Rapid Characterization of the Genetic Loci Controlling Commodity Traits of Chinese Hami Melon (Cucumis melo var. Saccharinensis Naud.) through Multiplexed Shotgun Genotyping
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials and DNA Extraction
2.2. Library Preparation and Sequencing
2.3. SNP Identification
2.4. Genotyping and Breakpoint Determination
2.5. Bin Map Construction
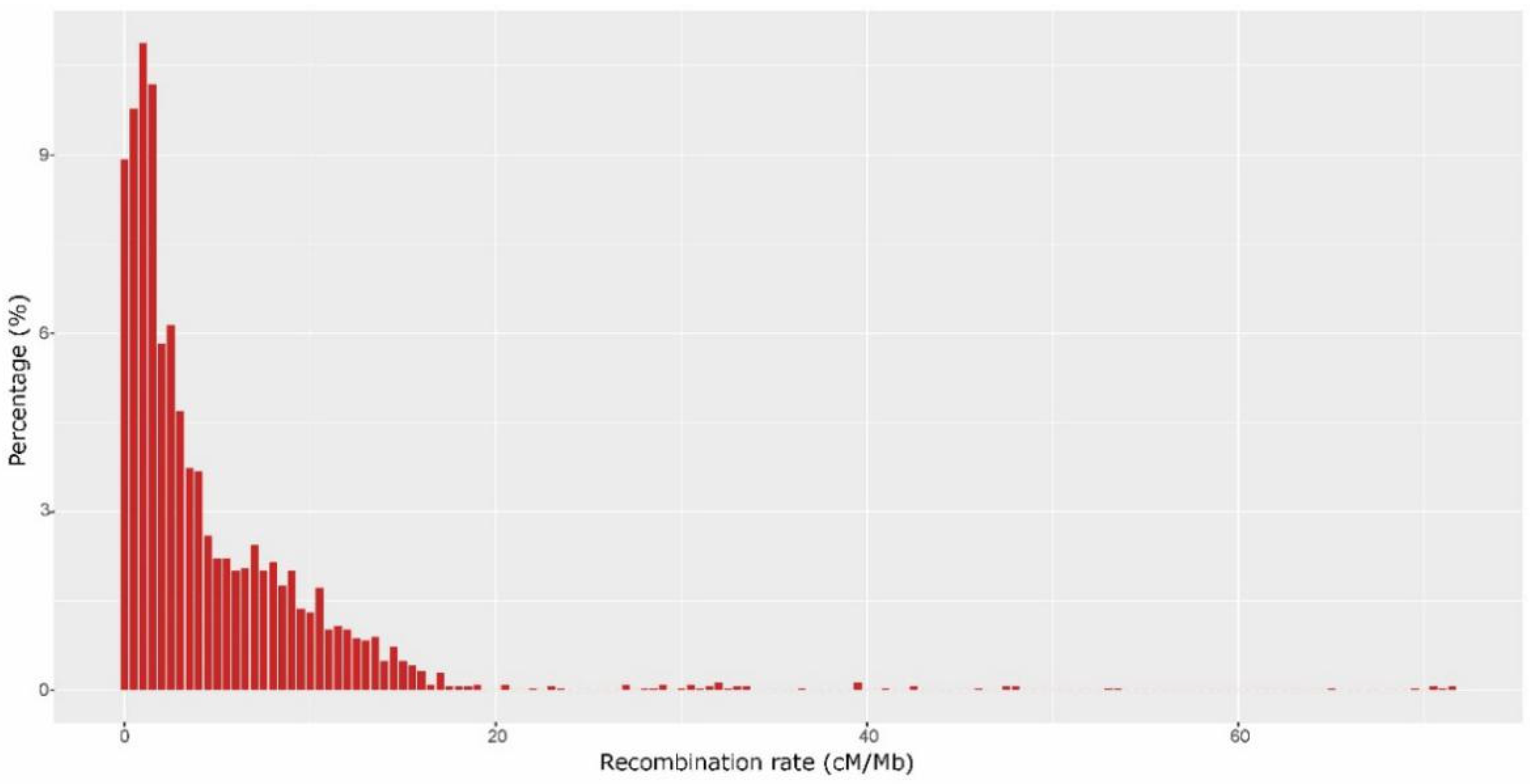
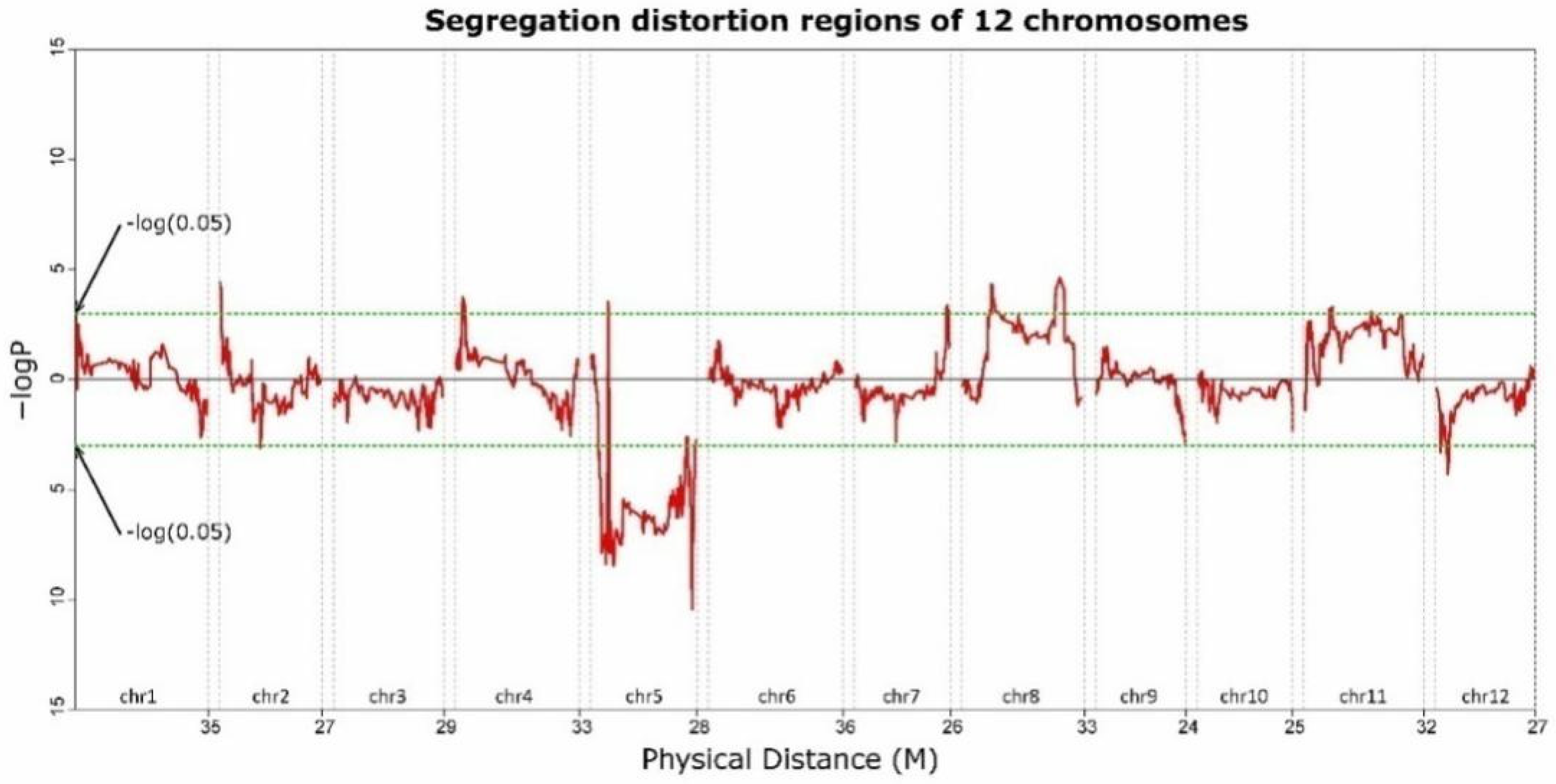
2.6. Segregation Distortion and Recombination Rate
2.7. Phenotype Assessment
2.8. QTL Mapping
3. Results
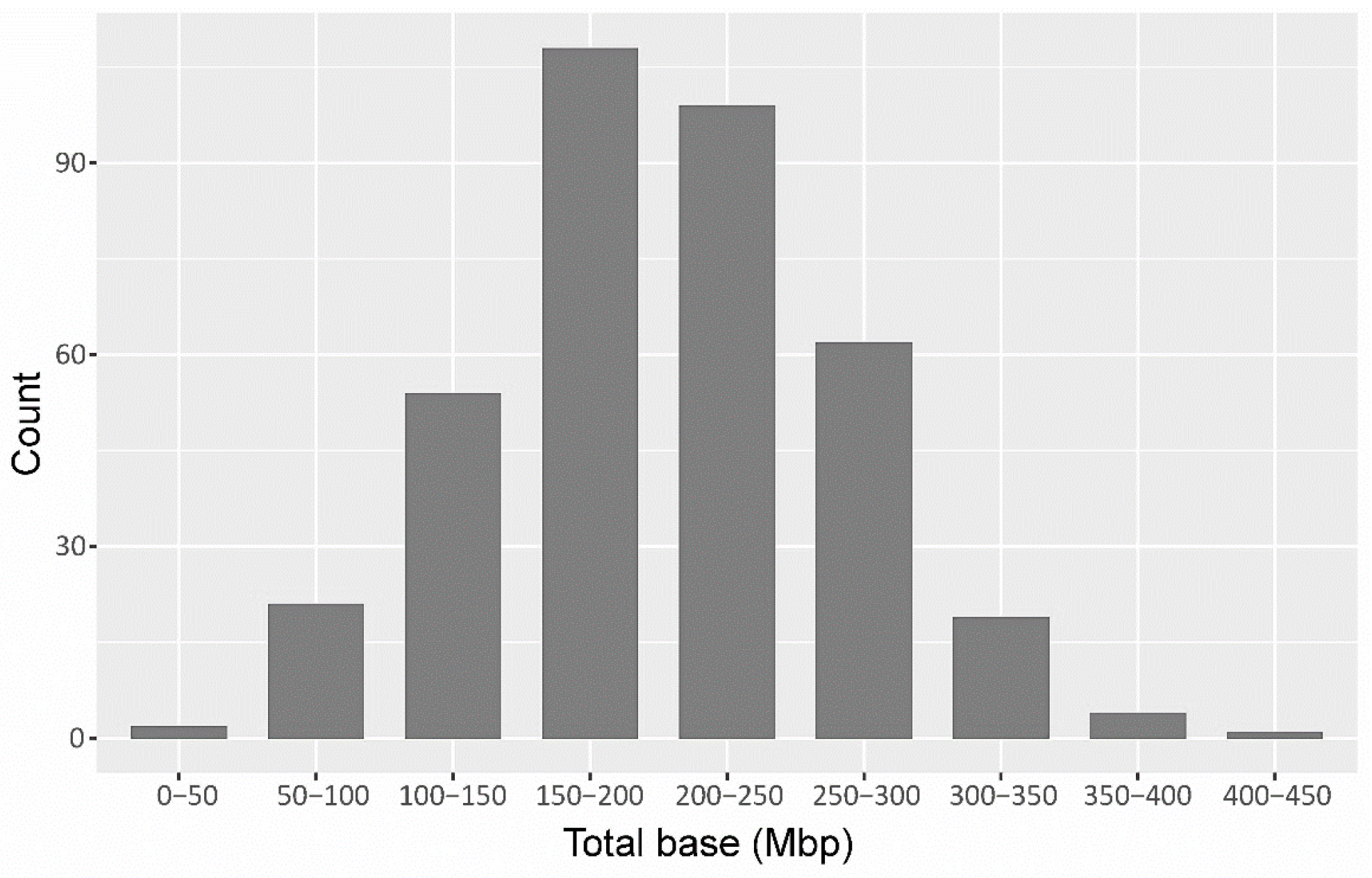
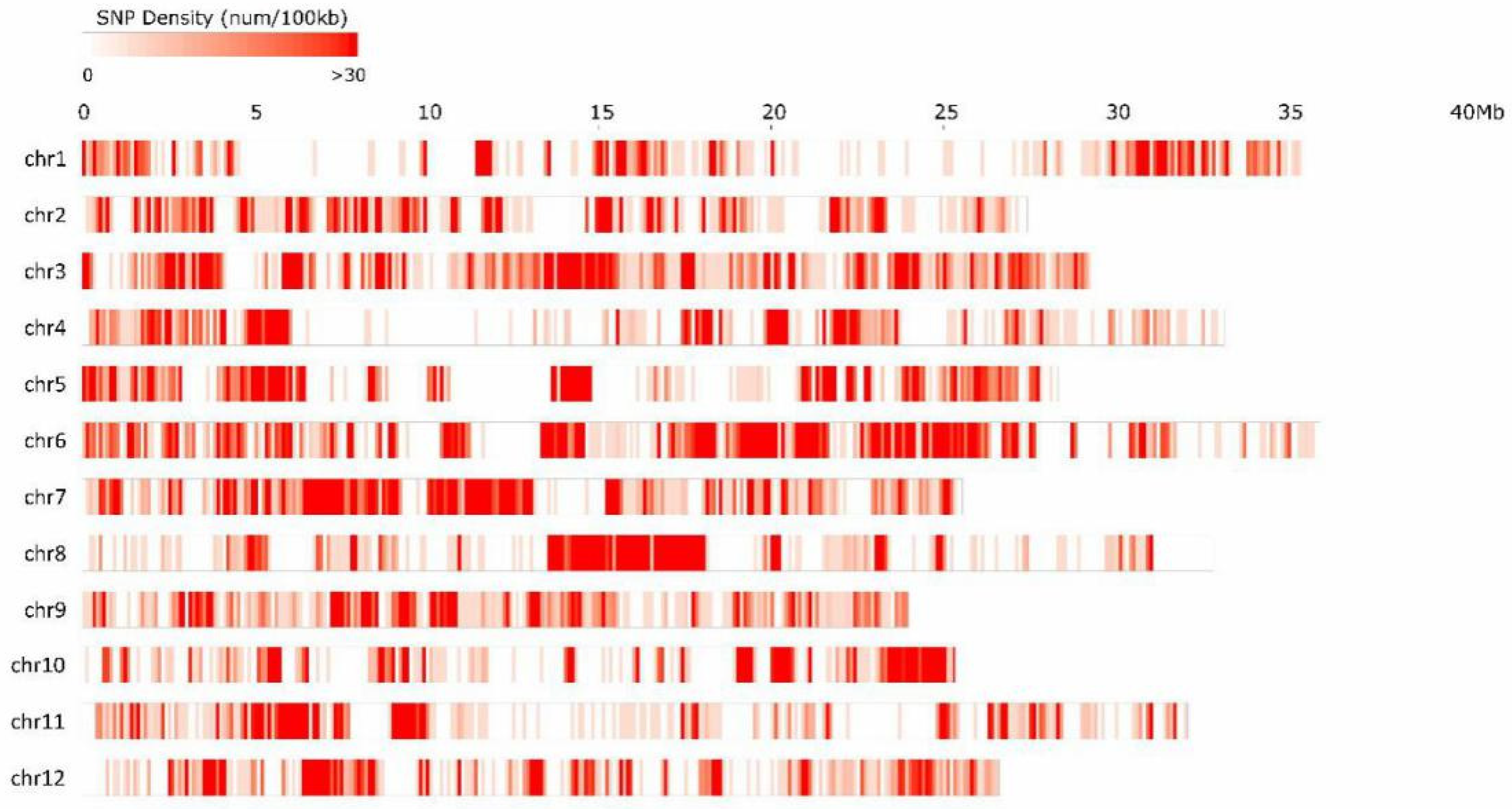
3.1. Sequencing and SNP Identification
3.2. Breakpoint Determination
3.3. Construction of Bin Maps and a High-Resolution Genetic Map
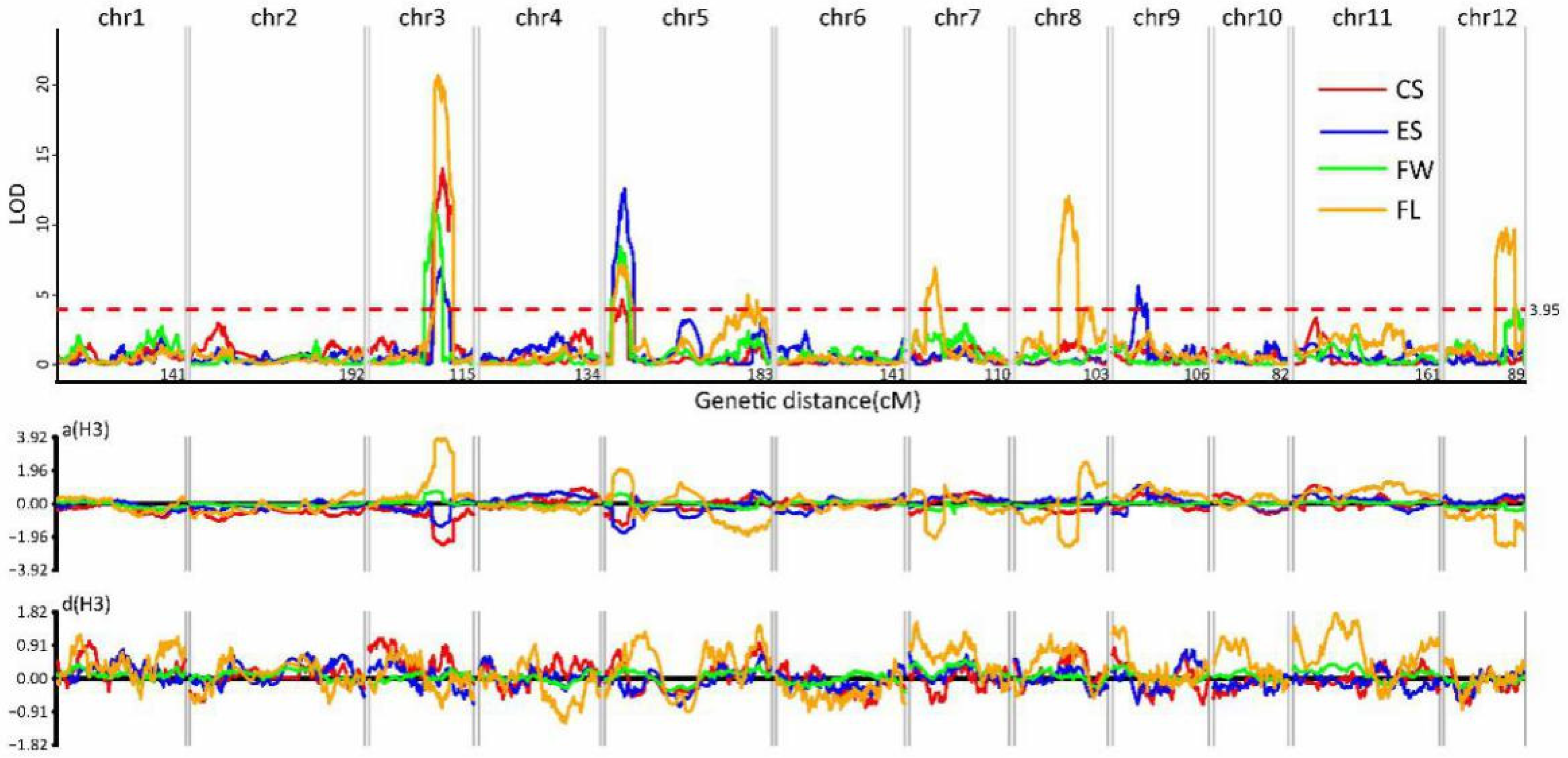
3.4. Mapping the Genetic Loci Controlling the Commodity Traits
4. Discussion
4.1. It Is Time- and Cost-Effective to Use MSG to Achieve Genetic Map Construction and QTL Mapping of an F2 Population in Melon
4.2. Chinese Hami Melon Has a Special Genetic Architecture
4.3. QTLs Identified in This Research Should Facilitate Chinese Hami Melon Breeding
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Availability of Data and Materials
Conflicts of Interest
References
- Food and agriculture Organization of the United Nations. Available online: http://www.fao.org/faostat/en/ (accessed on 4 August 2019).
- Escribano, S.; Lazaro, A. Physicochemical and nutritional evaluation of Spanish melon landraces. Plant Genet. Resour. C. 2015, 15, 177–186. [Google Scholar] [CrossRef]
- Saini, R.K.; Assefa, A.D.; Keum, Y.S. Fatty acid and carotenoid composition of bitter melon (Momordicacharantia L.) seed arils: A potentially valuable source of lycopene. J. Food Meas. Charact. 2017, 11, 1266–1273. [Google Scholar] [CrossRef]
- Garcia-Mas, J.; Benjak, A.; Sanseverino, W.; Bourgeois, M.; Mir, G.; Gonzalez, V.M.; Hénaff, E.; Câmara, F.; Cozzuto, L.; Lowy, E.; et al. The genome of melon (Cucumismelo L.). Proc. Natl. Acad. Sci. USA 2012, 109, 11872–11877. [Google Scholar] [CrossRef] [PubMed]
- Pech, J.C.; Bouzayen, M.; Latche, A. Climacteric fruit ripening: Ethylene-dependent and independent regulation of ripening pathways in melon fruit. Plant Sci. 2008, 175, 114–120. [Google Scholar] [CrossRef] [Green Version]
- Saladié, M.; Cañizares, J.; Phillips, M.A.; Rodriguez-Concepcion, M.; Larrigaudière, C.; Gibon, Y.; Stitt, M.; Lunn, J.E.; Garcia-Mas, J. Comparative transcriptional profiling analysis of developing melon (Cucumismelo L.) fruit from climacteric and non-climacteric varieties. BMC Genom. 2015, 16, 440. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.; Troadec, C.; Boualem, A.; Rajab, M.; Fernandez, R.; Morin, H.; Pitrat, M.; Dogimont, C.; Bendahmane, A. A transposon-induced epigenetic change leads to sex determination in melon. Nature 2009, 461, 1135–1138. [Google Scholar] [CrossRef] [PubMed]
- Ibdah, M.; Azulay, Y.; Portnoy, V.; Wasserman, B.; Bar, E.; Meir, A.; Burger, Y.; Hirschberg, J.; Schaffer, A.A.; Katzir, N.; et al. Functional characterization of CmCCD1, a carotenoid cleavage dioxygenase from melon. Phytochemistry 2006, 67, 1579–1589. [Google Scholar] [CrossRef] [PubMed]
- Harel-Beja, R.; Tzuri, G.; Portnoy, V.; Lotan-Pompan, M.; Lev, S.; Cohen, S.; Dai, N.; Yeselson, L.; Meir, A.; Libhaber, S.E.; et al. A genetic map of melon highly enriched with fruit quality QTLs and EST markers, including sugar and carotenoid metabolism genes. Theor. Appl. Genet. 2010, 121, 511–533. [Google Scholar] [CrossRef] [PubMed]
- Tzuri, G.; Zhou, X.; Chayut, N.; Yuan, H.; Portnoy, V.; Meir, A.; Sa’ar, U.; Baumkoler, F.; Mazourek, M.; Lewinsohn, E.; et al. A ‘golden’ SNP in CmOr governs the fruit flesh color of melon (Cucumismelo L.). Plant J. 2015, 82, 267–279. [Google Scholar] [CrossRef] [PubMed]
- Kirkbride, J.H.J. Biosystematic Monograph of the Genus Cucumis (Cucurbitaceae): Botanical Identification of Cucumbers and Melons; Parkway Publishers: Boone, NC, USA, 1993. [Google Scholar]
- Hammer, K.; Gladis, T. Notes on infraspecific nomenclature and classifications of cultivated plants in Compositae, Cruciferae, Cucurbitaceae, Gramineae (with a remark on Triticumdicoccon Schrank) and Leguminosae. Genet. Resour. Crop Evol. 2014, 61, 1455–1467. [Google Scholar] [CrossRef]
- Tanaka, K.; Akashi, Y.; Fukunaga, K.; Yamamoto, T.; Aierken, Y.; Nishida, H.; Long, C.L.; Yoshino, H.; Sato, Y.I.; Kato, K. Diversification and genetic differentiation of cultivated melon inferred from sequence polymorphism in the chloroplast genome. Breed. Sci. 2013, 63, 183–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Renner, S.S.; Schaefer, H.; Kocyan, A. Phylogenetics of Cucumis (Cucurbitaceae):Cucumber (C-sativus) belongs in an Asian/Australian clade far from melon (C-melo). BMC Evolut. Biol. 2007, 7, 58. [Google Scholar] [CrossRef] [PubMed]
- Robinson, R.W.; Decker-Walters, D.S. Cucurbits; Cab International: Wallingford, UK, 1997. [Google Scholar]
- Luan, F.; Delannay, I.; Staub, J.E. Chinese melon (Cucumismelo L.) diversity analyses provide strategies for germplasm curation, genetic improvement, and evidentiary support of domestication patterns. Euphytica 2008, 164, 445–461. [Google Scholar] [CrossRef]
- Malik, A.A.; Vashisht, V.K.; Singh, K.; Sharma, A.; Singh, D.K.; Singh, H.; Monforte, A.J.; McCreight, J.D.; Dhillon, N.P. Diversity among melon (Cucumismelo L.) landraces from the Indo-Gangetic plains of India and their genetic relationship with USA melon cultivars. Genet. Resour. Crop Evol. 2014, 61, 1189–1208. [Google Scholar] [CrossRef]
- Aierken, Y.; Akashi, Y.; Nhi, P.T.P.; Halidan, Y.; Tanaka, K.; Long, B.; Nishida, H.; Long, C.; Wu, M.Z.; Kato, K.; et al. Molecular analysis of the genetic diversity of chinese Hami melon and its relationship to the melon germplasm from central and south Asia. J. Jpn. Soc. Hortic. Sci. 2011, 80, 52–65. [Google Scholar] [CrossRef]
- RagHami, M.; Lopez-Sese, A.I.; Hasandokht, M.R.; Zamani, Z.; Moghadam, M.R.F.; Kashi, A. Genetic diversity among melon accessions from Iran and their relationships with melon germplasm of diverse origins using microsatellite markers. Plant Syst. Evol. 2014, 300, 139–151. [Google Scholar] [CrossRef]
- Zhang, Y.P.; Yang, S.J.; Chen, Y.Y. Studies on genetic relationship analysis and purity identification of hybrids of melon cultivars by SRAP markers. In Cucurbitaceae 2012. Proceedings of the Xth EUCARPIA Meeting on Genetics and Breeding of Cucurbitaceae, Antalya, Turkey, 5–18 October 2012; ZiraatFakultesi: Antalya, Turkey, 2012; pp. 459–465. [Google Scholar]
- Rodriguez-Moreno, L.; Gonzalez, V.M.; Benjak, A.; Marti, M.C.; Puigdomenech, P.; Aranda, M.A.; Garcia-Mas, J. Determination of the melon chloroplast and mitochondrial genome sequences reveals that the largest reported mitochondrial genome in plants contains a significant amount of DNA having a nuclear origin. BMC Genom. 2011, 12, 424. [Google Scholar] [CrossRef]
- Chang, C.W.; Wang, Y.H.; Tung, C.W. Genome-wide single nucleotide polymorphism discovery and the construction of a high-density genetic map for melon (Cucumismelo L.) using genotyping-by-sequencing. Front.Plant Sci. 2017, 8, 125. [Google Scholar] [CrossRef]
- Baudracco-Arnas, S.; Pitrat, M. A genetic map of melon (Cucumismelo L.) with RFLP, RAPD, isozyme, disease resistance and morphological markers. Theor. Appl. Genet. 1996, 93, 57–64. [Google Scholar] [CrossRef]
- Oliver, M.; Garcia-Mas, J.; Cardus, M.; Pueyo, N.; López-Sesé, A.I.; Arroyo, M.; Gomez-Paniagua, H.; Arus, P.; Vicente, M.D. Construction of a reference linkage map for melon. Genome 2001, 44, 836–845. [Google Scholar] [CrossRef]
- Perin, C.; Hagen, L.S.; Giovinazzo, N.; Besombes, D.; Dogimont, C.; Pitrat, M. Genetic control of fruit shape acts prior to anthesis in melon (Cucumismelo L.). Mol. Genet. Genom. 2002, 266, 933–941. [Google Scholar]
- Cuevas, H.E.; Staub, J.E.; Simon, P.W.; Zalapa, J.E. A consensus linkage map identifies genomic regions controlling fruit maturity and beta-carotene-associated flesh color in melon (Cucumismelo L.). Theoret. Appl. Genet. 2009, 119, 741–756. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, H.E.; Staub, J.E.; Simon, P.W.; Zalapa, J.E.; McCreight, J.D. Mapping of genetic loci that regulate quantity of beta-carotene in fruit of US Western Shipping melon (Cucumismelo L.). Theor. Appl. Genet. 2008, 117, 1345–1359. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Silva, I.; Eduardo, I.; Blanca, J.; Esteras, C.; Pico, B.; Nuez, F.; Arus, P.; Garcia-Mas, J.; Monforte, A.J. Bin mapping of genomic and EST-derived SSRs in melon (Cucumismelo L.). Theor. Appl. Genet. 2008, 118, 139–150. [Google Scholar] [CrossRef] [PubMed]
- Deleu, W.; Esteras, C.; Roig, C.; González-To, M.; Fernández-Silva, I.; Gonzalez-Ibeas, D.; Blanca, J.; Aranda, M.A.; Arús, P.; Nuez, F.; et al. A set of EST-SNPs for map saturation and cultivar identification in melon. BMC Plant Biol. 2009, 9, 90. [Google Scholar] [CrossRef] [PubMed]
- Boissot, N.; Thomas, S.; Sauvion, N.; Marchal, C.; Pavis, C.; Dogimont, C. Mapping and validation of QTLs for resistance to aphids and whiteflies in melon. Theor. Appl. Genet. 2010, 121, 9–20. [Google Scholar] [CrossRef]
- Lu, F.; Xu, Y.; Zhao, Y.; Cao, D.; Feng, J.; Guo, S.; Gong, G.; Yi, H.; Wu, M.; Zhang, H. Construction of permanent genetic mapand comparative analysis of Xinjiang Hami melon (Cucumismelo L. ssp.elo. convar. ameri (Pang.) Greb). Acta Hortic. 2009, 36, 1767–1774. [Google Scholar]
- Perin, C.; Hagen, L.; De Conto, V.; Katzir, N.; Danin-Poleg, Y.; Portnoy, V.; Baudracco-Arnas, S.; Chadoeuf, J.; Dogimont, C.; Pitrat, M. A reference map of Cucumismelo based on two recombinant inbred line populations. Theor. Appl. Genet. 2002, 104, 1017–1034. [Google Scholar] [CrossRef]
- Diaz, A.; Fergany, M.; Formisano, G.; Ziarsolo, P.; Blanca, J.; Fei, Z.; Staub, J.E.; Zalapa, J.E.; Cuevas, H.E.; Dace, G.; et al. A consensus linkage map for molecular markers and quantitative trait loci associated with economically important traits in melon (Cucumismelo L.). BMC Plant Biol. 2011, 11, 111. [Google Scholar] [CrossRef]
- Stange, M.; Utz, H.F.; Schrag, T.A.; Melchinger, A.E.; Wurschum, T. High-density genotyping: An overkill for QTL mapping Lessons learned from a case study in maize and simulations. Theor. Appl. Genet. 2013, 126, 2563–2574. [Google Scholar] [CrossRef]
- Almannai, M.; Marom, R.; Sutton, V.R. Newborn screening: A review of history, recent advancements, and future perspectives in the era of next generation sequencing. Curr. Opin. Pediatr. 2016, 28, 694–699. [Google Scholar] [CrossRef] [PubMed]
- Andolfatto, P.; Davison, D.; Erezyilmaz, D.; Hu, T.T.; Mast, J.; Sunayama-Morita, T.; Stern, D.L. Multiplexed shotgun genotyping for rapid and efficient genetic mapping. Genome Res. 2011, 21, 610–617. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baird, N.A.; Etter, P.D.; Atwood, T.S.; Currey, M.C.; Shiver, A.L.; Lewis, Z.A.; Selker, E.U.; Cresko, W.A.; Johnson, E.A. Rapid SNP discovery and genetic mapping using sequencedRAD markers. PLoS ONE 2008, 3, e3376. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Feng, Q.; Qian, Q.; Zhao, Q.; Wang, L.; Wang, A.; Guan, J.; Fan, D.; Weng, Q.; Huang, T.; et al. High-throughput genotyping bywhole-genome resequencing. Genome Res. 2009, 19, 1068–1076. [Google Scholar] [CrossRef] [PubMed]
- Porebski, S.; Bailey, L.G.; Baum, B.R. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep. 1997, 15, 8–15. [Google Scholar] [CrossRef]
- Li, R.; Yu, C.; Li, Y.; Lam, T.W.; Yiu, S.M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Li, Y.; Fang, X.; Yang, H.; Wang, J.; Kristiansen, K.; Wang, J. SNP detection for massively parallel whole-genome resequencing. Genome Res. 2009, 19, 1124–1132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, X.; Liu, X.; Ge, S.; Jensen, J.D.; Hu, F.; Li, X.; Dong, Y.; Gutenkunst, R.N.; Fang, L.; Huang, L.; et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 2012, 30, 105–111. [Google Scholar] [CrossRef]
- Duan, M.; Sun, Z.; Shu, L.; Tan, Y.; Yu, D.; Sun, X.; Liu, R.; Li, Y.; Gong, S.; Yuan, D. Genetic analysis of an elite super-hybrid rice parent using high-density SNP markers. Rice 2013, 6, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Van Os, H.; Andrzejewski, S.; Bakker, E.; Barrena, I.; Bryan, G.J.; Caromel, B.; Ghareeb, B.; Isidore, E.; De Jong, W.; Van Koert, P.; et al. Construction of a 10,000-marker ultradense genetic recombination map of potato:Providing a framework for accelerated gene isolation and a genomewide physical map. Genetics 2006, 173, 1075–1087. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.H.; Bhat, P.R.; Close, T.J.; Lonardi, S. Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet. 2008, 4, e1000212. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Basten, C.J.; Zeng, Z.B. Windows QTL Cartographer2.5 Department of Statistics. North Carolina StateUniversity: Raleigh, 2006. Available online: http://statgen.ncsu.edu/qtlcart/WQTLCart.htm (accessed on 1 August 2012).
- Zeng, Z.B. Theoretical basis for separation of multiple linked gene effects in mapping quantitative trait loci. Proc. Natl. Acad. Sci. USA 1993, 90, 10972–10976. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.B. Precision mapping of quantitative trait loci. Genetics 1994, 136, 1457–1468. [Google Scholar] [PubMed]
- Fazio, G.; Staub, J.E.; Stevens, M.R. Genetic mapping and QTL analysis of horticultural traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Tagtheor. Appl. Genet. 2003, 107, 864–874. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Zhao, H.; Kou, Q.; Jiang, J.; Guo, S.; Zhang, H.; Hou, W.; Zou, X.; Sun, H.; Gong, G.; et al. A high resolution genetic map anchoring scaffolds of the sequenced watermelon genome. PLoS ONE 2012, 7, e29453. [Google Scholar] [CrossRef]
- Pan, Q.C. Ali, F.; Yang, X.H.; Li, J.S.; Yan, J.B. Exploring the genetic characteristics of two recombinant inbred line populations via high-density SNP markers in maize. PLoS ONE 2012, 7, e52777. [Google Scholar] [CrossRef]
- Wang, Y.H.; Thomas, C.E.; Dean, R.A. A genetic map of melon (Cucumismelo L.) based on amplified fragment length polymorphism (AFLP) markers. Theor. Appl. Genet. 1997, 95, 791–798. [Google Scholar] [CrossRef]
- Ramamurthy, R.K.; Waters, B.M. Identification of fruit quality and morphology QTLs in melon (Cucumismelo L.) using a population derived from flexuosus and cantalupensis botanical groups. Euphytica 2015, 204, 163–177. [Google Scholar] [CrossRef]

| Total Base (bp) | Mapped Base (bp) | Alignment Rate (%) | Coverage (%) | Depth (X) | ||
|---|---|---|---|---|---|---|
| K1-7 | 535,301,928 | 368,700,864 | 68.88 | 8.99 | 9.75 | |
| K3-92 | 373,295,832 | 217,478,520 | 58.26 | 7.63 | 6.82 | |
| F2 population | Average | 202,861,119.2 | 136,117,273.5 | 66.52 | 5.86 | 4.91 |
| Max | 443,498,832 | 277,653,180 | 76.82 | 8.20 | 11.78 | |
| Min | 49,756,056 | 30,243,360 | 49.01 | 2.59 | 2.31 | |
| Chromosome | Physical Distance (bp) | Genetic Distance (cM) | #SNPs | Kb/SNP | Recombination Sites | MB/Recombination Sites | Recombination Rate (cM/Mb) |
|---|---|---|---|---|---|---|---|
| chr01 | 35,476,783 | 141.683 | 2930 | 12.10812 | 253 | 0.14 | 3.99 |
| chr02 | 27,452,605 | 197.253 | 3797 | 7.230078 | 213 | 0.128 | 6.57 |
| chr03 | 29,387,469 | 115.126 | 5199 | 5.652523 | 274 | 0.107 | 3.92 |
| chr04 | 33,172,220 | 134.606 | 3206 | 10.34692 | 224 | 0.147 | 4.06 |
| chr05 | 28,337,775 | 183.362 | 3759 | 7.538647 | 260 | 0.109 | 6.47 |
| chr06 | 35,939,859 | 141.273 | 6715 | 5.352176 | 321 | 0.112 | 3.93 |
| chr07 | 25,560,174 | 110.021 | 5366 | 4.763357 | 226 | 0.113 | 4.3 |
| chr08 | 32,847,821 | 103.359 | 3753 | 8.752417 | 178 | 0.184 | 3.14 |
| chr09 | 23,991,886 | 105.985 | 3307 | 7.254879 | 216 | 0.111 | 4.17 |
| chr10 | 25,362,315 | 82.488 | 3087 | 8.215845 | 143 | 0.176 | 3.25 |
| chr11 | 32,125,309 | 168.343 | 3246 | 9.896891 | 235 | 0.136 | 5.24 |
| chr12 | 26,635,610 | 89.455 | 3244 | 8.210731 | 186 | 0.142 | 3.35 |
| Entire genome | 356,289,826 | 1572.954 | 47,609 | 7.483665 | 2729 | 0.13 | 4.41 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, S.; Ni, X.; Xia, Q.; Li, Y.; Dong, X.; Hou, J.; Li, Z.; Cheng, S.; Cao, D.; Zhang, Z.; et al. Rapid Characterization of the Genetic Loci Controlling Commodity Traits of Chinese Hami Melon (Cucumis melo var. Saccharinensis Naud.) through Multiplexed Shotgun Genotyping. Agronomy 2019, 9, 430. https://doi.org/10.3390/agronomy9080430
Li S, Ni X, Xia Q, Li Y, Dong X, Hou J, Li Z, Cheng S, Cao D, Zhang Z, et al. Rapid Characterization of the Genetic Loci Controlling Commodity Traits of Chinese Hami Melon (Cucumis melo var. Saccharinensis Naud.) through Multiplexed Shotgun Genotyping. Agronomy. 2019; 9(8):430. https://doi.org/10.3390/agronomy9080430
Chicago/Turabian StyleLi, Shiming, Xuemei Ni, Qiuju Xia, Yunfei Li, Xiao Dong, Junliang Hou, Zehua Li, Shu Cheng, Dong Cao, Zhenyu Zhang, and et al. 2019. "Rapid Characterization of the Genetic Loci Controlling Commodity Traits of Chinese Hami Melon (Cucumis melo var. Saccharinensis Naud.) through Multiplexed Shotgun Genotyping" Agronomy 9, no. 8: 430. https://doi.org/10.3390/agronomy9080430




