Upregulated GATA3/miR205-5p Axis Inhibits MFNG Transcription and Reduces the Malignancy of Triple-Negative Breast Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Cells and Tissue Specimens
2.2. Generation of Stable Cells That Overexpress MFNG and GATA3
2.3. Genomic DNA, RNA Isolation, shRNA, PCR, and RT-qPCR
2.4. Western Blot Assay
2.5. Luciferase Reporter Assay
2.6. Gene Expression and Correlation Analysis of the Public Database
2.7. Cell Growth Assay
2.8. Transwell Migration and Wound-Healing Assays
2.9. Colony Formation Assay
2.10. Immunohistochemistry
2.11. Chromatin Immunoprecipitation (ChIP) Assay
2.12. Nanoparticles-miR205-5p Preparation
2.13. Tumor Xenograft Model in Mice
2.14. Statistical Analysis
3. Results
3.1. MFNG Expression Was Significantly Increased in TNBC and Positively Associated with Poor Prognosis of TNBC Patients
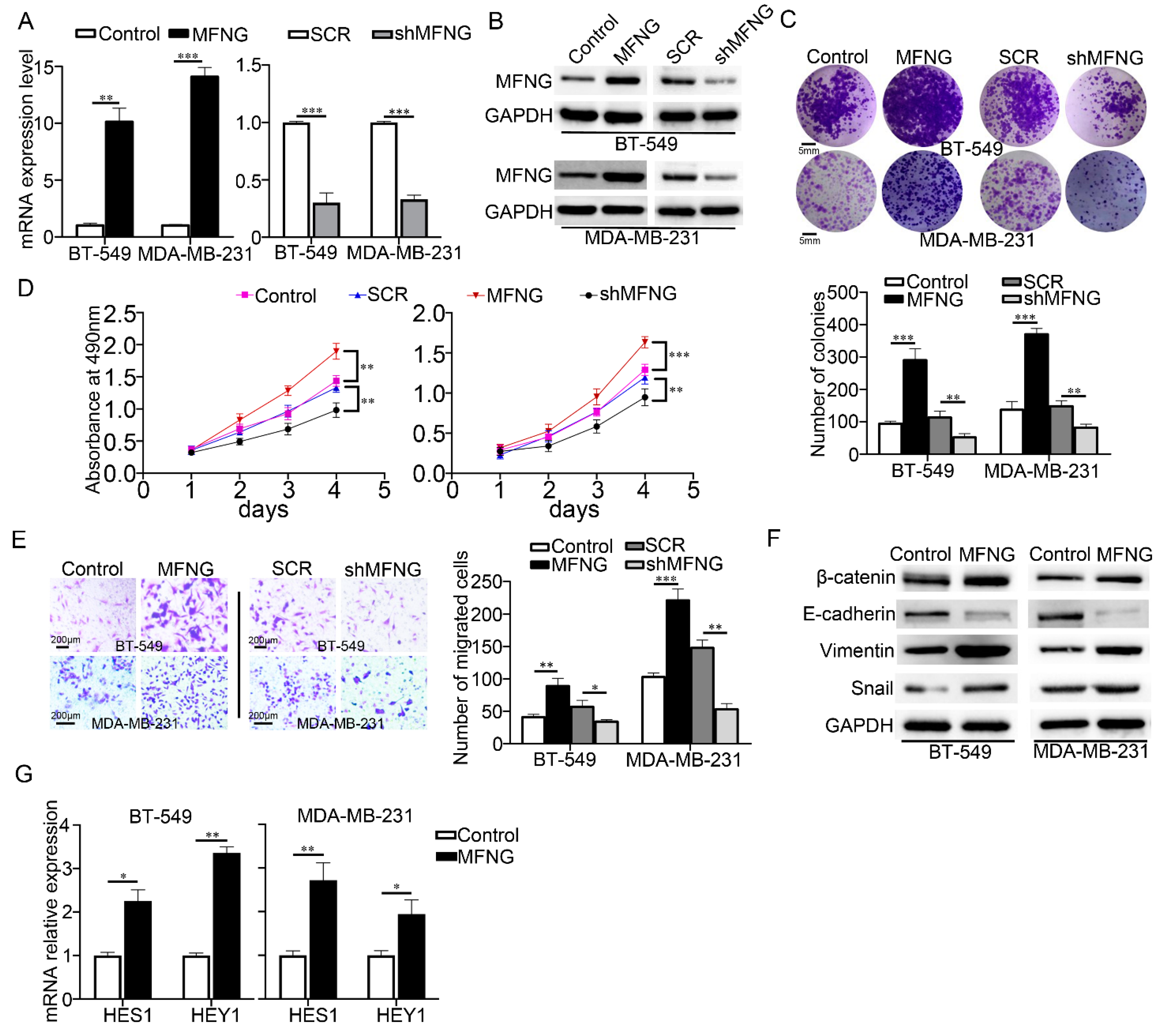
3.2. MFNG Expression Enhanced the Cell Growth and Migration of TNBC Cells
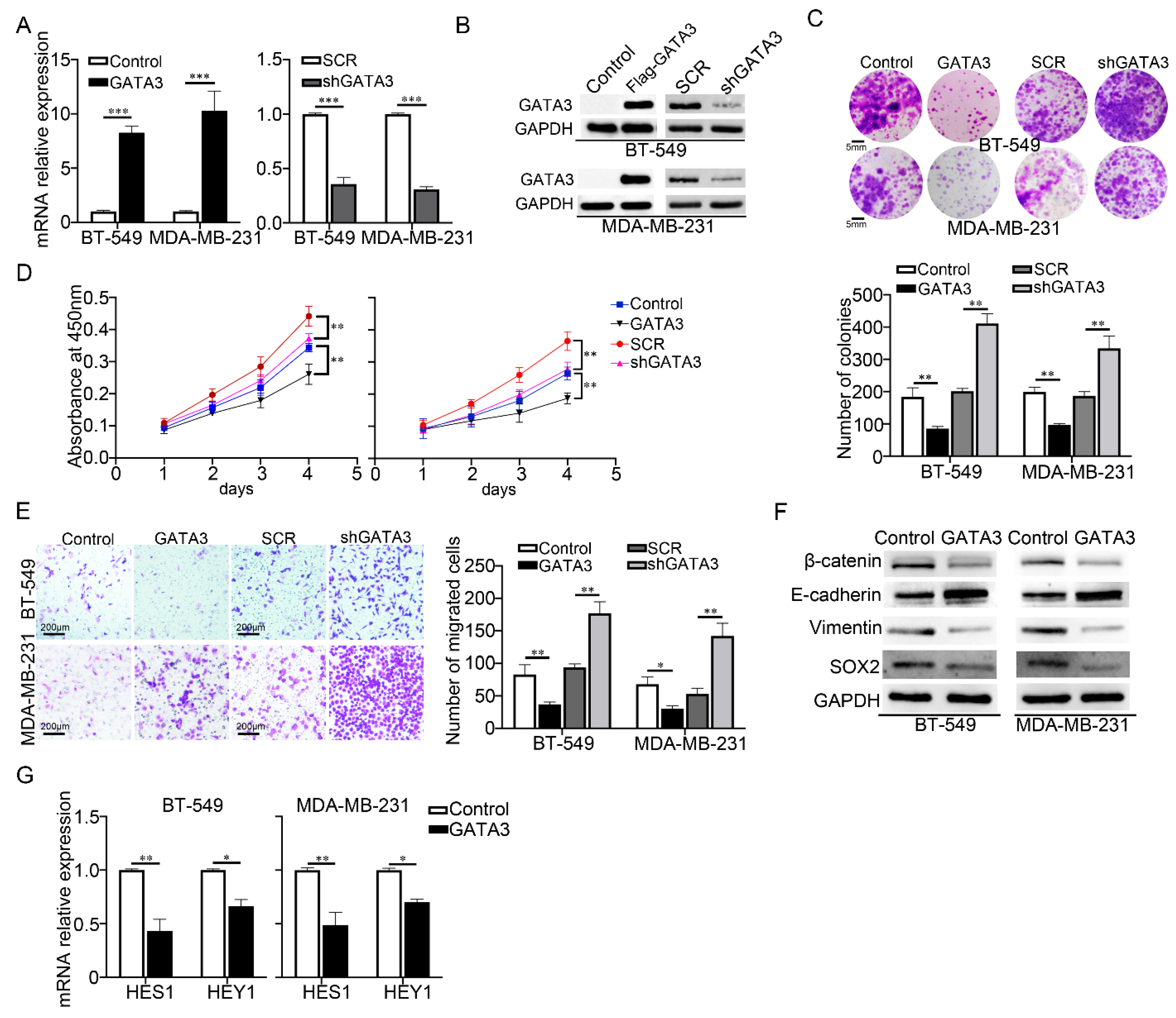
3.3. Decreased GATA3 Elevated the Malignancy of TNBC Cells and Promoted the Activity of Notch Signaling
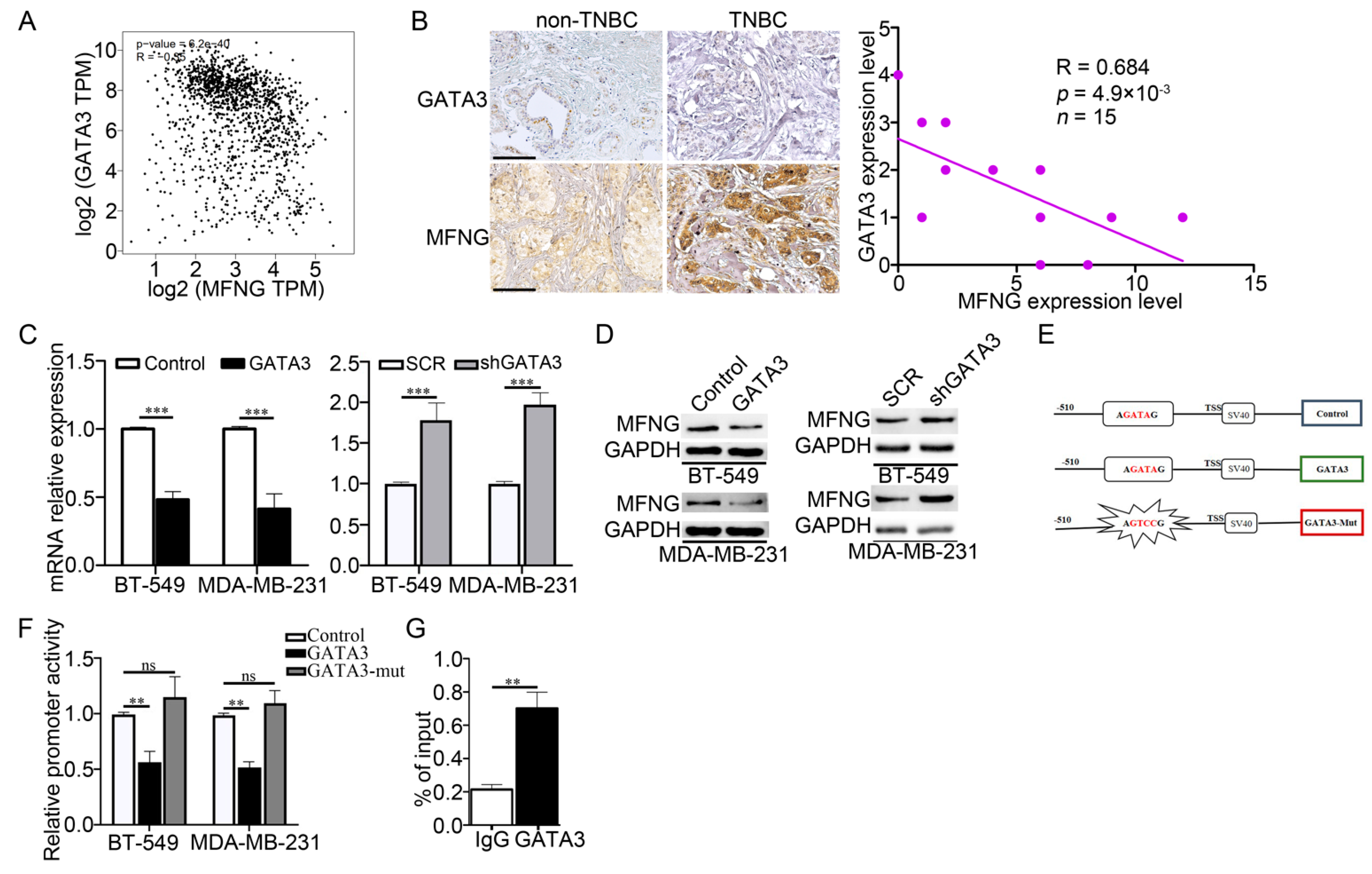
3.4. GATA3 Negatively Regulated MFNG Transcription by Abolishing Its Regulatory Activity in TNBC
3.5. miR205-5p Reduced the MFNG mRNA Level by Directly Binding to Its 3′ UTR Region and Inhibited the Malignancy of TNBC Cells
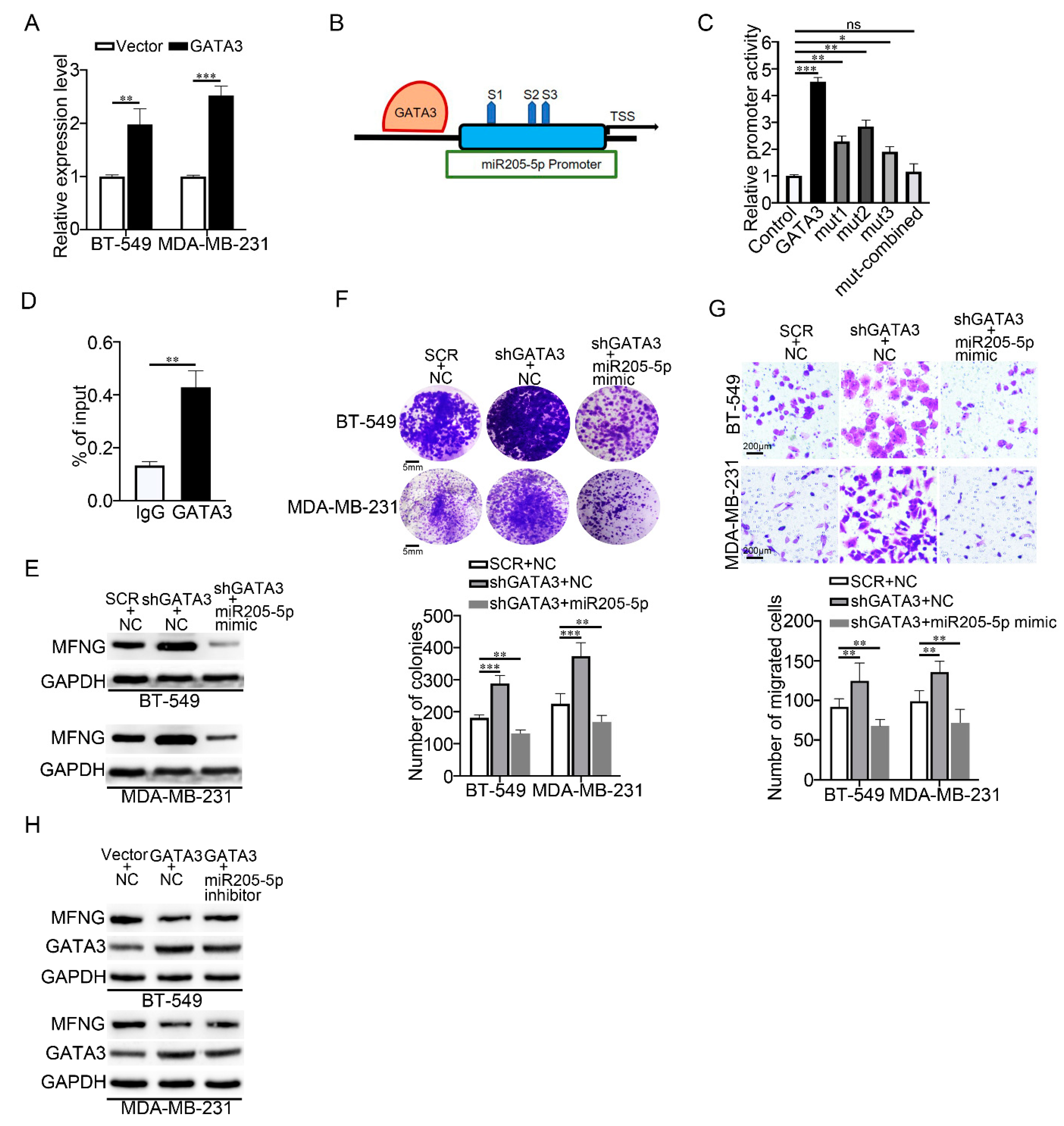
3.6. GATA3 Activated miR205-5p Transcription and Formed a Feed-Forward Loop with miR205-5p and MFNG in TNBC
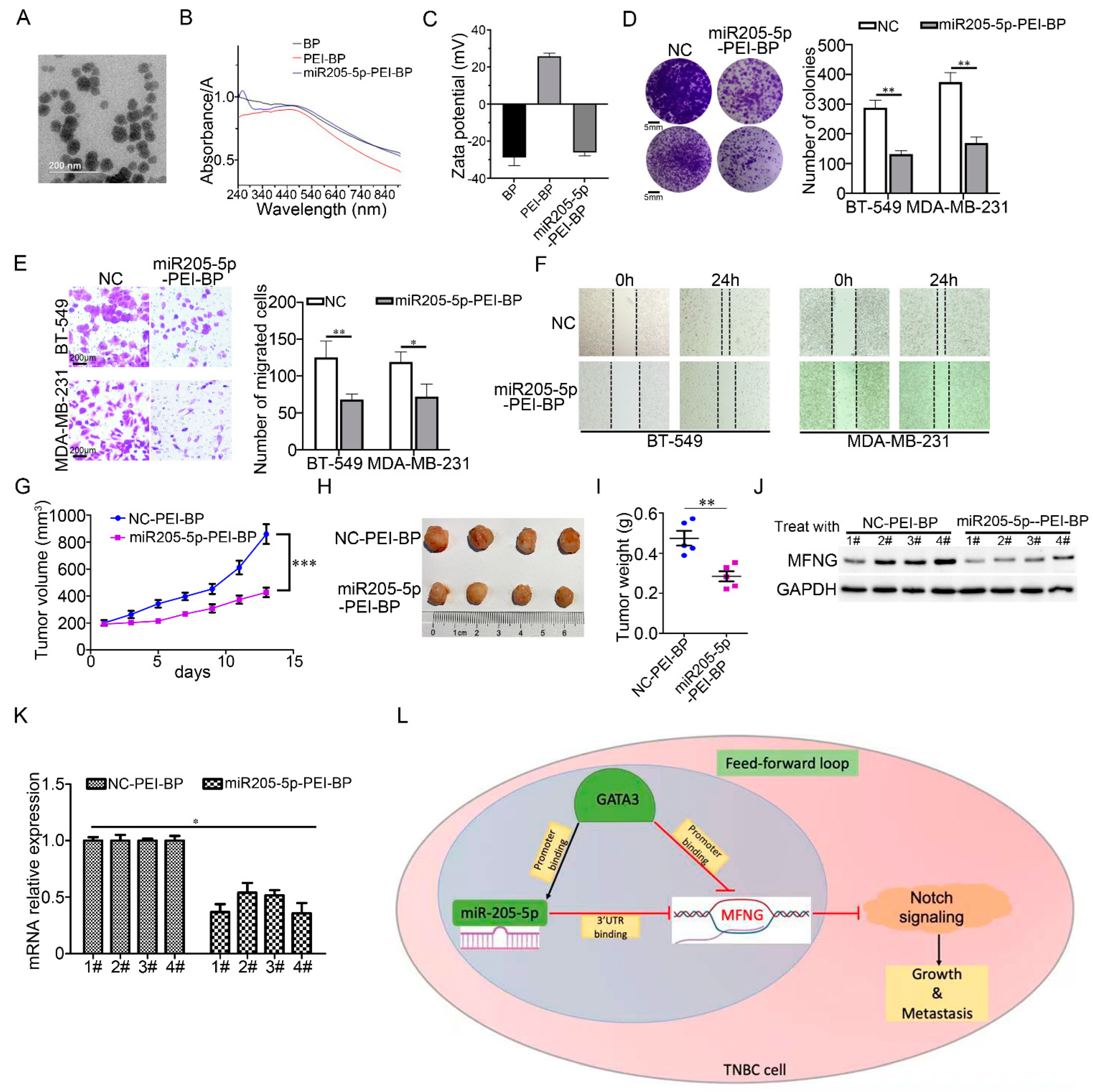
3.7. miR205-5p Exerted Effective Inhibition on The Malignancy of TNBC Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M. Molecular Stratification of Triple-Negative Breast Cancers. Oncologist 2011, 16 (Suppl. S1), 61–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Foulkes, W.D.; Smith, I.E.; Reis-Filho, J.S. Triple-Negative Breast Cancer. N. Engl. J. Med. 2010, 363, 1938–1948. [Google Scholar] [CrossRef] [Green Version]
- Brent, J.R.; Savjani, N.; Lewis, E.A.; Haigh, S.J.; Lewis, D.J.; O’Brien, P. Production of few-layer phosphorene by liquid exfoliation of black phosphorus. Chem. Commun. 2014, 50, 13338–13341. [Google Scholar] [CrossRef] [Green Version]
- Yang, K.; Feng, L.; Shi, X.; Liu, Z. Nano-graphene in biomedicine: Theranostic applications. Chem. Soc. Rev. 2013, 42, 530–547. [Google Scholar] [CrossRef]
- Choi, J.R.; Yong, K.W.; Choi, J.Y.; Nilghaz, A.; Lin, Y.; Xu, J.; Lu, X. Black Phosphorus and its Biomedical Applications. Theranostics 2018, 8, 1005–1026. [Google Scholar] [CrossRef]
- Miao, K.; Lei, J.H.; Valecha, M.V.; Zhang, A.; Xu, J.; Wang, L.; Lyu, X.; Chen, S.; Miao, Z.; Zhang, X.; et al. NOTCH1 activation compensates BRCA1 deficiency and promotes triple-negative breast cancer formation. Nat. Commun. 2020, 11, 3256. [Google Scholar] [CrossRef]
- Johnston, S.; Rauskolb, C.; Wilson, R.; Prabhakaran, B.; Irvine, K.; Vogt, T. A family of mammalian Fringe genes implicated in boundary determination and the Notch pathway. Development 1997, 124, 2245–2254. [Google Scholar] [CrossRef]
- Rampal, R.; Li, A.S.; Moloney, D.J.; Georgiou, S.A.; Luther, K.B.; Nita-Lazar, A.; Haltiwanger, R.S. Lunatic Fringe, Manic Fringe, and Radical Fringe Recognize Similar Specificity Determinants in O-Fucosylated Epidermal Growth Factor-like Repeats. J. Biol. Chem. 2005, 280, 42454–42463. [Google Scholar] [CrossRef] [Green Version]
- Yi, F.; Amarasinghe, B.; Dang, T.P. Manic fringe inhibits tumor growth by suppressing Notch3 degradation in lung cancer. Am. J. Cancer Res. 2013, 3, 490–499. [Google Scholar] [PubMed]
- Cheng, W.K.; Kaur, G.; Sjöberg, E.; Frödin, M.; Egevad, L.; Harmenberg, U.; Li, J.-L.; Oon, C.E. Nuclear and stromal expression of Manic fringe in renal cell carcinoma. Exp. Mol. Pathol. 2021, 122, 104667. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Chung, W.-C.; Wu, G.; Egan, S.E.; Miele, L.; Xu, K. Manic Fringe Promotes a Claudin-Low Breast Cancer Phenotype through Notch-Mediated PIK3CG Induction. Cancer Res. 2015, 75, 1936–1943. [Google Scholar] [CrossRef] [Green Version]
- Kouros-Mehr, H.; Slorach, E.M.; Sternlicht, M.D.; Werb, Z. GATA-3 Maintains the Differentiation of the Luminal Cell Fate in the Mammary Gland. Cell 2006, 127, 1041–1055. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, X.; Chen, Y.; Du, E.; Yang, K.; Zhang, Z.; Qi, S.; Xu, Y. GATA3-driven expression of miR-503 inhibits prostate cancer progression by repressing ZNF217 expression. Cell. Signal. 2016, 28, 1216–1224. [Google Scholar] [CrossRef] [PubMed]
- Shahi, P.; Wang, C.-Y.; Chou, J.; Hagerling, C.; Velozo, H.G.; Ruderisch, A.; Yu, Y.; Lai, M.-D.; Werb, Z. GATA3 targets semaphorin 3B in mammary epithelial cells to suppress breast cancer progression and metastasis. Oncogene 2017, 36, 5567–5575. [Google Scholar] [CrossRef] [Green Version]
- Krings, G.; Nystrom, M.; Mehdi, I.; Vohra, P.; Chen, Y.-Y. Diagnostic utility and sensitivities of GATA3 antibodies in triple-negative breast cancer. Hum. Pathol. 2014, 45, 2225–2232. [Google Scholar] [CrossRef]
- Neve, R.M.; Chin, K.; Fridlyand, J.; Yeh, J.; Baehner, F.L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.-P.; Tong, F.; et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef] [Green Version]
- Prat, A.; Parker, J.S.; Karginova, O.; Fan, C.; Livasy, C.; Herschkowitz, J.I.; He, X.; Perou, C.M. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 2010, 12, R68. [Google Scholar] [CrossRef] [Green Version]
- Solomon, E.R.; Caldwell, K.K.; Allan, A.M. A novel method for the normalization of ChIP-qPCR data. MethodsX 2021, 8, 101504. [Google Scholar] [CrossRef]
- Jiang, Y.-Z.; Yu, K.-D.; Zuo, W.-J.; Peng, W.-T.; Shao, Z.-M. GATA3 mutations define a unique subtype of luminal-like breast cancer with improved survival. Cancer 2014, 120, 1329–1337. [Google Scholar] [CrossRef]
- Abba, M.C.; Nunez, M.I.; Colussi, A.G.; Croce, M.V.; Segal-Eiras, A.; Aldaz, C.M. GATA3 protein as a MUC1 transcriptional regulator in breast cancer cells. Breast Cancer Res. 2006, 8, R64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, W.; Huang, W.; Yang, Y.; Qiu, R.; Zeng, Y.; Hou, Y.; Sun, G.; Shi, H.; Leng, S.; Feng, D.; et al. GATA3 recruits UTX for gene transcriptional activation to suppress metastasis of breast cancer. Cell Death Dis. 2019, 10, 832. [Google Scholar] [CrossRef] [PubMed]
- Tkocz, D.; Crawford, N.T.; Buckley, N.E.; Berry, F.B.; Kennedy, R.D.; Gorski, J.J.; Harkin, D.P.; Mullan, P.B. BRCA1 and GATA3 corepress FOXC1 to inhibit the pathogenesis of basal-like breast cancers. Oncogene 2011, 31, 3667–3678. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, E.-H.; Zhou, Q.; Liu, K.-S.; Wei, W.; Wang, C.-M.; Liu, X.-F.; Lu, C.; Ma, D.-Y. Screening miRNAs related to different subtypes of breast cancer with miRNAs microarray. Eur. Rev. Med. Pharmacol. Sci. 2014, 18, 2783–2788. [Google Scholar] [PubMed]
- Cava, C.; Colaprico, A.; Bertoli, G.; Bontempi, G.; Mauri, G.; Castiglioni, I. How interacting pathways are regulated by miRNAs in breast cancer subtypes. BMC Bioinform. 2016, 17, 111–133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plantamura, I.; Cataldo, A.; Cosentino, G.; Iorio, M.V. miR-205 in Breast Cancer: State of the Art. Int. J. Mol. Sci. 2020, 22, 27. [Google Scholar] [CrossRef]
- Chou, J.; Lin, J.H.; Brenot, A.; Kim, J.-W.; Provot, S.; Werb, Z. GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat. Cell Biol. 2013, 15, 201–213. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Zhang, Q.; Li, D.; Ching, K.; Zhang, C.; Zheng, X.; Ozeck, M.; Shi, S.; Li, X.; Wang, H.; et al. PEST Domain Mutations in Notch Receptors Comprise an Oncogenic Driver Segment in Triple-Negative Breast Cancer Sensitive to a γ-Secretase Inhibitor. Clin. Cancer Res. 2015, 21, 1487–1496. [Google Scholar] [CrossRef] [Green Version]
- Veeraraghavalu, K.; Pett, M.; Kumar, R.V.; Nair, P.; Rangarajan, A.; Stanley, M.A.; Krishna, S. Papillomavirus-Mediated Neoplastic Progression Is Associated with Reciprocal Changes in Jagged1 and Manic Fringe Expression Linked to Notch Activation. J. Virol. 2004, 78, 8687–8700. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Evans, T.; Reitman, M.; Felsenfeld, G. An erythrocyte-specific DNA-binding factor recognizes a regulatory sequence common to all chicken globin genes. Proc. Natl. Acad. Sci. USA 1988, 85, 5976–5980. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asch-Kendrick, R.; Cimino-Mathews, A. The role of GATA3 in breast carcinomas: A review. Hum. Pathol. 2015, 48, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Yan, W.; Cao, Q.J.; Arenas, R.B.; Bentley, B.; Shao, R. GATA3 Inhibits Breast Cancer Metastasis through the Reversal of Epithelial-Mesenchymal Transition. J. Biol. Chem. 2010, 285, 14042–14051. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takaku, M.; Grimm, S.A.; De Kumar, B.; Bennett, B.D.; Wade, P.A. Cancer-specific mutation of GATA3 disrupts the transcriptional regulatory network governed by Estrogen Receptor alpha, FOXA1 and GATA3. Nucleic Acids Res. 2020, 48, 4756–4768. [Google Scholar] [CrossRef] [PubMed]
- Sereesongsaeng, N.; McDowell, S.H.; Burrows, J.F.; Scott, C.J.; Burden, R.E. Cathepsin V suppresses GATA3 protein expression in luminal A breast cancer. Breast Cancer Res. 2020, 22, 139. [Google Scholar] [CrossRef]
- Farmer, P.; Bonnefoi, H.; Becette, V.; Tubiana-Hulin, M.; Fumoleau, P.; Larsimont, D.; MacGrogan, G.; Bergh, J.; Cameron, D.; Goldstein, D.; et al. Identification of molecular apocrine breast tumours by microarray analysis. Oncogene 2005, 24, 4660–4671. [Google Scholar] [CrossRef] [Green Version]
- Mehra, R.; Varambally, S.; Ding, L.; Shen, R.; Sabel, M.S.; Ghosh, D.; Chinnaiyan, A.M.; Kleer, C.G. Identification of GATA3 as a Breast Cancer Prognostic Marker by Global Gene Expression Meta-analysis. Cancer Res. 2005, 65, 11259–11264. [Google Scholar] [CrossRef] [Green Version]
- Cheng, C.-W.; Liao, W.-L.; Chen, P.-M.; Yu, J.-C.; Shiau, H.-P.; Hsieh, Y.-H.; Lee, H.-J.; Cheng, Y.-C.; Wu, P.-E.; Shen, C.-Y. MiR-139 Modulates Cancer Stem Cell Function of Human Breast Cancer through Targeting CXCR4. Cancers 2021, 13, 2582. [Google Scholar] [CrossRef]
- Deshmukh, S.K.; Srivastava, S.K.; Zubair, H.; Khan, M.A.; Singh, A.P.; Singh, S. Resistin Induces LIN28A-Mediated Let-7a Repression in Breast Cancer Cells Leading to IL-6 and STAT3 Upregulation. Cancers 2021, 13, 4498. [Google Scholar] [CrossRef]
- Shen, Y.; Xu, Y.; Huang, L.; Chi, Y.; Meng, L. MiR-205 suppressed the malignant behaviors of breast cancer cells by targeting CLDN11 via modulation of the epithelial-to-mesenchymal transition. Aging 2021, 13, 13073–13086. [Google Scholar] [CrossRef]
- Lin, L.-F.; Li, Y.-T.; Han, H.; Lin, S.-G. MicroRNA-205-5p targets the HOXD9-Snail1 axis to inhibit triple negative breast cancer cell proliferation and chemoresistance. Aging 2021, 13, 3945–3956. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Kang, F.-B.; Wang, J.; Yang, C.; He, D.-W. Downregulation of miR-205 contributes to epithelial–mesenchymal transition and invasion in triple-negative breast cancer by targeting HMGB1–RAGE signaling pathway. Anti-Cancer Drugs 2019, 30, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Dadwal, A.; Baldi, A.; Kumar Narang, R. Nanoparticles as carriers for drug delivery in cancer. Artif. Cells Nanomed. Biotechnol. 2018, 46, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Bobo, D.; Robinson, K.J.; Islam, J.; Thurecht, K.J.; Corrie, S.R. Nanoparticle-Based Medicines: A Review of FDA-Approved Materials and Clinical Trials to Date. Pharm. Res. 2016, 33, 2373–2387. [Google Scholar] [CrossRef]
- Cao, J.; Qi, J.; Lin, X.; Xiong, Y.; He, F.; Deng, W.; Liu, G. Biomimetic Black Phosphorus Nanosheet-Based Drug Delivery System for Targeted Photothermal-Chemo Cancer Therapy. Front. Bioeng. Biotechnol. 2021, 9, 707208. [Google Scholar] [CrossRef]
- Qin, G.; Mallik, S.; Mitra, R.; Li, A.; Jia, P.; Eischen, C.M.; Zhao, Z. MicroRNA and transcription factor co-regulatory networks and subtype classification of seminoma and non-seminoma in testicular germ cell tumors. Sci. Rep. 2020, 10, 852. [Google Scholar] [CrossRef] [Green Version]
- Afshar, A.S.; Xu, J.; Goutsias, J. Integrative Identification of Deregulated MiRNA/TF-Mediated Gene Regulatory Loops and Networks in Prostate Cancer. PLoS ONE 2014, 9, e100806. [Google Scholar] [CrossRef]
- Zhao, M.; Sun, J.; Zhao, Z. Synergetic regulatory networks mediated by oncogene-driven microRNAs and transcription factors in serous ovarian cancer. Mol. BioSyst. 2013, 9, 3187–3198. [Google Scholar] [CrossRef] [Green Version]
- Piovan, C.; Palmieri, D.; Di Leva, G.; Braccioli, L.; Casalini, P.; Nuovo, G.; Tortoreto, M.; Sasso, M.; Plantamura, I.; Triulzi, T.; et al. Oncosuppressive role of p53-induced miR-205 in triple negative breast cancer. Mol. Oncol. 2012, 6, 458–472. [Google Scholar] [CrossRef]
- Radojicic, J.; Zaravinos, A.; Vrekoussis, T.; Kafousi, M.; Spandidos, D.A.; Stathopoulos, E.N. MicroRNA expression analysis in triple-negative (ER, PR and Her2/neu) breast cancer. Cell Cycle 2011, 10, 507–517. [Google Scholar] [CrossRef] [Green Version]
- Emmanuel, N.; Lofgren, K.A.; Peterson, E.A.; Meier, D.R.; Jung, E.H.; Kenny, P.A. Mutant GATA3 Actively Promotes the Growth of Normal and Malignant Mammary Cells. Anticancer Res. 2018, 38, 4435–4441. [Google Scholar] [CrossRef] [PubMed]
- Gustin, J.P.; Miller, J.; Farag, M.; Rosen, D.M.; Thomas, M.; Scharpf, R.B.; Lauring, J. GATA3 frameshift mutation promotes tumor growth in human luminal breast cancer cells and induces transcriptional changes seen in primary GATA3 mutant breast cancers. Oncotarget 2017, 8, 103415–103427. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Afzaljavan, F.; Sadr, A.S.; Savas, S.; Pasdar, A. GATA3 somatic mutations are associated with clinicopathological features and expression profile in TCGA breast cancer patients. Sci. Rep. 2021, 11, 1679. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mugisha, S.; Di, X.; Wen, D.; Zhao, Y.; Wu, X.; Zhang, S.; Jiang, H. Upregulated GATA3/miR205-5p Axis Inhibits MFNG Transcription and Reduces the Malignancy of Triple-Negative Breast Cancer. Cancers 2022, 14, 3057. https://doi.org/10.3390/cancers14133057
Mugisha S, Di X, Wen D, Zhao Y, Wu X, Zhang S, Jiang H. Upregulated GATA3/miR205-5p Axis Inhibits MFNG Transcription and Reduces the Malignancy of Triple-Negative Breast Cancer. Cancers. 2022; 14(13):3057. https://doi.org/10.3390/cancers14133057
Chicago/Turabian StyleMugisha, Samson, Xiaotang Di, Doudou Wen, Yuetao Zhao, Xusheng Wu, Shubing Zhang, and Hao Jiang. 2022. "Upregulated GATA3/miR205-5p Axis Inhibits MFNG Transcription and Reduces the Malignancy of Triple-Negative Breast Cancer" Cancers 14, no. 13: 3057. https://doi.org/10.3390/cancers14133057





