Simple Summary
This study explored how genetic changes influence treatment effectiveness and survival in patients with advanced pancreatic cancer. By examining tumors from 142 patients, researchers identified specific genetic mutations that can predict how well a patient responds to chemotherapy. Findings showed that mutations in certain genetic pathways are associated with longer survival and better treatment outcomes. This research highlights the importance of tailoring cancer treatment based on individual genetic profiles, potentially leading to more personalized and effective therapies for pancreatic cancer patients.
Abstract
(1) Background: Pancreatic ductal adenocarcinoma (PDAC) has low survival rates despite treatment advancements. Aim: This study aims to show how molecular profiling could possibly guide personalized treatment strategies, which may help improve survival outcomes in patients with PDAC. (2) Materials and Methods: A retrospective analysis of 142 PDAC patients from a single academic center was conducted. Patients underwent chemotherapy and next-generation sequencing for molecular profiling. Key oncogenic pathways were identified using the Reactome pathway database. Survival analysis was performed using Kaplan–Meier curves and Cox Proportional Hazards Regression. (3) Results: Patients mainly received FOLFIRINOX (n = 62) or gemcitabine nab-paclitaxel (n = 62) as initial chemotherapy. The median OS was 13.6 months. Longer median OS was noted in patients with NOTCH (15 vs. 12.3 months, p = 0.007) and KIT pathway mutations (21.3 vs. 12.12 months, p = 0.04). Combinatorial pathway analysis indicated potential synergistic effects on survival. In the PFS, PI3K pathway (6.6 vs. 5.7 months, p = 0.03) and KIT pathway (10.3 vs. 6.2 months, p = 0.03) mutations correlated with improved PFS within the gemcitabine nab-paclitaxel subgroup. (4) Conclusions: Molecular profiling could play a role in PDAC for predicting outcomes and responses to therapies like FOLFIRINOX and gemcitabine nab-paclitaxel. Integrating genomic data into clinical decision-making can benefit PDAC treatment, though further validation is needed to fully utilize precision oncology in PDAC management.
1. Introduction
Pancreatic cancer presents a significant challenge as it is the third-leading cause of cancer-related deaths [1], with a rising incidence that is expected to make it the second-leading cause by 2030 [2]. Despite advancements in therapy, the 5-year survival rate remains low at 11.5% [3]. While surgical resection offers a curative option, it is feasible for only a small percentage of patients because this cancer often presents at an advanced stage, limiting the potential for curative interventions. [4,5]. Despite extensive study, immune–oncology agents have had little benefit in pancreatic ductal adenocarcinoma (PDAC), and targeted treatment based on molecular pathological tumor characteristics has yet to be as broadly beneficial as they have been in other tumor types [6]. Aggressive cytotoxic chemotherapy remains the primary treatment option for unresectable pancreatic tumors, despite its relatively low response rate [7]. Treatment regimens for PDAC vary based on the patient’s performance status [8]. FOLFIRINOX, which includes infusional 5-fluorouracil, leucovorin, irinotecan, and oxaliplatin, is preferred for patients with good performance status (ECOG 0-1) due to its superior efficacy compared to gemcitabine alone. For patients with a slightly lower performance status (ECOG 1-2), the combination of gemcitabine plus nab-paclitaxel is another frontline option.
PDAC exhibits significant molecular heterogeneity, with an average of from 60 to 70 genetic changes observed [9]. Efforts have been made to categorize pancreatic cancer based on molecular activity and phenotypical aspects. However, due to the complexity of the disease, a consensus has not been reached [10,11,12,13,14]. Nonetheless, profiling studies have identified potentially actionable molecular alterations in up to 25% of pancreatic cancers [15]. The advent of next-generation sequencing (NGS) has revolutionized genomic profiling and made it widely accessible, and it plays a crucial role in pancreatic cancer research and clinical decision-making [16,17,18]. Leveraging NGS technology can lead to a more comprehensive understanding of the molecular landscape of pancreatic cancer, paving the way for improved treatments and future advancements in the field [11].
In this study, we retrospectively examined the molecular profiles of unresectable PDAC patients, aiming to elucidate the relationship between specific genetic alterations, natural history of disease, and treatment responses. Our focus was on understanding how the presence of mutations in key oncogenic pathways might influence the overall survival and efficacy of first-line chemotherapy regimens, such as FOLFIRINOX and gemcitabine nab-paclitaxel. By integrating molecular data with clinical outcomes, this study seeks to contribute to the burgeoning field of precision oncology, offering insights into personalized treatment strategies that could potentially improve survival and quality of life for patients with this formidable disease.
2. Materials and Methods
This study is a retrospective chart review designed to investigate the role of molecular profiling in predicting treatment response and survival in patients with unresectable pancreatic ductal adenocarcinoma.
Study Population: The study population consisted of patients diagnosed with PDAC who presented to the medical oncology clinic of a single center (Beth Israel Deaconess Medical Center, Boston MA) between the years 2013 and 2022. Only those patients with next-generation tumor sequencing who underwent cytotoxic chemotherapy were selected for inclusion. The sample size was calculated to ensure a power of 0.80 and a significance level of 0.05. Patients who were included followed these characteristics: diagnosed with unresectable PDAC (Stage III or IV) [19], underwent NGS for molecular profiling, and received first-line systemic cytotoxic chemotherapy (FOLFIRINOX or gemcitabine nab-paclitaxel). Of those, we obtained a sample of 142 patients.
Data Collection: A targeted chart review was performed which captured demographic information, age at diagnosis, performance status at diagnosis, tumor histology, and first-line chemotherapy regimen. Although we captured all first-line chemotherapy regimens, most patients received standard-of-care treatment with either FOLFIRINOX [20,21] or gemcitabine nab-paclitaxel [20,22]. Disease progression was assessed using the Response Evaluation Criteria in Solid Tumors (RECIST) criteria, as determined by provider documentation [23].
Molecular Profiling: The genetic profile of each tumor sample was obtained through next-generation tumor sequencing using the Foundation Medicine CDX Report [24]. This analysis includes 324 genes that are potentially cancer-related, of which 225 were altered in at least one sample within our selected patient population. These genes were grouped into 14 different pathways (Appendix A) based on their biological function using the Reactome Pathway database [25]; some genes may be members of multiple pathways. Each patient sample was classified by the altered genes present, and then by the pathways represented by those mutations. We recorded whether mutations were present in these pathways as a binary yes-or-no variable. Any insertion, deletion, substitution, duplication, rearrangement, truncation, or amplification was considered a mutated state [26]. All data were collected and managed using REDCap electronic data capture hosted at Beth Israel Deaconess Medical Center [27,28]. This study was approved by the BIDMC IRB and determined to be exempt from informed consent requirements.
Statistical Analysis: We conducted two distinct survival analyses based on the presence or absence of specific mutated pathways: overall survival and progression-free survival. We performed a sub-group analysis of the patients who received FOLFIRINOX or gemcitabine nab-paclitaxel as a first-line therapy. We used Kaplan–Meier curves and log-rank tests to compare survival between groups with and without each specific mutated pathway. We also performed a Cox Proportional Hazards Regression analysis to adjust for potential confounding variables. The threshold for statistical significance was set at p < 0.05. All analyses were performed using R Statistical Software (v4.2.2; R Core Team 2021) via the survival package [29,30].
3. Results
3.1. Characteristics of the Patient Population
This study included 142 patients with a median age of 66 years who were balanced in gender (49% male, 51% female). Most patients were diagnosed at advanced stages (37% Stage III, 63% Stage IV). The majority underwent standard first-line chemotherapy, with equal distribution between FOLFIRINOX (44%) and gemcitabine nab-paclitaxel (44%) (Table 1).

Table 1.
Demographic and sample characteristics.
3.2. Characteristics of the Pathology Samples
The majority of samples were tissue samples obtained from the primary tumor (68, 48%) or a metastatic site (68, 48%), while a small number were from peripheral blood (6, 4%). Tumor mutation burden (TMB) status was known for 110 patients, with a median value of 2.5 (IQR 1.3–3.8), and 4 patients classified as TMB-High. MSI status was known for 140 patients, with 1 classified as MSI-H.
The comprehensive genomic analysis of the 142 PDAC patient samples revealed a significant degree of mutational burden, with an average of 7.83 mutations per sample, ranging from 1 to 35 mutations. Predominantly, mutations were observed in the MAPK-RAS pathway in 94% of patients (n = 134) and the TP53 pathway in 87% (n = 123). Other frequently altered pathways included the Cell Cycle (CC) pathway in 76% of patients (n = 108), DNA Damage Repair (DDR) in 49% (n = 70), TGF-beta pathway in 46% (n = 65), PI3K pathway in 37% (n = 52), and NOTCH pathway in 36% (n = 51). Less commonly altered pathways comprised FGFR (23%, n = 32), WNT (20%, n = 29), PDGFR (18%, n = 25), KIT (16%, n = 23), ERBB2 (14%, n = 20), ALK (13%, n = 19), and FTL3 (6%, n = 9). These findings highlight the extensive molecular diversity present in PDAC and underscore the potential for targeted therapeutic strategies based on individual mutational profiles.
3.3. Overall Survival Analysis
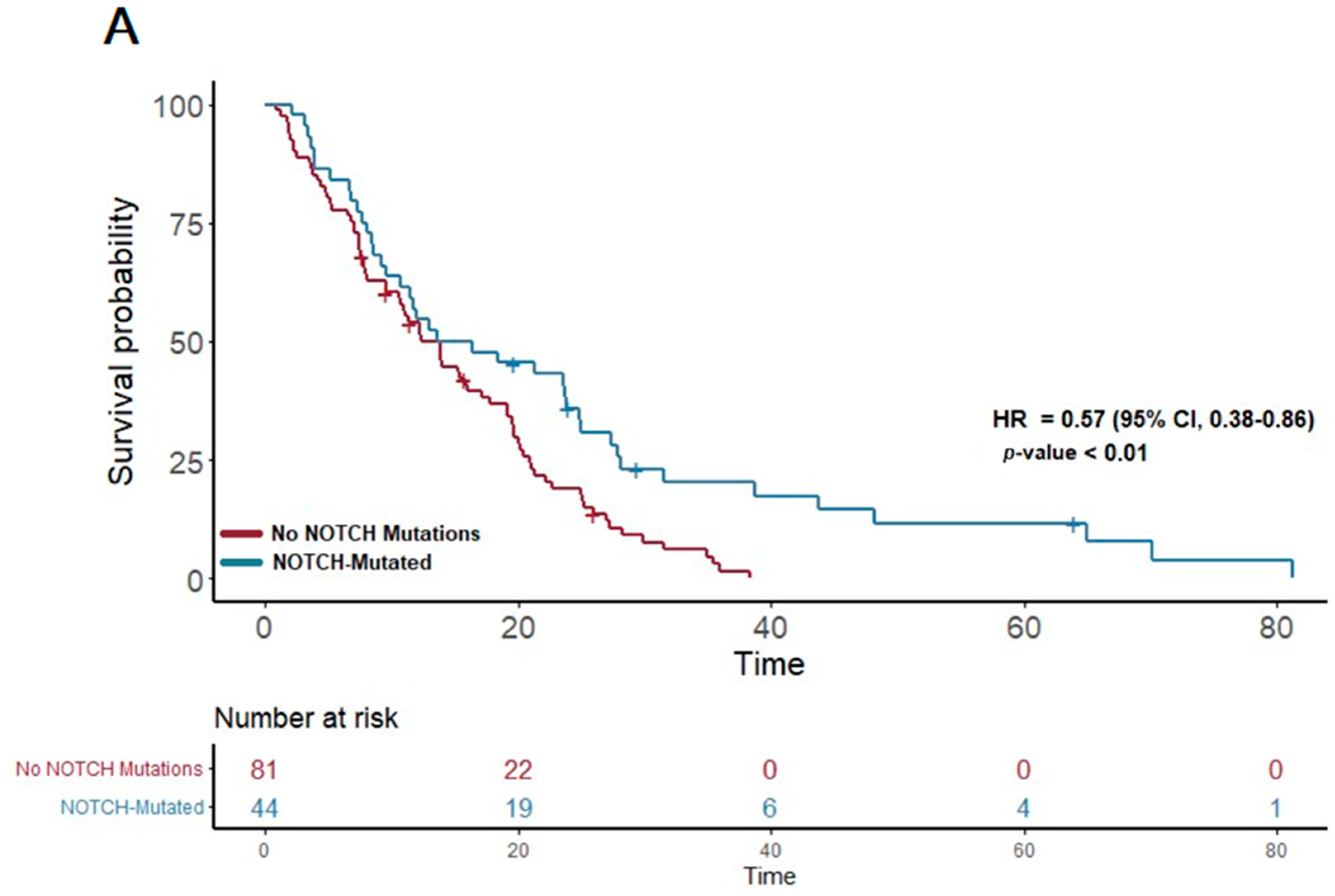
In our study cohort, the median overall survival (OS) was found to be 13.6 months. A detailed analysis of the genetic profiles revealed significant survival advantages in specific mutational pathways. Notably, patients exhibiting mutations in the NOTCH pathway (n = 51) demonstrated a median OS of 15 months, markedly longer than the 12.3 months observed in those without NOTCH mutations (p = 0.007; HR 0.57; 95% CI 0.38–0.86) (Figure 1A). This suggests a potential protective effect of NOTCH pathway alterations on survival.
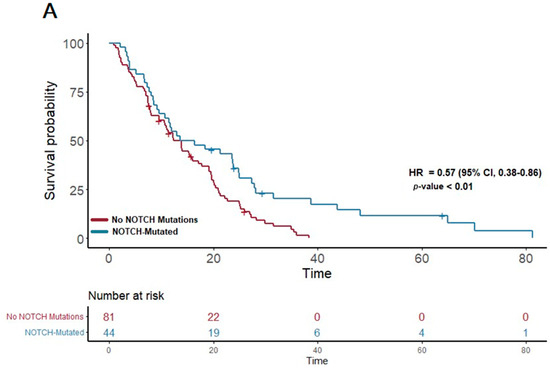
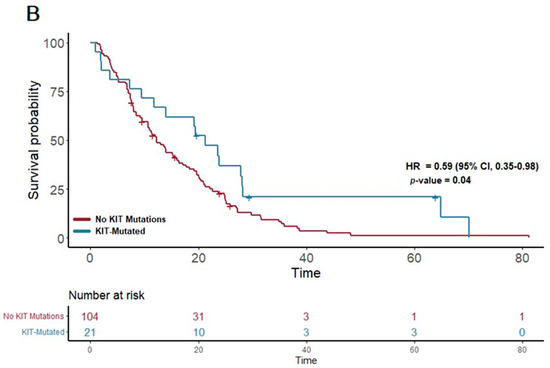
Figure 1.
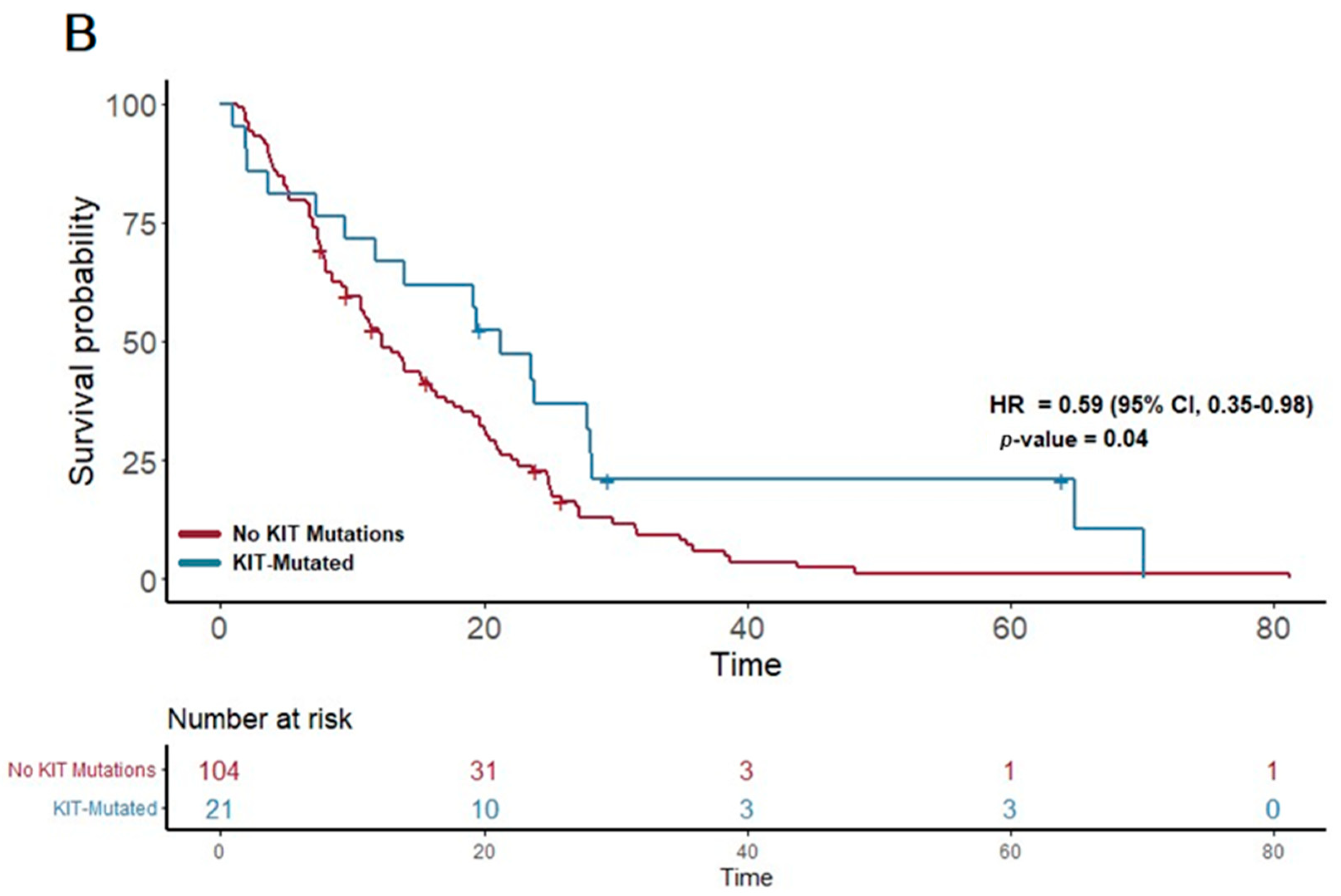
(A) The first panel shows overall survival (OS) for patients with NOTCH mutations (median OS: 15 months) versus those without (median OS: 12.3 months) (p = 0.007; HR 0.57; 95% CI 0.38–0.86). (B) The second panel shows OS for patients with KIT mutations (median OS: 21.3 months) versus those without (median OS: 12.12 months) (p = 0.04; HR 0.59; 95% CI 0.35–0.98).
Furthermore, a similar trend was observed in the KIT pathway mutations. Among the 23 patients with KIT pathway mutations, the median OS was significantly extended to 21.3 months, compared to 12.12 months in patients without these mutations (p = 0.04; HR 0.59; 95% CI 0.35–0.98) (Figure 1B). This finding suggests a notable survival benefit associated with KIT pathway alterations.
In contrast, other individual pathways did not show a statistically significant association with OS in our cohort. However, an interesting non-significant trend towards improved OS was noted in patients with mutations in the ALK pathway, where the median OS was 16.7 months compared to the overall cohort median of 13.6 months (p = 0.06, HR = 0.57, 95% CI 0.32–1.03). This observation warrants further investigation to elucidate the potential role of ALK pathway mutations in PDAC survival.
We further tested all two-way combinations of mutations among the 14 pathways. In the two-way combination analysis (Table 2 and Figure 2), mutations in the ALK, KIT, ERBB2, and PDGFR pathways could be synergistic with NOTCH, showing increases in overall survival (median OS of 25, 24, 25, and 23 months vs. 15 months for NOTCH overall). Combinatoric analysis with the KIT pathway revealed a potentially negative effect of combination with the DDR pathway (median OS of 11 months vs. 21 months for KIT overall). Although the ALK pathway overall did not show a statistically significant increase in OS, in combination with TGFB (median OS 28 months) and TP53 (median OS 24 months) it did achieve significance in the combination analysis.

Table 2.
Overall survival of mutations in patient samples and combination analysis.
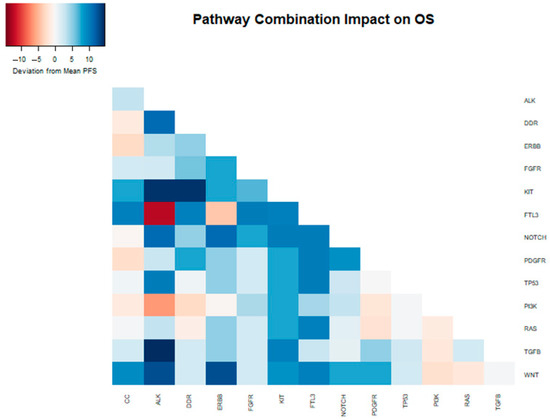
Figure 2.
Heatmap depicting the deviation from the median OS (13.6 months) within the entire sample, attributed to the presence of distinct mutations and their combinations in the analyzed samples.
3.4. Progression-Free Survival Analysis
In our study, we examined the progression-free survival (PFS) among the 142 patients with PDAC, specifically analyzing differences in PFS based on first-line chemotherapy regimens and genetic mutations. The median PFS for patients treated with FOLFIRINOX (n = 62) was 9.1 months, while those receiving gemcitabine nab-paclitaxel (n = 62) had a median PFS of 6.3 months.
For the FOLFIRINOX cohort, the analysis did not reveal any significant correlation between the presence of specific mutational pathways and improvements in PFS. Even with the exploration of potential two-way interactions between different pathways, no statistically significant results emerged that suggested a benefit in PFS from combined mutations in the FOLFIRINOX-treated group.
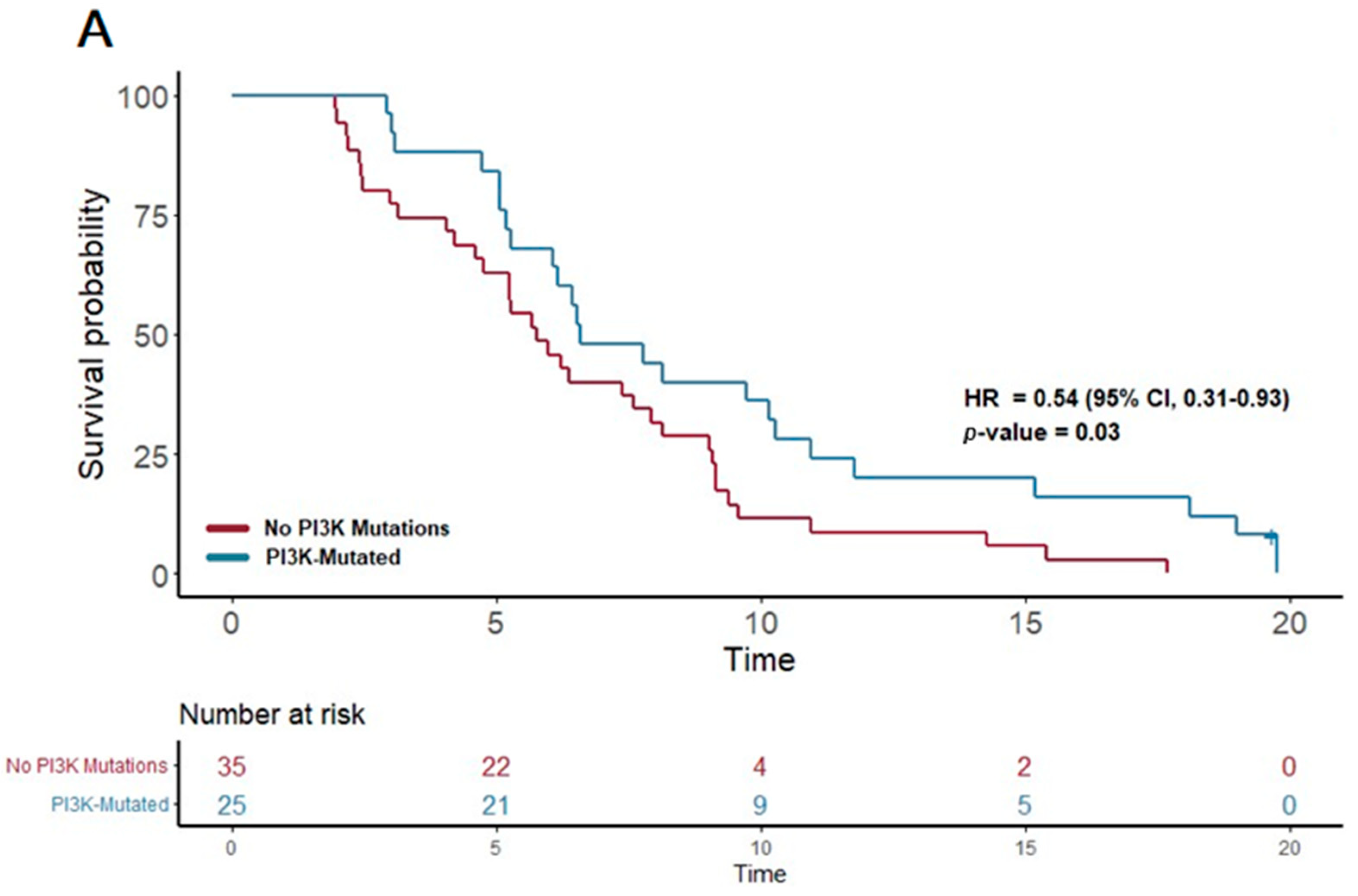
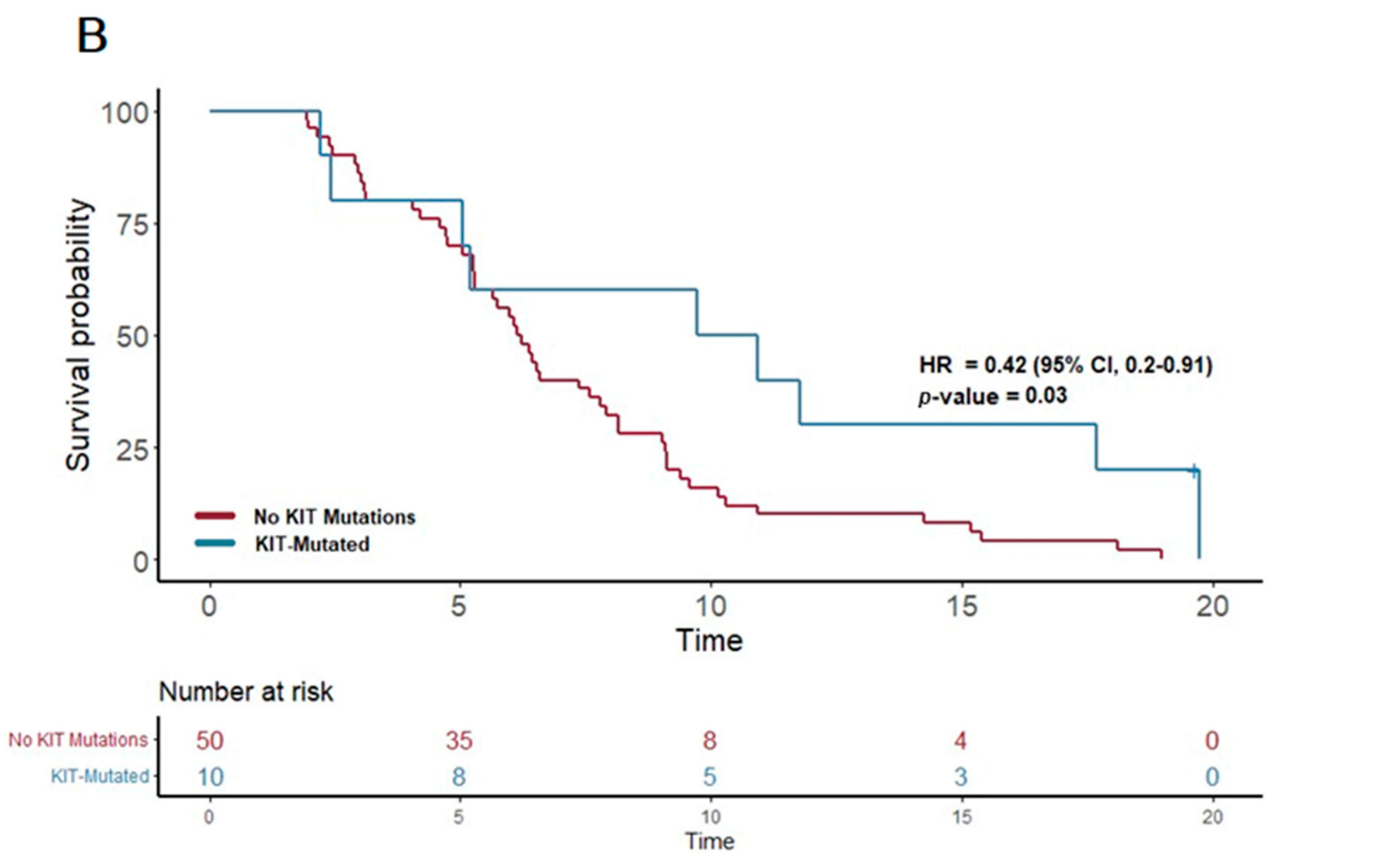
In contrast, the patients treated with gemcitabine nab-paclitaxel exhibited notable associations between certain genetic alterations and PFS. Notably, the presence of a PI3K pathway mutation (found in 25 patients) was associated with a significantly extended PFS of 6.6 months, compared to 5.7 months in patients without such mutations (p = 0.03; HR = 0.54, 95% CI 0.31–0.93). Furthermore, mutations in the KIT pathway (observed in 10 patients) correlated with an even more substantial increase in PFS, reaching 10.3 months versus 6.2 months in non-mutated cases (p = 0.03; HR 0.42, 95% CI 0.2–0.91). Additionally, mutations in the FTL3 pathway (n = 4) were linked to the longest median PFS of 13.3 months, a significant improvement compared to 6.2 months in those without FTL3 mutations (p = 0.02; HR 0.21, 95% CI 0.05–0.89).
These results highlight the potential of certain genetic mutations, particularly within the PI3K, KIT, and FTL3 pathways, to predict longer progression-free survival in PDAC patients receiving gemcitabine nab-paclitaxel as their first-line treatment. (Figure 3) This insight underscores the value of molecular profiling in guiding therapy choices and tailoring treatment strategies to individual patient profiles.
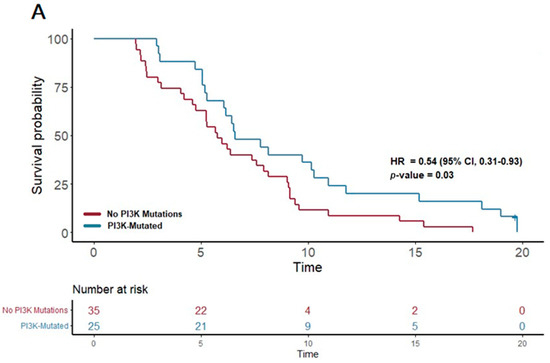
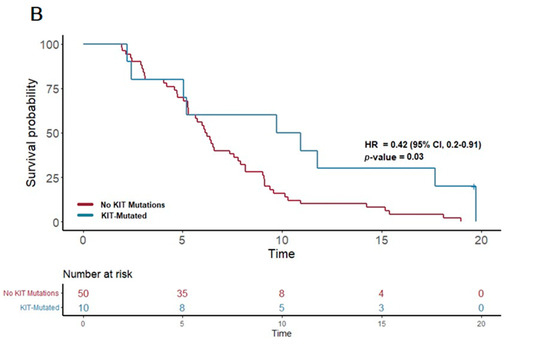
Figure 3.
Kaplan–Meier plots for PFS in patients that received first-line therapy gemcitabine nab-paclitaxel. (A) The first panel shows PFS for patients with PI3K mutations (median PFS: 6.6 months) versus those without (median PFS: 5.7 months) (p = 0.03; HR 0.54; 95% CI 0.31–0.93). (B) The second panel shows PFS for patients with KIT mutations (median PFS: 10.3 months) versus those without (median PFS: 6.2 months) (p = 0.03; HR 0.42; 95% CI 0.2–0.91).
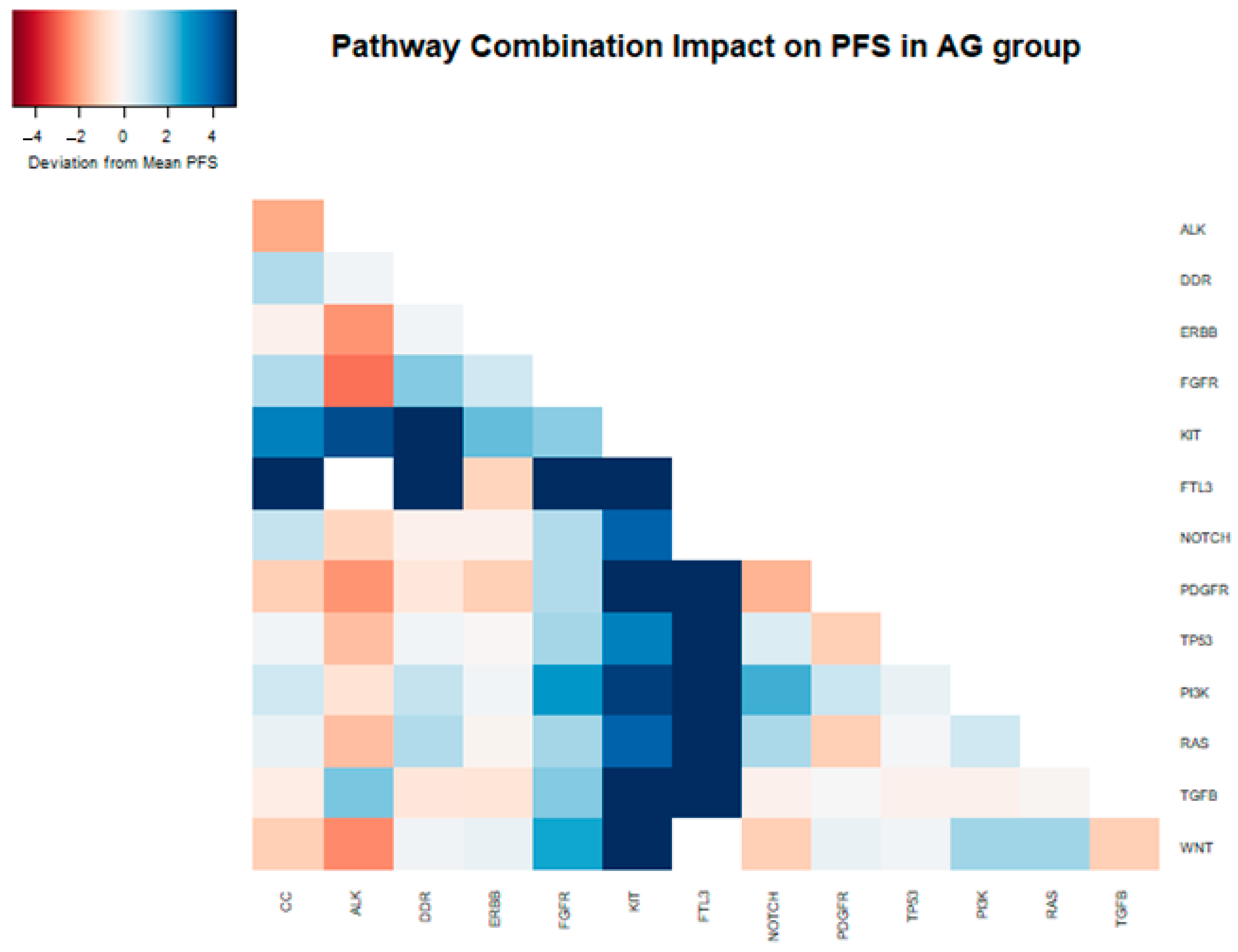
In a two-way combination analysis, these three pathways (PI3K, KIT, and FTL3) could be synergistic for response to gemcitabine nab-paclitaxel, with a longer PFS in combination (PI3K + KIT 10.9 months, PI3K + FTL3 13.1 months, KIT + FTL3 19.7 months). In addition, DDR pathway mutations in combination with KIT or FTL3 (but not PI3K) appeared to provide additional PFS benefits (DDR + KIT 14.7 months, DDR + FTL3 19.7 months) (Table 3 and Figure 4).

Table 3.
Progression-free survival of mutations and combination analysis in patient samples with gemcitabine nab-paclitaxel as first-line therapy.
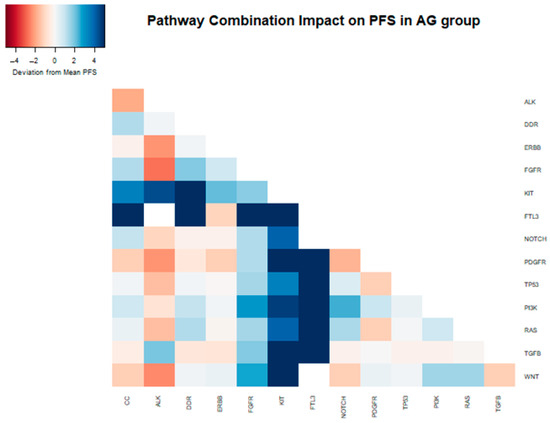
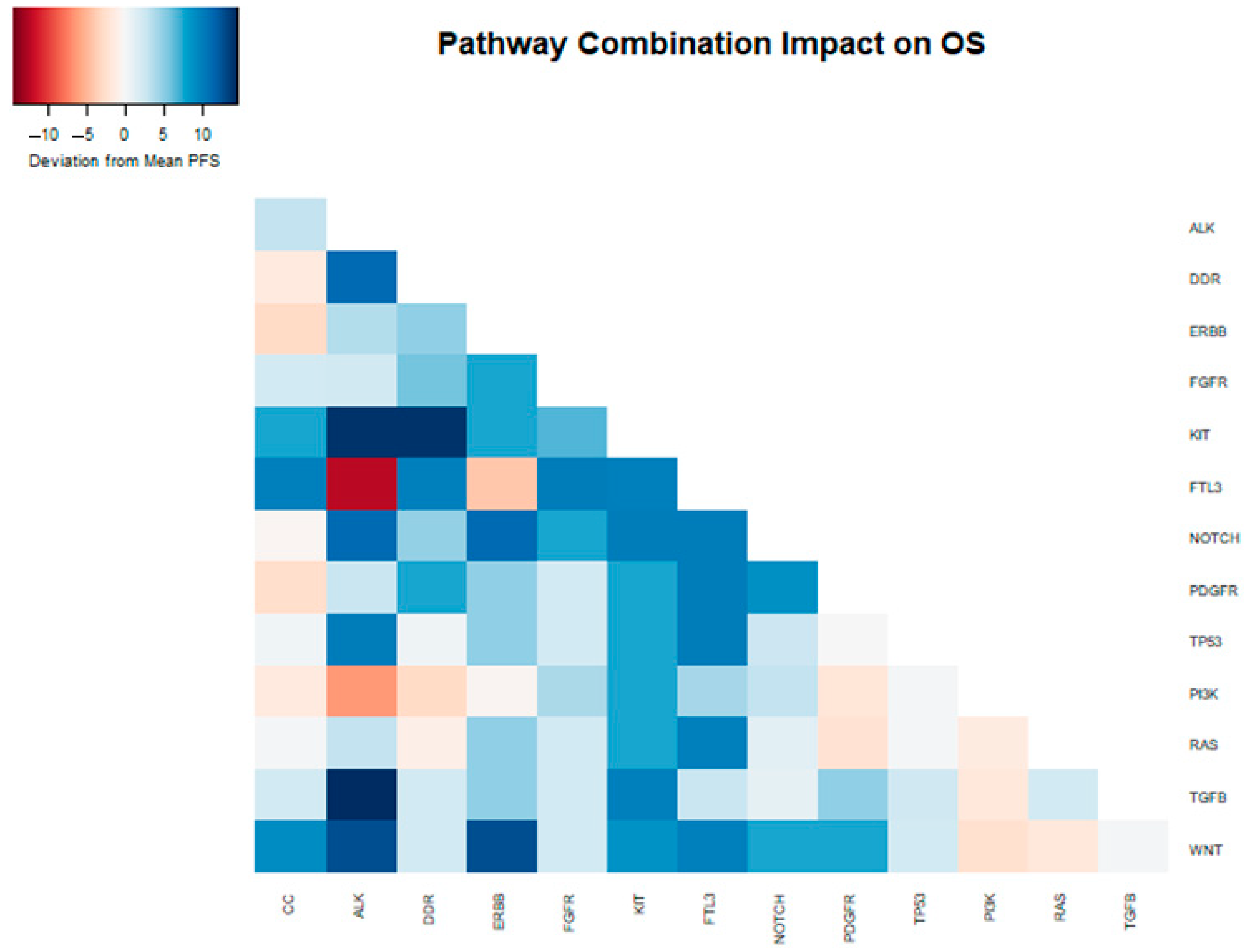
Figure 4.
Heatmap depicting the deviation from the median PFS (6.29 months) within the samples of patients that received gemcitabine nab-paclitaxel as their first-line therapy, attributed to the presence of distinct mutations and their combinations in the analyzed samples.
4. Discussion
Our study aimed to comprehensively investigate the molecular profiles and clinical characteristics of patients with PDAC. We placed particular emphasis on identifying potential predictive biomarkers and evaluating their significance in treatment selection and outcomes. This analysis consisted of 142 PDAC patients, with a relatively equal gender distribution and a median age of 66 years. In line with current treatment guidelines, the most frequently administered first-line chemotherapy regimens in our cohort were gemcitabine nab-paclitaxel and FOLFIRINOX [20]. Within our study, a small subset of patients exhibited TMB-H or MSI-H characteristics, a known characteristic that contributes to the limited efficacy of immunotherapy in pancreatic cancer [31,32,33,34].
In the current study, tumors with NOTCH-mutated pathways exhibited a longer OS compared to tumors without mutations in the NOTCH pathway. The role of NOTCH signaling in pancreatic cancer is complex and context-dependent. It appears to have both oncogenic and tumor-suppressive functions, with different NOTCH receptors playing distinct roles [35,36,37]. Individual NOTCH receptors can exhibit opposing activities and lead to diverse cellular responses [38]. Furthermore, NOTCH signaling is intricately intertwined with other signaling pathways and mechanisms, further complicating our understanding of its involvement in oncogenic processes [39,40]. The NOTCH signaling pathway has emerged as a crucial regulator in triggering epithelial-to-mesenchymal transition (EMT) [41], a transformative process that intensifies tumor aggressiveness by enhancing cellular motility and metastatic potential [42]. In addition, the NOTCH pathway has been linked to chemotherapy resistance in tumors [43,44], and therapeutic interventions targeting the NOTCH signaling pathway have shown potential for reversing chemoresistance [45]. We did not see a PFS difference between groups in our study. Interestingly, in our cohort, this improvement in OS with NOTCH-mutated pathways persisted even when combined with other mutated pathways such as TP53, which are typically associated with poor prognosis [46,47].
The consistent superior performance of NOTCH-mutated pathways raises intriguing questions. Some studies have proposed that highly mutated NOTCH samples exhibit significantly enhanced immunogenicity and immune response [48]. Additionally, other studies have suggested that the specific genes mutated within the NOTCH pathway may also play a role in determining prognosis [49]. The therapeutic targeting of the NOTCH pathway has been investigated in various trials using medications that globally inhibit the pathway [50,51]. However, other studies [52,53] suggest that the therapeutic targeting of the NOTCH pathway may be more complex than initially believed. The prognostic and predictive implications of NOTCH pathway mutations, as well as the potential targeting of this pathway for therapeutic benefit, will be a topic of substantial interest.
In our cohort, we identified a more favorable OS and PFS in samples exhibiting mutations within the KIT pathway. Furthermore, our findings indicate a synergistic effect as the combined presence of NOTCH and KIT mutations was associated with significantly prolonged OS compared to the analysis of these mutations individually. While the existing knowledge on pancreatic cancer is limited, certain studies have suggested that KIT may serve as a valuable prognostic marker for favorable outcomes in specific solid tumors [54]. However, the majority of available evidence suggests KIT as a marker for poorer prognosis [55,56]. Conversely, ALK has been proposed as a potential biomarker for favorable prognosis in lung cancer [57]. Although our results align with this finding, the understanding of ALK’s role in pancreatic cancer prognosis remains constrained due to limited data. Lastly, we observed that PI3K mutations were associated with improved PFS compared to samples lacking such mutations. Despite disappointing evidence regarding its therapeutic targeting [58], PI3K has been shown to interact with other pathways and influence survival outcomes [59].
Finally, this analysis highlights the potential role of chemotherapy selection, specifically noting that gemcitabine nab-paclitaxel may exhibit better performance in the presence of certain mutation pathways. Platinum-based therapy has been recognized for its effectiveness in tumors with homologous combination defects [60]. However, there is limited clinical data available for gemcitabine or taxanes in this context [61]. To predict in vivo tumor response to treatment, several studies have utilized PDX and organoid models [62], where the mutational profile serves as a readily available tool. This hypothesis-generating information suggests that mutational profile analysis should be considered for inclusion in future clinical trials.
This study has several limitations that should be taken into account when interpreting the findings. Firstly, the retrospective study design introduces inherent limitations. Data collected retrospectively from chart reviews may be influenced by biases and limitations in data availability, quality, and completeness. Secondly, this study’s single-center nature restricts the generalizability of the results. Moreover, this study’s sample size is relatively small, and the presence of missing data for OS or PFS analysis, as well as the subdivision of patients by treatment, further reduced the sample size. This limited sample size may affect the statistical power and precision of the findings. Finally, it is important to acknowledge that the genetic analysis was based on pathways that grouped several genetic mutations, thus limiting the scope of the analysis. This approach may have overlooked important genetic alterations that could specifically impact treatment response and survival. A more comprehensive and broader genetic profiling approach may provide additional insights into the study’s subject matter.
5. Conclusions
These findings contribute to our understanding of the clinical characteristics and treatment patterns of pancreatic cancer patients and highlight the potential for personalized treatment approaches based on molecular profiling. The integration of molecular profiling, such as next-generation sequencing, holds great promise for identifying actionable targets and guiding personalized therapeutic approaches in PDAC. However, further studies are needed to validate these findings and determine their clinical implications.
Author Contributions
Conceptualization, R.P.d.l.F. and M.L.B.P.; data curation, R.P.d.l.F.; formal analysis, R.P.d.l.F.; funding acquisition, M.L.B.P.; investigation, R.P.d.l.F., S.S. and C.P.; methodology, R.P.d.l.F. and M.L.B.P.; project administration, R.P.d.l.F. and M.L.B.P.; resources, M.L.B.P.; software, R.P.d.l.F.; supervision, M.L.B.P.; validation, A.A.A.R. and M.L.B.P.; visualization, R.P.d.l.F.; writing—original draft, R.P.d.l.F.; writing—review and editing, R.P.d.l.F., A.A.A.R. and M.L.B.P. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Cancer Institute, grant number K08CA248473.
Institutional Review Board Statement
This study was conducted in accordance with the Declaration of Helsinki, and approved by the Institutional Review Board (or Ethics Committee) of Beth Israel Deaconess Medical Center (protocol code 2006P000305 and date of approval 15 June 2011, with annual renewals most recently 2/21/24).
Informed Consent Statement
Patient consent was not required for this study as it involves a retrospective chart review with de-identified patient data, minimizing any risk of harm or breach of privacy. This consent waiver was approved by the Ethics Committee of Beth Israel Deaconess Medical Center.
Data Availability Statement
The data supporting the findings of this study are available from the corresponding author, Mary Linton B. Peters, upon reasonable request. The data are not publicly available due to privacy or ethical restrictions.
Conflicts of Interest
The authors declare no conflicts of interest related to this study. Mary Linton B. Peters has received institutional funding from NuCana, Bristol Myers Squibb, and Genentech outside the submitted work. No other relationships or activities that could appear to have influenced the submitted work exist.
Abbreviations
A list of the abbreviations used in this manuscript.
| PDAC | Pancreatic ductal adenocarcinoma |
| ECOG | Eastern Cooperative Oncology Group |
| NGS | Next-generation sequencing |
| RECIST | Response Evaluation Criteria in Solid Tumors |
| TMB | Tumor mutation burden |
| MSI-H | Microsatellite instability-high |
| OS | Overall survival |
| PFS | Progression-free survival |
| CC | Cell cycle |
| DDR | DNA damage repair |
| PI3K | Phosphoinositide 3-kinase |
| KIT | KIT proto-oncogene |
| NOTCH | Neurogenic locus notch homolog protein |
| ALK | Anaplastic lymphoma kinase |
| ERBB2 | Erb-B2 receptor tyrosine kinase 2 |
| TP53 | Tumor protein p53 |
| TGFB | Transforming growth factor beta |
| PDGFR | Platelet-derived growth factor receptor |
| RAS | Rat sarcoma |
| FTL3 | Fms-like tyrosine kinase 3 |
| FGFR | Fibroblast growth factor receptor |
| WNT | Wnt signaling pathway |
Appendix A
The appendix includes a table specifying the mutations included in each pathway.

Table A1.
Classification of gene mutations.
Table A1.
Classification of gene mutations.
| Pathway | Gene Mutations Included in the Pathway [63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96] | |||||
|---|---|---|---|---|---|---|
| Cell Cycle mutation | ||||||
| CCND1 | MYC | BCOR | BRD4 | RAD54L | AURKA | |
| CCND2 | STK11 | CIC | BTG2 | PRKCI | AURKB | |
| CCND3 | TERT | DAXX | BCL2L2 | TOP2A | XPO1 | |
| CCNE1 | BCL2L1 | ERG | BCL6 | TERC | PTEN | |
| CDK6 | PPP2R2A | ESR1 | JUN | TET2 | ALK | |
| CDKN2A | PPP2R1A | EWSR1 | IDH1 | HNF1A | ||
| CDKN2B | CASP8 | EZH2 | IDH2 | AR | ||
| RB1 | FAS | CDH1 | HDAC1 | PARP1 | ||
| FAT1 | APC | CDKN2C | GNA13 | ATR | ||
| ATRX | MDM2 | CDKN1B | MPL | RANBP2 | ||
| DNA Damage repair mutations | ||||||
| BLM | XRCC2 | WHSC1 | JUN | POLD1 | KRAS | |
| BRCA2 | FAT3 | CUL4A | H3F3A | RAD51L3 | ALK | |
| MLH1 | BRIP1 | FANCL | GEN1 | TP53BP1 | ABL1 | |
| MSH3 | FANCA | FANCM | FUBP1 | CHEK2 | ||
| PARP2 | FANCC | ERCC4 | EMSY | MSH2 | ||
| PMS2 | FANCD2 | CHEK1 | MPL | AR | ||
| POLE | FANCG | CDH1 | MRE11A | PARP1 | ||
| PRKDC | PALB2 | BRCA1 | MSH6 | ATR | ||
| RAD51C | ATM | BARD1 | MUTYH | SF3B1 | ||
| RAD52 | APC | BCL2L2 | NBN | FH | ||
| NOTCH signaling pathway | ||||||
| CREBBP | GATA6 | |||||
| EP300 | EGFR | |||||
| NCOR1 | CBFB | |||||
| NOTCH1 | FBXW7 | |||||
| NOTCH2 | CDK8 | |||||
| NOTCH3 | IKZF1 | |||||
| NOTCH4 | FLT4 | |||||
| SPEN | FH | |||||
| JAK2 | MDM2 | |||||
| RBM10 | BCOR | |||||
| PI3K/AKT signaling pathway | ||||||
| AKT2 | NTRK2 | IGF1R | ||||
| INPP4B | PDK1 | EZH2 | ||||
| MTOR | PHLPP2 | FGF4 | ||||
| PIK3CA | PIK3C2B | BCL2L2 | ||||
| PIK3R1 | PIK3C2G | RPTOR | ||||
| PPP2R1A | PIK3CB | AKT3 | ||||
| PTEN | PPARG | ERBB2 | ||||
| STK11 | PREX2 | ERBB3 | ||||
| TSC2 | RICTOR | ERBB4 | ||||
| MET | TYRO3 | |||||
| KIT signaling pathway | ||||||
| KIT | ||||||
| ROS1 | ||||||
| FGFR3 | ||||||
| MTOR | ||||||
| SRC | ||||||
| LYN | ||||||
| CBL | ||||||
| FTL3 | ||||||
| RAS/RAF/MAPK signaling pathway | ||||||
| BRAF | MAP2K1 | JUN | FGFR3 | |||
| KRAS | MAP2K4 | QKI | ||||
| MAP2K2 | MAP3K1 | RAF1 | ||||
| MET | MAP3K13 | PTPN11 | ||||
| NF1 | MAPK1 | ARAF | ||||
| NTRK1 | PPP2R1A | CBL | ||||
| NTRK2 | IGF1R | RET | ||||
| NTRK3 | DDR2 | ERBB2 | ||||
| XPO1 | CRKL | ERBB3 | ||||
| FGF4 | HRAS | ERBB4 | ||||
| TGF-beta Receptor | ||||||
| ACVR1B | AR | |||||
| SMAD2 | PARP1 | |||||
| SMAD3 | XPO1 | |||||
| SMAD4 | GATA6 | |||||
| TGFRB2 | ||||||
| APC | ||||||
| MYC | ||||||
| CDK8 | ||||||
| JUN | ||||||
| MEN1 | ||||||
| TP53 Activity Alteration | ||||||
| TP53 | BCL6 | AURKA | ||||
| ATM | MDM4 | AURKB | ||||
| CDK12 | PRDM1 | |||||
| MDM2 | PRKCI | |||||
| BCOR | RAD51L3 | |||||
| EP300 | SGK1 | |||||
| CIC | TAF1 | |||||
| DAXX | RPTOR | |||||
| CHEK12 | CHEK2 | |||||
| BCL2L2 | ATR | |||||
| WNT signaling pathway | ||||||
| APC | TRRAP | |||||
| AXIN1 | WT1 | |||||
| CHD4 | FAM123B | |||||
| CTNNB1 | XPO1 | |||||
| RNF43 | BCORL1 | |||||
| MYC | MED12 | |||||
| PPP2R1A | ||||||
| CDC73 | ||||||
| GSK3B | ||||||
| TNKS | ||||||
| ERBB2 signaling pathway | ||||||
| NF2 | ||||||
| EGFR | ||||||
| ERBB2 | ||||||
| ERBB3 | ||||||
| ERBB4 | ||||||
| CBL | ||||||
| BRAF | ||||||
| PDGFR signaling pathway | ||||||
| PDGFRA | ||||||
| PDGFRB | ||||||
| KDR | ||||||
| EGFR | ||||||
| IDH1 | ||||||
| LRP1B | ||||||
| AR | ||||||
| FH | ||||||
| KIT | ||||||
| NF1 | ||||||
| FGFR signaling pathway | ||||||
| FGFR1 | FGF6 | |||||
| FGFR3 | BCL2L2 | |||||
| MTOR | SOX2 | |||||
| PPP2R1A | FGFR2 | |||||
| PTEN | CBL | |||||
| ERBB3 | FH | |||||
| FGF19 | ||||||
| FGF23 | ||||||
| FGF3 | ||||||
| FGF4 | ||||||
| FTL3 signaling pathway | ||||||
| FLT3 | ||||||
| CBL | ||||||
| PIM1 | ||||||
| RAF1 | ||||||
| SRC | ||||||
| SYK | ||||||
| FH | ||||||
| PDGFRB | ||||||
| ALK signaling pathway | ||||||
| ALK | ||||||
| STAT3 | ||||||
| LTK | ||||||
| EGFR | ||||||
| ROS1 | ||||||
References
- Siegel, R.L.; Giaquinto, A.N.; Jemal, A. Cancer statistics, 2024. CA Cancer J. Clin. 2024, 74, 12–49, Erratum in CA Cancer J. Clin. 2024, 74, 203. [Google Scholar] [CrossRef] [PubMed]
- Rahib, L.; Smith, B.D.; Aizenberg, R.; Rosenzweig, A.B.; Fleshman, J.M.; Matrisian, L.M. Projecting Cancer Incidence and Deaths to 2030: The Unexpected Burden of Thyroid, Liver, and Pancreas Cancers in the United States. Cancer Res. 2014, 74, 2913–2921. [Google Scholar] [CrossRef] [PubMed]
- Cronin, K.A.; Scott, S.; Firth, A.U.; Sung, H.; Henley, S.J.; Sherman, R.L.; Siegel, R.L.; Anderson, R.N.; Kohler, B.A.; Benard, V.B.; et al. Annual report to the nation on the status of cancer, part 1: National cancer statistics. Cancer 2022, 128, 4251–4284. [Google Scholar] [CrossRef] [PubMed]
- Maloney, S.; Itchins, M.; Arena, J.; Sahni, S.; Howell, V.M.; Hayes, S.A.; Gill, A.J.; Clarke, S.J.; Samra, J.; Mittal, A.; et al. Optimal Upfront Treatment in Surgically Resectable Pancreatic Cancer Candidates: A High-Volume Center Retrospective Analysis. J. Clin. Med. 2021, 10, 2700. [Google Scholar] [CrossRef]
- Chhoda, A.; Vodusek, Z.; Wattamwar, K.; Mukherjee, E.; Gunderson, C.; Grimshaw, A.; Sharma, A.; Ahuja, N.; Kastrinos, F.; Farrell, J.J. Late-stage pancreatic cancer detected during high-risk individual surveillance: A systematic review and meta-analysis. Gastroenterology 2022, 162, 786–798. [Google Scholar] [CrossRef] [PubMed]
- Sarantis, P.; Koustas, E.; Papadimitropoulou, A.; Papavassiliou, A.G.; Karamouzis, M.V. Pancreatic ductal adenocarcinoma: Treatment hurdles, tumor microenvironment and immunotherapy. World J. Gastrointest. Oncol. 2020, 12, 173–181. [Google Scholar] [CrossRef]
- Principe, D.R.; Underwood, P.W.; Korc, M.; Trevino, J.G.; Munshi, H.G.; Rana, A. The Current Treatment Paradigm for Pancreatic Ductal Adenocarcinoma and Barriers to Therapeutic Efficacy. Front. Oncol. 2021, 11, 688377. [Google Scholar] [CrossRef]
- De Dosso, S.; Siebenhüner, A.R.; Winder, T.; Meisel, A.; Fritsch, R.; Astaras, C.; Szturz, P.; Borner, M. Treatment landscape of metastatic pancreatic cancer. Cancer Treat. Rev. 2021, 96, 102180. [Google Scholar] [CrossRef] [PubMed]
- Damanakis, A.I.; Gebauer, F.; Bruns, C.J. Molekulare Prädiktoren für den Krankheitsverlauf und die individualisierte Therapie beim Pankreaskarzinom. Chirurg 2020, 91, 642–649. [Google Scholar] [CrossRef]
- Bailey, P.; Chang, D.K.; Nones, K.; Johns, A.L.; Patch, A.M.; Gingras, M.C.; Miller, D.K.; Christ, A.N.; Bruxner, T.J.; Quinn, M.C.; et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016, 531, 47–52. [Google Scholar] [CrossRef]
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495–501. [Google Scholar] [CrossRef]
- Connor, A.A.; Denroche, R.E.; Jang, G.H.; Timms, L.; Kalimuthu, S.N.; Selander, I.; McPherson, T.; Wilson, G.W.; Chan-Seng-Yue, M.A.; Borozan, I.; et al. Association of Distinct Mutational Signatures with Correlates of Increased Immune Activity in Pancreatic Ductal Adenocarcinoma. JAMA Oncol. 2017, 3, 774–783. [Google Scholar] [CrossRef]
- Moffitt, R.A.; Marayati, R.; Flate, E.L.; Volmar, K.E.; Loeza, S.G.; Hoadley, K.A.; Rashid, N.U.; Williams, L.A.; Eaton, S.C.; Chung, A.H.; et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 2015, 47, 1168–1178. [Google Scholar] [CrossRef]
- Collisson, E.A.; Sadanandam, A.; Olson, P.; Gibb, W.J.; Truitt, M.; Gu, S.; Cooc, J.; Weinkle, J.; Kim, G.E.; Jakkula, L.; et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat. Med. 2011, 17, 500–503. [Google Scholar] [CrossRef]
- Pishvaian, M.J.; Blais, E.M.; Brody, J.R.; Lyons, E.; DeArbeloa, P.; Hendifar, A.; Mikhail, S.; Chung, V.; Sahai, V.; Sohal, D.P.S.; et al. Overall survival in patients with pancreatic cancer receiving matched therapies following molecular profiling: A retrospective analysis of the Know Your Tumor registry trial. Lancet Oncol. 2020, 21, 508–518. [Google Scholar] [CrossRef]
- Shen, G.Q.; Aleassa, E.M.; Walsh, R.M.; Morris-Stiff, G. Next-Generation Sequencing in Pancreatic Cancer. Pancreas 2019, 48, 739–748. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Qiu, X.; Lu, C.; Zhu, Y.; Kong, W.; Xu, M.; Jiang, W.; Wang, Y.; Li, Y.; Zhang, W.; et al. Molecular Landscape and Prognostic Biomarker Analysis of Advanced Pancreatic Cancer and Predictors of Treatment Efficacy of AG Chemotherapy. Front. Oncol. 2022, 12, 844527. [Google Scholar] [CrossRef]
- Shaya, J.; Kato, S.; Adashek, J.J.; Patel, H.; Fanta, P.T.; Botta, G.P.; Sicklick, J.K.; Kurzrock, R. Personalized matched targeted therapy in advanced pancreatic cancer: A pilot cohort analysis. NPJ Genom. Med. 2023, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Bilimoria, K.Y.; Bentrem, D.J.; Ko, C.Y.; Ritchey, J.; Stewart, A.K.; Winchester, D.P.; Talamonti, M.S.; Jennings, G.; Bhayani, N.H.; Stamos, M.J. Validation of the 6th Edition AJCC Pancreatic Cancer Staging System: Report from the National Cancer Database. Cancer 2007, 110, 738–744. [Google Scholar] [CrossRef]
- Referenced with Permission from the NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Pancreatic Adenocarcinoma V.1.2022. National Comprehensive Cancer Network, Inc. 2022. Available online: www.NCCN.org (accessed on 6 June 2023).
- Conroy, T.; Desseigne, F.; Ychou, M.; Bouché, O.; Guimbaud, R.; Bécouarn, Y.; Adenis, A.; Raoul, J.L.; Gourgou-Bourgade, S.; de la Fouchardière, C.; et al. FOLFIRINOX versus Gemcitabine for Metastatic Pancreatic Cancer. N. Engl. J. Med. 2011, 364, 1817–1825. [Google Scholar] [CrossRef]
- Von Hoff, D.D.; Ervin, T.; Arena, F.P.; Chiorean, E.G.; Infante, J.; Moore, M.; Seay, T.; Tjulandin, S.A.; Ma, W.W.; Saleh, M.N.; et al. Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N. Engl. J. Med. 2013, 369, 1691–1703. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Eisenhauer, E.A.; Therasse, P.; Bogaerts, J.; Schwartz, L.H.; Sargent, D.; Ford, R.; Dancey, J.; Arbuck, S.; Gwyther, S.; Mooney, M.; et al. New response evaluation criteria in solid tumours: Revised RECIST guideline (version 1.1). Eur. J. Cancer 2009, 45, 228–247. [Google Scholar] [CrossRef] [PubMed]
- Milbury, C.A.; Creeden, J.; Yip, W.-K.; Smith, D.L.; Pattani, V.; Maxwell, K.; Sawchyn, B.; Gjoerup, O.; Meng, W.; Skoletsky, J.; et al. Clinical and analytical validation of FoundationOne® CDx, a comprehensive genomic profiling assay for solid tumors. PLoS ONE 2022, 17, e0264138. [Google Scholar] [CrossRef]
- Gillespie, M.; Jassal, B.; Stephan, R.; Milacic, M.; Rothfels, K.; Senff-Ribeiro, A.; Griss, J.; Sevilla, C.; Matthews, L.; Gong, C.; et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 2022, 50, D687–D692. [Google Scholar] [CrossRef]
- Pös, O.; Radvanszky, J.; Buglyó, G.; Pös, Z.; Rusnakova, D.; Nagy, B.; Szemes, T.; Eisenhauer, E.A.; Therasse, P.; Bogaerts, J.; et al. DNA copy number variation: Main characteristics, evolutionary significance, and pathological aspects. Biomed. J. 2021, 44, 548–559. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Harris, P.A.; Taylor, R.; Thielke, R.; Payne, J.; Gonzalez, N.; Conde, J.G. Research electronic data capture (REDCap)—A metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 2009, 42, 377–381. [Google Scholar] [CrossRef]
- Harris, P.A.; Taylor, R.; Minor, B.L.; Elliott, V.; Fernandez, M.; O’Neal, L.; McLeod, L.; Delacqua, G.; Delacqua, F.; Kirby, J.; et al. The REDCap consortium: Building an international community of software platform partners. J. Biomed. Inform. 2019, 95, 103208. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.R-project.org/ (accessed on 6 June 2023).
- Therneau, T. A Package for Survival Analysis in R. R Package Version 3.4-0. Available online: https://CRAN.R-project.org/package=survival (accessed on 6 June 2023).
- Marabelle, A.; Fakih, M.; Lopez, J.; Shah, M.; Shapira-Frommer, R.; Nakagawa, K.; Chung, H.C.; Kindler, H.L.; Lopez-Martin, J.A.; Miller, W.H., Jr.; et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: Prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 2020, 21, 1353–1365. [Google Scholar] [CrossRef]
- Ghidini, M.; Lampis, A.; Mirchev, M.B.; Okuducu, A.F.; Ratti, M.; Valeri, N.; Hahne, J.C. Immune-Based Therapies and the Role of Microsatellite Instability in Pancreatic Cancer. Genes 2020, 12, 33. [Google Scholar] [CrossRef]
- Patnaik, A.; Kang, S.P.; Rasco, D.; Papadopoulos, K.P.; Elassaiss-Schaap, J.; Beeram, M.; Drengler, R.; Chen, C.; Smith, L.; Espino, G.; et al. Phase I Study of Pembrolizumab (MK-3475; Anti-PD-1 Monoclonal Antibody) in Patients with Advanced Solid Tumors. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2015, 21, 4286–4293. [Google Scholar] [CrossRef]
- Rosenberg, A.; Mahalingam, D. Immunotherapy in pancreatic adenocarcinoma-overcoming barriers to response. J. Gastrointest. Oncol. 2018, 9, 143–159. [Google Scholar] [CrossRef] [PubMed]
- Demehri, S.; Turkoz, A.; Kopan, R. Epidermal Notch1 loss promotes skin tumorigenesis by impacting the stromal microenvironment. Cancer Cell 2009, 16, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Mullendore, M.E.; Koorstra, J.B.; Li, Y.M.; Offerhaus, G.J.; Fan, X.; Henderson, C.M.; Matsui, W.; Eberhart, C.G.; Maitra, A.; Feldmann, G.; et al. Ligand-dependent Notch signaling is involved in tumor initiation and tumor maintenance in pancreatic cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 2009, 15, 2291–2301. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.J.; Sanborn, Z.; Arnett, K.L.; Bayston, L.J.; Liao, W.; Proby, C.M.; Leigh, I.M.; Collisson, E.A.; Gordon, P.B.; Jakkula, L.; et al. Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proc. Natl. Acad. Sci. USA 2011, 108, 17761–17766. [Google Scholar] [CrossRef] [PubMed]
- Graziani, I.; Eliasz, S.; De Marco, M.A.; Chen, Y.; Pass, H.I.; De May, R.M.; Strack, P.R.; Miele, L.; Bocchetta, M. Opposite effects of Notch-1 and Notch-2 on mesothelioma cell survival under hypoxia are exerted through the Akt pathway. Cancer Res. 2008, 68, 9678–9685. [Google Scholar] [CrossRef]
- De La, O.J.P.; Emerson, L.L.; Goodman, J.L.; Froebe, S.C.; Illum, B.E.; Curtis, A.B.; Murtaugh, L.C. Notch and Kras reprogram pancreatic acinar cells to ductal intraepithelial neoplasia. Proc. Natl. Acad. Sci. USA 2008, 105, 18907–18912. [Google Scholar] [CrossRef]
- Misiorek, J.O.; Przybyszewska-Podstawka, A.; Kałafut, J.; Paziewska, B.; Rolle, K.; Rivero-Müller, A.; Nees, M. Context Matters: NOTCH Signatures and Pathway in Cancer Progression and Metastasis. Cells 2021, 10, 94. [Google Scholar] [CrossRef]
- Zavadil, J.; Cermak, L.; Soto-Nieves, N.; Böttinger, E.P. Integration of TGF-beta/Smad and Jagged1/Notch signalling in epithelial-to-mesenchymal transition. EMBO J. 2004, 23, 1155–1165. [Google Scholar] [CrossRef]
- Wang, Z.; Li, Y.; Kong, D.; Sarkar, F.H. The role of Notch signaling pathway in epithelial-mesenchymal transition (EMT) during development and tumor aggressiveness. Curr. Drug Targets 2010, 11, 745–751. [Google Scholar] [CrossRef]
- Lombardo, Y.; Faronato, M.; Filipovic, A.; Vircillo, V.; Magnani, L.; Coombes, R.C. Nicastrin and Notch4 drive endocrine therapy resistance and epithelial to mesenchymal transition in MCF7 breast cancer cells. Breast Cancer Res. 2014, 16, R62. [Google Scholar] [CrossRef]
- Jia, Y.; Xie, J. Promising molecular mechanisms responsible for gemcitabine resistance in cancer. Genes Dis. 2015, 2, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.; Li, J.; Sun, H.; Liu, S.; Cui, Y.; Li, F. HES 1 is essential for chemoresistance induced by stellate cells and is associated with poor prognosis in pancreatic cancer. Oncol. Rep. 2015, 33, 1883–1889. [Google Scholar] [CrossRef]
- Chanrion, M.; Kuperstein, I.; Barrière, C.; El Marjou, F.; Cohen, D.; Vignjevic, D.; Stimmer, L.; Paul-Gilloteaux, P.; Bièche, I.; Tavares Sdos, R.; et al. Concomitant Notch activation and p53 deletion trigger epithelial-to-mesenchymal transition and metastasis in mouse gut. Nat. Commun. 2014, 5, 5005. [Google Scholar] [CrossRef] [PubMed]
- Dotto, G.P. Crosstalk of Notch with p53 and p63 in cancer growth control. Nat. Rev. Cancer 2009, 9, 587–595. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, Y.; Li, X.; Feng, G.; Hu, S.; Bai, Y. The Impact of NOTCH Pathway Alteration on Tumor Microenvironment and Clinical Survival of Immune Checkpoint Inhibitors in NSCLC. Front. Immunol. 2021, 12, 638763. [Google Scholar] [CrossRef] [PubMed]
- Takam Kamga, P.; Collo, G.D.; Resci, F.; Bazzoni, R.; Mercuri, A.; Quaglia, F.M.; Tanasi, I.; Delfino, P.; Visco, C.; Bonifacio, M.; et al. Notch Signaling Molecules as Prognostic Biomarkers for Acute Myeloid Leukemia. Cancers 2019, 11, 1958. [Google Scholar] [CrossRef] [PubMed]
- Maraver, A.; Fernández-Marcos, P.J.; Herranz, D.; Muñoz-Martin, M.; Gomez-Lopez, G.; Cañamero, M.; Mulero, F.; Megías, D.; Sanchez-Carbayo, M.; Shen, J.; et al. Therapeutic effect of γ-secretase inhibition in KrasG12V-driven non-small cell lung carcinoma by derepression of DUSP1 and inhibition of ERK. Cancer Cell 2012, 22, 222–234. [Google Scholar] [CrossRef]
- Massard, C.; Azaro, A.; Soria, J.C.; Lassen, U.; Le Tourneau, C.; Sarker, D.; Smith, C.; Ohnmacht, U.; Oakley, G.; Patel, B.K.R.; et al. First-in-human study of LY3039478, an oral Notch signaling inhibitor in advanced or metastatic cancer. Ann. Oncol. 2018, 29, 1911–1917. [Google Scholar] [CrossRef] [PubMed]
- Aste-Amézaga, M.; Zhang, N.; Lineberger, J.E.; Arnold, B.A.; Toner, T.J.; Gu, M.; Huang, L.; Vitelli, S.; Vo, K.T.; Haytko, P.; et al. Characterization of Notch1 antibodies that inhibit signaling of both normal and mutated Notch1 receptors. PLoS ONE 2010, 5, e9094. [Google Scholar] [CrossRef]
- Hu, Z.I.; Bendell, J.C.; Bullock, A.; LoConte, N.K.; Hatoum, H.; Ritch, P.; Hool, H.; Leach, J.W.; Sanchez, J.; Sohal, D.P.S.; et al. A randomized phase II trial of nab-paclitaxel and gemcitabine with tarextumab or placebo in patients with untreated metastatic pancreatic cancer. Cancer Med. 2019, 8, 5148–5157. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dias-Santagata, D.; Selim, M.A.; Su, Y.; Peng, Y.; Vollmer, R.; Chłopik, A.; Tell-Marti, G.; Paral, K.M.; Shalin, S.C.; Shea, C.R.; et al. KIT mutations and CD117 overexpression are markers of better progression-free survival in vulvar melanomas. Br. J. Dermatol. 2017, 177, 1376–1384. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, S.; Chi, S.; Minami, Y.; Fukushima, K.; Shibayama, H.; Hosono, N.; Yamauchi, T.; Morishita, T.; Kondo, T.; Yanada, M.; et al. Mutated KIT Tyrosine Kinase as a Novel Molecular Target in Acute Myeloid Leukemia. Int. J. Mol. Sci. 2022, 23, 4694. [Google Scholar] [CrossRef] [PubMed]
- Han, G.; Jiang, Z.; Zhang, J.; Li, Z.; Liu, Y.; Wang, D. A meta-analysis of prognostic value of KIT mutation status in gastrointestinal stromal tumors. OncoTargets Ther. 2016, 9, 3387–3398. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Qian, Y.; Wang, Y.; Fang, F.; Wu, G. Prognostic value of Beclin 1, EGFR and ALK in non-squamous non-small cell lung cancer. Discov. Oncol. 2022, 13, 127. [Google Scholar] [CrossRef] [PubMed]
- Junttila, M.R.; Devasthali, V.; Cheng, J.H.; Castillo, J.; Metcalfe, C.; Clermont, A.C.; Otter, D.D.; Chan, E.; Bou-Reslan, H.; Cao, T.; et al. Modeling targeted inhibition of MEK and PI3 kinase in human pancreatic cancer. Mol. Cancer Ther. 2015, 14, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Diehl, A.C.; Hannan, L.M.; Zhen, D.B.; Coveler, A.L.; King, G.; Cohen, S.A.; Harris, W.P.; Shankaran, V.; Wong, K.M.; Green, S.; et al. KRAS Mutation Variants and Co-occurring PI3K Pathway Alterations Impact Survival for Patients with Pancreatic Ductal Adenocarcinomas. Oncologist 2022, 27, 1025–1033. [Google Scholar] [CrossRef] [PubMed]
- Konstantinopoulos, P.A.; Spentzos, D.; Karlan, B.Y.; Taniguchi, T.; Fountzilas, E.; Francoeur, N.; Levine, D.A.; Cannistra, S.A. Gene expression profile of BRCAness that correlates with responsiveness to chemotherapy and with outcome in patients with epithelial ovarian cancer. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2010, 28, 3555–3561. [Google Scholar] [CrossRef] [PubMed]
- Liao, G.; Jiang, Z.; Yang, Y.; Zhang, C.; Jiang, M.; Zhu, J.; Xu, L.; Xie, A.; Yan, M.; Zhang, Y.; et al. Combined homologous recombination repair deficiency and immune activation analysis for predicting intensified responses of anthracycline, cyclophosphamide and taxane chemotherapy in triple-negative breast cancer. BMC Med. 2021, 19, 190. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Bockorny, B.; Paul, I.; Akshinthala, D.; Frappart, P.O.; Gandarilla, O.; Bose, A.; Sanchez-Gonzalez, V.; Rouse, E.E.; Lehoux, S.D.; et al. PDX-derived organoids model in vivo drug response and secrete biomarkers. JCI Insight 2020, 5, e135544. [Google Scholar] [CrossRef]
- Fabregat, A.; Sidiropoulos, K.; Viteri, G.; Marin-Garcia, P.; Ping, P.; Stein, L.; D’Eustachio, P.; Hermjakob, H. Reactome diagram viewer: Data structures and strategies to boost performance. Bioinformatics 2018, 34, 1208–1214. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Stacey, D.W. Cyclin D1 serves as a cell cycle regulatory switch in actively proliferating cells. Curr. Opin. Cell Biol. 2003, 15, 158–163. [Google Scholar] [CrossRef] [PubMed]
- Tadesse, S.; Yu, M.; Kumarasiri, M.; Le, B.T.; Wang, S. Targeting CDK6 in cancer: State of the art and new insights. Cell Cycle 2015, 14, 3220–3230. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Agarwal, P.; Sandey, M.; DeInnocentes, P.; Bird, R.C. Tumor suppressor gene p16/INK4A/CDKN2A-dependent regulation into and out of the cell cycle in a spontaneous canine model of breast cancer. J. Cell Biochem. 2013, 114, 1355–1363. [Google Scholar] [CrossRef] [PubMed]
- Su, B.; Lim, D.; Qi, C.; Zhang, Z.; Wang, J.; Zhang, F.; Dong, C.; Feng, Z. VPA mediates bidirectional regulation of cell cycle progression through the PPP2R2A-Chk1 signaling axis in response to HU. Cell Death Dis. 2023, 14, 114. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- de Miranda, N.F.; Peng, R.; Georgiou, K.; Wu, C.; Falk Sörqvist, E.; Berglund, M.; Chen, L.; Gao, Z.; Lagerstedt, K.; Lisboa, S.; et al. DNA repair genes are selectively mutated in diffuse large B cell lymphomas. J. Exp. Med. 2013, 210, 1729–1742. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ronen, A.; Glickman, B.W. Human DNA repair genes. Environ. Mol. Mutagen. 2001, 37, 241–283. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, C.; Bernstein, H.; Payne, C.M.; Garewal, H. DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: Fail-safe protection against carcinogenesis. Mutat. Res. 2002, 511, 145–178. [Google Scholar] [CrossRef] [PubMed]
- Toh, M.; Ngeow, J. Homologous recombination deficiency: Cancer predispositions and treatment implications. Oncologist 2021, 26, e1526–e1537. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Mekonnen, N.; Yang, H.; Shin, Y.K. Homologous recombination deficiency in ovarian, breast, colorectal, pancreatic, non-small cell lung and prostate cancers, and the mechanisms of resistance to PARP inhibitors. Front. Oncol. 2022, 12, 880643. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ohashi, S.; Natsuizaka, M.; Yashiro-Ohtani, Y.; Kalman, R.A.; Nakagawa, M.; Wu, L.; Klein-Szanto, A.J.; Herlyn, M.; Diehl, J.A.; Katz, J.P.; et al. NOTCH1 and NOTCH3 coordinate esophageal squamous differentiation through a CSL-dependent transcriptional network. Gastroenterology 2010, 139, 2113–2123. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ho, D.M.; Guruharsha, K.G.; Artavanis-Tsakonas, S. The Notch Interactome: Complexity in Signaling Circuitry. Adv. Exp. Med. Biol. 2018, 1066, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Knobbe, C.B.; Reifenberger, G. Genetic alterations and aberrant expression of genes related to the phosphatidyl-inositol-3′-kinase/protein kinase B (Akt) signal transduction pathway in glioblastomas. Brain Pathol. 2003, 13, 507–518. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bonin, S.; Pracella, D.; Barbazza, R.; Dotti, I.; Boffo, S.; Stanta, G. PI3K/AKT Signaling in Breast Cancer Molecular Subtyping and Lymph Node Involvement. Dis. Markers 2019, 2019, 7832376. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Phung, B.; Sun, J.; Schepsky, A.; Steingrimsson, E.; Rönnstrand, L. C-KIT signaling depends on microphthalmia-associated transcription factor for effects on cell proliferation. PLoS ONE 2011, 6, e24064. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bosbach, B.; Rossi, F.; Yozgat, Y.; Loo, J.; Zhang, J.Q.; Berrozpe, G.; Warpinski, K.; Ehlers, I.; Veach, D.; Kwok, A.; et al. Direct engagement of the PI3K pathway by mutant KIT dominates oncogenic signaling in gastrointestinal stromal tumor. Proc. Natl. Acad. Sci. USA 2017, 114, E8448–E8457. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yaeger, R.; Corcoran, R.B. Targeting Alterations in the RAF-MEK Pathway. Cancer Discov. 2019, 9, 329–341. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Rocca, A.; Braga, L.; Volpe, M.C.; Maiocchi, S.; Generali, D. The Predictive and Prognostic Role of RAS-RAF-MEK-ERK Pathway Alterations in Breast Cancer: Revision of the Literature and Comparison with the Analysis of Cancer Genomic Datasets. Cancers 2022, 14, 5306. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Nakao, A.; Imamura, T.; Souchelnytskyi, S.; Kawabata, M.; Ishisaki, A.; Oeda, E.; Tamaki, K.; Hanai, J.; Heldin, C.H.; Miyazono, K.; et al. TGF-beta receptor-mediated signalling through Smad2, Smad3 and Smad4. EMBO J. 1997, 16, 5353–5362. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gomes, Á.N.M.; Oliveira, K.K.; Marchi, F.A.; Bettim, B.B.; Germano, J.N.; Gonçalves Filho, J.; Pinto, C.A.L.; Lourenço, S.V.; Coutinho-Camillo, C.M. TGFβ signaling pathway in salivary gland tumors. Arch. Oral Biol. 2024, 162, 105943. [Google Scholar] [CrossRef] [PubMed]
- Basu, S.; Murphy, M.E. Genetic Modifiers of the p53 Pathway. Cold Spring Harb. Perspect. Med. 2016, 6, a026302. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yoon, H.; Liyanarachchi, S.; Wright, F.A.; Davuluri, R.; Lockman, J.C.; de la Chapelle, A.; Pellegata, N.S. Gene expression profiling of isogenic cells with different TP53 gene dosage reveals numerous genes that are affected by TP53 dosage and identifies CSPG2 as a direct target of p53. Proc. Natl. Acad. Sci. USA 2002, 99, 15632–15637. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liu, Y.; Zhang, X.; Han, C.; Wan, G.; Huang, X.; Ivan, C.; Jiang, D.; Rodriguez-Aguayo, C.; Lopez-Berestein, G.; Rao, P.H.; et al. TP53 loss creates therapeutic vulnerability in colorectal cancer. Nature 2015, 520, 697–701, Erratum in Nature 2021, 597, E6. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Aghabozorgi, A.S.; Ebrahimi, R.; Bahiraee, A.; Tehrani, S.S.; Nabizadeh, F.; Setayesh, L.; Jafarzadeh-Esfehani, R.; Ferns, G.A.; Avan, A.; Rashidi, Z. The genetic factors associated with Wnt signaling pathway in colorectal cancer. Life Sci. 2020, 256, 118006. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.A.; Michalski, M.N.; Stevens, P.D.; Sall, E.A.; Williams, B.O. Regulation of Wnt receptor activity: Implications for therapeutic development in colon cancer. J. Biol. Chem. 2021, 296, 100782. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kikuchi, A.; Kishida, S.; Yamamoto, H. Regulation of Wnt signaling by protein-protein interaction and post-translational modifications. Exp. Mol. Med. 2006, 38, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Wolff, A.C.; Hammond, M.E.; Schwartz, J.N.; Hagerty, K.L.; Allred, D.C.; Cote, R.J.; Dowsett, M.; Fitzgibbons, P.L.; Hanna, W.M.; Langer, A.; et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J. Clin. Oncol. 2007, 25, 118–145. [Google Scholar] [CrossRef] [PubMed]
- Black, L.E.; Longo, J.F.; Carroll, S.L. Mechanisms of Receptor Tyrosine-Protein Kinase ErbB-3 (ERBB3) Action in Human Neoplasia. Am. J. Pathol. 2019, 189, 1898–1912. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Funa, K.; Sasahara, M. The roles of PDGF in development and during neurogenesis in the normal and diseased nervous system. J. Neuroimmune Pharmacol. 2014, 9, 168–181. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Rogers, M.A.; Campaña, M.B.; Long, R.; Fantauzzo, K.A. PDGFR dimer-specific activation, trafficking and downstream signaling dynamics. J. Cell Sci. 2022, 135, jcs259686. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ho, B.B.; Bergwitz, C. FGF23 signalling and physiology. J. Mol. Endocrinol. 2021, 66, R23–R32. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wang, K.; Ji, W.; Yu, Y.; Li, Z.; Niu, X.; Xia, W.; Lu, S. FGFR1-ERK1/2-SOX2 axis promotes cell proliferation, epithelial-mesenchymal transition, and metastasis in FGFR1-amplified lung cancer. Oncogene 2018, 37, 5340–5354, Erratum in Oncogene 2020, 39, 6619–6620. [Google Scholar] [CrossRef] [PubMed]
- Robinson, L.J.; Xue, J.; Corey, S.J. Src family tyrosine kinases are activated by Flt3 and are involved in the proliferative effects of leukemia-associated Flt3 mutations. Exp. Hematol. 2005, 33, 469–479. [Google Scholar] [CrossRef] [PubMed]
- Reindl, C.; Quentmeier, H.; Petropoulos, K.; Greif, P.A.; Benthaus, T.; Argiropoulos, B.; Mellert, G.; Vempati, S.; Duyster, J.; Buske, C.; et al. CBL exon 8/9 mutants activate the FLT3 pathway and cluster in core binding factor/11q deletion acute myeloid leukemia/myelodysplastic syndrome subtypes. Clin. Cancer Res. 2009, 15, 2238–2247. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).