Methyladenosine Modification in RNAs: From Regulatory Roles to Therapeutic Implications in Cancer
Abstract
:Simple Summary
Abstract
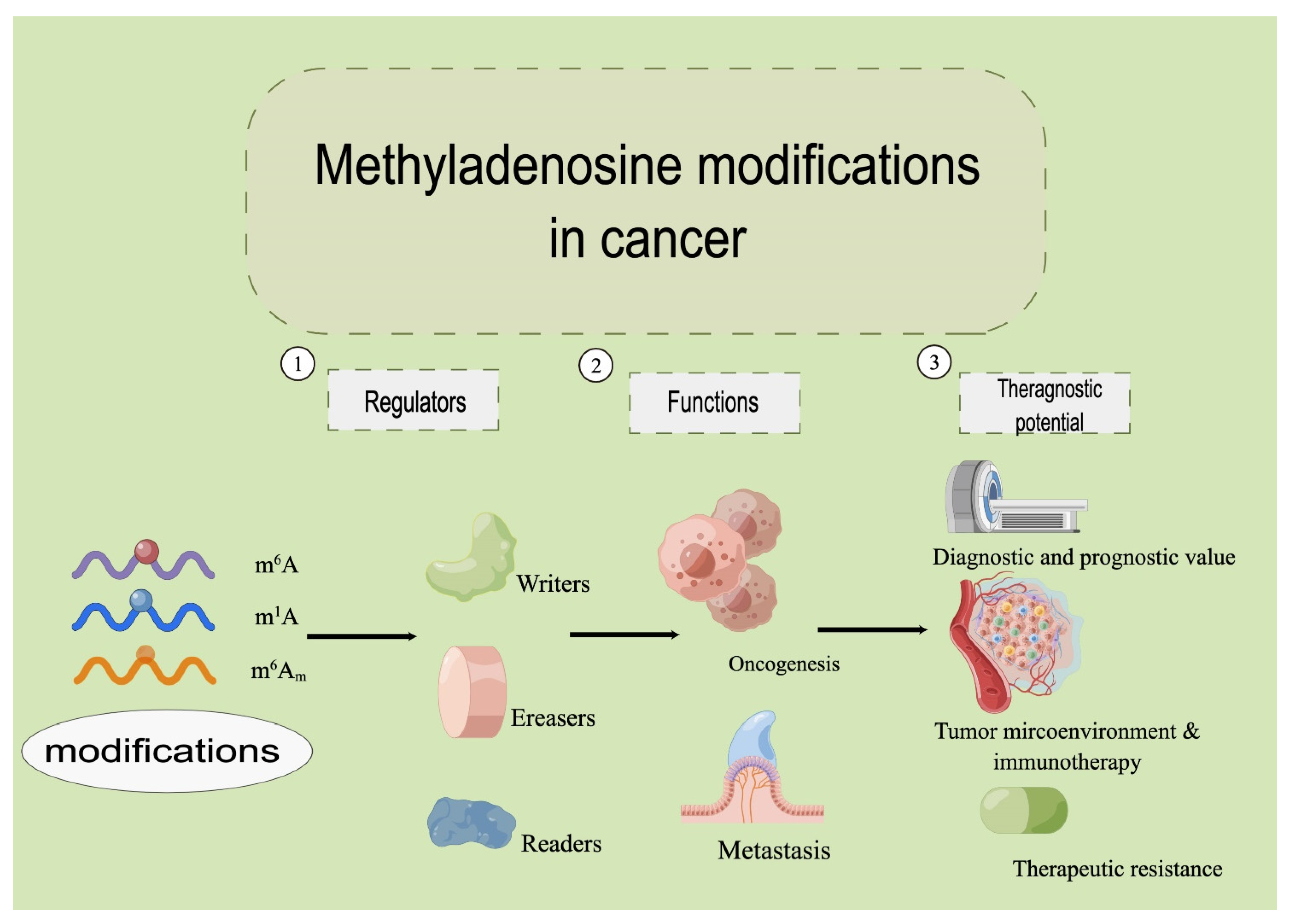
1. Introduction
2. Methyladenosine Modifications in Eukaryotic RNAs
3. Writers, Erasers, and Readers in the Coregulation of the Dynamic Adjustment of Methyladenosine Modification
3.1. Writers
3.2. Erasers
3.3. Readers
4. Methyladenosine Modification: An Inseparable Part of Cancers
4.1. Writers in Oncogenesis
4.2. Erasers in Oncogenesis
4.3. Readers in Oncogenesis
4.4. Methyladenosine Modifications in Metastasis
5. Theragnostic Potential of Methyladenosine Modifications in Cancer
5.1. Acting as Diagnostic Markers and Prognostic Predictors
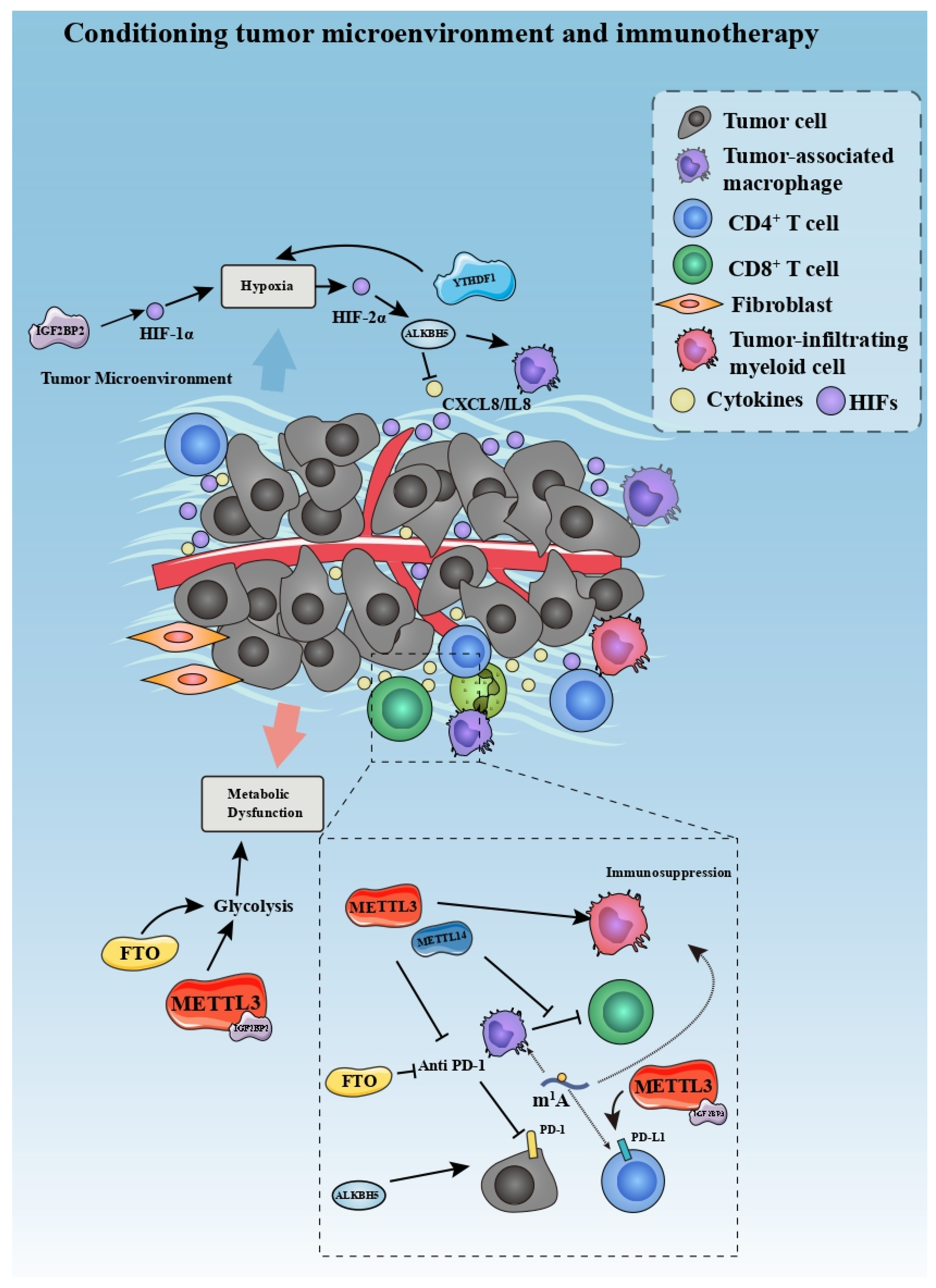
5.2. Conditioning Tumor Microenvironment and Immunotherapy
5.3. Modulating Therapeutic Resistance and Self-Renewal of Cancers
6. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Hou, J.; Zhou, Y.; Li, Z.; Cao, X. The RNA helicase DDX46 inhibits innate immunity by entrapping mA-demethylated antiviral transcripts in the nucleus. Nat. Immunol. 2017, 18, 1094–1103. [Google Scholar] [CrossRef] [PubMed]
- Bartosovic, M.; Molares, H.C.; Gregorova, P.; Hrossova, D.; Kudla, G.; Vanacova, S. N6-methyladenosine demethylase FTO targets pre-mRNAs and regulates alternative splicing and 3’-end processing. Nucleic Acids Res. 2017, 45, 11356–11370. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.S.; Roundtree, I.A.; He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 2017, 18, 31–42. [Google Scholar] [CrossRef] [Green Version]
- Oerum, S.; Meynier, V.; Catala, M.; Tisné, C. A comprehensive review of m6A/m6Am RNA methyltransferase structures. Nucleic Acids Res. 2021, 49, 7239–7255. [Google Scholar] [CrossRef]
- Safra, M.; Sas-Chen, A.; Nir, R.; Winkler, R.; Nachshon, A.; Bar-Yaacov, D.; Matthias, E.; Walter, R.; Noam, S.-G.; Schraga, S. The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature 2017, 551, 251–255. [Google Scholar] [CrossRef]
- Rong, D.; Sun, G.; Wu, F.; Cheng, Y.; Sun, G.; Jiang, W.; Li, X.; Zhong, Y.; Wu, L.; Zhang, C.; et al. Epigenetics: Roles and therapeutic implications of non-coding RNA modifications in human cancers. Mol. Ther. Nucleic Acids 2021, 25, 67–82. [Google Scholar] [CrossRef]
- Jiang, X.; Liu, B.; Nie, Z.; Duan, L.; Xiong, Q.; Jin, Z.; Yang, C.; Chen, Y. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target. Ther. 2021, 6, 74. [Google Scholar] [CrossRef]
- Huang, H.; Weng, H.; Chen, J. mA Modification in Coding and Non-coding RNAs: Roles and Therapeutic Implications in Cancer. Cancer Cell 2020, 37, 270–288. [Google Scholar] [CrossRef]
- Meyer, K.D.; Saletore, Y.; Zumbo, P.; Elemento, O.; Mason, C.E.; Jaffrey, S.R. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 2012, 149, 1635–1646. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Geng, X.; Li, Q.; Xu, J.; Tan, Y.; Xiao, M.; Song, J.; Liu, F.; Fang, C.; Wang, H. m6A modification in RNA: Biogenesis, functions and roles in gliomas. J. Exp. Clin. Cancer Res. 2020, 39, 192. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Liu, S.; Peng, Y.; Ge, R.; Su, R.; Senevirathne, C.; Harada, B.T.; Dai, Q.; Wei, J.; Zhang, L.; et al. m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome. Nat. Biotechnol. 2022, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Perry, R.P.; Kelley, D.E. Existence of methylated messenger RNA in mouse L cells. Cell 1974, 1, 37–42. [Google Scholar] [CrossRef]
- Desrosiers, R.; Friderici, K.; Rottman, F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc. Natl. Acad. Sci. USA 1974, 71, 3971–3975. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dominissini, D.; Moshitch-Moshkovitz, S.; Schwartz, S.; Salmon-Divon, M.; Ungar, L.; Osenberg, S.; Cesarkas, K.; Jacob-Hirsch, J.; Amariglio, N.; Kupiec, M.; et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 2012, 485, 201–206. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Campos, M.A.; Edelheit, S.; Toth, U.; Safra, M.; Shachar, R.; Viukov, S.; Winkler, R.; Nir, R.; Lasman, L.; Brandis, A.; et al. Deciphering the "mA Code" via Antibody-Independent Quantitative Profiling. Cell 2019, 178, 731–747. [Google Scholar] [CrossRef]
- Liu, N.; Dai, Q.; Zheng, G.; He, C.; Parisien, M.; Pan, T. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 2015, 518, 560–564. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, W.; Arnold, H.H.; Kersten, H. Biosynthetic pathway of ribothymidine in B. subtilis and M. lysodeikticus involving different coenzymes for transfer RNA and ribosomal RNA. Nucleic Acids Res. 1975, 2, 1043–1051. [Google Scholar] [CrossRef] [Green Version]
- Linder, B.; Grozhik, A.V.; Olarerin-George, A.O.; Meydan, C.; Mason, C.E.; Jaffrey, S.R. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 2015, 12, 767–772. [Google Scholar] [CrossRef]
- Zheng, G.; Dahl, J.A.; Niu, Y.; Fedorcsak, P.; Huang, C.-M.; Li, C.J.; Vågbø, C.B.; Shi, Y.; Wang, W.-L.; Song, S.-H.; et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 2013, 49, 18–29. [Google Scholar] [CrossRef] [Green Version]
- Kim, G.-W.; Siddiqui, A. N6-methyladenosine modification of HCV RNA genome regulates cap-independent IRES-mediated translation via YTHDC2 recognition. Proc. Natl. Acad. Sci. USA 2021, 118, e2022024118. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Li, X.; Yi, C. N-methyladenosine methylome in messenger RNA and non-coding RNA. Curr. Opin. Chem. Biol. 2018, 45, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.; Yi, C.; Jian, X.; Zheng, G.; He, C. Structure determination of DNA methylation lesions N1-meA and N3-meC in duplex DNA using a cross-linked protein-DNA system. Nucleic Acids Res. 2010, 38, 4415–4425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, G.; Fu, Y.; Zhao, X.; Dai, Q.; Zheng, G.; Yang, Y.; Yi, C.; Lindahl, T.; Pan, T.; Yang, Y.-G.; et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 2011, 7, 885–887. [Google Scholar] [CrossRef]
- Zaccara, S.; Ries, R.J.; Jaffrey, S.R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 2019, 20, 608–624. [Google Scholar] [CrossRef]
- Wang, P.; Doxtader, K.A.; Nam, Y. Structural Basis for Cooperative Function of Mettl3 and Mettl14 Methyltransferases. Mol. Cell 2016, 63, 306–317. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Feng, J.; Xue, Y.; Guan, Z.; Zhang, D.; Liu, Z.; Gong, Z.; Huang, J.; Tang, C.; Zou, T.; et al. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature 2016, 534, 575–578. [Google Scholar] [CrossRef]
- Liu, J.; Yue, Y.; Han, D.; Wang, X.; Fu, Y.; Zhang, L.; Jia, G.; Yu, M.; Lu, Z.; Deng, X.; et al. A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 2014, 10, 93–95. [Google Scholar] [CrossRef] [Green Version]
- Schwartz, S.; Mumbach, M.R.; Jovanovic, M.; Wang, T.; Maciag, K.; Bushkin, G.G.; Mertins, P.; Ter-Ovanesyan, D.; Habib, N.; Cacchiarelli, D.; et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep. 2014, 8, 284–296. [Google Scholar] [CrossRef] [Green Version]
- Akichika, S.; Hirano, S.; Shichino, Y.; Suzuki, T.; Nishimasu, H.; Ishitani, R.; Sugita, A.; Hirose, Y.; Iwasaki, S.; Nureki, O.; et al. Cap-specific terminal -methylation of RNA by an RNA polymerase II-associated methyltransferase. Science 2019, 363. [Google Scholar] [CrossRef]
- Zhang, X.; Wei, L.-H.; Wang, Y.; Xiao, Y.; Liu, J.; Zhang, W.; Yan, N.; Amu, G.; Tang, X.; Zhang, L.; et al. Structural insights into FTO’s catalytic mechanism for the demethylation of multiple RNA substrates. Proc. Natl. Acad. Sci. USA 2019, 116, 2919–2924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mauer, J.; Luo, X.; Blanjoie, A.; Jiao, X.; Grozhik, A.V.; Patil, D.P.; Linder, B.; Pickering, B.F.; Vasseur, J.-J.; Chen, Q.; et al. Reversible methylation of mA in the 5’ cap controls mRNA stability. Nature 2017, 541, 371–375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Zhao, B.; Roundtree, I.A.; Lu, Z.; Han, D.; Ma, H.; Weng, X.; Chen, K.; Shi, H.; He, C. N(6)-methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 2015, 161, 1388–1399. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Lu, Z.; Gomez, A.; Hon, G.C.; Yue, Y.; Han, D.; Fu, Y.; Parisien, M.; Dai, Q.; Jia, G.; et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 2014, 505, 117–120. [Google Scholar] [CrossRef]
- Shi, H.; Wang, X.; Lu, Z.; Zhao, B.S.; Ma, H.; Hsu, P.J.; Liu, C.; He, C. YTHDF3 facilitates translation and decay of N-methyladenosine-modified RNA. Cell Res. 2017, 27, 315–328. [Google Scholar] [CrossRef] [Green Version]
- Kretschmer, J.; Rao, H.; Hackert, P.; Sloan, K.E.; Höbartner, C.; Bohnsack, M.T. The mA reader protein YTHDC2 interacts with the small ribosomal subunit and the 5′-3′ exoribonuclease XRN1. RNA 2018, 24, 1339–1350. [Google Scholar] [CrossRef] [Green Version]
- Dai, X.; Wang, T.; Gonzalez, G.; Wang, Y. Identification of YTH Domain-Containing Proteins as the Readers for N1-Methyladenosine in RNA. Anal. Chem. 2018, 90, 6380–6384. [Google Scholar] [CrossRef]
- Meyer, K.D. DART-seq: An antibody-free method for global mA detection. Nat. Methods 2019, 16, 1275–1280. [Google Scholar] [CrossRef]
- Lin, S.; Choe, J.; Du, P.; Triboulet, R.; Gregory, R.I. The m(6)A Methyltransferase METTL3 Promotes Translation in Human Cancer Cells. Mol. Cell 2016, 62, 335–345. [Google Scholar] [CrossRef] [Green Version]
- Choe, J.; Lin, S.; Zhang, W.; Liu, Q.; Wang, L.; Ramírez-Moya, J.; Du, P.; Kim, W.; Tang, S.; Sliz, P.; et al. mRNA circularization by METTL3-eIF3h enhances translation and promotes oncogenesis. Nature 2018, 561, 556–560. [Google Scholar] [CrossRef] [Green Version]
- Shen, C.; Xuan, B.; Yan, T.; Ma, Y.; Xu, P.; Tian, X.; Zhang, X.; Cao, Y.; Ma, D.; Zhu, X.; et al. mA-dependent glycolysis enhances colorectal cancer progression. Mol. Cancer 2020, 19, 72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, D.; Tang, W.; Xu, Y.; Xu, Y.; Xu, B.; Fu, S.; Wang, Y.; Chen, F.; Chen, Y.; Han, Y.; et al. METTL3/YTHDF2 m6A axis accelerates colorectal carcinogenesis through epigenetically suppressing YPEL5. Mol. Oncol. 2021, 15, 2172–2184. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Wang, J.-Z.; Yang, X.; Yu, H.; Zhou, R.; Lu, H.-C.; Yuan, W.-B.; Lu, J.-C.; Zhou, Z.-J.; Lu, Q.; et al. METTL3 promote tumor proliferation of bladder cancer by accelerating pri-miR221/222 maturation in m6A-dependent manner. Mol. Cancer 2019, 18, 110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, H.; Wang, H.; Zhao, W.; Fu, S.; Li, Y.; Ni, W.; Xin, Y.; Li, W.; Yang, C.; Bai, Y.; et al. SUMO1 modification of methyltransferase-like 3 promotes tumor progression via regulating Snail mRNA homeostasis in hepatocellular carcinoma. Theranostics 2020, 10, 5671–5686. [Google Scholar] [CrossRef] [PubMed]
- Tassinari, V.; Cesarini, V.; Tomaselli, S.; Ianniello, Z.; Silvestris, D.A.; Ginistrelli, L.C.; Martini, M.; De Angelis, B.; De Luca, G.; Vitiani, L.R.; et al. ADAR1 is a new target of METTL3 and plays a pro-oncogenic role in glioblastoma by an editing-independent mechanism. Genome Biol. 2021, 22, 51. [Google Scholar] [CrossRef]
- Chai, R.-C.; Wu, F.; Wang, Q.-X.; Zhang, S.; Zhang, K.-N.; Liu, Y.-Q.; Zhao, Z.; Jiang, T.; Wang, Y.-Z.; Kang, C.-S. mA RNA methylation regulators contribute to malignant progression and have clinical prognostic impact in gliomas. Aging 2019, 11, 1204–1225. [Google Scholar] [CrossRef]
- Huo, F.-C.; Zhu, Z.-M.; Zhu, W.-T.; Du, Q.-Y.; Liang, J.; Mou, J. METTL3-mediated mA methylation of SPHK2 promotes gastric cancer progression by targeting KLF2. Oncogene 2021, 40, 2968–2981. [Google Scholar] [CrossRef]
- Yang, N.; Wang, T.; Li, Q.; Han, F.; Wang, Z.; Zhu, R.; Zhou, J. HBXIP drives metabolic reprogramming in hepatocellular carcinoma cells via METTL3-mediated m6A modification of HIF-1α. J. Cell Physiol. 2021, 236, 3863–3880. [Google Scholar] [CrossRef]
- Wang, H.; Xu, B.; Shi, J. N6-methyladenosine METTL3 promotes the breast cancer progression via targeting Bcl-2. Gene 2020, 722, 144076. [Google Scholar] [CrossRef]
- Vu, L.P.; Pickering, B.F.; Cheng, Y.; Zaccara, S.; Nguyen, D.; Minuesa, G.; Chou, T.; Chow, A.; Saletore, Y.; MacKay, M.; et al. The N-methyladenosine (mA)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 2017, 23, 1369–1376. [Google Scholar] [CrossRef]
- Weng, H.; Huang, H.; Wu, H.; Qin, X.; Zhao, B.S.; Dong, L.; Shi, H.; Skibbe, J.; Shen, C.; Hu, C.; et al. METTL14 Inhibits Hematopoietic Stem/Progenitor Differentiation and Promotes Leukemogenesis via mRNA mA Modification. Cell Stem. Cell 2018, 22, 191–205. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, T.; Wu, Z.; Wang, X.; Wang, Y.; Hu, X.; Qin, W.; Lu, S.; Xu, D.; Wu, Y.; Chen, Q.; et al. LNC942 promoting METTL14-mediated mA methylation in breast cancer cell proliferation and progression. Oncogene 2020, 39, 5358–5372. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Zhang, S.; He, C.; Xue, P.; Zhang, L.; He, Z.; Zang, L.; Feng, B.; Sun, J.; Zheng, M. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol. Cancer 2020, 19, 46. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Xu, M.; Xu, X.; Zeng, K.; Liu, X.; Sun, L.; Pan, B.; He, B.; Pan, Y.; Sun, H.; et al. METTL14 Suppresses CRC Progression via Regulating N6-Methyladenosine-Dependent Primary miR-375 Processing. Mol. Ther. 2020, 28, 599–612. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, H.-N.; Chen, Z.-Y.; Chen, X.-Y.; Chen, M.; Yi, Y.-C.; Zhu, J.-S.; Zhang, J. METTL14-mediated mA modification of circORC5 suppresses gastric cancer progression by regulating miR-30c-2-3p/AKT1S1 axis. Mol. Cancer 2022, 21, 51. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Yang, S.; Cui, Y.-H.; Wei, J.; Shah, P.; Park, G.; Cui, X.; He, C.; He, Y.-Y. METTL14 facilitates global genome repair and suppresses skin tumorigenesis. Proc. Natl. Acad. Sci. USA 2021, 118, e2025948118. [Google Scholar] [CrossRef]
- Chen, Y.; Peng, C.; Chen, J.; Chen, D.; Yang, B.; He, B.; Hu, W.; Zhang, Y.; Liu, H.; Dai, L.; et al. WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol. Cancer 2019, 18, 127. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.-X.; Zheng, Z.-Q.; Yang, P.-Y.; Lin, L.; Zhou, G.-Q.; Lv, J.-W.; Zhang, L.-L.; Chen, F.; Li, Y.-Q.; Wu, C.-F.; et al. WTAP-mediated m6A modification of lncRNA DIAPH1-AS1 enhances its stability to facilitate nasopharyngeal carcinoma growth and metastasis. Cell Death Differ. 2022, 29, 1137–1151. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Li, X.; Xiong, X.; Wang, J.; Zhou, Z.; Zhu, X.; Gu, Y.; Dominissini, D.; He, L.; et al. N-methyladenosine methylation in tRNA drives liver tumourigenesis by regulating cholesterol metabolism. Nat. Commun. 2021, 12, 6314. [Google Scholar] [CrossRef]
- Su, Z.; Monshaugen, I.; Wilson, B.; Wang, F.; Klungland, A.; Ougland, R.; Dutta, A. TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer. Nat. Commun. 2022, 13, 2165. [Google Scholar] [CrossRef]
- Li, Z.; Weng, H.; Su, R.; Weng, X.; Zuo, Z.; Li, C.; Huang, H.; Nachtergaele, S.; Dong, L.; Hu, C.; et al. FTO Plays an Oncogenic Role in Acute Myeloid Leukemia as a N-Methyladenosine RNA Demethylase. Cancer Cell 2017, 31, 127–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Su, R.; Deng, X.; Chen, Y.; Chen, J. FTO in cancer: Functions, molecular mechanisms, and therapeutic implications. Trends Cancer 2022, 8, 598–614. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Zhao, J.; Han, L.; Xu, M.; Wang, K.; Shi, J.; Dong, Z. Retrospective study of gene signatures and prognostic value of m6A regulatory factor in non-small cell lung cancer using TCGA database and the verification of FTO. Aging 2020, 12, 17022–17037. [Google Scholar] [CrossRef]
- Liu, J.; Ren, D.; Du, Z.; Wang, H.; Zhang, H.; Jin, Y. mA demethylase FTO facilitates tumor progression in lung squamous cell carcinoma by regulating MZF1 expression. Biochem. Biophys. Res. Commun. 2018, 502, 456–464. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Liao, Y.; Zhang, M.; Zhu, Y.; Wang, W.; Cai, H.; Liang, J.; Song, F.; Hou, C.; Huang, S.; et al. N6-methyladenosine demethyltransferase FTO-mediated autophagy in malignant development of oral squamous cell carcinoma. Oncogene 2021, 40, 3885–3898. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Lin, Z.; Wan, A.; Chen, H.; Liang, H.; Sun, L.; Wang, Y.; Li, X.; Xiong, X.-F.; Wei, B.; et al. RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Mol. Cancer 2019, 18, 46. [Google Scholar] [CrossRef] [Green Version]
- Cui, Y.; Zhang, C.; Ma, S.; Li, Z.; Wang, W.; Li, Y.; Ma, Y.; Fang, J.; Wang, Y.; Cao, W.; et al. RNA m6A demethylase FTO-mediated epigenetic up-regulation of LINC00022 promotes tumorigenesis in esophageal squamous cell carcinoma. J. Exp. Clin. Cancer Res. 2021, 40, 294. [Google Scholar] [CrossRef]
- Zeng, J.; Zhang, H.; Tan, Y.; Wang, Z.; Li, Y.; Yang, X. m6A demethylase FTO suppresses pancreatic cancer tumorigenesis by demethylating and inhibiting Wnt signaling. Mol. Ther. Nucleic Acids 2021, 25, 277–292. [Google Scholar] [CrossRef]
- Huang, J.; Sun, W.; Wang, Z.; Lv, C.; Zhang, T.; Zhang, D.; Dong, W.; Shao, L.; He, L.; Ji, X.; et al. FTO suppresses glycolysis and growth of papillary thyroid cancer via decreasing stability of APOE mRNA in an N6-methyladenosine-dependent manner. J. Exp. Clin. Cancer Res. 2022, 41, 42. [Google Scholar] [CrossRef]
- Shen, C.; Sheng, Y.; Zhu, A.C.; Robinson, S.; Jiang, X.; Dong, L.; Chen, H.; Su, R.; Yin, Z.; Li, W.; et al. RNA Demethylase ALKBH5 Selectively Promotes Tumorigenesis and Cancer Stem Cell Self-Renewal in Acute Myeloid Leukemia. Cell Stem. Cell 2020, 27, 64–80.e9. [Google Scholar] [CrossRef]
- Guo, X.; Li, K.; Jiang, W.; Hu, Y.; Xiao, W.; Huang, Y.; Feng, Y.; Pan, Q.; Wan, R. RNA demethylase ALKBH5 prevents pancreatic cancer progression by posttranscriptional activation of PER1 in an m6A-YTHDF2-dependent manner. Mol. Cancer 2020, 19, 91. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Wei, Q.; Jin, J.; Luo, Q.; Liu, Y.; Yang, Y.; Cheng, C.; Li, L.; Pi, J.; Si, Y.; et al. The m6A reader YTHDF1 promotes ovarian cancer progression via augmenting EIF3C translation. Nucleic Acids Res. 2020, 48, 3816–3831. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pi, J.; Wang, W.; Ji, M.; Wang, X.; Wei, X.; Jin, J.; Liu, T.; Qiang, J.; Qi, Z.; Li, F.; et al. YTHDF1 Promotes Gastric Carcinogenesis by Controlling Translation of. Cancer Res. 2021, 81, 2651–2665. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Sheng, H.; Ma, F.; Wu, Q.; Huang, J.; Chen, Q.; Sheng, L.; Zhu, X.; Zhu, X.; Xu, M. RNA mA reader YTHDF2 facilitates lung adenocarcinoma cell proliferation and metastasis by targeting the AXIN1/Wnt/β-catenin signaling. Cell Death Dis. 2021, 12, 479. [Google Scholar] [CrossRef]
- Zhang, L.; Wan, Y.; Zhang, Z.; Jiang, Y.; Gu, Z.; Ma, X.; Nie, S.; Yang, J.; Lang, J.; Cheng, W.; et al. IGF2BP1 overexpression stabilizes PEG10 mRNA in an m6A-dependent manner and promotes endometrial cancer progression. Theranostics 2021, 11, 1100–1114. [Google Scholar] [CrossRef]
- Müller, S.; Glaß, M.; Singh, A.K.; Haase, J.; Bley, N.; Fuchs, T.; Lederer, M.; Dahl, A.; Huang, H.; Chen, J.; et al. IGF2BP1 promotes SRF-dependent transcription in cancer in a m6A- and miRNA-dependent manner. Nucleic Acids Res. 2019, 47, 375–390. [Google Scholar] [CrossRef] [PubMed]
- Yue, B.; Song, C.; Yang, L.; Cui, R.; Cheng, X.; Zhang, Z.; Zhao, G. METTL3-mediated N6-methyladenosine modification is critical for epithelial-mesenchymal transition and metastasis of gastric cancer. Mol. Cancer 2019, 18, 142. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Pan, C.; Wang, X.; Xu, D.; Ma, Y.; Hu, J.; Chen, P.; Xiang, Z.; Rao, Q.; Han, X. Silencing of METTL3 effectively hinders invasion and metastasis of prostate cancer cells. Theranostics 2021, 11, 7640–7657. [Google Scholar] [CrossRef]
- Peng, W.; Li, J.; Chen, R.; Gu, Q.; Yang, P.; Qian, W.; Ji, D.; Wang, Q.; Zhang, Z.; Tang, J.; et al. Upregulated METTL3 promotes metastasis of colorectal Cancer via miR-1246/SPRED2/MAPK signaling pathway. J. Exp. Clin. Cancer Res. 2019, 38, 393. [Google Scholar] [CrossRef] [Green Version]
- Bi, X.; Lv, X.; Liu, D.; Guo, H.; Yao, G.; Wang, L.; Liang, X.; Yang, Y. METTL3 promotes the initiation and metastasis of ovarian cancer by inhibiting CCNG2 expression via promoting the maturation of pri-microRNA-1246. Cell Death Discov. 2021, 7, 237. [Google Scholar] [CrossRef]
- Dahal, U.; Le, K.; Gupta, M. RNA m6A methyltransferase METTL3 regulates invasiveness of melanoma cells by matrix metallopeptidase 2. Melanoma Res. 2019, 29, 382–389. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Xu, M.; Xu, X.; Zeng, K.; Liu, X.; Pan, B.; Li, C.; Sun, L.; Qin, J.; Xu, T.; et al. METTL14-mediated N6-methyladenosine modification of SOX4 mRNA inhibits tumor metastasis in colorectal cancer. Mol. Cancer 2020, 19, 106. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.-Z.; Yang, F.; Zhou, C.-C.; Liu, F.; Yuan, J.-H.; Wang, F.; Wang, T.-T.; Xu, Q.-G.; Zhou, W.-P.; Sun, S.-H. METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N -methyladenosine-dependent primary MicroRNA processing. Hepatology 2017, 65, 529–543. [Google Scholar] [CrossRef]
- Chen, S.; Yang, C.; Wang, Z.-W.; Hu, J.-F.; Pan, J.-J.; Liao, C.-Y.; Zhang, J.-Q.; Chen, J.-Z.; Huang, Y.; Huang, L.; et al. CLK1/SRSF5 pathway induces aberrant exon skipping of METTL14 and Cyclin L2 and promotes growth and metastasis of pancreatic cancer. J. Hematol. Oncol. 2021, 14, 60. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Liu, J.; Zhao, Y.; He, R.; Xu, X.; Guo, X.; Li, X.; Xu, S.; Miao, J.; Guo, J.; et al. Upregulation of METTL14 mediates the elevation of PERP mRNA N adenosine methylation promoting the growth and metastasis of pancreatic cancer. Mol. Cancer 2020, 19, 130. [Google Scholar] [CrossRef]
- Li, B.-Q.; Liang, Z.-Y.; Seery, S.; Liu, Q.-F.; You, L.; Zhang, T.-P.; Guo, J.-C.; Zhao, Y.-P. WT1 associated protein promotes metastasis and chemo-resistance to gemcitabine by stabilizing Fak mRNA in pancreatic cancer. Cancer Lett. 2019, 451, 48–57. [Google Scholar] [CrossRef]
- Xu, Y.; Ye, S.; Zhang, N.; Zheng, S.; Liu, H.; Zhou, K.; Wang, L.; Cao, Y.; Sun, P.; Wang, T. The FTO/miR-181b-3p/ARL5B signaling pathway regulates cell migration and invasion in breast cancer. Cancer Commun. 2020, 40, 484–500. [Google Scholar] [CrossRef]
- Wang, D.; Qu, X.; Lu, W.; Wang, Y.; Jin, Y.; Hou, K.; Yang, B.; Li, C.; Qi, J.; Xiao, J.; et al. N-Methyladenosine RNA Demethylase FTO Promotes Gastric Cancer Metastasis by Down-Regulating the m6A Methylation of ITGB1. Front. Oncol. 2021, 11, 681280. [Google Scholar] [CrossRef]
- Ruan, D.-Y.; Li, T.; Wang, Y.-N.; Meng, Q.; Li, Y.; Yu, K.; Wang, M.; Lin, J.-F.; Luo, L.-Z.; Wang, D.-S.; et al. FTO downregulation mediated by hypoxia facilitates colorectal cancer metastasis. Oncogene 2021, 40, 5168–5181. [Google Scholar] [CrossRef]
- Hu, Y.; Gong, C.; Li, Z.; Liu, J.; Chen, Y.; Huang, Y.; Luo, Q.; Wang, S.; Hou, Y.; Yang, S.; et al. Demethylase ALKBH5 suppresses invasion of gastric cancer via PKMYT1 m6A modification. Mol. Cancer 2022, 21, 34. [Google Scholar] [CrossRef]
- Jin, D.; Guo, J.; Wu, Y.; Yang, L.; Wang, X.; Du, J.; Dai, J.; Chen, W.; Gong, K.; Miao, S.; et al. mA demethylase ALKBH5 inhibits tumor growth and metastasis by reducing YTHDFs-mediated YAP expression and inhibiting miR-107/LATS2-mediated YAP activity in NSCLC. Mol. Cancer 2020, 19, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, X.; Dai, M.; Li, J.; Cai, J.; Zuo, Z.; Ni, S.; Zhang, Q.; Zhou, Z. m(6)A demethylase ALKBH5 inhibits cell proliferation and the metastasis of colorectal cancer by regulating the FOXO3/miR-21/SPRY2 axis. Am. J. Transl. Res. 2021, 13, 11209–11222. [Google Scholar] [PubMed]
- Zhu, K.; Li, Y.; Xu, Y. The FTO mA demethylase inhibits the invasion and migration of prostate cancer cells by regulating total mA levels. Life Sci. 2021, 271, 119180. [Google Scholar] [CrossRef] [PubMed]
- Su, T.; Huang, M.; Liao, J.; Lin, S.; Yu, P.; Yang, J.; Cai, Y.; Zhu, S.; Xu, L.; Peng, Z.; et al. Insufficient Radiofrequency Ablation Promotes Hepatocellular Carcinoma Metastasis Through N6-Methyladenosine mRNA Methylation-Dependent Mechanism. Hepatology 2021, 74, 1339–1356. [Google Scholar] [CrossRef]
- Wang, X.; Liu, C.; Zhang, S.; Yan, H.; Zhang, L.; Jiang, A.; Liu, Y.; Feng, Y.; Li, D.; Guo, Y.; et al. N-methyladenosine modification of MALAT1 promotes metastasis via reshaping nuclear speckles. Dev. Cell 2021, 56, 702–715.e8. [Google Scholar] [CrossRef]
- Chang, G.; Shi, L.; Ye, Y.; Shi, H.; Zeng, L.; Tiwary, S.; Huse, J.T.; Huo, L.; Ma, L.; Ma, Y.; et al. YTHDF3 Induces the Translation of mA-Enriched Gene Transcripts to Promote Breast Cancer Brain Metastasis. Cancer Cell 2020, 38, 857–871. [Google Scholar] [CrossRef]
- Zhao, T.; Wang, M.; Zhao, X.; Weng, S.; Qian, K.; Shi, K.; Gu, Y.; Ying, W.; Qian, X.; Zhang, Y. YTHDF2 Inhibits the Migration and Invasion of Lung Adenocarcinoma by Negatively Regulating the FAM83D-TGFβ1-SMAD2/3 Pathway. Front. Oncol. 2022, 12, 763341. [Google Scholar] [CrossRef]
- Bergers, G.; Benjamin, L.E. Tumorigenesis and the angiogenic switch. Nat. Rev. Cancer 2003, 3, 401–410. [Google Scholar] [CrossRef]
- Lee, E.Y.H.P.; Muller, W.J. Oncogenes and tumor suppressor genes. Cold Spring Harb. Perspect. Biol. 2010, 2, a003236. [Google Scholar] [CrossRef] [Green Version]
- Condorelli, D.F.; Privitera, A.P.; Barresi, V. Chromosomal Density of Cancer Up-Regulated Genes, Aberrant Enhancer Activity and Cancer Fitness Genes Are Associated with Transcriptional Cis-Effects of Broad Copy Number Gains in Colorectal Cancer. Int. J. Mol. Sci. 2019, 20, 4652. [Google Scholar] [CrossRef] [Green Version]
- Condorelli, D.F.; Spampinato, G.; Valenti, G.; Musso, N.; Castorina, S.; Barresi, V. Positive Caricature Transcriptomic Effects Associated with Broad Genomic Aberrations in Colorectal Cancer. Sci. Rep. 2018, 8, 14826. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Li, L.; Huang, Y.; Ma, J.; Min, J. Readers, writers and erasers of N-methylated adenosine modification. Curr. Opin. Struct. Biol. 2017, 47, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Gerken, T.; Girard, C.A.; Tung, Y.-C.L.; Webby, C.J.; Saudek, V.; Hewitson, K.S.; Yeo, G.S.H.; McDonough, M.A.; Cunliffe, S.; McNeill, L.A.; et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 2007, 318, 1469–1472. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zuidhof, H.R.; Calkhoven, C.F. Oncogenic and tumor-suppressive functions of the RNA demethylase FTO. Cancer Res. 2022, 82, 2201–2212. [Google Scholar] [CrossRef] [PubMed]
- Roomi, M.W.; Monterrey, J.C.; Kalinovsky, T.; Rath, M.; Niedzwiecki, A. In vitro modulation of MMP-2 and MMP-9 in human cervical and ovarian cancer cell lines by cytokines, inducers and inhibitors. Oncol. Rep. 2010, 23, 605–614. [Google Scholar]
- Huang, W.; Qi, C.-B.; Lv, S.-W.; Xie, M.; Feng, Y.-Q.; Yuan, B.-F. Determination of DNA and RNA Methylation in Circulating Tumor Cells by Mass Spectrometry. Anal. Chem. 2016, 88, 1378–1384. [Google Scholar] [CrossRef]
- Xiao, H.; Fan, X.; Zhang, R.; Wu, G. Upregulated N6-Methyladenosine RNA in Peripheral Blood: Potential Diagnostic Biomarker for Breast Cancer. Cancer Res. Treat. 2021, 53, 399–408. [Google Scholar] [CrossRef]
- Ge, L.; Zhang, N.; Chen, Z.; Song, J.; Wu, Y.; Li, Z.; Chen, F.; Wu, J.; Li, D.; Li, J.; et al. Level of N6-Methyladenosine in Peripheral Blood RNA: A Novel Predictive Biomarker for Gastric Cancer. Clin. Chem. 2020, 66, 342–351. [Google Scholar] [CrossRef]
- Pei, Y.; Lou, X.; Li, K.; Xu, X.; Guo, Y.; Xu, D.; Yang, Z.; Xu, D.; Cui, W.; Zhang, D. Peripheral Blood Leukocyte N6-methyladenosine is a Noninvasive Biomarker for Non-small-cell Lung Carcinoma. Onco Targets Ther. 2020, 13, 11913–11921. [Google Scholar] [CrossRef]
- Zhang, B.; Chen, Z.; Tao, B.; Yi, C.; Lin, Z.; Li, Y.; Shao, W.; Lin, J.; Chen, J. mA target microRNAs in serum for cancer detection. Mol. Cancer 2021, 20, 170. [Google Scholar] [CrossRef]
- Wang, Q.; Chen, C.; Ding, Q.; Zhao, Y.; Wang, Z.; Chen, J.; Jiang, Z.; Zhang, Y.; Xu, G.; Zhang, J.; et al. METTL3-mediated mA modification of HDGF mRNA promotes gastric cancer progression and has prognostic significance. Gut 2020, 69, 1193–1205. [Google Scholar] [CrossRef] [PubMed]
- Cai, C.; Long, J.; Huang, Q.; Han, Y.; Peng, Y.; Guo, C.; Liu, S.; Chen, Y.; Shen, E.; Long, K.; et al. M6A “Writer” Gene: A Favorable Prognostic Biomarker and Correlated With Immune Infiltrates in Rectal Cancer. Front. Oncol. 2021, 11, 615296. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Huang, X.; Li, Y.; Chen, Y.; Lin, L. mA-related lncRNAs are potential biomarkers for predicting prognoses and immune responses in patients with LUAD. Mol. Ther. Nucleic Acids 2021, 24, 780–791. [Google Scholar] [CrossRef]
- Tu, Z.; Wu, L.; Wang, P.; Hu, Q.; Tao, C.; Li, K.; Huang, K.; Zhu, X. N6-Methylandenosine-Related lncRNAs Are Potential Biomarkers for Predicting the Overall Survival of Lower-Grade Glioma Patients. Front. Cell Dev. Biol. 2020, 8, 642. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Meng, Q.; Ma, B. Characterization of the Prognostic m6A-Related lncRNA Signature in Gastric Cancer. Front. Oncol. 2021, 11, 630260. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhu, B.; He, M.; Cai, Y.; Ying, X.; Jiang, C.; Ji, W.; Zeng, J. N6-Methylandenosine-Related lncRNAs Predict Prognosis and Immunotherapy Response in Bladder Cancer. Front. Oncol. 2021, 11, 710767. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Zhang, R.; Jiang, Y.; Li, Y.; Lin, L.; Liu, Z.; Zhao, Y.; Shen, H.; Hu, Z.; Wei, Y.; et al. Comprehensive analyses of m6A regulators and interactive coding and non-coding RNAs across 32 cancer types. Mol. Cancer 2021, 20, 67. [Google Scholar] [CrossRef]
- Zheng, Q.; Yu, X.; Zhang, Q.; He, Y.; Guo, W. Genetic characteristics and prognostic implications of m1A regulators in pancreatic cancer. Biosci. Rep. 2021, 41. [Google Scholar] [CrossRef]
- Zhao, M.; Shen, S.; Xue, C. A Novel m1A-Score Model Correlated With the Immune Microenvironment Predicts Prognosis in Hepatocellular Carcinoma. Front. Immunol. 2022, 13, 805967. [Google Scholar] [CrossRef]
- Xiao, Y.; Yu, D. Tumor microenvironment as a therapeutic target in cancer. Pharmacol. Ther. 2021, 221, 107753. [Google Scholar] [CrossRef]
- Gu, Y.; Wu, X.; Zhang, J.; Fang, Y.; Pan, Y.; Shu, Y.; Ma, P. The evolving landscape of N-methyladenosine modification in the tumor microenvironment. Mol. Ther. 2021, 29, 1703–1715. [Google Scholar] [CrossRef] [PubMed]
- Subarsky, P.; Hill, R.P. The hypoxic tumour microenvironment and metastatic progression. Clin. Exp. Metastasis 2003, 20, 237–250. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Samanta, D.; Lu, H.; Bullen, J.W.; Zhang, H.; Chen, I.; He, X.; Semenza, G.L. Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m⁶A-demethylation of NANOG mRNA. Proc. Natl. Acad. Sci. USA 2016, 113, E2047–E2056. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, F.; Qin, X.; Wang, B.; Li, Q.; Hu, J.; Cheng, X.; Guo, D.; Cheng, F.-L.; Fang, C.; Tan, Y.; et al. ALKBH5 Facilitates Hypoxia-Induced Paraspeckle Assembly and IL8 Secretion to Generate an Immunosuppressive Tumor Microenvironment. Cancer Res. 2021, 81, 5876–5888. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Fan, S.; Wu, M.; Zuo, Z.; Li, X.; Jiang, L.; Shen, Q.; Xu, P.; Zeng, L.; Zhou, Y.; et al. YTHDF1 links hypoxia adaptation and non-small cell lung cancer progression. Nat. Commun. 2019, 10, 4892. [Google Scholar] [CrossRef] [Green Version]
- Liberti, M.V.; Locasale, J.W. The Warburg Effect: How Does it Benefit Cancer Cells? Trends Biochem. Sci. 2016, 41, 211–218. [Google Scholar] [CrossRef] [Green Version]
- Zhang, T.; Suo, C.; Zheng, C.; Zhang, H. Hypoxia and Metabolism in Metastasis. Adv. Exp. Med. Biol. 2019, 1136, 87–95. [Google Scholar]
- Gnoni, A.; Brunetti, O.; Longo, V.; Calabrese, A.; Argentiero, A.-l.; Calbi, R.; Solimando Antonio, G.; Licchetta, A. Immune system and bone microenvironment: Rationale for targeted cancer therapies. Oncotarget 2020, 11, 480–487. [Google Scholar]
- Morony, S.; Capparelli, C.; Sarosi, I.; Lacey, D.L.; Dunstan, C.R.; Kostenuik, P.J. Osteoprotegerin inhibits osteolysis and decreases skeletal tumor burden in syngeneic and nude mouse models of experimental bone metastasis. Cancer Res. 2001, 61, 4432–4436. [Google Scholar]
- Wong, S.K.; Mohamad, N.-V.; Giaze, T.R.; Chin, K.-Y.; Mohamed, N.; Ima-Nirwana, S. Prostate Cancer and Bone Metastases: The Underlying Mechanisms. Int. J. Mol. Sci. 2019, 20, 2587. [Google Scholar] [CrossRef] [Green Version]
- Fang, C.; He, M.; Li, D.; Xu, Q. YTHDF2 mediates LPS-induced osteoclastogenesis and inflammatory response via the NF-κB and MAPK signaling pathways. Cell Signal 2021, 85, 110060. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Qiao, S.-C.; Wu, X.-B.; Sun, B.; Yang, J.-G.; Li, X.; Zhang, X.; Qian, S.-J.; Gu, Y.-X.; Lai, H.-C. Circ_0008542 in osteoblast exosomes promotes osteoclast-induced bone resorption through m6A methylation. Cell Death Dis. 2021, 12, 628. [Google Scholar] [CrossRef] [PubMed]
- Previte, D.M.; Martins, C.P.; O’Connor, E.C.; Marre, M.L.; Coudriet, G.M.; Beck, N.W.; Menk, A.V.; Wright, R.H.; Tse, H.M.; Delgoffe, G.M.; et al. Lymphocyte Activation Gene-3 Maintains Mitochondrial and Metabolic Quiescence in Naive CD4 T Cells. Cell Rep. 2019, 27, 129–141.e4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bengsch, B.; Johnson, A.L.; Kurachi, M.; Odorizzi, P.M.; Pauken, K.E.; Attanasio, J.; Stelekati, E.; McLane, L.M.; Paley, M.A.; Delgoffe, G.M.; et al. Bioenergetic Insufficiencies Due to Metabolic Alterations Regulated by the Inhibitory Receptor PD-1 Are an Early Driver of CD8(+) T Cell Exhaustion. Immunity 2016, 45, 358–373. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Cao, X. Immunosuppressive cells in tumor immune escape and metastasis. J. Mol. Med. 2016, 94, 509–522. [Google Scholar] [CrossRef]
- Li, M.; Zha, X.; Wang, S. The role of N6-methyladenosine mRNA in the tumor microenvironment. Biochim. Biophys. Acta Rev. Cancer 2021, 1875, 188522. [Google Scholar] [CrossRef]
- Patsoukis, N.; Bardhan, K.; Chatterjee, P.; Sari, D.; Liu, B.; Bell, L.N.; Karoly, E.D.; Freeman, G.J.; Petkova, V.; Seth, P.; et al. PD-1 alters T-cell metabolic reprogramming by inhibiting glycolysis and promoting lipolysis and fatty acid oxidation. Nat. Commun. 2015, 6, 6692. [Google Scholar] [CrossRef] [Green Version]
- Xiong, J.; He, J.; Zhu, J.; Pan, J.; Liao, W.; Ye, H.; Wang, H.; Song, Y.; Du, Y.; Cui, B.; et al. Lactylation-driven METTL3-mediated RNA mA modification promotes immunosuppression of tumor-infiltrating myeloid cells. Mol. Cell 2022, 82, 1660–1677. [Google Scholar] [CrossRef]
- Dong, L.; Chen, C.; Zhang, Y.; Guo, P.; Wang, Z.; Li, J.; Liu, Y.; Liu, J.; Chang, R.; Li, Y.; et al. The loss of RNA N-adenosine methyltransferase Mettl14 in tumor-associated macrophages promotes CD8 T cell dysfunction and tumor growth. Cancer Cell 2021, 39, 945–957.e10. [Google Scholar] [CrossRef]
- Qiu, X.; Yang, S.; Wang, S.; Wu, J.; Zheng, B.; Wang, K.; Shen, S.; Jeong, S.; Li, Z.; Zhu, Y.; et al. MA Demethylase ALKBH5 Regulates PD-L1 Expression and Tumor Immunoenvironment in Intrahepatic Cholangiocarcinoma. Cancer Res. 2021, 81, 4778–4793. [Google Scholar] [CrossRef]
- Wang, L.; Hui, H.; Agrawal, K.; Kang, Y.; Li, N.; Tang, R.; Yuan, J.; Rana, T.M. m A RNA methyltransferases METTL3/14 regulate immune responses to anti-PD-1 therapy. EMBO J. 2020, 39, e104514. [Google Scholar] [CrossRef] [PubMed]
- Wan, W.; Ao, X.; Chen, Q.; Yu, Y.; Ao, L.; Xing, W.; Guo, W.; Wu, X.; Pu, C.; Hu, X.; et al. METTL3/IGF2BP3 axis inhibits tumor immune surveillance by upregulating N-methyladenosine modification of PD-L1 mRNA in breast cancer. Mol. Cancer 2022, 21, 60. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Wei, J.; Cui, Y.-H.; Park, G.; Shah, P.; Deng, Y.; Aplin, A.E.; Lu, Z.; Hwang, S.; He, C.; et al. mA mRNA demethylase FTO regulates melanoma tumorigenicity and response to anti-PD-1 blockade. Nat. Commun. 2019, 10, 2782. [Google Scholar] [CrossRef]
- Li, N.; Kang, Y.; Wang, L.; Huff, S.; Tang, R.; Hui, H.; Agrawal, K.; Gonzalez, G.M.; Wang, Y.; Patel, S.P.; et al. ALKBH5 regulates anti-PD-1 therapy response by modulating lactate and suppressive immune cell accumulation in tumor microenvironment. Proc. Natl. Acad. Sci. USA 2020, 117, 20159–20170. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhu, L.; Lu, C.; Wang, C.; Wang, H.; Jin, H.; Ma, X.; Cheng, Z.; Yu, C.; Wang, S.; et al. circNDUFB2 inhibits non-small cell lung cancer progression via destabilizing IGF2BPs and activating anti-tumor immunity. Nat. Commun. 2021, 12, 295. [Google Scholar] [CrossRef]
- Liu, Z.; Zhong, J.; Zeng, J.; Duan, X.; Lu, J.; Sun, X.; Liu, Q.; Liang, Y.; Lin, Z.; Zhong, W.; et al. Characterization of the m6A-Associated Tumor Immune Microenvironment in Prostate Cancer to Aid Immunotherapy. Front. Immunol. 2021, 12, 735170. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Wang, H.; Li, H.; Ye, X.; Xia, Y.; Yuan, S.; Lu, J.; Xie, X.; Wang, L.; Zhang, J. Integrated analyses of mA regulator-mediated modification patterns in tumor microenvironment-infiltrating immune cells in colon cancer. Oncoimmunology 2021, 10, 1936758. [Google Scholar] [CrossRef]
- Shriwas, O.; Mohapatra, P.; Mohanty, S.; Dash, R. The Impact of m6A RNA Modification in Therapy Resistance of Cancer: Implication in Chemotherapy, Radiotherapy, and Immunotherapy. Front. Oncol. 2020, 10, 612337. [Google Scholar] [CrossRef]
- Xiao, L.; Li, X.; Mu, Z.; Zhou, J.; Zhou, P.; Xie, C.; Jiang, S. FTO Inhibition Enhances the Antitumor Effect of Temozolomide by Targeting MYC-miR-155/23a Cluster-MXI1 Feedback Circuit in Glioma. Cancer Res. 2020, 80, 3945–3958. [Google Scholar] [CrossRef]
- Jin, D.; Guo, J.; Wu, Y.; Du, J.; Yang, L.; Wang, X.; Di, W.; Hu, B.; An, J.; Kong, L.; et al. mA mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. J. Hematol. Oncol. 2019, 12, 135. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Li, Q.; Li, G.; Zhang, Q.; Zhuo, L.; Han, X.; Zhang, M.; Chen, X.; Pan, T.; Yan, L.; et al. The mechanism of mA methyltransferase METTL3-mediated autophagy in reversing gefitinib resistance in NSCLC cells by β-elemene. Cell Death Dis. 2020, 11, 969. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Hong, X.; Li, S.; Meng, P.; Xiao, F. METTL3 promotes adriamycin resistance in MCF-7 breast cancer cells by accelerating pri-microRNA-221-3p maturation in a m6A-dependent manner. Exp. Mol. Med. 2021, 53, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Fukumoto, T.; Zhu, H.; Nacarelli, T.; Karakashev, S.; Fatkhutdinov, N.; Wu, S.; Liu, P.; Kossenkov, A.V.; Showe, L.C.; Jean, S.; et al. N-Methylation of Adenosine of mRNA Contributes to PARP Inhibitor Resistance. Cancer Res. 2019, 79, 2812–2820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Privitera, A.P.; Barresi, V.; Condorelli, D.F. Aberrations of Chromosomes 1 and 16 in Breast Cancer: A Framework for Cooperation of Transcriptionally Dysregulated Genes. Cancers 2021, 13, 1585. [Google Scholar] [CrossRef]
- Lin, Z.; Niu, Y.; Wan, A.; Chen, D.; Liang, H.; Chen, X.; Sun, L.; Zhan, S.; Chen, L.; Cheng, C.; et al. RNA m A methylation regulates sorafenib resistance in liver cancer through FOXO3-mediated autophagy. EMBO J. 2020, 39, e103181. [Google Scholar] [CrossRef]
- Wu, P.; Fang, X.; Liu, Y.; Tang, Y.; Wang, W.; Li, X.; Fan, Y. N6-methyladenosine modification of circCUX1 confers radioresistance of hypopharyngeal squamous cell carcinoma through caspase1 pathway. Cell Death Dis. 2021, 12, 298. [Google Scholar] [CrossRef]
- Cui, Q.; Shi, H.; Ye, P.; Li, L.; Qu, Q.; Sun, G.; Sun, G.; Lu, Z.; Huang, Y.; Yang, C.-G.; et al. mA RNA Methylation Regulates the Self-Renewal and Tumorigenesis of Glioblastoma Stem Cells. Cell Rep. 2017, 18, 2622–2634. [Google Scholar] [CrossRef]
- Dixit, D.; Prager, B.C.; Gimple, R.C.; Poh, H.X.; Wang, Y.; Wu, Q.; Qiu, Z.; Kidwell, R.L.; Kim, L.J.Y.; Xie, Q.; et al. The RNA m6A Reader YTHDF2 Maintains Oncogene Expression and Is a Targetable Dependency in Glioblastoma Stem Cells. Cancer Discov. 2021, 11, 480–499. [Google Scholar] [CrossRef]
- Su, R.; Dong, L.; Li, Y.; Gao, M.; Han, L.; Wunderlich, M.; Deng, X.; Li, H.; Huang, Y.; Gao, L.; et al. Targeting FTO Suppresses Cancer Stem Cell Maintenance and Immune Evasion. Cancer Cell 2020, 38, 79–96. [Google Scholar] [CrossRef]
- Relier, S.; Ripoll, J.; Guillorit, H.; Amalric, A.; Achour, C.; Boissière, F.; Vialaret, J.; Attina, A.; Debart, F.; Choquet, A.; et al. FTO-mediated cytoplasmic mA demethylation adjusts stem-like properties in colorectal cancer cell. Nat. Commun. 2021, 12, 1716. [Google Scholar] [CrossRef]
- Huang, H.; Wang, Y.; Kandpal, M.; Zhao, G.; Cardenas, H.; Ji, Y.; Chaparala, A.; Tanner, E.J.; Chen, J.; Davuluri, R.V.; et al. FTO-Dependent -Methyladenosine Modifications Inhibit Ovarian Cancer Stem Cell Self-Renewal by Blocking cAMP Signaling. Cancer Res. 2020, 80, 3200–3214. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Yang, H.; Yang, C.; Bao, Y.-L.; Yang, S.-M.; Liu, J.; Xiao, Y.-F. Translation of noncoding RNAs and cancer. Cancer Lett. 2021, 497, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Yankova, E.; Blackaby, W.; Albertella, M.; Rak, J.; De Braekeleer, E.; Tsagkogeorga, G.; Pilka, E.S.; Aspris, D.; Leggate, D.; Hendrick, A.G.; et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature 2021, 593, 597–601. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-H.; Choi, N.; Kim, S.; Jin, M.S.; Shen, H.; Kim, Y.-C. Eltrombopag as an Allosteric Inhibitor of the METTL3-14 Complex Affecting the mA Methylation of RNA in Acute Myeloid Leukemia Cells. Pharmaceuticals 2022, 15, 440. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Su, R.; Sheng, Y.; Dong, L.; Dong, Z.; Xu, H.; Ni, T.; Zhang, Z.S.; Zhang, T.; Li, C.; et al. Small-Molecule Targeting of Oncogenic FTO Demethylase in Acute Myeloid Leukemia. Cancer Cell 2019, 35, 677–691. [Google Scholar] [CrossRef] [Green Version]
- Sabnis, R.W. Novel Small Molecule RNA m6A Demethylase AlkBH5 Inhibitors for Treating Cancer. ACS Med. Chem. Lett. 2021, 12, 856–857. [Google Scholar] [CrossRef]



| Regulator | Target | Function | Reference |
|---|---|---|---|
| Writers | |||
| METTL3 | m6A | Binding to SAM; catalytic methylation | [26,27] |
| METTL14 | m6A | Almost no methyltransferase activity; increases enzyme activity of METTL3 by stabilizing the conformation of METTL3; involved in recognition of bases | [26,27] |
| WTAP | m6A | Regulatory factors of MTX of METTL3-METTL14 complex | [28,29] |
| KIAA1429 | m6A | Regulatory factors of MTX of METTL3-METTL14 complex | [29] |
| TRMT6/TRMT61A complex | m1A | Methyltransferases for mRNA in the cytosol | [6] |
| TRMT10C | m1A | Methyltransferases for mitochondrial mRNA | [6] |
| PCIF1 | m6Am | Methyltransferases | [30] |
| Erasers | |||
| FTO | m6A | Catalyzes the reaction of oxidative demethylation of m6A on ssRNAs | [31] |
| m6Am | Demethylates m6Am on snRNAs | [16] | |
| m1A | Demethylates m1A in RNAs with loop structure and tRNAs | [31] | |
| ALKBH5 | m1A | Demethylates specifically m6A on ssRNAs | [20,32] |
| Readers | |||
| YTHDF1 | m6A | Accelerate the translation process by promoting ribosome binding to target transcripts and increasing translation efficiency | [33] |
| YTHDF2 | m6A | Affect mRNA degradation by transporting mRNA with m6A modification to P bodies and other RNA decay sites | [34] |
| YTHDF3 | m6A | Promote protein translation in coordination with YTHDF1; accelerate mRNA degradation together with YTHDF2 | [35] |
| YTHDC2 | m6A | May involve in the regulation of RNA stability by binding to 5′-3′ exonuclease; affecting translation process by liaising m6A-containing transcripts and ribosomes | [36] |
| YTHDF1 | m1A | Promote the translation of transcripts with an m1A modification | [37] |
| TYHDF2 | m1A | Accelerate the deg-radiation of modified mRNAs | [37] |
| Regulator | Modification | Role in Cancer | Cancer Type | Functional Pathway | References |
|---|---|---|---|---|---|
| METTL3 | m6A | Oncogenesis | Human cancer cells | EGFR, TAZ | [39] |
| METTL3 | m6A | Oncogenesis | Human lung cancer cells | BRD4, eIF3h-dependent | [40] |
| METTL3 | m6A | Oncogenesis | Colorectal cancer (CRC) | HK2, SLC2A1, m6A-IGF2BP2/3-dependent | [41] |
| METTL3 | m6A | Oncogenesis | CRC | YPEL5, m6A-YTHDF2-dependent | [42] |
| METTL3 | m6A | Oncogenesis | Bladder cancer cells | PTEN, pri-miR221/222 | [43] |
| METTL3 | m6A | Oncogenesis | Hepatocellular carcinoma (HCC) | UBC9/SUMOylated METTL3/SNAIL axis | [44] |
| METTL3 | m6A | Oncogenesis | Glioma | [45,46] | |
| METTL3 | m6A | Oncogenesis | Gastric cancer | [47] | |
| METTL3 | m6A | Oncogenesis | HCC | [48] | |
| METTL3 | m6A | Oncogenesis | Breast cancer (BRCA) | [49] | |
| METTL3 | m6A | Oncogenesis | Acute myeloid leukemia (AML) | [50] | |
| METTL4 | m6A | Oncogenesis | AML | Self-renewal of leukemia stem cells, initiation of AML | [51] |
| METTL4 | m6A | Oncogenesis | BRCA | CXCR4, CYP1B1 | [52] |
| METTL4 | m6A | Anti-Oncogenesis | CRC | XIST | [53] |
| METTL4 | m6A | Anti-Oncogenesis | CRC | miR375/YAP1 pathway | [54] |
| METTL4 | m6A | Anti-Oncogenesis | Gastric cancer | miR-30c-2-3p/AKT1S1 axis | [55] |
| METTL4 | m6A | Anti-Oncogenesis | Skin oncogenesis induced by UVB | DDB2 | [56] |
| WTAP | m6A | Oncogenesis | HCC | ETS1, HuR/p21/p27-dpendent | [57] |
| WTAP | m6A | Oncogenesis | Nasopharyngeal carcinoma | DIAPH1-AS1, MTDH-LASP1 complex, LASP1 | [58] |
| TRMT6/TRMT61A | m1A | Oncogenesis | HCC | PPARδ, Cholesterol synthesis, Hedgehog signaling | [59] |
| TRMT6/TRMT62A | m1A | Oncogenesis | bladder cancer | Targetome of tRNA fragments, Unfolded protein response, Genes silence | [60] |
| FTO | m6A | Oncogenesis | AML | [61,62] | |
| FTO | m6A | Oncogenesis | Lung cancer | KRAS ang MZF1 signaling | [63,64] |
| FTO | m6A | Oncogenesis | Oral squamous cell carcinoma | eIF4G1 | [65] |
| FTO | m6A | Oncogenesis | BRCA | BNIP3, m6A-YTHDF2-dependent | [66] |
| FTO | m6A | Oncogenesis | Esophageal squamous Cell carcinoma | LINC00022 | [67] |
| FTO | m6A | Anti-Oncogenesis | Pancreatic cancer | Wnt signaling, PJA2 | [68] |
| FTO | m6A | Anti-Oncogenesis | Papillary thyroid cancer (PTC) | APOE, m6A-IGF2BP2-dependent | [69] |
| ALKBH5 | m6A | Oncogenesis | AML | [70] | |
| ALKBH5 | m6A | Anti-Oncogenesis | Pancreatic cancer | PER1, m6A-YTHDF2-dependent | [71] |
| YTHDF1 | Oncogenesis | Ovarian cancer | eIF3C | [72] | |
| YTHDF1 | Oncogenesis | Gastric cancer | FZD7 | [73] | |
| YTHDF2 | Oncogenesis | Lung cancer | AXIN1 | [74] | |
| IGF2BP1 | m6A | Oncogenesis | Endometrial cancer | PEG10 mRNA | [75] |
| IGF2BP1 | Oncogenesis | Lung, Ovarian and Liver cancer | SRF, FOXK1, PDZ, PDLIM7 | [76] | |
| METTL3 | m6A | Metastasis | Gastric cancer | ZMYM1, CtBP/LSD1/CoREST complex | [77] |
| METTL3 | Metastasis | Prostate cancer | A2696, USP4, ELAVL1 | [78] | |
| METTL3 | Metastasis | CRC | pri-miR-1246, SPRED2/MAPK signaling pathway | [79] | |
| METTL3 | Metastasis | Ovarian cancer | pri-miR-1246, CCNG2 pathway | [80] | |
| METTL3 | Metastasis | Melanoma cells | MMPs | [81] | |
| METTL14 | m6A | Anti-Metastasis | CRC | SOX4, PI3K/Akt signaling | [82] |
| METTL14 | Anti-Metastasis | HCC | pri-miR-126, DGCR8 | [83] | |
| METTL14 | Anti-Metastasis | Pancreatic cancer | CLK1/SRSF5 pathway | [84] | |
| METTL14 | Metastasis | Pancreatic cancer | p53, PERP mRNA | [85] | |
| WTAP | Metastasis | Pancreatic cancer | Fak mRNA, Fak-related pathways | [86] | |
| FTO | Metastasis | BRCA | miR-181b-3p, ARL5B | [87] | |
| FTO | m6A | Metastasis | Gastric cancer | ITGB1 | [88] |
| FTO | Anti-Metastasis | CRC | MTA1, IGF2BP2 | [89] | |
| AKJBH5 | m6A | Anti-Metastasis | Gastric cancer | PKMYT1, IGF2BP3-m6A-mediated | [90] |
| AKJBH5 | Anti-Metastasis | Prostate cancer, CRC and non-small-cell lung cancer | [91,92,93] | ||
| YTHDF1 | m6A | Metastasis | HCC (after insufficient radiofrequency ablation) | EGFR | [94] |
| YTHDC1 | m6A | Metastasis | Esophageal cancer cells (ESCCs) | MALAT1 | [95] |
| YTHDF3 | m6A | Metastasis | BRCA | ST6GALNAC5, GJA1 and EGFR | [96] |
| YTHDF2 | Anti-Metastasis | Lung adenocarcinoma | [97] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qu, X.; Zhang, Y.; Sang, X.; Ren, D.; Zhao, H.; Wong, S.T.C. Methyladenosine Modification in RNAs: From Regulatory Roles to Therapeutic Implications in Cancer. Cancers 2022, 14, 3195. https://doi.org/10.3390/cancers14133195
Qu X, Zhang Y, Sang X, Ren D, Zhao H, Wong STC. Methyladenosine Modification in RNAs: From Regulatory Roles to Therapeutic Implications in Cancer. Cancers. 2022; 14(13):3195. https://doi.org/10.3390/cancers14133195
Chicago/Turabian StyleQu, Xiaolin, Yongqiu Zhang, Xianzheng Sang, Ding Ren, Hong Zhao, and Stephen T. C. Wong. 2022. "Methyladenosine Modification in RNAs: From Regulatory Roles to Therapeutic Implications in Cancer" Cancers 14, no. 13: 3195. https://doi.org/10.3390/cancers14133195






