Simple Summary
Emerging evidence highlights long non-coding RNAs as important regulators of epithelial–mesenchymal transition. Numerous studies have attempted to define their possible diagnostic, prognostic and therapeutic values in various human cancers. The aim of this review is to summarize long non-coding RNAs involved in the regulation of epithelial–mesenchymal transition in colorectal carcinoma. Additional candidate long non-coding RNAs are identified through a bioinformatics analysis.
Abstract
Epithelial–mesenchymal transition (EMT) plays a pivotal role in carcinogenesis, influencing cancer progression, metastases, stemness, immune evasion, metabolic reprogramming and therapeutic resistance. EMT in most carcinomas, including colorectal carcinoma (CRC), is only partial, and can be evidenced by identification of the underlying molecular drivers and their regulatory molecules. During EMT, cellular reprogramming is orchestrated by core EMT transcription factors (EMT-TFs), namely ZEB1/2, TWIST1/2, SNAI1 (SNAIL) and SNAI2 (SLUG). While microRNAs have been clearly defined as regulators of EMT, the role of long non-coding RNAs (lncRNAs) in EMT is poorly defined and controversial. Determining the role of lncRNAs in EMT remains a challenge, because they are involved in a number of cellular pathways and are operating through various mechanisms. Adding to the complexity, some lncRNAs have controversial functions across different tumor types, acting as EMT promotors in some tumors and as EMT suppressors in others. The aim of this review is to summarize the role of lncRNAs involved in the regulation of EMT-TFs in human CRC. Additional candidate lncRNAs were identified through a bioinformatics analysis.
1. Introduction
Epithelial–mesenchymal transition (EMT) is a process of cellular transdifferentiation whereby stationary epithelial cells acquire a motile mesenchymal phenotype [1]. It is generally accepted that EMT is an important mechanism in human carcinogenesis, but despite extensive research, its role in cancer development and progression is not completely understood yet. Cells undergoing EMT gain the ability to migrate and invade beyond the invasive tumor front, promoting tumor progression and metastasis [2]. Additionally, studies indicate that EMT plays a pivotal role in cancer cell stemness, immune evasion, metabolic reprogramming and therapeutic resistance [3,4].
Full EMT involves a complete transition from an epithelial to a mesenchymal cell, evidenced by an altered morphology and a change in EMT-related molecular biomarkers [5,6]. Full EMT is rare in human cancers, but it does occur, for example in spindle cell carcinoma and carcinosarcoma [7]. EMT in most other carcinomas including colorectal cancer (CRC) is only partial, giving rise to a spectrum of intermediate cellular transition states [8,9]. Traditional biomarkers based on protein expression characteristic of either epithelial (E-cadherin, cytokeratins) or mesenchymal (N-cadherin, vimentin, fibronectin) cells are insufficient for defining the various states of partial EMT. Therefore, evidence of partial EMT relies on identification of molecular drivers governing EMT, transcription factors and their regulatory molecules [10,11].
EMT is orchestrated by EMT transcription factors (EMT-TF), which can both repress epithelial genes involved in cellular adhesion, polarity and cytoskeleton reorganization as well as activate genes associated with a mesenchymal phenotype. Among the most investigated EMT-TFs are ZEB1/2, TWIST1/2, SNAI1 (SNAIL) and SNAI2 (SLUG). These are crucial for maintaining normal epithelium structure and their dysregulation induces EMT [12,13,14,15]. Over the past years, studies on various different cancers in vitro and in vivo also highlighted their role in other aspects of tumorigenesis, such as regulation of stemness, therapy resistance, immune evasion, DNA integrity and metabolic reprogramming. EMT-TFs usually cooperate with one another and likely act in a tumor- and dosage-specific manner, consequently exhibiting different expression patterns and functions across tumor types [3,16,17,18]. Their expression is regulated on multiple cellular levels including non-coding RNAs as one of the epigenetic mechanisms. While microRNAs have been clearly defined as regulators of EMT, less is known about the role of long non-coding RNAs (lncRNAs) in EMT in human cancers [19,20,21,22].
LncRNAs have emerged as important regulators of EMT in human cancers and an increasing number of studies have attempted to define their possible diagnostic, prognostic and therapeutic values. LncRNAs are RNA transcripts longer than 200 base pairs that do not encode proteins. Instead, they influence various cellular processes by different modes of action: as mediators of gene transcription; as decoy molecules for proteins, which results in dysregulation of DNA and mRNA; as partners in competing endogenous RNA (ceRNA), which results in miRNA depletion and increasing expression of their targeted genes; as guide molecules for proteins; and as scaffold molecules resulting in the assembly of macromolecular complexes that regulate the expression of their target genes [23,24].
LncRNAs may play a critical role in tumor progression and metastasis due to their ability to regulate EMT-TFs. The regulation can take place in the nucleus at epigenetic or transcription level or as regulation of pre-mRNA alternative splicing. The regulation can also take place in the cytoplasm at the post-transcriptional level, which includes mRNA translation and stability, protein stability and the ceRNA network [24]. However, because their regulation is diverse, determining the effect of lncRNAs on the process of EMT remains a challenge [21,24,25]. Adding to the complexity, some lncRNAs have controversial functions across different tumor types, acting as EMT promotors in some tumors and as EMT suppressors in others [26,27,28,29].
A thorough understanding of EMT-TF regulation is a prerequisite to defining complete and partial EMT. For this reason, we aim to investigate the role of lncRNAs involved in the regulation of EMT-TFs in human CRC. The study provides candidate lncRNAs identified through a systematic literature review as well as by performing a bioinformatics analysis.
2. Methods
2.1. Systematic Literature Review
2.1.1. Study Design
This study was conducted using the “Preferred Reporting Items for Systematic Reviews and Meta-analyses” (PRISMA) guidelines [30]. The study is registered with the Research Registry and its unique identifying number (UIN) is: reviewregistry1311. The online PubMed (NCBI, Bethesda, MD, USA) database was systematically searched for all articles relating to lncRNAs and their role in relation to EMT-TFs in colorectal adenocarcinoma for the period from 1 January 1990 to 30 March 2022. No language restrictions were applied. The search was conducted using the following terms: »colorectal«, »colon«, »long non-coding« and »lncRNA« in combination with each of the following transcription factors: »SNAIL«, »SNAI1«, »SLUG«, »SNAI2«, »TWIST«, »TWIST1«, »TWIST2«, »ZEB«, »ZEB1« and »ZEB2« present in either the title or the abstract of articles. »AND« was used to connect main research terms whereas »OR« was used to incorporate synonym words.
2.1.2. Inclusion and Exclusion Criteria
Studies were included if they provided insight into the molecular signaling between lncRNAs and any of the core EMT-TFs: SNAI1 (SNAIL), SNAI2 (SLUG), TWIST1 (TWIST), TWIST2, ZEB1 or ZEB2 in human CRC samples, in vitro human CRC cell lines and/or animal models. Studies were excluded if (i) they did not pertain to CRC, (ii) they were focused on other signaling pathways without evaluating the expression of the above-mentioned TFs and/or lncRNAs (iii) they were reviews, conference abstracts, unpublished manuscripts and letters, or (iv) full-text articles were not available.
2.1.3. Data Extraction and Outcomes
Following the conducted search, the titles and abstracts of articles were screened by two reviewers (AP and NZ). For the sake of completion, the references of the articles were manually searched for additional studies that, based on their titles, appeared relevant to this review, but were not found with the primary database search. These articles were added to the screening process and were then further categorized based on the inclusion and exclusion criteria with the rest of the articles. The following data were extracted from each article: authors, year and journal of publication, patient characteristics (overall survival, TNM stage, lymphovascular invasion), model type (human tissue, CRC cell lines, experimental animals), main study findings including expression of lncRNA and EMT-TF in CRC, function and the proposed mechanism of lncRNA in CRC.
2.2. Bioinformatics Analysis
A bioinformatics analysis of lncRNA and TF correlations on TCGA datasets of colon adenocarcinoma and rectum adenocarcinoma was performed using The Cancer Genome Atlas (TCGA) with TCGAbiolinks package in R programming language [31].
3. Results
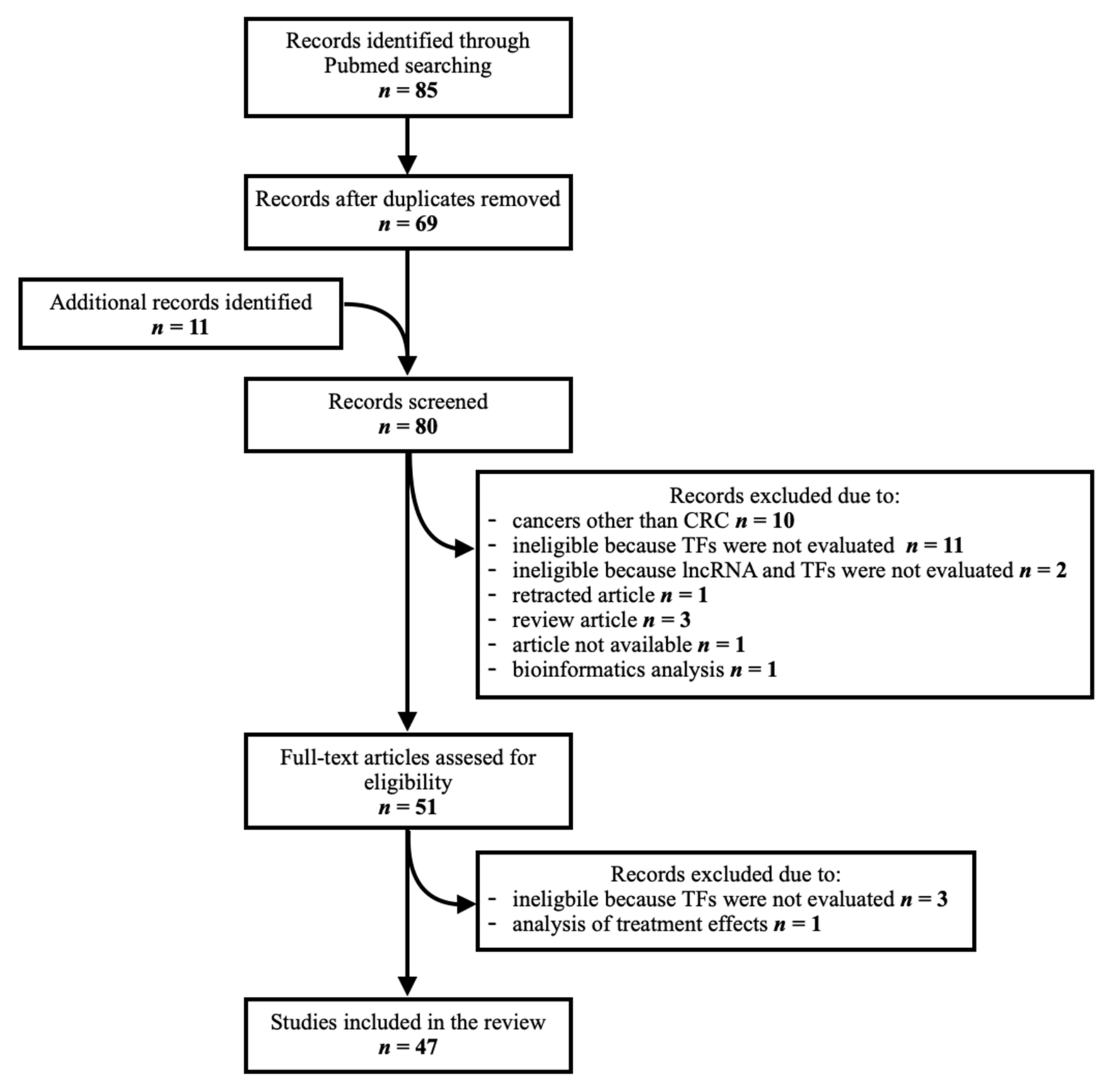
Eighty-five articles were identified through a PubMed database search. Duplicates were removed (n = 16). Additional articles pertaining to this topic (n = 11) were manually searched from the references of records identified through the database search. They were screened along with those identified through the database search to assess their relevance, as some contained relevant data despite not being found with the initial search. In total, eighty articles were screened. Thirty-three were excluded due to the following reasons: research on cancers other than colorectal (n = 10), ineligible because the TFs and/or lncRNAs of interest were not evaluated (n = 16), retracted article (n = 1), review article (n = 3), unavailable article (n = 1), analysis of treatment effect (n = 1) and bioinformatics analysis (n = 1), leaving 47 studies for inclusion in the final analysis. A flow chart of the article identification and selection process is provided in Figure 1.
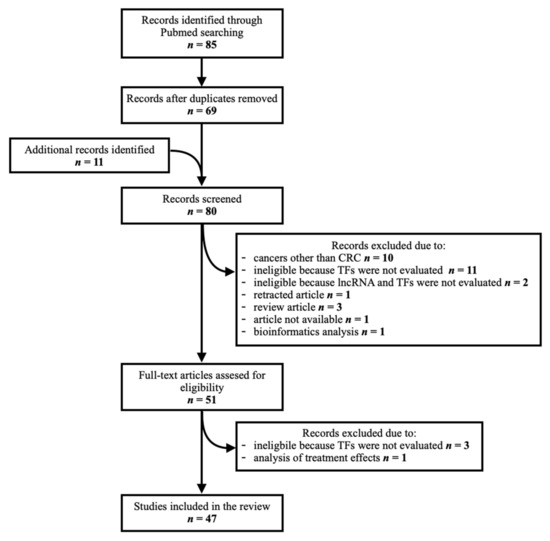
Figure 1.
Schematic representation of article identification and selection process.
3.1. LncRNAs Regulating a Single EMT-TF
Fourty-one lncRNAs acting as regulators of one or more EMT-TFs were identified. Twenty-five lncRNAs were shown to regulate a single EMT-TF; ZEB1 (n = 13), SNAI1 (n = 4), SNAI2 (n = 3) and TWIST1 (n = 5). A short summary of their cellular effects is provided in Table 1. None of the lncRNAs were shown to influence the expression of ZEB2 only.

Table 1.
Identified lncRNAs regulating a single EMT-TF and their major effects. Legend: CSC, cancer stem cell properties; Invas., invasion; LVI, lymphovascular invasion; Migr., migration; M, metastasis stage; Met., distant metastasis; Morph., change in morphology; N, node stage; OS, overall patient survival; Prolif., proliferation; T, tumor stage; ↓, decreased; ↑, increased.
3.2. ZEB1
The greatest number of studies researching lncRNAs as regulators of EMT-TFs pertained to ZEB1. Twenty lncRNAs modulating the activity of ZEB1 have been identified so far. All exhibit a pro-EMT effect. Thirteen lncRNAs were shown to influence the expression of ZEB1 only. Their activation was mostly associated with a shorter overall survival and a higher TNM stage. In vitro studies were performed for all 13 lncRNAs. Apart from DUXAP8, which was shown to promote cell proliferation only, the effects of all other lncRNAs were similar as observed in vitro. They mainly affected cell migration and, in many cases, also cell invasion. Metastatic potential was additionally confirmed in vivo for five out of seven lncRNAs (AC010789.1, AK000053, LINC01413, N-BLR, RP11 and XIST). Even though ZFAS exhibited migratory and invasive traits in vitro, it only promoted tumor growth and not metastasis formation in vivo. Interestingly, AK000053 and RP11 were also found to influence cell morphology promoting a mesenchymal phenotype, further highlighting their role in EMT. AK000053 was the only lncRNA from this group, which was also shown to evoke stem cell properties.
3.3. SNAI1
SNAI1 is frequently regulated alongside other TFs as a result of various lncRNA activations (n = 13), whereas only four lncRNAs seem to regulate SNAI1 only. Two of them, namely GNAT1-1 and SATB2-AS1, exhibited an anti-EMT effect. Their activation was associated with an increased overall patient survival and a lower TNM stage. Both were also shown to decrease migration and cell invasion in vitro and decrease the incidence of metastasis in vivo. The other two lncRNAs modulating SNAI1, namely LDLRAD4-AS1 and TRERNA1, exhibited a pro-EMT effect, with cellular migration and invasion observed in vitro. They were both associated with nodal and distant metastases in CRC patients. High expression of TRERNA1 was also associated with metastatic traits in vitro; however, metastatic potential in vivo was confirmed only for LDLRAD4-AS1.
3.4. TWIST1
Five lncRNAs (AK027294, CHRF, LINCOO467, LINC00941 and SNHG11) were shown to stimulate EMT by influencing the expression of TWIST1 only. All of them were associated with an increased migratory and/or invasive ability of cells in vitro. In vivo studies were available for CHRF, LINC00941 and SNHG11, all of which were shown to be associated with an increased metastatic potential. In addition, high expression of CHRF and LINC00941 also triggered a change in cell morphology, a feature of a complete EMT.
3.5. SNAI2
SNAI2 was solely regulated by three lncRNAs, namely GAPLINC, SNHG15 and XLOC_010588, all exerting a pro-EMT effect, with cellular migration and/or invasion observed in vitro. Interestingly, none of these lncRNAs promoted metastasis formation either in vitro or in vivo.
3.6. LncRNAs Regulating Multiple EMT-TFs
Sixteen lncRNAs were shown to regulate either two (n = 9) or three (n = 7) EMT-TFs (Table 2). The identified lncRNAs and a short summary of their effects are provided in Table 3.

Table 2.
LncRNAs regulating multiple EMT-TFs.

Table 3.
Identified lncRNAs regulating multiple EMT-TFs and their major effects. Legend: CSC, cancer stem cell properties; Invas., invasion; LVI, lymphovascular invasion; Migr., migration; M, metastasis stage; Met., distant metastasis; Morph., change in morphology; N, node stage; OS, overall patient survival; Prolif., proliferation; T, tumor stage; ↓, decreased; ↑, increased.
3.7. ZEB1 and ZEB2
The effects of lncRNAs regulating both ZEB1 and ZEB2 (DLEU2, LINC01296) were similar to those of lncRNAs regulating ZEB1 alone. Both correlated with distant metastasis in patients with CRC and a cellular invasive phenotype observed in vitro. As of yet, no functional analyses have been performed in vivo.
3.8. ZEB1 and SNAI1
Two lncRNAs were shown to regulate both ZEB1 and SNAI1, namely VIM-AS1 and ZEB2-AS1. While they were both associated with a shorter survival, observed in vitro effects were different. VIM-AS1 promoted cellular proliferation and migration and had an effect on cell morphology, while ZEB2-AS1 was linked to cell migration and invasion only. Their effects have not yet been investigated in vivo.
3.9. ZEB1, ZEB2 and SNAI1
H19 has been shown to affect the expression of ZEB1, ZEB2 and SNAI1. Expression of H19 in CRC correlated with a shorter survival and an increased TNM stage. The effects of H19 in vitro were assessed by two study groups, both reporting an increased cellular proliferation and migration of CRC cells. One of the groups also detected a change in cell morphology while the other observed an invasive and metastasizing cellular phenotype. H19 has been shown to stimulate tumor growth in vivo.
3.10. ZEB1, SNAI1 and SNAI2
SNHG6 and XIST have been found to influence the expressions of ZEB1, SNAI1 and SNAI2 in CRC. Both have been linked to a decreased overall survival. Their expression promoted proliferation, migration and invasion of cancer cells in vitro. SNHG6 also evoked metastatic cellular traits in vitro; however, its metastatic potential was not confirmed in vivo. On the contrary, higher expression of XIST both increased metastasis formation in vivo and evoked stem cell properties and the spindle morphology of cells in vitro.
3.11. ZEB2, SNAI1 and SNAI2
UICLM influenced the expression of ZEB2, SNAI1 and SNAI2. Its expression was associated with a shorter overall survival and a higher TNM stage. In vitro, it stimulated cell proliferation and invasion as well as promoted stem cell properties. In vivo, its expression increased tumor size and promoted metastasis formation.
3.12. ZEB2 and SNAI2
B3GALT5-AS1 regulating ZEB2 and SNAI2 has been shown to have an anti-EMT effect in CRC. Its expression was associated with an increased overall patient survival and inhibited migratory and invasive cell abilities in vitro. In vivo studies also suggest that high B3GALT5-AS1 expression prevents metastasis formation.
3.13. ZEB2, SNAI1 and TWIST1
The effect of TUG1 on EMT-TF in CRC has been investigated by two groups. One found that TUG1 augmented proliferation and the invasive traits of CRC cells in vitro by regulating ZEB2 and SNAI1. The other observed no increase in cancer cell proliferation in vitro, but rather an increase in migratory as well as invasive cell abilities. By regulating TWIST1 they also showed that TUG1 expression influences metastasis formation in vivo.
3.14. SNAI1 and SNAI2
MAPKAPK5-AS1 and HOTAIR have been shown to promote EMT in CRC by regulating SNAI1 and SNAI2. Their expression exerted a similar effect. Both lncRNAs promoted cell migration and invasion in vitro and stimulated the formation of distant metastasis in vivo. The expression of MAPKAPK5-AS1 has also been linked to an increased TNM stage and a shorter overall survival, whereas this research has not been conducted for HOTAIR.
3.15. SNAI1 and TWIST1
SNAI1 and TWIST1 are regulated by lncRNAs CTD-903 and PANDAR. CTD-903 was shown to suppress EMT in vitro, as evidenced by decreased cell proliferation, migration and invasion. It also promoted an epithelioid cellular phenotype. The expression of PANDAR increased cell proliferation, migration and invasion in vitro. The in vivo effects of neither CTD-903 nor PANDAR have been investigated.
3.16. SNAI1, SNAI2 and TWIST1
HOXA11-AS and MIR4435-2HG have been shown to regulate three EMT-TFs, namely SNAI1, SNAI2 and TWIST1. High expression of both HOXA11-AS and MIR4435-2HG was associated with a higher rate of metastasis in CRC patients and they were both shown to evoke migration and invasion of cells in vitro. Additionally, MIR4435-2HG also promoted tumor growth and distant metastasis formation in vivo, whereas such studies have not been performed yet for HOXA11-AS.
3.17. Bioinformatics Analysis
Data were available for 609 tumor samples and 51 normal samples. We obtained a list of 18,855 known lncRNAs from (http://genome.igib.res.in/lncRNome, accessed on 27 January 2022). The intersect between the list of known lncRNAs and TCGA data yielded 3418 lncRNAs. For the purpose of this study, we calculated Pearson’s correlation with the package “sigr” among all the lncRNAs included in TCGA and EMT-TFs: ZEB1, ZEB1, SNAI1, SNAI2, TWIST1 and TWIST2 [78].
Significant correlations (p < 0.05) were divided into groups designated as “very strong”, “strong”, “moderate” and “weak” correlations, based on correlation coefficients of at least |0.8|, |0.6|, |0.4|, and |0.2|, respectively. The analysis yielded 258 significant correlations of lncRNAs-TF pairs in tumor samples (n = 2 very strong; n = 11 strong; n = 41 moderate; n = 204 weak). The significant strong and very strong correlations are presented in Table 4, while all the results are presented in Supplementary Table S1.

Table 4.
Statistically significant correlations of selected lncRNAs and EMT-TFs in tumor of TCGA colon and rectum adenocarcinoma datasets. ↑↑↑, very strong positive correlation; ↑↑, strong positive correlation.
4. Discussion
The aim of this review was to summarize and identify lncRNAs involved in the process of EMT in CRC and examine their functions. Through a systematic literature review and a bioinformatics analysis, we identified lncRNAs shown to regulate core EMT-TFs responsible for cellular reprogramming during EMT in CRC, namely ZEB1/2, TWIST1/2 and SNAI1/2.
Through a systematic literature review, we identified 41 lncRNAs shown to regulate one or more core EMT-TFs in CRC. With regard to clinical relevance, the expression of the most lncRNAs significantly correlated with overall survival and TNM stages. The majority of lncRNAs were upregulated in CRC and promoted the “classical” traits of EMT. Following their knock-out in vitro and in vivo, the proliferative, migratory and invasive capabilities of cancer cells diminished. Frequently, the expression of investigated lncRNAs also enhanced the metastatic potential of CRC cells. Importantly, the effects of lncRNAs in CRC cell lines did not always correlate to the ones on animal models. Studies containing both in vitro and in vivo results were available for 24 lncRNAs. Most of lncRNAs were shown to trigger invasive cancer cell traits in vitro and concordantly promoted metastasis formation in vivo. A few lncRNAs were not associated with cell invasion in vitro or increased metastatic potential in vivo. By contrast, certain lncRNAs (ZFAS1, GAPLINC, SNHG6) promoted invasive or even metastatic potential on CRC cells in vitro and yet were not observed to enhance the metastatic potential of tumors in vivo [45,50,71]. Therefore, conclusions regarding the role of lncRNAs in CRC progression should not be based solely on in vitro studies as further validation is required before establishing any possible prognostic role of lncRNAs in the clinical setting.
Other features of EMT were less frequently associated with lncRNAs. Firstly, silencing certain lncRNAs prompted a change in cell morphology, a feature of full EMT [26,34,42,54,56,59,61,76]. This is consistent with previous studies linking ZEB1, SNAI1 and TWIST1 with a mesenchymal phenotype of tumor cells in CRC. ZEB1 was overexpressed particularly in tumor cells at the invasive front and its knock-down promoted an epithelioid phenotype in CRC [79]. In CRC cells overexpressing SNAI1, a clear switch to a mesenchymal phenotype was observed microscopically [80]. The presence of TWIST1 expression in CRC was reported mainly in cancer cells with a mesenchymal phenotype located in the tumor stroma. Additionally, cell lines transfected with TWIST1 acquired characteristics of activated cancer-associated fibroblasts [81,82]. Secondly, AK000053, XIST and UICLM were shown to evoke stem cell properties in CRC [26,34,75]. This corroborates previous findings linking their EMT-TF targets (ZEB1, SNAI1 and SNAI2) with cancer stem cells in CRC. SNAI1 was shown to induce a stem-cell-like phenotype [80,83], while ZEB1 [84] and SNAI2 [85] were suggested to play critical roles in the maintenance of self-renewal capacity in CRC. Lastly, some studies also examined the role of lncRNAs in therapy resistance. By affecting the expression of ZEB1, N-BLR and LINC00460 induced chemoresistance and contributed to the radioresistance of CRC, respectively [39,41]. This is consistent with previous findings indicating that ZEB1 plays a critical role in the EMT-regulated oxaliplatin resistance of CRC [86].
It is important to note that none of the investigated lncRNAs were shown to influence ZEB2 only. We can only speculatively suggest possible explanations. Firstly, similarly to squamous cell carcinoma of the head and neck, ZEB1 and ZEB2 could be co-expressed and regulated together [87]. Secondly, it is possible that ZEB2 activation occurs incidentally during the regulation of ZEB1 or SLUG, as frequently observed with miRNAs [88]. Thirdly, TFs might activate and co-regulate each other to provide a certain phenotype, similarly to stem cell markers or members of one miRNA family. The activation of the EMT program might depend on the co-regulation of different EMT-TFs (e.g., SNAI2 and ZEB2) [89]. Lastly, lncRNAs influencing only ZEB2 have yet to be discovered.
Accurate evaluation of the role of lncRNAs in EMT requires further research taking into account their possible differential expression in the tumor microenvironment. The majority of studies identified in this review utilized whole-tumor RNA isolation, thus disregarding the important concept of intra-tumor heterogeneity. While the genetic landscape of the entire tumor may be very similar, the distribution and frequency of epigenetic changes vary within a tumor [90]. Accordingly, it has been shown that partial EMT is mainly activated at the invasive tumor front, especially in the areas of basement membrane disintegration and in the areas of tumor budding [91,92,93,94]. Assessing the expression of lncRNAs at these critical sites could provide insight into the mechanism of their formation and possibly clarify their role in cancer progression.
The bioinformatics analysis uncovered 258 significant correlations of lncRNAs to EMT-TFs on colon and rectum adenocarcinoma datasets. Among the lncRNAs with either strong or very strong correlations to EMT-TFs, only two lncRNAs, namely MAGI2-AS3 and TP73-AS1 have been further validated on CRC cell lines and mouse models [95,96,97]. MAGI2-AS3 promoted CRC progression by facilitating the proliferation and migration of cancer cells in vitro and promoted tumor growth in vivo [95]. Similarly, TP73-AS1 promoted the proliferation, migration and invasion of cancer cells in vitro as well as tumor growth in vivo [96,97]. Overexpression of TP73-AS1 was also associated with metastasis and advanced clinical stages in CRC patients [97]. Interestingly, only ZEB1, ZEB2 and SLUG had strong or very strong correlations with lncRNAs. We can only speculate the reason for this finding. Certain EMT-TFs may have different functions in different types of tissue or may be regulated by other factors with varying intensity. Therefore, it is possible that ZEB1, ZEB2 and SLUG are involved in the regulation of EMT in CRC more strongly than other EMT-TFs, or the correlated lncRNAs have tissue-specific expressions influencing these three EMT-TFs more strongly than others [89,98].
The bioinformatics analysis yielded a high number of correlations between lncRNAs and EMT-TFs in CRC; however, their significance in EMT is uncertain. Any correlation, either positive or negative, suggests that EMT-TFs might be regulated by lncRNAs or vice versa. EMT-TFs are regulated either directly (binding of lncRNAs to the EMT-TF or binding of EMT-TF to the promotor of lncRNAs) or indirectly (e.g., through sponging miRNAs by lncRNAs). There are a huge number of possibilities to be explored in the regulation of EMT-TFs by lncRNAs or vice versa thus influencing the activation of EMT or leading to its partial or full EMT phenotype. In turn, all these possible regulations might be also associated with clinicopathological characteristics that are consequences of EMT [98]. Further validation in vitro and in vivo is required to evaluate their function in EMT and pinpoint those with a potential diagnostic, prognostic and therapeutic value.
There are a few limitations of this review. Firstly, all the studies providing any evidence of lncRNAs mediating an effect on one or more EMT-TFs were included, even though the exact mechanism of action of many lncRNAs was not shown, particularly in cases reporting a change in the expression of multiple EMT-TFs. It is uncertain whether they are all in fact direct targets of the investigated lncRNA or if the regulation is indirect, since it is well known that many EMT-TFs regulate each other as well [10]. Secondly, this review also focuses only on the regulation of core EMT-TFs. Other TFs have also been implicated in the regulation of EMT; however, their functions are less well established. They have either not been shown to induce EMT in cell cultures and animal models or there is insufficient information on whether they induce EMT directly or through the activation of core EMT-TFs [10,99]. Therefore, they were not included in this review.
5. Conclusions
Emerging evidence highlights lncRNAs as regulators of EMT, potentially making them valuable diagnostic markers or therapeutic targets and thus warranting further research. It remains to be seen if their expression patterns and mechanisms of action may contribute to the unresolved issues of EMT, particularly regarding partial EMT states.
Supplementary Materials
The following supporting information can be downloaded at: www.mdpi.com/article/10.3390/cancers14092280/s1. Table S1: Correlations between lncRNAs and TFs obtained with a bioinformatics analysis.
Author Contributions
Conceptualization, A.P. and N.Z.; methodology, A.P. and E.B.; software, N.H.; validation, A.P., E.B., N.H. and N.Z.; formal analysis, A.P. and N.H.; investigation, A.P. and N.H.; resources, E.B. and N.Z.; data curation, A.P. and N.H.; writing—original draft preparation, A.P.; writing—review and editing, A.P., E.B., N.H. and N.Z.; visualization, A.P. and E.B.; supervision, N.Z.; project administration, N.Z.; funding acquisition, E.B. and N.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Slovenian Research Agency (ARRS) under research core funding numbers P3-0054 and J3-1754.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Thiery, J.P.; Acloque, H.; Huang, R.Y.; Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 2009, 139, 871–890. [Google Scholar] [CrossRef] [PubMed]
- Pearson, G.W. Control of invasion by epithelial-to-mesenchymal transition programs during metastasis. J. Clin. Med. 2019, 8, 646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brabletz, S.; Schuhwerk, H.; Brabletz, T.; Stemmler, M.P. Dynamic EMT: A multi-tool for tumor progression. EMBO J. 2021, 40, e108647. [Google Scholar] [CrossRef] [PubMed]
- Dongre, A.; Weinberg, R.A. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat. Rev. Mol. Cell Biol. 2019, 20, 69–84. [Google Scholar] [CrossRef] [PubMed]
- Tarin, D.; Thompson, E.W.; Newgreen, D.F. The fallacy of epithelial mesenchymal transition in neoplasia. Cancer Res. 2005, 65, 6000–6001. [Google Scholar] [CrossRef] [Green Version]
- Zidar, N.; Boštjančič, E.; Gale, N.; Kojc, N.; Poljak, M.; Glavač, D.; Cardesa, A. Down-regulation of microRNAs of the miR-200 family and miR-205, and an altered expression of classic and desmosomal cadherins in spindle cell carcinoma of the head and neck—Hallmark of epithelial-mesenchymal transition. Hum. Pathol. 2011, 42, 482–488. [Google Scholar] [CrossRef]
- Zidar, N.; Boštjančič, E.; Malgaj, M.; Gale, N.; Dovšak, T.; Didanovič, V. The role of epithelial-mesenchymal transition in squamous cell carcinoma of the oral cavity. Virchows Arch. 2018, 472, 237–245. [Google Scholar] [CrossRef]
- Bakir, B.; Chiarella, A.M.; Pitarresi, J.R.; Rustgi, A.K. EMT, MET, plasticity, and tumor metastasis. Trends Cell Biol. 2020, 30, 764–776. [Google Scholar] [CrossRef]
- Pastushenko, I.; Brisebarre, A.; Sifrim, A.; Fioramonti, M.; Revenco, T.; Boumahdi, S.; van Keymeulen, A.; Brown, D.; Moers, V.; Lemaire, S.; et al. Identification of the tumour transition states occurring during EMT. Nature 2018, 556, 463–468. [Google Scholar] [CrossRef]
- Kaller, M.; Hermeking, H. Interplay between transcription factors and microRNAs regulating epithelial-mesenchymal transitions in colorectal cancer. Adv. Exp. Med. Biol. 2016, 937, 71–92. [Google Scholar] [CrossRef]
- Ranković, B.; Zidar, N.; Žlajpah, M.; Boštjančič, E. Epithelial-mesenchymal transition-related microRNAs and their target genes in colorectal cancerogenesis. J. Clin. Med. 2019, 8, 1603. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, H.; Xu, E.; Liu, H.; Wan, L.; Lai, M. Epithelial-mesenchymal transition in colorectal cancer metastasis: A system review. Pathol. Res. Pract. 2015, 211, 557–569. [Google Scholar] [CrossRef] [PubMed]
- Nieto, M.A.; Huang, R.Y.; Jackson, R.A.; Thiery, J.P. EMT: 2016. Cell 2016, 166, 21–45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vu, T.; Datta, P.K. Regulation of EMT in colorectal cancer: A culprit in metastasis. Cancers 2017, 9, 171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davalos, V.; Moutinho, C.; Villanueva, A.; Boque, R.; Silva, P.; Carneiro, F.; Esteller, M. Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene 2012, 31, 2062–2074. [Google Scholar] [CrossRef]
- Goossens, S.; Vandamme, N.; van Vlierberghe, P.; Berx, G. EMT transcription factors in cancer development re-evaluated: Beyond EMT and MET. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 584–591. [Google Scholar] [CrossRef]
- Yang, J.; Antin, P.; Berx, G.; Blanpain, C.; Brabletz, T.; Bronner, M.; Campbell, K.; Cano, A.; Casanova, J.; Christofori, G.; et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2020, 21, 341–352. [Google Scholar] [CrossRef] [Green Version]
- Teeuwssen, M.; Fodde, R. Cell heterogeneity and phenotypic plasticity in metastasis formation: The case of colon cancer. Cancers 2019, 11, 1368. [Google Scholar] [CrossRef] [Green Version]
- O’Brien, S.J.; Bishop, C.; Hallion, J.; Fiechter, C.; Scheurlen, K.; Paas, M.; Burton, J.; Galandiuk, S. Long non-coding RNA (lncRNA) and epithelial-mesenchymal transition (EMT) in colorectal cancer: A systematic review. Cancer Biol. Ther. 2020, 21, 769–781. [Google Scholar] [CrossRef]
- Cavallari, I.; Ciccarese, F.; Sharova, E.; Urso, L.; Raimondi, V.; Silic-Benussi, M.; D’Agostino, D.M.; Ciminale, V. The miR-200 family of microRNAs: Fine tuners of epithelial-mesenchymal transition and circulating cancer biomarkers. Cancers 2021, 13, 5874. [Google Scholar] [CrossRef]
- Gugnoni, M.; Ciarrocchi, A. Long noncoding RNA and epithelial mesenchymal transition in cancer. Int. J. Mol. Sci. 2019, 20, 1924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uhan, S.; Hauptman, N. Metastatic EMT phenotype is governed by microRNA-200-mediated competing endogenous RNA networks. Cells 2021, 11, 73. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, N.; Li, Y.; Li, J.; Gao, Z.; Yang, Z.; Li, Y.; Liu, H.; Fan, T. Long Non-Coding RNAs: The Regulatory Mechanisms, Research Strategies, and Future Directions in Cancers. Front. Oncol. 2020, 10, 598817. [Google Scholar] [CrossRef] [PubMed]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long noncoding RNA and cancer: A new paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef] [Green Version]
- Chen, D.L.; Chen, L.Z.; Lu, Y.X.; Zhang, D.S.; Zeng, Z.L.; Pan, Z.Z.; Huang, P.; Wang, F.H.; Li, Y.H.; Ju, H.Q.; et al. Long noncoding RNA XIST expedites metastasis and modulates epithelial-mesenchymal transition in colorectal cancer. Cell Death Dis. 2017, 8, e3011. [Google Scholar] [CrossRef]
- Cheng, Y.; Chang, Q.; Zheng, B.; Xu, J.; Li, H.; Wang, R. LncRNA XIST promotes the epithelial to mesenchymal transition of retinoblastoma via sponging miR-101. Eur. J. Pharmacol. 2019, 843, 210–216. [Google Scholar] [CrossRef]
- Zheng, R.; Lin, S.; Guan, L.; Yuan, H.; Liu, K.; Liu, C.; Ye, W.; Liao, Y.; Jia, J.; Zhang, R. Long non-coding RNA XIST inhibited breast cancer cell growth, migration, and invasion via miR-155/CDX1 axis. Biochem. Biophys. Res. Commun. 2018, 498, 1002–1008. [Google Scholar] [CrossRef]
- Du, Y.; Weng, X.D.; Wang, L.; Liu, X.H.; Zhu, H.C.; Guo, J.; Ning, J.Z.; Xiao, C.C. LncRNA XIST acts as a tumor suppressor in prostate cancer through sponging miR-23a to modulate RKIP expression. Oncotarget 2017, 8, 94358–94370. [Google Scholar] [CrossRef] [Green Version]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. PLoS Med. 2009, 6, e1000097. [Google Scholar] [CrossRef] [Green Version]
- Colaprico, A.; Silva, T.C.; Olsen, C.; Garofano, L.; Cava, C.; Garolini, D.; Sabedot, T.S.; Malta, T.M.; Pagnotta, S.M.; Castiglioni, I.; et al. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016, 44, e71. [Google Scholar] [CrossRef] [PubMed]
- Duan, W.; Kong, X.; Li, J.; Li, P.; Zhao, Y.; Liu, T.; Binang, H.B.; Wang, Y.; Du, L.; Wang, C. LncRNA AC010789.1 promotes colorectal cancer progression by targeting microRNA-432-3p/ZEB1 axis and the wnt/beta-catenin signaling pathway. Front. Cell Dev. Biol. 2020, 8, 565355. [Google Scholar] [CrossRef] [PubMed]
- Bo, H.; Fan, L.; Li, J.; Liu, Z.; Zhang, S.; Shi, L.; Guo, C.; Li, X.; Liao, Q.; Zhang, W.; et al. High expression of lncRNA AFAP1-AS1 promotes the progression of colon cancer and predicts poor prognosis. J. Cancer 2018, 9, 4677–4683. [Google Scholar] [CrossRef] [PubMed]
- Yan, T.T.; Ren, L.L.; Shen, C.Q.; Wang, Z.H.; Yu, Y.N.; Liang, Q.; Tang, J.Y.; Chen, Y.X.; Sun, D.F.; Zgodzinski, W.; et al. miR-508 defines the stem-like/mesenchymal subtype in colorectal cancer. Cancer Res. 2018, 78, 1751–1765. [Google Scholar] [CrossRef] [Green Version]
- Yue, B.; Qiu, S.; Zhao, S.; Liu, C.; Zhang, D.; Yu, F.; Peng, Z.; Yan, D. LncRNA-ATB mediated E-cadherin repression promotes the progression of colon cancer and predicts poor prognosis. J. Gastroenterol. Hepatol. 2016, 31, 595–603. [Google Scholar] [CrossRef]
- Liang, H.; Wang, J.; Zhang, P.; Yang, W.; Yang, Y.; Zhi, Y.; Wu, W.; Dong, X. Long non-coding RNA Duxap8 facilitates cell proliferation and induces apoptosis in colorectal cancer via miR-519b/ZNF277 axis. Onco Targets Ther. 2021, 14, 4693–4703. [Google Scholar] [CrossRef]
- Yang, C.; Sun, J.; Liu, W.; Yang, Y.; Chu, Z.; Yang, T.; Gui, Y.; Wang, D. Long noncoding RNA HCP5 contributes to epithelial-mesenchymal transition in colorectal cancer through ZEB1 activation and interacting with miR-139-5p. Am. J. Transl. Res. 2019, 11, 953–963. [Google Scholar]
- Bian, Y.; Gao, G.; Zhang, Q.; Qian, H.; Yu, L.; Yao, N.; Qian, J.; Liu, B.; Qian, X. KCNQ1OT1/miR-217/ZEB1 feedback loop facilitates cell migration and epithelial-mesenchymal transition in colorectal cancer. Cancer Biol. Ther. 2019, 20, 886–896. [Google Scholar] [CrossRef]
- Zhang, J.; Ding, L.; Sun, G.; Ning, H.; Huang, R. Suppression of LINC00460 mediated the sensitization of HCT116 cells to ionizing radiation by inhibiting epithelial-mesenchymal transition. Toxicol. Res. 2020, 9, 107–116. [Google Scholar] [CrossRef]
- Ji, L.; Li, X.; Zhou, Z.; Zheng, Z.; Jin, L.; Jiang, F. LINC01413/hnRNP-K/ZEB1 axis accelerates cell proliferation and EMT in colorectal cancer via inducing YAP1/TAZ1 translocation. Mol. Ther. Nucleic Acids 2020, 19, 546–561. [Google Scholar] [CrossRef]
- Rigoutsos, I.; Lee, S.K.; Nam, S.Y.; Anfossi, S.; Pasculli, B.; Pichler, M.; Jing, Y.; Rodriguez-Aguayo, C.; Telonis, A.G.; Rossi, S.; et al. N-BLR, a primate-specific non-coding transcript leads to colorectal cancer invasion and migration. Genome Biol. 2017, 18, 98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, Y.; Yang, X.; Chen, Z.; Tian, L.; Jiang, G.; Chen, F.; Li, J.; An, P.; Lu, L.; Luo, N.; et al. m(6)A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol. Cancer 2019, 18, 87. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Xue, M.; Zhao, Y.; Han, Y.; Li, C.; Zhang, S.; Zhang, J.; Xu, J. Long noncoding RNA ZEB1-AS1 acts as a sponge of miR-141-3p to inhibit cell proliferation in colorectal cancer. Int. J. Med. Sci. 2020, 17, 1589–1597. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.C.; Han, N.; Wu, N.; Zhao, K.L.; Han, C.; Wang, H.X.; Ping, G.F.; Zheng, P.F.; Feng, H.; Qin, L.; et al. Interplay between long noncoding RNA ZEB1-AS1 and miR-101/ZEB1 axis regulates proliferation and migration of colorectal cancer cells. Am. J. Transl. Res. 2018, 10, 605–617. [Google Scholar] [PubMed]
- O’Brien, S.J.; Fiechter, C.; Burton, J.; Hallion, J.; Paas, M.; Patel, A.; Patel, A.; Rochet, A.; Scheurlen, K.; Gardner, S.; et al. Long non-coding RNA ZFAS1 is a major regulator of epithelial-mesenchymal transition through miR-200/ZEB1/E-cadherin, vimentin signaling in colon adenocarcinoma. Cell Death Discov. 2021, 7, 61. [Google Scholar] [CrossRef]
- Ye, C.; Shen, Z.; Wang, B.; Li, Y.; Li, T.; Yang, Y.; Jiang, K.; Ye, Y.; Wang, S. A novel long non-coding RNA lnc-GNAT1-1 is low expressed in colorectal cancer and acts as a tumor suppressor through regulating RKIP-NF-κB-Snail circuit. J. Exp. Clin. Cancer Res. 2016, 35, 187. [Google Scholar] [CrossRef] [Green Version]
- Mo, S.; Zhang, L.; Dai, W.; Han, L.; Wang, R.; Xiang, W.; Wang, Z.; Li, Q.; Yu, J.; Yuan, J.; et al. Antisense lncRNA LDLRAD4-AS1 promotes metastasis by decreasing the expression of LDLRAD4 and predicts a poor prognosis in colorectal cancer. Cell Death Dis. 2020, 11, 155. [Google Scholar] [CrossRef]
- Wang, Y.Q.; Jiang, D.M.; Hu, S.S.; Zhao, L.; Wang, L.; Yang, M.H.; Ai, M.L.; Jiang, H.J.; Han, Y.; Ding, Y.Q.; et al. SATB2-AS1 suppresses colorectal carcinoma aggressiveness by inhibiting SATB2-dependent Snail transcription and epithelial-mesenchymal transition. Cancer Res. 2019, 79, 3542–3556. [Google Scholar] [CrossRef] [Green Version]
- Wang, W.; Tang, X.; Qu, H.; He, Q. Translation regulatory long non-coding RNA 1 represents a potential prognostic biomarker for colorectal cancer. Oncol. Lett. 2020, 19, 4077–4087. [Google Scholar] [CrossRef] [Green Version]
- Yang, P.; Chen, T.; Xu, Z.; Zhu, H.; Wang, J.; He, Z. Long noncoding RNA GAPLINC promotes invasion in colorectal cancer by targeting SNAI2 through binding with PSF and NONO. Oncotarget 2016, 7, 42183–42194. [Google Scholar] [CrossRef]
- Jiang, H.; Li, T.; Qu, Y.; Wang, X.; Li, B.; Song, J.; Sun, X.; Tang, Y.; Wan, J.; Yu, Y.; et al. Long non-coding RNA SNHG15 interacts with and stabilizes transcription factor Slug and promotes colon cancer progression. Cancer Lett. 2018, 425, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhao, L.; Zhang, Y.; Guan, L.; Zhang, H.; Zhou, H.; Gao, T.; Miao, P.; Sun, M. Downregulation of the long non-coding RNA XLOC_010588 inhibits the invasion and migration of colorectal cancer. Oncol. Rep. 2018, 39, 1619–1630. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Niu, H.; Hu, Z.; Liu, H.; Hu, G.; Yang, B.; Wu, S.; Li, F. Long non-coding RNA AK027294 involves in the process of proliferation, migration, and apoptosis of colorectal cancer cells. Tumour Biol. 2016, 37, 10097–10105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tao, Y.; Han, T.; Zhang, T.; Ma, C.; Sun, C. LncRNA CHRF-induced miR-489 loss promotes metastasis of colorectal cancer via TWIST1/EMT signaling pathway. Oncotarget 2017, 8, 36410–36422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, X.; Li, S.; Yu, B.; Kuang, G.; Wu, Y.; Zhang, M.; He, Y.; Ou, C.; Cao, P. Up-regulation of LINC00467 promotes the tumourigenesis in colorectal cancer. J. Cancer 2019, 10, 6405–6413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, N.; Jiang, M.; Liu, H.; Chu, Y.; Wang, D.; Cao, J.; Wang, Z.; Xie, X.; Han, Y.; Xu, B. LINC00941 promotes CRC metastasis through preventing SMAD4 protein degradation and activating the TGF-β/SMAD2/3 signaling pathway. Cell Death Differ. 2021, 28, 219–232. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Huan, L.; Guo, T.; Wu, Y.; Liu, Y.; Wang, Q.; Huang, S.; Xu, Y.; Liang, L.; He, X. LncRNA SNHG11 facilitates tumor metastasis by interacting with and stabilizing HIF-1alpha. Oncogene 2020, 39, 7005–7018. [Google Scholar] [CrossRef]
- Wang, L.; Wei, Z.; Wu, K.; Dai, W.; Zhang, C.; Peng, J.; He, Y. Long noncoding RNA B3GALT5-AS1 suppresses colon cancer liver metastasis via repressing microRNA-203. Aging Albany N. Y. 2018, 10, 3662–3682. [Google Scholar] [CrossRef]
- Yuan, Z.; Yu, X.; Ni, B.; Chen, D.; Yang, Z.; Huang, J.; Wang, J.; Chen, D.; Wang, L. Overexpression of long non-coding RNA-CTD903 inhibits colorectal cancer invasion and migration by repressing Wnt/β-catenin signaling and predicts favorable prognosis. Int. J. Oncol. 2016, 48, 2675–2685. [Google Scholar] [CrossRef] [Green Version]
- He, X.; Yu, B.; Kuang, G.; Wu, Y.; Zhang, M.; Cao, P.; Ou, C. Long noncoding RNA DLEU2 affects the proliferative and invasive ability of colorectal cancer cells. J. Cancer 2021, 12, 428–437. [Google Scholar] [CrossRef]
- Liang, W.C.; Fu, W.M.; Wong, C.W.; Wang, Y.; Wang, W.M.; Hu, G.X.; Zhang, L.; Xiao, L.J.; Wan, D.C.; Zhang, J.F.; et al. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 2015, 6, 22513–22525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhong, M.E.; Chen, Y.; Zhang, G.; Xu, L.; Ge, W.; Wu, B. LncRNA H19 regulates PI3K-Akt signal pathway by functioning as a ceRNA and predicts poor prognosis in colorectal cancer: Integrative analysis of dysregulated ncRNA-associated ceRNA network. Cancer Cell Int. 2019, 19, 148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.W.; Zhu, J.; Ma, J.; Zhang, J.L.; Zuo, S.; Chen, G.W.; Wang, X.; Pan, Y.S.; Liu, Y.C.; Wang, P.Y. Overexpression of long non-coding RNA H19 is associated with unfavorable prognosis in patients with colorectal cancer and increased proliferation and migration in colon cancer cells. Oncol. Lett. 2017, 14, 2446–2452. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Li, C.; Zhao, T.; Li, D.; Yang, L.; Zhang, B. LncRNA H19/miR-29b-3p/PGRN axis promoted epithelial-mesenchymal transition of colorectal cancer cells by acting on wnt signaling. Mol. Cells 2018, 41, 423–435. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Pan, Y.L.; Zhang, J.; Cao, P.G. LncRNA HOTAIR recruits SNAIL to inhibit the transcription of HNF4α and promote the viability, migration, invasion and EMT of colorectal cancer. Transl. Oncol. 2021, 14, 101036. [Google Scholar] [CrossRef]
- Chen, D.; Zhang, M.; Ruan, J.; Li, X.; Wang, S.; Cheng, X.; Zhao, H.; Zeng, Y.; Liu, J.; He, K.; et al. The long non-coding RNA HOXA11-AS promotes epithelial mesenchymal transition by sponging miR-149-3p in colorectal cancer. J. Cancer 2020, 11, 6050–6058. [Google Scholar] [CrossRef]
- Sun, Z.; Shao, B.; Liu, Z.; Dang, Q.; Guo, Y.; Chen, C.; Guo, Y.; Chen, Z.; Liu, J.; Hu, S.; et al. LINC01296/miR-141-3p/ZEB1-ZEB2 axis promotes tumor metastasis via enhancing epithelial-mesenchymal transition process. J. Cancer 2021, 12, 2723–2734. [Google Scholar] [CrossRef]
- Yang, T.; Chen, W.C.; Shi, P.C.; Liu, M.R.; Jiang, T.; Song, H.; Wang, J.Q.; Fan, R.Z.; Pei, D.S.; Song, J. Long noncoding RNA MAPKAPK5-AS1 promotes colorectal cancer progression by cis-regulating the nearby gene MK5 and acting as a let-7f-1-3p sponge. J. Exp. Clin. Cancer Res. 2020, 39, 139. [Google Scholar] [CrossRef]
- Dong, X.; Yang, Z.; Yang, H.; Li, D.; Qiu, X. Long non-coding RNA MIR4435-2HG promotes colorectal cancer proliferation and metastasis through miR-206/YAP1 axis. Front. Oncol. 2020, 10, 160. [Google Scholar] [CrossRef] [Green Version]
- Lu, M.; Liu, Z.; Li, B.; Wang, G.; Li, D.; Zhu, Y. The high expression of long non-coding RNA PANDAR indicates a poor prognosis for colorectal cancer and promotes metastasis by EMT pathway. J. Cancer Res. Clin. Oncol. 2017, 143, 71–81. [Google Scholar] [CrossRef]
- Wang, X.; Lai, Q.; He, J.; Li, Q.; Ding, J.; Lan, Z.; Gu, C.; Yan, Q.; Fang, Y.; Zhao, X.; et al. LncRNA SNHG6 promotes proliferation, invasion and migration in colorectal cancer cells by activating TGF-β/Smad signaling pathway via targeting UPF1 and inducing EMT via regulation of ZEB1. Int. J. Med. Sci. 2019, 16, 51–59. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, M.; Duan, W.; Sun, W. LncRNA SNHG6 promotes the migration, invasion, and epithelial-mesenchymal transition of colorectal cancer cells by miR-26a/EZH2 axis. Onco Targets Ther. 2019, 12, 3349–3360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, Z.; Bi, M.; Zhang, Q.; Song, Y.; Hong, S. LncRNA TUG1 promotes the progression of colorectal cancer via the miR-138-5p/ZEB2 axis. Biosci. Rep. 2020, 40, BSR20201025. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.; Hu, X.; Mao, J.; Wu, Y.; Liu, H.; Shen, J.; Yu, J.; Chen, W. The long noncoding RNA TUG1 is required for TGF-β/TWIST1/EMT-mediated metastasis in colorectal cancer cells. Cell Death Dis. 2020, 11, 65. [Google Scholar] [CrossRef] [Green Version]
- Chen, D.L.; Lu, Y.X.; Zhang, J.X.; Wei, X.L.; Wang, F.; Zeng, Z.L.; Pan, Z.Z.; Yuan, Y.F.; Wang, F.H.; Pelicano, H.; et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics 2017, 7, 4836–4849. [Google Scholar] [CrossRef]
- Rezanejad Bardaji, H.; Asadi, M.H.; Yaghoobi, M.M. Long noncoding RNA VIM-AS1 promotes colorectal cancer progression and metastasis by inducing EMT. Eur. J. Cell Biol. 2018, 97, 279–288. [Google Scholar] [CrossRef]
- Jiang, Y.; Liu, G.; Ye, W.; Xie, J.; Shao, C.; Wang, X.; Li, X. ZEB2-AS1 accelerates epithelial/mesenchymal transition through miR-1205/CRKL pathway in colorectal cancer. Cancer Biother. Radiopharm. 2020, 35, 153–162. [Google Scholar] [CrossRef] [Green Version]
- Mount, J.; Zumel, N. Sigr: Succinct and Correct Statistical Summaries for Reports. 2017. Available online: https://rdrr.io/cran/sigr/ (accessed on 11 January 2022).
- Spaderna, S.; Schmalhofer, O.; Hlubek, F.; Berx, G.; Eger, A.; Merkel, S.; Jung, A.; Kirchner, T.; Brabletz, T. A transient, EMT-linked loss of basement membranes indicates metastasis and poor survival in colorectal cancer. Gastroenterology 2006, 131, 830–840. [Google Scholar] [CrossRef]
- Fan, F.; Samuel, S.; Evans, K.W.; Lu, J.; Xia, L.; Zhou, Y.; Sceusi, E.; Tozzi, F.; Ye, X.C.; Mani, S.A.; et al. Overexpression of snail induces epithelial-mesenchymal transition and a cancer stem cell-like phenotype in human colorectal cancer cells. Cancer Med. 2012, 1, 5–16. [Google Scholar] [CrossRef]
- García-Palmero, I.; Torres, S.; Bartolomé, R.A.; Peláez-García, A.; Larriba, M.J.; Lopez-Lucendo, M.; Peña, C.; Escudero-Paniagua, B.; Muñoz, A.; Casal, J.I. Twist1-induced activation of human fibroblasts promotes matrix stiffness by upregulating palladin and collagen α1(VI). Oncogene 2016, 35, 5224–5236. [Google Scholar] [CrossRef]
- Celesti, G.; di Caro, G.; Bianchi, P.; Grizzi, F.; Basso, G.; Marchesi, F.; Doni, A.; Marra, G.; Roncalli, M.; Mantovani, A.; et al. Presence of Twist1-positive neoplastic cells in the stroma of chromosome-unstable colorectal tumors. Gastroenterology 2013, 145, 647–657.e615. [Google Scholar] [CrossRef] [PubMed]
- Hwang, W.L.; Yang, M.H.; Tsai, M.L.; Lan, H.Y.; Su, S.H.; Chang, S.C.; Teng, H.W.; Yang, S.H.; Lan, Y.T.; Chiou, S.H.; et al. SNAIL regulates interleukin-8 expression, stem cell-like activity, and tumorigenicity of human colorectal carcinoma cells. Gastroenterology 2011, 141, 279–291. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Wang, L.; Pappan, L.; Galliher-Beckley, A.; Shi, J. IL-1beta promotes stemness and invasiveness of colon cancer cells through Zeb1 activation. Mol. Cancer 2012, 11, 87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, C.; Su, L.; Shan, J.; Zhu, C.; Liu, L.; Liu, C.; Xu, Y.; Yang, Z.; Bian, X.; Shao, J.; et al. IGF/STAT3/NANOG/Slug signaling axis simultaneously controls epithelial-mesenchymal transition and stemness maintenance in colorectal cancer. Stem Cells 2016, 34, 820–831. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Ma, J.; Deng, G.; Qu, Y.; Yin, L.; Li, Y.; Han, Y.; Cai, C.; Shen, H.; Zeng, S. ZEB1 promotes oxaliplatin resistance through the induction of epithelial—Mesenchymal transition in colon cancer cells. J. Cancer 2017, 8, 3555–3566. [Google Scholar] [CrossRef] [Green Version]
- Chu, P.Y.; Hu, F.W.; Yu, C.C.; Tsai, L.L.; Yu, C.H.; Wu, B.C.; Chen, Y.W.; Huang, P.I.; Lo, W.L. Epithelial-mesenchymal transition transcription factor ZEB1/ZEB2 co-expression predicts poor prognosis and maintains tumor-initiating properties in head and neck cancer. Oral Oncol. 2013, 49, 34–41. [Google Scholar] [CrossRef]
- Diener, C.; Keller, A.; Meese, E. Emerging concepts of miRNA therapeutics: From cells to clinic. Trends Genet. 2022. [Google Scholar] [CrossRef]
- Sinha, D.; Saha, P.; Samanta, A.; Bishayee, A. Emerging concepts of hybrid epithelial-to-mesenchymal transition in cancer progression. Biomolecules 2020, 10, 1561. [Google Scholar] [CrossRef]
- Blank, A.; Roberts, D.E., 2nd; Dawson, H.; Zlobec, I.; Lugli, A. Tumor heterogeneity in primary colorectal cancer and corresponding metastases. Does the apple fall far from the tree? Front. Med. 2018, 5, 234. [Google Scholar] [CrossRef] [Green Version]
- Pavlic, A.; Urh, K.; Stajer, K.; Bostjancic, E.; Zidar, N. Epithelial-mesenchymal transition in colorectal carcinoma: Comparison between primary tumor, lymph node and liver metastases. Front. Oncol. 2021, 11, 662806. [Google Scholar] [CrossRef]
- Paterson, E.L.; Kazenwadel, J.; Bert, A.G.; Khew-Goodall, Y.; Ruszkiewicz, A.; Goodall, G.J. Down-regulation of the miRNA-200 family at the invasive front of colorectal cancers with degraded basement membrane indicates EMT is involved in cancer progression. Neoplasia 2013, 15, 180–191. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, K.N.; Lindebjerg, J.; Nielsen, B.S.; Hansen, T.F.; Sorensen, F.B. MicroRNA-200b is downregulated in colon cancer budding cells. PLoS ONE 2017, 12, e0178564. [Google Scholar] [CrossRef] [PubMed]
- Hur, K.; Toiyama, Y.; Takahashi, M.; Balaguer, F.; Nagasaka, T.; Koike, J.; Hemmi, H.; Koi, M.; Boland, C.R.; Goel, A. MicroRNA-200c modulates epithelial-to-mesenchymal transition (EMT) in human colorectal cancer metastasis. Gut 2013, 62, 1315–1326. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, H.; Li, Z.; Tang, Z.; Li, J.; Lang, X. Long noncoding MAGI2-AS3 promotes colorectal cancer progression through regulating miR-3163/TMEM106B axis. J. Cell. Physiol. 2020, 235, 4824–4833. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Yan, P.; Zhang, G.; Yang, W.; Wang, H.; Cheng, X. Long non-coding RNA TP73-AS1 sponges miR-194 to promote colorectal cancer cell proliferation, migration and invasion via up-regulating TGFα. Cancer Biomark. 2018, 23, 145–156. [Google Scholar] [CrossRef]
- Chen, C.; Shu, L.; Zou, W. Role of long non-coding RNA TP73-AS1 in cancer. Biosci. Rep. 2019, 39, BSR20192274. [Google Scholar] [CrossRef]
- Jia, Z.; An, J.; Liu, Z.; Zhang, F. Non-coding RNAs in colorectal cancer: Their functions and mechanisms. Front. Oncol. 2022, 12, 783079. [Google Scholar] [CrossRef]
- Takahashi, Y.; Sawada, G.; Kurashige, J.; Uchi, R.; Matsumura, T.; Ueo, H.; Takano, Y.; Akiyoshi, S.; Eguchi, H.; Sudo, T.; et al. Paired related homoeobox 1, a new EMT inducer, is involved in metastasis and poor prognosis in colorectal cancer. Br. J. Cancer 2013, 109, 307–311. [Google Scholar] [CrossRef] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).