Abstract
Alfalfa is a perennial herbaceous forage legume that is significantly and adversely affected by monocropping. Crop rotation is the most effective measure to overcome continuous cropping obstacles. However, the mechanisms of how bacterial communities are affected and the potential links between these effects and cropping systems remain poorly understood. Based on a long-term field experiments with continuous alfalfa and forage crops with alfalfa rotation in the black soil region of the western Songnen Plain in Northeast China, the alterations in soil bacterial community structure using high-throughput sequencing of the 16S rRNA gene and soil chemical properties and enzyme activities were analyzed. The alfalfa–forage oats–silage maize–alfalfa and alfalfa–silage maize–forage oats–alfalfa system significantly increase the levels of total phosphorus and available phosphorus, and promote the activities of acid phosphatase, β-glucosidase, leucine aminopeptidase, and N-acetyl-β-glucosaminidase in comparison to continuous alfalfa. While alfalfa crop rotation did not affect the α-diversity of soil bacteria, it significantly altered the bacterial community composition and structure. Some key taxa are significantly enriched in the crop rotation system soils, including Bacillus, Sphingobium, Paenibacillus, Hydrogenispora, Rubrobacter, Haliangium, and Rubellimicrobium. Additionally, crop rotation with alfalfa increased the stability and complexity of the soil bacterial co-occurrence network. Based on our findings, we recommend promoting the alfalfa–forage oats–silage maize–alfalfa and alfalfa–silage maize–forage oats–alfalfa rotation systems as ideal practices for overcoming the challenges associated with continuous cropping of alfalfa. These systems not only enhance soil nutrient content and enzyme activities but also foster a beneficial microbial community, ultimately improving soil functionality and crop performance.
1. Introduction
The practice of growing the same crop on the same land year after year without interruption is a common farming practice due to economic benefit or natural agroecological conditions [1]. However, this approach usually leads to severe continuous cropping obstacles (CCOs), with poor crop growth, increased incidence of pests and diseases, and a decrease in both yield and quality [2,3]. Continuous cropping obstacles affect several crops, including medicinal plants [4], watermelon [5], cotton [6], and alfalfa [7]. Numerous studies have demonstrated that the development of CCOs for crops is highly complex, involving factors such as damaged soil property, reduced soil enzyme activity, imbalance in soil microbial community structure and allelopathic-autotoxic effects [8]. Decades of continuous sugarcane cropping resulted in soil acidification, a reduction in the carbon–nitrogen ratio, an increase in alkali hydrolyzable nitrogen, a decline in organic matter as well as total sulfur content when compared to newly planted fields [9]. Additionally, the accumulation of self-toxic substances, such as phenolic acids, in soils from continuous cropping inhibits the growth of crops such as cucumbers, strawberries, and tobacco [10,11,12]. Many studies have also shown that continuous cropping significantly increases the relative abundance of fungal pathogens, leading to severe diseases in corn, soybeans, and melons [13,14,15].
Alfalfa (Medicago sativa L.) is a perennial leguminous plant renowned as the “Queen of forages” for its high yield, rich protein content, excellent palatability and wide adaptability [16,17]. Furthermore, alfalfa exhibits a high ability of root nodulation with N2-fixing Rhizobia and produces numerous fibrous roots, which enhances soil organic matter and improves soil aggregate structure [18]. The Songnen Plain in northeast China serves as a crucial region for the development of alfalfa industry, is also an important commodity grain base and livestock husbandry development base. This region plays an important role in safeguarding national food security and regional ecological stability [19,20]. However, in order to meet the demand for animal feed, the widespread cultivation of alfalfa in the region has led to an increasingly common phenomenon of continuous cropping, consequently resulting in a gradual decline in production capacity year by year [21]. The reduced production capacity is demonstrated by a number of factors, such as declined root activity, decreased seed germination rates, increased incidences of root rot, and challenges in resowing or replanting [22]. This phenomenon is a typical CCOs of alfalfa, which is accompanied by a decline in soil quality. This, in turn, is transforming the black soil area in the western part of the Songnen Plain from a “functional ecological zone” to a “vulnerable ecological zone”. Therefore, urgent measures are needed to address the adverse effects of CCOs on alfalfa yield and ecosystem health.
Crop rotation is an effective approach to alleviate the CCOs. Rational crop rotation not only enhances ecological and economic benefits but also effectively improves soil physical and chemical properties, thereby achieving increased yields and income [23,24]. For example, the implementation of a spring wheat–alfalfa crop rotation can obviously reduce the loss of soil nutrients in semi-arid areas, thereby promoting the sustainability of agroecosystems [25]. Alfalfa crop rotation compared with continuous corn is a means to greatly reduce the leaching of NO3−-N to the aquifer and improve the surface soil, with the potential to increase soil organic carbon sequestration [26]. Previous studies have pointed out that, compared with continuous cropping, crop rotation could increase soil microbial biomass, alter microbial diversity, and mediate community composition [27,28,29]. The incorporation of alfalfa into a tomato–maize rotation has been demonstrated to result in the recruitment of a diverse array of bacterial and fungal taxa, including arbuscular mycorrhizal fungi, and taxa associated with plant growth promoting and biocontrol abilities [30]. However, most research available primarily focuses on the rotational patterns of alfalfa with crops such as corn, soybeans, and wheat, whereas studies on the rotational benefits of alfalfa with other forage crops are relatively limited. Furthermore, investigations into the formation process of soil bacterial communities with what key bacteria are recruited by alfalfa under different rotational systems remain unexplored.
This study conducted field experiments on alfalfa continuous cropping and different alfalfa crop rotation systems in the black soil region of the western Songnen Plain, China. The changes in soil physicochemical properties and soil enzyme activities were analyzed, and the effects of different alfalfa crop rotation systems on soil bacterial community structure were investigated by using high-throughput sequencing technology. The objective of this study is to delve into the interrelationship between different alfalfa crop rotation systems and soil microbial community structure. The findings will provide crucial insights for the design of strategies to enrich microbial communities, and thus alleviating the obstacles of alfalfa monocropping.
2. Materials and Methods
2.1. Experimental Design
This study established a continuous cropping/crop rotation platform of alfalfa in Fulaerji district, Qiqihar city, Heilongjiang province, China (47°15′ N, 123°41′ E). The area belongs to the black soil region of the western Songnen Plain, with an average annual temperature of 3 °C and an average annual precipitation of 450 mm. The soil studied is aeolian sandy soil in a semiarid region, with an average pH value above 7.0 and salinity of approximately 0.24%.
The experimental plots were established in 2011, and a continuous cropping of alfalfa was conducted for eight years until 2019. Then, in 2019, the land was ploughed and a crop rotation system was implemented. By 2023, the first crop rotation cycle was completed. The experiment consisted of four crop rotation treatments: alfalfa–silage maize–silage maize–alfalfa (ACCA); alfalfa–silage maize–forage oats–alfalfa (ACOA); alfalfa–forage oats–forage oats–alfalfa (AOOA); and alfalfa–forage oats–silage maize–alfalfa (AOCA). Continuous alfalfa cropping served as the control, denoted as 8 years of alfalfa (AAAA). There was a total of 5 treatments, each covering an area of 300 m2. Six replicates were set up for each treatment, with a total of 30 plots. Before the start of the experiment, alfalfa seeds were sown at a density of 18 kg·ha−1, forage oats at 180 kg·ha−1, and silage maize at 22.5 kg·ha−1. The field layout and crop rotation schedule are shown in Table S1.
2.2. Soil Sample Collection and Soil Parameter Determination
In the final year of the rotation cycle, soil sampling was collected from each treatment during the alfalfa flowering stage. Five replicate samples were collected in a five-point method and then homogenized to provide one composite sample per replicated plot. Alfalfa roots were randomly uprooted and clipped from crown. Using the soil shaking method, soil adhering to the alfalfa roots was gently shaken off, with soil within 2 mm of the roots considered rhizosphere soil [31]. The bulk soil was collected simultaneously, and the bulk soil collection depth was 20 cm. After mixing each type of soil, they were sieved through a 2 mm mesh. Part of the soil was air-dried for subsequent analysis of soil physicochemical properties, while part was stored at −80 °C for soil DNA extraction.
The soil pH was determined using a pH meter in a soil water suspension (1:2.5 w/v). The soil total carbon (TC) and total nitrogen (TN) contents were measured using an elemental analyzer (VarioEL III, Elementar, Hanau, Germany) [32]. Soil total phosphorus (TP) digested with H2SO4-HClO4, available phosphorus (AP) extracted with 0.5 M NaHCO3, and ammonium nitrogen (NH4+-N) and nitrate nitrogen (NO3−-N) extracted with 2.0 M KCl were assayed using a continuous flow analytical system (SAN, SKALAR, Breda, The Netherlands) [33,34]. Available potassium (AK) extracted with 1.0M CH3COONH4 was quantified using inductively coupled plasma-atomic emission spectrometry (ICPS-7500, Shimadzu, Kyoto, Japan) [35].
Four soil enzymes examined were acid phosphatase (ACP), β-glucosidase (βGC), leucine aminopeptidase (LAP), and N-acetyl-β-D-glucosaminidase (NAG). These enzymes primarily participate in the cycling of elements such as carbon, nitrogen, and phosphorus, and could be used to assess the health and biological activity of soil ecosystems. ACP activity was determined by the p-nitrophenol method [36]. βGC activity was assessed through incubation of the enzyme with 5 mM para-nitrophenyl-d-glucopyranoside (pNPG) in 50 mM citrate buffer (pH 6.0) within a 0.55 mL reaction mixture, maintained at 50 °C for 15 min [37]. The activities of LAP and NAG were determined using a microplate fluorescence method, with L-leucine-7-amido-4-methylcoumarin (L-leucine-AMC) and 4-methylumbelliferyl-N-acetyl-β-d-glucosaminide as fluorogenic substrates, respectively [38,39]. The four enzyme activities were all expressed in µmol day−1 g−1.
2.3. Soil DNA Extraction, PCR Amplification and High-Throughput Sequencing
Soil DNA was extracted from the frozen soil samples (0.5 g wet weight) using a E.Z.N.A Soil DNA Kit (OMEGA, Norcross, GA, USA) according to the manufacturer’s instructions [40]. The extracted DNA was stored in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0). The quality and concentration of DNA were determined using a NanoDrop® ND-2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). The bacterial 16S rRNA gene of the V4-V5 hypervariable regions was amplified using 515F (5′-GTGCCAGCMGCCGCGG-3′) and 907R (5′-CCGTCAATTCMTTTRAGTTT-3′) [41]. Each 25 µL amplification system contains 10 µL of SYBR Premix Ex TaqTM (TaKaRa, Dalian, China), 0.5 µL of forward and reverse primers at a concentration of 10 µM, 2 µL of DNA sample (1–10 ng/µL) as the template, and 12 µL of sterile water to make up the total volume to 25 µL. PCR amplification cycling was initial denaturation at 95 °C for 5 min, followed by 30 cycles (95 °C for 1 min, 63 °C for 1 min, 72 °C for 1 min) with a final extension at 72 °C for 5 min.
Using the QIAEX Agarose Gel Extraction Kit (II) (QIAGEN, Shanghai, China), PCR products were purified. The purified products were quantified using the Quantus™ Fluorometer (Promega, Madison, WI, USA). Each PCR product was purified and pooled in equimolar amounts, and then prepared for sequencing using an Illumina MiSeq platform (PE 300).
2.4. Processing of Bacterial 16S rRNA Sequencing Data
The raw sequences were processed using QIIME (v 1.9.1) (http://qiime.org/tutorials/tutorial.html) (accessed on 15 May 2023) [42]. FLASH software (http://www.cbcb.umd.edu/software/flash) (accessed on 15 May 2023) was employed for merging paired reads [43]. Then, the deblur algorithm was applied to eliminate low-quality sequences, and subsequent quality control involved the removal of chimeric sequences using the Uchime algorithm [44,45]. High-quality sequences were clustered into operational taxonomic units (OTUs) based on a 97% similarity threshold [46]. Taxonomic assignments for bacterial OTU representative sequences were conducted using the RDP naïve Bayesian classifier, following the SILVA (v 128) database [47,48]. The raw sequence data were deposited into the Sequence Read Archive (SRA) database at the National Center for Biotechnology Information (NCBI) under accession number [PRJNA1085785].
2.5. Statistical Analysis
The Shannon diversity index, representing alpha diversity, was computed using the QIIME Pipeline (v 1.9.1). For the beta diversity, principal coordinate analysis (PCoA) based on the Bray–Curtis distance using the “vegan” package in R (v 4.1.0) was conducted to compare the differences in bacterial community structures among different treatments [49,50]. Differential analysis at the OTU level between the continuous cropping and alfalfa crop rotation was conducted using “DESeq2” package in R (v 4.1.0) by fitting a generalized linear model with a negative binomial distribution [51]. OTUs with p < 0.05 and |log2(fold change) | ≥ 1 were considered a significant difference. Venn diagrams for differential enriched OTUs in different treatments were constructed using tool Venny (v 2.1.0). Co-occurrence networks were utilized to assess the relationships between each bacterial OTU with a relative abundance greater than 0.1%. The Spearman coefficient between OTUs was determined using the “Hmisc” and “igraph” packages in R (v 4.1.0) [52]. When the Spearman coefficient between OTUs was greater than 0.8 or less than −0.8, with a significance level of p < 0.05, the OTU was included in the network analysis [53]. The networks were explored and visualized using Gephi (v 0.8.2) [54]. Redundancy analysis (RDA) and Pearson correlation analysis were utilized to determine which soil parameter was significantly correlated with the bacterial community structure [55]. The statistical significance of the soil-related indicators, bacterial α-diversity, and bacterial relative abundance at various taxonomic levels were tested using an analysis of variance (ANOVA) and pairwise comparisons with Tukey’s HDS.
3. Results
3.1. Soil Chemical Properties and Soil Enzyme Activities
Soil TP and AP contents from different alfalfa crop rotation systems were significantly higher than AAAA (Table 1). The value of alfalfa crop rotations for TP was ACOA > AOOA > AOCA > ACCA > AAAA, while that of alfalfa crop rotations for AP was AOOA > ACCA > AOCA > ACOA > AAAA. On the contrary, the alfalfa crop rotation treatments showed average reductions of 5.58%, 7.11%, and 24.2% in TC, TN, and NO3−-N, respectively, compared to AAAA. Nevertheless, the soil pH, NH4+-N, and AK contents in the alfalfa crop rotation treatments were similar to those in the AAAA.

Table 1.
Effects of continuous cropping/crop rotation of alfalfa on soil chemical properties.
Different alfalfa crop rotation systems exhibited a significant effect on the activities of four enzymes (Table 2). In detail, all alfalfa crop rotation systems resulted in an augmentation of ACP, βGC, LAP, and NAG activities, with ACOA and AOCA showing the most pronounced enhancements. ACOA demonstrated a notable increase in the activities of ACP, βGC, LAP, and NAG by 30.0%, 56.3%, 136%, and 94.9%, respectively, compared to AAAA. Similarly, AOCA exhibited a significant increase in the activities of ACP, βGC, LAP, and NAG by 21.4%, 54.0%, 126%, and 96.0%, respectively, compared to AAAA.

Table 2.
Effects of continuous cropping/crop rotation of alfalfa on soil enzyme activities.
3.2. Soil Bacterial Diversity and Structure
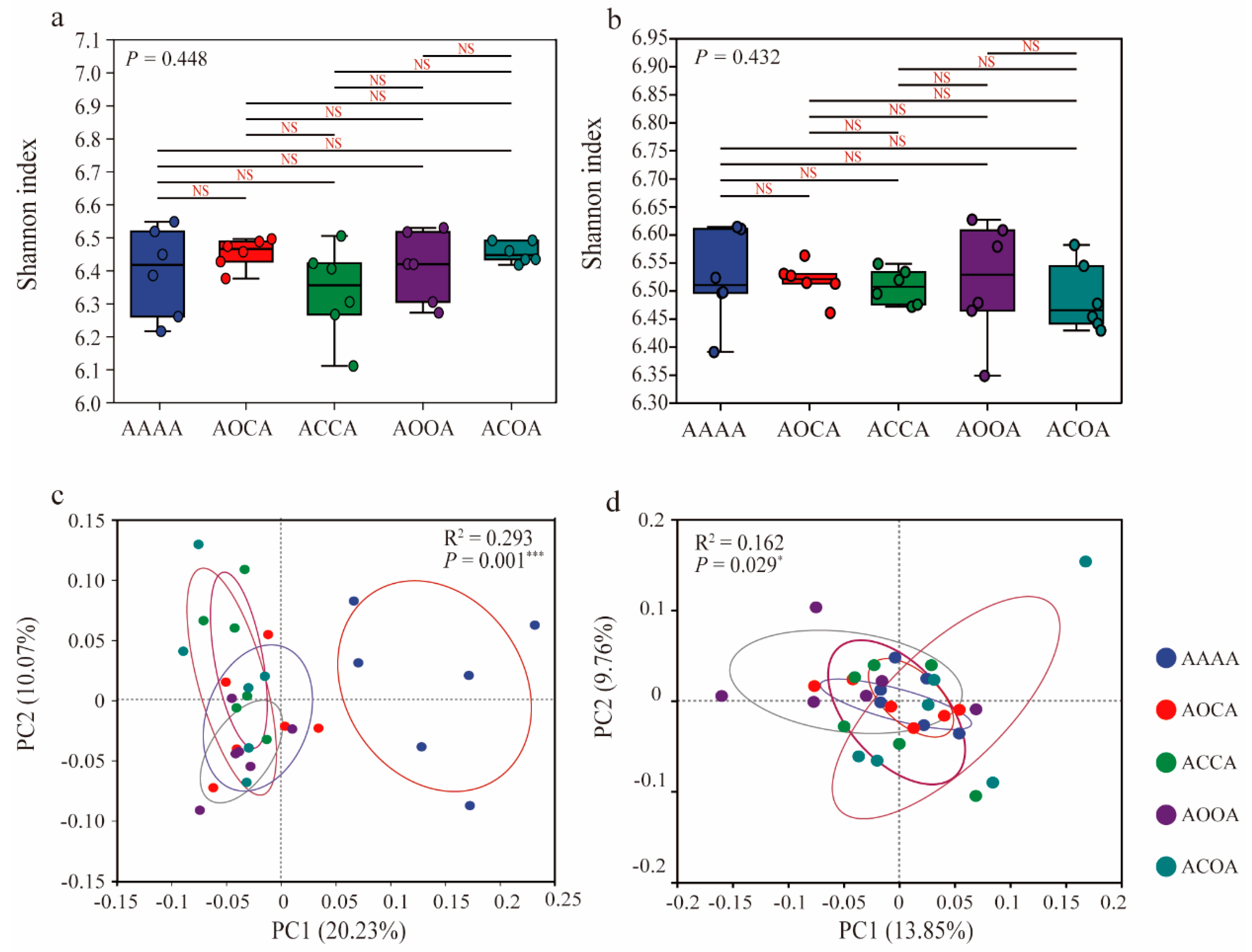
A total of 3,954,125 high-quality 16S rRNA sequences were generated in this study and identified as 13,530 bacterial OTUs. There was no significant difference in bacterial Shannon index among all treatments, regardless of bulk soil or rhizosphere soil (Figure 1a,b). Principal coordinate analysis (PCoA) was employed to analyze the β-diversity of bacterial communities (Figure 1c,d), and showed a significant discrimination between alfalfa crop rotation samples and the continuous cropping samples. This indicates that rotation practices have a notable effect on both bulk soil (p = 0.0001) and rhizosphere soil (p = 0.029) bacterial communities.
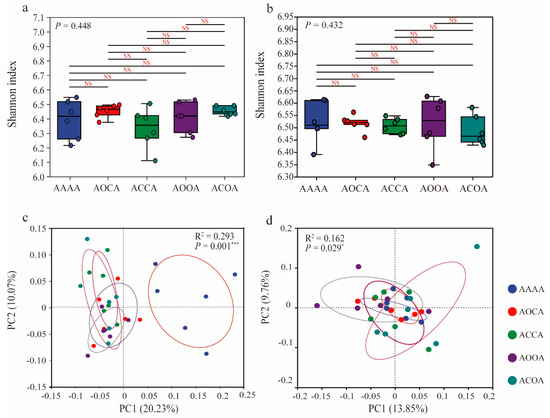
Figure 1.
Effects of continuous cropping/crop rotation of alfalfa on soil bacterial diversity. Bacterial Shannon diversity in bulk soil (a) and rhizosphere soil (b). Principal coordinate analysis (PCoA) based on Bray−Curtis distance for differences in the composition of bacterial communities in bulk soil (c) and rhizosphere soil (d). * p < 0.05; *** p < 0.001; NS: not significant.
Across all bulk soil samples, the dominant bacterial phyla (relative abundance higher than 5%) were Proteobacteria (27.3–31.6%), Actinobacteria (21.7–25.2%), Acidobacteria (11.1–13.3%), and Chloroflexi (6.07–7.05%). Additionally, relative abundance at phyla level in alfalfa crop rotation was improved compared to continuous cropping (Figure 2a). For example, compared to continuous cropping, alfalfa crop rotation resulted in higher richness of Acidobacteriota, Chloroflexi, Gemmatimonadota, and Firmicutes. Conversely, the relative abundance of Proteobacteria and Bacteroidota decreased significantly in the alfalfa crop rotation treatment. In terms of all rhizosphere soil samples, it was found that Proteobacteria had the highest relative abundance (17.8–21.2%), which was significantly lower in ACOA than that in AAAA (Figure 2b). Furthermore, compared to AAAA, an increase in the relative abundance of Firmicutes and Myxococcota was found among different alfalfa crop rotation treatments, although the differences were not significant.
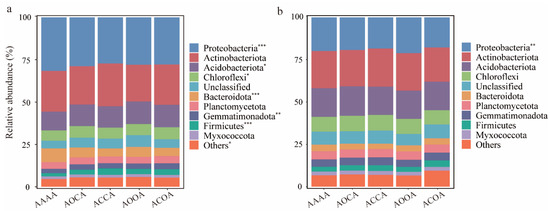
Figure 2.
The relative abundance of soil bacterial communities at phylum level. Changes in the relative abundance of bacterial phylum in bulk soil (a) and rhizosphere soil (b) under different alfalfa continuous cropping/crop rotation treatments. Significance levels are as follows: * p < 0.05; ** p < 0.01; *** p < 0.001.
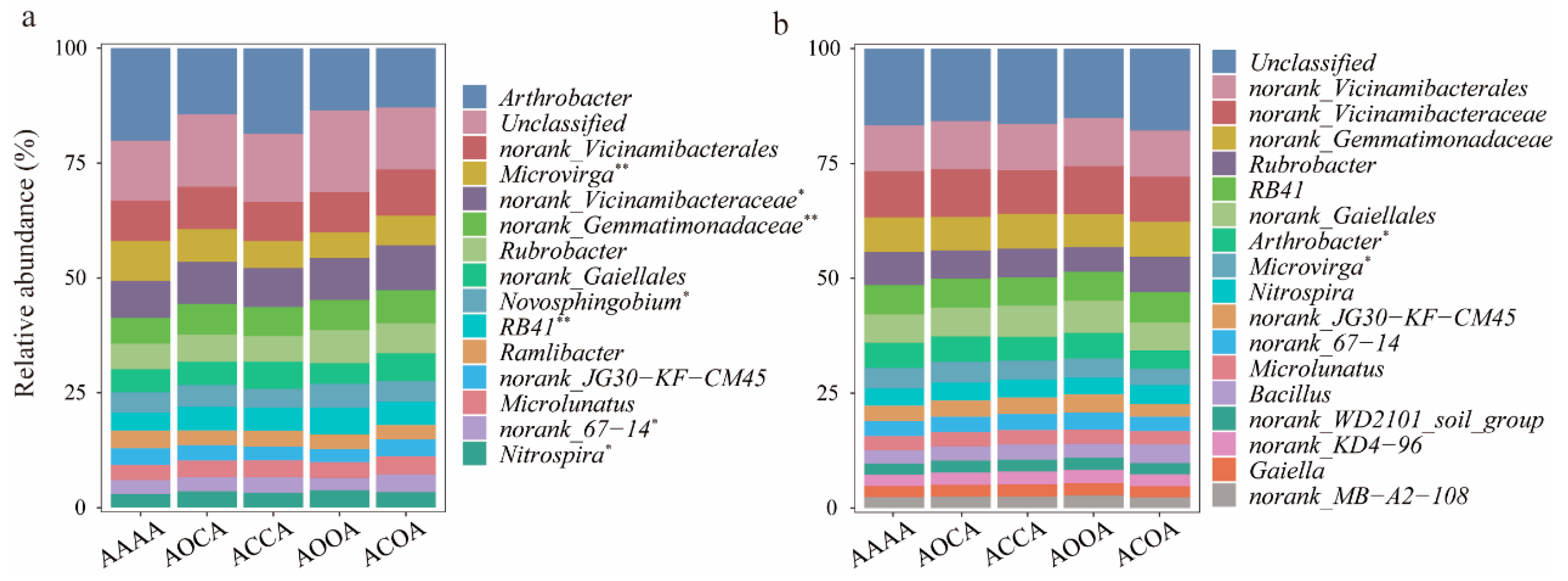
The relative abundance exceeding 1% was considered to be dominant bacterial genera (Figure 3). With regards to bulk soil bacterial communities, the alfalfa crop rotation treatment exhibited higher relative abundance of norank_Vicinamibacteraceae, norank_Gemmatimonadaceae, Novosphingobium, RB41, and Nitrospira, but lower relative abundance of Microvirga and Ramlibacter. For rhizosphere bacterial communities, it was noted that ACOA significantly reduced the relative abundance of Arthrobacter and Microvirga in comparison with AAAA.
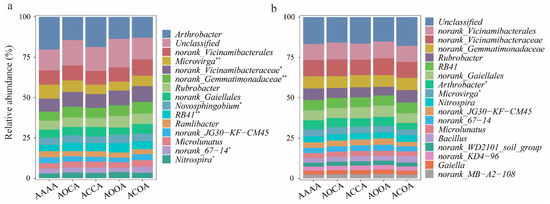
Figure 3.
The relative abundance of soil bacterial communities at genus level. Changes in the relative abundance of dominant genera in bulk soil (a) and rhizosphere soil (b) under different alfalfa continuous cropping/crop rotation treatments. Significance levels are as follows: * p < 0.05; ** p < 0.01.
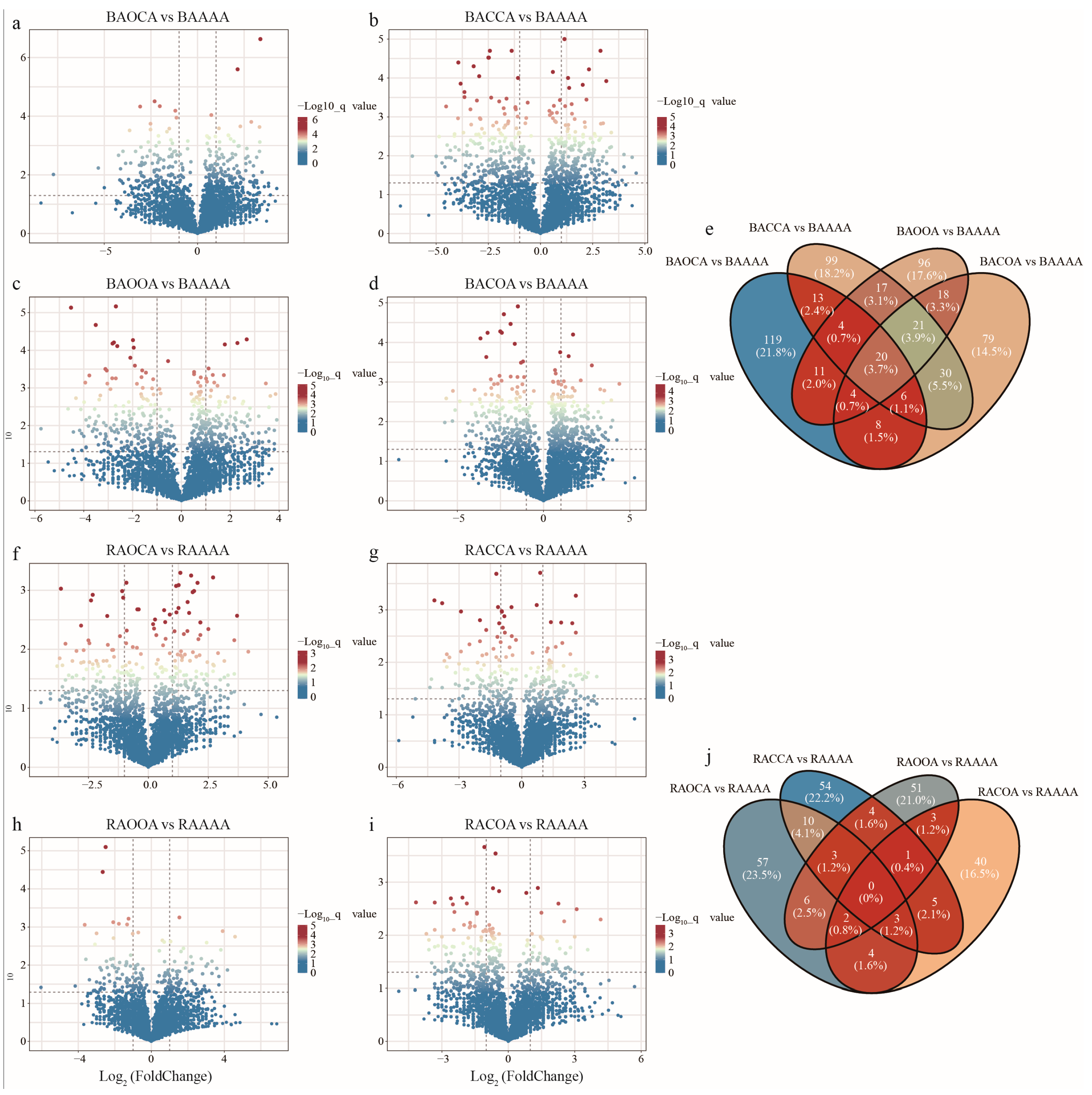
The differences in bacterial species between alfalfa continuous cropping and alfalfa crop rotation at the OTU level were investigated (Table S2). In bulk soil, compared to AAAA, both ACCA and ACOA showed a significantly higher number of depleted OTUs than enriched OTUs, with enriched/depleted OTU numbers being 210/231 and 186/208, respectively (Figure 4a–d). For AOCA and AOOA, the numbers of significantly enriched OTUs were close to the numbers of depleted OTUs, with enriched/depleted OTU numbers being 185/184 and 191/190, respectively. In rhizosphere soil, compared to in AAAA, AOCA exhibited a significantly higher number of enriched OTUs than depleted OTUs, with enriched/depleted OTU numbers being 85/75. Conversely, ACOA showed the opposite pattern, with enriched/depleted OTU numbers being 58/80. Furthermore, both ACCA (80/78) and AOOA (70/70) demonstrated a similar number of significantly enriched and depleted OTUs (Figure 4f–i).
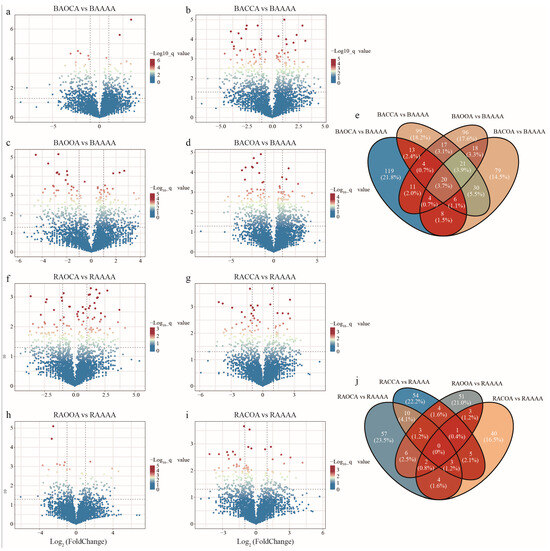
Figure 4.
Certain OTUs are enriched and depressed under alfalfa continuous cropping/crop rotation. Volcano plot was used to describe OTU differences in AOCA (a), ACCA (b), AOOA (c), and ACOA (d) in bulk soil was expressed as BAAAA, BAOCA, BACCA, BAOOA, and BACOA, respectively. The OTU differences in AOCA (f), ACCA (g), AOOA (h), and ACOA (i) in rhizosphere soil was expressed as RAAAA, RAOCA, RACCA, RAOOA and RACOA, respectively. Each point represents an individual OTU, and the position along the y axis represents the abundance fold change compared with the abundance fold change in AAAA. Number of significantly enriched differential OTUs between continuous cropping and rotation treatments in bulk soil (e) and rhizosphere soil (j).
To further explore the relationships among differential OTUs under different planting systems, the unique and shared enriched OTUs between different alfalfa crop rotation and continuous cropping treatments were presented. The Venn diagrams revealed an overlap of 20 OTUs among different rotation treatments in bulk soil (Figure 4e). These OTUs were affiliated with taxa such as Pontibacter, Sphingobium, Bacillus, Parasegetibacter and norank_Vicinamibacteraceae (Table S3), suggesting their potential importance in plant growth under various crop rotation treatments. Additionally, a considerable number of unique OTUs were observed in AOCA, ACCA, AOOA, and ACOA treatments, numbering 119, 99, 96, and 79, respectively. However, the enriched OTUs in rhizosphere soil had no overlap among different crop rotation treatments, indicating a lower likelihood of mutual colonization among these OTUs (Figure 4j). Furthermore, in AOCA treatment, the proportion of unique OTUs was higher, reaching 23.5%. These OTUs belonged to taxa such as Paenibacillus, norank_Ardenticatenales, Sporosarcina, Rubrobacter and Rubellimicrobium, indicating the prevalence of these taxa in response to the AOCA treatment. Meanwhile, specific OTUs such as Pirellula, Phenylobacterium, norank_Roseiflexaceae, Pajaroellobacter, Hydrogenispora, Bacillus, and Segetibacter were significantly enriched in other crop rotation treatments (Table S3).
3.3. Soil Bacterial Co-Occurrence Network
The interrelated characteristics of bacteria in both bulk soil and rhizosphere soil were investigated through the utilization of co-occurrence network analysis at the OTU level (with a relevance abundance threshold of more than 0.1%), then the network topological features of these networks for each treatment were calculated (Figure 5; Table 3). Compared to the BAAAA network, the alfalfa crop rotation network exhibited significantly higher values in average path length, average clustering coefficient, average weighted degree, and modularity, while the number of nodes, edges, and average degree was lower. These results suggest that the alfalfa crop rotation network in the bulk soil is more complex (Figure 5a–e; Table 3). Among each network, BAOCA, BACCA, and BAOOA had similar numbers of nodes, edges, graph density, and average path length. BAOOA showed the highest modularity and average weighted degree.
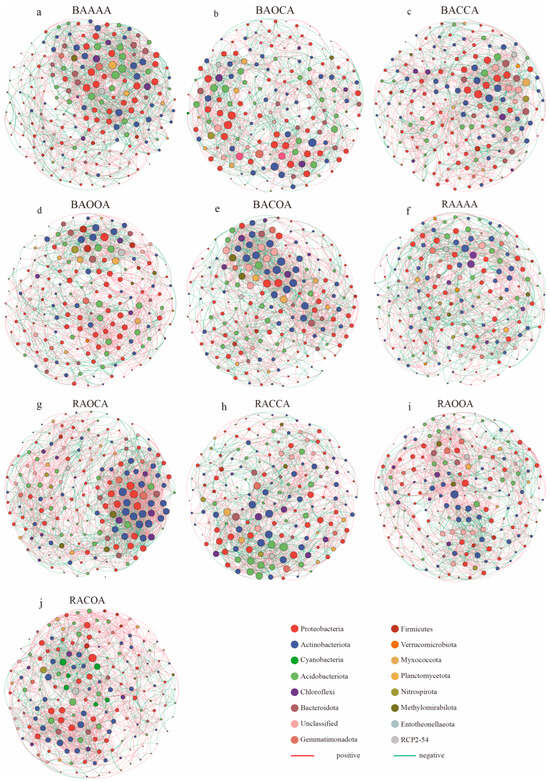
Figure 5.
Effects of continuous cropping/crop rotation of alfalfa on soil bacterial co-occurrence network. The bacterial co-occurrence network of AAAA (a), AOCA (b), ACCA (c), AOOA (d) and ACOA (e) in bulk soil was expressed as BAAAA, BAOCA, BACCA, BAOOA, and BACOA, respectively. The bacterial co-occurrence network of AAAA (f), AOCA (g), ACCA (h), AOOA (i), and ACOA (j) in rhizosphere soil was expressed as RAAAA, RAOCA, RACCA, RAOOA, and RACOA, respectively. The nodes are colored according to bacterial phylum. Node size indicates the degree of connection. Edge color represents positive (red) and negative (green) correlations. The key taxa are annotated on the network.

Table 3.
The co-occurrence network characteristics in the alfalfa continuous cropping/crop rotation system.
Moreover, there were discernible variations in the response of rhizospheric bacterial community networks to different alfalfa crop rotation treatments (Figure 5f–j; Table 3). Specifically, compared to the RAAAA network, both RAOCA and RACOA had similar node numbers but possessed more edges, positive/negative correlations, average degree, graph density and average weighted degree. They also showed lower average path length, average clustering coefficient, and modularity. In contrast, RACCA displayed opposite characteristics with fewer nodes and edges, lower negative correlations, as well as higher average clustering coefficient and modularity. Additionally, RAAAA was similar to RAOOA with slight differences in the number of nodes and edges, but in terms of positive and negative correlations, RAOOA had more positive correlations. Overall, alfalfa crop rotation led to an increased complexity in the bacterial network structure in the rhizosphere soils, while RACOA and RAOCA exhibited even more intricate network structures.
In order to evaluate the possible key role of taxa in the network, closeness centrality and betweenness centrality were used to describe the topological role of bacterial OTU in the network (Table S4). It was found that most of these key taxa belonged to Proteobacteria, Actinobacteriota, and Acidobacteriota. In bulk soil, key taxa from different alfalfa crop rotation treatments included OTU1123 (Lysobacter), OTU7993 (Terrimonas), OTU2302 (Mycetocola), OTU2114 (norank_Gaiellales), and OTU2189 (Acidibacter). In rhizosphere soil, key taxa from different alfalfa crop rotation treatments were OTU2294 (Allorhizobium-Neorhizobium-Pararhizobium-Rhizobium), OTU1426 (norank_Gemmatimonadaceae), OTU1142 (Nocardioides), OTU1042 (Pedomicrobium), and OTU784 (norank_Xanthobacteraceae).
3.4. Correlations between Soil Parameters and Soil Bacterial Community Structure
The redundancy analysis (RDA) showed that the soil parameters (i.e., pH, TC, TN, AP, ACP, and LAP) had different effects on bacterial communities. Apart from ACP, pH, and AK, all other soil parameters significantly altered the structure of bacterial community in bulk soil (Figure 6a). In the rhizosphere soil, only AK was the vital factor that significantly affected the bacterial community structure (p = 0.007, Figure 6b).
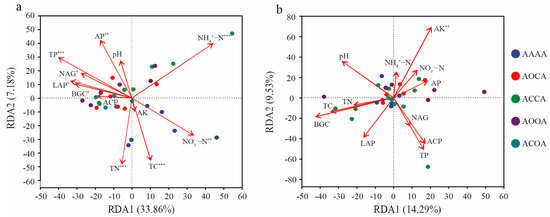
Figure 6.
Redundancy analysis (RDA) of bacterial OTUs and soil parameters. The relationship between bacterial OTUs and soil parameters in bulk soil (a) and rhizosphere soil (b). Significance levels are as follows: * p < 0.05; ** p < 0.01; *** p < 0.001.
The relationship between bacterial phyla and soil parameters was further analyzed (Table S5). For the bulk soil, most of the bacterial phyla were significantly associated with soil nutrients and enzyme activities, and responses to these parameters varied among phyla. For example, Proteobacteria and Bacteroidota were negatively correlated with TP but positively correlated with NO3−−N. In contrast, four other bacterial phyla including Chloroflexi were positively correlated with TP but negatively correlated with NO3−-N. Additionally, Proteobacteria and Bacteroidota also showed a positive response to AP, while Firmicutes and Acidobacteriota exhibited a negative response to AP. Bacteroidota, Chloroflexi, Gemmatimonadota, Firmicutes, and Acidobacteriota demonstrated significant associations with soil enzyme activities. Specifically, Chloroflexi was positively correlated with βGC activity. Bacteroidota was negatively correlated with LAP and NAG activity, while other phyla were positively correlated with these enzyme activities. Only Planctomycetota was negatively correlated with NH4+-N (Figure 7a).

Figure 7.
Heatmap showing the strength of correlations between soil parameters and soil bacterial communities at phylum level in bulk soil (a) and rhizosphere soil (b). Significance levels are as follows: * p < 0.05; ** p < 0.011. Color legend represents the values of correlation coefficients (r). Negative values indicate negative correlations, while positive values indicate positive correlations.
In terms of the rhizosphere soil, the relative abundance of Chloroflexi was negatively correlated with TN and TC. Similarly, the relative abundance of Proteobacteria was also negatively correlated with βGC and LAP activity. However, Actinobacteriota and Firmicutes were positively correlated with AK and LAP activity, respectively. Additionally, Bacteroidota was negatively correlated with soil pH but positively correlated with TC (Figure 7b).
4. Discussion
Crop rotation, as an essential farming management practice, has been demonstrated to effectively enhance the soil microenvironment, increase microbial diversity, and promote the growth of beneficial microorganisms [56,57]. This study demonstrated that alfalfa crop rotation leads to the improvement of soil chemical properties, with significantly higher soil TP and AP than continuous cropping system (Table 1). These improvements in soil chemical properties contribute to the establishment of a more stable and healthier soil ecosystem, thereby reducing the occurrence of soil-borne diseases [58]. However, the present study found that the contents of TC, TN, and NO3− are significantly lower in the alfalfa crop rotation system compared to the continuous cropping system. This indicates that alfalfa continuous cropping increases some soil nutrient indicators. This finding is consistent with the research by Chen et al. (2015), who observed that the soil TC, TN, AN, and AP was significantly increased after continuous cropping. This might be due to the long-term acclimatization where continuous cropping specifically enriched certain microbes that positively contributed to the soil nutrient status [59]. For instance, Wang et al. (2017) discovered that continuous cropping significantly enriched certain microbial species capable of degrading cellulose [60].
Although alfalfa crop rotation significantly altered the microbial community structures in both bulk soils and rhizosphere soils (Figure 1), we found that continuous cropping has a greater impact on bulk soil than on rhizosphere soil, which is inconsistent with previous studies [61]. This discrepancy might be due to the differences in root exudates among various crops, the duration of continuous cropping and soil types, which could affect microbial community structures differently, leading to varying effects observed across different cropping systems [62]. The present study found that AP, TP, NAG, LAP, βGC, TN, TC, NO3−-N, and NH4+-N are the most significant factors influencing the bacterial community structure in different alfalfa crop rotation modes (Figure 6). This aligned with some findings from Lian et al. (2019) [63] and Liu et al. (2020) [61], who discovered that certain soil enzymes, along with TN and TC, played crucial roles in shaping microbial community structures.
In crop rotation systems, the role of complex interactions among soil microorganisms extends beyond maintaining soil health; it directly influences crop growth and resilience [57]. Our study underscores that alfalfa crop rotation fosters stronger inter-microbial relationships, culminating in intricate and resilient ecological networks (Figure 5). The higher numbers of nodes and edges of RAOCA and RACOA of bacterial network in the rhizospheric soils than those of RAAAA soils indicate that RAOCA and RACOA exhibit a greater network size and more microbial taxa involved in the potential microbial interactions than those in RAAAA treatment. These networks, by bolstering soil resilience, significantly contribute to its adaptability to environmental shifts. Similar results have been reported by Fan et al. (2018), that crop rotation promoted soil ecosystem stability and functionality [64].
The increased presence of beneficial soil microorganisms by alfalfa crop rotation, such as nitrogen-fixing bacteria and plant growth-promoting rhizobacteria has significant effects in mitigating the adverse effects of continuous cropping. In fact, Bacillus niacin is responsible for phosphate solubilization and siderophore production [65], while Paenibacillus has been reported to be associated with nitrogen fixation [66]. This not only could contribute to the reduction in the use of chemical pesticides but also naturally boost the health and yield of crops, thereby promoting the development of sustainable agriculture [67]. Thus, our results further highlight the indispensable role of microbial ecosystems in sustaining agricultural productivity and resilience through intricate woven fabric [68].
5. Conclusions
In conclusion, compared to continuous cropping, the alfalfa–forage oats–silage maize–alfalfa and alfalfa–silage maize–forage oats–alfalfa system could significantly increase the contents of TP, AP, ACP, and promote the βGC, LAP, and NAG activities, which is beneficial for enhancing soil nutrient supply and promoting crop growth and development. Although alfalfa crop rotation does not affect the α-diversity of soil bacteria, it significantly alters the bacterial community structure, enriching beneficial bacterial taxa such as Bacillus, Sphingobium, Paenibacillus, Hydrogenispora, Rubrobacter, and Rubellimicrobium. Furthermore, the alfalfa crop rotation systems enhance the stability and complexity of soil bacterial co-occurrence networks. Therefore, it is recommended to promote the adoption of the alfalfa–forage oats–silage maize–alfalfa and alfalfa–silage maize–forage oats–alfalfa system as an ideal crop rotation system for alleviating alfalfa continuous cropping obstacles in the region. This study underscores the vital role of alfalfa crop rotation in mitigating the challenges of continuous cropping, enhancing soil nutrient content, promoting the diversity and abundance of beneficial bacteria, and thus improving soil functionality.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agronomy14071349/s1. Table S1: Field layout of alfalfa continuous cropping/crop rotation system. Note: A represents alfalfa, C represents silage maize, and O represents forage oats; Table S2: Bacterial OTUs that are significantly abundance in the bulk soil and rhizosphere soil (crop rotation vs continues cropping, p < 0.05). Note: B represents the bulk soil samples. R represents the rhizosphere soil samples; Table S3: Common and unique bacterial OTUs in bulk soil and rhizosphere soil samples (crop rotation vs continues cropping). Note: B represents the bulk soil samples. R represents the rhizosphere soil samples; Table S4: Topological characteristics of bacterial hubs observed in the bulk soil and rhizosphere soils across the alfalfa continuous cropping/crop rotation system. Note: B represents the bulk soil samples. R represents the rhizosphere soil samples; Table S5: Pearson correlation analysis of bacterial phylum and soil parameters in bulk soil and rhizosphere soil.
Author Contributions
Y.X.: investigation, formal analysis, funding acquisition, writing—original draft; Z.L.: investigation, formal analysis, writing—original draft; Z.S.: writing—review and editing, supervision; Z.Y.: conceptualization, data curation, formal analysis. X.F.: investigation, resources, data curation; X.W.: data curation, visualization. S.L.: data curation, visualization; H.C.: investigation, methodology, resources; R.W.: conceptualization, methodology, resources, data curation; X.L.: conceptualization, methodology, resources; J.L.: methodology, writing—review and editing, supervision, conceptualization. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Outstanding Youth Fund of Heilongjiang Academy of Agricultural Sciences (CX23JQ05), the Grass-field Rotation Scientist Studio of Heilongjiang Province (202004), the National Natural Science Foundation of China (42177105), the Postdoctoral Fellowship Program of CPSF (GZC20232643).
Data Availability Statement
The original data presented in the study are openly available in the NCBI short-read archive under accession number PRJNA1085785.
Conflicts of Interest
We declare that we do not have any commercial or associative interests that represent conflicts of interest in connection with the work submitted here.
References
- Liu, X.; Herbert, S.J. Fifteen years of research examining cultivation of continuous soybean in northeast China: A review. Field Crop Res. 2002, 79, 1–7. [Google Scholar] [CrossRef]
- Chen, S.; Qi, G.; Luo, T.; Zhang, H.; Jiang, Q.; Wang, R.; Zhao, X. Continuous-cropping tobacco caused variance of chemical properties and structure of bacterial network in soils. Land. Degrad. Dev. 2018, 29, 4106–4120. [Google Scholar] [CrossRef]
- Pervaiz, Z.H.; Iqbal, J.; Zhang, Q.; Chen, D.; Wei, H.; Saleem, M. Continuous cropping alters multiple biotic and abiotic indicators of soil health. Soil Syst. 2020, 4, 59. [Google Scholar] [CrossRef]
- Liao, J.; Xia, P. Continuous cropping obstacles of medicinal plants: Focus on the plant-soil-microbe interaction system in the rhizosphere. Sci. Hortic. 2024, 328, 112927. [Google Scholar] [CrossRef]
- Zheng, X.; Wei, L.; Lv, W.; Zhang, H.; Zhang, Y.; Zhang, H.; Zhang, H.; Zhu, Z.; Ge, T.; Zhang, W. Long-term bioorganic and organic fertilization improved soil quality and multifunctionality under continuous cropping in watermelon. Agric. Ecosyst. Environ. 2024, 359, 108721. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, W.; Li, Y.; Sun, Y.; Bian, X. Effects of long-term continuous cropping system of cotton on soil physical-chemical properties and activities of soil enzyme in oasis in Xinjiang. Sci. Agric. Sin. 2009, 42, 725–733. [Google Scholar]
- Xu, Y.; Liu, J.; Liu, X.; Li, H.; Yang, Z.; Wang, H.; Huang, X.; Lan, L.; An, Y.; Li, L. Continuous cropping of alfalfa (Medicago sativa L.) reduces bacterial diversity and simplifies cooccurrence networks in aeolian sandy soil. Soil Ecol. Lett. 2022, 4, 113–143. [Google Scholar] [CrossRef]
- Zeeshan Ul Haq, M.; Yu, J.; Yao, G.; Yang, H.; Iqbal, H.A.; Tahir, H.; Cui, H.; Liu, Y.; Wu, Y. A systematic review on the continuous cropping obstacles and control strategies in medicinal plants. Int. J. Mol. Sci. 2023, 24, 12470. [Google Scholar] [CrossRef]
- Pang, Z.; Tayyab, M.; Kong, C.; Liu, Q.; Liu, Y.; Hu, C.; Huang, J.; Weng, P.; Islam, W.; Lin, W. Continuous sugarcane planting negatively impacts soil microbial community structure, soil fertility, and sugarcane agronomic parameters. Microorganisms 2021, 9, 2008. [Google Scholar] [CrossRef]
- Chen, D.M.; Yang, Y.H.; Jin, Y.; Wang, H.B.; Duan, Y.Q.; You, C.H. Constituents of autotoxic chemical from rhizosphere soil under flue-cured tobacco continuous cropping. Pratacultural Sci. 2011, 28, 1766–1769. [Google Scholar]
- Li, H.Q.; Zhang, L.L.; Jiang, X.W.; Liu, Q.Z. Allelopathic effects of phenolic acids on the growth and physiological characteristics of strawberry plants. Allelopathy J. 2015, 35, 61. [Google Scholar]
- Wu, F.; Wang, X.; Xue, C. Effect of cinnamic acid on soil microbial characteristics in the cucumber rhizosphere. Eur. J. Soil Biol. 2009, 45, 356–362. [Google Scholar] [CrossRef]
- Bai, L.; Cui, J.; Jie, W.; Cai, B. Analysis of the community compositions of rhizosphere fungi in soybeans continuous cropping fields. Microbiol. Res. 2015, 180, 49–56. [Google Scholar] [CrossRef]
- Li, M.; Wang, J.; Zhou, Q.; Yasen, M. Effects of continuous melon cropping on rhizospheric fungal communities. Rhizosphere 2023, 27, 100726. [Google Scholar] [CrossRef]
- Strom, N.; Hu, W.; Haarith, D.; Chen, S.; Bushley, K. Interactions between soil properties, fungal communities, the soybean cyst nematode, and crop yield under continuous corn and soybean monoculture. Appl. Soil Ecol. 2020, 147, 103388. [Google Scholar] [CrossRef]
- McDonald, I.; Baral, R.; Min, D. Effects of alfalfa and alfalfa-grass mixtures with nitrogen fertilization on dry matter yield and forage nutritive value. J. Anim. Sci. Technol. 2021, 63, 305. [Google Scholar] [CrossRef]
- Suwignyo, B.; Rini, E.A.; Helmiyati, S. The profile of tropical alfalfa in Indonesia: A review. Saudi J. Biol. Sci. 2023, 30, 103504. [Google Scholar]
- Niu, Y.; Luo, Z.; Cai, L.; Coulter, J.A.; Zhang, Y.; Berti, M. Continuous Monoculture of Alfalfa and Annual Crops Influence Soil Organic Matter and Microbial Communities in the Rainfed Loess Plateau of China. Agronomy 2020, 10, 1054. [Google Scholar] [CrossRef]
- Wang, T.; Zhang, W. Priorities for the development of alfalfa pasture in northern China. Fundam. Res. 2023, 3, 225–228. [Google Scholar] [CrossRef]
- Yang, H.; Feng, X.; Wang, H.; Yan, H.; Zhao, P.; Gao, F.; Guo, X.; Xie, B. Long time-series variation of crop yield under drought stress and drought vulnerability curves in Songnen Plain, Northeast China. Ecol. Indic. 2023, 154, 110624. [Google Scholar] [CrossRef]
- Yang, Z.; Xu, Y.; Li, H.; Li, S.; Wang, X.; Chai, H. Difference of bacterial community structure in the Meadow, Maize, and continuous cropped Alfalfa in Northeast China. Front. Microbiol. 2022, 13, 794848. [Google Scholar] [CrossRef]
- Wang, R.; Liu, J.; Jiang, W.; Ji, P.; Li, Y. Metabolomics and microbiomics reveal impacts of rhizosphere metabolites on alfalfa continuous cropping. Front. Microbiol. 2022, 13, 833968. [Google Scholar]
- Kiani, M.; Hernandez-Ramirez, G.; Quideau, S.; Smith, E.; Janzen, H.; Larney, F.J.; Puurveen, D. Quantifying sensitive soil quality indicators across contrasting long-term land management systems: Crop rotations and nutrient regimes. Agric. Ecosyst. Environ. 2017, 248, 123–135. [Google Scholar] [CrossRef]
- Shah, K.K.; Modi, B.; Pandey, H.P.; Subedi, A.; Aryal, G.; Pandey, M.; Shrestha, J.; Fahad, S. Diversified Crop Rotation: An Approach for Sustainable Agriculture Production. Adv. Agric. 2021, 2021, 8924087. [Google Scholar] [CrossRef]
- Li, A.; Wu, Y.; Tai, X.; Cao, S.; Gao, T. Effects of Alfalfa Crop Rotation on Soil Nutrients and Loss of Soil and Nutrients in Semi-Arid Regions. Sustainability 2023, 15, 15164. [Google Scholar] [CrossRef]
- Singh, A.; Afzal, T.; Woodbury, B.; Wortmann, C.; Iqbal, J. Alfalfa in rotation with annual crops reduced nitrate leaching potential. J. Environ. Qual. 2023, 52, 930–938. [Google Scholar] [CrossRef]
- Liu, Q.; Zhao, Y.; Li, T.; Chen, L.; Chen, Y.; Sui, P. Changes in soil microbial biomass, diversity, and activity with crop rotation in cropping systems: A global synthesis. Appl. Soil Ecol. 2023, 186, 104815. [Google Scholar] [CrossRef]
- Stamenov, D.; Đurić, S.; Hajnal, J.T.; Šeremešić, S. Fertilization and crop rotation effects on the number of different groups of microorganisms. Ratar. I Povrt. 2016, 53, 96–100. [Google Scholar]
- Venter, Z.S.; Jacobs, K.; Hawkins, H. The impact of crop rotation on soil microbial diversity: A meta-analysis. Pedobiologia 2016, 59, 215–223. [Google Scholar] [CrossRef]
- Samaddar, S.; Schmidt, R.; Tautges, N.E.; Scow, K. Adding alfalfa to an annual crop rotation shifts the composition and functional responses of tomato rhizosphere microbial communities. Appl. Soil Ecol. 2021, 167, 104102. [Google Scholar] [CrossRef]
- Brtnicky, M.; Kintl, A.; Hammerschmiedt, T.; Mustafa, A.; Elbl, J.; Kucerik, J.; Vyhnanek, T.; Skladanka, J.; Hunady, I.; Holatko, J. Clover Species Specific Influence on Microbial Abundance and Associated Enzyme Activities in Rhizosphere and Non-Rhizosphere Soils. Agronomy 2021, 11, 2214. [Google Scholar] [CrossRef]
- Jones, D.L.; Willett, V.B. Experimental evaluation of methods to quantify dissolved organic nitrogen (DON) and dissolved organic carbon (DOC) in soil. Soil Biol. Biochem. 2006, 38, 991–999. [Google Scholar] [CrossRef]
- Dospatliev, L.; Ivanova, M. Inductively coupled plasma optical emission spectrometry determination of total phosphorus and sulphur in virginia tobacco leaves. Oxid. Commun. 2022, 45, 115. [Google Scholar]
- Miranda, K.M.; Espey, M.G.; Wink, D.A. A Rapid, Simple Spectrophotometric Method for Simultaneous Detection of Nitrate and Nitrite. Nitric Oxide 2001, 5, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Dahlquist, R.L.; Knoll, J.W. Inductively coupled plasma-atomic emission spectrometry: Analysis of biological materials and soils for major, trace, and ultra-trace elements. Appl. Spectrosc. 1978, 32, 1–30. [Google Scholar] [CrossRef]
- Dick, W.A.; Cheng, L.; Wang, P. Soil acid and alkaline phosphatase activity as pH adjustment indicators. Soil Biol. Biochem. 2000, 32, 1915–1919. [Google Scholar]
- Nidhi, A.; Sneha, S.; Annamma, A.; Arvind, L.; Shams, Y.S. Specific Fusion of β-1,4-Endoglucanase and β-1,4-Glucosidase Enhances Cellulolytic Activity and Helps in Channeling of Intermediates. Appl. Environ. Microb. 2012, 78, 7447–7454. [Google Scholar] [CrossRef]
- Linko-Löppönen, S.; Mäkinen, M. A microtiter plate assay for N-acetyl-β-d-glucosaminidase using a fluorogenic substrate. Anal. Biochem. 1985, 148, 50–53. [Google Scholar] [CrossRef]
- Wang, J.; Huang, S.; He, Q.; Bing, H.; Chen, X.; Zhang, X.; Tian, X.; Zhou, J.; Wilcke, W.; Wu, Y. Microplate fluorimetric assay of soil leucine aminopeptidase activity: Alkalization is not needed before fluorescence reading. Biol. Fert. Soils 2020, 56, 281–285. [Google Scholar] [CrossRef]
- Dineen, S.M.; Aranda, R., IV.; Anders, D.L.; Robertson, J.M. An evaluation of commercial DNA extraction kits for the isolation of bacterial spore DNA from soil. J. Appl. Microbiol. 2010, 109, 1886–1896. [Google Scholar] [CrossRef]
- Chen, L.; Luo, Y.; Xu, J.; Yu, Z.; Zhang, K.; Brookes, P.C. Assessment of bacterial communities and predictive functional profiling in soils subjected to short-term fumigation-incubation. Microb. Ecol. 2016, 72, 240–251. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Jeffrey, I.G. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef]
- Amir, A.; McDonald, D.; Navas-Molina, J.A.; Kopylova, E.; Morton, J.T.; Zech Xu, Z.; Kightley, E.P.; Thompson, L.R.; Hyde, E.R.; Gonzalez, A. Deblur rapidly resolves single-nucleotide community sequence patterns. Msystems 2017, 2, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Preheim, S.P.; Perrotta, A.R.; Martin-Platero, A.M.; Gupta, A.; Alm, E.J. Distribution-based clustering: Using ecology to refine the operational taxonomic unit. Appl. Environ. Microb. 2013, 79, 6593–6603. [Google Scholar] [CrossRef] [PubMed]
- Almeida, A.; Mitchell, A.L.; Tarkowska, A.; Finn, R.D. Benchmarking taxonomic assignments based on 16S rRNA gene profiling of the microbiota from commonly sampled environments. Gigascience 2018, 7, giy054. [Google Scholar] [CrossRef] [PubMed]
- McDonald, D.; Price, M.N.; Goodrich, J.; Nawrocki, E.P.; DeSantis, T.Z.; Probst, A.; Andersen, G.L.; Knight, R.; Hugenholtz, P. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 2012, 6, 610–618. [Google Scholar] [CrossRef] [PubMed]
- Lkhagva, E.; Chung, H.; Hong, J.; Tang, W.H.W.; Lee, S.; Hong, S.; Lee, S. The regional diversity of gut microbiome along the GI tract of male C57BL/6 mice. BMC Microbiol. 2021, 21, 1–13. [Google Scholar] [CrossRef]
- Team, R. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2009; Volume 14, pp. 12–21. [Google Scholar]
- Shi, Z.; Shi, S.; Li, W.; Wang, C.; Hu, F. Bacterial community characteristics in epigeic and anecic earthworm vermicompartments within soil-earthworm systems. Pedosphere 2023, 9, 1–21. [Google Scholar] [CrossRef]
- Rojas-Jimenez, K.; Grossart, H.; Cordes, E.; Cortés, J. Fungal communities in sediments along a depth gradient in the Eastern Tropical Pacific. Front. Microbiol. 2020, 11, 575207. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Yang, T.; Bao, Y.; He, P.; Yang, K.; Mei, X.; Wei, Z.; Xu, Y.; Shen, Q.; Banerjee, S. Network analysis and subsequent culturing reveal keystone taxa involved in microbial litter decomposition dynamics. Soil Biol. Biochem. 2021, 157, 108230. [Google Scholar] [CrossRef]
- Cherven, K. Mastering Gephi Network Visualization; Packt Publishing Ltd.: Birmingham, UK, 2015. [Google Scholar]
- Böer, S.I.; Hedtkamp, S.I.; Van Beusekom, J.E.; Fuhrman, J.A.; Boetius, A.; Ramette, A. Time-and sediment depth-related variations in bacterial diversity and community structure in subtidal sands. ISME J. 2009, 3, 780–791. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Singh, U.B.; Sahu, P.K.; Paul, S.; Kumar, A.; Malviya, D.; Singh, S.; Kuppusamy, P.; Singh, P.; Paul, D. Linking soil microbial diversity to modern agriculture practices: A review. Int. J. Environ. Res. Public Health 2022, 19, 3141. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Plaza-Bonilla, D.; Coulter, J.A.; Kutcher, H.R.; Beckie, H.J.; Wang, L.; Floc’H, J.; Hamel, C.; Siddique, K.H.; Li, L. Diversifying crop rotations enhances agroecosystem services and resilience. Adv. Agron. 2022, 173, 299–335. [Google Scholar]
- Stirling, G.; Hayden, H.; Pattison, T.; Stirling, M. Soil Health, Soil Biology, Soilborne Diseases and Sustainable Agriculture: A Guide; Csiro Publishing: Clayton, Australia, 2016. [Google Scholar]
- Chen, T.; Li, J.; Wu, L.K.; Lin, S.; Wang, J.H.; Li, Z.F.; Zhang, Z.Y.; Lin, W.X. Effects of continuous monoculture of Achyranthes bidentata on microbial community structure and functional diversity in soil. Allelopath. J. 2015, 36, 197. [Google Scholar]
- Wang, J.Y.; Xu, J.H.; Wu, L.K.; Wu, H.M.; Zhu, Q.; Kong, L.F.; Lin, W.X. Analysis of physicochemical properties and microbial diversity in rhizosphere soil of Achyranthes bidentata under different cropping years. Acta Ecol. Sin. 2017, 37, 5621–5629. [Google Scholar]
- Liu, Z.; Liu, J.; Yu, Z.; Yao, Q.; Li, Y.; Liang, A.; Zhang, W.; Mi, G.; Jin, J.; Liu, X. Long-term continuous cropping of soybean is comparable to crop rotation in mediating microbial abundance, diversity and community composition. Soil Tillage Res. 2020, 197, 104503. [Google Scholar] [CrossRef]
- Lian, T.; Cheng, L.; Liu, Q.; Yu, T.; Cai, Z.; Nian, H.; Hartmann, M. Potential relevance between soybean nitrogen uptake and rhizosphere prokaryotic communities under waterlogging stress. ISME Commun. 2023, 3, 71. [Google Scholar] [CrossRef]
- Lian, T.; Ma, Q.; Shi, Q.; Cai, Z.; Zhang, Y.; Cheng, Y.; Nian, H. High aluminum stress drives different rhizosphere soil enzyme activities and bacterial community structure between aluminum-tolerant and aluminum-sensitive soybean genotypes. Plant Soil 2019, 440, 409–425. [Google Scholar] [CrossRef]
- Fan, K.; Weisenhorn, P.; Gilbert, J.A.; Chu, H. Wheat rhizosphere harbors a less complex and more stable microbial co-occurrence pattern than bulk soil. Soil Biol. Biochem. 2018, 125, 251–260. [Google Scholar] [CrossRef]
- Bag, S.; Mondal, A.; Banik, A. Exploring tea (Camellia sinensis) microbiome: Insights into the functional characteristics and their impact on tea growth promotion. Microbiol. Res. 2022, 254, 126890. [Google Scholar] [CrossRef] [PubMed]
- Grady, E.N.; MacDonald, J.; Liu, L.; Richman, A.; Yuan, Z. Current knowledge and perspectives of Paenibacillus: A review. Microb. Cell Fact. 2016, 15, 203. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Yu, T.; Shi, Q.; Han, D.; Yu, K.; Wang, L.; Wang, S.; Xiang, H.; Wen, R.; Nian, H. Rhizosphere soil bacterial communities of continuous cropping-tolerant and sensitive soybean genotypes respond differently to long-term continuous cropping in mollisols. Front. Microbiol. 2021, 12, 729047. [Google Scholar] [CrossRef]
- Rastogi, M.; Verma, S.; Kumar, S.; Bharti, S.; Kumar, G.; Azam, K.; Singh, V. Soil health and sustainability in the age of organic amendments: A review. Int. J. Environ. Clim. Chang. 2023, 13, 2088–2102. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).