Genomic Signature of Oral Squamous Cell Carcinomas from Non-Smoking Non-Drinking Patients
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
3. Results
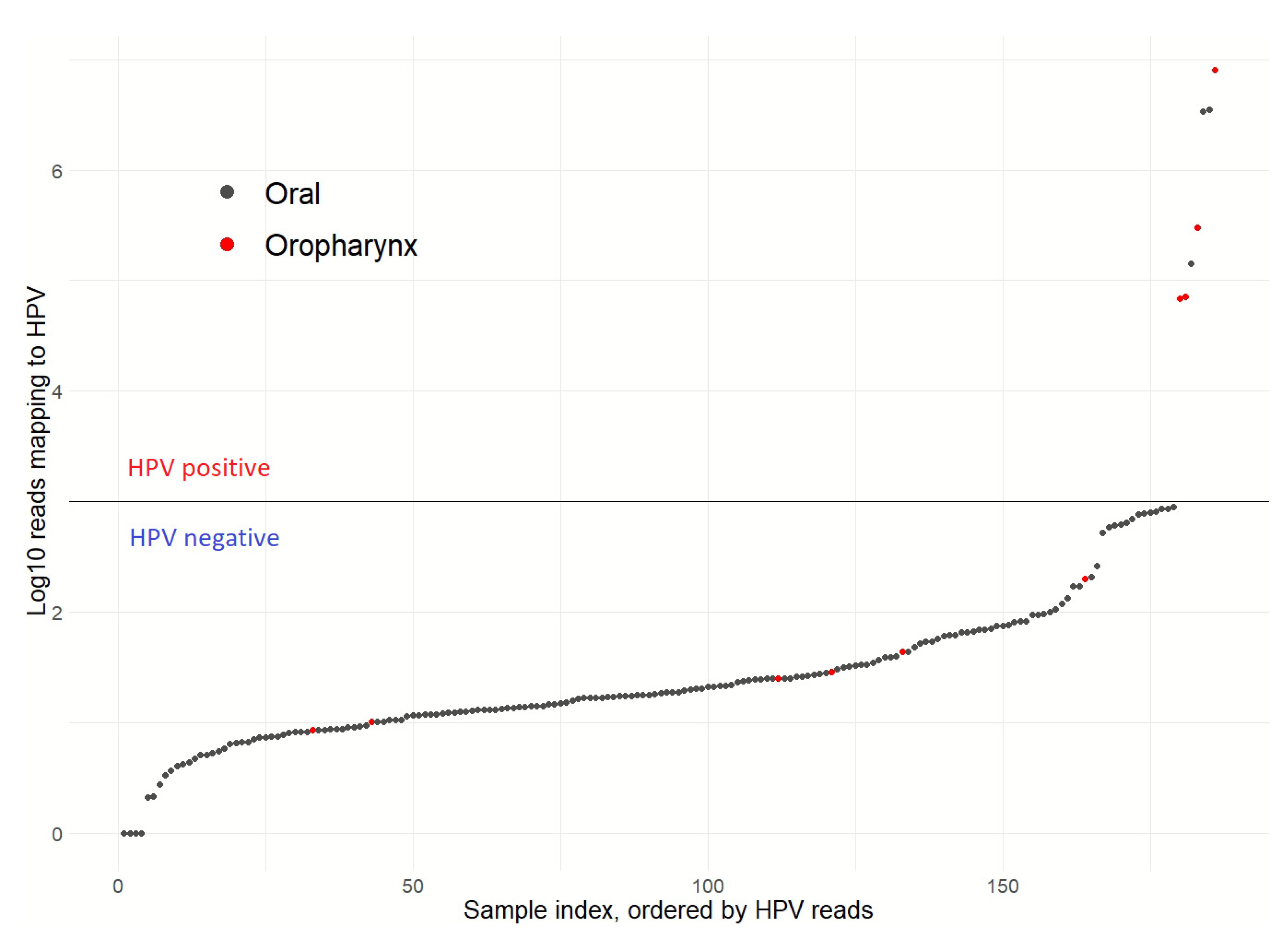
3.1. Patient Clinical Characteristics and HPV Status
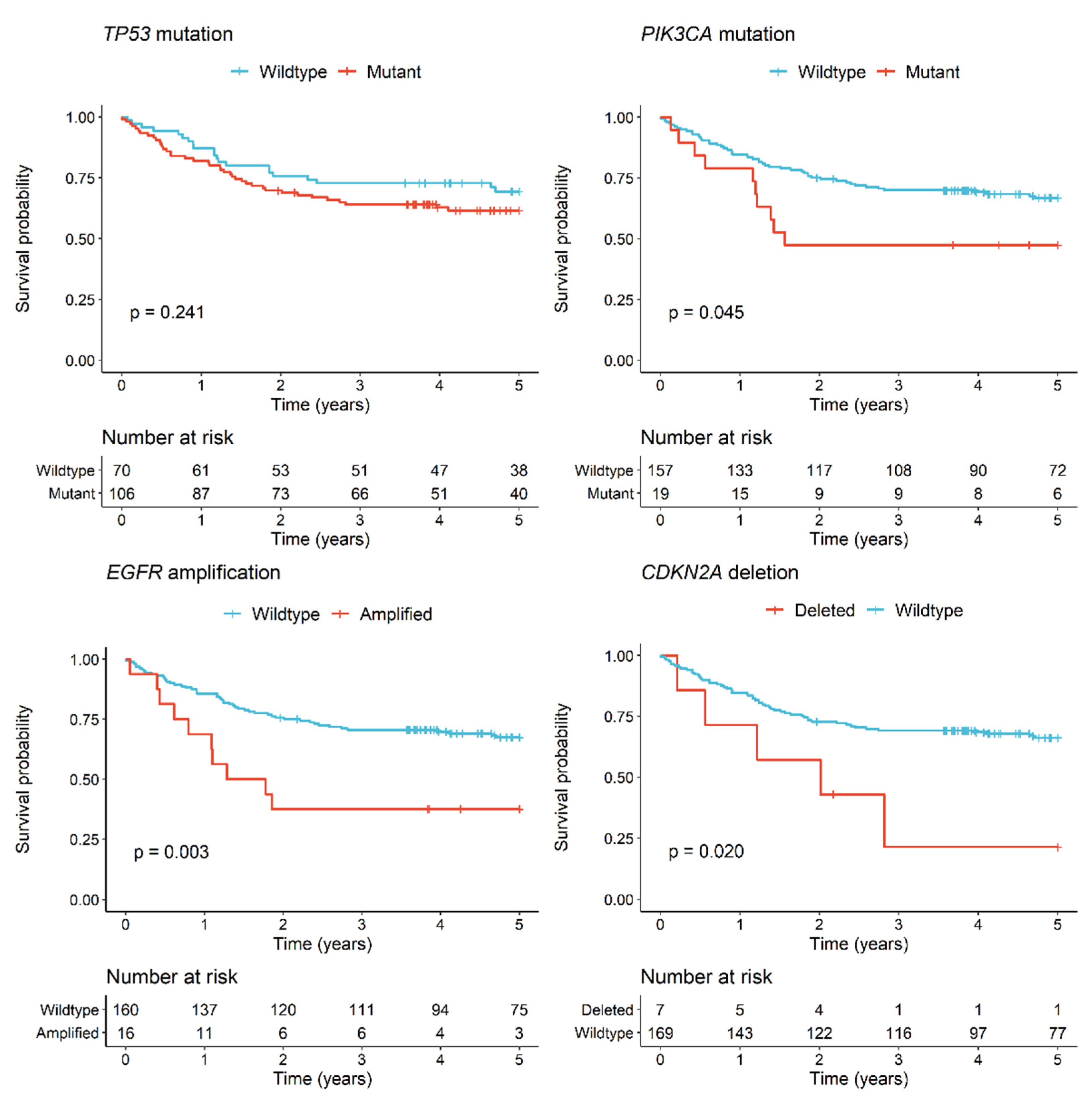
3.2. Genomic Alterations and Clinical Associations for OSCC Patients
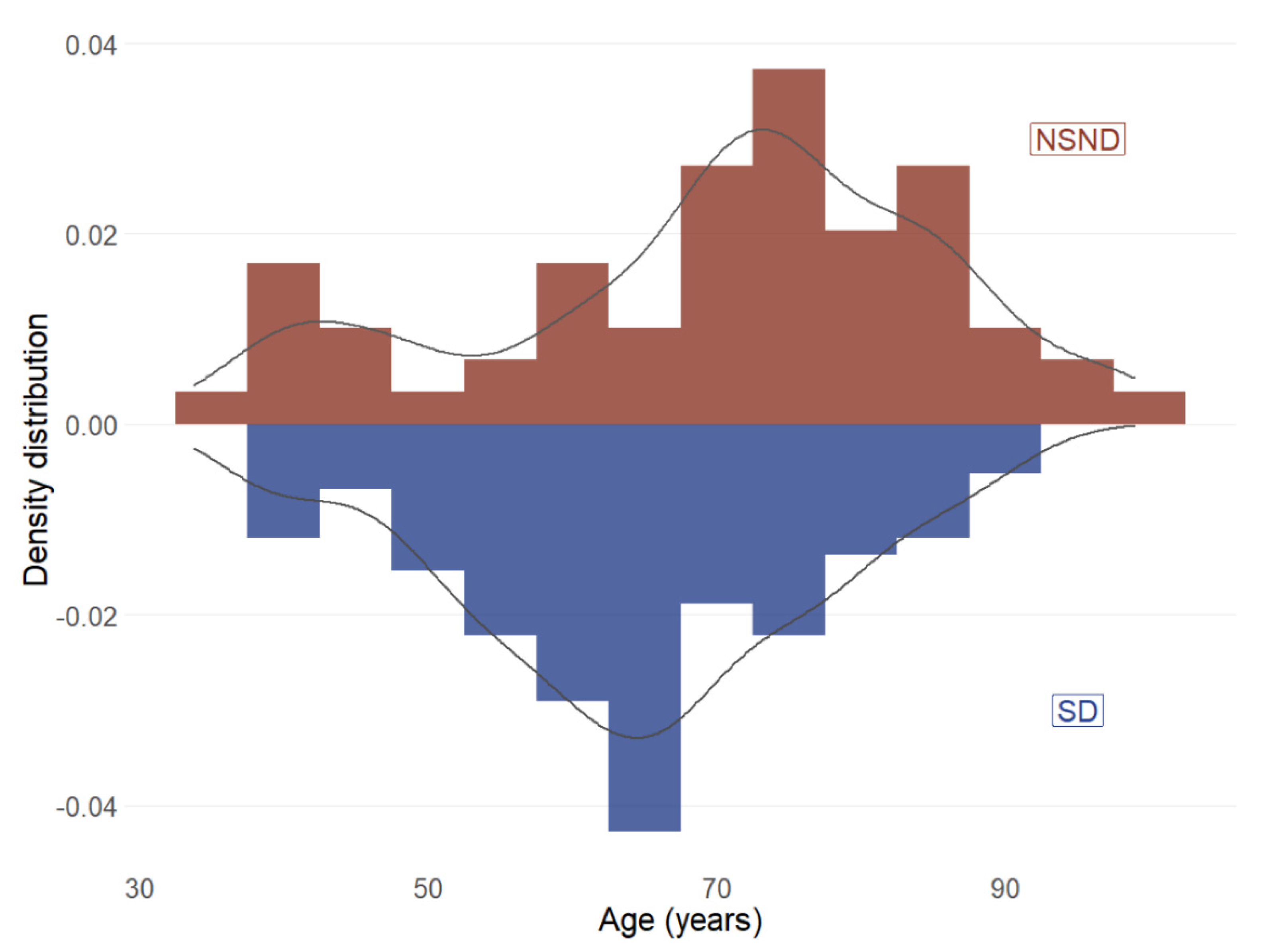
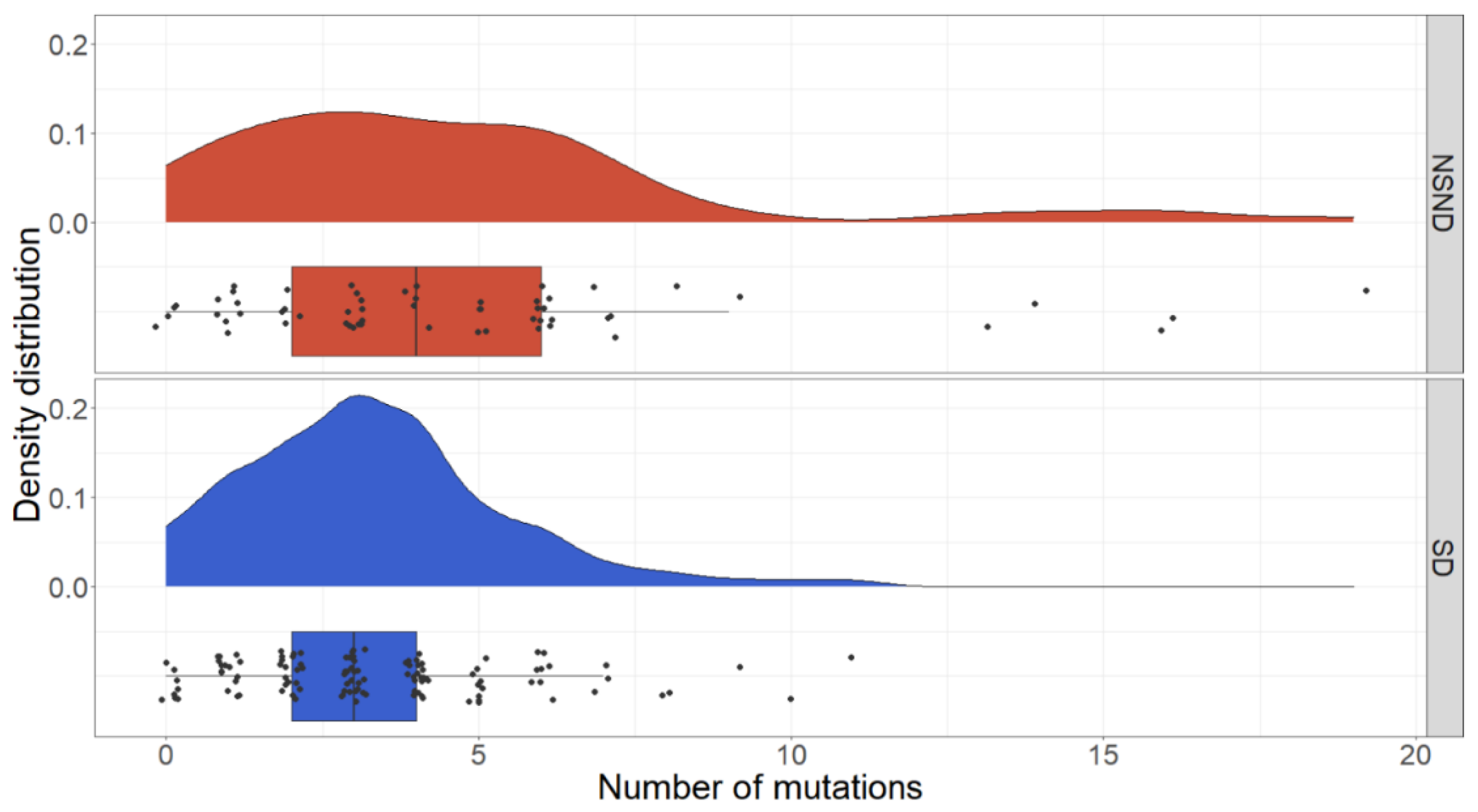
3.3. Mutation Differences between NSND and SD Patients
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control (CDC). Smoking and cancer. MMWR Morb. Mortal. Wkly Rep. 1982, 31, 77–80. [Google Scholar]
- Tuyns, A.J. Epidemiology of alcohol and cancer. Cancer Res. 1979, 39 Pt 2, 2840–2843. [Google Scholar]
- Chen, F.; Wang, J.; Chen, J.; Yan, L.; Hu, Z.; Wu, J.; Bao, X.; Lin, L.; Wang, R.; Cai, L.; et al. Serum copper and zinc levels and the risk of oral cancer: A new insight based on large-scale case-control study. Oral Dis. 2019, 25, 80–86. [Google Scholar] [CrossRef] [Green Version]
- Ang, K.K.; Harris, J.; Wheeler, R.; Weber, R.; Rosenthal, D.I.; Nguyen-Tan, P.F.; Westra, W.H.; Chung, C.H.; Jordan, R.C.; Lu, C.; et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N. Engl. J. Med. 2010, 363, 24–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; Human Papillomaviruses: Lyon, France, 2007; Volume 90.
- Koo, K.; Barrowman, R.; McCullough, M.; Iseli, T.; Wiesenfeld, D. Non-smoking non-drinking elderly females: A clinically distinct subgroup of oral squamous cell carcinoma patients. Int. J. Oral. Maxillofac. Surg. 2013, 42, 929–933. [Google Scholar] [CrossRef]
- DeAngelis, A.; Breik, O.; Koo, K.; Iseli, T.; Nastri, A.; Fua, T.; Rischin, D.; McCullough, M.; Wiesenfeld, D. Non-smoking, non-drinking elderly females, a 5year follow-up of a clinically distinct cohort of oral squamous cell carcinoma patients. Oral Oncol. 2018, 86, 113–120. [Google Scholar] [CrossRef]
- Wiseman, S.M.; Swede, H.; Stoler, D.L.; Anderson, G.R.; Rigual, N.R.; Hicks, W.L., Jr.; Douglas, W.G.; Tan, D.; Loree, T.R. Squamous cell carcinoma of the head and neck in nonsmokers and nondrinkers: An analysis of clinicopathologic characteristics and treatment outcomes. Ann. Surg. Oncol. 2003, 10, 551–557. [Google Scholar] [CrossRef]
- Farshadpour, F.; Hordijk, G.J.; Koole, R.; Slootweg, P.J. Non-smoking and non-drinking patients with head and neck squamous cell carcinoma: A distinct population. Oral Dis. 2007, 13, 239–243. [Google Scholar] [CrossRef]
- Dahlstrom, K.R.; Little, J.A.; Zafereo, M.E.; Lung, M.; Wei, Q.; Sturgis, E.M. Squamous cell carcinoma of the head and neck in never smoker-never drinkers: A descriptive epidemiologic study. Head Neck 2008, 30, 75–84. [Google Scholar] [CrossRef]
- Harris, S.L.; Kimple, R.J.; Hayes, D.N.; Couch, M.E.; Rosenman, J.G. Never-smokers, never-drinkers: Unique clinical subgroup of young patients with head and neck squamous cell cancers. Head Neck 2010, 32, 499–503. [Google Scholar] [CrossRef] [PubMed]
- Bachar, G.; Hod, R.; Goldstein, D.P.; Irish, J.C.; Gullane, P.J.; Brown, D.; Gilbert, R.W.; Hadar, T.; Feinmesser, R.; Shpitzer, T. Outcome of oral tongue squamous cell carcinoma in patients with and without known risk factors. Oral Oncol. 2011, 47, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, R.; Lopez-Lopez, J.; Mari-Roig, A.; Jane-Salas, E.; Rosello-Llabres, X.; Santos, J.R. Oral tongue squamous cell carcinoma (OTSCC): Alcohol and tobacco consumption versus non-consumption. A study in a Portuguese population. Braz. Dent. J. 2011, 22, 517–521. [Google Scholar] [CrossRef] [Green Version]
- Kruse, A.L.; Bredell, M.; Luebbers, H.T.; Gratz, K.W. Head and neck cancer in the elderly: A retrospective study over 10 years (1999–2008). Head Neck Oncol. 2010, 2, 25. [Google Scholar] [CrossRef] [Green Version]
- Laco, J.; Vosmikova, H.; Novakova, V.; Celakovsky, P.; Dolezalova, H.; Tucek, L.; Nekvindova, J.; Vosmik, M.; Cermakova, E.; Ryska, A. The role of high-risk human papillomavirus infection in oral and oropharyngeal squamous cell carcinoma in non-smoking and non-drinking patients: A clinicopathological and molecular study of 46 cases. Virchows Arch. 2011, 458, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Lechner, M.; Frampton, G.M.; Fenton, T.; Feber, A.; Palmer, G.; Jay, A.; Pillay, N.; Forster, M.; Cronin, M.T.; Lipson, D.; et al. Targeted next-generation sequencing of head and neck squamous cell carcinoma identifies novel genetic alterations in HPV+ and HPV-tumors. Genome Med. 2013, 5, 49. [Google Scholar] [CrossRef] [Green Version]
- Seiwert, T.Y.; Zuo, Z.; Keck, M.K.; Khattri, A.; Pedamallu, C.S.; Stricker, T.; Brown, C.; Pugh, T.J.; Stojanov, P.; Cho, J.; et al. Integrative and comparative genomic analysis of HPV-positive and HPV-negative head and neck squamous cell carcinomas. Clin. Cancer Res. 2015, 21, 632–641. [Google Scholar] [CrossRef] [Green Version]
- van Ginkel, J.H.; de Leng, W.W.; de Bree, R.; van Es, R.J.; Willems, S.M. Targeted sequencing reveals TP53 as a potential diagnostic biomarker in the post-treatment surveillance of head and neck cancer. Oncotarget 2016, 7, 61575–61586. [Google Scholar] [CrossRef] [Green Version]
- Chung, C.H.; Guthrie, V.B.; Masica, D.L.; Tokheim, C.; Kang, H.; Richmon, J.; Agrawal, N.; Fakhry, C.; Quon, H.; Subramaniam, R.M.; et al. Genomic alterations in head and neck squamous cell carcinoma determined by cancer gene-targeted sequencing. Ann. Oncol. 2015, 26, 1216–1223. [Google Scholar] [CrossRef]
- Chen, S.J.; Liu, H.; Liao, C.T.; Huang, P.J.; Huang, Y.; Hsu, A.; Tang, P.; Chang, Y.S.; Chen, H.C.; Yen, T.C. Ultra-deep targeted sequencing of advanced oral squamous cell carcinoma identifies a mutation-based prognostic gene signature. Oncotarget 2015, 6, 18066–18080. [Google Scholar] [CrossRef] [Green Version]
- Tinhofer, I.; Budach, V.; Saki, M.; Konschak, R.; Niehr, F.; Johrens, K.; Weichert, W.; Linge, A.; Lohaus, F.; Krause, M.; et al. Targeted next-generation sequencing of locally advanced squamous cell carcinomas of the head and neck reveals druggable targets for improving adjuvant chemoradiation. Eur. J. Cancer 2016, 57, 78–86. [Google Scholar] [CrossRef] [Green Version]
- Er, T.K.; Wang, Y.Y.; Chen, C.C.; Herreros-Villanueva, M.; Liu, T.C.; Yuan, S.S. Molecular characterization of oral squamous cell carcinoma using targeted next-generation sequencing. Oral Dis. 2015, 21, 872–878. [Google Scholar] [CrossRef]
- Al-Hebshi, N.N.; Li, S.; Nasher, A.T.; El-Setouhy, M.; Alsanosi, R.; Blancato, J.; Loffredo, C. Exome sequencing of oral squamous cell carcinoma in users of Arabian snuff reveals novel candidates for driver genes. Int. J. Cancer 2016, 139, 363–372. [Google Scholar] [CrossRef] [Green Version]
- Pickering, C.R.; Zhang, J.; Yoo, S.Y.; Bengtsson, L.; Moorthy, S.; Neskey, D.M.; Zhao, M.; Ortega Alves, M.V.; Chang, K.; Drummond, J.; et al. Integrative genomic characterization of oral squamous cell carcinoma identifies frequent somatic drivers. Cancer Discov. 2013, 3, 770–781. [Google Scholar] [CrossRef] [Green Version]
- Stransky, N.; Egloff, A.M.; Tward, A.D.; Kostic, A.D.; Cibulskis, K.; Sivachenko, A.; Kryukov, G.V.; Lawrence, M.S.; Sougnez, C.; McKenna, A.; et al. The mutational landscape of head and neck squamous cell carcinoma. Science 2011, 333, 1157–1160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Agrawal, N.; Frederick, M.J.; Pickering, C.R.; Bettegowda, C.; Chang, K.; Li, R.J.; Fakhry, C.; Xie, T.X.; Zhang, J.; Wang, J.; et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science 2011, 333, 1154–1157. [Google Scholar] [CrossRef] [Green Version]
- India Project Team of the International Cancer Genome Consortium. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat. Commun. 2013, 4, 2873. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015, 517, 576–582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakagaki, T.; Tamura, M.; Kobashi, K.; Omori, A.; Koyama, R.; Idogawa, M.; Ogi, K.; Hiratsuka, H.; Tokino, T.; Sasaki, Y. Targeted next-generation sequencing of 50 cancer-related genes in Japanese patients with oral squamous cell carcinoma. Tumour. Biol. 2018, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Su, S.C.; Lin, C.W.; Liu, Y.F.; Fan, W.L.; Chen, M.K.; Yu, C.P.; Yang, W.E.; Su, C.W.; Chuang, C.Y.; Li, W.H.; et al. Exome Sequencing of Oral Squamous Cell Carcinoma Reveals Molecular Subgroups and Novel Therapeutic Opportunities. Theranostics 2017, 7, 1088–1099. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, L.B.; Ju, Y.S.; Haase, K.; Van Loo, P.; Martincorena, I.; Nik-Zainal, S.; Totoki, Y.; Fujimoto, A.; Nakagawa, H.; Shibata, T.; et al. Mutational signatures associated with tobacco smoking in human cancer. Science 2016, 354, 618–622. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gillison, M.L.; Akagi, K.; Xiao, W.; Jiang, B.; Pickard, R.K.L.; Li, J.; Swanson, B.J.; Agrawal, A.D.; Zucker, M.; Stache-Crain, B.; et al. Human papillomavirus and the landscape of secondary genetic alterations in oral cancers. Genome Res. 2019, 29, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Chang, J.; Tan, W.; Ling, Z.; Xi, R.; Shao, M.; Chen, M.; Luo, Y.; Zhao, Y.; Liu, Y.; Huang, X.; et al. Genomic analysis of oesophageal squamous-cell carcinoma identifies alcohol drinking-related mutation signature and genomic alterations. Nat. Commun. 2017, 8, 15290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Letouze, E.; Shinde, J.; Renault, V.; Couchy, G.; Blanc, J.F.; Tubacher, E.; Bayard, Q.; Bacq, D.; Meyer, V.; Semhoun, J.; et al. Mutational signatures reveal the dynamic interplay of risk factors and cellular processes during liver tumorigenesis. Nat. Commun. 2017, 8, 1315. [Google Scholar] [CrossRef] [Green Version]
- Pickering, C.R.; Zhang, J.; Neskey, D.M.; Zhao, M.; Jasser, S.A.; Wang, J.; Ward, A.; Tsai, C.J.; Ortega Alves, M.V.; Zhou, J.H.; et al. Squamous cell carcinoma of the oral tongue in young non-smokers is genomically similar to tumors in older smokers. Clin. Cancer Res. 2014, 20, 3842–3848. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edge, S.; Byrd, D.R.; Compton, C.C.; Fritz, A.G.; Greene, F.; Trotti, A. (Eds.) AJCC Cancer Staging Manual, 7th ed.; Springer: New York, NY, USA, 2010. [Google Scholar]
- Tamborero, D.; Gonzalez-Perez, A.; Lopez-Bigas, N. OncodriveCLUST: Exploiting the positional clustering of somatic mutations to identify cancer genes. Bioinformatics 2013, 29, 2238–2244. [Google Scholar] [CrossRef]
- Lawrence, M.S.; Stojanov, P.; Polak, P.; Kryukov, G.V.; Cibulskis, K.; Sivachenko, A.; Carter, S.L.; Stewart, C.; Mermel, C.H.; Roberts, S.A.; et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 2013, 499, 214–218. [Google Scholar] [CrossRef] [PubMed]
- DePristo, M.A.; Banks, E.; Poplin, R.; Garimella, K.V.; Maguire, J.R.; Hartl, C.; Philippakis, A.A.; del Angel, G.; Rivas, M.A.; Hanna, M.; et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 2011, 43, 491–498. [Google Scholar] [CrossRef]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar]
- Cingolani, P.; Platts, A.; Wang le, L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly Austin 2012, 6, 80–92. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.; Li, J.; Li, C.I.; Long, J.; Samuels, D.C.; Shyr, Y. The effect of strand bias in Illumina short-read sequencing data. BMC Genom. 2012, 13, 666. [Google Scholar] [CrossRef] [Green Version]
- Mouradov, D.; Sloggett, C.; Jorissen, R.N.; Love, C.G.; Li, S.; Burgess, A.W.; Arango, D.; Strausberg, R.L.; Buchanan, D.; Wormald, S.; et al. Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. Cancer Res. 2014, 74, 3238–3247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adzhubei, I.; Jordan, D.M.; Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 2013, 76, 7–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plagnol, V.; Curtis, J.; Epstein, M.; Mok, K.Y.; Stebbings, E.; Grigoriadou, S.; Wood, N.W.; Hambleton, S.; Burns, S.O.; Thrasher, A.J.; et al. A robust model for read count data in exome sequencing experiments and implications for copy number variant calling. Bioinformatic 2012, 28, 2747–2754. [Google Scholar] [CrossRef] [Green Version]
- Roca, I.; Gonzalez-Castro, L.; Fernandez, H.; Couce, M.L.; Fernandez-Marmiesse, A. Free-access copy-number variant detection tools for targeted next-generation sequencing data. Mutat. Res. 2019, 779, 114–125. [Google Scholar] [CrossRef]
- Rajagopalan, R.; Murrell, J.R.; Luo, M.; Conlin, L.K. A highly sensitive and specific workflow for detecting rare copy-number variants from exome sequencing data. Genome Med. 2020, 12, 14. [Google Scholar] [CrossRef] [Green Version]
- Derrien, T.; Estelle, J.; Marco Sola, S.; Knowles, D.G.; Raineri, E.; Guigo, R.; Ribeca, P. Fast computation and applications of genome mappability. PLoS ONE 2012, 7, e30377. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Zeileis, A.; Kleiber, C.; Jackman, S. Regression Models for Count Data in R. J. Stat. Softw. 2008, 27, 25. [Google Scholar] [CrossRef] [Green Version]
- DrinkWise Australia. Australian Drinking Habits: 2007 vs. 2017. Available online: https://drinkwise.org.au/our-work/australian-drinking-habits-2007-vs-2017 (accessed on 17 September 2020).
- Belobrov, S.; Cornall, A.M.; Young, R.J.; Koo, K.; Angel, C.; Wiesenfeld, D.; Rischin, D.; Garland, S.M.; McCullough, M. The role of human papillomavirus in p16-positive oral cancers. J. Oral Pathol. Med. 2018, 47, 18–24. [Google Scholar] [CrossRef] [Green Version]
- Kreimer, A.R.; Clifford, G.M.; Boyle, P.; Franceschi, S. Human papillomavirus types in head and neck squamous cell carcinomas worldwide: A systematic review. Cancer Epidemiol. Biomark. Prev. 2005, 14, 467–475. [Google Scholar] [CrossRef] [Green Version]
- Scheffner, M.; Werness, B.A.; Huibregtse, J.M.; Levine, A.J.; Howley, P.M. The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell 1990, 63, 1129–1136. [Google Scholar] [CrossRef]
- Tam, K.W.; Zhang, W.; Soh, J.; Stastny, V.; Chen, M.; Sun, H.; Thu, K.; Rios, J.J.; Yang, C.; Marconett, C.N.; et al. CDKN2A/p16 inactivation mechanisms and their relationship to smoke exposure and molecular features in non-small-cell lung cancer. J. Thorac. Oncol. 2013, 8, 1378–1388. [Google Scholar] [CrossRef] [Green Version]
- Sivarajah, S.; Kostiuk, M.; Lindsay, C.; Puttagunta, L.; O’Connell, D.A.; Harris, J.; Seikaly, H.; Biron, V.L. EGFR as a biomarker of smoking status and survival in oropharyngeal squamous cell carcinoma. J. Otolaryngol. Head Neck Surg. 2019, 48, 1. [Google Scholar] [CrossRef] [PubMed]
- Rudin, C.M.; Avila-Tang, E.; Harris, C.C.; Herman, J.G.; Hirsch, F.R.; Pao, W.; Schwartz, A.G.; Vahakangas, K.H.; Samet, J.M. Lung cancer in never smokers: Molecular profiles and therapeutic implications. Clin. Cancer Res. 2009, 15, 5646–5661. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sandulache, V.C.; Michikawa, C.; Kataria, P.; Gleber-Netto, F.O.; Bell, D.; Trivedi, S.; Rao, X.; Wang, J.; Zhao, M.; Jasser, S.; et al. High-Risk TP53 Mutations Are Associated with Extranodal Extension in Oral Cavity Squamous Cell Carcinoma. Clin. Cancer Res. 2018, 24, 1727–1733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, W.S.; Bindra, R.S.; Mo, A.; Hayman, T.; Husain, Z.; Contessa, J.N.; Gaffney, S.G.; Townsend, J.P.; Yu, J.B. CDKN2A Copy Number Loss Is an Independent Prognostic Factor in HPV-Negative Head and Neck Squamous Cell Carcinoma. Front. Oncol. 2018, 8, 95. [Google Scholar] [CrossRef] [Green Version]
- Michikawa, C.; Uzawa, N.; Sato, H.; Ohyama, Y.; Okada, N.; Amagasa, T. Epidermal growth factor receptor gene copy number aberration at the primary tumour is significantly associated with extracapsular spread in oral cancer. Br. J. Cancer 2011, 104, 850–855. [Google Scholar] [CrossRef] [Green Version]
- Bissinger, O.; Kolk, A.; Drecoll, E.; Straub, M.; Lutz, C.; Wolff, K.D.; Gotz, C. EGFR and Cortactin: Markers for potential double target therapy in oral squamous cell carcinoma. Exp. Ther. Med. 2017, 14, 4620–4626. [Google Scholar]
- Chen, I.H.; Chang, J.T.; Liao, C.T.; Wang, H.M.; Hsieh, L.L.; Cheng, A.J. Prognostic significance of EGFR and Her-2 in oral cavity cancer in betel quid prevalent area cancer prognosis. Br. J. Cancer 2003, 89, 681–686. [Google Scholar] [CrossRef] [Green Version]
- Huang, S.F.; Cheng, S.D.; Chien, H.T.; Liao, C.T.; Chen, I.H.; Wang, H.M.; Chuang, W.Y.; Wang, C.Y.; Hsieh, L.L. Relationship between epidermal growth factor receptor gene copy number and protein expression in oral cavity squamous cell carcinoma. Oral Oncol. 2012, 48, 67–72. [Google Scholar] [CrossRef]
- Chau, N.G.; Perez-Ordonez, B.; Zhang, K.; Pham, N.A.; Ho, J.; Zhang, T.; Ludkovski, O.; Wang, L.; Chen, E.X.; Tsao, M.S.; et al. The association between EGFR variant III, HPV, p16, c-MET, EGFR gene copy number and response to EGFR inhibitors in patients with recurrent or metastatic squamous cell carcinoma of the head and neck. Head Neck Oncol. 2011, 3, 11. [Google Scholar] [CrossRef] [Green Version]
- Beaty, B.T.; Moon, D.H.; Shen, C.J.; Amdur, R.J.; Weiss, J.; Grilley-Olson, J.; Patel, S.; Zanation, A.; Hackman, T.G.; Thorp, B.; et al. PIK3CA Mutation in HPV-Associated OPSCC Patients Receiving Deintensified Chemoradiation. J. Natl. Cancer Inst. 2020, 112, 855–858. [Google Scholar] [CrossRef] [PubMed]


| Characteristic | All Patients (n = 176) | |
|---|---|---|
| Gender | Female | 76 (43.2) |
| Male | 100 (56.8) | |
| Age | Median (range) | 66 (33–98) |
| Non-smoker | 86 (48.9) | |
| Non-drinker | 79 (44.9) | |
| NSND | 59 (33.5) | |
| T stage | 1 | 39 (22.2) |
| 2 | 66 (37.5) | |
| 3 | 14 (8.0) | |
| 4 | 57 (32.4) | |
| N stage | N0 | 115 (65.3) |
| N+ | 61 (34.7) | |
| AJCC stage | I | 32 (18.2) |
| II | 50 (28.4) | |
| III | 37 (21.0) | |
| IV | 57 (32.4) | |
| Perineural invasion | Present | 32 (18.2) |
| Absent | 144 (81.8) | |
| Lymphovascular invasion | Present | 18 (10.2) |
| Absent | 158 (89.8) | |
| Extracapsular spread | Present | 20 (11.4) |
| Absent | 156 (88.6) | |
| HPV status | Positive | 3 (1.7) |
| Negative | 173 (98.3) | |
| Radiotherapy | Yes | 110 (62.5) |
| No | 66 (37.5) | |
| Chemotherapy | Yes | 39 (22.2) |
| No | 137 (77.8) |
| Group 1 vs. Group 2 | Gene | Group 1 n (%) | Group 2 n (%) | OR (95% CI) | p |
|---|---|---|---|---|---|
| Male vs. Female | TP53 mut | 67/100 (67.0) | 39/76 (51.3) | 1.9 (1.0–3.7) | 0.043 * |
| CASP8 mut | 3/100 (3.0) | 11/76 (14.5) | 0.2 (0.0–0.7) | 0.009 * | |
| Smokers vs. Non-smokers | CDKN2A mut | 15/90 (16.7) | 27/86 (31.4) | 0.4 (0.2–0.9) | 0.033 * |
| Drinkers vs. Non-drinkers | CASP8 mut | 4/97 (4.1) | 10/79 (12.7) | 0.3 (0.1–1.1) | 0.050 * |
| LAMA4 mut | 1/97 (1.0) | 8/79 (10.1) | 0.1 (0.0–0.7) | 0.012 * | |
| NSND vs. SD | CDKN2A mut | 21/59 (35.6) | 21/117 (17.9) | 2.5 (1.2–5.5) | 0.014 * |
| EGFR amp | 10/59 (16.9) | 6/117 (5.1) | 3.7 (1.2–13.3) | 0.023 * | |
| BRCA2 del | 7/59 (11.9) | 2/117 (1.7) | 7.6 (1.4–77.8) | 0.007 * | |
| T3/4 tumours vs. T1/2 tumours | CDKN2A mut | 26/71 (36.6) | 16/105 (15.2) | 3.2 (1.5–7.1) | 0.002 * |
| CDKN2A del | 7/71 (9.9) | 0/105 (0) | Inf (2.3–Inf) | 0.001 * | |
| BRCA2 del | 8/71 (11.3) | 1/105 (1.0) | 13 (1.7–590.0) | 0.003 * | |
| LN+ vs. LN− | TP53 mut | 48/61 (78.7) | 58/115 (50.4) | 3.6 (1.7–8.1) | <0.001 * |
| BRCA2 del | 7/61 (11.5) | 2/115 (1.7) | 7.2 (1.3–73.5) | 0.009 * | |
| LVI+ vs. LVI− | TP53 mut | 15/18 (83.3) | 91/158 (57.6) | 3.7 (1.0–20.0) | 0.042 * |
| NCOR2 mut | 4/18 (22.2) | 8/156 (5.1) | 5.3 (1.0–22.9) | 0.023 * | |
| PNI+ vs. PNI− | EGFR amp | 7/32 (21.9) | 9/144 (6.2) | 4.2 (1.2–13.9) | 0.012 * |
| ECS+ vs. ECS− | CDKN2A mut | 9/20 (45.0) | 33/156 (21.2) | 3 (1.0–8.8) | 0.026 * |
| TP53 mut | 19/20 (95.0) | 87/156 (55.8) | 15 (2.3–633.0) | <0.001 * | |
| EGFR amp | 5/20 (25.0) | 11/156 (7.1) | 4.3 (1.0–16.0) | 0.022 * | |
| BRCA2 del | 4/20 (20.0) | 5/156 (3.2) | 7.4 (1.3–38.4) | 0.011 * |
| Univariate Analysis | Multivariate Analysis | |||||
|---|---|---|---|---|---|---|
| HR | 95% CI | p | AHR | 95% CI | p | |
| PIK3CA Mutation | 2.0 | 1.0–3.9 | 0.050 * | 1.4 | 0.7–2.9 | 0.303 |
| Male vs. female | 0.8 | 0.5–1.3 | 0.406 | 1.1 | 0.6–1.9 | 0.808 |
| Age (in decades) | 1.7 | 1.2–1.8 | <0.001 * | 1.6 | 1.3–2.0 | <0.001 * |
| NSND vs. SD | 1.7 | 1.0–2.8 | 0.050 * | 1.2 | 0.6–2.1 | 0.630 |
| T3/4 vs. T1/2 | 2.9 | 1.7–5.0 | <0.001 * | 2.6 | 1.5–4.4 | 0.001 * |
| LN+ vs. LN- | 2.3 | 1.4–3.8 | 0.001 * | 2.0 | 1.1–3.6 | 0.019 * |
| PNI+ vs. PNI- | 1.7 | 1.0–3.1 | 0.064 | 1.5 | 0.8–2.7 | 0.211 |
| LVI+ vs. LVI- | 2.0 | 1.0–4.0 | 0.064 | 1.4 | 0.6–3.1 | 0.443 |
| EGFR Amplification | 2.7 | 1.4–5.4 | 0.004 * | 1.8 | 0.9–3.6 | 0.118 |
| Male vs. female | 0.8 | 0.5–1.3 | 0.406 | 1.1 | 0.6–1.9 | 0.861 |
| Age (in decades) | 1.7 | 1.2–1.8 | <0.001 * | 1.6 | 1.3–2.1 | <0.001 * |
| NSND vs. SD | 1.7 | 1.0–2.8 | 0.050 * | 1.1 | 0.6–2.0 | 0.829 |
| T3/4 vs. T1/2 | 2.9 | 1.7–5.0 | <0.001 * | 2.4 | 1.4–4.2 | 0.001 * |
| LN+ vs. LN- | 2.3 | 1.4–3.8 | 0.001 * | 2.0 | 1.1–3.7 | 0.016 * |
| PNI+ vs. PNI- | 1.7 | 1.0–3.1 | 0.064 | 1.4 | 0.7–2.6 | 0.301 |
| LVI+ vs. LVI- | 2.0 | 1.0–4.0 | 0.064 | 1.5 | 0.6–3.3 | 0.360 |
| CDKN2A Deletion | 2.8 | 1.1–7.1 | 0.026 * | 1.8 | 0.6–5.0 | 0.261 |
| Male vs. female | 0.8 | 0.5–1.3 | 0.406 | 1.0 | 0.6–1.9 | 0.932 |
| Age (in decades) | 1.7 | 1.2–1.8 | <0.001 * | 1.6 | 1.3–2.0 | <0.001 * |
| NSND vs. SD | 1.7 | 1.0–2.8 | 0.050 * | 1.2 | 0.6–2.3 | 0.556 |
| T3/4 vs. T1/2 | 2.9 | 1.7–5.0 | <0.001 * | 2.3 | 1.3–4.1 | 0.004 * |
| LN+ vs. LN- | 2.3 | 1.4–3.8 | 0.001 * | 2.2 | 1.2–4.1 | 0.009 * |
| PNI+ vs. PNI- | 1.7 | 1.0–3.1 | 0.064 | 1.5 | 0.8–2.8 | 0.172 |
| LVI+ vs. LVI- | 2.0 | 1.0–4.0 | 0.064 | 1.2 | 0.5–2.8 | 0.654 |
| Alteration | SD (n = 117, 434 Mutations) | NSND, All (n = 59, 366 Mutations) | NSND, Low Mutation Group (n = 54, 233 Mutations) | NSND, High Mutation Group (n = 5, 133 Mutations) |
|---|---|---|---|---|
| C > A | 60 (13.8) | 39 (10.7) | 27 (11.6) | 12 (9.6) |
| C>G | 44 (10.1) | 42 (11.5) | 27 (11.6) | 15 (12.0) |
| C>T | 178 (41.0) | 164 (44.8) | 107 (45.9) | 57 (45.6) |
| T>A | 33 (7.6) | 12 (3.3) | 6 (2.6) | 6 (4.8) |
| T > C | 47 (10.8) | 61 (16.7) | 35 (15.0) | 26 (20.8) |
| T>G | 18 (4.1) | 16 (4.4) | 7 (3.0) | 9 (7.2) |
| Indel | 54 (12.4) | 32 (8.7) | 24 (10.3) | 8 (6.0) |
| Compared to SD | p = 0.010 * | p = 0.067 | p = 0.019 * | |
| Compared to NSND | p = 0.297 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koo, K.; Mouradov, D.; Angel, C.M.; Iseli, T.A.; Wiesenfeld, D.; McCullough, M.J.; Burgess, A.W.; Sieber, O.M. Genomic Signature of Oral Squamous Cell Carcinomas from Non-Smoking Non-Drinking Patients. Cancers 2021, 13, 1029. https://doi.org/10.3390/cancers13051029
Koo K, Mouradov D, Angel CM, Iseli TA, Wiesenfeld D, McCullough MJ, Burgess AW, Sieber OM. Genomic Signature of Oral Squamous Cell Carcinomas from Non-Smoking Non-Drinking Patients. Cancers. 2021; 13(5):1029. https://doi.org/10.3390/cancers13051029
Chicago/Turabian StyleKoo, Kendrick, Dmitri Mouradov, Christopher M. Angel, Tim A. Iseli, David Wiesenfeld, Michael J. McCullough, Antony W. Burgess, and Oliver M. Sieber. 2021. "Genomic Signature of Oral Squamous Cell Carcinomas from Non-Smoking Non-Drinking Patients" Cancers 13, no. 5: 1029. https://doi.org/10.3390/cancers13051029








