Transcriptome Insights into Resistance Mechanisms Against Soybean Mosaic Virus Strain SC4 in Soybean
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials, SMV Strain, and SMV Inoculation
2.2. Serological Determination of Sequenced Materials
2.3. Total RNA Extraction and cDNA Library Construction for Transcriptome Analysis
2.4. Principal Component Analysis
2.5. Differentially Expressed Genes (DEGs)
2.6. Gene Ontology Analysis of DEGs
2.7. Pathway Enrichment Analysis
3. Results
3.1. Transcriptome or RNA-Seq Analysis
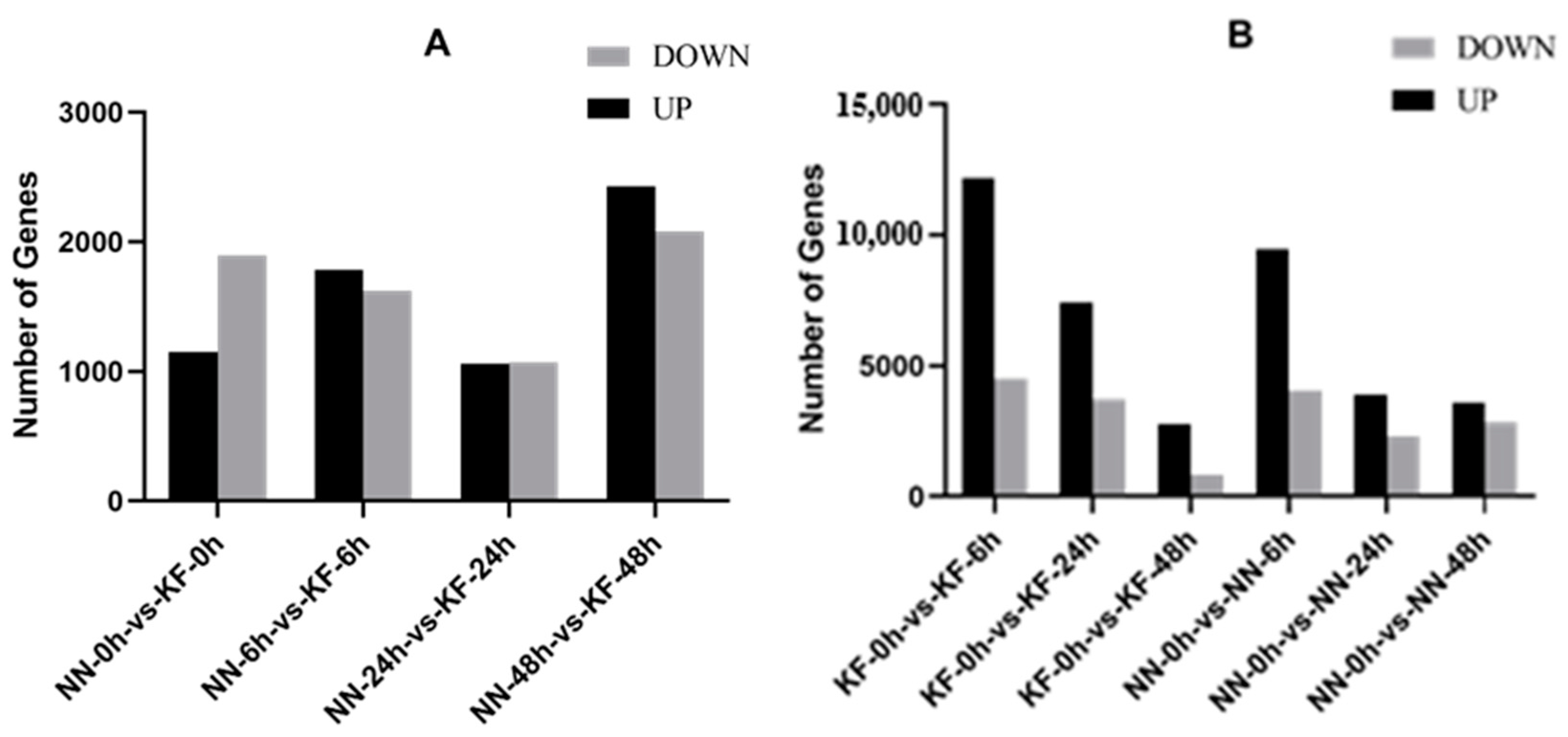
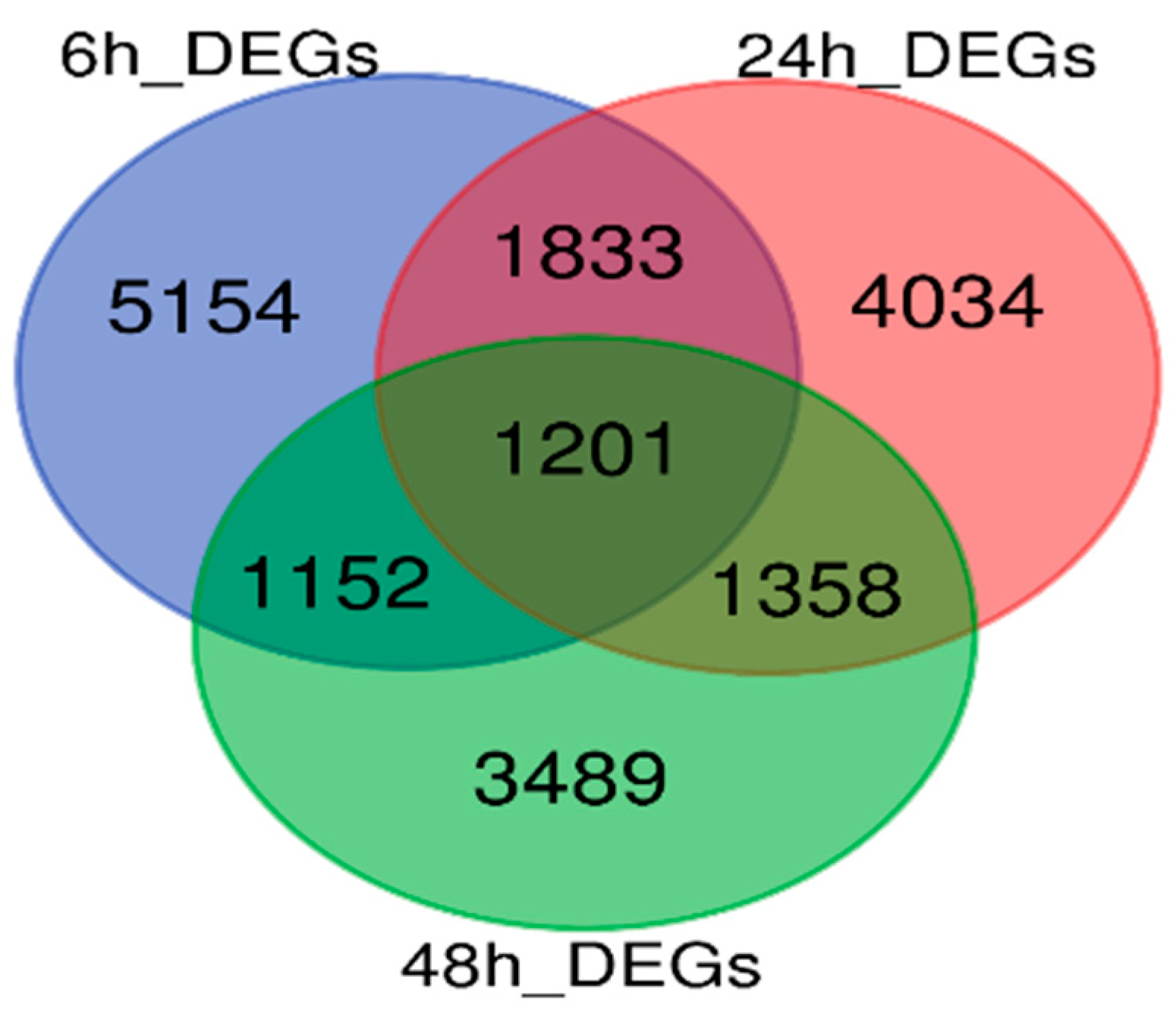
3.2. Identification of DEGs at Four-Time Stages After SC4-Inoculation
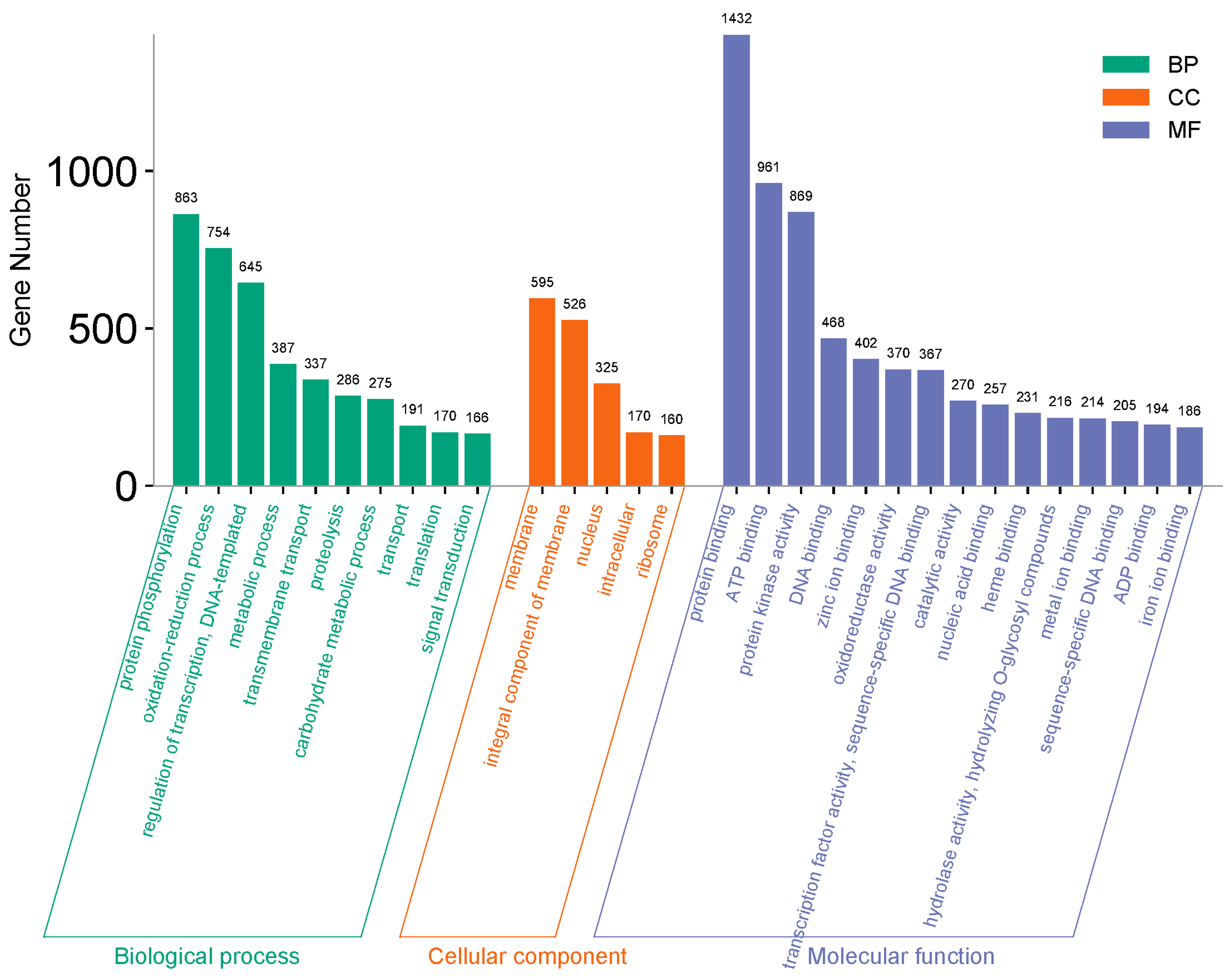
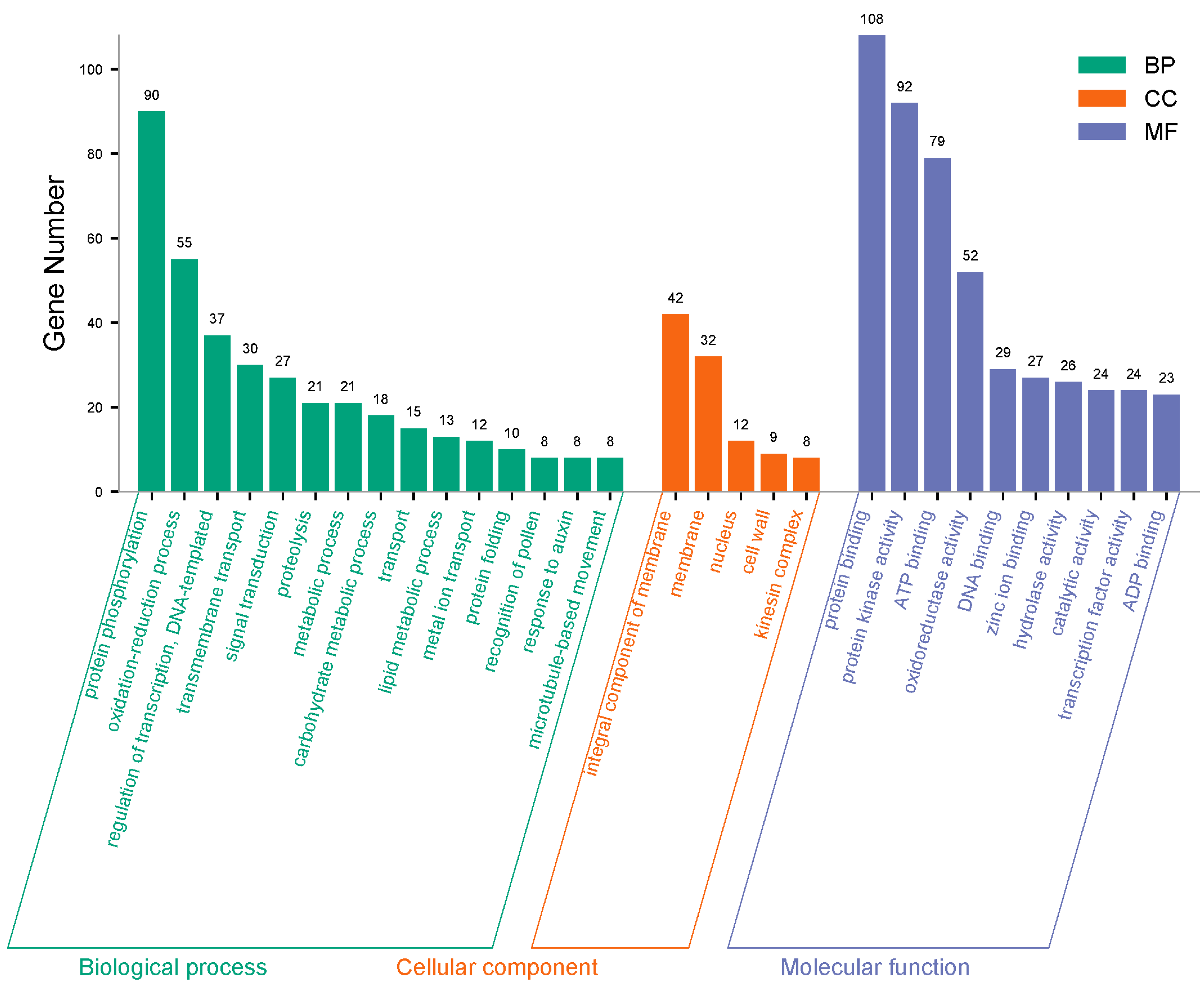
3.3. GO Enrichment Analysis of the Total 18,221 DEGs and 1201 Joint DEGs Among Stages
3.4. Protein Phosphorylation, Signal Transduction and Protein Kinase Activity in Early Defense Response to SMV
3.5. Oxidation-Reduction and Oxidoreductase Activity in Resistance
3.6. Genes Related to Signal Transduction and Response to Auxin in Soybean
4. Discussion
4.1. Transcriptome Study on the Defense Mechanism Against Soybean Mosaic Virus
4.2. Resistance to SMV Is Mediated by a Network of Processes and Signaling Pathways
4.3. Role of Hormones in the Defense Mechanism
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Adams, M.J.; Antoniw, J.F.; Beaudoin, F. Overview and Analysis of the Polyprotein Cleavage Sites in the Family Potyviridae. Mol. Plant Pathol. 2005, 6, 471–487. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Yang, Q.H.; Zhi, H.J.; Gai, J.Y. Identification and Distribution of Soybean Mosaic Virus Strains in Southern China. Plant Dis. 2010, 94, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Tian, Z.; Li, K.; Li, H.; Huang, Z.; Hu, G.; Zhi, H. Identification and Variation Analysis of Soybean Mosaic Virus Strains in Shandong, Henan and Anhui Provinces of China. Soybean Sci. 2013, 32, 806–809. [Google Scholar]
- Cho, E.K.; Goodman, R.M. Strains of Soybean Mosaic Virus: Classification Based on Virulence in Resistant Soybean Cultivars. Phytopathology 1979, 69, 467–470. [Google Scholar]
- Cho, E.K.; Goodman, R.M. Evaluation of Resistance in Soybeans to Soybean Mosaic Virus Strains 1. Crop Sci. 1982, 22, 1133–1136. [Google Scholar] [CrossRef]
- Takahashi, K.; Tanaka, T.; Iida, W.; Tsuda, Y. Studies on Virus Diseases and Causal Viruses of Soybean in Japan. Bull. Tohoku Natl. Agric. Exp. Stn. 1980, 62, 1–130. [Google Scholar]
- Yu, X.-Q.; Niu, H.-Q.; Liu, C.; Wang, H.-L.; Yin, W.; Xia, X. PTI-ETI Synergistic Signal Mechanisms in Plant Immunity. Plant Biotechnol. J. 2024, 22, 2113–2128. [Google Scholar] [CrossRef]
- Lin, H.; Wang, M.; Chen, Y.; Nomura, K.; Hui, S.; Gui, J.; Zhang, X.; Wu, Y.; Liu, J.; Li, Q.; et al. An MKP-MAPK Protein Phosphorylation Cascade Controls Vascular Immunity in Plants. Sci. Adv. 2022, 8, eabg8723. [Google Scholar] [CrossRef]
- Li, P.; Zhao, L.; Qi, F.; Htwe, N.M.P.S.; Li, Q.; Zhang, D.; Lin, F.; Shang-Guan, K.; Liang, Y. The Receptor-like Cytoplasmic Kinase RIPK Regulates Broad-Spectrum ROS Signaling in Multiple Layers of Plant Immune System. Mol. Plant 2021, 14, 1652–1667. [Google Scholar] [CrossRef]
- Ding, L.N.; Li, Y.T.; Wu, Y.Z.; Li, T.; Geng, R.; Cao, J.; Zhang, W.; Tan, X.L. Plant Disease Resistance-Related Signaling Pathways: Recent Progress and Future Prospects. Int. J. Mol. Sci. 2022, 23, 16200. [Google Scholar] [CrossRef]
- Liu, J.Z.; Lam, H.M. Signal Transduction Pathways in Plants for Resistance against Pathogens. Int. J. Mol. Sci. 2019, 20, 2335. [Google Scholar] [CrossRef] [PubMed]
- Nogia, P.; Pati, P.K. Plant Secondary Metabolite Transporters: Diversity, Functionality, and Their Modulation. Front. Plant Sci. 2021, 12, 758202. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Bao, J.; Lu, X.; Ma, L.; Zhao, Y.; Lan, S.; Cao, J.; Ma, S.; Li, S. The Mechanism of Ca2+ Signal Transduction in Plants Responding to Abiotic Stresses. Environ. Exp. Bot. 2023, 216, 105514. [Google Scholar] [CrossRef]
- Mazzoni-Putman, S.M.; Brumos, J.; Zhao, C.; Alonso, J.M.; Stepanova, A.N. Auxin Interactions with Other Hormones in Plant Development. Cold Spring Harb. Perspect. Biol. 2021, 13, a039990. [Google Scholar] [CrossRef]
- Gomes, G.L.B.; Scortecci, K.C. Auxin and Its Role in Plant Development: Structure, Signalling, Regulation and Response Mechanisms. Plant Biol. 2021, 23, 894–904. [Google Scholar] [CrossRef]
- Wang, D.; Ma, Y.; Liu, N.; Yang, Z.; Zheng, G.; Zhi, H. Fine Mapping and Identification of the Soybean R SC4 Resistance Candidate Gene to Soybean Mosaic Virus. Plant Breed. 2011, 130, 653–659. [Google Scholar] [CrossRef]
- Rui, R.; Liu, S.; Karthikeyan, A.; Wang, T.; Niu, H.; Yin, J.; Yang, Y.; Wang, L.; Yang, Q.; Zhi, H.; et al. Fine-Mapping and Identification of a Novel Locus Rsc15 Underlying Soybean Resistance to Soybean Mosaic Virus. Theor. Appl. Genet. 2017, 130, 2395–2410. [Google Scholar] [CrossRef]
- Zheng, G.; Yang, Y.; MA, Y.; Yang, X.; Chen, S.; Ren, R.; Wang, D.; Yang, Z.; Zhi, H. Fine Mapping and Candidate Gene Analysis of Resistance Gene R to Soybean Mosaic Virus in Qihuang 1. J. Integr. Agric. 2014, 13, 2608–2615. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor Package for Differential Expression Analysis of Digital Gene Expression Data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Kanehisa, M. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Suman, S.; Bagal, D.; Jain, D.; Singh, R.; Singh, I.K.; Singh, A. Biotic Stresses on Plants: Reactive Oxygen Species Generation and Antioxidant Mechanism. In Frontiers in Plant-Soil Interaction; Aftab, T., Hakeem, R.K., Eds.; Academic Press: London, UK, 2021; pp. 381–411. [Google Scholar]
- Sharma, I.; Ahmad, P. Catalase. In Oxidative Damage to Plants; Academic Press: London, UK, 2014; pp. 131–148. [Google Scholar]
- Padder, B.A. Plant Disease Resistance Genes: From Perception to Signal Transduction. In Plant Signaling: Understanding the Molecular Crosstalk; Hakeem, K., Rehman, R., Tahir, I., Eds.; Springer: New Delhi, India, 2014; pp. 345–354. [Google Scholar]
- Bigeard, J.; Colcombet, J.; Hirt, H. Signaling Mechanisms in Pattern-Triggered Immunity (PTI). Mol. Plant 2015, 8, 521–539. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Ngou, B.P.M.; Ding, P.; Xin, X.F. PTI-ETI Crosstalk: An Integrative View of Plant Immunity. Curr. Opin. Plant Biol. 2021, 62, 102030. [Google Scholar] [CrossRef]
- Ijaz, S.; Haq, I.U.; Babar, M.; Nasir, B. Disease Resistance Genes’ Identification, Cloning, and Characterization in Plants. In Cereal Diseases: Nanobiotechnological Approaches for Diagnosis and Management; Springer Nature: Singapore, 2022; pp. 249–269. [Google Scholar]
- Ameline Torregrosa, C.; Wang, B.B.; O’Bleness, M.S.; Deshpande, S.; Zhu, H.; Roe, B.; Young, N.D.; Cannon, S.B. Identification and Characterization of Nucleotide-Binding Site-Leucine-Rich Repeat Genes in the Model Plant Medicago Truncatula. Plant Physiol. 2008, 146, 5–21. [Google Scholar] [CrossRef]
- Bai, S.; Liu, J.; Chang, C.; Zhang, L.; Maekawa, T.; Wang, Q.; Xiao, W.; Liu, Y.; Chai, J.; Takken, F.L.W.; et al. Structure-Function Analysis of Barley NLR Immune Receptor MLA10 Reveals Its Cell Compartment Specific Activity in Cell Death and Disease Resistance. PLoS Pathog. 2012, 8, e1002752. [Google Scholar] [CrossRef]
- Cao, Y.; Mo, W.; Li, Y.; Xiong, Y.; Wang, H.; Zhang, Y.; Lin, M.; Zhang, L.; Li, X. Functional Characterization of NBS-LRR Genes Reveals an NBS-LRR Gene That Mediates Resistance against Fusarium Wilt. BMC Biol. 2024, 22, 45. [Google Scholar] [CrossRef]
- Liu, C.; Liu, Q.; Mou, Z. Redox Signaling and Oxidative Stress in Systemic Acquired Resistance. J. Exp. Bot. 2024, 75, 4535–4548. [Google Scholar] [CrossRef]
- Weralupitiya, C.; Eccersall, S.; Meisrimler, C.-N. Shared Signals, Different Fates: Calcium and ROS in Plant PRR and NLR Immunity. Cell Rep. 2024, 43, 114910. [Google Scholar] [CrossRef]
- Shafqat, A.; Abbas, S.; Ambreen, M.; Siddiqa Bhatti, A.; Kausar, H.; Gull, T. Exploring the Vital Role of Phytohormones and Plant Growth Regulators in Orchestrating Plant Immunity. Physiol. Mol. Plant Pathol. 2024, 133, 102359. [Google Scholar] [CrossRef]
- Kalsi, H.S.; Karkhanis, A.A.; Natarajan, B.; Bhide, A.J.; Banerjee, A.K. AUXIN RESPONSE FACTOR 16 (StARF16) Regulates Defense Gene StNPR1 upon Infection with Necrotrophic Pathogen in Potato. Plant Mol. Biol. 2022, 109, 13–28. [Google Scholar] [CrossRef]
- Liu, L.; Yahaya, B.S.; Li, J.; Wu, F. Enigmatic Role of Auxin Response Factors in Plant Growth and Stress Tolerance. Front. Plant Sci. 2024, 15, 1398818. [Google Scholar] [CrossRef]
- Bouzroud, S.; Gouiaa, S.; Hu, N.; Bernadac, A.; Mila, I.; Bendaou, N.; Smouni, A.; Bouzayen, M.; Zouine, M. Auxin Response Factors (ARFs) Are Potential Mediators of Auxin Action in Tomato Response to Biotic and Abiotic Stress (Solanum lycopersicum). PLoS ONE 2018, 13, e0193517. [Google Scholar] [CrossRef]
- Ghanashyam, C.; Jain, M. Role of Auxin-Responsive Genes in Biotic Stress Responses. Plant Signal. Behav. 2009, 4, 846–848. [Google Scholar] [CrossRef] [PubMed]
- Guilfoyle, T.J.; Hagen, G. Auxin Response Factors. Curr. Opin. Plant Biol. 2007, 10, 453–460. [Google Scholar] [CrossRef]
- Werghi, S.; Herrero, F.A.; Fakhfakh, H.; Gorsane, F. Auxin Drives Tomato Spotted Wilt Virus (TSWV) Resistance through Epigenetic Regulation of Auxin Response Factor ARF8 Expression in Tomato. Gene 2021, 804, 145905. [Google Scholar]
- Vaisman, M.; Hak, H.; Arazi, T.; Spiegelman, Z. The Impact of Tobamovirus Infection on Root Development Involves Induction of Auxin Response Factor 10a in Tomato. Plant Cell Physiol. 2023, 63, 1980–1993. [Google Scholar] [CrossRef]
- Schepetilnikov, M.; Ryabova, L.A. Auxin Signaling in Regulation of Plant Translation Reinitiation. Front. Plant Sci. 2017, 8, 1014. [Google Scholar] [CrossRef]
- Schepetilnikov, M.; Ryabova, L.A. Recent Discoveries on the Role of TOR (Target of Rapamycin) Signaling in Translation in Plants. Plant Physiol. 2018, 176, 1095–1105. [Google Scholar] [CrossRef]
- Niu, J.; Zhao, J.; Guo, Q.; Zhang, H.; Yue, A.; Zhao, J.; Yin, C.; Wang, M.; Du, W. WGCNA Reveals Hub Genes and Key Gene Regulatory Pathways of the Response of Soybean to Infection by Soybean Mosaic Virus. Genes 2024, 15, 566. [Google Scholar] [CrossRef]
- Zhu, H.; Li, R.; Fang, Y.; Zhao, X.; Teng, W.; Li, H.; Han, Y. Weighted Gene Co-Expression Network Analysis Uncovers Critical Genes and Pathways Involved in Soybean Response to Soybean Mosaic Virus. Agronomy 2024, 14, 2455. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raza, M.M.; Jia, H.; Gu, S.; Gai, J.; Li, K. Transcriptome Insights into Resistance Mechanisms Against Soybean Mosaic Virus Strain SC4 in Soybean. Agronomy 2025, 15, 906. https://doi.org/10.3390/agronomy15040906
Raza MM, Jia H, Gu S, Gai J, Li K. Transcriptome Insights into Resistance Mechanisms Against Soybean Mosaic Virus Strain SC4 in Soybean. Agronomy. 2025; 15(4):906. https://doi.org/10.3390/agronomy15040906
Chicago/Turabian StyleRaza, Muhammad Muzzafar, Huiying Jia, Shengyu Gu, Junyi Gai, and Kai Li. 2025. "Transcriptome Insights into Resistance Mechanisms Against Soybean Mosaic Virus Strain SC4 in Soybean" Agronomy 15, no. 4: 906. https://doi.org/10.3390/agronomy15040906
APA StyleRaza, M. M., Jia, H., Gu, S., Gai, J., & Li, K. (2025). Transcriptome Insights into Resistance Mechanisms Against Soybean Mosaic Virus Strain SC4 in Soybean. Agronomy, 15(4), 906. https://doi.org/10.3390/agronomy15040906






