A Machine Learning Framework to Predict the Tensile Stress of Natural Rubber: Based on Molecular Dynamics Simulation Data
Abstract
1. Introduction
2. Molecular Modeling
2.1. NR Modeling
2.2. Molecular Dynamics Simulation Methods
3. Proposed Machine Learning Framework
3.1. MD Experimental Data Collection
3.2. Data Preprocessing and Feature Engineering
3.2.1. Data Expansion
3.2.2. Solving Sample Imbalance
3.3. Model Establishment and Improvement
3.4. Model Performance
4. Results and Discussion
4.1. Performance Analysis of MD-XGB
4.2. Analysis of Variable Feature Importance
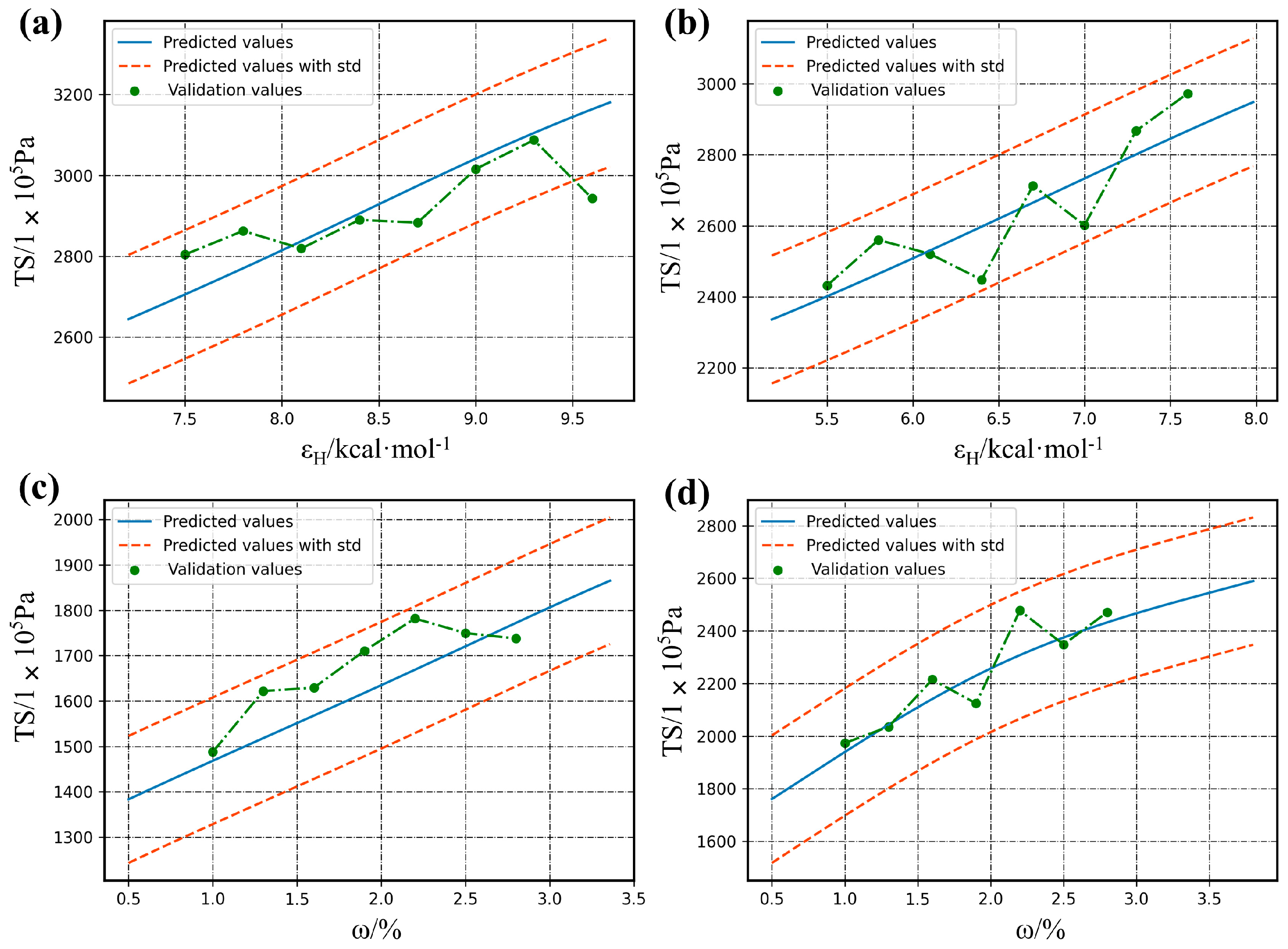
4.3. Model Visualization and External Validation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cornish, K. Similarities and Differences in Rubber Biochemistry among Plant Species. Phytochemistry 2001, 57, 1123–1134. [Google Scholar] [CrossRef]
- Toki, S.; Sics, I.; Ran, S.; Liu, L.; Hsiao, B.S.; Murakami, S.; Senoo, K.; Kohjiya, S. New Insights into Structural Development in Natural Rubber during Uniaxial Deformation by In Situ Synchrotron X-ray Diffraction. Macromolecules 2002, 35, 6578–6584. [Google Scholar] [CrossRef]
- Trabelsi, S.; Albouy, P.-A.; Rault, J. Stress-Induced Crystallization around a Crack Tip in Natural Rubber. Macromolecules 2002, 35, 10054–10061. [Google Scholar] [CrossRef]
- Le Cam, J.-B.; Huneau, B.; Verron, E.; Gornet, L. Mechanism of Fatigue Crack Growth in Carbon Black Filled Natural Rubber. Macromolecules 2004, 37, 5011–5017. [Google Scholar] [CrossRef]
- Ikeda, Y.; Yasuda, Y.; Hijikata, K.; Tosaka, M.; Kohjiya, S. Comparative Study on Strain-Induced Crystallization Behavior of Peroxide Cross-Linked and Sulfur Cross-Linked Natural Rubber. Macromolecules 2008, 41, 5876–5884. [Google Scholar] [CrossRef]
- Men, X.; Wang, F.; Chen, G.-Q.; Zhang, H.-B.; Xian, M. Biosynthesis of Natural Rubber: Current State and Perspectives. Int. J. Mol. Sci. 2018, 20, 50. [Google Scholar] [CrossRef]
- Sakdapipanich, J.; Kalah, R.; Nimpaiboon, A.; Ho, C.C. Influence of Mixed Layer of Proteins and Phospholipids on the Unique Film Formation Behavior of Hevea Natural Rubber Latex. Colloids Surf. A Physicochem. Eng. Asp. 2015, 466, 100–106. [Google Scholar] [CrossRef]
- Wei, Y.-C.; Liu, G.-X.; Zhang, L.; Xu, W.-Z.; Liao, S.; Luo, M.-C. Mimicking the Mechanical Robustness of Natural Rubber Based on a Sacrificial Network Constructed by Phospholipids. ACS Appl. Mater. Interfaces 2020, 12, 14468–14475. [Google Scholar] [CrossRef]
- Jong, L. Toughness of Natural Rubber Composites Reinforced with Hydrolyzed and Modified Wheat Gluten Aggregates. J. Polym. Environ. 2015, 23, 541–550. [Google Scholar] [CrossRef]
- Jong, L. Influence of Protein Hydrolysis on the Mechanical Properties of Natural Rubber Composites Reinforced with Soy Protein Particles. Ind. Crops Prod. 2015, 65, 102–109. [Google Scholar] [CrossRef]
- Kosugi, K.; Kawahara, S. Natural Rubber with Nanomatrix of Non-Rubber Components Observed by Focused Ion Beam-Scanning Electron Microscopy. Colloid Polym. Sci. 2015, 293, 135–141. [Google Scholar] [CrossRef]
- Sliozberg, Y.R.; Yeh, I.-C.; Kröger, M.; Masser, K.A.; Lenhart, J.L.; Andzelm, J.W. Ordering and Crystallization of Entangled Polyethylene Melts under Uniaxial Tension: A Molecular Dynamics Study. Macromolecules 2018, 51, 9635–9648. [Google Scholar] [CrossRef]
- Hamer, M.J.; Iyer, B.V.S.; Yashin, V.V.; Kowalewski, T.; Matyjaszewski, K.; Balazs, A.C. Modeling Polymer Grafted Nanoparticle Networks Reinforced by High-Strength Chains. Soft Matter 2014, 10, 1374–1383. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Xu, Q.; Jin, Y.; Qian, X.; Liu, L.; Liu, J.; Ganesan, V. Design of End-to-End Assembly of Side-Grafted Nanorods in a Homopolymer Matrix. Macromolecules 2018, 51, 4143–4157. [Google Scholar] [CrossRef]
- Shen, J.; Li, X.; Shen, X.; Liu, J. Insight into the Dispersion Mechanism of Polymer-Grafted Nanorods in Polymer Nanocomposites: A Molecular Dynamics Simulation Study. Macromolecules 2017, 50, 687–699. [Google Scholar] [CrossRef]
- Li, W.; Bačová, P.; Behbahani, A.F.; Burkhart, C.; Polińska, P.; Harmandaris, V.; Doxastakis, M. Tailoring Interfacial Properties in Polymer–Silica Nanocomposites via Surface Modification: An Atomistic Simulation Study. ACS Appl. Polym. Mater. 2021, 3, 2576–2587. [Google Scholar] [CrossRef]
- Zhang, Z.; Liu, J.; Li, S.; Gao, K.; Ganesan, V.; Zhang, L. Constructing Sacrificial Multiple Networks to Toughen Elastomer. Macromolecules 2019, 52, 4154–4168. [Google Scholar] [CrossRef]
- Wan, H.; Shen, J.; Gao, N.; Liu, J.; Gao, Y.; Zhang, L. Tailoring the Mechanical Properties by Molecular Integration of Flexible and Stiff Polymer Networks. Soft Matter 2018, 14, 2379–2390. [Google Scholar] [CrossRef]
- Gao, N.; Hou, G.; Liu, J.; Shen, J.; Gao, Y.; Lyulin, A.V.; Zhang, L. Tailoring the Mechanical Properties of Polymer Nanocomposites: Via Interfacial Engineering. Phys. Chem. Chem. Phys. 2019, 21, 18714–18726. [Google Scholar] [CrossRef]
- Butler, K.T.; Davies, D.W.; Cartwright, H.; Isayev, O.; Walsh, A. Machine Learning for Molecular and Materials Science. Nature 2018, 559, 547–555. [Google Scholar] [CrossRef]
- Liu, Y.; Niu, C.; Wang, Z.; Gan, Y.; Zhu, Y.; Sun, S.; Shen, T. Machine Learning in Materials Genome Initiative: A Review. J. Mater. Sci. Technol. 2020, 57, 113–122. [Google Scholar] [CrossRef]
- Schütt, K.T.; Arbabzadah, F.; Chmiela, S.; Müller, K.R.; Tkatchenko, A. Quantum-Chemical Insights from Deep Tensor Neural Networks. Nat. Commun. 2017, 8, 13890. [Google Scholar] [CrossRef] [PubMed]
- von Lilienfeld, O.A.; Ramakrishnan, R.; Rupp, M.; Knoll, A. Fourier Series of Atomic Radial Distribution Functions: A Molecular Fingerprint for Machine Learning Models of Quantum Chemical Properties. Int. J. Quantum Chem. 2015, 115, 1084–1093. [Google Scholar] [CrossRef]
- Chmiela, S.; Tkatchenko, A.; Sauceda, H.E.; Poltavsky, I.; Schütt, K.T.; Müller, K.-R. Machine Learning of Accurate Energy-Conserving Molecular Force Fields. Sci. Adv. 2017, 3, e1603015. [Google Scholar] [CrossRef]
- Rupp, M.; Ramakrishnan, R.; von Lilienfeld, O.A. Machine Learning for Quantum Mechanical Properties of Atoms in Molecules. J. Phys. Chem. Lett. 2015, 6, 3309–3313. [Google Scholar] [CrossRef]
- Chen, C.; Lu, Z.; Ciucci, F. Data Mining of Molecular Dynamics Data Reveals Li Diffusion Characteristics in Garnet Li7La3Zr2O12. Sci. Rep. 2017, 7, 40769. [Google Scholar] [CrossRef]
- Lu, S.; Zhou, Q.; Ouyang, Y.; Guo, Y.; Li, Q.; Wang, J. Accelerated Discovery of Stable Lead-Free Hybrid Organic-Inorganic Perovskites via Machine Learning. Nat. Commun. 2018, 9, 3405. [Google Scholar] [CrossRef]
- Rahman, A.; Deshpande, P.; Radue, M.S.; Odegard, G.M.; Gowtham, S.; Ghosh, S.; Spear, A.D. A Machine Learning Framework for Predicting the Shear Strength of Carbon Nanotube-Polymer Interfaces Based on Molecular Dynamics Simulation Data. Compos. Sci. Technol. 2021, 207, 108627. [Google Scholar] [CrossRef]
- Wang, Y.; Yao, Q.; Kwok, J.T.; Ni, L.M. Generalizing from a Few Examples. ACM Comput. Surv. 2021, 53, 1–34. [Google Scholar] [CrossRef]
- Xie, B.-G.; Wang, H.; Lu, R.-L.; Wang, H.; Xia, R.; Chen, P.; Qian, J.-S. A Combined Simulation and Experiment Study on Polyisoprene Rubber Composites. Compos. Sci. Technol. 2020, 200, 108398. [Google Scholar] [CrossRef]
- Kroll, D.M.; Croll, S.G. Influence of Crosslinking Functionality, Temperature and Conversion on Heterogeneities in Polymer Networks. Polymer 2015, 79, 82–90. [Google Scholar] [CrossRef]
- Bennemann, C.; Paul, W.; Baschnagel, J.; Binder, K. Investigating the Influence of Different Thermodynamic Paths on the Structural Relaxation in a Glass-Forming Polymer Melt. J. Phys. Condens. Matter 1999, 11, 2179–2192. [Google Scholar] [CrossRef]
- Everaers, R. Constrained Fluctuation Theories of Rubber Elasticity: General Results and an Exactly Solvable Model. Eur. Phys. J. B 1998, 4, 341–350. [Google Scholar] [CrossRef]
- Plimpton, S. Fast Parallel Algorithms for Short-Range Molecular Dynamics. J. Comput. Phys. 1995, 117, 1–19. [Google Scholar] [CrossRef]
- Jiang, N.; Wang, L. Quantum Image Scaling Using Nearest Neighbor Interpolation. Quantum Inf. Process. 2015, 14, 1559–1571. [Google Scholar] [CrossRef]
- Dunlop, G.R. A Rapid Computational Method for Improvements to Nearest Neighbour Interpolation. Comput. Math. Appl. 1980, 6, 349–353. [Google Scholar] [CrossRef][Green Version]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic Minority Over-Sampling Technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Liu, T.; Liu, L.; Cui, F.; Ding, F.; Zhang, Q.; Li, Y. Predicting the Performance of Polyvinylidene Fluoride, Polyethersulfone and Polysulfone Filtration Membranes Using Machine Learning. J. Mater. Chem. A 2020, 8, 21862–21871. [Google Scholar] [CrossRef]
- Lundberg, S.; Lee, S.-I. A Unified Approach to Interpreting Model Predictions. Adv. Neural Inf. Process. Syst. 2017. [Google Scholar] [CrossRef]
- Chen, Q.; Zhang, Z.; Huang, Y.; Zhao, H.; Chen, Z.; Gao, K.; Yue, T.; Zhang, L.; Liu, J. Structure–Mechanics Relation of Natural Rubber: Insights from Molecular Dynamics Simulations. ACS Appl. Polym. Mater. 2022. [Google Scholar] [CrossRef]
- Piegl, L.A.; Tiller, W. Algorithm for Finding All k Nearest Neighbors. Comput. Des. 2002, 34, 167–172. [Google Scholar] [CrossRef]
- Montavon, G.; Orr, G.B.; Mueller, K.-R. Neural Networks: Tricks of the Trade, 2nd ed.; Springer: Berlin, Germany, 2012; pp. 639–655. [Google Scholar]








| Hydrogen Bond Interaction | ||||
|---|---|---|---|---|
| Hydrogen Bond Pair | ||||
| Phosphate group | Phospholipid | 3.80 | 4.89 | 12.5 |
| Polypeptide | Protein | 3.80 | 4.89 | 12.5 |
| Phospholipid | Phospholipid | 3.80 | 4.89 | 12.5 |
| Protein | Protein | 3.80 | 4.89 | 12.5 |
| Non-Hydrogen Bond Interaction | ||||
|---|---|---|---|---|
| Non-Hydrogen Bond Pair | ||||
| cis-1,4-polyisoprene repeating unit | Protein | 0.38 | 4.89 | 12.5 |
| cis-1,4-polyisoprene repeating unit | Phospholipid | 0.38 | 4.89 | 12.5 |
| Metrics | Training Set | Testing Set | ||||
|---|---|---|---|---|---|---|
| XGB | SVR | MLR | XGB | SVR | MLR | |
| 0.968 | 0.905 | 0.826 | 0.964 | 0.876 | 0.792 | |
| COV | 0.886 | 0.828 | 0.800 | 0.878 | 0.835 | 0.787 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, Y.; Chen, Q.; Zhang, Z.; Gao, K.; Hu, A.; Dong, Y.; Liu, J.; Cui, L. A Machine Learning Framework to Predict the Tensile Stress of Natural Rubber: Based on Molecular Dynamics Simulation Data. Polymers 2022, 14, 1897. https://doi.org/10.3390/polym14091897
Huang Y, Chen Q, Zhang Z, Gao K, Hu A, Dong Y, Liu J, Cui L. A Machine Learning Framework to Predict the Tensile Stress of Natural Rubber: Based on Molecular Dynamics Simulation Data. Polymers. 2022; 14(9):1897. https://doi.org/10.3390/polym14091897
Chicago/Turabian StyleHuang, Yongdi, Qionghai Chen, Zhiyu Zhang, Ke Gao, Anwen Hu, Yining Dong, Jun Liu, and Lihong Cui. 2022. "A Machine Learning Framework to Predict the Tensile Stress of Natural Rubber: Based on Molecular Dynamics Simulation Data" Polymers 14, no. 9: 1897. https://doi.org/10.3390/polym14091897
APA StyleHuang, Y., Chen, Q., Zhang, Z., Gao, K., Hu, A., Dong, Y., Liu, J., & Cui, L. (2022). A Machine Learning Framework to Predict the Tensile Stress of Natural Rubber: Based on Molecular Dynamics Simulation Data. Polymers, 14(9), 1897. https://doi.org/10.3390/polym14091897





