Abstract
Circulating microRNAs (miRNA) are small noncoding RNA molecules that can be detected in bodily fluids without the need for major invasive procedures on patients. miRNAs have shown great promise as biomarkers for tumors to both assess their presence and to predict their type and subtype. Recently, thanks to the availability of miRNAs datasets, machine learning techniques have been successfully applied to tumor classification. The results, however, are difficult to assess and interpret by medical experts because the algorithms exploit information from thousands of miRNAs. In this work, we propose a novel technique that aims at reducing the necessary information to the smallest possible set of circulating miRNAs. The dimensionality reduction achieved reflects a very important first step in a potential, clinically actionable, circulating miRNA-based precision medicine pipeline. While it is currently under discussion whether this first step can be taken, we demonstrate here that it is possible to perform classification tasks by exploiting a recursive feature elimination procedure that integrates a heterogeneous ensemble of high-quality, state-of-the-art classifiers on circulating miRNAs. Heterogeneous ensembles can compensate inherent biases of classifiers by using different classification algorithms. Selecting features then further eliminates biases emerging from using data from different studies or batches, yielding more robust and reliable outcomes. The proposed approach is first tested on a tumor classification problem in order to separate 10 different types of cancer, with samples collected over 10 different clinical trials, and later is assessed on a cancer subtype classification task, with the aim to distinguish triple negative breast cancer from other subtypes of breast cancer. Overall, the presented methodology proves to be effective and compares favorably to other state-of-the-art feature selection methods.
1. Introduction
MicroRNAs (miRNAs) are noncoding RNA molecules of 18–25 nucleotides in length that regulate the expression of more than one third of human genes [1,2]. Since the discovery of the first miRNA in Caenorhabditis elegans [3], these molecules have been found in many organisms and tissue types. miRNAs have been shown to play an important role in cell biology, including differentiation, proliferation and apoptosis [4]. To date, there is evidence that miRNAs regulate different aspects of cancer development [5].
The biogenesis of miRNAs starts with a stem loop precursor created by RNA polymerase II, called primary precursor miRNA (pri-miRNA), that is cleaved by Drosa and DGCR8 proteins to obtain the precursor miRNAs (pre-miRNA) [6]. Finally, the pre-miRNA is cleaved by the Dicer/TRBP complex to create miRNA that represses or degrades the target mRNAs [7,8]. This machinery is altered in cancer cells, perturbing miRNA expression and accelerating the process of tumorigenesis. The discovery of cell-free circulating miRNAs in body fluids (blood, plasma, serum, urine, and cerebrospinal liquid) has put miRNAs in the focus of current research as promising cancer biomarkers [1,2,7,9,10,11,12]. Because the histological examination of tissues is an invasive and comparatively risky procedure, studying miRNAs in biological fluids offers a useful alternative for diagnosis, typing and management of cancer patients.
miRNA expression levels have proven to substantially vary relative to cell types. That makes it possible to use miRNAs to distinguish between cell types [13]. Furthermore, molecular signatures can be useful to differentiate between cancer types in general [14,15]. Another particularity is that these molecules are stable in extracellular environments: for example, they are resistant to pH and heat changes. Nowadays, the use of microarrays, real-time polymerase chain reaction (PCR) and next generation sequencing (NGS) technologies and the creation of databases give us the opportunity to study miRNAs as cancer biomarkers. Several studies have exploited the biomarker properties of miRNAs for cancer detection and classification, using machine learning techniques [16,17,18,19,20].
These works typically analyze thousands of different miRNAs, amounts that would make it impossible for medical experts to manually validate the results or to obtain novel insights. Furthermore, employing thousands of miRNAs in machine learning approaches translates into operating in feature spaces of thousands of dimensions, which nurtures the usual issues linked to the curse of dimensionality. Therefore, in addition to enhancing the interpretability of results, determining small, actionable subselections of features warrants approaches that are insensitive to biases emerging from batch effects (due to processing data from multiple studies, for example), from the use of sets of classifiers that vary in terms of their strengths and weaknesses or just from the nature of their technical foundations. Finding the smallest subset of circulating miRNAs that can identify the presence of cancer or the type of tumor is therefore of utmost practical importance.
In this work, we propose a new methodology to reduce the number of significant circulating miRNAs needed by machine learning techniques to detect and identify cancer types using 16 miRNA datasets from clinical trials. The technique relies on a heterogeneous ensemble of classifiers to provide more robust results than single algorithms or even homogeneous ensembles. The presented approach is first used to identify 10 different types of cancer, and then, in a second experiment, the same technique is applied to separate tumor subtypes in breast cancer. The methodology not only is proven to be effective but also compares favorably to current state-of-the-art techniques.
While a similar technique was presented in [21,22], the approach we propose features several improvements and important innovations that set it apart from previous contributions: (i) previous works did not select for circulating miRNAs, and thus, resulting signatures could not be easily measured in clinical practice; (ii) previous techniques needed extra parameters to be defined by the user (for example, a desired number of features), while the novel approach we propose does not require users to arbitrarily set values for thresholds; and (iii) finally, the amount of data used in the experimental verification greatly increased, getting a total of 16 gene expression omnibus (GEO) datasets.
2. Materials and Methods
First, we compiled a list of circulating miRNAs (mature sequence) based on 5 reviews of circulating miRNAs from cancer studies [1,2,23,24,25]. Next, from this list, we consider only the miRNAs that appear in blood, serum, urine, plasma and saliva. To narrow it further, we focus on the miRNAs that can be detected by Affymetrix platforms Affy-1 (GPL8786), Affy-2 (GPL14613) and Affy-3 (GPL16384). Our choice of restricting to datasets from Affymetrix platforms GPL8786, GPL14613 and GPL16384 has the aim of avoiding the known issue of miRNA expression levels being platform- and technology-dependent [26,27,28]. After this selection, a total of 253 miRNAs remain. The detailed list is included in Appendix A.
2.1. Feature Selection
As our objective is to select the most meaningful miRNAs to correctly classify the cancer types, we used a recursive ensemble feature selection algorithm where features in our datasets are expression values of different miRNAs. The idea behind recursive feature selection is to score each feature depending on its usefulness for the classification process, resorting to a classifier. Features with the lowest scores are then removed, and the process is iterated with the remaining features until the overall classification accuracy drops below a given threshold or when a user-defined number of features is reached. While this technique is effective, it still relies on a classification algorithm to score the features, and a single algorithm might be affected by bias when it assigns scores to the features. A way to reduce the bias is to exploit an ensemble of classification algorithms with different topologies, an idea that is proven to be effective for different problems [29,30,31].
For the ensemble, we selected 8 classifiers from the sci-kit learn toolbox [32] that all were proven to be effective for cancer classification using miRNAs [18] and that are able to score features according to their importance: Stochastic Gradient Descent (SGD), Support Vector Machine classifier (SVC), gradient boosting, random forest, logistic regression, passive aggressive classifier, ridge classifier and bagging. Parameters for each classifier, when different from the default, were taken from [18].
Different algorithms assess feature importance differently, as the scoring depends on the computational particularities of the algorithms. Bagging, gradient boosting and random forest use ensembles of classification trees. In these cases, we count the features that appear in the splits of the trees and rank them by frequency. For SVC, SGD, passive aggressive, logistic regression and ridge, the feature importance is given by the absolute value of the coefficients associated to each feature. Therefore, the ranking is based on the value of these coefficients.
As the ranking of each classifier has a different meaning, it is necessary to aggregate this information into an ensemble ranking. Each feature f is assigned a simple score , where is the number of times that feature appears among the top S over all classifier instances, while is the number of classifier instances used. Each classification algorithm has 10 instances, produced by a 10-fold stratified cross-validation (). The cross-validation is used to increase generality of the results. We selected a stratified cross-validation because it preserves the same ratio of samples for each class in the training and test. Next, the recursive feature algorithm will reduce the number of features S by at each iteration. For our experiments, we decided to stop the procedure when the global average accuracy among all classifiers drops to less than . The complete procedure is summarized by Algorithm 1.
| Algorithm 1: Recursive ensemble feature selection. |
 |
2.2. Cancer Type Classification
From the gene expression omnibus (GEO) repository [33], we selected 16 datasets for 10 different types of cancer based on clinical studies: Breast (BRCA), esophageal (ESCA), head and neck squamous cell (HNSC), liver hepatocellular (LIHC), prostate (PRCA), glioblastoma (GBM), colorectal (CRC), non-small-cell lung (NSCLC), gastric (GC) and ovarian (OVC), as summarized in Table 1. For each dataset, we downloaded the raw data and processed it using the function Affyrma() from the Matlab bioinformatics toolbox™. This function processes the probe intensity values using RMA background adjustment, quantile normalization and summarizing procedures and then outputs expression (nondimensional). The resulting aggregated dataset for our multi-class classification problem presents 845 samples, 253 features and 10 different tumor classes. Next, we applied Z-score normalization on the dataset to then run the feature selection algorithm in a 10-fold stratified cross-validation scheme.

Table 1.
Gene expression omnibus (GEO) repository datasets of miRNA cancer studies used in the project for platforms GPL8786, GPL14613 and GPL16384. BRCA: breast cancer; ESCA: esophageal cancer; HSNC: head and neck squamous cell cancer; LIHC: liver hepatocellular cancer; PRCA: prostate cancer; GBM: gliobastoma; CRC: colorectal cancer; NSCLC: non-small-cell lung cancer; GC: gastric cancer; OVC: ovarian cancer.
Then, we compared our results against two current state-of-the-art feature selection methodologies: a homogeneous ensemble classifier exploiting variations of SVC [29] and a feature selection tool based on genetic algorithms, called GALGO [43]. Since each algorithm contains stochastic elements, we run each algorithm 10 times and keep the set of features with the best average accuracy.
The homogeneous ensemble uses several runs of SVC to rank the features by weight and reduces the number of features by a given percentage at each step. In this case, we used the same parameters as for Algorithm 1: 20% step reduction and 90% accuracy as stop parameters. In contrast, for GALGO to obtain a fair comparison, the requested number of features is set to the resulting number of features from the heterogeneous ensemble feature selection classifier.
Finally, we analyzed the genes targeted by the candidate miRNAs using miRNet [44]. The parameters for the miRNet analysis are target genes as main function with a 0.05 betweenness filter and pathway enrichment analysis using the Kyoto Encyclopedia of Genes and Genomes [45] (KEGG) and Gene Ontology-Biological Process [46] (GO:BP). Using a betweenness filter implies that the genes must be targeted by at least 2 miRNAs.
2.3. Triple-Negative Breast Cancer Classification
Cancer tumors are divided into tumor subtypes, which can be treated by different strategies depending on their classification. From the available data in the GEO repository, we were able to compile a dataset to assess the possibility of classifying tumor subtypes (luminal A, luminal B, HER2-enriched, triple-negative and normal [47]) in breast cancer (BRCA) using circulating miRNAs. Then, we selected datasets GSE86277, GSE86278, GSE86281 and GSE46823, which are BRCA studies with subtype information. From the BRCA subtypes, triple-negative has the worst prognosis, as it is resistant to hormone therapies [48]. For this reason, we set the labels of the resulting dataset to separate the triple-negative subtype from the rest. Although making an analysis of all the subtypes would have been more interesting, the unbalance in the subtype samples found in the original data makes it impossible; thus, more precisely, the resulting dataset has 139 triple-negative samples and only 44 from the rest of the subtypes, for a total of 183 samples, 253 features and 2 classes (triple-negative/Other).
Next, we applied the function Affyrma from the Matlab bioinformatics toolbox™. Then, we applied Z-score normalization on the dataset to run the feature selection algorithm in a 10-fold stratified cross-validation scheme. As in the previous experiment, the feature selection algorithm was set to identify the smallest miRNA subset sufficient to obtain a 90% accuracy. In addition, we compared our results with the 31-miRNA signature proposed by Romero et al. [35] to separate Triple-Negative Breast Cancer (TNBC) from other subtypes of BRCA using miRNA–mRNA integrative analysis in TNBC tumors based on the differential expressed transcripts. It is important to take into consideration that this 31-miRNA list considers noncirculating miRNAs that are not included in our method and could potentially access more information. Finally, we ran miRNet using the candidate miRNAs, as in Section 2.2.
3. Results
3.1. Cancer Type Classification
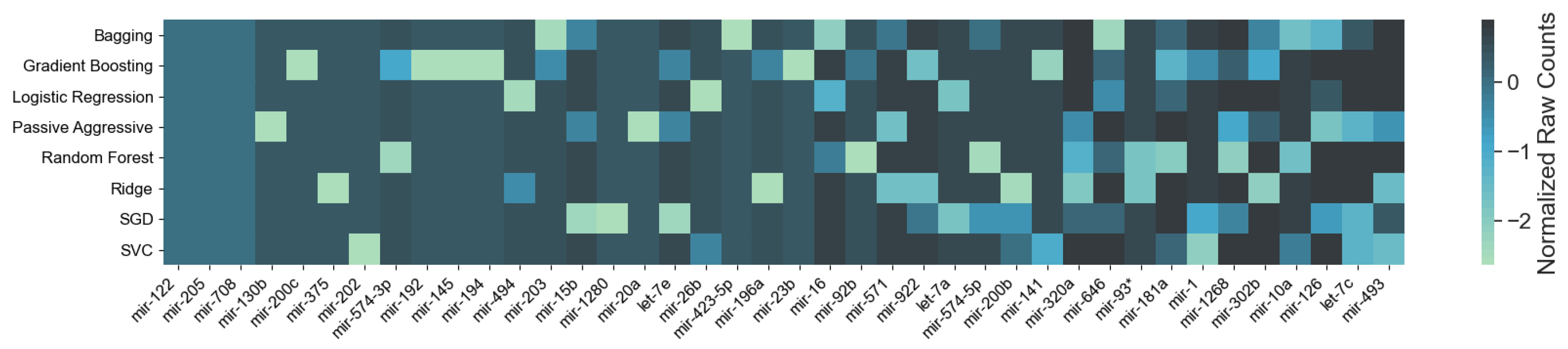
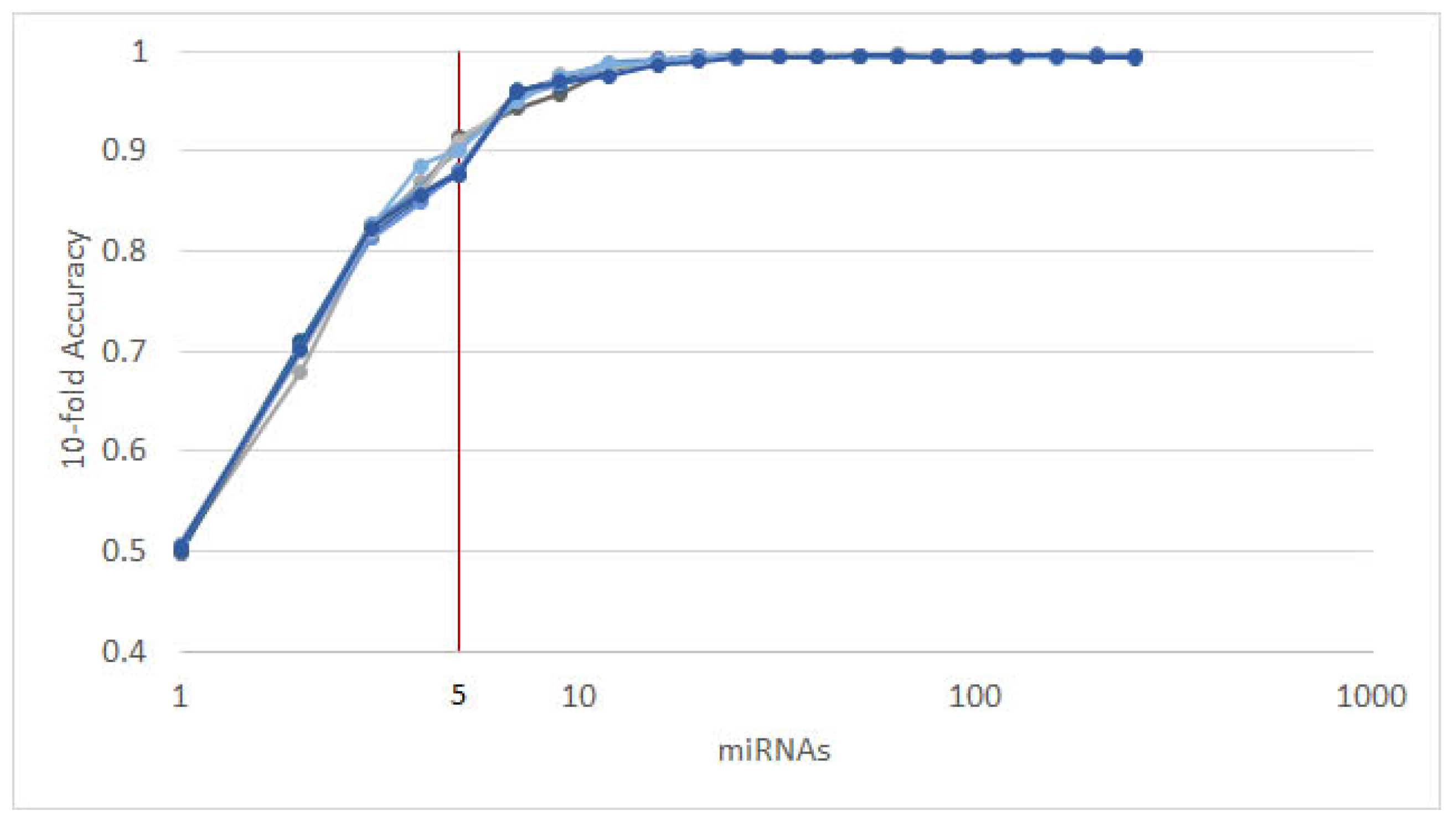
As mentioned in Section 2.1, each of the classifiers ranks the features differently. When applied to the 253 circulating miRNAs, the top features obtained by each classifier appear as in Figure 1. From visual inspection, it is easy to observe how each classifier ranks the features differently. Nevertheless, the features where the classifiers concur are the most important. From the feature selection algorithm, we reduced the original 253 miRNA to 5, while maintaining an average accuracy of 90% over the selected classifiers (Figure 2).
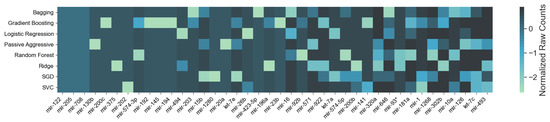
Figure 1.
Feature importance by classifier: On the horizontal axis, the top features are reported, following their ensemble ranking. The intensity of the color in each square represents the frequency of appearance of that particular feature in the 10 instances of the same classifier produced by cross-validation; the darker the color, the more frequent the appearance of that feature among the most important. It is noticeable how different classifiers rank features differently. For this figure, we report the top 42 features only for visualization purposes.
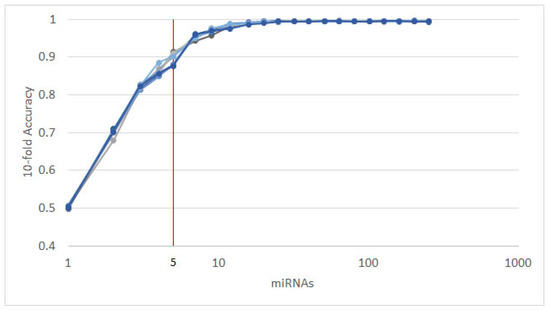
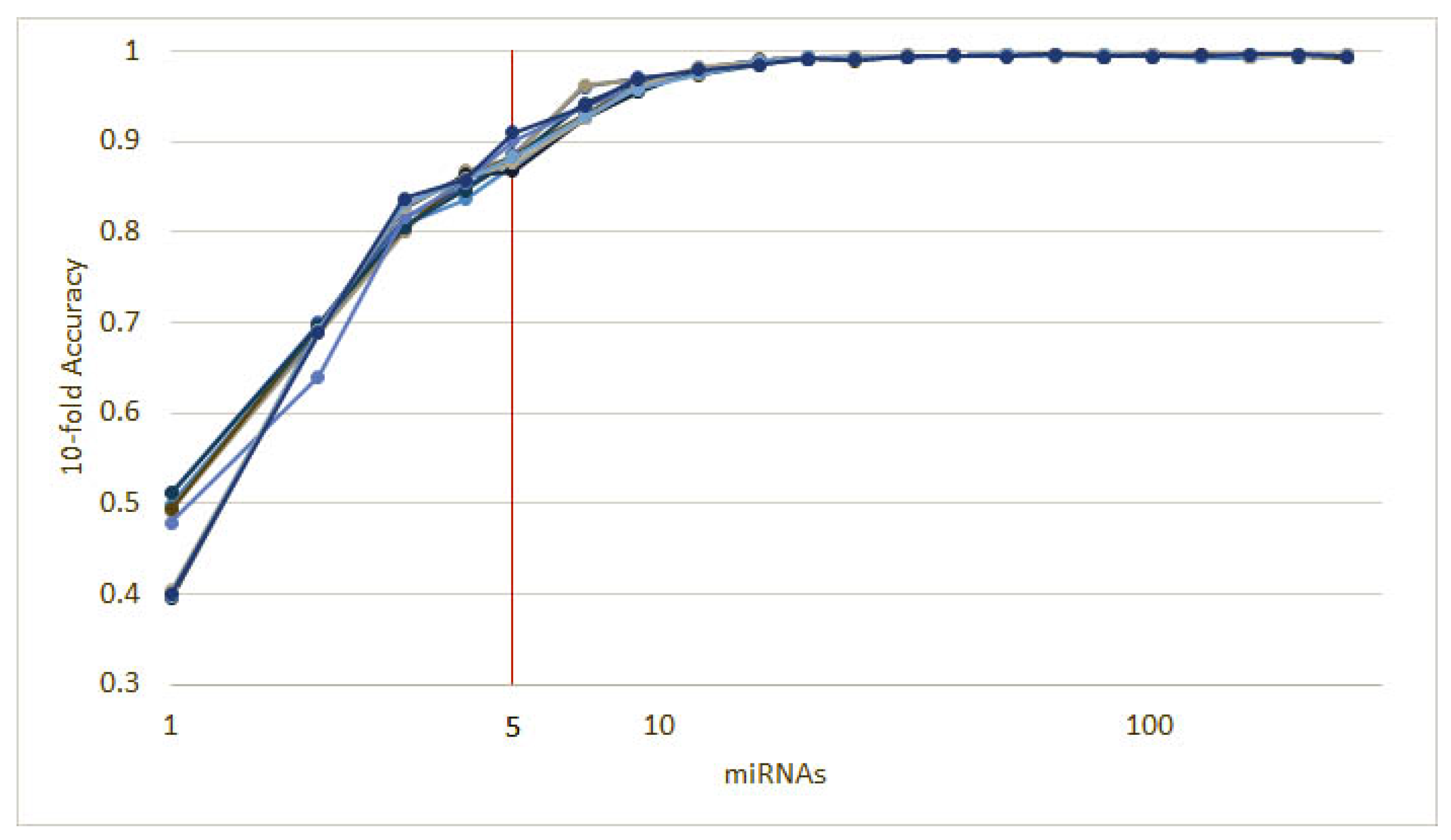
Figure 2.
The results of 10 runs of the recursive ensemble feature selection for cancer type classification: The x axis cuts at 5 variables, where all runs cross the 90% average accuracy stop parameter (we computed the subsequent values as a reference).
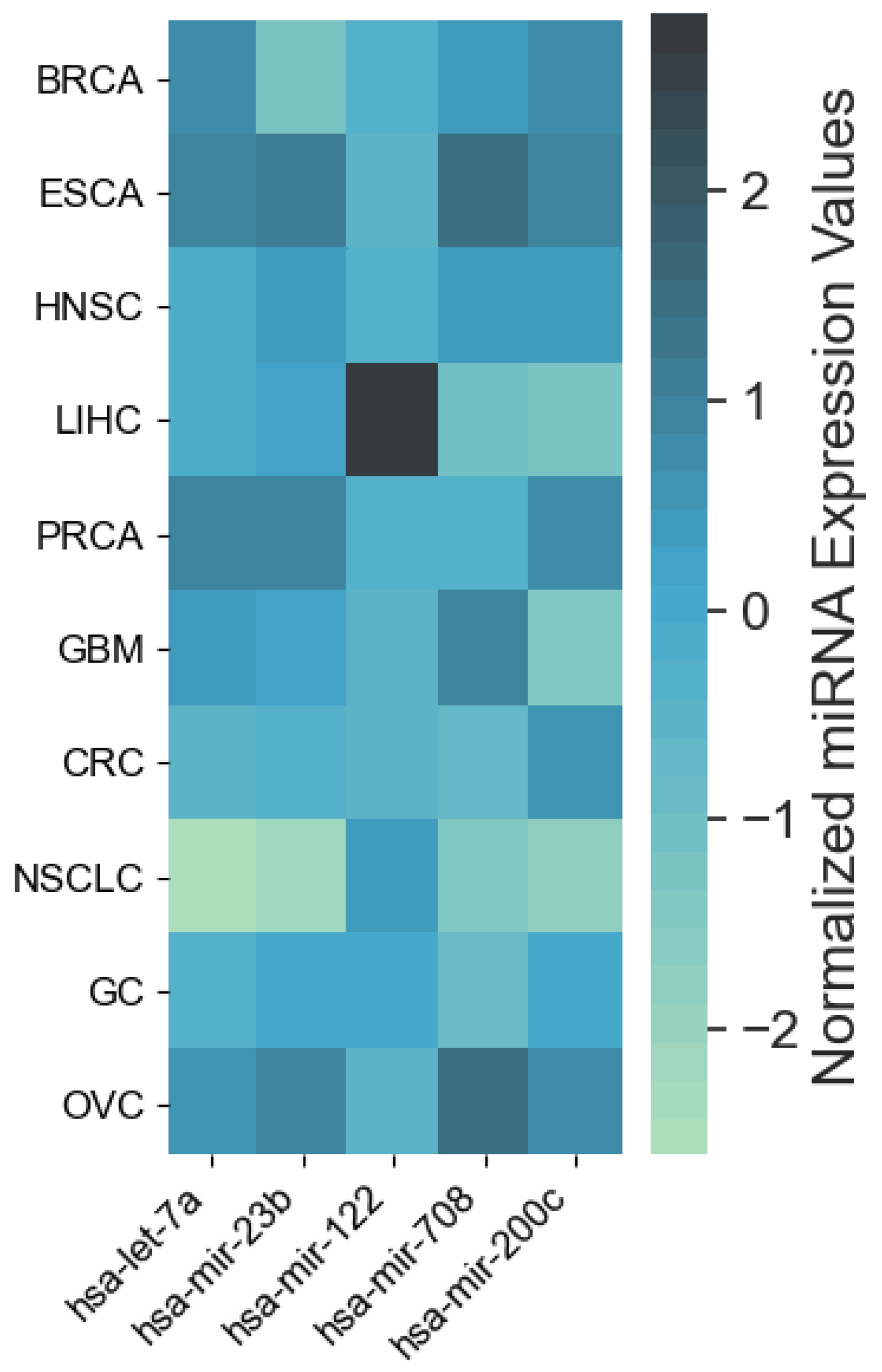
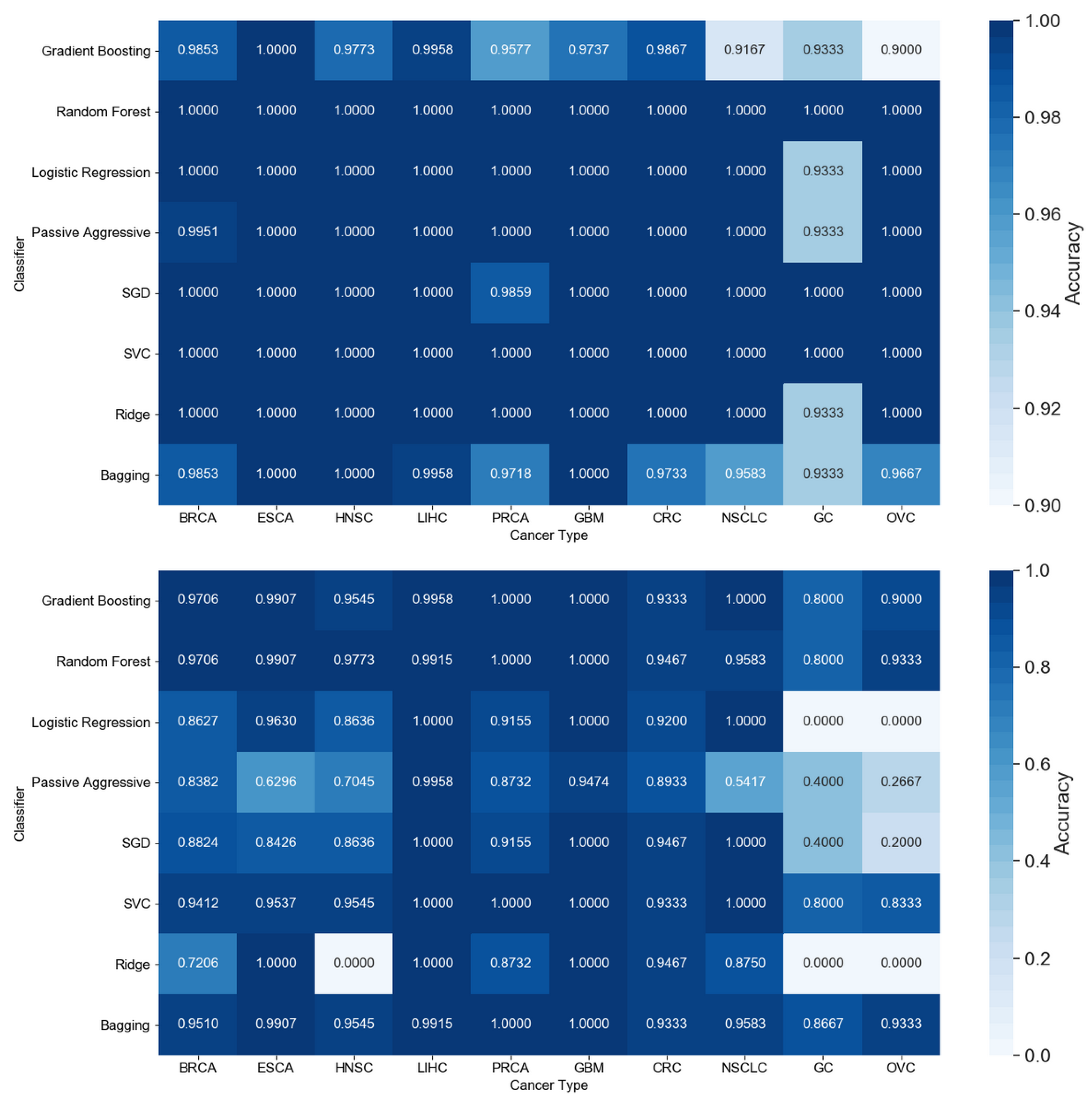
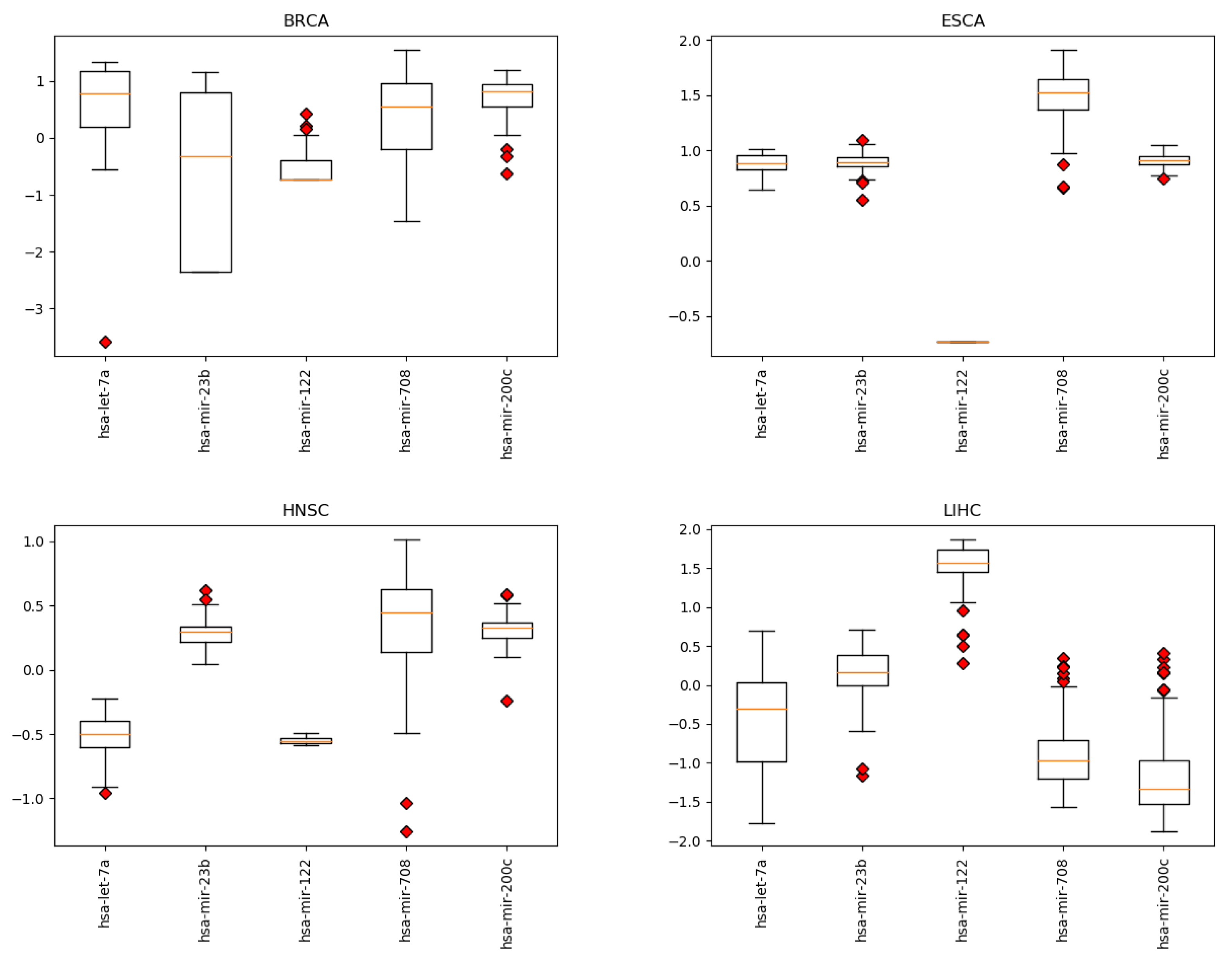
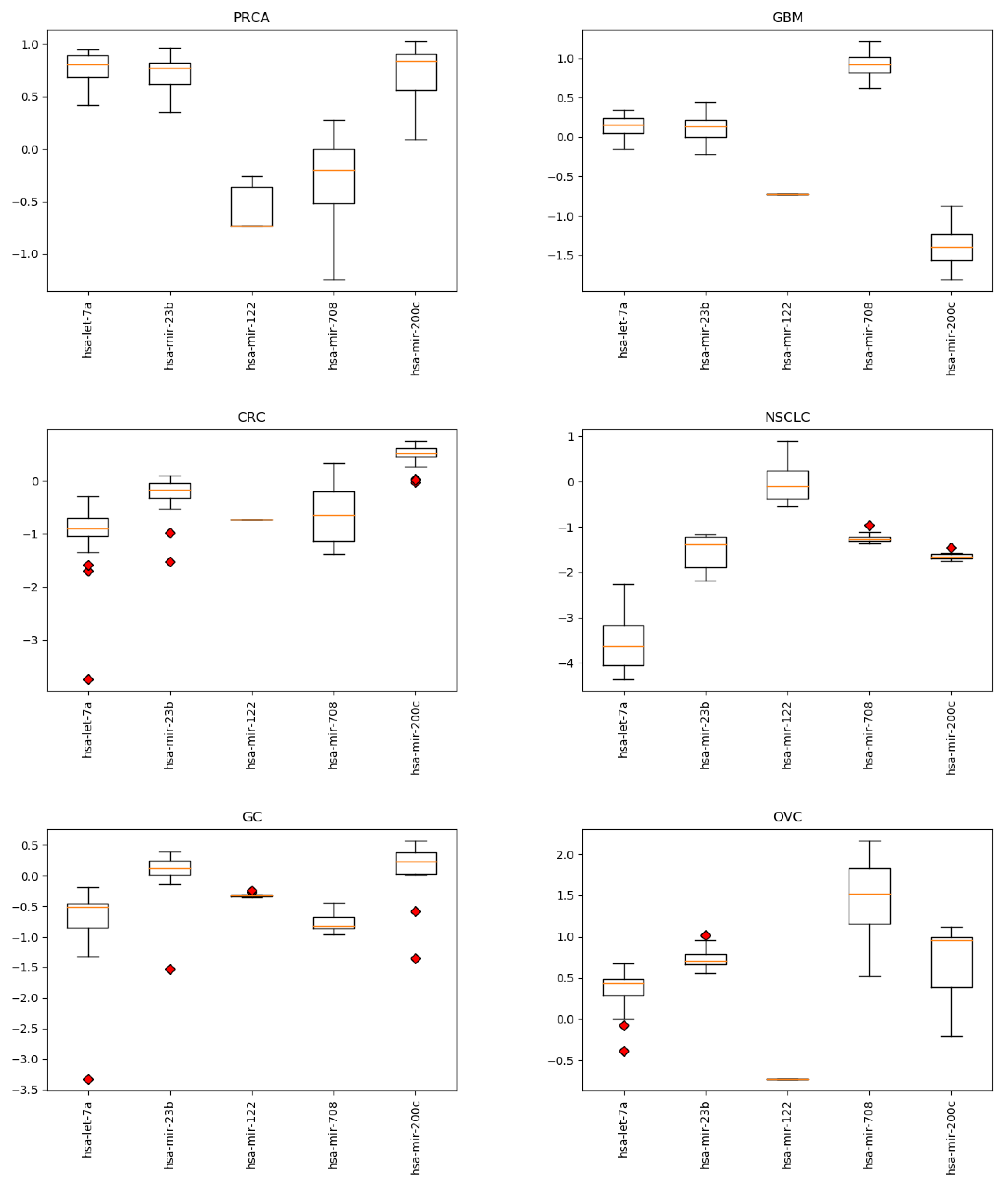
The resulting most significant 5 features uncovered by the presented algorithm are hsa-let-7a, hsa-miR-23b, hsa-miR-122, hsa-miR-708 and hsa-miR-200c, with seemingly different expression levels for each cancer type (Figure 3). For more detailed expression values by cancer type, see Appendix B. The classifiers gradient boosting, random forest, SVC and bagging seem to work in a satisfying way for all tumor types using only 5 miRNAs, whereas the rest have issues classifying the types of cancer HNSC, GC and OVC while having better performance when using the full 253 miRNAs, as shown in Figure 4. Interestingly, hsa-let-7 and hsa-miR-200c were also discovered by the homogeneous ensemble while GALGO’s performance seems to be less effective and has no miRNAs in common with our approach. From the comparison with GALGO and the homogeneous ensemble classifier with SVC, we can notice how the proposed heterogeneous ensemble classifier outperforms the other feature selection techniques in Table 2.
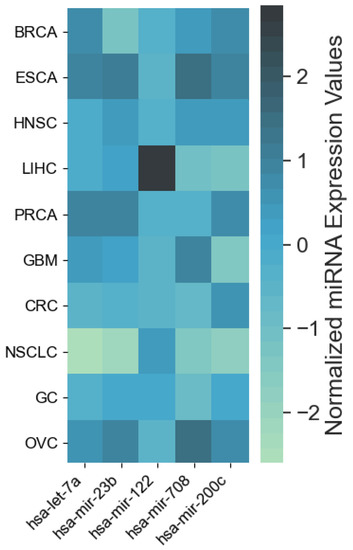
Figure 3.
Heatmap of average expression levels by cancer type for the 5 miRNAs identified by the proposed approach. Cancer types: breat (BRCA); esophageal (ESCA); head and neck squamous cell (HNSC); liver hepatocelluar (LIHC); prostate (PRCA); gliobastoma (GBM); colorectal (CRC); non-small-cell lung (NSCLC); gastric (GC); ovarian (OVC).
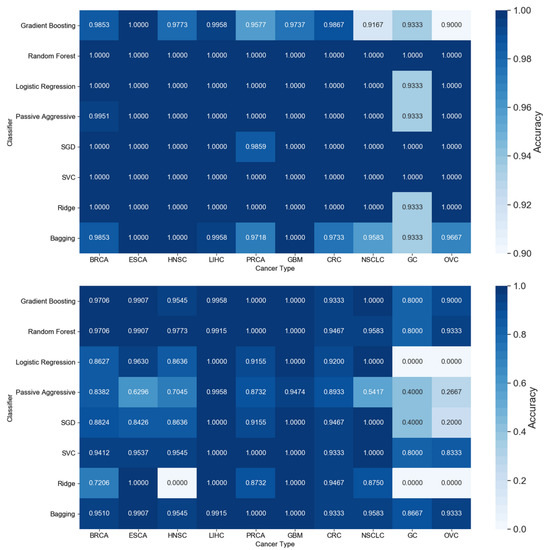
Figure 4.
Comparison of accuracy by classifier and tumor type for all 253 features (top) and the 5 features identified by the proposed approach (bottom). Cancer types: breast (BRCA); esophageal (ESCA); head and neck squamous cell (HNSC); liver hepatocellular (LIHC); prostate (PRCA); gliobastoma (GBM); colorectal (CRC); non-small-cell lun g (NSCLC); gastric (GC); ovarian (OVC).

Table 2.
Comparison of the results of the different feature selection algorithms, reduced from the initial 253 to 5 features to differentiate cancer types.
3.1.1. Numerical Validation
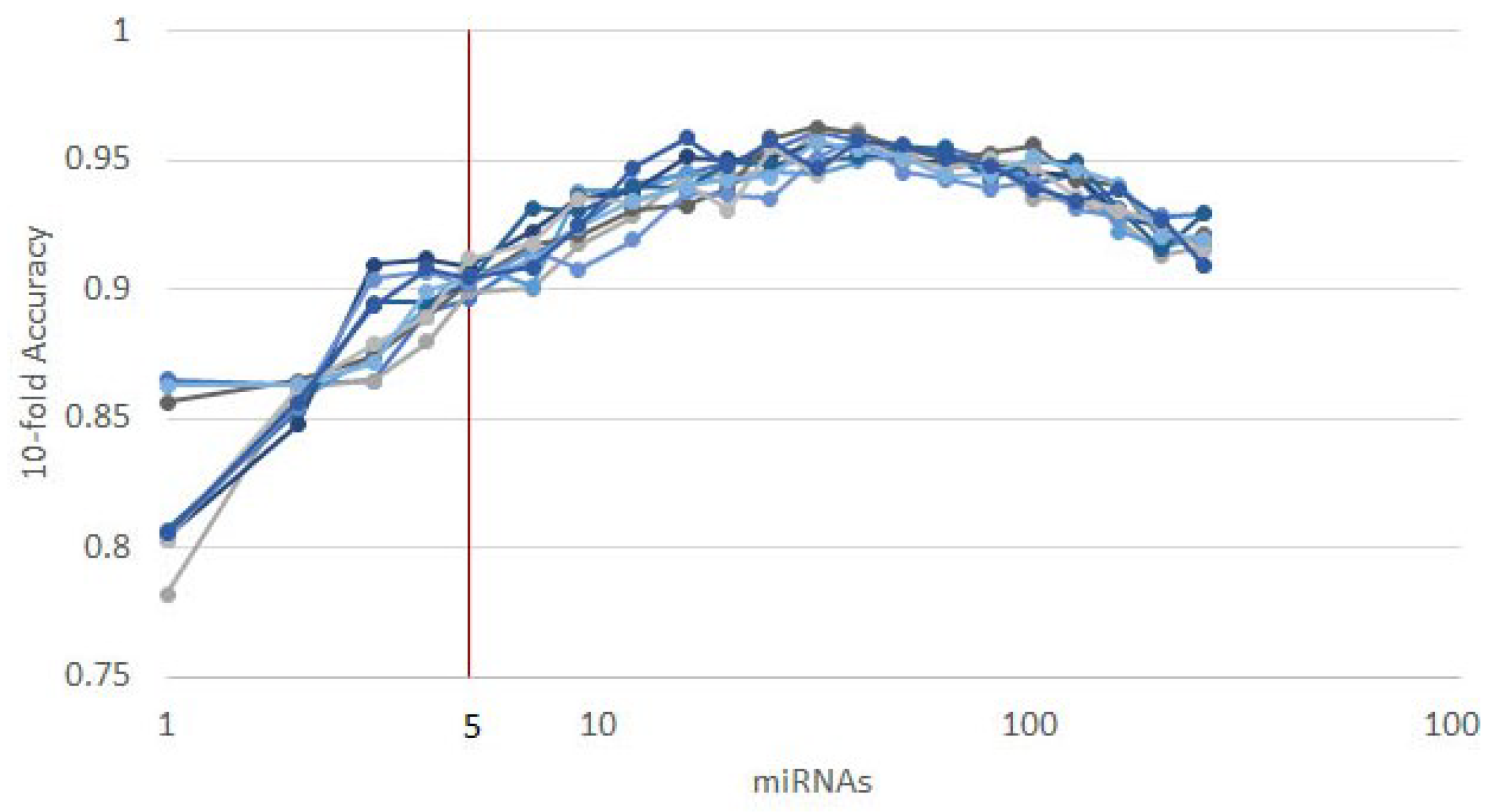
To further validate our methodology, we split the dataset described in Section 2.2 into a training (90%) and a validation (10%) subsets. Then, we ran 10 instances of the recursive ensemble feature selection algorithm with 90% of the data in a 10-fold cross-validation, which yields the curve in Figure 5.
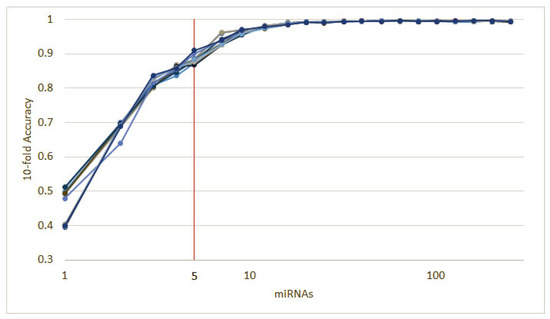
Figure 5.
Ten runs of the heterogeneous ensemble recursive selection algorithm. From the 10 runs, the minimum number of necessary miRNA to have an accuracy above 90% is 5: hsa-let-7a, hsa-miR-23b, hsa-miR-122, hsa-miR-708, and hsa-miR-200c.
Next, we selected the smallest signature that provided an accuracy of 90% or above, having as a result hsa-let-7a, hsa-mir-122, hsa-mir-200c, hsa-mir-708 and hsa-mir-23b, the same miRNAs identified in the previous experiment described in Section 3.1. Then, we tested this signature on the 10% subset, comparing against signatures identified by other approaches: homogeneous ensemble feature selection, GALGO, K-best univariate feature selection (using f-score) and 3 random selected subsets. In addition, we shuffled the test set’s labels to verify the proper workings of the classifiers (Table 3). Finally, we calculated the Matthews Correlation Coefficient (MCC) for all of the signatures and classifiers [49] (Table 4).

Table 3.
Accuracy on the 10% data for testing the feature selection algorithm: Results for the signatures found by the heterogenous recursive ensemble feature selection algorithm, the homogeneous recursive ensemble feature selection algorithm, K-Best feature selection algorithm using f-score as selection citeria, 3 random feature subsets and a random shuffle of the test labels.

Table 4.
Matthews correlation coefficient values for the 10% data left for testing the feature selection algorithm: The results for the heterogenous recursive ensemble feature selection algorithm, the homogeneous recursive ensemble feature selection algorithm, K-Best feature selection algorithm using f-score as selection citeria, 3 random feature subsets and a random shuffle of the test labels.
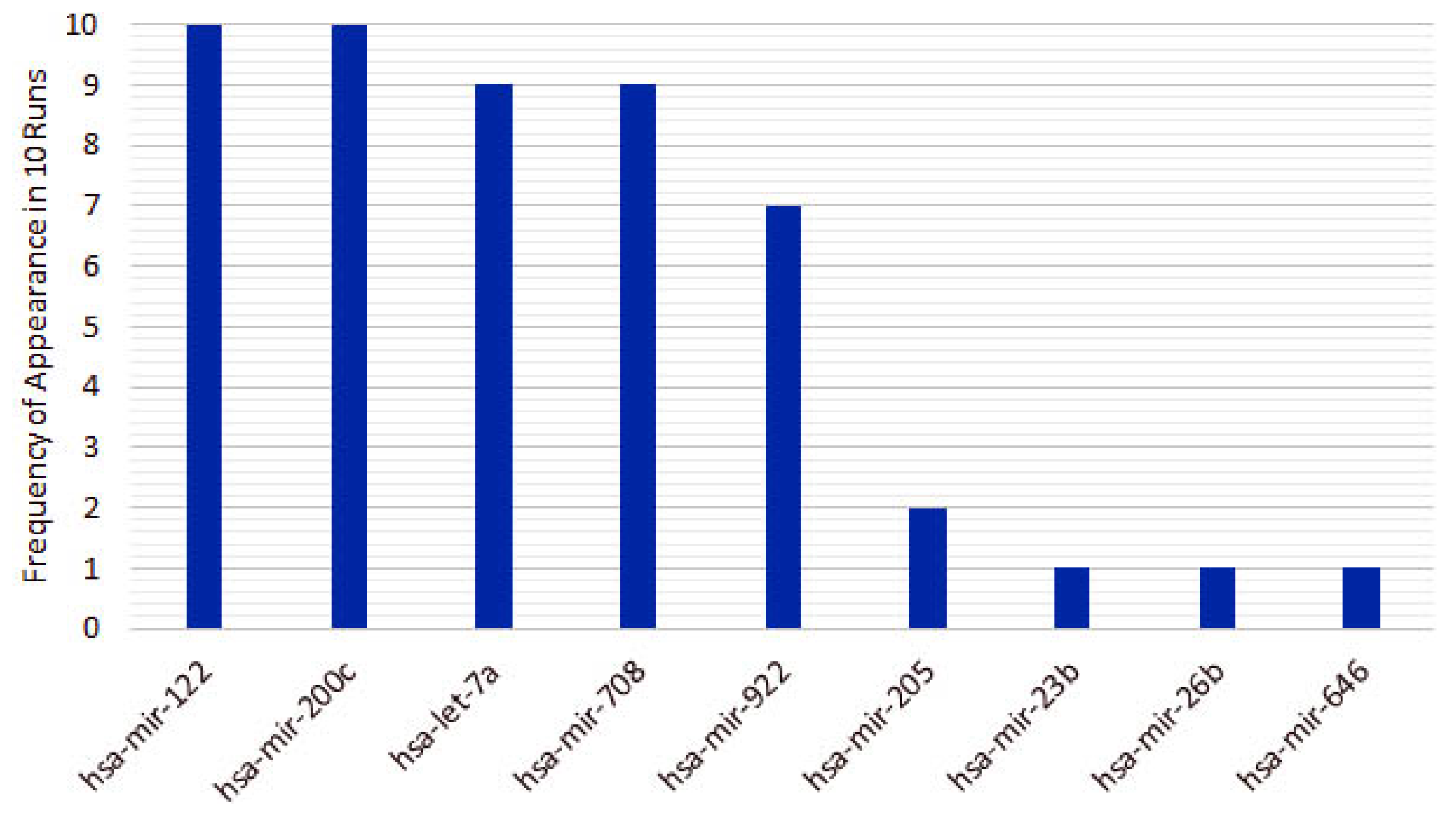
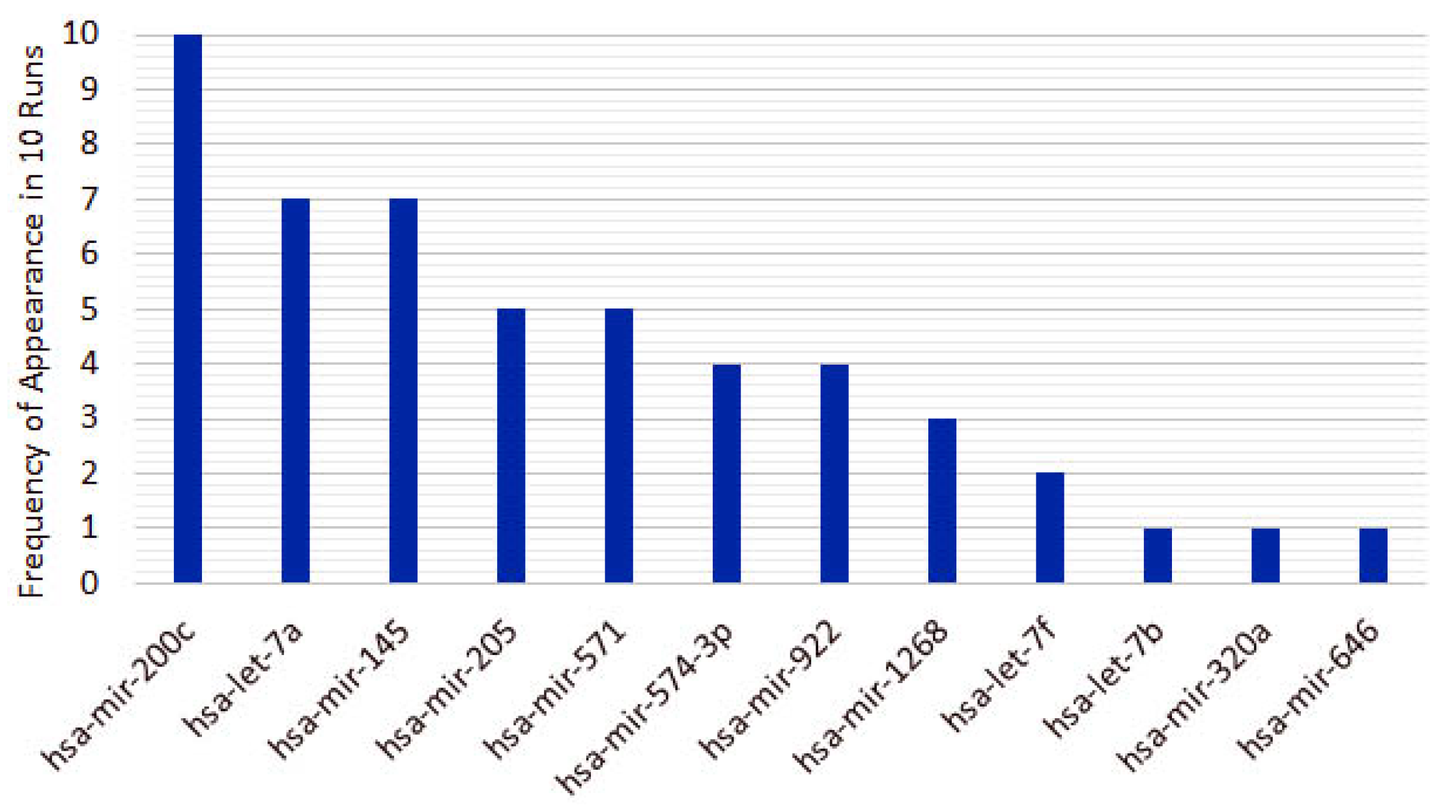
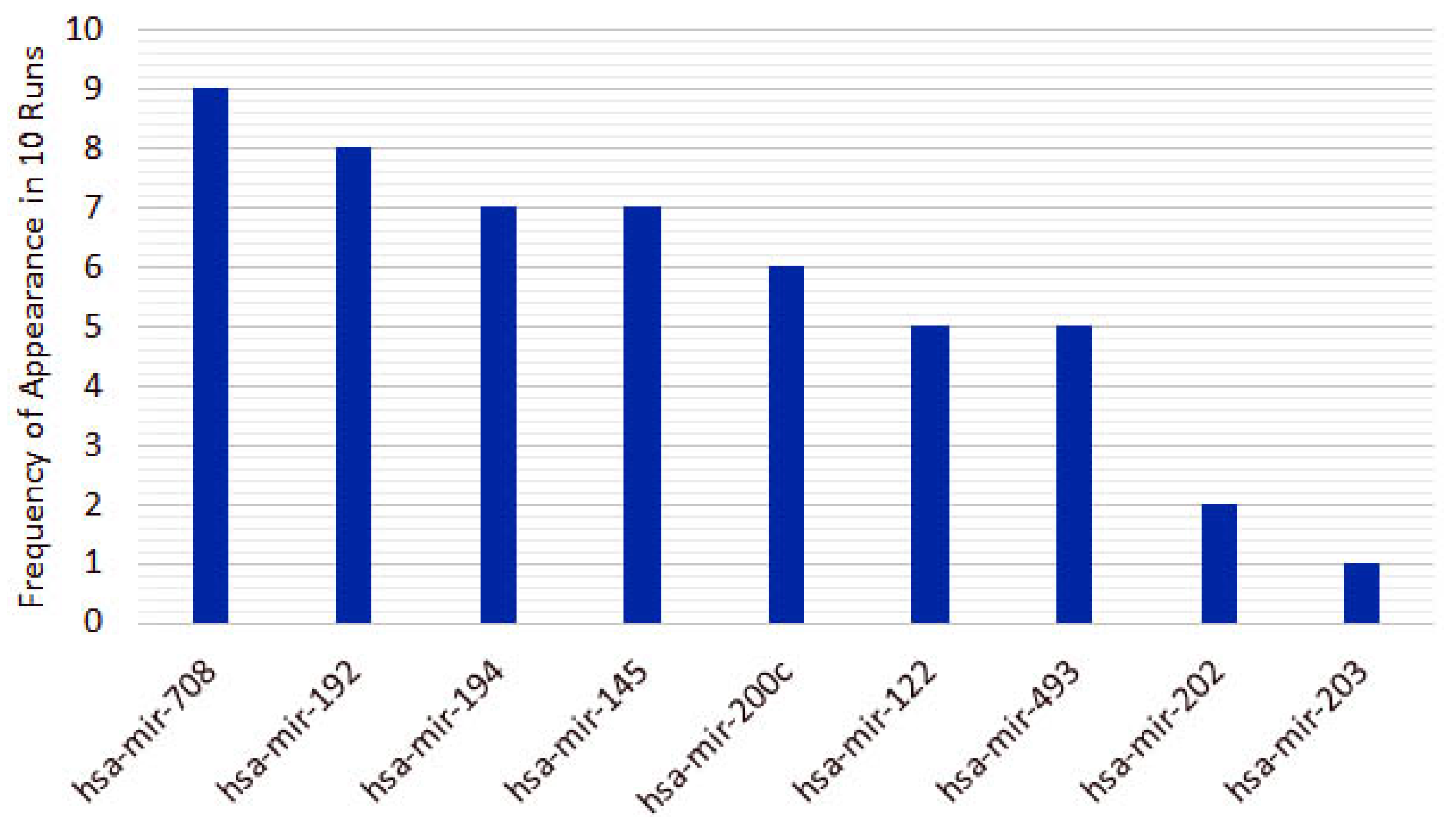
From the 10 instances, we then measured the frequency of appearance of miRNAs in the top 5 features for each run. From the original 253 features, only 10 features appear in the top 5 for the heterogeneous recursive ensemble feature selection algorithm, with the frequencies presented in Figure 6. We repeated the same procedure for 10 runs of the homogeneous ensemble feature selection algorithm (feature frequency presented in Figure 7) and GALGO (feature frequency presented in Figure 8). The variability of the output signature for each algorithm reflected that the average and standard deviations for accuracy and MCC for the proposed heterogeneous recursive ensemble feature selection algorithm are better than the homogeneous recursive ensemble feature selection algorithm and GALGO (see Table 5).
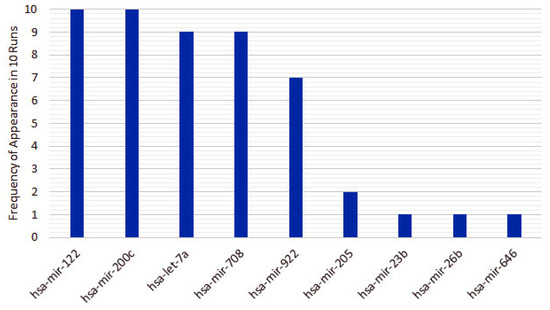
Figure 6.
Ten recurrent features in the 5-feature signature for the heterogeneous ensemble feature selection algorithm.
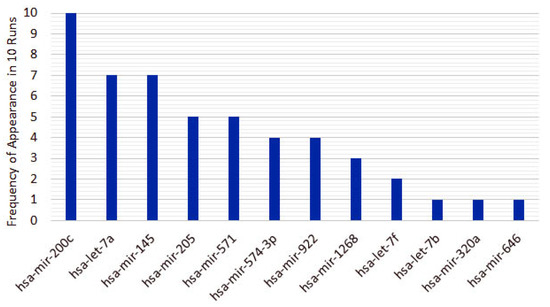
Figure 7.
Twelve recurrent features in the 5-feature signature for the homogeneous ensemble feature selection algorithm.
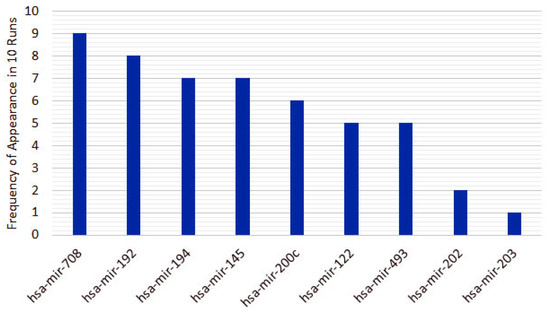
Figure 8.
Nine recurrent features in the 5-feature signature for the GALGO.

Table 5.
and for accuracy and MCC over 10 runs using the top 5 features, for each algorithm.
3.1.2. Pathway Analysis
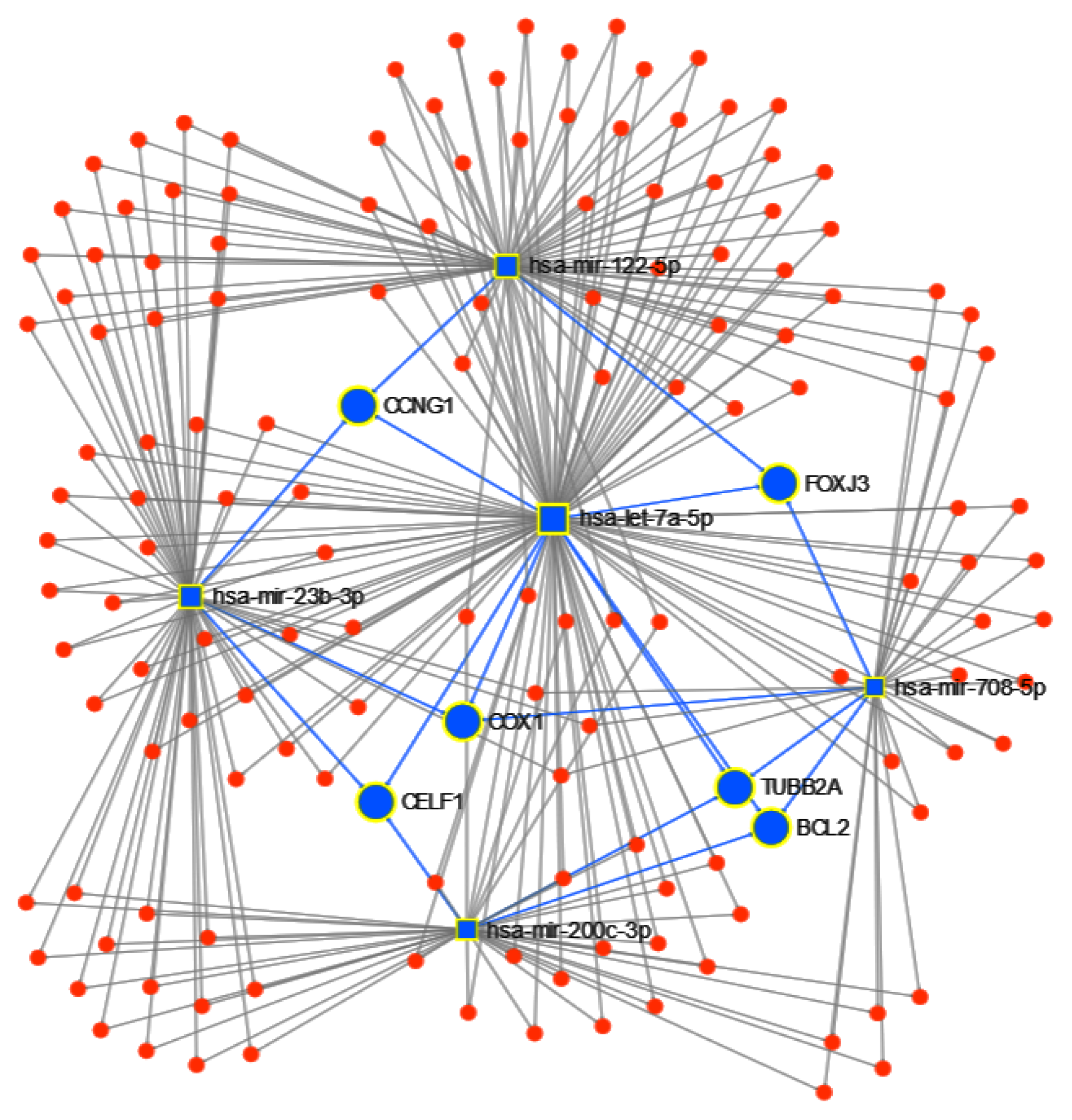
Next, using the 5 candidate miRNAs identified by the proposed approach to separate the tumor type, we ran miRNet to identify the targeted genes, obtaining a total of 1732 genes. After we apply a 0.05 betweenness filter, we reduced the list to 156 genes. From these genes, BCL2, CCNG1, COX1, TUBB2A, CELF1 and FOXJ3 are targeted by at least 3 of the 5 miRNAs (Figure 9). Finally, using the function explorer of miRNet, we performed a functional enrichment analysis with a hypergeometric test of the genes from the KEGG database and GO:BP. In Table 6 and Table 7, we show the results of the top 10 functional enrichment analyses for KEGG and GO:BP respectively. The first result in KEGG is the P53 signaling pathway. The P53 protein is a tumor suppressor protein, and it is involved in several anticancer mechanisms [50]. In the GO:BP database, the first result is the cellular response to stress, with 44 of the genes in the pathway; cellular stress is a component of the P53-mediated tumor suppression [51].
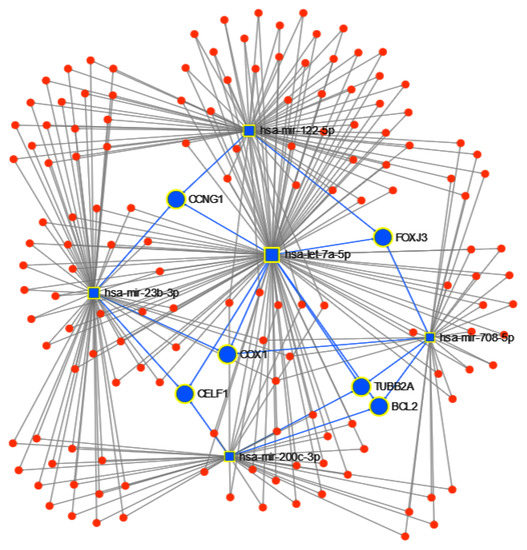
Figure 9.
miRNET targeted genes analysis, showing genes targeted by at least 3 of the 5 miRNAs to classify cancer type: BCL2, CCNG1, COX1, TUBB2A, CELF1 and FOXJ3.

Table 6.
Top 10 miRNet enrichment analyses in the Kyoto Encyclopedia of Genes and Genomes (KEGG) dataset for miRNAs hsa-miR-122, hsa-let-7a, hsa-miR-23b, hsa-miR-708 and hsa-miR-200c.

Table 7.
Top 10 miRNet enrichment analyses in the Gene Ontology-Biological Process (GO:BP) dataset for miRNAs hsa-miR-122, hsa-let-7a, hsa-miR-23b, hsa-miR-708 and hsa-miR-200c.
3.2. Triple-Negative Breast Cancer Classification
We ran the heterogeneous ensemble algorithm 10 times, identifying 5 meaningful miRNA features for separating triple-negative BRCA from the other subtypes (Figure 10). The resulting miRNAs are hsa-miR-378*, hsa-miR-221, hsa-miR-342-3p, hsa-miR-630 and hsa-miR-145. The corresponding expression levels for the identified miRNAs in TNBC and non-TNBC are reported in Figure 11.
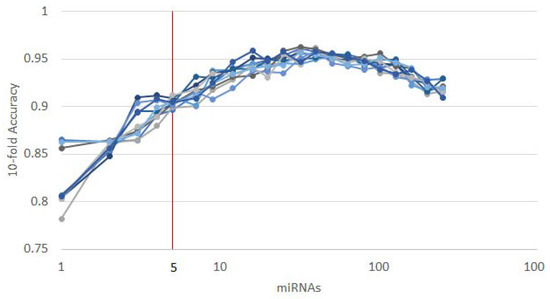
Figure 10.
Results of 10 runs of the recursive ensemble feature selection for the TNBC discrimination example: The x axis cuts at 5 variables, which is where most evaluations cross the average 0.90 accuracy stop parameter.
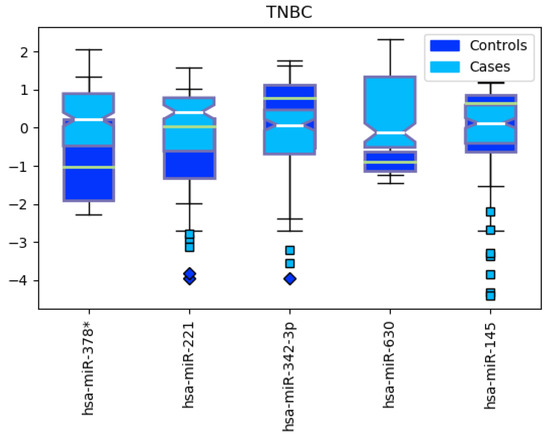
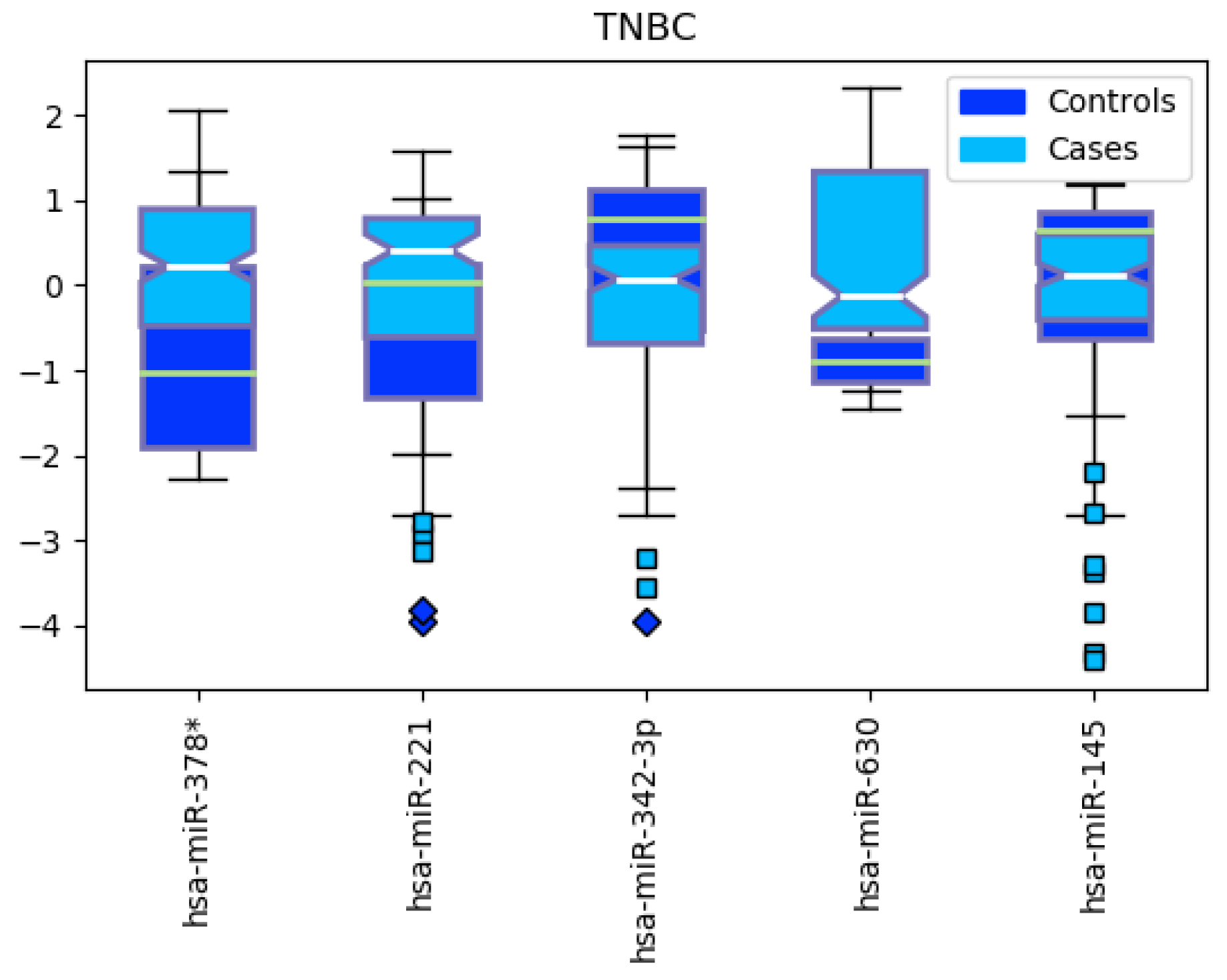
Figure 11.
Boxplot for the expression levels between Triple Negative Breast Cancer (TNBC, cases) and other subtypes (controls).
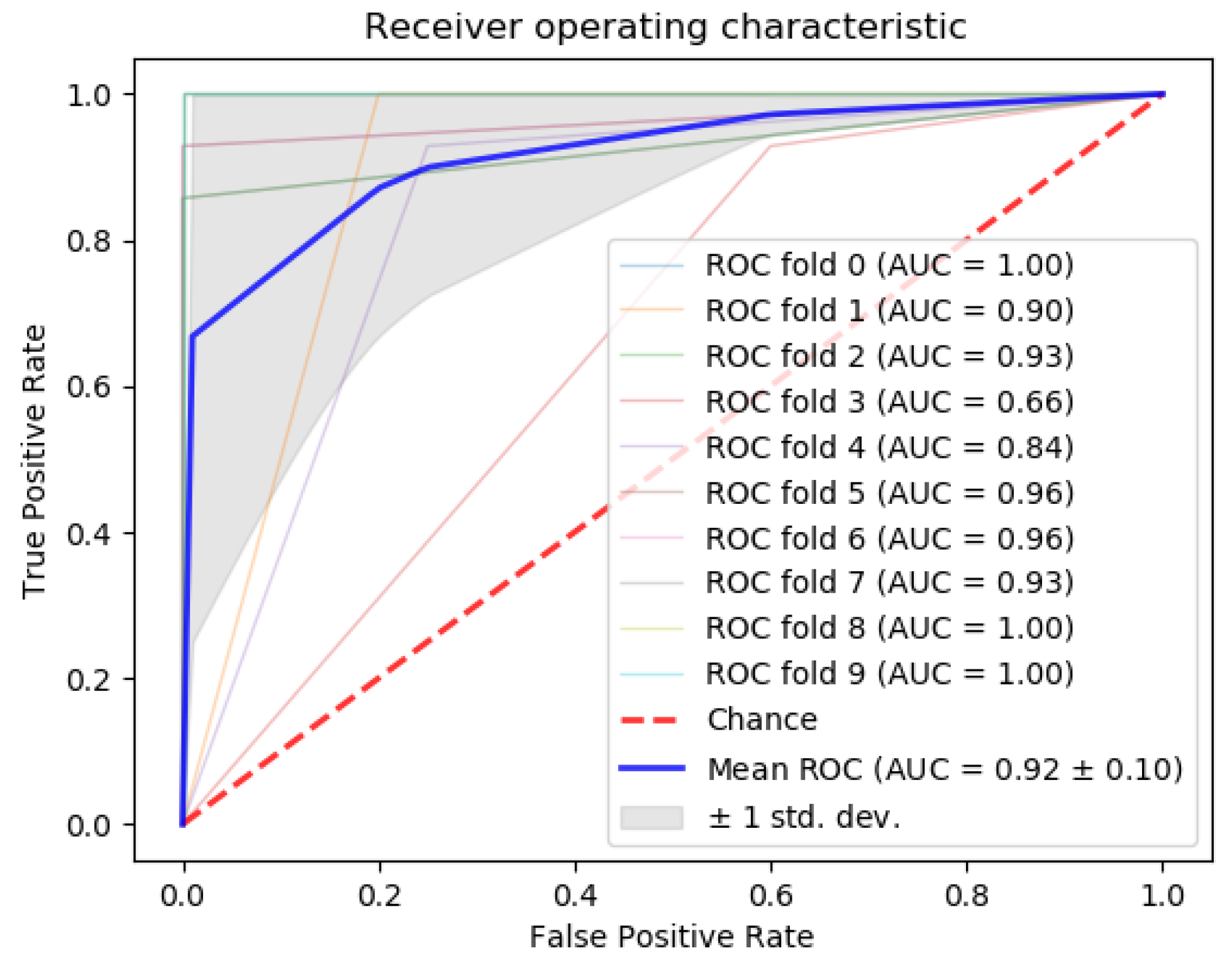
Next, we compared the accuracy between classifiers using all 253 miRNAs in the dataset, our 5-miRNA signature, and the 31-miRNA signature proposed by Romero et al. for distinguishing TNBC from other cancers (Table 8). From the results, our algorithm outperforms the 31-miRNA signature. In addition, the area under the curve (AUC) of the results (Figure 12) calculated with the gradient boosting classifier is above 90%. This is considered outstanding results following the guidelines in [52,53] for clinical use of algorithmic methodologies.

Table 8.
Accuracy comparison for all classifiers, using all 253 features, the 5-miRNA signature found by the proposed approach, and the 31-miRNA signature from Romero et al. for separating triple-negative from the rest of the BRCA subtypes.
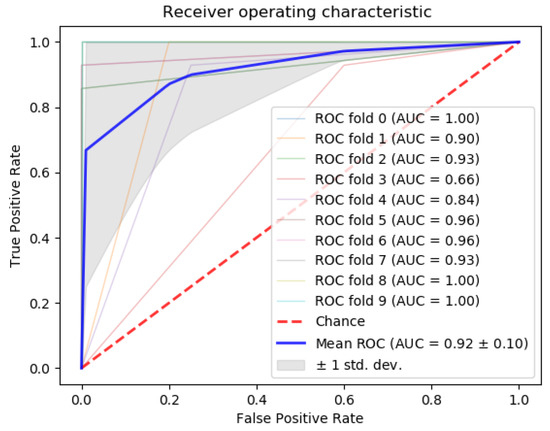
Figure 12.
ROC curve using the gradient boosting classifier to separate Triple Negative Breast Cancer (TNBC) from the rest of the breast cancer subtypes.
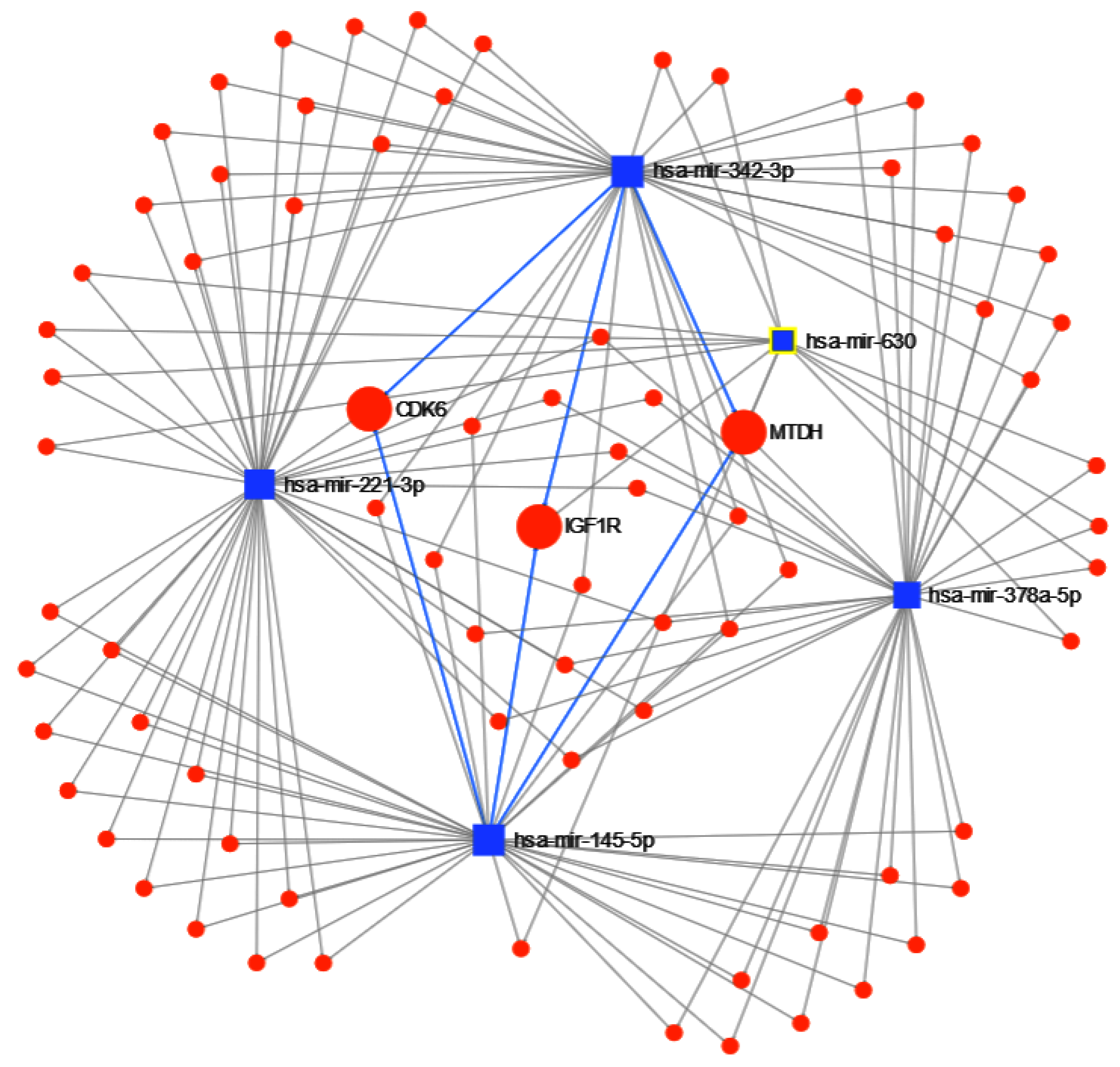
Finally, the results of miRNet found 1294 genes targeted by the 5 miRNAs, with 79 having at least 2 miRNAs in common. From those 79, metastasis gene metadherin-positive (MTDH) is targeted by 4 miRNAs, while type 1 insulin-like growth factor receptor-positive (IGF1R) and cyclin-dependent kinase 6-positive (CDK6) are targeted by 3; see Figure 13. From the enrichment analysis, the most important functional pathway in the KEGG database (Table 9) is the p53 signaling pathway (the same identified in the previous experiments for separating cancer types), and in GO:BP (Table 10), the negative regulation of cell proliferation, with 12 of the 79 genes followed by regulation of cell proliferation and just cell proliferation. These results show an important involvement of cell proliferation in TNBC.
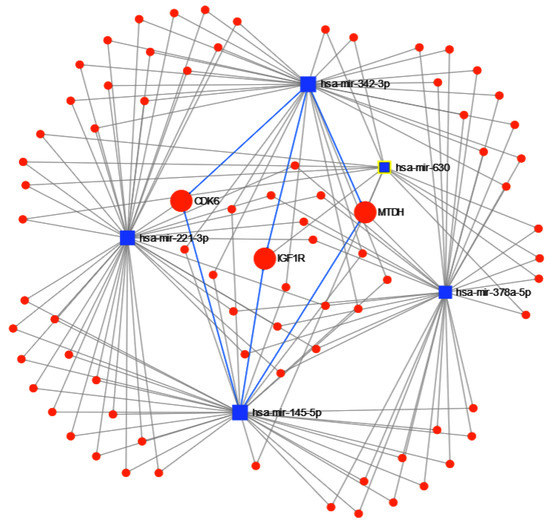
Figure 13.
miRNET targeted genes analysis, showing genes targeted by at least 3 of the 5 miRNAs to separate Triple Negative Breast Cancer (TNBC) from other breast cancer subtypes: metastasis gene metadherin-positive (MTDH), type 1 insulin-like growth factor receptor-positive (IGF1R) and cyclin-dependent kinase 6-positive (CDK6).

Table 9.
Top 10 miRNet enrichment analysis results for miRNAs hsa-miR-378*, hsa-miR-221, hsa-miR-342-3p, hsa-miR-630 and hsa-miR-145 using the KEGG database.

Table 10.
Top 10 miRNet enrichment analysis results for miRNAs hsa-miR-378*, hsa-miR-221, hsa-miR-342-3p, hsa-miR-630 and hsa-miR-145 using the GO:BP database.
4. Discussion
In this section, we perform an analysis of the candidate miRNAs identified by the proposed feature selection method, using the available literature in cancer studies.
4.1. miRNAs from Cancer Type Classification
The five circulating miRNAs identified by our method as the most informative for cancer type classification are hsa-miR-122, hsa-let-7a, hsa-miR-23b, hsa-miR-708 and hsa-miR-200c.
hsa-miR-122 is a 22-nucleotide RNA molecule that plays an important role in liver functions [54]. It is related to regulation of cholesterol, fatty acid metabolism, and hepatocytes differentiation. Evidence indicates that hsa-miR-122 acts like a tumor suppressor, and its depletion is related to liver inflammation and hepatocellular cancer in mice [54,55]. In breast cancer, hsa-miR-122 has different expression patterns according to the subtype [56]. In addition, miR-122 promotes aggression and epithelial-mesenchymal transition in TNBC [57] and cell survival in radio-resistance cells [58]. High plasma miR-122 levels have been detected in AFP-producing gastric cancer [59].
The let-7 miRNAs show a high evolutionary conservation between organisms. Vertebrates have multiple let-7 isoforms and play an important role in development and tumor suppression [60]. hsa-let-7a is a member of the family and shows a downregulated expression in many tumor types like breast cancer [61,62], lung adenocarcinoma [63] and gastric cancer [64].
hsa-miR-23b is known to target tumor suppressor and cancer promoter genes. hsa-miR-23b is dis-regulated in proliferation, invasion, migration, apoptosis, autophagy and cell survival [65]. As a circulating biomarker, hsa-miR-23b is downregulated in colon cancer measured in plasma [66]. In contrast, it is upregulated in gastric cancer [67], lung cancer [68] and pancreatic cancer [69].
hsa-miR-708, also known as miR-708-5p, is a microRNA encoded within an intron of the ODZ4 gene. It can be found in different tissues with varying expression patterns like reproductive, secretory, muscle, gastrointestinal, nervous and lung [70]. hsa-miR-708 acts as a tumor suppressor or oncogene according to the cancer type. It has been associated with poor prognosis in lung adenocarcinoma [71] and carcinogenesis in colon [72] and bladder [73]. On the other hand, normal levels of hsa-miR-708 decrease cell growth and invasion and increase apoptosis in renal cancer cells [74].
hsa-miR-200c has been identified in lung, gastric, breast, ovarian and endometrial cancer with different expression patterns related to prognosis, aggressiveness and chemoresistance [75,76]. Moreover, hsa-miR-200c is involved in signaling cascades such as TGF-, PI3K/Akt, Notch, VEGF, and NF-B making it a candidate biomarker in cancer [77].
The result with the smallest p-value from the enrichment analysis in the KEGG dataset identified a strong relationship between the P53 signaling pathway and hsa-miR-122, hsa-let-7a, hsa-miR-23b, hsa-miR-708 and hsa-miR-200c. P53 is an important tumor suppressor that regulates the expression of many genes and is one of the most common mutated genes in cancer. Many miRNAs work as direct and indirect mediators of the P53 activity and the components of its pathway [78,79]. Moreover, the normal function of this tumor suppressor helps the maturation of some miRNAs with growth-suppressing function [80].
On the other hand, the first result in the enrichment analysis in the GO:BP dataset was cellular stress response. In normal cells, there is a balance between the activation of survival and cell death pathways, according to the type and duration of stress [81]. Cancer cells develop molecular mechanisms that facilitate their adaptation to different conditions like oxidative, metabolic, mechanical and genotoxic stresses, avoiding the restriction of the growth and increasing cell proliferation [82]. Importantly, miRNAs have the capacity to modify the stress response in cancer by making cells more susceptible or resistant to chemotherapy [83]. These findings prove that miRNAs play an important role in cancer biology and could be used as powerful circulating biomarkers for diagnosis and prognosis in human malignancies.
4.2. miRNAs from Triple-Negative Breast Cancer Classification
From our analysis, we selected 5 candidate miRNAs that are the most informative to separate cancer TNBC from the other subtypes in BRCA: hsa-miR-378*, hsa-miR-221, hsa-miR-342-3p, hsa-miR-630 and hsa-miR-145. All of them had already been shown to have potential as circulating cancer biomarkers in cancer studies, e.g., [84,85,86,87,88,89,90,91,92].
hsa-miR-378* is considered an onco-miRNA for its relationship with tumor growth and cell renewal. It is associated with the progression of breast cancer and the Warburg effect. Furthermore, hsa-miR-378* is capable of discriminating between breast cancer patients and controls [84,85].
Evidence indicates that hsa-miR-221 is upregulated and that its expression is related to proliferative pathways [93,94]. Several studies have linked the microRNA cluster 221/222 with chemoresistance. The miR-221/222 expression participates in the clinically aggressive basal-like subtype [95] and tamoxifen resistance in ER-positive breast cancer cells [87,88]. Furthermore, this cluster interfers with ER expression [87] and miR-221/222 knockdown induces growth arrest and apoptosis in cells exposed to tamoxifen [88].
On the other hand, hsa-miR-342-3p expression correlates with ER mRNA expression and its downregulation is related to tamoxifen resistance. hsa-miR-342-3p plays an important role in the therapy response of tamoxifen in ER-positive breast cancer [86,89]. Moreover, hsa-miR-342-3p activity affects some metabolic pathways like lactate and glucose fluxes in TNBC [35].
hsa-miR-630 is considerably suppressed in BRCA [90]. From in vitro experiments in which hsa-miR-630 mimics was transfected into MDA-MB-231 cells, it could be detected that the expression of hsa-miR-630 was decreased. miR-630 was also capable in inhibiting MDA-MB-231 cell migration and invasion targeting SOX4-3’-UT. Additionally, the SOX4 overexpression plasmid was transfected to further confirm that hsa-miR-630 played its role by downregulation [96].
Finally, hsa-miR-145 acts as a tumor suppressor through the inhibition of different proteins like ERBB3 and RTKN [91,92]. Additionally, hsa-miR-145 cooperates with P53 and has a proapoptotic effect in patients with breast cancer [97].
The miRNet enrichment analysis yields that P53 and the negative regulation of cell proliferation were the signaling pathways mostly involved with these miRNAs. Furthermore, the MTDH, IGF1R and CDK6 genes are targeted by at least 3 of the 5 miRNAs used to identify TNBC. Zare et al. [98] described the interplay of methilation patterns in miRNAs and the epithelial-mesenchymal transition. They identified that some genes like MTDH, IGF1R and CDK6 can be affected by miRNAs and can modify cellular processes in breast cancer.
5. Conclusions
miRNAs are known to play important roles in cellular biology processes such as differentiation, proliferation and apoptosis. Several research lines suggest that miRNAs are involved in different aspects of cancer, and recent studies indicate that there is potential in using their expression profiles as molecular signatures in clinically relevant settings.
miRBase (v22.1) consists of 1917 stem-loop sequences and 2657 mature sequences for human miRNAs [99]. Only some of these 2657 mature sequences are circulating miRNAs, and from that quantity, only 253 can be measured in blood, urine, plasma, serum or saliva (excluding pancreatic juice and cerebrospinal fluid). In this paper, our aim has been to reduce as much as possible the number of miRNAs necessary to classify cancer tumor types and to identify TNBC in BRCA. Our proposed approach consists in applying a recursive ensemble feature selection algorithm to reduce the original 253 miRNAs to 5 for each case study considered while, at the same time, ensuring high-quality classification (>90% mean classification accuracy over all the ensemble). It is important to state that our results are based on readily available clinical studies from the GEO repository.
Using the identified 5-miRNA signature for tumor classification, the classifier random forest obtains a mean accuracy of 97.61% in a 10-fold cross-validation, providing both results of high quality and a compact, human-interpretable list of miRNAs. When compared to the state-of-the-art in feature selection, our methodology was proven to be better than GALGO and ensemble-based approaches with an homogeneous topology, with a significant statistical difference ( using a standard Welch’s T-test). In the TNBC example, the signature obtained by our methodology outperforms the 31-miRNA signature from [35]. These remarkable results stem from the use of machine learning algorithms which are able to consider the influence of groups of features (in this case miRNAs) at the same time, while previous works only employed univariate statistics. Such an outcome is consistent with Mootha et al. [100], which makes the case for considering gene sets instead of individual genes. This methodology can be applied in other problems, such as differentiating between tumors with and without metastasis (Appendix C), and it is not restricted to only miRNAs but can also be used in mRNA data. In contrast to other methods such as Saha et al. [20], it is not limited by the number of variables (Appendix D).
This analysis is a first step towards assembling new approaches for cancer detection using circulating miRNAs, as measuring only 5 miRNAs levels is not only easier but also more resistant to measurement errors than to try and measure all available miRNAs levels. This research line might help the development of new concepts for prevention, secondary prevention and novel therapies.
Author Contributions
A.L.-R. suggested the problem, wrote, built the datasets and coded. M.M.-A. and L.M.-M. helped with the writing, miRNAs concepts and the bibliographic analysis. A.S., A.D.K. and J.G. wrote and conceived the validation, performed the comparison to other methods and secured funding. A.T. wrote and coded. All authors have read and agreed to the published version of the manuscript.
Funding
The funding for the study was provided by the division of Pharmacology, Department of Pharmaceutical Sciences, Faculty of Science, Utrecht University. This work was carried out on the Dutch national e-infrastructure with the support of SURF Cooperative.
Conflicts of Interest
J.G. is employed by the company Danone Nutricia Research. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Sample Availability
The datasets generated/analyzed for this study can be found in the GitHub repository https://github.com/steppenwolf0/circulating.
Appendix A. Circulating miRNAs
In Table A1, we present the list of all 253 circulating miRNAs identified in the dataset, using an analysis of the available literature.

Table A1.
List of all circulating miRNAs.
Table A1.
List of all circulating miRNAs.
| let-7a | miR-140-3p | miR-19b | miR-335 | miR-513a-3p |
| let-7a* | miR-141 | miR-200a | miR-338-3p | miR-516b |
| let-7b | miR-142-3p | miR-200b | miR-338-5p | miR-518b |
| let-7c | miR-143 | miR-200c | miR-339-3p | miR-520a-3p |
| let-7d | miR-144 | miR-202 | miR-339-5p | miR-548b-5p |
| let-7e | miR-145 | miR-203 | miR-340* | miR-557 |
| let-7f | miR-146a | miR-205 | miR-342-3p | miR-564 |
| let-7g | miR-146b-3p | miR-206 | miR-345 | miR-566 |
| let-7i | miR-146b-5p | miR-20a | miR-346 | miR-571 |
| miR-1 | miR-148a | miR-20b | miR-34a | miR-574-3p |
| miR-100 | miR-148b | miR-21 | miR-34b | miR-574-5p |
| miR-101 | miR-150 | miR-210 | miR-361-3p | miR-587 |
| miR-106b | miR-150* | miR-212 | miR-365 | miR-589 |
| miR-107 | miR-151-5p | miR-214 | miR-371-5p | miR-595 |
| miR-10a | miR-152 | miR-215 | miR-372 | miR-601 |
| miR-10b | miR-155 | miR-218 | miR-373 | miR-616* |
| miR-1182 | miR-15a | miR-22 | miR-375 | miR-618 |
| miR-122 | miR-15b | miR-221 | miR-376a | miR-622 |
| miR-122* | miR-15b* | miR-222 | miR-376c | miR-625 |
| miR-1224-5p | miR-16 | miR-223 | miR-377 | miR-625* |
| miR-1229 | miR-16-2* | miR-23a | miR-378 | miR-628-3p |
| miR-1231 | miR-17 | miR-23b | miR-378* | miR-629 |
| miR-1245 | miR-181a | miR-24 | miR-379 | miR-630 |
| miR-1246 | miR-181a-2* | miR-25 | miR-382 | miR-638 |
| miR-1254 | miR-181b | miR-26a | miR-409-3p | miR-646 |
| miR-125b | miR-181d | miR-26b | miR-409-5p | miR-650 |
| miR-125b-2* | miR-182 | miR-27a | miR-410 | miR-652 |
| miR-126 | miR-1825 | miR-27b | miR-411 | miR-654-3p |
| miR-1260 | miR-183 | miR-296-5p | miR-421 | miR-656 |
| miR-1268 | miR-184 | miR-298 | miR-423-5p | miR-668 |
| miR-127-3p | miR-185 | miR-299-5p | miR-425 | miR-675 |
| miR-1275 | miR-186 | miR-29a | miR-425* | miR-7 |
| miR-128 | miR-187 | miR-29b | miR-429 | miR-708 |
| miR-1280 | miR-187* | miR-29c | miR-431 | miR-744 |
| miR-1284 | miR-18a | miR-301a | miR-431* | miR-744* |
| miR-1285 | miR-18b | miR-302b | miR-432 | miR-760 |
| miR-1288 | miR-18b* | miR-30a | miR-451 | miR-874 |
| miR-1290 | miR-190b | miR-30b | miR-452 | miR-885-5p |
| miR-1295 | miR-191 | miR-30c | miR-454 | miR-922 |
| miR-129-5p | miR-192 | miR-30c-1* | miR-454* | miR-92a |
| miR-1304 | miR-193a-3p | miR-30d | miR-483-3p | miR-92a-2* |
| miR-130a* | miR-193b | miR-30e | miR-483-5p | miR-92b |
| miR-130b | miR-194 | miR-31 | miR-484 | miR-93 |
| miR-1323 | miR-195 | miR-32 | miR-486-3p | miR-93* |
| miR-133a | miR-196a | miR-320a | miR-486-5p | miR-936 |
| miR-133b | miR-196b | miR-320c | miR-487b | miR-939 |
| miR-134 | miR-197 | miR-320d | miR-493 | miR-942 |
| miR-138 | miR-198 | miR-324-3p | miR-494 | miR-99a |
| miR-138-2* | miR-199a-3p | miR-326 | miR-497 | miR-99b |
| miR-139-3p | miR-199a-5p | miR-328 | miR-502-5p | |
| miR-139-5p | miR-19a | miR-331-3p | miR-504 |
Appendix B. miRNA Levels
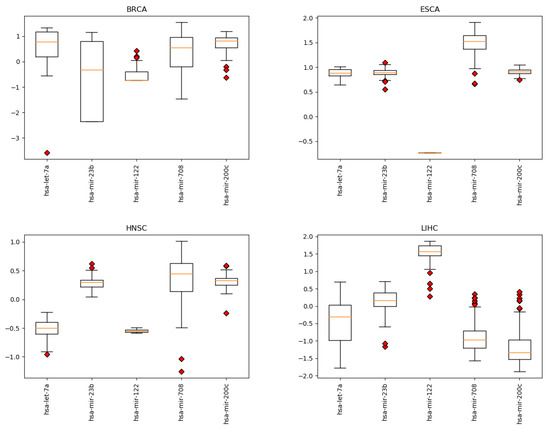
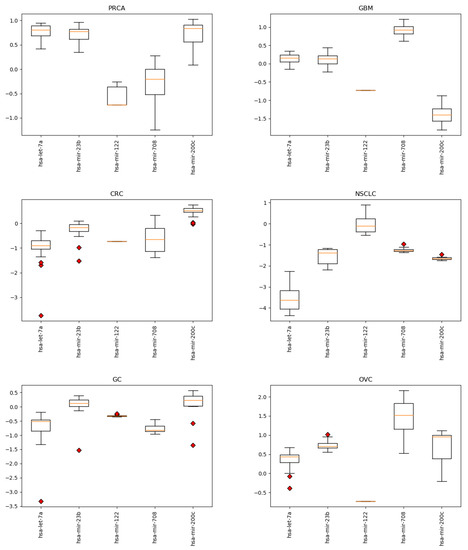
Figure A1.
Expression levels by cancer type for the 5 miRNAs identified by the proposed approach. Cancer types: breat (BRCA); esophageal (ESCA); head and neck squamous cell (HNSC); liver hepatocelluar (LIHC); prostate (PRCA); gliobastoma (GBM); colorectal (CRC); non-small-cell lung (NSCLC); gastric (GC); ovarian (OVC).
Figure A1.
Expression levels by cancer type for the 5 miRNAs identified by the proposed approach. Cancer types: breat (BRCA); esophageal (ESCA); head and neck squamous cell (HNSC); liver hepatocelluar (LIHC); prostate (PRCA); gliobastoma (GBM); colorectal (CRC); non-small-cell lung (NSCLC); gastric (GC); ovarian (OVC).
Appendix C. miRNA Levels in CRC with and without Metastasis
To provide evidence that the proposed methodology can be used not only to classify tumors but also more in general to answer specific questions related to tumors, where miRNAs can be informative, we applied our algorithm to dataset GSE53159 [101], separating metastasized tumors from those which are not. This dataset is composed of 32 samples and 16 colorectal cancer (CRC) samples with liver metastasis and 16 CRC samples without liver metastasis for platform GPL8786. After applying our method, we obtain a 4-miRNA signature, with the differentially expressed hsa-mir-486-3p, hsa-mir-21, hsa-mir-1285, hsa-miR-708 and hsa-mir-638. The final average accuracy is 0.9312, with 0.8625 MCC.
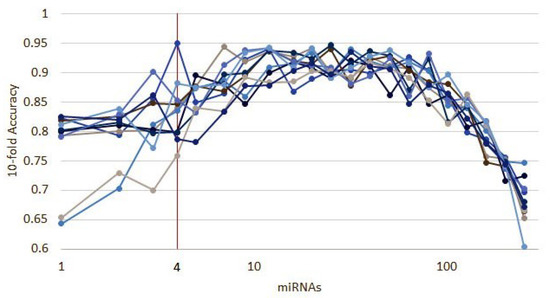
Figure A2.
Ten runs of the heterogeneous ensemble recursive selection algorithm: From the 10 runs, the highest accuracy is given by 4 miRNAs: hsa-mir-486-3p, hsa-mir-21, hsa-mir-1285, hsa-miR-708 and hsa-mir-638.
Figure A2.
Ten runs of the heterogeneous ensemble recursive selection algorithm: From the 10 runs, the highest accuracy is given by 4 miRNAs: hsa-mir-486-3p, hsa-mir-21, hsa-mir-1285, hsa-miR-708 and hsa-mir-638.
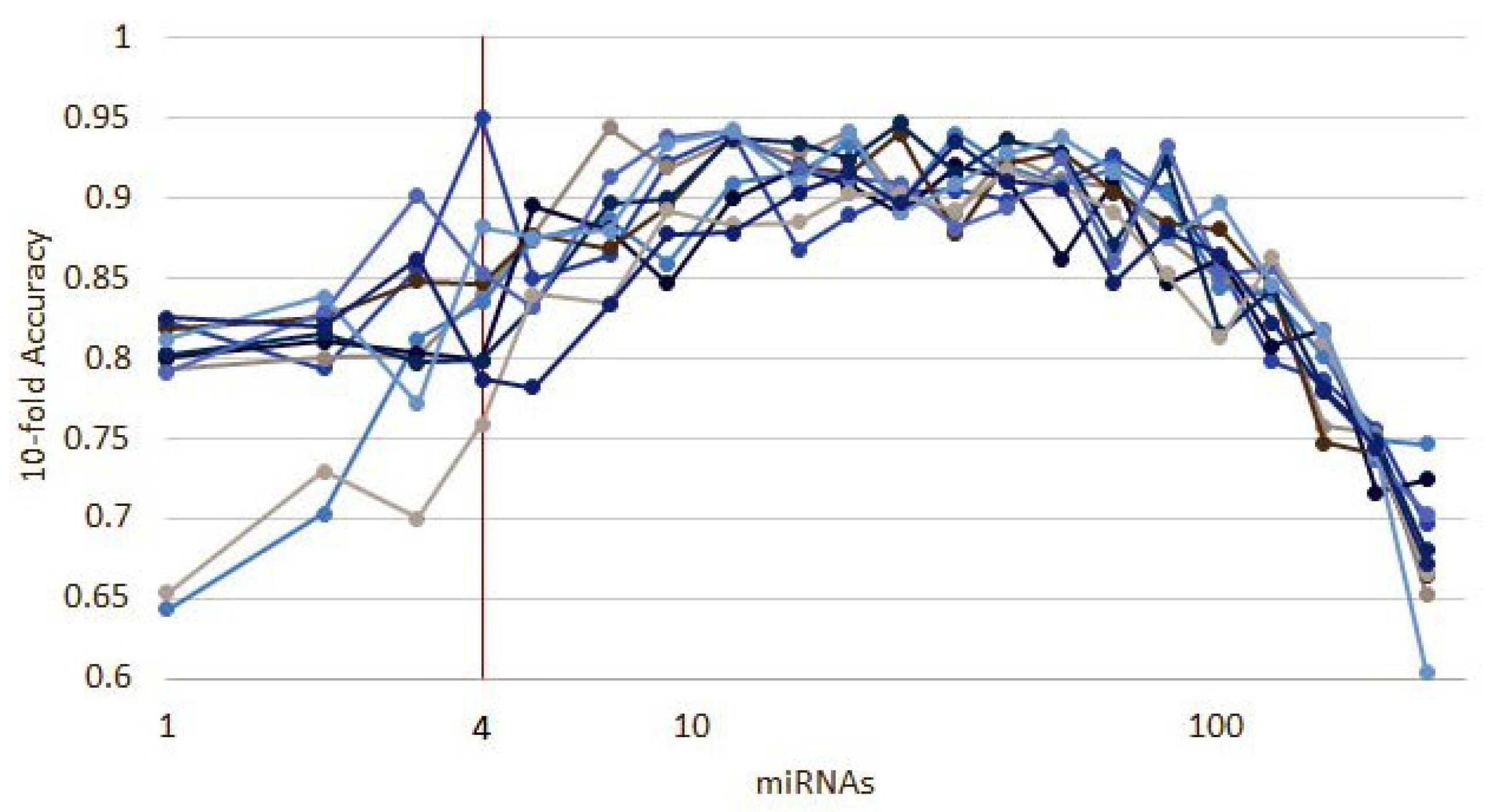
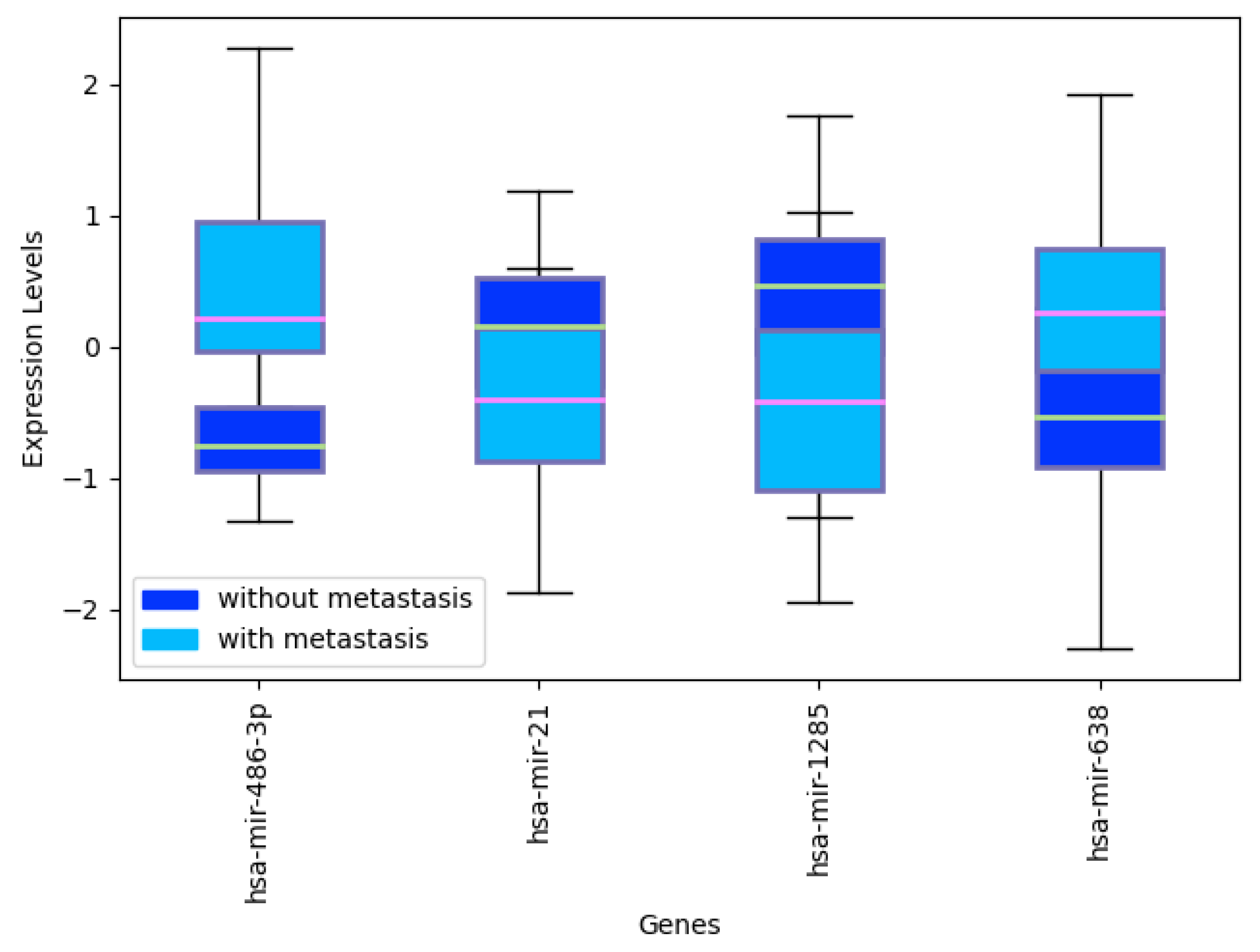
Figure A3.
Expression levels of selectedmiRNAs: hsa-mir-486-3p, hsa-mir-21, hsa-mir-1285, hsa-miR-708 and hsa-mir-638 in CRC samples with and without metastasis.
Figure A3.
Expression levels of selectedmiRNAs: hsa-mir-486-3p, hsa-mir-21, hsa-mir-1285, hsa-miR-708 and hsa-mir-638 in CRC samples with and without metastasis.
Appendix D. Recursive Ensemble Feature Selection in mRNA Data
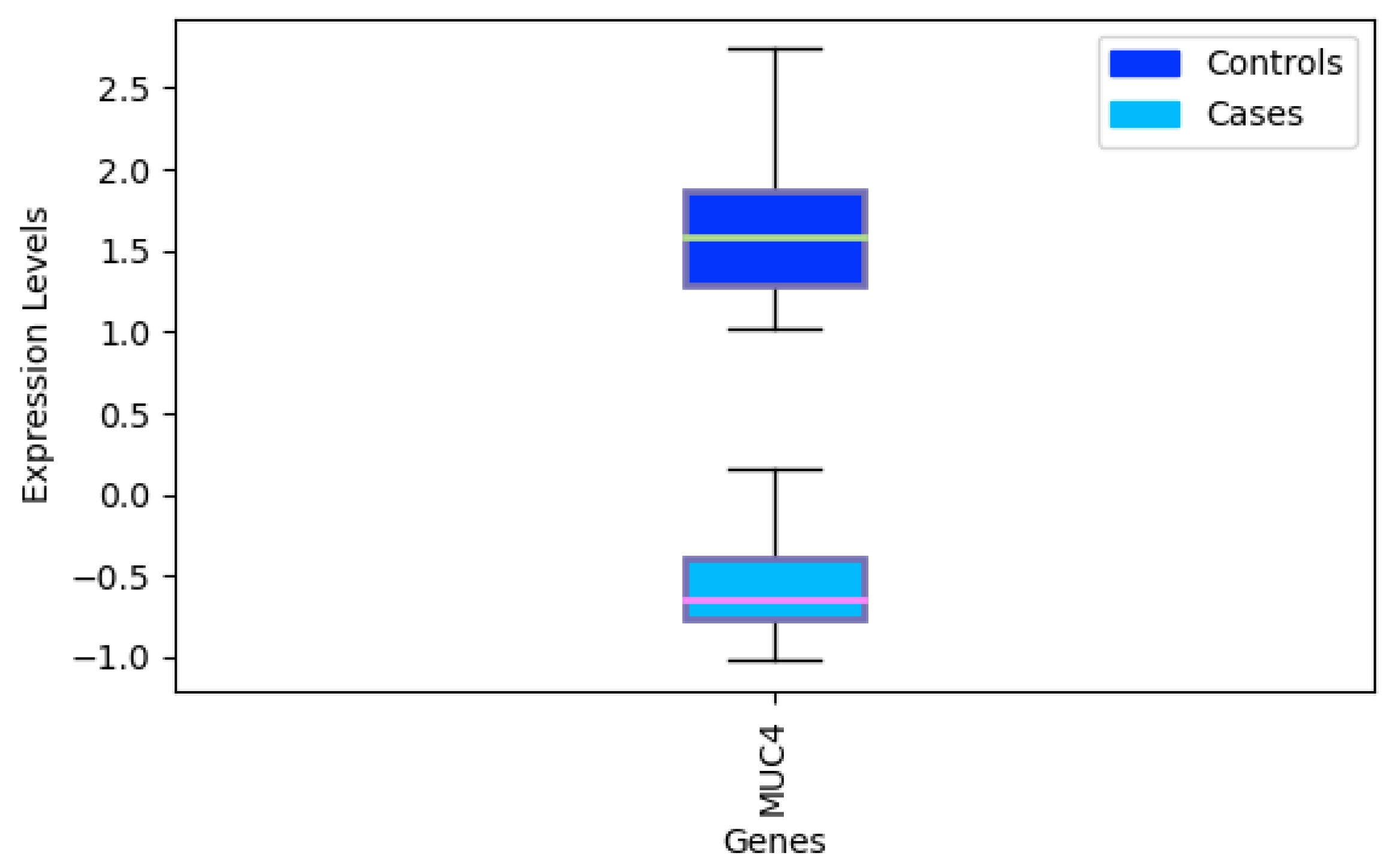
While other techniques such as [20] can also be effective to identify signatures for bioinformatic applications, they are usually limited to working with a few hundreds of features. In order to show how our algorithm can be effective even with a large number of features, we apply it to dataset GSE12452 [102] that contains 54,675 features related to messenger RNA (mRNA). This dataset is composed of 41 samples: 31 samples are nasopharyngeal tumor tissue and 10 are normal nasopharyngeal healthy controls for platform GPL570. After applying our methodology, we obtain a signature composed of just one gene, MUC4, differentially expressed to separate tumor and healthy tissue. This is consistent with studies that point out MUC4 as a cancer biomarker [103,104]. Overall, the signature identified has a global accuracy of 1.0, with 1.0 MCC.
Figure A4.
Ten runs of the heterogeneous ensemble recursive selection algorithm: From the 10 runs, the highest accuracy is given by MUC4 gene expression alone.
Figure A4.
Ten runs of the heterogeneous ensemble recursive selection algorithm: From the 10 runs, the highest accuracy is given by MUC4 gene expression alone.
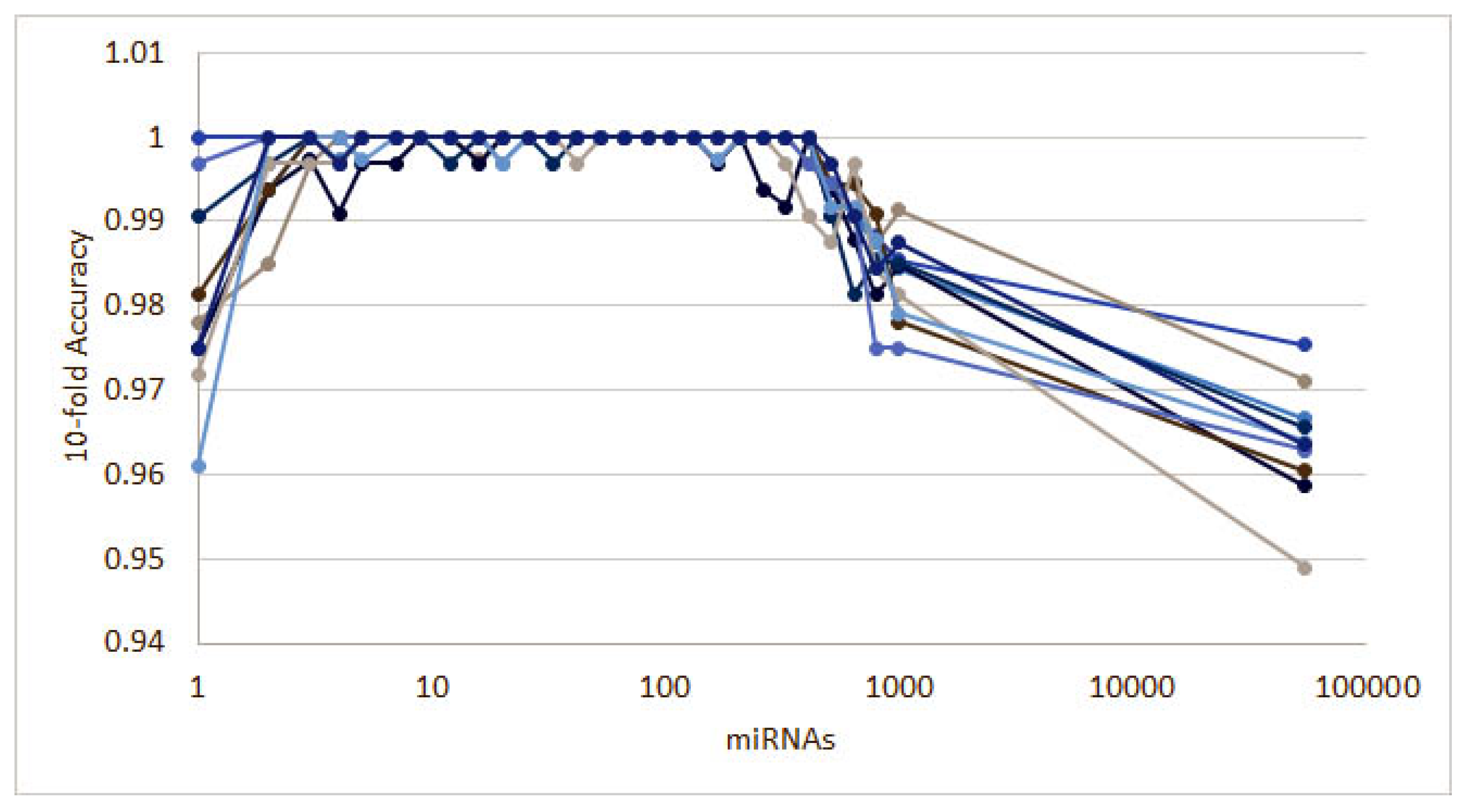
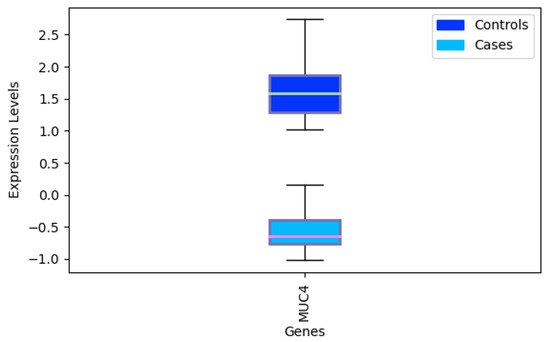
Figure A5.
Expression levels of MUC4 and difference between tumor tissue and controls.
Figure A5.
Expression levels of MUC4 and difference between tumor tissue and controls.
References
- Larrea, E.; Sole, C.; Manterola, L.; Goicoechea, I.; Armesto, M.; Arestin, M.; Caffarel, M.M.; Araujo, A.M.; Araiz, M.; Fernandez-Mercado, M.; et al. New concepts in cancer biomarkers: Circulating miRNAs in liquid biopsies. Int. J. Mol. Sci. 2016, 17, 627. [Google Scholar] [CrossRef]
- He, Y.; Lin, J.; Kong, D.; Huang, M.; Xu, C.; Kim, T.K.; Etheridge, A.; Luo, Y.; Ding, Y.; Wang, K. Current state of circulating microRNAs as cancer biomarkers. Clin. Chem. 2015, 61, 1138–1155. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Jeon, K.; Lee, J.T.; Kim, S.; Kim, V.N. MicroRNA maturation: Stepwise processing and subcellular localization. EMBO J. 2002, 21, 4663–4670. [Google Scholar] [CrossRef] [PubMed]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Price, C.; Chen, J. MicroRNAs in cancer biology and therapy: Current status and perspectives. Genes Dis. 2014, 1, 53–63. [Google Scholar] [CrossRef]
- Han, J.; Lee, Y.; Yeom, K.H.; Kim, Y.K.; Jin, H.; Kim, V.N. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004, 18, 3016–3027. [Google Scholar] [CrossRef]
- Cheng, C.; Wang, Q.; You, W.; Chen, M.; Xia, J. MiRNAs as biomarkers of myocardial infarction: A meta-analysis. PLoS ONE 2014, 9, e88566. [Google Scholar] [CrossRef]
- Huang, J.T.; Wang, J.; Srivastava, V.; Sen, S.; Liu, S.M. MicroRNA machinery genes as novel biomarkers for cancer. Front. Oncol. 2014, 4, 113. [Google Scholar] [CrossRef]
- Zhang, Y.H.; Jin, M.; Li, J.; Kong, X. Identifying circulating miRNA biomarkers for early diagnosis and monitoring of lung cancer. Biochim. Biophys. Acta (BBA) Mol. Basis Dis. 2020, 165847. [Google Scholar] [CrossRef]
- Zheng, D.; Ding, Y.; Ma, Q.; Zhao, L.; Guo, X.; Shen, Y.; He, Y.; Wei, W.; Liu, F. Identification of serum microRNAs as novel biomarkers in esophageal squamous cell carcinoma using feature selection algorithms. Front. Oncol. 2019, 8, 674. [Google Scholar] [CrossRef]
- Fehlmann, T.; Kahraman, M.; Ludwig, N.; Backes, C.; Galata, V.; Keller, V.; Geffers, L.; Mercaldo, N.; Hornung, D.; Weis, T.; et al. Evaluating the Use of Circulating MicroRNA Profiles for Lung Cancer Detection in Symptomatic Patients. JAMA Oncol. 2020, 6, 714–723. [Google Scholar] [CrossRef] [PubMed]
- Ning, Z.; Feng, C.; Song, C.; Liu, W.; Shang, D.; Li, M.; Wang, Q.; Zhao, J.; Liu, Y.; Chen, J.; et al. Topologically inferring active miRNA-mediated subpathways toward precise cancer classification by directed random walk. Mol. Oncol. 2019, 13, 2211–2226. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, C.C.; Cheng, H.H.; Tewari, M. MicroRNA profiling: Approaches and considerations. Nat. Rev. Genet. 2012, 13, 358–369. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 2007, 21, 1025–1030. [Google Scholar] [CrossRef] [PubMed]
- Al-Sheikh, Y.A.; Ghneim, H.K.; Softa, K.I.; Al-Jobran, A.A.; Al-Obeed, O.; Mohamed, M.A.; Abdulla, M.; Aboul-Soud, M.A. Expression profiling of selected microRNA signatures in plasma and tissues of Saudi colorectal cancer patients by qPCR. Oncol. Lett. 2016, 11, 1406–1412. [Google Scholar] [CrossRef]
- Adam, L.; Wszolek, M.F.; Liu, C.G.; Jing, W.; Diao, L.; Zien, A.; Zhang, J.D.; Jackson, D.; Dinney, C.P. Plasma microRNA profiles for bladder cancer detection. In Urologic Oncology: Seminars and Original Investigations; Elsevier: Amsterdam, The Netherlands, 2013; Volume 31, pp. 1701–1708. [Google Scholar]
- Liao, Z.; Li, D.; Wang, X.; Li, L.; Zou, Q. Cancer diagnosis through IsomiR expression with machine learning method. Curr. Bioinform. 2018, 13, 57–63. [Google Scholar] [CrossRef]
- Rincon, A.L.; Tonda, A.; Elati, M.; Schwander, O.; Piwowarski, B.; Gallinari, P. Evolutionary Optimization of Convolutional Neural Networks for Cancer miRNA Biomarkers Classification. Appl. Soft Comput. 2018. [Google Scholar] [CrossRef]
- Yang, S.; Guo, L.; Shao, F.; Zhao, Y.; Chen, F. A systematic evaluation of feature selection and classification algorithms using simulated and real miRNA sequencing data. Comput. Math. Methods Med. 2015, 2015. [Google Scholar] [CrossRef]
- Saha, S.; Mitra, S.; Yadav, R.K. A stack-based ensemble framework for detecting cancer microRNA biomarkers. Genom. Proteom. Bioinform. 2017, 15, 381–388. [Google Scholar] [CrossRef]
- Lopez-Rincon, A.; Martinez-Archundia, M.; Martinez-Ruiz, G.U.; Tonda, A. Ensemble Feature Selection and Meta-Analysis of Cancer miRNA Biomarkers. bioRxiv 2018. [Google Scholar] [CrossRef]
- Lopez-Rincon, A.; Martinez-Archundia, M.; Martinez-Ruiz, G.U.; Schoenhuth, A.; Tonda, A. Automatic discovery of 100-miRNA signature for cancer classification using ensemble feature selection. BMC Bioinform. 2019, 20, 480. [Google Scholar] [CrossRef]
- Calore, F.; Lovat, F.; Garofalo, M. Non-coding RNAs and cancer. Int. J. Mol. Sci. 2013, 14, 17085–17110. [Google Scholar] [CrossRef] [PubMed]
- Cheng, G. Circulating miRNAs: Roles in cancer diagnosis, prognosis and therapy. Adv. Drug Deliv. Rev. 2015, 81, 75–93. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, K.Y.; Liu, S.M.; Sen, S. Tumor-associated circulating microRNAs as biomarkers of cancer. Molecules 2014, 19, 1912–1938. [Google Scholar] [CrossRef] [PubMed]
- Leshkowitz, D.; Horn-Saban, S.; Parmet, Y.; Feldmesser, E. Differences in microRNA detection levels are technology and sequence dependent. RNA 2013, 19, 527–538. [Google Scholar] [CrossRef] [PubMed]
- Del Vescovo, V.; Meier, T.; Inga, A.; Denti, M.A.; Borlak, J. A cross-platform comparison of affymetrix and Agilent microarrays reveals discordant miRNA expression in lung tumors of c-Raf transgenic mice. PLoS ONE 2013, 8, e78870. [Google Scholar] [CrossRef]
- Bassani, N.; Ambrogi, F.; Biganzoli, E. Assessing agreement between miRNA microarray platforms. Microarrays 2014, 3, 302–321. [Google Scholar] [CrossRef]
- Abeel, T.; Helleputte, T.; Van de Peer, Y.; Dupont, P.; Saeys, Y. Robust biomarker identification for cancer diagnosis with ensemble feature selection methods. Bioinformatics 2009, 26, 392–398. [Google Scholar] [CrossRef]
- Saeys, Y.; Abeel, T.; Van de Peer, Y. Robust feature selection using ensemble feature selection techniques. In Proceedings of the Joint European Conference on Machine Learning and Knowledge Discovery in Databases, Antwerp, Belgium, 15–19 September 2008; pp. 313–325. [Google Scholar]
- Seijo-Pardo, B.; Porto-Diaz, I.; Bolon-Canedo, V.; Alonso-Betanzos, A. Ensemble feature selection: Homogeneous and heterogeneous approaches. Knowl.-Based Syst. 2017, 118, 124–139. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Edgar, R.; Domrachev, M.; Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef] [PubMed]
- Peña-Chilet, M.; Martínez, M.T.; Pérez-Fidalgo, J.A.; Peiró-Chova, L.; Oltra, S.S.; Tormo, E.; Alonso-Yuste, E.; Martinez-Delgado, B.; Eroles, P.; Climent, J.; et al. MicroRNA profile in very young women with breast cancer. BMC Cancer 2014, 14, 529. [Google Scholar] [CrossRef] [PubMed]
- Romero-Cordoba, S.L.; Rodriguez-Cuevas, S.; Bautista-Pina, V.; Maffuz-Aziz, A.; D’Ippolito, E.; Cosentino, G.; Baroni, S.; Iorio, M.V.; Hidalgo-Miranda, A. Loss of function of miR-342-3p results in MCT1 over-expression and contributes to oncogenic metabolic reprogramming in triple negative breast cancer. Sci. Rep. 2018, 8, 12252. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.J.; Lee, H.S.; Burt, B.M.; Lee, G.K.; Yoon, K.A.; Park, Y.Y.; Sohn, B.H.; Kim, S.B.; Kim, M.S.; Lee, J.M.; et al. Integrated genomic analysis of recurrence-associated small non-coding RNAs in oesophageal cancer. Gut 2017, 66, 215–225. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.C.; Chiu, Y.L.; Banerjee, S.; Park, K.; Mosquera, J.M.; Giannopoulou, E.; Alves, P.; Tewari, A.K.; Gerstein, M.B.; Beltran, H.; et al. Epigenetic repression of miR-31 disrupts androgen receptor homeostasis and contributes to prostate cancer progression. Cancer Res. 2013, 73, 1232–1244. [Google Scholar] [CrossRef]
- Casanova-Salas, I.; Rubio-Briones, J.; Calatrava, A.; Mancarella, C.; Masiá, E.; Casanova, J.; Fernández-Serra, A.; Rubio, L.; Ramírez-Backhaus, M.; Armiñán, A.; et al. Identification of miR-187 and miR-182 as biomarkers of early diagnosis and prognosis in patients with prostate cancer treated with radical prostatectomy. J. Urol. 2014, 192, 252–259. [Google Scholar] [CrossRef]
- Hermansen, S.K.; Sørensen, M.D.; Hansen, A.; Knudsen, S.; Alvarado, A.G.; Lathia, J.D.; Kristensen, B.W. A 4-miRNA signature to predict survival in glioblastomas. PLoS ONE 2017, 12, e0188090. [Google Scholar] [CrossRef]
- Jepsen, R.K.; Novotny, G.W.; Klarskov, L.L.; Bang-Berthelsen, C.H.; Haakansson, I.T.; Hansen, A.; Christensen, I.J.; Riis, L.B.; Høgdall, E. Early metastatic colorectal cancers show increased tissue expression of miR-17/92 cluster members in the invasive tumor front. Hum. Pathol. 2018, 80, 231–238. [Google Scholar] [CrossRef]
- Zhang, X.; Ni, Z.; Duan, Z.; Xin, Z.; Wang, H.; Tan, J.; Wang, G.; Li, F. Overexpression of E2F mRNAs associated with gastric cancer progression identified by the transcription factor and miRNA co-regulatory network analysis. PLoS ONE 2015, 10, e0116979. [Google Scholar] [CrossRef]
- Elgaaen, B.V.; Olstad, O.K.; Haug, K.B.F.; Brusletto, B.; Sandvik, L.; Staff, A.C.; Gautvik, K.M.; Davidson, B. Global miRNA expression analysis of serous and clear cell ovarian carcinomas identifies differentially expressed miRNAs including miR-200c-3p as a prognostic marker. BMC Cancer 2014, 14, 80. [Google Scholar]
- Trevino, V.; Falciani, F. GALGO: An R package for multivariate variable selection using genetic algorithms. Bioinformatics 2006, 22, 1154–1156. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Siklenka, K.; Arora, S.K.; Ribeiro, P.; Kimmins, S.; Xia, J. miRNet-dissecting miRNA-target interactions and functional associations through network-based visual analysis. Nucleic Acids Res. 2016, 44, W135–W141. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2016, 45, D353–D361. [Google Scholar] [CrossRef] [PubMed]
- Falcon, S.; Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 2006, 23, 257–258. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Li, T.; Bai, Z.; Yang, Y.; Liu, X.; Zhan, J.; Shi, B. Breast cancer intrinsic subtype classification, clinical use and future trends. Am. J. Cancer Res. 2015, 5, 2929. [Google Scholar] [PubMed]
- Weiss, M. Your Guide to the Breast Cancer Pathology Report; Breastcancer. org: Ardmore, PA, USA, 2013. [Google Scholar]
- Jurman, G.; Riccadonna, S.; Furlanello, C. A comparison of MCC and CEN error measures in multi-class prediction. PLoS ONE 2012, 7, e41882. [Google Scholar] [CrossRef]
- Stegh, A.H. Targeting the p53 signaling pathway in cancer therapy–the promises, challenges and perils. Expert Opin. Ther. Targets 2012, 16, 67–83. [Google Scholar] [CrossRef]
- Collado, M.; Blasco, M.A.; Serrano, M. Cellular senescence in cancer and aging. Cell 2007, 130, 223–233. [Google Scholar] [CrossRef]
- Šimundić, A.M. Measures of diagnostic accuracy: Basic definitions. eJIFCC 2009, 19, 203. [Google Scholar]
- Mandrekar, J.N. Receiver operating characteristic curve in diagnostic test assessment. J. Thorac. Oncol. 2010, 5, 1315–1316. [Google Scholar] [CrossRef]
- Thakral, S.; Ghoshal, K. miR-122 is a unique molecule with great potential in diagnosis, prognosis of liver disease, and therapy both as miRNA mimic and antimir. Curr. Gene Ther. 2015, 15, 142–150. [Google Scholar] [CrossRef] [PubMed]
- Bandiera, S.; Pfeffer, S.; Baumert, T.F.; Zeisel, M.B. miR-122—A key factor and therapeutic target in liver disease. J. Hepatol. 2015, 62, 448–457. [Google Scholar] [CrossRef] [PubMed]
- Ergün, S.; Ulasli, M.; Igci, Y.Z.; Igci, M.; Kırkbes, S.; Borazan, E.; Balik, A.; Yumrutaş, Ö.; Camci, C.; Cakmak, E.A.; et al. The association of the expression of miR-122-5p and its target ADAM10 with human breast cancer. Mol. Biol. Rep. 2015, 42, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, X. miR-122-5p promotes aggression and epithelial-mesenchymal transition in triple-negative breast cancer by suppressing charged multivesicular body protein 3 through mitogen-activated protein kinase signaling. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]
- Perez-Añorve, I.X.; Gonzalez-De la Rosa, C.H.; Soto-Reyes, E.; Beltran-Anaya, F.O.; Del Moral-Hernandez, O.; Salgado-Albarran, M.; Angeles-Zaragoza, O.; Gonzalez-Barrios, J.A.; Landero-Huerta, D.A.; Chavez-Saldaña, M.; et al. New insights into radioresistance in breast cancer identify a dual function of miR-122 as a tumor suppressor and oncomiR. Mol. Oncol. 2019, 13, 1249–1267. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, S.; Furuya, S.; Shiraishi, K.; Shimizu, H.; Akaike, H.; Hosomura, N.; Kawaguchi, Y.; Amemiya, H.; Kawaida, H.; Sudo, M.; et al. miR-122-5p as a novel biomarker for alpha-fetoprotein-producing gastric cancer. World J. Gastrointest. Oncol. 2018, 10, 344. [Google Scholar] [CrossRef]
- Lee, H.; Han, S.; Kwon, C.S.; Lee, D. Biogenesis and regulation of the let-7 miRNAs and their functional implications. Protein Cell 2016, 7, 100–113. [Google Scholar] [CrossRef]
- Liu, K.; Zhang, C.; Li, T.; Ding, Y.; Tu, T.; Zhou, F.; Qi, W.; Chen, H.; Sun, X. Let-7a inhibits growth and migration of breast cancer cells by targeting HMGA1. Int. J. Oncol. 2015, 46, 2526–2534. [Google Scholar] [CrossRef]
- Khalighfard, S.; Alizadeh, A.M.; Irani, S.; Omranipour, R. Plasma miR-21, miR-155, miR-10b, and Let-7a as the potential biomarkers for the monitoring of breast cancer patients. Sci. Rep. 2018, 8, 17981. [Google Scholar] [CrossRef]
- Zhao, W.; Hu, J.X.; Hao, R.M.; Zhang, Q.; Guo, J.Q.; Li, Y.J.; Xie, N.; Liu, L.Y.; Wang, P.Y.; Zhang, C.; et al. Induction of microRNA-let-7a inhibits lung adenocarcinoma cell growth by regulating cyclin D1. Oncol. Rep. 2018, 40, 1843–1854. [Google Scholar] [CrossRef]
- Yang, Q.; Jie, Z.; Cao, H.; Greenlee, A.R.; Yang, C.; Zou, F.; Jiang, Y. Low-level expression of let-7a in gastric cancer and its involvement in tumorigenesis by targeting RAB40C. Carcinogenesis 2011, 32, 713–722. [Google Scholar] [CrossRef]
- Grossi, I.; Salvi, A.; Baiocchi, G.; Portolani, N.; De Petro, G. Functional role of microRNA-23b-3p in cancer biology. MicroRNA 2018, 7, 156–166. [Google Scholar] [CrossRef] [PubMed]
- Kou, C.H.; Zhou, T.; Han, X.L.; Zhuang, H.J.; Qian, H.X. Downregulation of mir-23b in plasma is associated with poor prognosis in patients with colorectal cancer. Oncol. Lett. 2016, 12, 4838–4844. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, K.; Han, K.; Tang, H.; Yin, X.; Zhang, J.; Zhang, X.; Zhang, L. Up-regulation of plasma miR-23b is associated with poor prognosis of gastric cancer. Med. Sci. Monit. Int. Med. J. Exp. Clin. Res. 2016, 22, 256. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Li, T.; Chen, G.; Yan, G.; Zhang, X.; Wan, Y.; Li, Q.; Zhu, B.; Zhuo, W. Identification of a serum microRNA expression signature for detection of lung cancer, involving miR-23b, miR-221, miR-148b and miR-423-3p. Lung Cancer 2017, 114, 6–11. [Google Scholar] [CrossRef]
- Chen, D.; Wu, X.; Xia, M.; Wu, F.; Ding, J.; Jiao, Y.; Zhan, Q.; An, F. Upregulated exosomic miR-23b-3p plays regulatory roles in the progression of pancreatic cancer. Oncol. Rep. 2017, 38, 2182–2188. [Google Scholar] [CrossRef]
- Monteleone, N.J.; Lutz, C.S. miR-708-5p: A microRNA with emerging roles in cancer. Oncotarget 2017, 8, 71292. [Google Scholar] [CrossRef]
- Jang, J.S.; Jeon, H.S.; Sun, Z.; Aubry, M.C.; Tang, H.; Park, C.H.; Rakhshan, F.; Schultz, D.A.; Kolbert, C.P.; Lupu, R.; et al. Increased miR-708 expression in NSCLC and its association with poor survival in lung adenocarcinoma from never smokers. Clin. Cancer Res. 2012, 18, 3658–3667. [Google Scholar] [CrossRef]
- Fedatto, P.F.; de Carvalho, T.I.; de Oliveira, J.C.; Antônio, D.S.M.; Pezuk, J.A.; da Cunha Tirapelli, D.P.; Féres, O.; da Rocha, J.J.R.; Scrideli, C.A.; Tone, L.G.; et al. MiR-708-5p as a Predictive Marker of Colorectal Cancer Prognosis. J. Anal. Oncol. 2016, 5, 14–23. [Google Scholar]
- Song, T.; Zhang, X.; Zhang, L.; Dong, J.; Cai, W.; Gao, J.; Hong, B. miR-708 promotes the development of bladder carcinoma via direct repression of Caspase-2. J. Cancer Res. Clin. Oncol. 2013, 139, 1189–1198. [Google Scholar] [CrossRef]
- Saini, S.; Yamamura, S.; Majid, S.; Shahryari, V.; Hirata, H.; Tanaka, Y.; Dahiya, R. MicroRNA-708 induces apoptosis and suppresses tumorigenicity in renal cancer cells. Cancer Res. 2011, 71, 6208–6219. [Google Scholar] [CrossRef] [PubMed]
- Cochrane, D.R.; Howe, E.N.; Spoelstra, N.S.; Richer, J.K. Loss of miR-200c: A marker of aggressiveness and chemoresistance in female reproductive cancers. J. Oncol. 2010, 2010. [Google Scholar] [CrossRef]
- Liu, X.G.; Zhu, W.Y.; Huang, Y.Y.; Ma, L.N.; Zhou, S.Q.; Wang, Y.K.; Zeng, F.; Zhou, J.H.; Zhang, Y.K. High expression of serum miR-21 and tumor miR-200c associated with poor prognosis in patients with lung cancer. Med. Oncol. 2012, 29, 618–626. [Google Scholar] [CrossRef] [PubMed]
- Mutlu, M.; Raza, U.; Saatci, Ö.; Eyüpoğlu, E.; Yurdusev, E.; Şahin, Ö. miR-200c: A versatile watchdog in cancer progression, EMT, and drug resistance. J. Mol. Med. 2016, 94, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zhang, C.; Zhao, Y.; Feng, Z. MicroRNA control of p53. J. Cell. Biochem. 2017, 118, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Takwi, A.; Li, Y. The p53 pathway encounters the microRNA world. Curr. Genom. 2009, 10, 194–197. [Google Scholar] [CrossRef]
- Suzuki, H.I.; Yamagata, K.; Sugimoto, K.; Iwamoto, T.; Kato, S.; Miyazono, K. Modulation of microRNA processing by p53. Nature 2009, 460, 529–533. [Google Scholar] [CrossRef]
- Fulda, S.; Gorman, A.M.; Hori, O.; Samali, A. Cellular stress responses: Cell survival and cell death. Int. J. Cell Biol. 2010, 2010. [Google Scholar] [CrossRef]
- Chen, M.; Xie, S. Therapeutic targeting of cellular stress responses in cancer. Thorac. Cancer 2018, 9, 1575–1582. [Google Scholar] [CrossRef]
- Babar, I.A.; Slack, F.J.; Weidhaas, J.B. miRNA modulation of the cellular stress response. Future Oncol. 2008. [Google Scholar] [CrossRef]
- Eichner, L.J.; Perry, M.C.; Dufour, C.R.; Bertos, N.; Park, M.; St-Pierre, J.; Giguère, V. miR-378 mediates metabolic shift in breast cancer cells via the PGC-1β/ERRγ transcriptional pathway. Cell Metab. 2010, 12, 352–361. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.Y.; Deng, Z.Q.; Liu, F.Q.; Qian, J.; Lin, J.; Tang, Q.; Wen, X.M.; Zhou, J.D.; Zhang, Y.Y.; Zhu, X.W. Association between mir-24 and mir-378 in formalin-fixed paraffin-embedded tissues of breast cancer. Int. J. Clin. Exp. Pathol. 2014, 7, 4261. [Google Scholar] [PubMed]
- He, Y.J.; Wu, J.Z.; Ji, M.H.; Ma, T.; Qiao, E.Q.; Ma, R.; Tang, J.H. miR-342 is associated with estrogen receptor-α expression and response to tamoxifen in breast cancer. Exp. Ther. Med. 2013, 5, 813–818. [Google Scholar] [CrossRef]
- Wei, Y.; Lai, X.; Yu, S.; Chen, S.; Ma, Y.; Zhang, Y.; Li, H.; Zhu, X.; Yao, L.; Zhang, J. Exosomal miR-221/222 enhances tamoxifen resistance in recipient ER-positive breast cancer cells. Breast Cancer Res. Treat. 2014, 147, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.J.; Lin, J.; Yang, H.; Kong, W.; He, L.; Ma, X.; Coppola, D.; Cheng, J.Q. MicroRNA-221/222 negatively regulates estrogen receptorα and is associated with tamoxifen resistance in breast cancer. J. Biol. Chem. 2008, 283, 31079–31086. [Google Scholar] [CrossRef] [PubMed]
- Cittelly, D.M.; Das, P.M.; Spoelstra, N.S.; Edgerton, S.M.; Richer, J.K.; Thor, A.D.; Jones, F.E. Downregulation of miR-342 is associated with tamoxifen resistant breast tumors. Mol. Cancer 2010, 9, 317. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.X.; Wang, C.L.; Yu, A.L.; Wang, Q.Y.; Zhan, M.N.; Tang, J.; Gong, X.F.; Yin, Q.Q.; He, M.; He, J.R.; et al. MiR-630 suppresses breast cancer progression by targeting metadherin. Oncotarget 2016, 7, 1288. [Google Scholar] [CrossRef]
- Wang, S.; Bian, C.; Yang, Z.; Bo, Y.; Li, J.; Zeng, L.; Zhou, H.; Zhao, R.C. miR-145 inhibits breast cancer cell growth through RTKN. Int. J. Oncol. 2009, 34, 1461–1466. [Google Scholar]
- Yan, X.; Chen, X.; Liang, H.; Deng, T.; Chen, W.; Zhang, S.; Liu, M.; Gao, X.; Liu, Y.; Zhao, C.; et al. miR-143 and miR-145 synergistically regulate ERBB3 to suppress cell proliferation and invasion in breast cancer. Mol. Cancer 2014, 13, 220. [Google Scholar] [CrossRef]
- Santolla, M.F.; Lappano, R.; Cirillo, F.; Rigiracciolo, D.C.; Sebastiani, A.; Abonante, S.; Tassone, P.; Tagliaferri, P.; Di Martino, M.T.; Maggiolini, M.; et al. miR-221 stimulates breast cancer cells and cancer- associated fibroblasts (CAFs) through selective interference with the A20/c-Rel/CTGF signaling. J. Exp. Clin. Cancer Res. 2018, 37, 94. [Google Scholar] [CrossRef]
- Chen, W.X.; Hu, Q.; Qiu, M.T.; Zhong, S.L.; Xu, J.J.; Tang, J.H.; Zhao, J.H. miR-221/222: Promising biomarkers for breast cancer. Tumor Biol. 2013, 34, 1361–1370. [Google Scholar] [CrossRef]
- Stinson, S.; Lackner, M.R.; Adai, A.T.; Yu, N.; Kim, H.J.; O’Brien, C.; Spoerke, J.; Jhunjhunwala, S.; Boyd, Z.; Januario, T.; et al. TRPS1 targeting by miR-221/222 promotes the epithelial-to-mesenchymal transition in breast cancer. Sci. Signal. 2011, 4, ra41. [Google Scholar]
- Liu, Y.X.; Zhao, L.P.; Zhang, Y.L.; Dong, Y.Y.; Ren, H.Y.; Diao, K.X.; Mi, X.Y. MiR-630 inhibits cells migration and invasion by targeting SOX4 in triple-negative breast cancer. Int. J. Clin. Exp. Pathol. 2016, 9, 9097–9105. [Google Scholar]
- Spizzo, R.; Nicoloso, M.; Lupini, L.; Lu, Y.; Fogarty, J.; Rossi, S.; Zagatti, B.; Fabbri, M.; Veronese, A.; Liu, X.; et al. miR-145 participates with TP53 in a death-promoting regulatory loop and targets estrogen receptor-α in human breast cancer cells. Cell Death Differ. 2010, 17, 246–254. [Google Scholar] [CrossRef]
- Zare, M.; Bastami, M.; Solali, S.; Alivand, M.R. Aberrant miRNA promoter methylation and EMT-involving miRNAs in breast cancer metastasis: Diagnosis and therapeutic implications. J. Cell. Physiol. 2018, 233, 3729–3744. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S.; Saini, H.K.; van Dongen, S.; Enright, A.J. miRBase: Tools for microRNA genomics. Nucleic Acids Res. 2007, 36, D154–D158. [Google Scholar] [CrossRef] [PubMed]
- Mootha, V.K.; Lindgren, C.M.; Eriksson, K.F.; Subramanian, A.; Sihag, S.; Lehar, J.; Puigserver, P.; Carlsson, E.; Ridderstråle, M.; Laurila, E.; et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 2003, 34, 267–273. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.L.; Wang, Z.Q.; Zeng, Z.L.; Wu, W.J.; Zhang, D.S.; Luo, H.Y.; Wang, F.; Qiu, M.Z.; Wang, D.S.; Ren, C.; et al. Identification of microRNA-214 as a negative regulator of colorectal cancer liver metastasis by way of regulation of fibroblast growth factor receptor 1 expression. Hepatology 2014, 60, 598–609. [Google Scholar] [CrossRef]
- Dodd, L.E.; Sengupta, S.; Chen, I.H.; Den Boon, J.A.; Cheng, Y.J.; Westra, W.; Newton, M.A.; Mittl, B.F.; McShane, L.; Chen, C.J.; et al. Genes involved in DNA repair and nitrosamine metabolism and those located on chromosome 14q32 are dysregulated in nasopharyngeal carcinoma. Cancer Epidemiol. Prev. Biomark. 2006, 15, 2216–2225. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Jain, M.; Sasson, A.R.; Batra, S.K. MUC4 as a diagnostic marker in cancer. Expert Opin. Med. Diagn. 2008, 2, 891–910. [Google Scholar] [CrossRef]
- Wei, Y.C.; Yang, S.F.; Chang, S.L.; Chen, T.J.; Lee, S.W.; Chen, H.S.; Lin, L.C.; Li, C.F. Periostin overexpression is associated with worse prognosis in nasopharyngeal carcinoma from endemic area: A cohort study. OncoTargets Ther. 2018, 11, 3205. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).