Utility of Bulk T-Cell Receptor Repertoire Sequencing Analysis in Understanding Immune Responses to COVID-19
Abstract
:1. Introduction
2. Overview of T-Cells in COVID-19
2.1. Immunological Memory and Longevity
2.2. T-Cell Cross-Reactivity
2.3. Pathogenic Effects of T Cells
2.4. T-Cell Tests in Current Clinical Use
3. The TCR Repertoire
3.1. Sample Cohort Building for TCR Repertoire Analysis
3.2. Laboratory Methods in ‘Bulk’ TCR Repertoire Sequencing
3.2.1. Substrate for Repertoire Sequencing
3.2.2. Methodological Considerations
3.3. Analysis of “Bulk” TCR Repertoire Sequencing
3.3.1. Principles of Analysis of CDR3 Sequences
3.3.2. Clonotypic Analysis
3.3.3. Diversity Profiling and Related Analyses
| Analytical Approach | Principles/Interpretation |
|---|---|
| CDR3 length profiles [59] |
|
| VDJ usage |
|
| Clonal abundance |
|
| Clonal frequency |
|
| Richness [62] |
|
| D50 diversity [64] |
|
| Simpson diversity [65,66,67] |
|
| Shannon diversity [61,67,69,70,71] |
|
| Hill’s diversity (Hill’s evenness) [73] |
|
| Pielou’s evenness index [64] |
|
| Parametric methods [74] |
|
3.3.4. Analyses Based on Sequence or Motif Identification
 indicates that the TCR clustering method uses the feature to define a cluster.
indicates that the TCR clustering method uses the feature to define a cluster.
 indicates that the TCR clustering method uses the feature to define a cluster.
indicates that the TCR clustering method uses the feature to define a cluster.| Method | Features | |||||||
|---|---|---|---|---|---|---|---|---|
| V(D)J Alignment | CDR3s | Short Motifs | Physio-Chemical Properties | Amino Acids | Nucleotides | Frequency | Enrichment | |
| GIANA [82] |  (V Only) (V Only) |  |  |  | ||||
| ALICE [83] |  |  |  |  | ||||
| clusTCR [84] |  |  | ||||||
| GLIPH2 [85] |  |  |  |  |  | |||
| iSMART [86] |  |  |  |  |  | |||
| TCRdist [87] |  (CDR1 and 2) (CDR1 and 2) |  |  | |||||
| TCRNET [83] |  (V and J) (V and J) |  |  |  (control samples required) (control samples required) | ||||
| ImmunoMap [88] |  |  |  | |||||
| MiXCR [51] |  (V and J) (V and J) |  |  | |||||
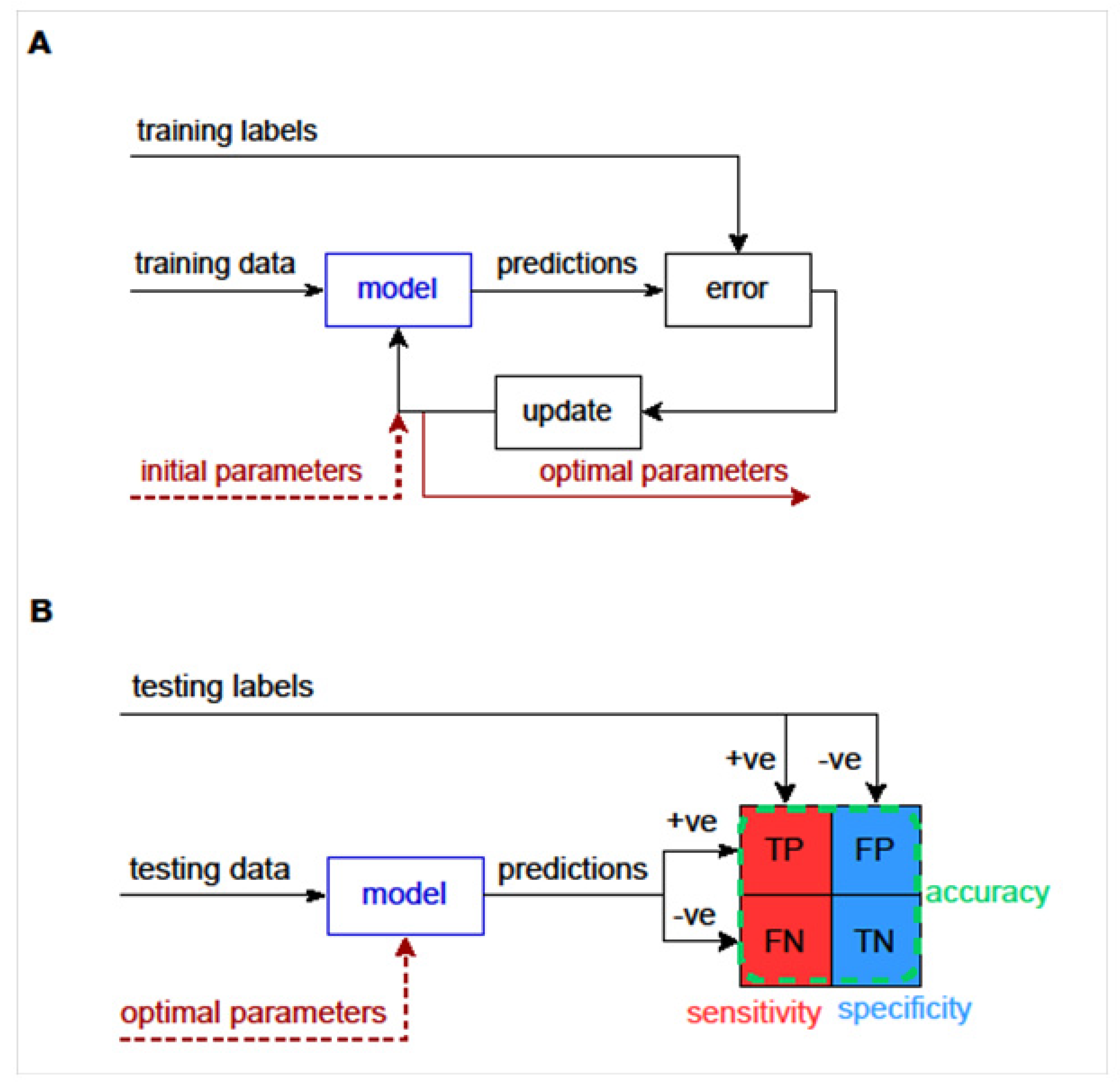
3.3.5. Machine Learning to Predict Diagnosis, Exposure to Infection or Outcome of Infection
3.3.6. Machine Learning to Identify New Antigen-Specific Sequences
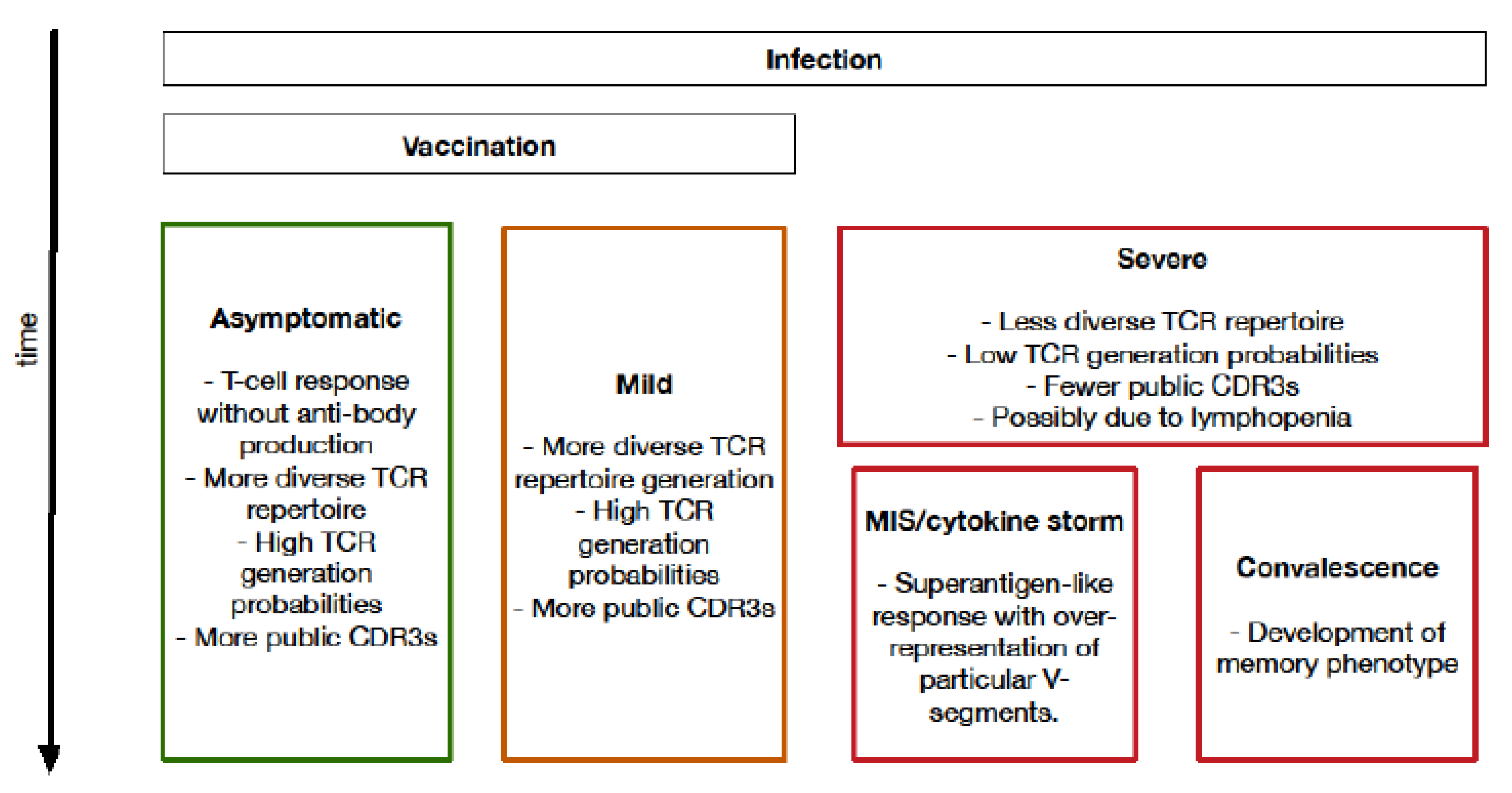
4. Biological Insights into COVID-19 from T-Cell Receptor Analysis
4.1. Relative Contributions of T Cells and B Cells to SARS-CoV-2 Immunity
4.2. Association between Higher Repertoire Diversity and Improved Outcomes
4.3. Kinetics of CD4 and CD8 T-Cell Responses
4.4. Importance of Specific V-, D- and J-Segment Usage
4.5. Importance of SARS-CoV-2 Specific TCR Sequences and Motifs
| First Author | Number of Samples | Cells | DNA/RNA | Loci | Key Points |
|---|---|---|---|---|---|
| Chang [97] |
| PBMCs | RNA | TRB |
|
| Cheng [5] |
| PBMCs | DNA | TRB |
|
| Minervina [99] |
| CD4+ and CD8+ T cells | RNA | TRB TRA |
|
| Niu [96] |
| PBMCs | RNA | TRB TRA TRG TRD |
|
| Rajeh [89] |
| PBMCs | RNA | TRB TRA |
|
| Schultheiss [95] |
| PBMCs | DNA | TRB |
|
| Shomuradova [94] |
| CD4+ and CD8+ T cells | RNA | TRB TRA |
|
| Shoukat [14] |
| PBMCs | DNA | TRB |
|
| Sidhom [92] |
| PBMCs | DNA | TRB |
|
| Swanson [99] |
| PBMCs | DNA | TRB |
|
| Wang [100] |
| CD4+ and CD8+ T cells | RNA | TRB TRA |
|
| Hu [101] |
| PBMCs | RNA | TRB |
|
| Simnica [102] |
| PBMCs Brain-derived T cells; | DNA | TRB |
|
| Shimizu [103] |
| CD8+ T cells | DNA | TRB TRA |
|
| Li [104] |
| PBMCs | RNA | TRB |
|
4.6. Vaccine-Induced T-Cell Responses in Comparison to Responses to Native Infection
5. Areas for Further Research
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Goldstein, J.R.; Lee, R.D. Demographic Perspectives on the Mortality of COVID-19 and Other Epidemics. Proc. Natl. Acad. Sci. USA 2020, 117, 22035–22041. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Wherry, E.J. T Cell Responses in Patients with COVID-19. Nat. Rev. Immunol. 2020, 20, 529–536. [Google Scholar] [CrossRef] [PubMed]
- DiPiazza, A.T.; Graham, B.S.; Ruckwardt, T.J. T Cell Immunity to SARS-CoV-2 Following Natural Infection and Vaccination. Biochem. Biophys. Res. Commun. 2021, 538, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Schub, D.; Klemis, V.; Schneitler, S.; Mihm, J.; Lepper, P.M.; Wilkens, H.; Bals, R.; Eichler, H.; Gartner, B.C.; Becker, S.L.; et al. High Levels of SARS-CoV-2-Specific T Cells with Restricted Functionality in Severe Courses of COVID-19. JCI Insight 2020, 5, e142167. [Google Scholar] [CrossRef] [PubMed]
- Cheng, M.H.; Zhang, S.; Porritt, R.A.; Noval Rivas, M.; Paschold, L.; Willscher, E.; Binder, M.; Arditi, M.; Bahar, I. Superantigenic Character of an Insert Unique to SARS-CoV-2 Spike Supported by Skewed TCR Repertoire in Patients with Hyperinflammation. Proc. Natl. Acad. Sci. USA 2020, 117, 25254–25262. [Google Scholar] [CrossRef] [PubMed]
- Le Bert, N.; Tan, A.T.; Kunasegaran, K.; Tham, C.Y.L.; Hafezi, M.; Chia, A.; Chng, M.H.Y.; Lin, M.; Tan, N.; Linster, M.; et al. SARS-CoV-2-Specific T Cell Immunity in Cases of COVID-19 and SARS, and Uninfected Controls. Nature 2020, 584, 457–462. [Google Scholar] [CrossRef]
- Dan, J.M.; Mateus, J.; Kato, Y.; Hastie, K.M.; Yu, E.D.; Faliti, C.E.; Grifoni, A.; Ramirez, S.I.; Haupt, S.; Frazier, A.; et al. Immunological Memory to SARS-CoV-2 Assessed for up to 8 Months after Infection. Science 2021, 371, eabf4063. [Google Scholar] [CrossRef]
- Sekine, T.; Perez-Potti, A.; Rivera-Ballesteros, O.; Stralin, K.; Gorin, J.B.; Olsson, A.; Llewellyn-Lacey, S.; Kamal, H.; Bogdanovic, G.; Muschiol, S.; et al. Robust T Cell Immunity in Convalescent Individuals with Asymptomatic or Mild COVID-19. Cell 2020, 183, 158–168.e14. [Google Scholar] [CrossRef]
- Redd, A.D.; Nardin, A.; Kared, H.; Bloch, E.M.; Pekosz, A.; Laeyendecker, O.; Abel, B.; Fehlings, M.; Quinn, T.C.; Tobian, A.A.R. CD8+ T-Cell Responses in COVID-19 Convalescent Individuals Target Conserved Epitopes from Multiple Prominent SARS-CoV-2 Circulating Variants. Open Forum Infect. Dis. 2021, 8, ofab143. [Google Scholar] [CrossRef]
- Tarke, A.; Sidney, J.; Methot, N.; Yu, E.D.; Zhang, Y.; Dan, J.M.; Goodwin, B.; Rubiro, P.; Sutherland, A.; Wang, E.; et al. Impact of SARS-CoV-2 Variants on the Total CD4(+) and CD8(+) T Cell Reactivity in Infected or Vaccinated Individuals. Cell Rep. Med. 2021, 2, 100355. [Google Scholar] [CrossRef]
- Jung, M.K.; Shin, E.C. Phenotypes and Functions of SARS-CoV-2-Reactive T Cells. Mol. Cells 2021, 44, 401–407. [Google Scholar] [CrossRef] [PubMed]
- Lipsitch, M.; Grad, Y.H.; Sette, A.; Crotty, S. Cross-Reactive Memory T Cells and Herd Immunity to SARS-CoV-2. Nat. Rev. Immunol. 2020, 20, 709–713. [Google Scholar] [CrossRef] [PubMed]
- Sheridan, C. COVID-19 Testing Turns to T Cells. Nat. Biotechnol. 2021, 39, 533–534. [Google Scholar] [CrossRef]
- Shoukat, M.S.; Foers, A.D.; Woodmansey, S.; Evans, S.C.; Fowler, A.; Soilleux, E.J. Use of Machine Learning to Identify a T Cell Response to SARS-CoV-2. Cell Rep. Med. 2021, 2, 100192. [Google Scholar] [CrossRef] [PubMed]
- Delves, P.J.; Roitt, I.M. The Immune System. First of two parts. N. Engl. J. Med. 2000, 343, 37–49. [Google Scholar] [CrossRef]
- Delves, P.J.; Roitt, I.M. The Immune System. Second of two parts. N. Engl. J. Med. 2000, 343, 108–117. [Google Scholar] [CrossRef]
- Foers, A.D.; Shoukat, M.S.; Welsh, O.E.; Donovan, K.; Petry, R.; Evans, S.C.; FitzPatrick, M.E.; Collins, N.; Klenerman, P.; Fowler, A.; et al. Classification of Intestinal T-Cell Receptor Repertoires Using Machine Learning Methods Can Identify Patients with Coeliac Disease Regardless of Dietary Gluten Status. J. Pathol. 2021, 253, 279–291. [Google Scholar] [CrossRef]
- Gutierrez, L.; Beckford, J.; Alachkar, H. Deciphering the TCR Repertoire to Solve the COVID-19 Mystery. Trends Pharmacol. Sci. 2020, 41, 518–530. [Google Scholar] [CrossRef]
- Mamedov, I.Z.; Britanova, O.V.; Zvyagin, I.V.; Turchaninova, M.A.; Bolotin, D.A.; Putintseva, E.V.; Lebedev, Y.B.; Chudakov, D.M. Preparing Unbiased T-Cell Receptor and Antibody CDNA Libraries for the Deep next Generation Sequencing Profiling. Front. Immunol. 2013, 4, 456. [Google Scholar] [CrossRef] [Green Version]
- Zeng, G.; Huang, Y.; Huang, Y.; Lyu, Z.; Lesniak, D.; Randhawa, P. Antigen-Specificity of T Cell Infiltrates in Biopsies with T Cell–Mediated Rejection and BK Polyomavirus Viremia: Analysis by Next Generation Sequencing. Am. J. Transplant. 2016, 16, 3131–3138. [Google Scholar] [CrossRef] [Green Version]
- Howie, B.; Sherwood, A.M.; Berkebile, A.D.; Berka, J.; Emerson, R.O.; Williamson, D.W.; Kirsch, I.; Vignali, M.; Rieder, M.J.; Carlson, C.S.; et al. High-Throughput Pairing of T Cell Receptor α and β Sequences. Sci. Transl. Med. 2015, 7, 301ra131. [Google Scholar] [CrossRef] [PubMed]
- Sims, J.S.; Grinshpun, B.; Feng, Y.; Ung, T.H.; Neira, J.A.; Samanamud, J.L.; Canoll, P.; Shen, Y.; Sims, P.A.; Bruce, J.N. Diversity and Divergence of the Glioma-Infiltrating T-Cell Receptor Repertoire. Proc. Natl. Acad. Sci. USA 2016, 113, E3529–E3537. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rossetti, M.; Spreafico, R.; Consolaro, A.; Leong, J.Y.; Chua, C.; Massa, M.; Saidin, S.; Magni-Manzoni, S.; Arkachaisri, T.; Wallace, C.A.; et al. TCR Repertoire Sequencing Identifies Synovial Treg Cell Clonotypes in the Bloodstream during Active Inflammation in Human Arthritis. Ann. Rheum. Dis. 2017, 76, 435–441. [Google Scholar] [CrossRef] [PubMed]
- Dziubianau, M.; Hecht, J.; Kuchenbecker, L.; Sattler, A.; Stervbo, U.; Rödelsperger, C.; Nickel, P.; Neumann, A.U.; Robinson, P.N.; Mundlos, S.; et al. TCR Repertoire Analysis by Next Generation Sequencing Allows Complex Differential Diagnosis of T Cell–Related Pathology. Am. J. Transplant. 2013, 13, 2842–2854. [Google Scholar] [CrossRef] [PubMed]
- Okino, S.T.; Kong, M.; Sarras, H.; Wang, Y. Evaluation of Bias Associated with High-Multiplex, Target-Specific Pre-Amplification. Biomol. Detect. Quantif. 2016, 6, 13–21. [Google Scholar] [CrossRef] [Green Version]
- Ndifon, W.; Gal, H.; Shifrut, E.; Aharoni, R.; Yissachar, N.; Waysbort, N.; Reich-Zeliger, S.; Arnon, R.; Friedman, N. Chromatin Conformation Governs T-Cell Receptor Jβ Gene Segment Usage. Proc. Natl. Acad. Sci. USA 2012, 109, 15865–15870. [Google Scholar] [CrossRef] [Green Version]
- Madi, A.; Shifrut, E.; Reich-Zeliger, S.; Gal, H.; Best, K.; Ndifon, W.; Chain, B.; Cohen, I.R.; Friedman, N. T-Cell Receptor Repertoires Share a Restricted Set of Public and Abundant CDR3 Sequences That Are Associated with Self-Related Immunity. Genome Res. 2014, 24, 1603–1612. [Google Scholar] [CrossRef] [Green Version]
- Akatsuka, Y.; Martin, E.G.; Madonik, A.; Barsoukov, A.A.; Hansen, J.A. Rapid Screening of T-Cell Receptor (TCR) Variable Gene Usage by Multiplex PCR: Application for Assessment of Clonal Composition. Tissue Antigens 1999, 53, 122–134. [Google Scholar] [CrossRef]
- Hettinger, K.; Fischer, S.; Panzer, S.; Panzer-Grümayer, E.R. Multiplex PCR for TCR Delta Rearrangements: A Rapid and Specific Approach for the Detection and Identification of Immature and Mature Rearrangements in ALL. Br. J. Haematol. 1998, 102, 1050–1054. [Google Scholar] [CrossRef] [Green Version]
- Shugay, M.; Britanova, O.V.; Merzlyak, E.M.; Turchaninova, M.A.; Mamedov, I.Z.; Tuganbaev, T.R.; Bolotin, D.A.; Staroverov, D.B.; Putintseva, E.V.; Plevova, K.; et al. Towards Error-Free Profiling of Immune Repertoires. Nat. Methods 2014, 11, 653–655. [Google Scholar] [CrossRef]
- Peng, Q.; Vijaya Satya, R.; Lewis, M.; Randad, P.; Wang, Y. Reducing Amplification Artifacts in High Multiplex Amplicon Sequencing by Using Molecular Barcodes. BMC Genom. 2015, 16, 589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turchaninova, M.A.; Davydov, A.; Britanova, O.V.; Shugay, M.; Bikos, V.; Egorov, E.S.; Kirgizova, V.I.; Merzlyak, E.M.; Staroverov, D.B.; Bolotin, D.A.; et al. High-Quality Full-Length Immunoglobulin Profiling with Unique Molecular Barcoding. Nat. Protoc. 2016, 11, 1599–1616. [Google Scholar] [CrossRef] [PubMed]
- Rapid Amplification of 5′ Complementary DNA Ends (5′RACE). Nat. Methods 2005, 2, 629–630. [CrossRef] [PubMed]
- Freeman, L.A. Cloning Full-Length Transcripts and Transcript Variants Using 5′ and 3′ RACE. Methods Mol. Biol. 2013, 1027, 3–17. [Google Scholar] [CrossRef]
- Matz, M.; Shagin, D.; Bogdanova, E.; Britanova, O.; Lukyanov, S.; Diatchenko, L.; Chenchik, A. Amplification of CDNA Ends Based on Template-Switching Effect and Step-out PCR. Nucleic Acids Res. 1999, 27, 1558–1560. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; He, L.; Cai, L. Transcriptome Sequencing: RNA-Seq. In Computational Systems Biology: Methods and Protocols; Huang, T., Ed.; Springer: New York, NY, USA, 2018; pp. 15–27. ISBN 978-1-4939-7717-8. [Google Scholar]
- Linnemann, C.; Heemskerk, B.; Kvistborg, P.; Kluin, R.J.C.; Bolotin, D.A.; Chen, X.; Bresser, K.; Nieuwland, M.; Schotte, R.; Michels, S.; et al. High-Throughput Identification of Antigen-Specific TCRs by TCR Gene Capture. Nat. Med. 2013, 19, 1534–1541. [Google Scholar] [CrossRef]
- Mulder, D.T.; Mahé, E.R.; Dowar, M.; Hanna, Y.; Li, T.; Nguyen, L.T.; Butler, M.O.; Hirano, N.; Delabie, J.; Ohashi, P.S.; et al. CapTCR-Seq: Hybrid Capture for T-Cell Receptor Repertoire Profiling. Blood Adv. 2018, 2, 3506–3514. [Google Scholar] [CrossRef]
- Kozarewa, I.; Armisen, J.; Gardner, A.F.; Slatko, B.E.; Hendrickson, C.L. Overview of Target Enrichment Strategies. Curr. Protoc. Mol. Biol. 2015, 112, 7.21.1–7.21.23. [Google Scholar] [CrossRef]
- Mamanova, L.; Coffey, A.J.; Scott, C.E.; Kozarewa, I.; Turner, E.H.; Kumar, A.; Howard, E.; Shendure, J.; Turner, D.J. Target-Enrichment Strategies for next-Generation Sequencing. Nat. Methods 2010, 7, 111–118. [Google Scholar] [CrossRef]
- Linnemann, C.; Mezzadra, R.; Schumacher, T.N.M. TCR Repertoires of Intratumoral T-Cell Subsets. Immunol. Rev. 2014, 257, 72–82. [Google Scholar] [CrossRef]
- Looney, T.J.; Topacio-Hall, D.; Lowman, G.; Conroy, J.; Morrison, C.; Oh, D.; Fong, L.; Zhang, L. TCR Convergence in Individuals Treated With Immune Checkpoint Inhibition for Cancer. Front. Immunol. 2019, 10, 2985. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, H.; Yamaguchi, R.; Liu, X.; Daigo, Y.; Yew, P.Y.; Tanikawa, C.; Matsuda, K.; Imoto, S.; Miyano, S.; Nakamura, Y. Quantitative T Cell Repertoire Analysis by Deep CDNA Sequencing of T Cell Receptor α and β Chains Using Next-Generation Sequencing (NGS). OncoImmunology 2014, 3, e968467. [Google Scholar] [CrossRef] [PubMed]
- Lay, L.; Stroup, B.; Payton, J.E. Validation and Interpretation of IGH and TCR Clonality Testing by Ion Torrent S5 NGS for Diagnosis and Disease Monitoring in B and T Cell Cancers. Pract. Lab. Med. 2020, 22, e00191. [Google Scholar] [CrossRef] [PubMed]
- van Schaik, B.; Klarenbeek, P.; Doorenspleet, M.; van Kampen, A.; Moody, D.B.; de Vries, N.; Van Rhijn, I. Discovery of Invariant T Cells by Next-Generation Sequencing of the Human TCR α-Chain Repertoire. J. Immunol. 2014, 193, 5338–5344. [Google Scholar] [CrossRef] [Green Version]
- Kitaura, K.; Shini, T.; Matsutani, T.; Suzuki, R. A New High-Throughput Sequencing Method for Determining Diversity and Similarity of T Cell Receptor (TCR) α and β Repertoires and Identifying Potential New Invariant TCR α Chains. BMC Immunol. 2016, 17, 38. [Google Scholar] [CrossRef] [Green Version]
- Rosati, E.; Dowds, C.M.; Liaskou, E.; Henriksen, E.K.K.; Karlsen, T.H.; Franke, A. Overview of Methodologies for T-Cell Receptor Repertoire Analysis. BMC Biotechnol. 2017, 17, 61. [Google Scholar] [CrossRef]
- Barennes, P.; Quiniou, V.; Shugay, M.; Egorov, E.S.; Davydov, A.N.; Chudakov, D.M.; Uddin, I.; Ismail, M.; Oakes, T.; Chain, B.; et al. Benchmarking of T Cell Receptor Repertoire Profiling Methods Reveals Large Systematic Biases. Nat. Biotechnol. 2021, 39, 236–245. [Google Scholar] [CrossRef]
- Glanville, J.; Huang, H.; Nau, A.; Hatton, O.; Wagar, L.E.; Rubelt, F.; Ji, X.; Han, A.; Krams, S.M.; Pettus, C.; et al. Identifying Specificity Groups in the T Cell Receptor Repertoire. Nature 2017, 547, 94–98. [Google Scholar] [CrossRef]
- Greiff, V.; Miho, E.; Menzel, U.; Reddy, S.T. Bioinformatic and Statistical Analysis of Adaptive Immune Repertoires. Trends Immunol. 2015, 36, 738–749. [Google Scholar] [CrossRef]
- Bolotin, D.A.; Poslavsky, S.; Mitrophanov, I.; Shugay, M.; Mamedov, I.Z.; Putintseva, E.V.; Chudakov, D.M. MiXCR: Software for Comprehensive Adaptive Immunity Profiling. Nat. Methods 2015, 12, 380–381. [Google Scholar] [CrossRef]
- Ye, J.; Ma, N.; Madden, T.L.; Ostell, J.M. IgBLAST: An Immunoglobulin Variable Domain Sequence Analysis Tool. Nucleic Acids Res. 2013, 41, W34–W40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alamyar, E.; Duroux, P.; Lefranc, M.-P.; Giudicelli, V. IMGT® Tools for the Nucleotide Analysis of Immunoglobulin (IG) and T Cell Receptor (TR) V-(D)-J Repertoires, Polymorphisms, and IG Mutations: IMGT/V-QUEST and IMGT/HighV-QUEST for NGS. In Immunogenetics: Methods and Applications in Clinical Practice; Methods in Molecular, Biology; Christiansen, F.T., Tait, B.D., Eds.; Humana Press: Totowa, NJ, USA, 2012; pp. 569–604. ISBN 978-1-61779-842-9. [Google Scholar]
- Smakaj, E.; Babrak, L.; Ohlin, M.; Shugay, M.; Briney, B.; Tosoni, D.; Galli, C.; Grobelsek, V.; D’Angelo, I.; Olson, B.; et al. Benchmarking Immunoinformatic Tools for the Analysis of Antibody Repertoire Sequences. Bioinformatics 2020, 36, 1731–1739. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shugay, M.; Bagaev, D.V.; Turchaninova, M.A.; Bolotin, D.A.; Britanova, O.V.; Putintseva, E.V.; Pogorelyy, M.V.; Nazarov, V.I.; Zvyagin, I.V.; Kirgizova, V.I.; et al. VDJtools: Unifying Post-Analysis of T Cell Receptor Repertoires. PLoS Comput. Biol. 2015, 11, e1004503. [Google Scholar] [CrossRef] [PubMed]
- Ni, Q.; Zhang, J.; Zheng, Z.; Chen, G.; Christian, L.; Grönholm, J.; Yu, H.; Zhou, D.; Zhuang, Y.; Li, Q.-J.; et al. VisTCR: An Interactive Software for T Cell Repertoire Sequencing Data Analysis. Front. Genet. 2020, 11, 771. [Google Scholar] [CrossRef] [PubMed]
- Bystry, V.; Reigl, T.; Krejci, A.; Demko, M.; Hanakova, B.; Grioni, A.; Knecht, H.; Schlitt, M.; Dreger, P.; Sellner, L.; et al. ARResT/Interrogate: An Interactive Immunoprofiler for IG/TR NGS Data. Bioinformatics 2017, 33, 435–437. [Google Scholar] [CrossRef] [Green Version]
- Scott, J.K.; Breden, F. The Adaptive Immune Receptor Repertoire Community as a Model for FAIR Stewardship of Big Immunology Data. Curr. Opin. Syst. Biol. 2020, 24, 71–77. [Google Scholar] [CrossRef]
- Pannetier, C.; Even, J.; Kourilsky, P. T-Cell Repertoire Diversity and Clonal Expansions in Normal and Clinical Samples. Immunol. Today 1995, 16, 176–181. [Google Scholar] [CrossRef]
- Li, B.; Li, T.; Pignon, J.-C.; Wang, B.; Wang, J.; Shukla, S.A.; Dou, R.; Chen, Q.; Hodi, F.S.; Choueiri, T.K.; et al. Landscape of Tumor-Infiltrating T Cell Repertoire of Human Cancers. Nat. Genet. 2016, 48, 725–732. [Google Scholar] [CrossRef]
- Cui, J.-H.; Lin, K.-R.; Yuan, S.-H.; Jin, Y.-B.; Chen, X.-P.; Su, X.-K.; Jiang, J.; Pan, Y.-M.; Mao, S.-L.; Mao, X.-F.; et al. TCR Repertoire as a Novel Indicator for Immune Monitoring and Prognosis Assessment of Patients With Cervical Cancer. Front. Immunol. 2018, 9, 2729. [Google Scholar] [CrossRef] [Green Version]
- Bolotin, D.A.; Mamedov, I.Z.; Britanova, O.V.; Zvyagin, I.V.; Shagin, D.; Ustyugova, S.V.; Turchaninova, M.A.; Lukyanov, S.; Lebedev, Y.B.; Chudakov, D.M. Next Generation Sequencing for TCR Repertoire Profiling: Platform-Specific Features and Correction Algorithms. Eur. J. Immunol. 2012, 42, 3073–3083. [Google Scholar] [CrossRef] [Green Version]
- Postow, M.A.; Manuel, M.; Wong, P.; Yuan, J.; Dong, Z.; Liu, C.; Perez, S.; Tanneau, I.; Noel, M.; Courtier, A.; et al. Peripheral T Cell Receptor Diversity Is Associated with Clinical Outcomes Following Ipilimumab Treatment in Metastatic Melanoma. J. Immunother. Cancer 2015, 3, 25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, P.; Xu, Z.; Zhou, W.; Jin, X.; Xu, C.; Luo, M.; Ma, K.; Cao, H.; Huang, Y.; Lin, X.; et al. Identification of Potential Vaccine Targets for COVID-19 by Combining Single-Cell and Bulk TCR Sequencing. Clin. Transl. Med. 2021, 11, e430. [Google Scholar] [CrossRef] [PubMed]
- Keylock, C.J. Simpson Diversity and the Shannon–Wiener Index as Special Cases of a Generalized Entropy. Oikos 2005, 109, 203–207. [Google Scholar] [CrossRef]
- Simpson, E.H. Measurement of Diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Venturi, V.; Kedzierska, K.; Turner, S.J.; Doherty, P.C.; Davenport, M.P. Methods for Comparing the Diversity of Samples of the T Cell Receptor Repertoire. J. Immunol. Methods 2007, 321, 182–195. [Google Scholar] [CrossRef]
- Magurran, A.E. Measuring Biological Diversity; John Wiley & Sons: Hoboken, NJ, USA, 2013; ISBN 978-1-118-68792-5. [Google Scholar]
- Shannon, C.E. A Mathematical Theory of Communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef] [Green Version]
- Stewart, J.J.; Lee, C.Y.; Ibrahim, S.; Watts, P.; Shlomchik, M.; Weigert, M.; Litwin, S. A Shannon Entropy Analysis of Immunoglobulin and T Cell Receptor. Mol. Immunol. 1997, 34, 1067–1082. [Google Scholar] [CrossRef]
- Keane, C.; Gould, C.; Jones, K.; Hamm, D.; Talaulikar, D.; Ellis, J.; Vari, F.; Birch, S.; Han, E.; Wood, P.; et al. The T-Cell Receptor Repertoire Influences the Tumor Microenvironment and Is Associated with Survival in Aggressive B-Cell Lymphoma. Clin. Cancer Res. 2017, 23, 1820–1828. [Google Scholar] [CrossRef] [Green Version]
- Willis, A.D. Rarefaction, Alpha Diversity, and Statistics. Front. Microbiol. 2019, 10, 2407. [Google Scholar] [CrossRef] [Green Version]
- Hill, M.O. Diversity and Evenness: A Unifying Notation and Its Consequences. Ecology 1973, 54, 427–432. [Google Scholar] [CrossRef] [Green Version]
- Greiff, V.; Bhat, P.; Cook, S.C.; Menzel, U.; Kang, W.; Reddy, S.T. A Bioinformatic Framework for Immune Repertoire Diversity Profiling Enables Detection of Immunological Status. Genome Med. 2015, 7, 49. [Google Scholar] [CrossRef] [PubMed]
- Kaplinsky, J.; Arnaout, R. Robust Estimates of Overall Immune-Repertoire Diversity from High-Throughput Measurements on Samples. Nat. Commun. 2016, 7, 11881. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mora, T.; Walczak, A.M.; Bialek, W.; Callan, C.G. Maximum Entropy Models for Antibody Diversity. Proc. Natl. Acad. Sci. USA 2010, 107, 5405–5410. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gielis, S.; Moris, P.; Bittremieux, W.; De Neuter, N.; Ogunjimi, B.; Laukens, K.; Meysman, P. Identification of Epitope-Specific T Cells in T-Cell Receptor Repertoires. Methods Mol. Biol. 2020, 2120, 183–195. [Google Scholar] [CrossRef]
- Montemurro, A.; Schuster, V.; Povlsen, H.R.; Bentzen, A.K.; Jurtz, V.; Chronister, W.D.; Crinklaw, A.; Hadrup, S.R.; Winther, O.; Peters, B.; et al. NetTCR-2.0 Enables Accurate Prediction of TCR-Peptide Binding by Using Paired TCRα and β Sequence Data. Commun. Biol. 2021, 4, 1060. [Google Scholar] [CrossRef] [PubMed]
- Tickotsky, N.; Sagiv, T.; Prilusky, J.; Shifrut, E.; Friedman, N. McPAS-TCR: A Manually Curated Catalogue of Pathology-Associated T Cell Receptor Sequences. Bioinformatics 2017, 33, 2924–2929. [Google Scholar] [CrossRef] [Green Version]
- Klinger, M.; Pepin, F.; Wilkins, J.; Asbury, T.; Wittkop, T.; Zheng, J.; Moorhead, M.; Faham, M. Multiplex Identification of Antigen-Specific T Cell Receptors Using a Combination of Immune Assays and Immune Receptor Sequencing. PLoS ONE 2015, 10, e0141561. [Google Scholar] [CrossRef]
- Shugay, M.; Bagaev, D.V.; Zvyagin, I.V.; Vroomans, R.M.; Crawford, J.C.; Dolton, G.; Komech, E.A.; Sycheva, A.L.; Koneva, A.E.; Egorov, E.S.; et al. VDJdb: A Curated Database of T-Cell Receptor Sequences with Known Antigen Specificity. Nucleic Acids Res. 2018, 46, D419–D427. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Zhan, X.; Li, B. GIANA Allows Computationally-Efficient TCR Clustering and Multi-Disease Repertoire Classification by Isometric Transformation. Nat. Commun. 2021, 12, 4699. [Google Scholar] [CrossRef]
- Pogorelyy, M.V.; Minervina, A.A.; Shugay, M.; Chudakov, D.M.; Lebedev, Y.B.; Mora, T.; Walczak, A.M. Detecting T Cell Receptors Involved in Immune Responses from Single Repertoire Snapshots. PLoS Biol. 2019, 17, e3000314. [Google Scholar] [CrossRef] [Green Version]
- Valkiers, S.; Van Houcke, M.; Laukens, K.; Meysman, P. ClusTCR: A Python Interface for Rapid Clustering of Large Sets of CDR3 Sequences with Unknown Antigen Specificity. Bioinformatics 2021, 37, 4865–4867. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Wang, C.; Rubelt, F.; Scriba, T.J.; Davis, M.M. Analyzing the Mycobacterium Tuberculosis Immune Response by T-Cell Receptor Clustering with GLIPH2 and Genome-Wide Antigen Screening. Nat. Biotechnol. 2020, 38, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, L.; Zhang, J.; Chen, J.; Ye, J.; Shukla, S.; Qiao, J.; Zhan, X.; Chen, H.; Wu, C.J.; et al. Investigation of Antigen-Specific T-Cell Receptor Clusters in Human Cancers. Clin. Cancer Res. 2020, 26, 1359–1371. [Google Scholar] [CrossRef] [PubMed]
- Dash, P.; Fiore-Gartland, A.J.; Hertz, T.; Wang, G.C.; Sharma, S.; Souquette, A.; Crawford, J.C.; Clemens, E.B.; Nguyen, T.H.O.; Kedzierska, K.; et al. Quantifiable Predictive Features Define Epitope-Specific T Cell Receptor Repertoires. Nature 2017, 547, 89–93. [Google Scholar] [CrossRef] [Green Version]
- Sidhom, J.-W.; Bessell, C.A.; Havel, J.J.; Kosmides, A.; Chan, T.A.; Schneck, J.P. ImmunoMap: A Bioinformatics Tool for T-Cell Repertoire Analysis. Cancer Immunol. Res. 2018, 6, 151–162. [Google Scholar] [CrossRef] [Green Version]
- Rajeh, A.; Wolf, K.; Schiebout, C.; Sait, N.; Kosfeld, T.; DiPaolo, R.J.; Ahn, T.-H. ICAT: Diagnostic Assessment Tool of Immunological History Using High-Throughput T-Cell Receptor Sequencing. F1000Research 2021, 10, 65. [Google Scholar] [CrossRef]
- Henikoff, S.; Henikoff, J.G. Amino Acid Substitution Matrices from Protein Blocks. Proc. Natl. Acad. Sci. USA 1992, 89, 10915–10919. [Google Scholar] [CrossRef] [Green Version]
- Sidhom, J.-W.; Larman, H.B.; Pardoll, D.M.; Baras, A.S. DeepTCR Is a Deep Learning Framework for Revealing Sequence Concepts within T-Cell Repertoires. Nat. Commun. 2021, 12, 1605. [Google Scholar] [CrossRef]
- Sidhom, J.-W.; Baras, A.S. Deep Learning Identifies Antigenic Determinants of Severe SARS-CoV-2 Infection within T-Cell Repertoires. Sci. Rep. 2021, 11, 14275. [Google Scholar] [CrossRef]
- Yohannes, D.A.; Kaukinen, K.; Kurppa, K.; Saavalainen, P.; Greco, D. Clustering Based Approach for Population Level Identification of Condition-Associated T-Cell Receptor β-Chain CDR3 Sequences. BMC Bioinform. 2021, 22, 159. [Google Scholar] [CrossRef]
- Shomuradova, A.S.; Vagida, M.S.; Sheetikov, S.A.; Zornikova, K.V.; Kiryukhin, D.; Titov, A.; Peshkova, I.O.; Khmelevskaya, A.; Dianov, D.V.; Malasheva, M.; et al. SARS-CoV-2 Epitopes Are Recognized by a Public and Diverse Repertoire of Human T Cell Receptors. Immunity 2020, 53, 1245–1257.e5. [Google Scholar] [CrossRef] [PubMed]
- Schultheiß, C.; Paschold, L.; Simnica, D.; Mohme, M.; Willscher, E.; von Wenserski, L.; Scholz, R.; Wieters, I.; Dahlke, C.; Tolosa, E.; et al. Next-Generation Sequencing of T and B Cell Receptor Repertoires from COVID-19 Patients Showed Signatures Associated with Severity of Disease. Immunity 2020, 53, 442–455.e4. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Li, S.; Li, P.; Pan, W.; Wang, Q.; Feng, Y.; Mo, X.; Yan, Q.; Ye, X.; Luo, J.; et al. Longitudinal Analysis of T and B Cell Receptor Repertoire Transcripts Reveal Dynamic Immune Response in COVID-19 Patients. Front. Immunol. 2020, 11, 582010. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.-M.; Feng, P.-H.; Wu, T.-H.; Alachkar, H.; Lee, K.-Y.; Chang, W.-C. Profiling of T Cell Repertoire in SARS-CoV-2-Infected COVID-19 Patients Between Mild Disease and Pneumonia. J. Clin. Immunol. 2021, 41, 1131–1145. [Google Scholar] [CrossRef] [PubMed]
- Minervina, A.A.; Komech, E.A.; Titov, A.; Bensouda Koraichi, M.; Rosati, E.; Mamedov, I.Z.; Franke, A.; Efimov, G.A.; Chudakov, D.M.; Mora, T.; et al. Longitudinal High-Throughput TCR Repertoire Profiling Reveals the Dynamics of T-Cell Memory Formation after Mild COVID-19 Infection. eLife 2021, 10, e63502. [Google Scholar] [CrossRef]
- Swanson, P.A., II; Padilla, M.; Hoyland, W.; McGlinchey, K.; Fields, P.A.; Bibi, S.; Faust, S.N.; McDermott, A.B.; Lambe, T.; Pollard, A.J.; et al. AZD1222/ChAdOx1 NCoV-19 Vaccination Induces a Polyfunctional Spike Protein-Specific Th1 Response with a Diverse TCR Repertoire. Sci. Transl. Med. 2021, 13, eabj7211. [Google Scholar] [CrossRef]
- Wang, Y.; Duan, F.; Zhu, Z.; Yu, M.; Jia, X.; Dai, H.; Wang, P.; Qiu, X.; Lu, Y.; Huang, J. Analysis of TCR Repertoire by High-Throughput Sequencing Indicates the Feature of T Cell Immune Response after SARS-CoV-2 Infection. Cells 2021, 11, 68. [Google Scholar] [CrossRef]
- Hu, W.; He, M.; Wang, X.; Sun, Q.; Kuang, M. Specific CD8+ TCR Repertoire Recognizing Conserved Antigens of SARS-CoV-2 in Unexposed Population: A Prerequisite for Broad-Spectrum CD8+ T Cell Immunity. Vaccines 2021, 9, 1093. [Google Scholar] [CrossRef]
- Simnica, D.; Schultheiß, C.; Mohme, M.; Paschold, L.; Willscher, E.; Fitzek, A.; Püschel, K.; Matschke, J.; Ciesek, S.; Sedding, D.G.; et al. Landscape of T-Cell Repertoires with Public COVID-19-Associated T-Cell Receptors in Pre-Pandemic Risk Cohorts. Clin. Transl. Immunol. 2021, 10, e1340. [Google Scholar] [CrossRef]
- Shimizu, K.; Iyoda, T.; Sanpei, A.; Nakazato, H.; Okada, M.; Ueda, S.; Kato-Murayama, M.; Murayama, K.; Shirouzu, M.; Harada, N.; et al. Identification of TCR Repertoires in Functionally Competent Cytotoxic T Cells Cross-Reactive to SARS-CoV-2. Commun. Biol. 2021, 4, 1365. [Google Scholar] [CrossRef]
- Li, Y.; Hu, J.; Wang, Y.; Liu, D.; Shi, Y.; Zhang, J.; Liu, Y.; Lin, D.; Lin, J.; Hu, W.; et al. T-Cell Repertoire Characteristics of Asymptomatic and Re-Detectable Positive COVID-19 Patients. Front. Immunol. 2021, 12, 769442. [Google Scholar] [CrossRef] [PubMed]

| Method | DNA/RNA | Principles | Advantages | Disadvantages | Examples of Manufacturers |
|---|---|---|---|---|---|
| 5′RACE | RNA |
|
|
| iRepertoire, Clontech/Takara, MiLaboratories |
| Multiplex PCR | DNA, RNA |
|
|
| Adaptive Biotechnologies (Immunoseq); Invivoscribe (Lymphotrack) |
| Hybridisation capture | DNA, RNA |
|
|
| Bespoke approaches |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kockelbergh, H.; Evans, S.; Deng, T.; Clyne, E.; Kyriakidou, A.; Economou, A.; Luu Hoang, K.N.; Woodmansey, S.; Foers, A.; Fowler, A.; et al. Utility of Bulk T-Cell Receptor Repertoire Sequencing Analysis in Understanding Immune Responses to COVID-19. Diagnostics 2022, 12, 1222. https://doi.org/10.3390/diagnostics12051222
Kockelbergh H, Evans S, Deng T, Clyne E, Kyriakidou A, Economou A, Luu Hoang KN, Woodmansey S, Foers A, Fowler A, et al. Utility of Bulk T-Cell Receptor Repertoire Sequencing Analysis in Understanding Immune Responses to COVID-19. Diagnostics. 2022; 12(5):1222. https://doi.org/10.3390/diagnostics12051222
Chicago/Turabian StyleKockelbergh, Hannah, Shelley Evans, Tong Deng, Ella Clyne, Anna Kyriakidou, Andreas Economou, Kim Ngan Luu Hoang, Stephen Woodmansey, Andrew Foers, Anna Fowler, and et al. 2022. "Utility of Bulk T-Cell Receptor Repertoire Sequencing Analysis in Understanding Immune Responses to COVID-19" Diagnostics 12, no. 5: 1222. https://doi.org/10.3390/diagnostics12051222







