcEF1a was used as an internal reference gene to calculate the relative expression of each target gene within a sample. The qualified RNA was employed as the template to construct an RT reaction solution in accordance with the TransScript® One-Step gDNA Removal and cDNA Synthesis SuperMix instructions. The final whole reaction system volume was 20 µL, which was inactivated at 85 °C for 5 s to produce cDNA. The reaction conditions were as follows: 94 °C for 30 s, 40 cycles of 94 °C for 5 s and 60 °C for 30 s, 72 °C for 10 s, 95 °C for 15 s, 60 °C for 1 min, and 95 °C for 1 s. Each reaction was completed for three technical replicates, and each biological replicate included three technical replications. The relative expression levels of all genes were determined using the 2−ΔΔCt method [32].
3. Results
3.1. Terpenoids as the First Principal Component of Four Cinnamomum Species’ Leaf EOs
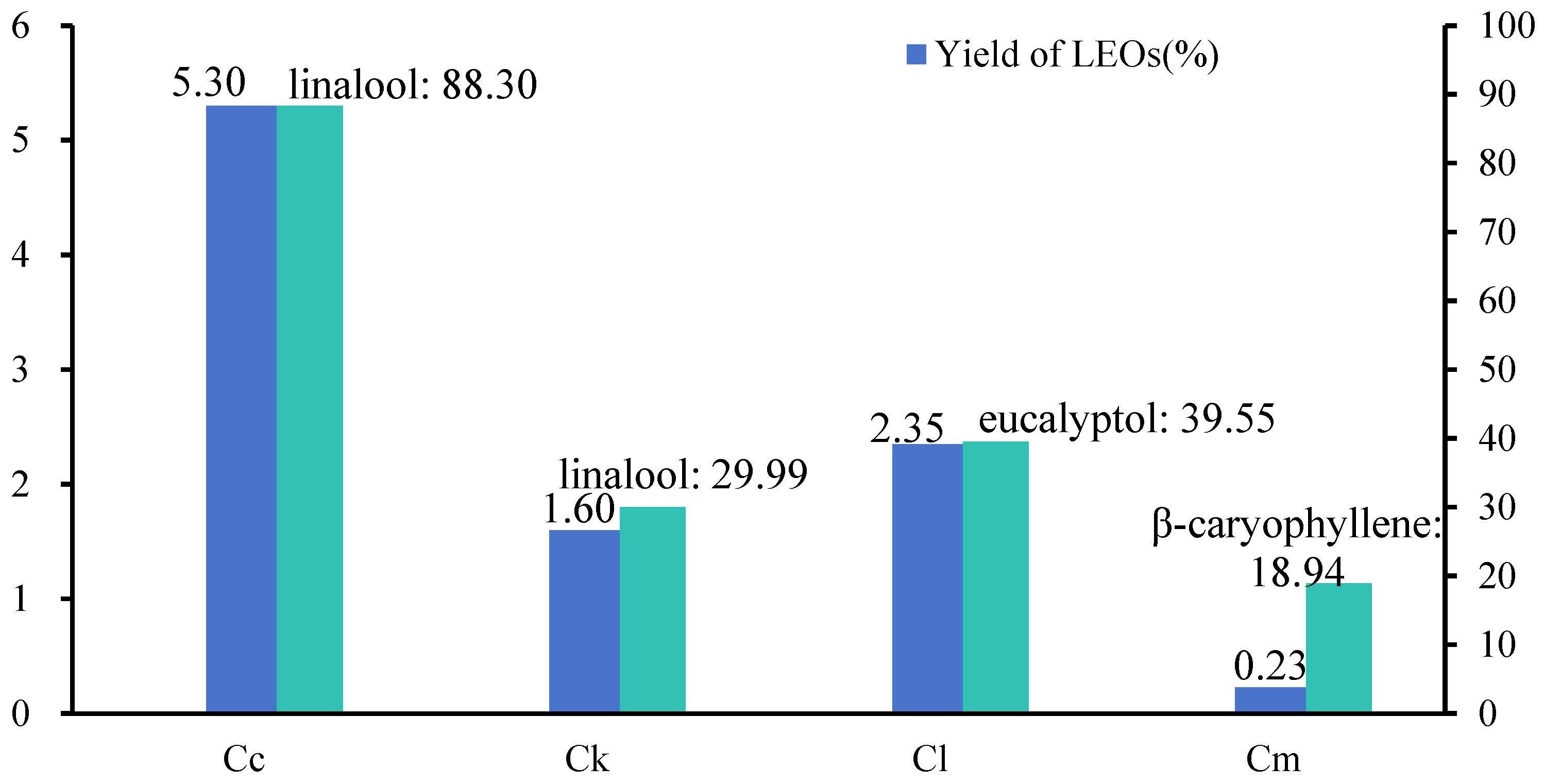
We extracted the EOs from the leaves of C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum using water distillation. The EO yield of the C. camphora var. linaloolifera leaves was the highest at 5.3%, followed by C. longipaniculatum (2.35%), C. kanehirae (1.6%), and C. micranthum (0.23%) (Figure 1). Subsequently, we determined the major composition of the EOs of these four species using GC-MS (Figure 1 and Table A2, Table A3, Table A4 and Table A5). The results indicated that the primary constituents of the EOs of C. camphora var. linaloolifera, C. kanehirae, and C. longipaniculatum were terpenoids, whereas those of C. micranthum were aliphatic aldehydes. Specifically, 30 compounds were identified in the EOs of the C. camphora var. linaloolifera leaves, with linalool (88.30%), α-terpineol (1.25%), and geraniol (0.81%) being the main components (Table A2). A total of 58 compounds were detected in the EOs of the C. kanehirae leaves, with linalool (29.99%), o-cymol (10.94%), a-phellandrene (7.14%), and eucalyptol (6.89%) being the main components (Table A3). A total of 56 compounds were detected in the EOs of the C. longipaniculatum leaves, predominantly eucalyptol (39.55%), β-thujene (15.80%), and α-terpineol (13.47) (Table A4). Lastly, a total of 89 compounds were detected in the EOs of the C. micranthum leaves, with β-caryophyllene (18.94%) as the primary component, followed by aliphatic aldehydes such as decanal (4.12%), nonanal (1.14%), and dodecanal (1.07%) (Table A5).
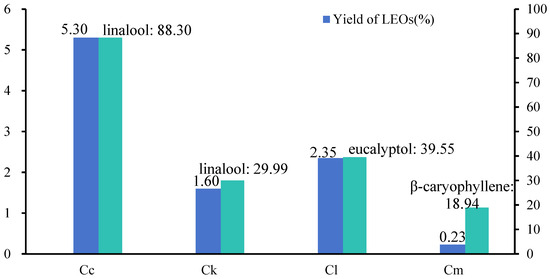
Figure 1.
Leaf EO content and principal components from four Cinnamomum species. Abbreviations: leaf essential oil (LEO), C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
3.2. Sequencing and Transcriptome Assembly
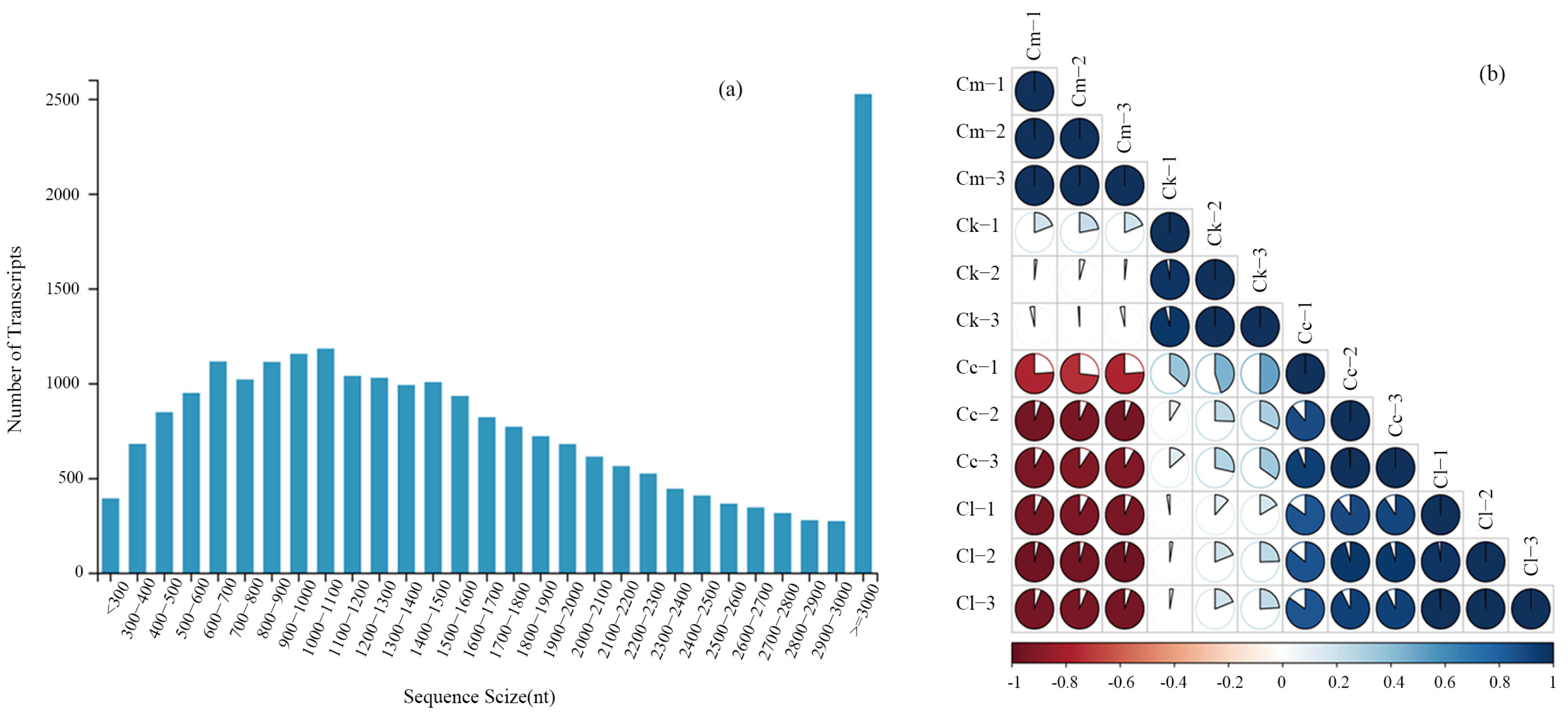
To identify the key genes involved in the synthesis of the major components of the EOs extracted from the fresh leaves of C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum and their expression profiles, transcriptome sequencing was performed on 12 samples. The Q30 was greater than 90.90%, and the genome mapping rate was greater than 83.30% for each sample (Table A6). We obtained transcriptome information from the 12 leaf samples, using the C. camphora var. linaloolifera genome sequence as the reference. A total of 6869 transcripts were identified with lengths ranging from 300 to 1000 nt, accounting for 43.45% of the total transcripts. Additionally, we found 9163 transcripts with lengths ranging from 1000 to 2000 nt, accounting for 57.96% of the total transcripts, and 2525 transcripts longer than 3000 nt, accounting for 15.97% of the total transcripts (Figure 2a). Furthermore, the Pearson correlation coefficient (PCC) analysis of the sample groups showed that the correlation within the same group was higher than that between different groups (Figure 2b), indicating good biological reproducibility. These results confirm that the transcriptome sequencing and assembly are of good quality and can be used for subsequent analyses.
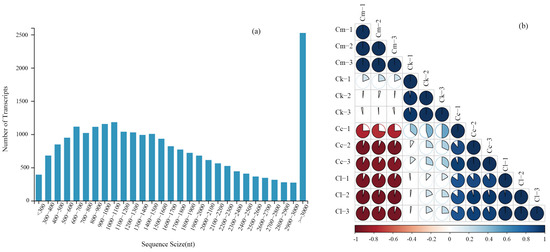
Figure 2.
Transcriptome sequencing quality assessment. (a) Length distribution of transcripts; (b) Pearson correlation coefficient (PCC) between samples. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
3.3. Analysis of Differentially Expressed Genes (DEGs)
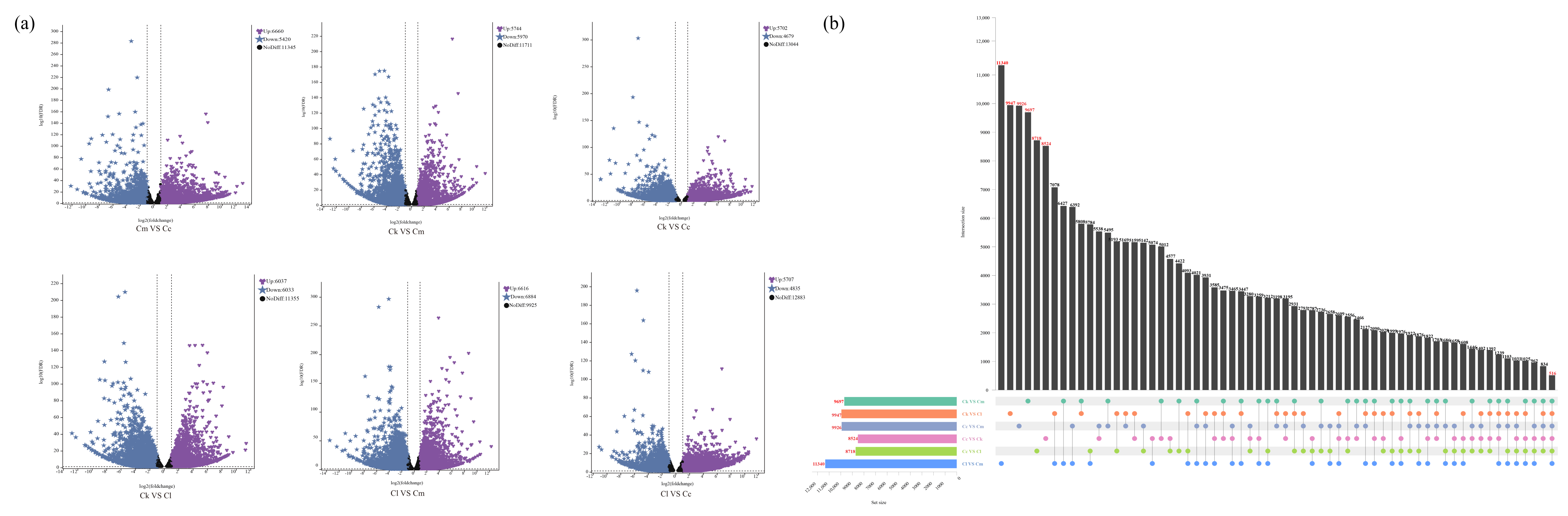
We screened the DEGs in the six groups of C. camphora var. linaloolifera vs. C. kanehirae (Cc vs. Ck), C. camphora var. linaloolifera vs. C. longipaniculatum (Cc vs. Cl), C. camphora var. linaloolifera vs. C. micranthum (Cc vs. Cm), C. kanehirae vs. C. longipaniculatum (Ck vs. Cl), C. kanehirae vs. C. micranthum (Ck vs. Cm), and C. longipaniculatum vs. C. micranthum (Cl vs. Cm) (Figure 3a). The results showed that there were 10,381 DEGs (up-regulated DEGs: 5702; down-regulated DEGs: 4679) for Cc vs. Ck, 12,080 (up-regulated DEGs: 6660; down-regulated DEGs: 5420) for Cc vs. Cm, 10,542 (up-regulated DEGs: 5707; down-regulated DEGs: 4835) for Cc vs. Cl, 11,714 (up-regulated DEGs: 5744; down-regulated DEGs: 5970) for Ck vs. Cm, 10,207 (up-regulated DEGs: 6037; down-regulated DEGs: 6033) for Ck vs. Cl, and 13,500 (up-regulated DEGs: 6616; down-regulated DEGs: 6884) for Cl vs. Cm. In addition, we identified 516 DEGs that were shared among the four Cinnamomum species, of which, 8524 DEGs were shared by Cc vs. Ck, 9926 DEGs shared by Cc vs. Cm, 8718 DEGs shared by Cc vs. Cl, 9697 DEGs shared by Ck vs. Cm, 9947 DEGs shared by Ck vs. Cl, and 11,340 DEGs shared by Cl vs. Cm (Figure 3b).
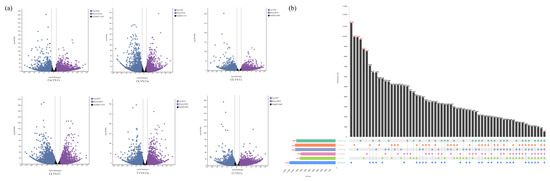
Figure 3.
Identification of DEGs. (a) Volcano plot of DEGs between groups; (b) Upset plots of DEGs between groups. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
3.4. Functional Annotation of DEGs
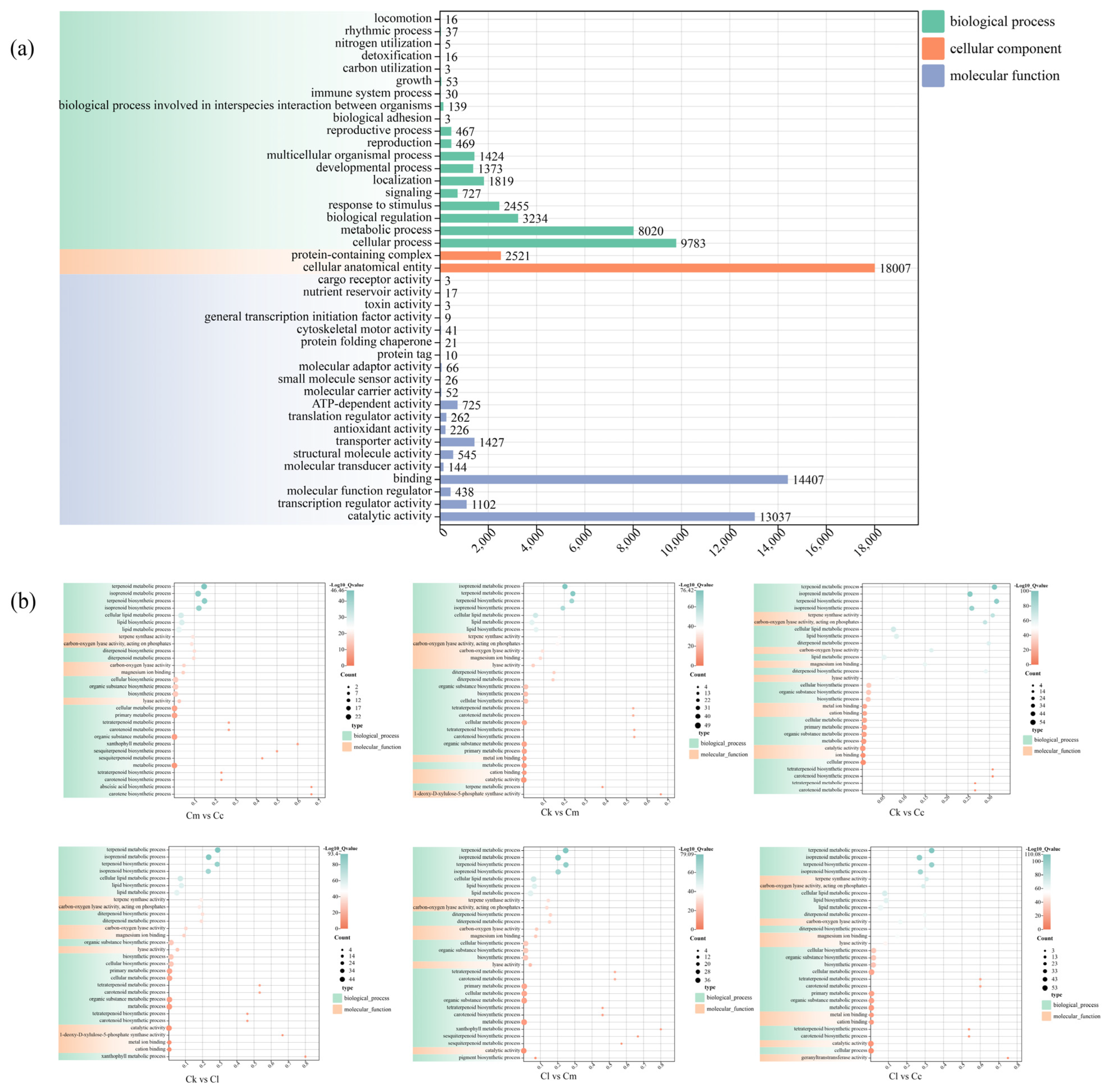
GO assignments were used to classify the functions of all the obtained DEGs, categorizing 83,152 DEGs under the categories of molecular function (32,561 DEGs, 39.15%), biological process (29,773 DEGs, 35.81%), and cellular component (20,528 DEGs, 24.68%) (Figure 4a). In the category of molecular functions, the DEGs were grouped into 20 classes, with the highest subcategory being ‘binding’ (14,407 DEGs), followed by ‘catalytic activity’ (13,037 DEGs). In the category of biological process, the DEGs were classified into 19 groups, with the highest subcategory being ‘cellular process’ (9783 DEGs), followed by ‘metabolic process’ (8020 DEGs). With regard to the cellular component category, the DEGs were divided into ‘cellular anatomical entity’ (18,007 DEGs) and ‘protein-containing complex’ (2521 DEGs) groups.
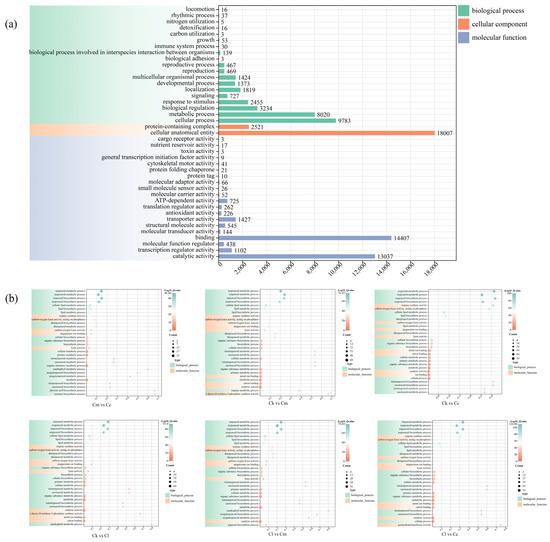
Figure 4.
GO enrichment distribution of DEGs. (a) GO enrichment of DEGs from the four Cinnamomum species; (b) GO enrichment of DEGs between groups. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
Subsequently, we performed GO functional enrichment analysis of the intergroup DEGs in Cc vs. Ck, Cc vs. Cl, Cc vs. Cm, Ck vs. Cl, Ck vs. Cm, and Cl vs. Cm (Figure 4b). The results showed that these DEGs were significantly enriched with terpene synthesis-related entries, such as ‘terpenoids metabolic/biosynthetic process’, ‘isoprenoids metabolic/biosynthetic process’, and ‘diterpenoid metabolic/biosynthetic process’.
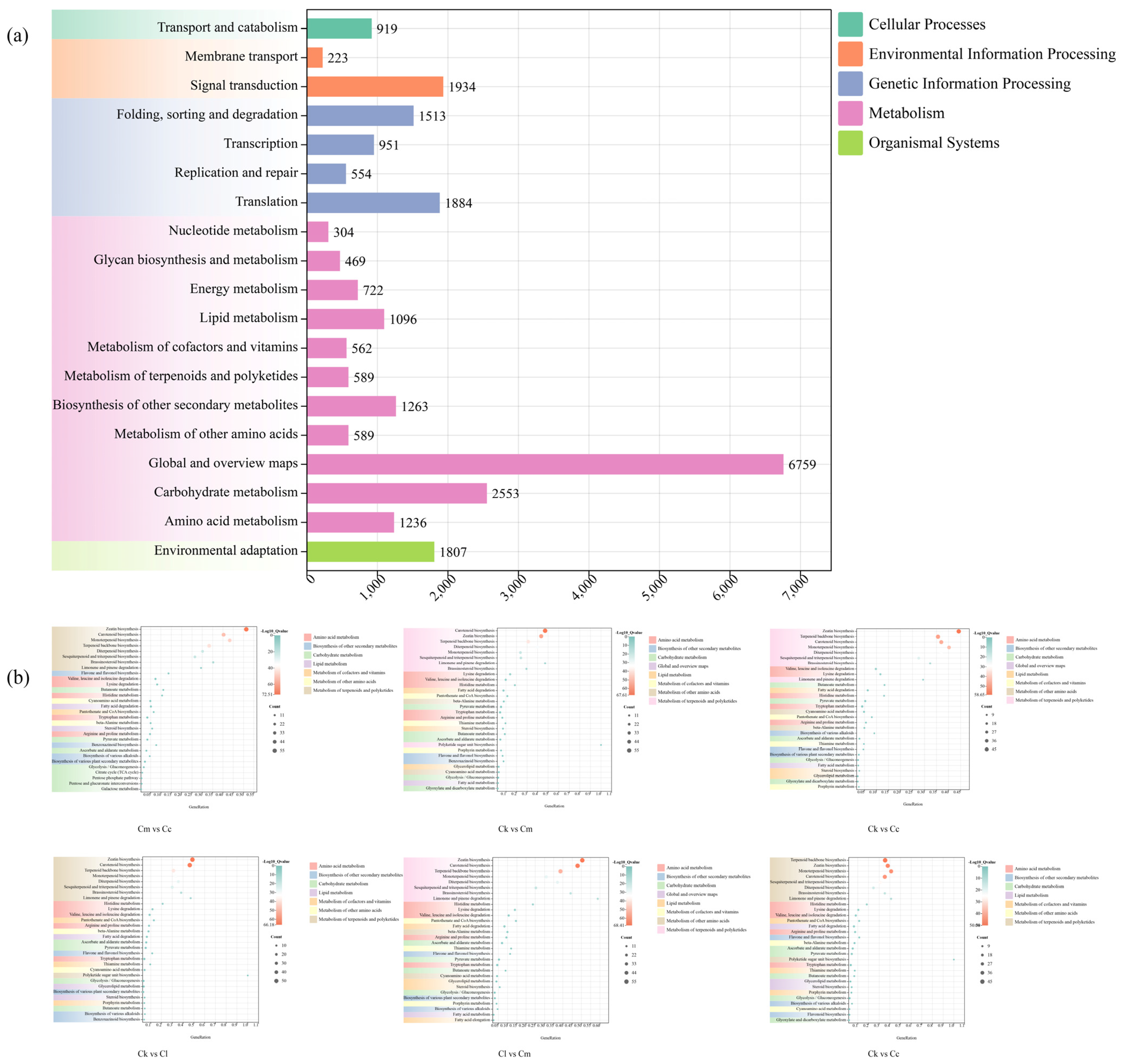
To understand the biological functions of the genes, we performed pathway analysis using the KEGG database. A total of 25,927 DEGs were annotated and categorized into metabolism (16,142 DEGs, 62.26%), genetic information processing (4902 DEGs, 18.91%), environmental information processing (2157 DEGs, 8.32%), organismal systems (1807 DEGs, 6.97%), and cellular processes (919 DEGs, 3.54%) (Figure 5a). Among these groups, metabolism was the most prominent, and it was further divided into 11 subgroups, with 589 DEGs annotated with ‘metabolism of terpenoids and polyketides’ (Figure 5a).
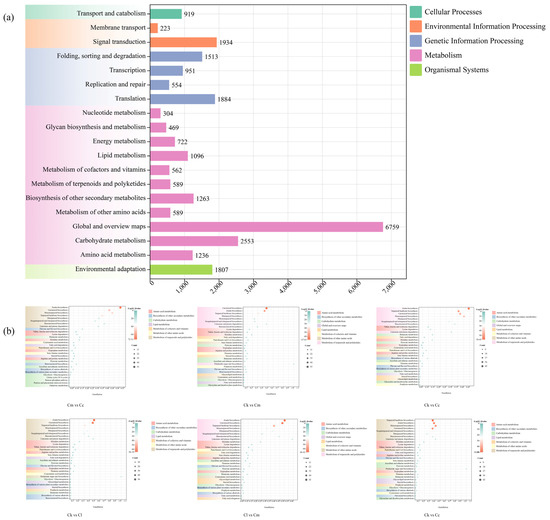
Figure 5.
KEGG enrichment distribution of DEGs. (a) KEGG enrichment of DEGs in the four Cinnamomum species; (b) KEGG enrichment of DEGs between groups. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
Similarly, we performed KEGG enrichment analysis of the intergroup DEGs in Cc vs. Ck, Cc vs. Cl, Cc vs. Cm, Ck vs. Cl, Ck vs. Cm, and Cl vs. Cm (Figure 5b). The results showed that the intergroup DEGs in these six groups were significantly enriched in pathways related to ‘terpenoids backbone biosynthesis’, ‘diterpenoids biosynthesis’, ‘sesquiterpenoid and triterpenoid biosynthesis’, and ‘monoterpenoid biosynthesis’.
3.5. Candidate Genes Involved in Terpenoid Biosynthesis
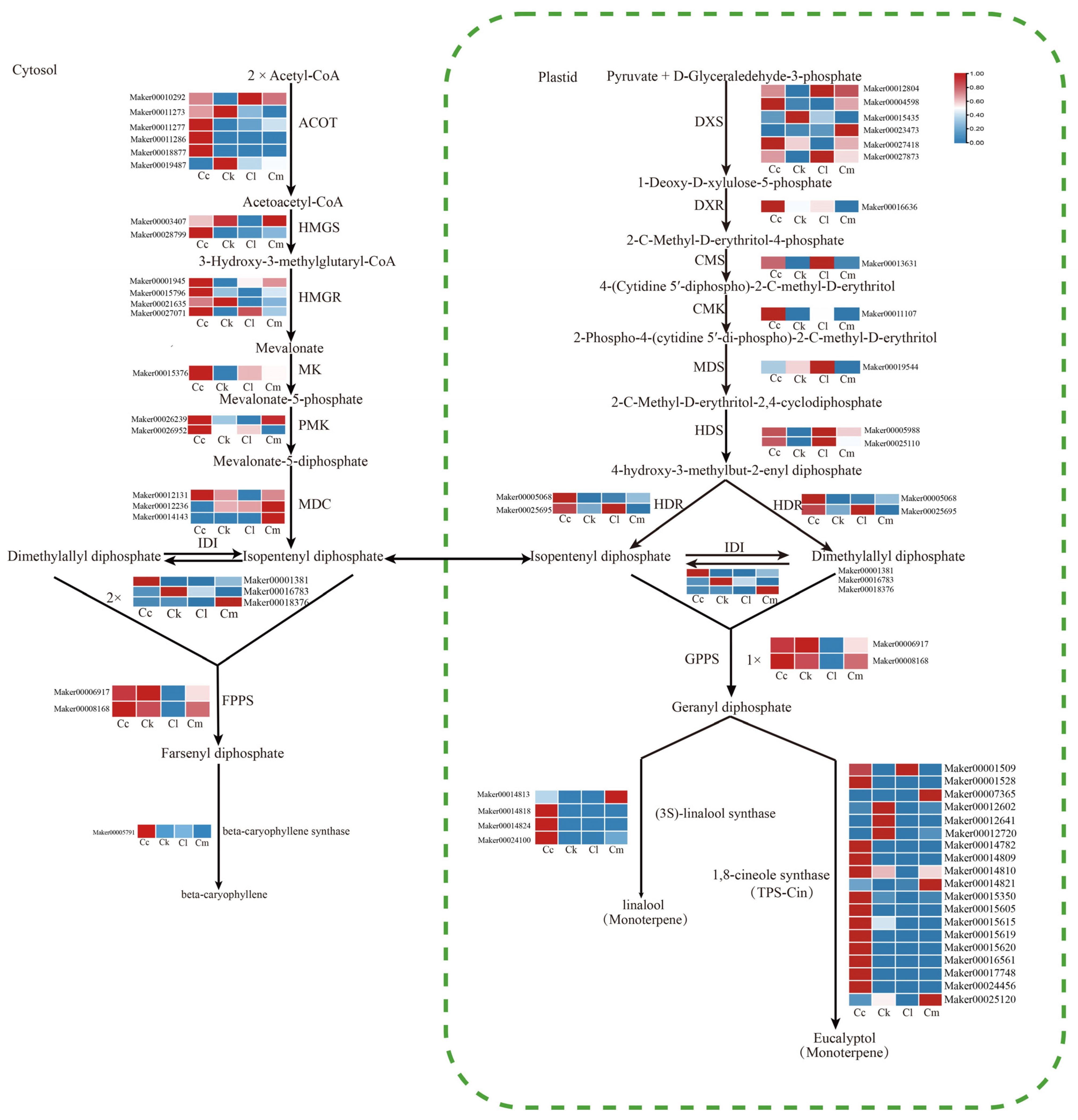
According to results of the GC-MS and DEG functional enrichment analyses, the main component of the four Cinnamomum leaf EOs were terpenoids. Consequently, we mapped the enriched DEGs to the terpenoid synthesis pathway and analyzed their expression. Terpene biosynthesis occurs through two distinct pathways: the cytosolic MVA pathway and the plastidial MEP pathway. In the MVA pathway, 24 DEGs were identified. These included eighteen DEGs involved in converting acetyl-CoA to IPP and DMAPP: six acetyl-CoA acetyltransferase (ACAT) genes, two hydroxymethylglutaryl-CoA synthase (HMGS) genes, four hydroxymethylglutaryl-CoA reductase (HMGR) genes, one mevalonate kinase (MK) gene, two phosphomevalonate kinase (PMK) genes, and three diphosphomevalonate decarboxylase (MDC) genes. Additionally, three isopentenyl diphosphate isomerase (IDI) genes (Maker00001381, Maker00016783, and Maker00018376) were identified that are involved in the isomeric conversion of IPP and DMAPP, and two farnesyl diphosphate synthase (FPPS) genes (Maker00006917 and Maker00008168) were identified that are involved in the synthesis of the direct precursor farnesyl diphosphate (Table A7). Fourteen DEGs were annotated as being involved in the MEP pathway and are responsible for converting pyruvate to IPP and DMAPP, including six 1-deoxy-D-xylulose-5-phosphate synthase (DXS) genes, one deoxyxylulose phosphate reductoisomerase (DXR) gene, one 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase (CMS) gene, one 4-diphosphocytidyl-2-C-methyl-D-erythritol kinase (CMK) gene, one 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (MDS) gene, two 4-hydroxy-3-methylbut-2-enyl diphosphate synthase (HDS) genes, and two 4-hydroxy-3-methylbut-2-enyl diphosphate reductase (HDR) genes. Furthermore, three IDI genes and two geranyl diphosphate synthase (GPPS) genes (Maker00006917 and Maker00008168) were identified that are involved in the synthesis of the direct precursor geranyl/geranylgeranyl diphosphate (Table A7).
Additionally, four candidate TPS genes (Maker00024100, Maker00014813, Maker00014818, and Maker00018424) were identified that are involved in the synthesis of cyclic monoterpenes. Nineteen candidate TPS genes were discovered that are involved in the synthesis of monoterpenes. One TPS gene (Maker00005791) was identified that is involved in the production of sesquiterpenes (Figure 6).
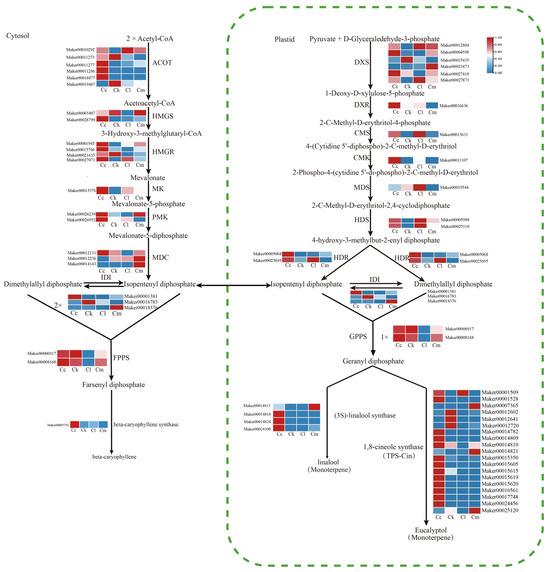
Figure 6.
Expression profiles of terpenoid biosynthesis-related genes from leaves of four Cinnamomum species. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
3.6. Expression Patterns of Terpenoid Synthesis Candidate Genes
The MEP pathway provides the precursors for the synthesis of monoterpenes and diterpenes, and the MVA pathway provides the precursors for the synthesis of sesquiterpenes [15]. The main constituents of the EOs of C. camphora var. linaloolifera, C. kanehirae, and C. longipaniculatum leaves were monoterpenes, while the main constituents of the EO of C. micranthum leaves was sesquiterpenes. An expression analysis showed pathway-specific patterns of genes involved in terpenoid synthesis across the four Cinnamomum species. C. camphora var. linaloolifera showed high expression of most structural genes in the MEP pathway except for the MDS gene. Similarly, in C. longipaniculatum, high expression levels were observed for most structural genes in the MEP pathway, except for DXR, CMK, and GPPS. In C. kanehirae, except for one DXS gene (Maker00015435), two GPPS genes (Maker00006917 and Maker00008168) were highly expressed, and the other structural genes of the MEP pathway were expressed at low levels (Figure 6). In the leaves of C. micranthum, one HMGS gene (Maker00003407), one PMK gene (Maker00026239), two MDC genes (Maker00012236 and Maker00014143), and one IDI gene (Maker00018376) of the MVA pathway were highly expressed, while the other structural genes were expressed at low levels (Figure 6). Notably, all structural genes of the MVA pathway were highly expressed in the leaves of C. camphora var. linaloolifera.
In addition, four linalool-synthesizing TPS genes were expressed at low levels in C. kanehirae, and among these, three of them (Maker00014818, Maker00014824, and Maker00024100) were highly expressed in C. camphora var. linaloolifera. Only 1 (Maker00001509) of the 19 eucalyptol-synthesizing candidate TPS genes was highly expressed in C. longipaniculatum, whereas one β-caryophyllene-synthesizing TPS gene (Maker00005791) was expressed at low levels in C. micranthum and at high levels in C. camphora var. linaloolifera (Figure 6).
3.7. Expression Validation by qRT-PCR
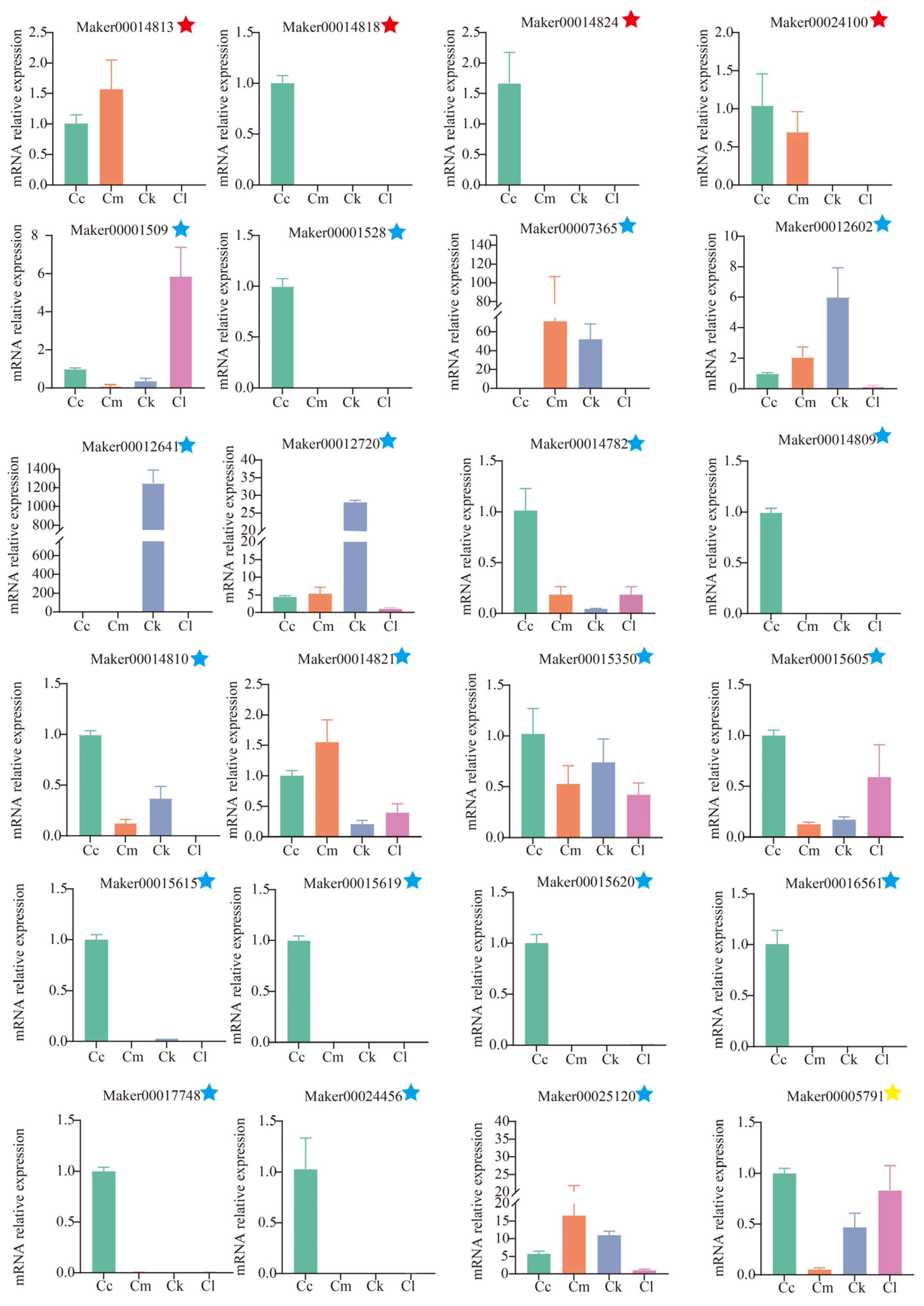
To verify the accuracy of the RNA-Seq expression data, a total of 24 candidate TPS genes were subjected to qRT-PCR analysis (Figure 7). Regarding linalool synthesis-related genes, four TPS genes were highly expressed in C. camphora var. linaloolifera, but showed very low expression in C. longipaniculatum and C. micranthum. Among these, Maker00014813 and Maker00014818 were also highly expressed in C. kanehirae. Regarding eucalyptol synthesis-related genes, 8 of the 19 candidate genes were expressed only in C. camphora var. linaloolifera and 6 were expressed in C. longipaniculatum, with Maker00001509 showing the highest expression in C. longipaniculatum. The β-caryophyllene-synthesizing gene, Maker00005791, was expressed in the four Cinnamomum species and showed high levels of expression in C. camphora var. linaloolifera and C. longipaniculatum, but low expression in C. micranthum. Overall, the qRT-PCR validation results aligned well with the transcriptome expression profiles. From this analysis, we identified four key genes involved in linalool synthesis (Maker00024100, Maker00014813, Maker00014818, and Maker00018424), one key gene involved in eucalyptol synthesis (Maker00001509), and one key gene involved in β-caryophyllene synthesis (Maker00005791).
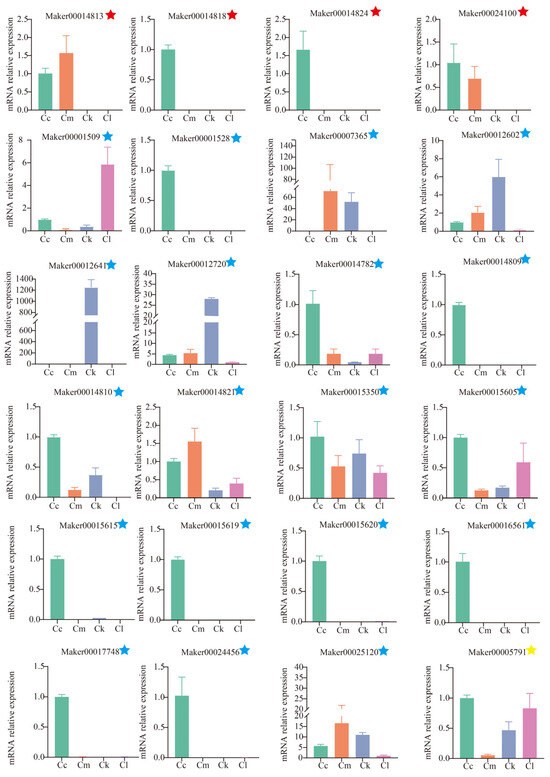
Figure 7.
Expression levels of TPS genes selected for validation. The red five-pointed star represents the candidate TPS gene involved in linalool synthesis, the blue five-pointed star represents the candidate TPS gene involved in eucalyptol synthesis, and the yellow-pointed star represents the candidate TPS gene involved in β-caryophyllene synthesis. Abbreviations: C. camphora var. linaloolifera (Cc), C. kanehirae (Ck), C. longipaniculatum (Cl), and C. micranthum (Cm).
4. Discussion
Cinnamomum species are rich in EOs, which are employed globally as fragrances, spices, and traditional herbal remedies. Our comparative analysis of the fresh leaf EO yields from C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum revealed that C. camphora var. linaloolifera exhibited the highest EO yield (5.3%), notably higher than those of other Cinnamomum species that were previously reported in the literature [33]. Chemotypic diversity is prevalent among Cinnamomum species. Our chemical compositional analysis showed that the acyclic monoterpene linalool was detected in the EOs of all four Cinnamomum species. The principal component of the EOs of C. camphora var. linaloolifera and C. kanehirae was linalool, comprising 88.3% and 29.99% of the EO, respectively. In this study, the higher EO yield and linalool content from C. camphora var. linaloolifera can be attributed to the use of the high-yield cultivar Nan’an 1 that was selected by our research group [34]. Eucalyptol, which is commonly added to flavorings, spices, and cosmetics, was identified in our study as the main component of the C. longipaniculatum EO, with a content of 39.55%. This contrasts with previous findings by Zhao et al. [21] who only detected seven terpenoids in C. longipaniculatum EOs and did not find eucalyptol. The discrepancy is likely due to the different chemotypes of C. longipaniculatum used in the two studies. Although a significant number of aliphatic aldehydes were detected in the EO of C. micranthum leaves, its primary constituent was the bicyclic sesquiterpene β-caryophyllene (18.94%). This compound is known for its strong medical value, including anti-inflammatory and analgesic properties [35]. Collectively, while the main constituents of the EOs from the four Cinnamomum species were terpenoids, the specific terpenoid content varied significantly among species.
Previous investigations into terpenoid biosynthesis in different chemotypes of Cinnamomum species relied on transcriptome data [9,19,20]. Numerous terpenoid synthases have been isolated and characterized from various plant species [36,37]. Transcriptome sequencing allows for the acquisition of a large number of transcripts, thereby providing a time-efficient and cost-effective approach for the rapid identification of functional genes. However, due to the temporal and spatial limitations of gene transcription, transcriptome sequencing is only capable of providing transient gene transcription information during RNA extraction, which may result in the omission of time-specific active functional genes. Whole-genome sequencing addresses this limitation by providing a comprehensive genetic resource. Recently, several C. camphora reference genomes have been released, including a new variety of C. camphora var. linaloolifera (Nan’an 1), which has a high linalool content [22,23,24,25]. In this study, we integrated genomic and transcriptomic data to elucidate the molecular regulatory mechanisms underlying the synthesis of the primary terpenoid components in C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum.
Six sets of DEGs were analyzed, revealing that Cl vs. Cm exhibited the highest number of DEGs and shared DEGs, whereas Cc vs. Ck showed the lowest number. This variation may be attributed to the distinct main terpenoid components in these species: C. longipaniculatum primarily produces the monoterpene eucalyptol, while C. micranthum produces the sesquiterpene β-caryophyllene. Conversely, both the C. camphora var. linaloolifera and C. kanehirae leaf EOs predominantly contained the cyclic monoterpene linalool. The functional enrichment analysis further revealed that the DEGs were significantly enriched in terpene-related pathways, consistent with the results of Zhao et al. [21]. Terpenoids are synthesized via the MVP pathway in plastids and the MVA pathway in the cytosol [38]. A total of 42 DEGs were found to be associated with the MVA and MEP pathways, indicating that the EOs in Cinnamomum species are synthesized via both pathways. The expression levels of the candidate genes in the MEP pathway were higher than those in the MVA pathway in both C. camphora var. linaloolifera and C. kanehirae, indicating a higher abundance of monoterpenes.
TPSs are the rate-limiting enzymes responsible for the production of various terpenoids [14,39]. Phylogenetic analyses have divided the TPS gene family into several subfamilies, of which, the TPS-b/g subfamily typically encodes monoterpene synthases, with TPS-g specifically encoding acyclic monoterpene synthases, while the TPS-a subfamily is involved in the biosynthesis of sesquiterpenes [40,41]. The expansion of the TPS family (especially the TPS-b subfamily) in the genomes of C. camphora and C. kanehirae partially explains the diversity of terpenoids in these Cinnamomum species [7,22,23,24]. In total, 24 TPS candidate genes were identified, with 19 belonging to the TPS-b subfamily. The expression validation indicated that only one TPS-b gene is involved in the biosynthesis of the monoterpene eucalyptol.
5. Conclusions
In this study, we systematically and comparatively analyzed the principal components of the leaf EOs of four Cinnamomum species: C. camphora var. linaloolifera, C. kanehirae, C. longipaniculatum, and C. micranthum. Terpenoids were identified as the primary constituents of the EOs of all four species. Specifically, linalool, an acyclic monoterpene, was the predominant constituent in the EOs of C. camphora var. linaloolifera and C. kanehirae, the monoterpene eucalyptol was the main constituent in the C. longipaniculatum EO, while the sesquiterpene β-caryophyllene was the main constituent in the C. micranthum EO. Notably, the leaf EO yield and linalool content of C. camphora var. linaloolifera were found to be higher than those of other Cinnamomum plants reported in the literature, suggesting that the leaves of C. camphora var. linaloolifera are an ideal sample for studying linalool biosynthesis and are a high-quality source of naturally occurring linalool for extraction. A total of 12 samples of mature leaves from four Cinnamomum species were subjected to transcriptome sequencing and assembly, using the C. camphora var. linaloolifera genome as the reference. We identified 66 candidate genes that are involved in the synthesis of the major EO components (linalool, eucalyptol, and β-caryophyllene) across the four species, including 42 structural genes belonging to the MEP and MVA pathways and 24 TPS genes. Furthermore, the qRT-PCR data revealed that the expression patterns of these genes varied among the different Cinnamomum species, correlating with the different principal components and their contents in the EOs. In conclusion, these results provide a theoretical basis for further exploration of the biosynthesis of the major components of EOs from different chemotypes of Cinnamomum.
Author Contributions
Conceptualization, W.S. and S.Z.; methodology, H.N.; software, W.S. and H.N.; validation, Z.Z. and H.N.; formal analysis, W.S. and H.N.; investigation, Z.Z.; resources, H.N.; data curation, W.S.; writing—original draft preparation, W.S.; writing—review and editing, W.S. and S.Z.; visualization, H.N.; supervision, S.Z.; project administration, S.Z.; funding acquisition, S.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Fujian Forestry Science and Technology Plan project “Salt-Tolerance Mechanism of Cinnamomum Camphor and its Application in Coastal Mountain Afforestation” (2024FKJ25) and “Application Demonstration of Cinnamomum Camphor Seed in Coastal Ecological Landscape Forest Transformation” (2024TG08).
Data Availability Statement
All the raw transcriptome sequences described in this manuscript were deposited in BioProject/GSA, National Genomics Data Center (NGDC), under accession No. PRJCA002001/CRA004220.
Conflicts of Interest
The authors declare no conflicts of interest.
Appendix A

Table A1.
Primers for real-time fluorescence quantification.
Table A1.
Primers for real-time fluorescence quantification.
| Gene ID | Primer Sequence (5′) | Primer Sequence (3′) |
|---|---|---|
| Maker00014818 | GTTCGGGAGTGCATATCGT | CCACAACCGCAGAATCGT |
| Maker00014824 | TTCCAATCAATCCGAGCAA | ATGTCAGGGTCTTGGGTGC |
| Maker00024100 | CCCATCAGGGAGATTCGTTA | TGCCACCATTATTTCCAGAGT |
| Maker00014813 | CAAGACCCTGACATGGATAC | GCCATTGCCACCATTAT |
| Maker00001528 | TGTCAAGGTGGTGGAAAGATTTGGG | TTGTGGCTCGGATGTCATTGCG |
| Maker00007365 | CAATCCGTTTCAAGTTTCC | CCTTGTAGGCACTCGTCA |
| Maker00012602 | GATTCTGATGGAAAGGGTG | TATGCCGTCTCATTGGTT |
| Maker00012641 | AACCAATGAGACGGCATA | CCCCAACTGAGACTAAAGC |
| Maker00012720 | CAAAGTAACTCGCACTATCG | CCATCAAAGTCATGATGGTACATCA |
| Maker00014782 | TGAGCCCTTCAATCCGCAAAGAC | TTCGAGTAGGTTGGAGCACTTGTTG |
| Maker00014809 | GCAAACGACATTCCAAAC | CCTCAAGATCCCATCTATCA |
| Maker00014810 | CGCTTGATTGTGGGTCT | CGGTCTTTGTTGGCTTT |
| Maker00014821 | TTCAGGTGGTGGAAGGAT | TGGCTCGGGTGTTATTG |
| Maker00015350 | TAGCATCAAGCAAACCAAC | CTATGTACCACCTAGCCTCC |
| Maker00015605 | ACAATCTCCCAGTCCGC | AAATCAATGTCCCAAGAGC |
| Maker00015615 | GGTCGGATTTGTTTTGC | CTTTGCTTGATGCTATTGTG |
| Maker00015619 | GTGTATGGTTCTTTGGAGGA | GGTTTCTTTGCTTGATGCT |
| Maker00015620 | TCGTGGGGAACATAGACC | GAAGCACTCCATCAGCCT |
| Maker00016561 | GGCAATAAGCCAACCCT | TTAGCAACCGAAGACCAA |
| Maker00017748 | GTTGGAACTCTTTACTGATGC | GACGACCACTTTATTAGGCT |
| Maker00024456 | TCCTTCCTCCCAAATCC | GCACAAAATCATAGTCCCAT |
| Maker00025120 | CAGACTATCACCCGACCTT | CCTTAATTGCGTGCTCC |
| CcEF1a | TCCAAGGCACGGTATGAT | CCTGAAGAGGGAGACGAA |

Table A2.
Chemical composition analysis of essential oil of C. camphora var. linaloolifera leaves.
Table A2.
Chemical composition analysis of essential oil of C. camphora var. linaloolifera leaves.
| Number | Retention Time (s) | Component | Chemical Formula | Relative Content (%) |
|---|---|---|---|---|
| 1 | 3.82 | Furan,2-methoxy | C5H6O2 | 0.16 |
| 2 | 6.74 | Alpha.-phellandrene | C10H16 | 0.05 |
| 3 | 7.32 | 2-Methylbicyclo [4.3.0]non-1(6)-ene | C10H16 | 0.16 |
| 4 | 9.23 | 2,4,6-Octatriene, 2,6-dimethyl | C10H16 | 0.06 |
| 5 | 9.47 | o-Cymene | C10H14 | 0.08 |
| 6 | 9.59 | D-Limonene | C10H16 | 0.56 |
| 7 | 9.87 | 3-Carene | C10H16 | 0.68 |
| 8 | 10.47 | Gamma.-Terpinene | C10H16 | 0.08 |
| 9 | 12.41 | Linalool | C10H18O | 88.3 |
| 10 | 12.71 | 1,3,8-p-Menthatriene | C10H14 | 0.22 |
| 11 | 13.2 | Camphor | C10H16O | 0.16 |
| 12 | 14.05 | Terpinen-4-ol | C10H18O | 0.25 |
| 13 | 14.42 | L-.alpha.-terpineol | C10H18O | 1.25 |
| 14 | 15.43 | 2,6-Octadien-1-ol, 3,7-dimethyl- | C10H18O | 0.45 |
| 15 | 16.14 | Geraniol | C10H18O | 0.81 |
| 16 | 17.08 | Safrole | C10H10O2 | 0.11 |
| 17 | 18.72 | Alpha.-cubebene | C15H24 | 0.46 |
| 18 | 19.84 | Cyclohexane, 1-ethenyl-1-methyl-2,4-bis(1-methylethenyl)-, [1S-(1.alpha.,2.beta.,4.beta.)] | C15H24 | 0.43 |
| 19 | 20.11 | Methyleugenol | C11H14O2 | 0.11 |
| 20 | 20.32 | Delta.-selinene | C15H24 | 0.13 |
| 21 | 20.89 | 7-Epi-.alpha.-selinene | C15H24 | 0.1 |
| 22 | 20.98 | Naphthalene, 1,2,3,5,6,7,8,8a-octahydro-1,8a-dimethyl-7-(1-methylethenyl)-, [1R-(1.alpha.,7.beta.,8a.alpha.)] | C15H24 | 0.97 |
| 23 | 21.45 | Alloaromadendrene | C15H24 | 0.23 |
| 24 | 21.74 | Gamma.-muurolene | C15H24 | 0.29 |
| 25 | 22.78 | Cubenene | C15H24 | 0.09 |
| 26 | 23.85 | Guaiol | C15H26O | 0.25 |
| 27 | 24.06 | 2-Cyclohexene-1-carboxaldehyde, 2,6-dimethyl-6-(4-methyl-3-pentenyl) | C15H24O | 0.21 |
| 28 | 24.31 | (−)-Spathulenol | C15H24O | 0.09 |
| 29 | 6.91 | (1R)-2,6,6-Trimethylbicyclo [3.1.1]hept-2-ene((1R)-2,6,6 | C10H16 | 0.48 |
| 30 | 8.5 | Bicyclo [3.1.1]heptane, 6,6-dimethyl-2-methylene-, (1S)-((1S) | C10H16 | 0.43 |

Table A3.
Chemical composition analysis of the essential oil of C. kanehirae leaves.
Table A3.
Chemical composition analysis of the essential oil of C. kanehirae leaves.
| Number | Retention Time (s) | Component | Chemical Formula | Relative Content (%) |
|---|---|---|---|---|
| 1 | 14.5494 | 1R-.Alpha.-pinene | C10H16 | 1.31 |
| 2 | 16.6985 | Beta.-thujene | C10H16 | 0.34 |
| 3 | 16.8461 | Beta.-pinene | C10H16 | 0.57 |
| 4 | 17.045 | 5-Hepten-2-one, 6-methyl | C8H14O | 0.12 |
| 5 | 17.2567 | Beta.-myrcene | C10H16 | 0.75 |
| 6 | 18.1677 | Alpha.-phellandrene | C10H16 | 7.14 |
| 7 | 18.2703 | (1S)-(+)-3-Carene | C10H16 | 0.76 |
| 8 | 18.6809 | (+)-4-Carene | C10H16 | 0.5 |
| 9 | 18.7579 | 4-Cymene | C10H14 | 0.3 |
| 10 | 19.0081 | o-Cymol | C10H14 | 10.94 |
| 11 | 19.2134 | Limonene | C10H16 | 2.81 |
| 12 | 19.316 | Beta.-phellandrene | C10H16 | 0.33 |
| 13 | 19.3801 | Eucalyptol | C10H18O | 6.89 |
| 14 | 19.8741 | Ocimene | C10H16 | 0.28 |
| 15 | 20.4579 | Gamma.-terpinene | C10H16 | 0.97 |
| 16 | 21.4331 | 2-Carene | C16H16 | 0.13 |
| 17 | 21.6063 | Terpinolene | C10H16 | 1.09 |
| 18 | 21.818 | Benzene, 1-methyl-4-(1-methylethenyl) | C10H12 | 0.15 |
| 19 | 22.1516 | Linalool | C10H18O | 29.99 |
| 20 | 22.5493 | Rose oxide | C10H18O | 0.03 |
| 21 | 24.1275 | (−)-Camphor | C10H16O | 0.27 |
| 22 | 25.0128 | Borneol | C10H18O | 0.1 |
| 23 | 25.2694 | (-)-4-Terpineol | C10H18O | 3.2 |
| 24 | 25.449 | Thymol | C10H14O | 0.26 |
| 25 | 25.757 | Alpha.-terpineol | C10H18O | 4.06 |
| 26 | 26.2253 | cis-Piperitol | C10H20O | 0.09 |
| 27 | 26.6359 | Citronellol | C10H20O | 3.23 |
| 28 | 31.6719 | Beta.-bourbonene | C15H24 | 0.07 |
| 29 | 31.6719 | Beta.-elemene | C15H24 | 0.15 |
| 30 | 32.3134 | (−)-.Alpha.-gurjunene | C15H24 | 0.12 |
| 31 | 32.5251 | (−)-.Alpha. | C15H24 | 0.01 |
| 32 | 32.6919 | Caryophyllene | C15H24 | 1.66 |
| 33 | 32.8651 | Trans-.alpha.-bergamotene | C15H24 | 0.01 |
| 34 | 33.1795 | (+)-Cyclosativene | C15H24 | 0.12 |
| 35 | 33.6029 | (−)-.Beta.-santalene | C15H24 | 0.48 |
| 36 | 33.6542 | Alpha.-caryophyllene | C15H24 | 0.47 |
| 37 | 33.7889 | (−)-Alloaromadendrene | C15H24 | 0.09 |
| 38 | 33.9942 | Gamma.-muurolene | C15H24 | 0.12 |
| 39 | 34.052 | 2-Isopropenyl-4a,8-dimethyl-1,2,3,4,4a,5,6,7-octahydronaphthalene | C15H24 | 0.25 |
| 40 | 34.3021 | Germacrene D | C15H24 | 0.29 |
| 41 | 34.6485 | Phenol, 2,5-bis(1,1-dimethylethyl) | C14H22O | 0.92 |
| 42 | 34.7127 | Gamma.-selinene | C15H24 | 0.6 |
| 43 | 35.181 | Delta.-cadinene | C15H24 | 1.54 |
| 44 | 35.3222 | Calamenene | C15H24 | 0.28 |
| 45 | 35.7135 | (−)-.Alpha.-cadinene | C15H24 | 0.1 |
| 46 | 36.0985 | Nerolidol 2 | C15H26O | 3.73 |
| 47 | 36.6758 | (+)-Aromadendrene((+) | C15H24 | 0.04 |
| 48 | 36.7656 | Oxalic acid, 1-menthyl pentyl ester | C4H6O4 | 0.09 |
| 49 | 36.9581 | Caryophyllene oxide | C15H24O | 0.16 |
| 50 | 37.1441 | Guaiol | C15H24O | 0.68 |
| 51 | 37.4457 | Ledol | C15H24O | 0.07 |
| 52 | 38.1449 | Alpha.-cadinol | C15H24O | 0.3 |
| 53 | 38.3759 | (−)-Alloisolongifolene | C15H24 | 0.36 |
| 54 | 38.4464 | Beta.-eudesmol | C15H24O | 0.17 |
| 55 | 38.5619 | Bulnesol | C15H24O | 0.75 |
| 56 | 38.9019 | 2-Dodecanone | C15H24O | 0.05 |
| 57 | 40.7559 | Alpha.-phellandrene | C10H16 | 0.08 |
| 58 | 36.9581 | Caryophyllene oxide | C15H24O | 0.16 |
| 59 | 37.1441 | Guaiol | C15H24O | 0.68 |

Table A4.
Chemical composition analysis of essential oil of C. longepaniculatum leaves.
Table A4.
Chemical composition analysis of essential oil of C. longepaniculatum leaves.
| Number | Retention Time (s) | Component | Chemical Formula | Relative Content (%) |
|---|---|---|---|---|
| 1 | 10.9376 | 4-Hexen-1-ol, (E) | C6H12O | 0.01 |
| 2 | 14.5494 | 1R-.Alpha.-pinene | C10H16 | 5.89 |
| 3 | 15.4796 | Camphene | C10H16 | 0.19 |
| 4 | 16.6985 | Beta.-thujene | C10H16 | 15.8 |
| 6 | 16.9359 | (S)-.Beta.-pinene | C10H16 | 4.39 |
| 7 | 17.2567 | Beta.-myrcene | C10H16 | 2.34 |
| 8 | 17.4299 | Alpha.-phellandrene | C10H16 | 0.12 |
| 9 | 18.6809 | (+)-4-Carene(+) | C10H16 | 1.22 |
| 10 | 19.0081 | o-Cymol | C10H14 | 0.04 |
| 11 | 19.2134 | Limonene | C10H16 | 0.38 |
| 12 | 19.316 | Beta.-phellandrene | C10H16 | 0.45 |
| 13 | 19.3801 | Eucalyptol | C10H18O | 39.55 |
| 14 | 19.8741 | Ocimene | C10H16 | 0.12 |
| 15 | 20.4579 | Gamma.-terpinene | C10H16 | 1.8 |
| 16 | 21.0417 | cis-.Beta.-terpineol | C10H18O | 0.74 |
| 17 | 21.6063 | Terpinolene | C10H16 | 0.53 |
| 18 | 22.1516 | Linalool | C10H18O | 0.13 |
| 19 | 23.0369 | Fenchol | C10H18O | 0.01 |
| 20 | 23.672 | (E,Z)-Alloocimene | C10H16 | 0.01 |
| 21 | 24.2622 | Neopentylidenecyclohexane | C11H20 | 0.01 |
| 22 | 24.9358 | (−)-.Alpha.-terpineol | C10H18O | 1.4 |
| 23 | 25.0128 | Borneol | C10H18O | 0.18 |
| 24 | 25.2694 | (−)-4-Terpineol | C10H18O | 3.06 |
| 25 | 25.757 | Alpha.-terpineol | C10H18O | 13.47 |
| 26 | 26.2253 | cis-Piperitol | C10H18 | 0.07 |
| 27 | 26.6359 | Citronellol | C10H20O | 0.12 |
| 28 | 30.1514 | Delta.-elemene | C15H24 | 0.01 |
| 29 | 31.2228 | Di-epi-.alpha.-cedrene | C15H24 | 0.02 |
| 30 | 31.4024 | Copaene | C15H24 | 0.02 |
| 31 | 31.6719 | Beta.-bourbonene | C15H24 | 0.02 |
| 32 | 31.7168 | Beta.-elemene | C15H24 | 0.12 |
| 33 | 32.3134 | (−)-.Alpha.-gurjunene | C15H24 | 0.01 |
| 34 | 32.5251 | (−)-.Alpha.-santalene | C15H24 | 0.01 |
| 35 | 32.6213 | (+)-Calarene | C15H24 | 0.01 |
| 36 | 32.6919 | Caryophyllene | C15H24 | 1.66 |
| 37 | 32.7945 | Gamma.-elemene | C15H24 | 0.02 |
| 38 | 32.8651 | Trans-.alpha.-bergamotene | C15H24 | 0.01 |
| 39 | 33.1987 | (−)-Alloaromadendrene | C15H24 | 0.06 |
| 40 | 33.6542 | Alpha.-caryophyllene | C15H24 | 0.98 |
| 41 | 34.1546 | Gamma.-cadinene | C15H24 | 0.02 |
| 42 | 34.3021 | Germacrene D | C15H24 | 0.35 |
| 43 | 34.3855 | Beta.-maaliene | C15H24 | 0.03 |
| 44 | 34.5523 | Beta.-selinene | C15H24 | 0.16 |
| 45 | 34.7127 | Gamma.-selinene | C15H24 | 1.74 |
| 46 | 34.8025 | (+)-.Alpha.-elemene | C15H24 | 0.03 |
| 47 | 35.181 | Delta.-cadinene | C15H24 | 0.04 |
| 48 | 35.3735 | Epizonarene | C15H24 | 0.02 |
| 49 | 36.0985 | Nerolidol 2 | C15H26O | 0.02 |
| 50 | 36.7913 | (−)-Spathulenol | C15H26O | 0.07 |
| 51 | 36.9581 | Caryophyllene oxide | C15H26O | 0.06 |
| 52 | 37.1441 | Guaiol | C15H26O | 0.08 |
| 53 | 38.3759 | (-)-Alloisolongifolene((−)-ALLO | C15H24 | 0.01 |
| 54 | 38.4464 | Beta.-eudesmol | C15H26O | 0.01 |
| 55 | 38.5619 | Bulnesol | C15H26O | 0.01 |
| 56 | 41.2499 | 2-Pentadecanone, 6,10,14-trimethyl | C18H36O | 0.02 |
| 57 | 41.3718 | 6,10,14-Trimethyl-pentadecan-2-ol | C18H36O | 0.01 |

Table A5.
Chemical composition analysis of essential oil of C. micranthum leaves.
Table A5.
Chemical composition analysis of essential oil of C. micranthum leaves.
| Number | Retention Time (s) | Component | Chemical Formula | Relative Content (%) |
|---|---|---|---|---|
| 1 | 5.3627 | Butanal, 3-methyl | C5H10O | 0.01 |
| 2 | 6.017 | Furan, 2-ethyl | C6H8O | 0.01 |
| 3 | 8.6858 | Octane | C8H18 | 0.01 |
| 4 | 8.8911 | Hexanal | C6H12O | 0.01 |
| 5 | 10.8478 | 2-Hexenal | C6H10O | 0.01 |
| 6 | 10.9119 | 3-Hexen-1-ol, (Z) | C6H12O | 0.01 |
| 7 | 12.7018 | Nonane | C9H20 | 0.08 |
| 8 | 12.8814 | Heptanal | C7H14O | 0.01 |
| 9 | 14.5815 | 1R-.Alpha.-pinene | C10H16 | 0.1 |
| 10 | 15.486 | Camphene | C10H16 | 0.07 |
| 11 | 16.8653 | (S)-.Beta.-pinene | C10H16 | 0.04 |
| 12 | 17.0385 | 5-Hepten-2-one, 6-methyl | C8H14O | 0.01 |
| 13 | 17.2695 | Beta.-myrcene | C10H16 | 0.02 |
| 14 | 17.9559 | Octanal | C8H16O | 0.11 |
| 15 | 18.1741 | Alpha.-phellandrene | C10H16 | 0.05 |
| 16 | 18.2831 | (1S)-(+)-3-Carene | C10H16 | 0.02 |
| 17 | 18.405 | 2-Hexen-1-ol, acetate | C8H14O2 | 0.01 |
| 18 | 19.2197 | D-Limonene | C10H16 | 0.02 |
| 19 | 19.3929 | 4-Carene | C10H16 | 0.13 |
| 20 | 19.8998 | Ocimene | C10H16 | 0.9 |
| 21 | 20.4643 | Gamma.-terpinene | C10H16 | 0.01 |
| 22 | 20.9198 | Formic acid, octyl ester | C9H18O2 | 0.03 |
| 23 | 20.9968 | 2-Pentene, 3-methyl | C6H12 | 0.03 |
| 24 | 21.6062 | 2-Carene | C10H16 | 0.02 |
| 25 | 21.741 | 1-Undecene | C11H22 | 0.04 |
| 26 | 21.8949 | Neral | C10H16O | 0.01 |
| 27 | 22.1323 | Linalool | C10H18O | 0.25 |
| 28 | 22.344 | Nonanal | C9H18O | 1.14 |
| 29 | 24.4161 | 2-Nonenal, (E) | C9H16O | 0.01 |
| 30 | 24.7305 | 1-Nonanol | C9H20O | 0.93 |
| 31 | 25.0897 | Bornel | C10H18O | 0.04 |
| 32 | 25.2886 | (−)-4-Terpineol | C10H18O | 0.03 |
| 33 | 25.5196 | 2-Decanone | C10H20O | 0.1 |
| 34 | 25.7569 | (+)-4-Carene | C10H16 | 0.03 |
| 35 | 26.3279 | Decanal | C10H20O | 4.12 |
| 36 | 26.3728 | Cyclohexane, hexyl | C12H24 | 0.03 |
| 37 | 26.6936 | Geranial | C10H16O | 0.01 |
| 38 | 27.0079 | Alpha.-terpinene | C10H16O | 0.01 |
| 39 | 27.1298 | Cyclohexanone, 2-butyl | C10H18O | 0.01 |
| 40 | 27.2453 | 2-Pentylcyclopentanone | C10H18O | 0.03 |
| 41 | 27.4249 | Nerol | C10H18O | 0.02 |
| 42 | 27.5917 | Acetaldehyde, (3,3-dimethylcyclohexylidene) | C10H16O | 0.01 |
| 43 | 27.7906 | Beta.-cyclohomocitral | C10H16O | 0.01 |
| 44 | 27.8419 | 2-Decenal | C10H18O | 0.03 |
| 45 | 28.0151 | Nonanoic acid | C9H18O2 | 0.12 |
| 46 | 28.0921 | Cyclooctane | C8H16 | 0.29 |
| 47 | 28.631 | D,L-Isobornyl acetate | C12H20O2 | 0.23 |
| 48 | 28.7401 | 2-Undecanone | C11H22O | 0.68 |
| 49 | 29.0095 | Phenol, 2-methyl-5-(1-methylethyl) | C10H14O | 0.02 |
| 50 | 29.2084 | Undecanal | C11H22O | 0.52 |
| 51 | 29.4201 | 2-Methoxy-4-vinylphenol | C9H10O2 | 0.01 |
| 52 | 29.5997 | 2,4-Dodecadienal, (E,E) | C12H20O | 0.01 |
| 53 | 29.9012 | 3-Undecene, (Z) | C11H22 | 0.03 |
| 54 | 30.0359 | 2-Hexene, 3-methyl-, (Z) | C7H14 | 0.01 |
| 55 | 30.0809 | (−)-Alloaromadendrene | C15H24 | 0.01 |
| 56 | 30.1514 | Delta.-elemene | C15H24 | 0.06 |
| 57 | 30.5107 | Alpha.-cubebene | C15H24 | 0.06 |
| 58 | 31.0816 | n-Decanoic acid | C10H20O2 | 1.3 |
| 59 | 31.2292 | Ylangene | C15H24 | 2.02 |
| 60 | 31.4088 | Copaene | C15H24 | 0.73 |
| 61 | 31.6718 | 2-Dodecanone | C12H24O | 0.65 |
| 62 | 31.7232 | Beta.-elemene | C15H24 | 0.16 |
| 63 | 31.922 | 1,2-Decanediol | C10H22O2 | 0.18 |
| 64 | 32.0054 | Acetic acid, decyl ester | C12H24O2 | 0.04 |
| 65 | 32.1337 | Dodecanal | C12H24O | 1.07 |
| 66 | 32.2621 | (−)-Isocaryophyllene | C15H24 | 0.11 |
| 67 | 32.3326 | (−)-.Alpha.-gurjunene | C15H24 | 0.6 |
| 68 | 32.3839 | Trans-.alpha.-bergamotene | C15H24 | 0.82 |
| 69 | 32.493 | Terpinolene | C10H16 | 0.16 |
| 70 | 32.5636 | (−)-.Alpha.-santalene | C14H26O | 0.39 |
| 71 | 32.7689 | β-Caryophyllene | C15H24 | 18.94 |
| 72 | 32.8907 | (+)-.Gamma.-maalien | C15H24 | 0.11 |
| 73 | 32.9485 | (+)-Epi-bicyclosesquiphellandrene | C15H24 | 0.21 |
| 74 | 33.0255 | (+)-Calarene | C15H24 | 0.08 |
| 75 | 33.2244 | (+)-Aromadendrene | C15H24 | 1.4 |
| 76 | 33.6798 | Alpha.-caryophyllene | C15H24 | 2.28 |
| 77 | 33.8017 | Decanoic acid, 2-hydroxy-1-(hydroxymethyl)ethyl ester | C13H26O4 | 2.23 |
| 78 | 34.193 | Alpha.-muurolene | C15H24 | 0.45 |
| 79 | 34.3214 | Germacrene D | C15H24 | 0.6 |
| 80 | 34.7383 | Alpha.-selinene | C15H24 | 0.36 |
| 81 | 34.8474 | Beta.-bisabolene | C15H24 | 1.9 |
| 82 | 34.9629 | Aromandendrene | C15H24 | 0.1 |
| 83 | 35.0591 | 3-Methylene-1,5,5-trimethylcyclohexene | C10H16 | 0.13 |
| 84 | 35.1489 | 4,9-Cadinadiene | C15H20O2 | 0.76 |
| 85 | 35.2388 | Delta.-cadinene | C15H24 | 2.29 |
| 86 | 35.367 | Calamenene | C15H22 | 1.66 |
| 87 | 35.4825 | Butylated hydroxytoluene | C15H24O | 0.16 |
| 88 | 35.5595 | Gamma.-elemene | C15H24 | 0.07 |
| 89 | 35.6622 | Alpha.-himachalene | C15H24 | 1.5 |

Table A6.
Transcriptome sequencing data statistics.
Table A6.
Transcriptome sequencing data statistics.
| Sample | Raw Reads (M) | Clean Reads (M) | Total Clean Bases (Gb) | Q30 (%) | Genome Mapping (%) | Mapping (%) |
|---|---|---|---|---|---|---|
| Cc-1 | 43.82 | 42.39 | 6.4 | 92.79 | 88.57 | 74.59 |
| Cc-2 | 43.82 | 42.61 | 6.42 | 93.08 | 87.2 | 75.26 |
| Cc-3 | 43.82 | 42.45 | 6.39 | 93.25 | 88.02 | 75.74 |
| Ck-1 | 43.82 | 42.3 | 6.34 | 94.26 | 86.48 | 68.45 |
| Ck-2 | 43.82 | 42.81 | 6.42 | 94.26 | 86.07 | 69.24 |
| Ck-3 | 43.82 | 43.02 | 6.45 | 93.78 | 87.7 | 69.82 |
| Cl-1 | 43.82 | 42.39 | 6.36 | 93.25 | 85.32 | 63.43 |
| Cl-2 | 43.82 | 42.61 | 6.39 | 93.1 | 86.38 | 65.03 |
| Cl-3 | 43.82 | 42.45 | 6.37 | 93.28 | 86.06 | 62.65 |
| Cm-1 | 43.82 | 42.12 | 6.32 | 89.25 | 86.31 | 68.04 |
| Cm-2 | 47.33 | 46.06 | 6.91 | 90.9 | 83.3 | 69.1 |
| Cm-3 | 45.57 | 44.64 | 6.7 | 91.24 | 87.88 | 70.26 |

Table A7.
Identification of structural genes of the MVA and MEP pathways.
Table A7.
Identification of structural genes of the MVA and MEP pathways.
| Pathway | Pathway Number | Gene ID | Gene Name | Abbreviation | Enzyme Number |
|---|---|---|---|---|---|
| MVA | K00626 | Maker00010292 | acetyl-CoA C-acetyltransferase | ACAT | 2.3.1.9 |
| Maker00011273 | |||||
| Maker00011277 | |||||
| Maker00011286 | |||||
| Maker00018877 | |||||
| Maker00019487 | |||||
| K01641 | Maker00003407 | hydroxymethylglutaryl-CoA synthase | HMGS | 2.3.3.10 | |
| Maker00028799 | |||||
| K00021 | Maker00001945 | hydroxymethylglutaryl-CoA reductase | HMGR | 1.1.1.34 | |
| Maker00015796 | |||||
| Maker00021635 | |||||
| Maker00027071 | |||||
| K00869 | Maker00015376 | mevalonate kinase | MK | 2.7.1.36 | |
| Maker00015398 | |||||
| K00938 | Maker00026239 | phosphomevalonate kinase | PMK | 2.7.4.2 | |
| Maker00026952 | |||||
| K01597 | Maker00012131 | diphosphomevalonate decarboxylase | MDC | 4.1.1.33 | |
| Maker00012132 | |||||
| Maker00012236 | |||||
| Maker00014143 | |||||
| Maker00014156 | |||||
| Maker00014205 | |||||
| K00787 | Maker00006917 | farnesyl diphosphate synthase | FDPS | 2.5.1.1 2.5.1.10 | |
| Maker00008168 | |||||
| MEP | K01662 | Maker00012804 | 1-deoxy-D-xylulose-5-phosphate synthase | DXS | 2.2.1.7 |
| Maker00004598 | |||||
| Maker00015435 | |||||
| Maker00023473 | |||||
| Maker00027418 | |||||
| Maker00027873 | |||||
| K00099 | Maker00016636 | 1-deoxy-D-xylulose-5-phosphate reductoisomerase | DXR | 1.1.1.267 | |
| K00991 | Maker00013631 | 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase | CMS | 2.7.7.60 | |
| K00919 | Maker00011107 | 4-diphosphocytidyl-2-C-methyl-D-erythritol kinase | CMK | 2.7.1.148 | |
| K01770 | Maker00019544 | 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase | MDS | 4.6.1.12 | |
| K03526 | Maker00005988 | (E)-4-hydroxy-3-methylbut-2-enyl-diphosphate synthase | HDS | 1.17.7.1 1.17.7.3 | |
| Maker00025110 | |||||
| K03527 | Maker00005068 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate reductase | HDR | 1.17.7.4 | |
| Maker00025695 | |||||
| K00787 | Maker00006917 | farnesyl diphosphate synthase | GPPS | 2.5.1.1 2.5.1.10 | |
| Maker00008168 | |||||
| K01823 | Maker00001381 | isopentenyl-diphosphate Delta-isomerase | IDI | 5.3.3.2 | |
| Maker00016783 | |||||
| Maker00018376 |
References
- Singh, M.K.; Sharma, M.; Sharma, C.L. Wood anatomy of some members of family Lauraceae with reference to their identification. J. Indian Acad. Wood Sci. 2015, 12, 137–144. [Google Scholar] [CrossRef]
- Chen, W.Q.; Liu, L.; He, G.X.; Zhang, D. Diverse essential oils from leaves of three chemotypes Cinnamomum camphora. Ekoloji Derg. 2019, 28, 87–91. [Google Scholar]
- Fu, C.; Liu, X.; Liu, Q.; Qiu, F.Y.; Yan, J.D.; Zhang, Y.T.; Zhang, T.; Li, J.N. Variations in essential oils from the leaves of Cinnamomum bodinieri in China. Molecules 2023, 28, 3659. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, B.; Marques, A.; Ramos, C.; Teixeira, B.; Marques, A.; Ramos, C.; Neng, N.R.; Nogueira, J.M.; Saraiva, J.A.; Nunes, M.L. Chemical composition and antibacterial and antioxidant properties of commercial essential oils. Ind Crops Prod. 2013, 43, 587–595. [Google Scholar] [CrossRef]
- Sara, B. Essential oils: Their antibacterial properties and potential applications in foods—A review. Int. J. Food Microbiol. 2004, 94, 223–253. [Google Scholar]
- Damasceno, C.S.B.; Fabri-Higaki, N.T.; Dias, J.F.G.; Miguel, M.D.; Miguel, O.G. Chemical composition and biological activities of essential oils in the family Lauraceae: A systematic review of the literature. Planta Med. 2019, 85, 1054–1072. [Google Scholar] [CrossRef]
- Wang, X.D.; Xu, C.Y.; Zheng, Y.J.; Wu, Y.F.; Zhang, Y.T.; Zhang, T.; Xiong, Z.Y.; Yang, H.K.; Li, J.; Fu, C.; et al. Chromosome-level genome assembly and resequencing of camphor tree (Cinnamomum camphora) provides insight into phylogeny and diversification of terpenoid and triglyceride biosynthesis of Cinnamomum. Hortic. Res. 2022, 9, uhac216. [Google Scholar] [CrossRef]
- Shi, W.Y.; He, W.; Wen, G.Y.; Guo, D.X.; Long, G.Y.; Lin, Y.G. Study on chemical constituents of the essential oil and classification of types from Cinnamomum camphora. J. Integr. Plant Biol. 1989, 31, 209–214. [Google Scholar]
- Chen, C.; Zheng, Y.; Zhong, Y.; Wu, Y.; Li, Z.; Xu, L.A.; Xu, M. Transcriptome analysis and identification of genes related to terpenoid biosynthesis in Cinnamomum camphora. BMC Genom. 2018, 19, 550. [Google Scholar] [CrossRef]
- Guo, S.; Geng, Z.; Zhang, W.; Liang, J.; Wang, C.; Deng, Z.; Du, S. The chemical composition of essential oils from Cinnamomum camphora and their insecticidal activity against the stored product pests. Int. J. Mol. Sci. 2016, 17, 1836. [Google Scholar] [CrossRef]
- Guo, X.L.; Cui, M.; Deng, M.; Liu, X.; Huang, X.; Zhang, X.; Luo, L. Molecular differentiation of five Cinnamomum camphora chemotypes using desorption atmospheric pressure chemical ionization mass spectrometry of raw leaves. Sci. Rep. 2017, 7, 46579. [Google Scholar] [CrossRef] [PubMed]
- Carretero-Paulet, L.; Ahumada, I.; Cunillera, N.; Rodrıguez-Concepción, M.; Ferrer, A.; Boronat, A.; Campos, N. Expression and molecular analysis of the Arabidopsis DXR gene encoding 1-deoxy-D-xylulose 5-phosphate reductoisomerase, the first committed enzyme of the 2-C-methyl-D-erythritol 4-phosphate pathway. Plant Physiol. 2002, 129, 1581–1591. [Google Scholar] [CrossRef]
- Aharoni, A.; Jongsma, M.A.; Kim, T.Y.; Ri, M.B.; Giri, A.P.; Verstappen, F.W.A.; Schwab, W.; Bouwmeester, H.J. Metabolic engineering of terpenoid biosynthesis in plants. Phytochem. Rev. 2006, 5, 49–58. [Google Scholar] [CrossRef]
- Tholl, D. Biosynthesis and biological functions of terpenoids in plants. Adv. Biochem. Eng. Biot. 2015, 148, 63. [Google Scholar]
- Vranova, E.; Coman, D.; Gruissem, W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu. Rev. Plant Biol. 2013, 64, 665–700. [Google Scholar] [CrossRef] [PubMed]
- Pu, X.; Dong, X.; Li, Q.; Chen, Z.; Liu, L. An update on the function and 579 regulation of methylerythritol phosphate and mevalonate pathways and their 580 evolutionary dynamics. J. Integr. Plant Biol. 2021, 63, 1211–1226. [Google Scholar] [CrossRef] [PubMed]
- Dillies, M.A.; Rau, A.; Aubert, J.; Hennequet-Antier, C.; Jeanmougin, M.; Servant, N.; Keime, C.; Marot, G.; Castel, D.; Estelle, J.; et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief Bioinform. 2013, 14, 671. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Y.; Wang, Y.H.; Shen, S.K. Transcriptome analysis of Cinnamomum chago: A revelation of candidate genes for abiotic stress response and terpenoid and fatty acid biosyntheses. Front. Genet. 2018, 9, 505. [Google Scholar] [CrossRef] [PubMed]
- Qiu, F.; Wang, X.; Zheng, Y.; Wang, H.; Liu, X.; Su, X. Full-length transcriptome sequencing and different chemotype expression profile analysis of genes related to monoterpenoid biosynthesis in Cinnamomum porrectum. Int. J. Mol. Sci. 2019, 20, 6230. [Google Scholar]
- Yang, Z.R.; Xie, C.Z.; Huang, Y.Y.; An, W.; Liu, S.; Huang, S.; Zheng, X. Metabolism and transcriptome profiling provides insight into the genes and transcription factors involved in monoterpene biosynthesis of borneol chemotype of Cinnamomum camphora induced by mechanical damage. PeerJ 2021, 9, e11465. [Google Scholar] [CrossRef]
- Zhao, X.; Yan, Y.; Zhou, W.H.; Feng, R.Z.; Shuai, Y.K.; Yang, L.; Liu, M.J.; He, X.Y.; Wei, Q. Transcriptome and metabolome reveal the accumulation of secondary metabolites in different varieties of Cinnamomum longepaniculatum. BMC Plant Biol. 2022, 22, 243. [Google Scholar] [CrossRef] [PubMed]
- Chaw, S.M.; Liu, Y.C.; Wu, Y.W.; Wang, H.Y.; Lin, C.Y.L.; Wu, C.S.; Ke, H.M.; Chang, L.Y.; Hsu, C.Y.; Yang, H.T.; et al. Stout camphor tree genome fills gaps in understanding of flowering plant genome evolution. Nat. Plants 2019, 5, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.H.; Xiang, S.; Zhang, Q.G.; Xiao, L.; Zhang, D.Y.; Zhang, P.L.; Chen, D.Q.; Hao, Y.; Liu, D.K.; Ding, L.; et al. The camphor tree genome enhances the understanding of magnoliid evolution. J. Genet. Genom. 2021, 49, 249–253. [Google Scholar] [CrossRef] [PubMed]
- Jiang, R.H.; Chen, X.L.; Liao, X.Z.; Peng, D.; Han, X.X.; Zhu, C.S.; Wang, P.; Hufnagel, D.E.; Wang, L.; Li, K.X.; et al. A chromosome-level genome of the camphor tree and the underlying genetic and climatic factors for its top-Geoherbalism. Front. Plant Sci. 2022, 13, 827890. [Google Scholar] [CrossRef]
- Li, D.; Lin, H.Y.; Wang, X.; Bi, B.; Gao, Y.; Shao, L.M.; Zhang, R.L.; Liang, Y.W.; Xia, Y.P.; Zhao, Y.P.; et al. Genome and whole-genome resequencing of Cinnamomum camphora elucidate its dominance in subtropical urban landscapes. BMC Biol. 2023, 21, 192. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Chen, Y.; Shi, C.; Huang, Z.B.; Zhang, Y.; Li, S.K.; Li, Y.; Ye, J.; Yu, C.; Li, Z.; et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 2018, 7, gix120. [Google Scholar] [CrossRef]
- Bolger, A.M.; Marc, L.; Bjoern, U. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Anders, S.; Pyl, P.; Huber, W. HTSeq: A python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zheng, Y.; Fu, C.; Yang, H.K.; Liu, X.L.; Qiu, F.Y.; Wang, X.D.; Wang, Z.D. Chemical variation and environmental influence on essential oil of Cinnamomum camphora. Molecules 2023, 28, 973. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Zhang, X.H.; Zou, S.Q.; Su, B.C.; Gao, J.X.; Zou, X.X. A new Cinnamomum camphora cultivar ‘Nanan 1’. Yuan Yi Xue Bao 2022, 49, 943–944. [Google Scholar]
- Hernandez-Leon, A.; González-Trujano, M.E.; Narváez-González, F.; Pérez-Ortega, G.; Rivero-Cruz, F.; Aguilar, M.I. Role of β-caryophyllene in the antinociceptive and anti-inflammatory effects of Tagetes lucida Cav. essential oil. Molecules 2020, 25, 675. [Google Scholar] [CrossRef]
- He, J.; Verstappen, F.; Jiao, A.; Dicke, M.; Bouwmeester, H.J.; Kappers, I.F. Terpene synthases in cucumber (Cucumis sativus) and their contribution to herbivore-induced volatile terpenoid emission. New Phytol. 2022, 233, 862–877. [Google Scholar] [CrossRef]
- Yang, Z.; Zhan, T.; Xie, C.; Huang, S.; Zheng, X. Genome-wide analyzation and functional characterization on the TPS family provide insight into the biosynthesis of mono-terpenes in the camphor tree. Plant Physiol. Biochem. 2023, 196, 55–64. [Google Scholar] [CrossRef]
- Moses, T.; Pollier, J.; Thevelein, J.M.; Goossens, A. Bioengineering of plant (tri)terpenoids: From metabolic engineering of plants to synthetic biology in vivo and in vitro. New Phytol. 2013, 200, 27–43. [Google Scholar] [CrossRef]
- Yu, Z.; Zhao, C.; Zhang, G. Genome-wide identification and expression profile of TPS gene family in Dendrobium officinale and the role of DoTPS10 in linalool biosynthesis. Int. J. Mol. Sci. 2020, 21, 5419. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Tholl, D.; Bohlmann, J.; Pichersky, E. The family of terpene synthases in plants: A mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J. 2011, 66, 212–229. [Google Scholar] [CrossRef]
- Chen, H.; Köllner, T.G.; Li, G.L.; Wei, G.; Chen, X.L.; Zeng, D.L.; Qian, Q.; Chen, F. Combinatorial evolution of a terpene synthase gene cluster explains terpene variations in Oryza. Plant Physiol. 2020, 182, 480–492. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).