COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2
Abstract
:1. Introduction
2. History of SARS-CoV-2 or Epidemiology
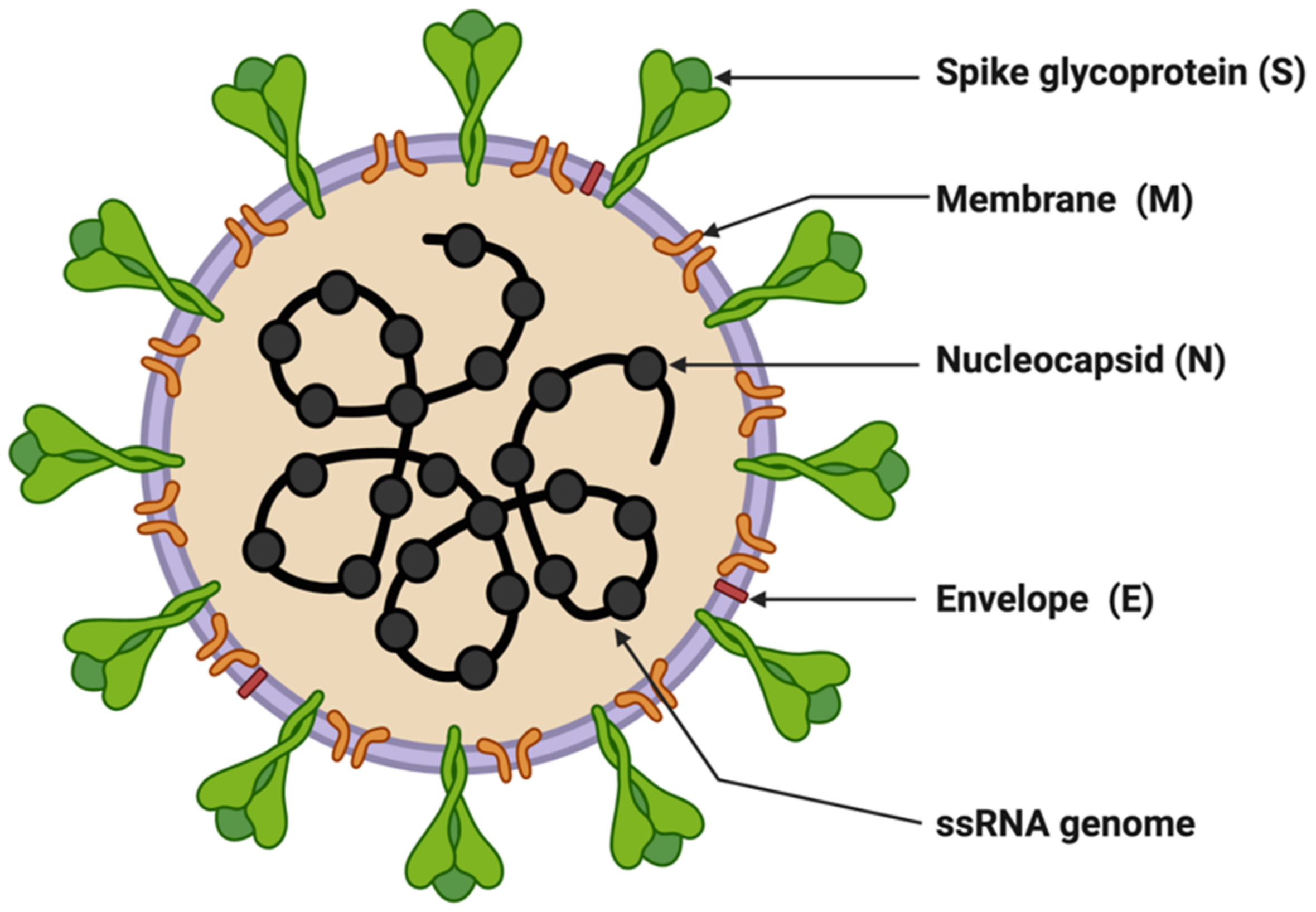
3. Molecular Biology of SARS-CoV-2
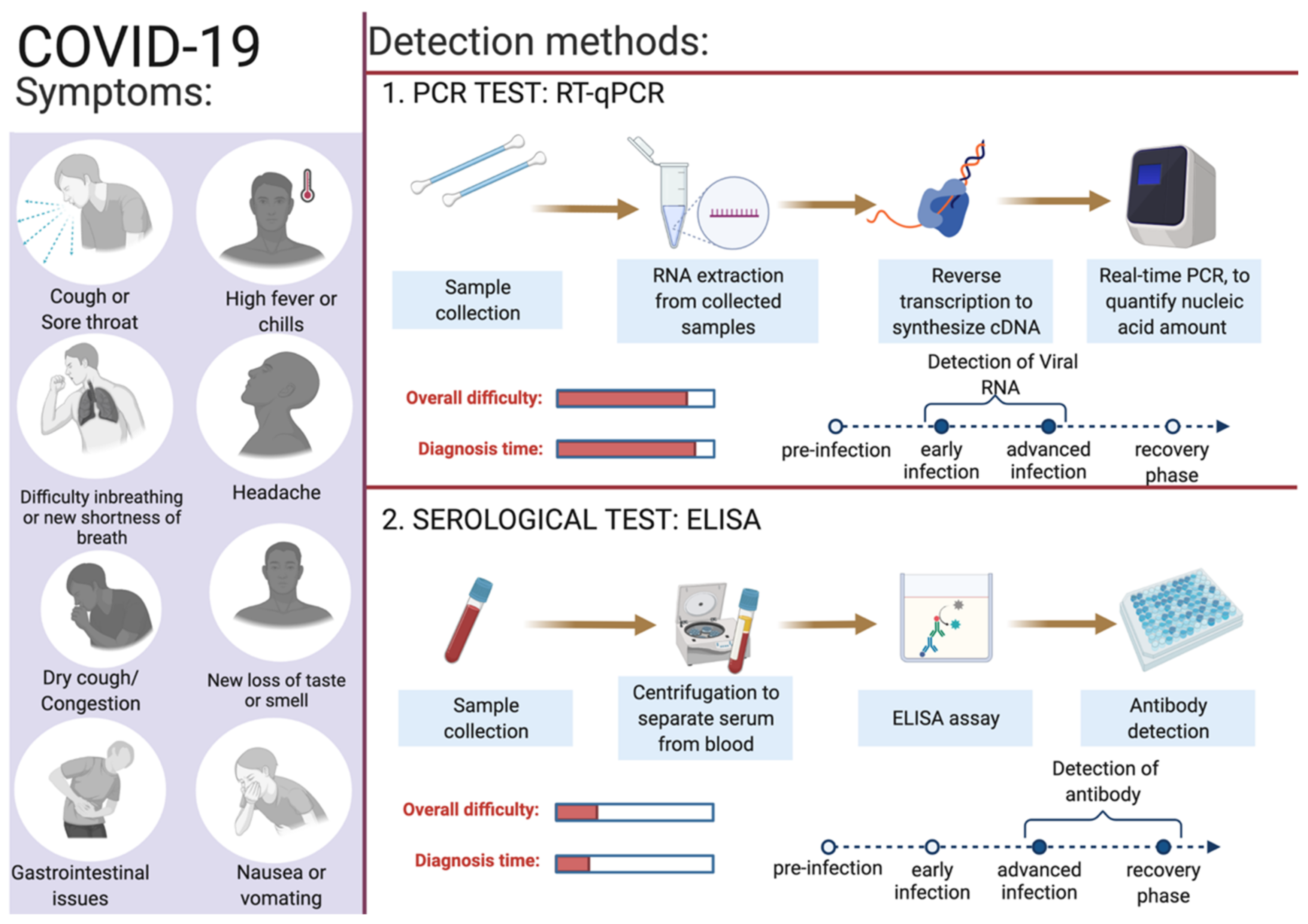
4. Diagnostics for COVID-19
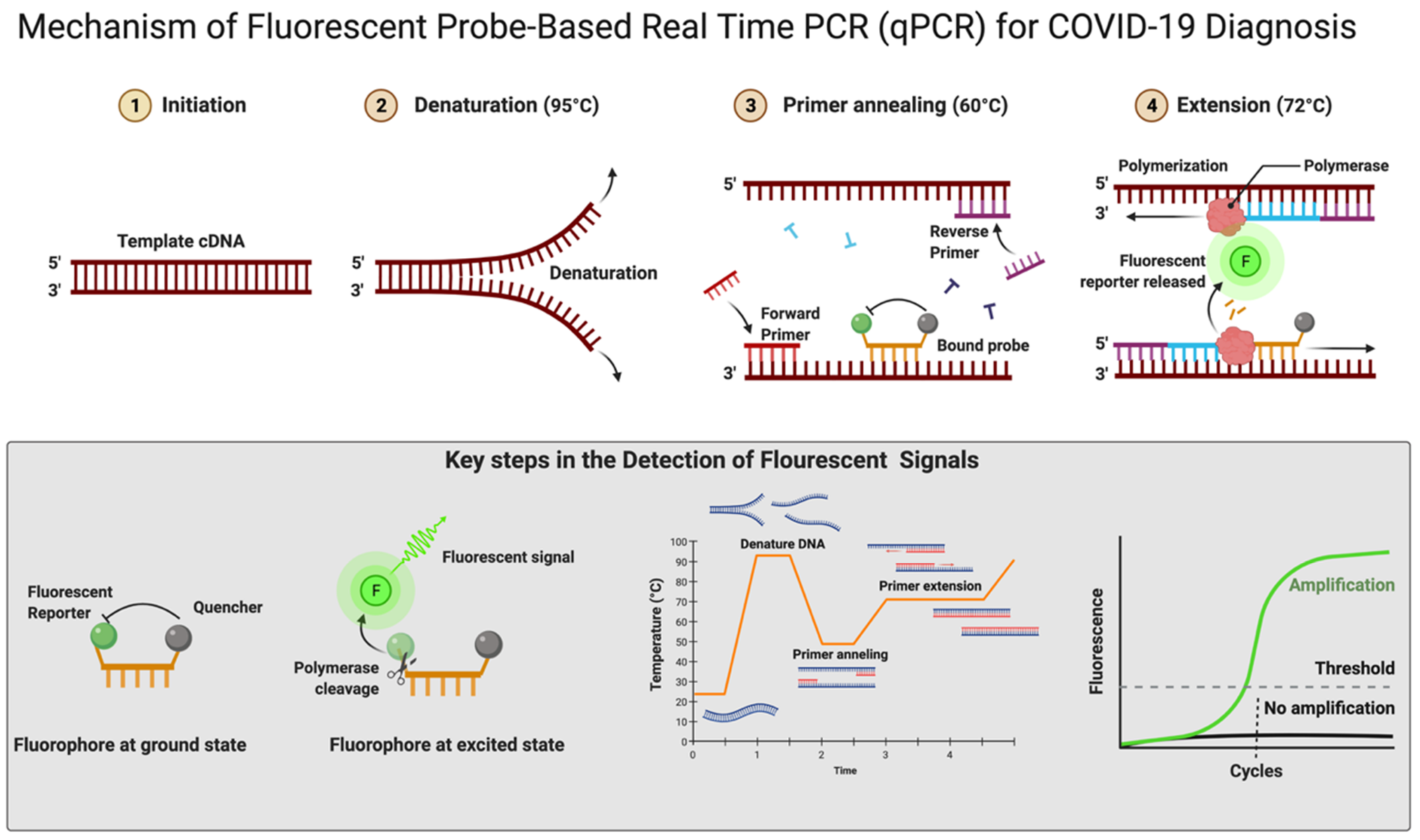
4.1. Reverse-Transcriptase PCR (RT-qPCR)
4.2. Specimens for Detection of SARS-CoV-2
4.3. Biomarkers/Genes Used for RT-qPCR
4.4. Reagents (Dyes)
4.5. Ct Value/Threshold Value
5. Limitations of RT-qPCR Detection Technique for SARS-CoV-2
6. Future Perspectives and Conclusions
Supplementary Materials
Funding
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Mackenzie, J.S.; Smith, D.W. COVID-19: A novel zoonotic disease caused by a coronavirus from China: What we know and What we don’t. Microbiol. Aust. 2020, 41, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta Biomed. 2020, 91, 157–160. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [Green Version]
- Sethuraman, N.; Jeremiah, S.S.; Ryo, A. Interpreting Diagnostic Tests for SARS-CoV-2. JAMA 2020, 323, 2249–2251. [Google Scholar] [CrossRef]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822–3835. [Google Scholar] [CrossRef] [Green Version]
- Escandon, K.; Rasmussen, A.L.; Bogoch, I.I.; Murray, E.J.; Escandon, K.; Popescu, S.V.; Kindrachuk, J. COVID-19 false dichotomies and a comprehensive review of the evidence regarding public health, COVID-19 symptomatology, SARS-CoV-2 transmission, mask wearing, and reinfection. BMC Infect. Dis. 2021, 21, 710. [Google Scholar] [CrossRef]
- Woloshin, S.; Patel, N.; Kesselheim, A.S. False Negative Tests for SARS-CoV-2 Infection—Challenges and Implications. N. Engl. J. Med. 2020, 383, e38. [Google Scholar] [CrossRef]
- Liu, R.; Han, H.; Liu, F.; Lv, Z.; Wu, K.; Liu, Y.; Feng, Y.; Zhu, C. Positive rate of RT-PCR detection of SARS-CoV-2 infection in 4880 cases from one hospital in Wuhan, China, from Jan to Feb 2020. Clin. Chim. Acta 2020, 505, 172–175. [Google Scholar] [CrossRef]
- Drame, M.; Tabue Teguo, M.; Proye, E.; Hequet, F.; Hentzien, M.; Kanagaratnam, L.; Godaert, L. Should RT-PCR be considered a gold standard in the diagnosis of COVID-19? J. Med. Virol. 2020, 92, 2312–2313. [Google Scholar] [CrossRef]
- Ksiazek, T.G.; Erdman, D.; Goldsmith, C.S.; Zaki, S.R.; Peret, T.; Emery, S.; Tong, S.; Urbani, C.; Comer, J.A.; Lim, W.; et al. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1953–1966. [Google Scholar] [CrossRef] [PubMed]
- Chau, C.H.; Strope, J.D.; Figg, W.D. COVID-19 Clinical Diagnostics and Testing Technology. Pharmacotherapy 2020, 40, 857–868. [Google Scholar] [CrossRef] [PubMed]
- Waller, J.V.; Kaur, P.; Tucker, A.; Lin, K.K.; Diaz, M.J.; Henry, T.S.; Hope, M. Diagnostic Tools for Coronavirus Disease (COVID-19): Comparing CT and RT-PCR Viral Nucleic Acid Testing. AJR Am. J. Roentgenol. 2020, 215, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Machhi, J.; Herskovitz, J.; Senan, A.M.; Dutta, D.; Nath, B.; Oleynikov, M.D.; Blomberg, W.R.; Meigs, D.D.; Hasan, M.; Patel, M.; et al. The Natural History, Pathobiology, and Clinical Manifestations of SARS-CoV-2 Infections. J. Neuroimmune Pharm. 2020, 15, 359–386. [Google Scholar] [CrossRef]
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef] [Green Version]
- Pal, M.; Berhanu, G.; Desalegn, C.; Kandi, V. Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2): An Update. Cureus 2020, 12, e7423. [Google Scholar] [CrossRef] [Green Version]
- Mousavizadeh, L.; Ghasemi, S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J. Microbiol. Immunol. Infect. 2021, 54, 159–163. [Google Scholar] [CrossRef]
- Bar-On, Y.M.; Flamholz, A.; Phillips, R.; Milo, R. SARS-CoV-2 (COVID-19) by the numbers. eLife 2020, 9, e57309. [Google Scholar] [CrossRef]
- Smith, E.C.; Denison, M.R. Implications of altered replication fidelity on the evolution and pathogenesis of coronaviruses. Curr. Opin. Virol. 2012, 2, 519–524. [Google Scholar] [CrossRef]
- Ye, Z.W.; Yuan, S.; Yuen, K.S.; Fung, S.Y.; Chan, C.P.; Jin, D.Y. Zoonotic origins of human coronaviruses. Int. J. Biol. Sci. 2020, 16, 1686–1697. [Google Scholar] [CrossRef] [Green Version]
- Liu, D.X.; Liang, J.Q.; Fung, T.S. Human Coronavirus-229E, -OC43, -NL63, and -HKU1 (Coronaviridae). Encycl. Virol. 2021, 2, 428–440. [Google Scholar] [CrossRef]
- Van der Hoek, L.; Pyrc, K.; Jebbink, M.F.; Vermeulen-Oost, W.; Berkhout, R.J.; Wolthers, K.C.; Wertheim-van Dillen, P.M.; Kaandorp, J.; Spaargaren, J.; Berkhout, B. Identification of a new human coronavirus. Nat. Med. 2004, 10, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Woo, P.C.; Huang, Y.; Lau, S.K.; Yuen, K.Y. Coronavirus genomics and bioinformatics analysis. Viruses 2010, 2, 1804–1820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, Y.X.; Ng, Y.L.; Tam, J.P.; Liu, D.X. Human Coronaviruses: A Review of Virus-Host Interactions. Diseases 2016, 4, 26. [Google Scholar] [CrossRef]
- Irwin, R.J.; McEwen, S.A.; Clarke, R.C.; Meek, A.H. The prevalence of verocytotoxin-producing Escherichia coli and antimicrobial resistance patterns of nonverocytotoxin-producing Escherichia coli and Salmonella in Ontario broiler chickens. Can. J. Vet. Res. 1989, 53, 411–418. [Google Scholar]
- Poon, L.L.; Chu, D.K.; Chan, K.H.; Wong, O.K.; Ellis, T.M.; Leung, Y.H.; Lau, S.K.; Woo, P.C.; Suen, K.Y.; Yuen, K.Y.; et al. Identification of a novel coronavirus in bats. J. Virol. 2005, 79, 2001–2009. [Google Scholar] [CrossRef] [Green Version]
- Peiris, J.S.; Lai, S.T.; Poon, L.L.; Guan, Y.; Yam, L.Y.; Lim, W.; Nicholls, J.; Yee, W.K.; Yan, W.W.; Cheung, M.T.; et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 2003, 361, 1319–1325. [Google Scholar] [CrossRef] [Green Version]
- Satija, N.; Lal, S.K. The molecular biology of SARS coronavirus. Ann. N. Y. Acad. Sci. 2007, 1102, 26–38. [Google Scholar] [CrossRef]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Hung, L.S. The SARS epidemic in Hong Kong: What lessons have we learned? J. R. Soc. Med. 2003, 96, 374–378. [Google Scholar] [CrossRef]
- Hung, E.C.; Chim, S.S.; Chan, P.K.; Tong, Y.K.; Ng, E.K.; Chiu, R.W.; Leung, C.B.; Sung, J.J.; Tam, J.S.; Lo, Y.M. Detection of SARS coronavirus RNA in the cerebrospinal fluid of a patient with severe acute respiratory syndrome. Clin. Chem. 2003, 49, 2108–2109. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Qi, T.; Liu, L.; Ling, Y.; Qian, Z.; Li, T.; Li, F.; Xu, Q.; Zhang, Y.; Xu, S.; et al. Clinical progression of patients with COVID-19 in Shanghai, China. J. Infect. 2020, 80, e1–e6. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, A.S.; Tomlinson, B.; Cockram, C.S.; Thomas, G.N. Lessons from the severe acute respiratory syndrome outbreak in Hong Kong. Emerg. Infect. Dis. 2003, 9, 1042–1045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cho, S.Y.; Kang, J.M.; Ha, Y.E.; Park, G.E.; Lee, J.Y.; Ko, J.H.; Lee, J.Y.; Kim, J.M.; Kang, C.I.; Jo, I.J.; et al. MERS-CoV outbreak following a single patient exposure in an emergency room in South Korea: An epidemiological outbreak study. Lancet 2016, 388, 994–1001. [Google Scholar] [CrossRef] [Green Version]
- Mackay, I.M.; Arden, K.E. MERS coronavirus: Diagnostics, epidemiology and transmission. Virol. J. 2015, 12, 222. [Google Scholar] [CrossRef] [Green Version]
- Vos, L.M.; Bruyndonckx, R.; Zuithoff, N.P.A.; Little, P.; Oosterheert, J.J.; Broekhuizen, B.D.L.; Lammens, C.; Loens, K.; Viveen, M.; Butler, C.C.; et al. Lower respiratory tract infection in the community: Associations between viral aetiology and illness course. Clin. Microbiol. Infect. 2021, 27, 96–104. [Google Scholar] [CrossRef]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef]
- Zhu, Z.; Lian, X.; Su, X.; Wu, W.; Marraro, G.A.; Zeng, Y. From SARS and MERS to COVID-19: A brief summary and comparison of severe acute respiratory infections caused by three highly pathogenic human coronaviruses. Respir. Res. 2020, 21, 224. [Google Scholar] [CrossRef]
- Al-Qahtani, A.A. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): Emergence, history, basic and clinical aspects. Saudi J. Biol. Sci. 2020, 27, 2531–2538. [Google Scholar] [CrossRef]
- Llanes, A.; Restrepo, C.M.; Caballero, Z.; Rajeev, S.; Kennedy, M.A.; Lleonart, R. Betacoronavirus Genomes: How Genomic Information has been Used to Deal with Past Outbreaks and the COVID-19 Pandemic. Int. J. Mol. Sci. 2020, 21, 4546. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [Green Version]
- International Committee on Taxonomy of Viruses Executive Committee. The new scope of virus taxonomy: Partitioning the virosphere into 15 hierarchical ranks. Nat. Microbiol. 2020, 5, 668–674. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [Green Version]
- Ren, L.L.; Wang, Y.M.; Wu, Z.Q.; Xiang, Z.C.; Guo, L.; Xu, T.; Jiang, Y.Z.; Xiong, Y.; Li, Y.J.; Li, X.W.; et al. Identification of a novel coronavirus causing severe pneumonia in human: A descriptive study. Chin. Med. J. 2020, 133, 1015–1024. [Google Scholar] [CrossRef]
- Lau, Y.L.; Peiris, J.S. Pathogenesis of severe acute respiratory syndrome. Curr. Opin. Immunol. 2005, 17, 404–410. [Google Scholar] [CrossRef]
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, M.I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [Green Version]
- Chan, J.F.; Kok, K.H.; Zhu, Z.; Chu, H.; To, K.K.; Yuan, S.; Yuen, K.Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef] [Green Version]
- Mariano, G.; Farthing, R.J.; Lale-Farjat, S.L.M.; Bergeron, J.R.C. Structural Characterization of SARS-CoV-2: Where We Are, and Where We Need to Be. Front. Mol. Biosci. 2020, 7, 605236. [Google Scholar] [CrossRef]
- Das, A.; Ahmed, R.; Akhtar, S.; Begum, K.; Banu, S. An overview of basic molecular biology of SARS-CoV-2 and current COVID-19 prevention strategies. Gene Rep. 2021, 23, 101122. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 2020, 11, 1620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, J.; Sun, P.D. High affinity binding of SARS-CoV-2 spike protein enhances ACE2 carboxypeptidase activity. J. Biol. Chem. 2020, 295, 18579–18588. [Google Scholar] [CrossRef]
- Peacock, T.P.; Goldhill, D.H.; Zhou, J.; Baillon, L.; Frise, R.; Swann, O.C.; Kugathasan, R.; Penn, R.; Brown, J.C.; Sanchez-David, R.Y.; et al. The furin cleavage site in the SARS-CoV-2 spike protein is required for transmission in ferrets. Nat. Microbiol. 2021, 6, 899–909. [Google Scholar] [CrossRef]
- Gobeil, S.M.; Janowska, K.; McDowell, S.; Mansouri, K.; Parks, R.; Stalls, V.; Kopp, M.F.; Manne, K.; Li, D.; Wiehe, K.; et al. Effect of natural mutations of SARS-CoV-2 on spike structure, conformation, and antigenicity. Science 2021, 373. [Google Scholar] [CrossRef]
- McCallum, M.; Bassi, J.; De Marco, A.; Chen, A.; Walls, A.C.; Di Iulio, J.; Tortorici, M.A.; Navarro, M.J.; Silacci-Fregni, C.; Saliba, C.; et al. SARS-CoV-2 immune evasion by the B.1.427/B.1.429 variant of concern. Science 2021, 373, 648–654. [Google Scholar] [CrossRef]
- Carroll, T.; Fox, D.; van Doremalen, N.; Ball, E.; Morris, M.K.; Sotomayor-Gonzalez, A.; Servellita, V.; Rustagi, A.; Yinda, C.K.; Fritts, L.; et al. The B.1.427/1.429 (epsilon) SARS-CoV-2 variants are more virulent than ancestral B.1 (614G) in Syrian hamsters. PLoS Pathog. 2022, 18, e1009914. [Google Scholar] [CrossRef]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 2021, 592, 438–443. [Google Scholar] [CrossRef]
- Simulundu, E.; Mupeta, F.; Chanda-Kapata, P.; Saasa, N.; Changula, K.; Muleya, W.; Chitanga, S.; Mwanza, M.; Simusika, P.; Chambaro, H.; et al. First COVID-19 case in Zambia—Comparative phylogenomic analyses of SARS-CoV-2 detected in African countries. Int. J. Infect. Dis. 2021, 102, 455–459. [Google Scholar] [CrossRef]
- Wink, P.L.; Volpato, F.C.Z.; Monteiro, F.L.; Willig, J.B.; Zavascki, A.P.; Barth, A.L.; Martins, A.F. First identification of SARS-CoV-2 lambda (C.37) variant in Southern Brazil. Infect. Control. Hosp. Epidemiol. 2021, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Darvishi, M.; Rahimi, F.; Talebi Bezmin Abadi, A. SARS-CoV-2 Lambda (C.37): An emerging variant of concern? Gene Rep. 2021, 25, 101378. [Google Scholar] [CrossRef] [PubMed]
- Romero, P.E.; Davila-Barclay, A.; Salvatierra, G.; Gonzalez, L.; Cuicapuza, D.; Solis, L.; Marcos-Carbajal, P.; Huancachoque, J.; Maturrano, L.; Tsukayama, P. The Emergence of Sars-CoV-2 Variant Lambda (C.37) in South America. Microbiol. Spectr. 2021, 9, e0078921. [Google Scholar] [CrossRef] [PubMed]
- Lee, R. B.1.1.7: What We Know about the Novel SARS-CoV-2 Variant. Available online: https://asm.org/Articles/2021/January/B-1-1-7-What-We-Know-About-the-Novel-SARS-CoV-2-Va (accessed on 12 June 2022).
- Tang, J.W.; Tambyah, P.A.; Hui, D.S. Emergence of a new SARS-CoV-2 variant in the UK. J. Infect. 2021, 82, e27–e28. [Google Scholar] [CrossRef]
- Wall, E.C.; Wu, M.; Harvey, R.; Kelly, G.; Warchal, S.; Sawyer, C.; Daniels, R.; Hobson, P.; Hatipoglu, E.; Ngai, Y.; et al. Neutralising antibody activity against SARS-CoV-2 VOCs B.1.617.2 and B.1.351 by BNT162b2 vaccination. Lancet 2021, 397, 2331–2333. [Google Scholar] [CrossRef]
- Lovelace, B., Jr. WHO Says Delta Is Becoming the Dominant COVID Variant Globally. Available online: https://www.cnbc.com/2021/06/18/who-says-delta-is-becoming-the-dominant-covid-variant-globally.html (accessed on 12 June 2022).
- Voloch, C.M.; da Silva Francisco, R., Jr.; de Almeida, L.G.P.; Cardoso, C.C.; Brustolini, O.J.; Gerber, A.L.; Guimaraes, A.P.C.; Mariani, D.; da Costa, R.M.; Ferreira, O.C., Jr.; et al. Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. J. Virol. 2021, 95, e00119-21. [Google Scholar] [CrossRef]
- Nonaka, C.K.V.; Graf, T.; Barcia, C.A.L.; Costa, V.F.; de Oliveira, J.L.; Passos, R.D.H.; Bastos, I.N.; de Santana, M.C.B.; Santos, I.M.; de Sousa, K.A.F.; et al. SARS-CoV-2 variant of concern P.1 (Gamma) infection in young and middle-aged patients admitted to the intensive care units of a single hospital in Salvador, Northeast Brazil, February 2021. Int. J. Infect. Dis. 2021, 111, 47–54. [Google Scholar] [CrossRef]
- Annavajhala, M.K.; Mohri, H.; Wang, P.; Nair, M.; Zucker, J.E.; Sheng, Z.; Gomez-Simmonds, A.; Kelley, A.L.; Tagliavia, M.; Huang, Y.; et al. Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York. Nature 2021, 597, 703–708. [Google Scholar] [CrossRef]
- Yang, W.; Greene, S.K.; Peterson, E.R.; Li, W.; Mathes, R.; Graf, L.; Lall, R.; Hughes, S.; Wang, J.; Fine, A. Epidemiological characteristics of the B.1.526 SARS-CoV-2 variant. Sci. Adv. 2022, 8, eabm0300. [Google Scholar] [CrossRef]
- Delli Compagni, E.; Mangone, I.; Bonfini, B.; Di Gennaro, A.; Teodori, L.; Leone, A.; Casaccia, C.; Portanti, O.; Averaimo, D.; Zilli, K.; et al. Whole-Genome Sequences of SARS-CoV-2 Lineage B.1.525 Strains (Variant eta) Detected from Patients in the Abruzzo Region (Central Italy) during Spring 2021. Microbiol. Resour. Announc. 2021, 10, e0061821. [Google Scholar] [CrossRef]
- Ozer, E.A.; Simons, L.M.; Adewumi, O.M.; Fowotade, A.A.; Omoruyi, E.C.; Adeniji, J.A.; Olayinka, O.A.; Dean, T.J.; Zayas, J.; Bhimalli, P.P.; et al. Multiple expansions of globally uncommon SARS-CoV-2 lineages in Nigeria. Nat. Commun. 2022, 13, 688. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, I.; Kemp, S.A.; Datir, R.; Saito, A.; Meng, B.; Rakshit, P.; Takaori-Kondo, A.; Kosugi, Y.; Uriu, K.; Kimura, I.; et al. SARS-CoV-2 B.1.617 Mutations L452R and E484Q Are Not Synergistic for Antibody Evasion. J. Infect. Dis. 2021, 224, 989–994. [Google Scholar] [CrossRef] [PubMed]
- Mlcochova, P.; Kemp, S.A.; Dhar, M.S.; Papa, G.; Meng, B.; Ferreira, I.; Datir, R.; Collier, D.A.; Albecka, A.; Singh, S.; et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 2021, 599, 114–119. [Google Scholar] [CrossRef] [PubMed]
- Haw, N.J.L.; Canal, E.M.R.; Zuasula, J., Jr.; Loreche, M.J.; Bernadas, J. Epidemiological characteristics of the SARS-CoV-2 Theta variant (P.3) in the Central Visayas region, Philippines, 30 October 2020-16 February 2021. West. Pac. Surveill. Response J. 2022, 13, 1–3. [Google Scholar] [CrossRef]
- Halfmann, P.J.; Kuroda, M.; Armbrust, T.; Theiler, J.; Balaram, A.; Moreno, G.K.; Accola, M.A.; Iwatsuki-Horimoto, K.; Valdez, R.; Stoneman, E.; et al. Characterization of the SARS-CoV-2 B.1.621 (Mu) variant. Sci. Transl. Med. 2022, eabm4908. [Google Scholar] [CrossRef]
- Barrera-Avalos, C.; Luraschi, R.; Acuna-Castillo, C.; Vidal, M.; Mella-Torres, A.; Inostroza-Molina, A.; Vera, R.; Vargas, S.; Hernandez, I.; Perez, C.; et al. Description of Symptoms Caused by the Infection of the SARS-CoV-2 B.1.621 (Mu) Variant in Patients With Complete CoronaVac Vaccination Scheme: First Case Report From Santiago of Chile. Front. Public Health 2022, 10, 797569. [Google Scholar] [CrossRef]
- Manouana, G.P.; Nzamba Maloum, M.; Bikangui, R.; Oye Bingono, S.O.; Ondo Nguema, G.; Honkpehedji, J.Y.; Rossatanga, E.G.; Zoa-Assoumou, S.; Pallerla, S.R.; Rachakonda, S.; et al. Emergence of B.1.1.318 SARS-CoV-2 viral lineage and high incidence of alpha B.1.1.7 variant of concern in the Republic of Gabon. Int. J. Infect. Dis. 2022, 114, 151–154. [Google Scholar] [CrossRef]
- Scheepers, C.; Everatt, J.; Amoako, D.G.; Tegally, H.; Wibmer, C.K.; Mnguni, A.; Ismail, A.; Mahlangu, B.; Lambson, B.E.; Richardson, S.I.; et al. Emergence and phenotypic characterization of C.1.2, a globally detected lineage that rapidly accumulated mutations of concern. medRxiv 2021. [Google Scholar] [CrossRef]
- Colson, P.; Delerce, J.; Burel, E.; Dahan, J.; Jouffret, A.; Fenollar, F.; Yahi, N.; Fantini, J.; La Scola, B.; Raoult, D. Emergence in Southern France of a new SARS-CoV-2 variant of probably Cameroonian origin harbouring both substitutions N501Y and E484K in the spike protein. medRxiv 2021. [Google Scholar] [CrossRef]
- Ren, S.Y.; Wang, W.B.; Gao, R.D.; Zhou, A.M. Omicron variant (B.1.1.529) of SARS-CoV-2: Mutation, infectivity, transmission, and vaccine resistance. World J. Clin. Cases 2022, 10, 1–11. [Google Scholar] [CrossRef]
- World Health Organization. Enhancing Response to Omicron SARS-CoV-2 Variant. Available online: https://www.who.int/publications/m/item/enhancing-readiness-for-omicron-(b.1.1.529)-technical-brief-and-priority-actions-for-member-states (accessed on 12 June 2022).
- Chen, J.; Wei, G.W. Omicron BA.2 (B.1.1.529.2): High Potential for Becoming the Next Dominant Variant. J. Phys. Chem. Lett. 2022, 13, 3840–3849. [Google Scholar] [CrossRef] [PubMed]
- Majumdar, S.; Sarkar, R. Mutational and phylogenetic analyses of the two lineages of the Omicron variant. J. Med. Virol. 2022, 94, 1777–1779. [Google Scholar] [CrossRef] [PubMed]
- Desingu, P.A.; Nagarajan, K.; Dhama, K. Emergence of Omicron third lineage BA.3 and its importance. J. Med. Virol. 2022, 94, 1808–1810. [Google Scholar] [CrossRef] [PubMed]
- European Centre for Disease Prevention and Control. Epidemiological Update: SARS-CoV-2 Omicron Sub-Lineages BA.4 and BA.5. Available online: https://www.ecdc.europa.eu/en/news-events/epidemiological-update-sars-cov-2-omicron-sub-lineages-ba4-and-ba5 (accessed on 12 June 2022).
- Worl Health Organization. Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/activities/tracking-SARS-CoV-2-variants (accessed on 12 June 2022).
- Khateeb, J.; Li, Y.; Zhang, H. Emerging SARS-CoV-2 variants of concern and potential intervention approaches. Crit. Care 2021, 25, 244. [Google Scholar] [CrossRef]
- Callaway, E.; Ledford, H. How bad is Omicron? What scientists know so far. Nature 2021, 600, 197–199. [Google Scholar] [CrossRef]
- Pulliam, J.R.C.; van Schalkwyk, C.; Govender, N.; von Gottberg, A.; Cohen, C.; Groome, M.J.; Dushoff, J.; Mlisana, K.; Moultrie, H. Increased risk of SARS-CoV-2 reinfection associated with emergence of the Omicron variant in South Africa. medRxiv 2021. [Google Scholar] [CrossRef]
- Oran, D.P.; Topol, E.J. Prevalence of Asymptomatic SARS-CoV-2 Infection: A Narrative Review. Ann. Intern. Med. 2020, 173, 362–367. [Google Scholar] [CrossRef]
- Richardson, S.; Hirsch, J.S.; Narasimhan, M.; Crawford, J.M.; McGinn, T.; Davidson, K.W.; The Northwell COVID-19 Research Consortium; Barnaby, D.P.; Becker, L.B.; Chelico, J.D.; et al. Presenting Characteristics, Comorbidities, and Outcomes Among 5700 Patients Hospitalized With COVID-19 in the New York City Area. JAMA 2020, 323, 2052–2059. [Google Scholar] [CrossRef]
- Wu, S.Y.; Yau, H.S.; Yu, M.Y.; Tsang, H.F.; Chan, L.W.C.; Cho, W.C.S.; Shing Yu, A.C.; Yuen Yim, A.K.; Li, M.J.W.; Wong, Y.K.E.; et al. The diagnostic methods in the COVID-19 pandemic, today and in the future. Expert Rev. Mol. Diagn. 2020, 20, 985–993. [Google Scholar] [CrossRef]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Xia, J.; Tong, J.; Liu, M.; Shen, Y.; Guo, D. Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection. J. Med. Virol. 2020, 92, 589–594. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carter, L.J.; Garner, L.V.; Smoot, J.W.; Li, Y.; Zhou, Q.; Saveson, C.J.; Sasso, J.M.; Gregg, A.C.; Soares, D.J.; Beskid, T.R.; et al. Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Cent. Sci. 2020, 6, 591–605. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. COVID-19 Testing: What You Need to Know. Available online: https://www.cdc.gov/coronavirus/2019-ncov/symptoms-testing/testing.html (accessed on 12 June 2022).
- Kashir, J.; Yaqinuddin, A. Loop mediated isothermal amplification (LAMP) assays as a rapid diagnostic for COVID-19. Med. Hypotheses 2020, 141, 109786. [Google Scholar] [CrossRef] [PubMed]
- Khan, P.; Aufdembrink, L.M.; Engelhart, A.E. Isothermal SARS-CoV-2 Diagnostics: Tools for Enabling Distributed Pandemic Testing as a Means of Supporting Safe Reopenings. ACS Synth. Biol. 2020, 9, 2861–2880. [Google Scholar] [CrossRef]
- James, A.S.; Alawneh, J.I. COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2. Diagnostics 2020, 10, 399. [Google Scholar] [CrossRef]
- Gorzalski, A.J.; Tian, H.; Laverdure, C.; Morzunov, S.; Verma, S.C.; VanHooser, S.; Pandori, M.W. High-Throughput Transcription-mediated amplification on the Hologic Panther is a highly sensitive method of detection for SARS-CoV-2. J. Clin. Virol. 2020, 129, 104501. [Google Scholar] [CrossRef]
- Rahman, M.R.; Hossain, M.A.; Mozibullah, M.; Mujib, F.A.; Afrose, A.; Shahed-Al-Mahmud, M.; Apu, M.A.I. CRISPR is a useful biological tool for detecting nucleic acid of SARS-CoV-2 in human clinical samples. Biomed. Pharm. 2021, 140, 111772. [Google Scholar] [CrossRef]
- Vindeirinho, J.M.; Pinho, E.; Azevedo, N.F.; Almeida, C. SARS-CoV-2 Diagnostics Based on Nucleic Acids Amplification: From Fundamental Concepts to Applications and Beyond. Front. Cell Infect. Microbiol. 2022, 12, 799678. [Google Scholar] [CrossRef]
- Lee, E.Y.P.; Ng, M.Y.; Khong, P.L. COVID-19 pneumonia: What has CT taught us? Lancet Infect. Dis. 2020, 20, 384–385. [Google Scholar] [CrossRef]
- Sidiq, Z.; Hanif, M.; Dwivedi, K.K.; Chopra, K.K. Benefits and limitations of serological assays in COVID-19 infection. Indian J. Tuberc 2020, 67, S163–S166. [Google Scholar] [CrossRef]
- Karp, D.G.; Danh, K.; Espinoza, N.F.; Seftel, D.; Robinson, P.V.; Tsai, C.T. A serological assay to detect SARS-CoV-2 antibodies in at-home collected finger-prick dried blood spots. Sci. Rep. 2020, 10, 20188. [Google Scholar] [CrossRef] [PubMed]
- Van Kasteren, P.B.; van der Veer, B.; van den Brink, S.; Wijsman, L.; de Jonge, J.; van den Brandt, A.; Molenkamp, R.; Reusken, C.; Meijer, A. Comparison of seven commercial RT-PCR diagnostic kits for COVID-19. J. Clin. Virol. 2020, 128, 104412. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Rothman, R.E. PCR-based diagnostics for infectious diseases: Uses, limitations, and future applications in acute-care settings. Lancet Infect. Dis. 2004, 4, 337–348. [Google Scholar] [CrossRef]
- Deepak, S.; Kottapalli, K.; Rakwal, R.; Oros, G.; Rangappa, K.; Iwahashi, H.; Masuo, Y.; Agrawal, G. Real-Time PCR: Revolutionizing Detection and Expression Analysis of Genes. Curr. Genom. 2007, 8, 234–251. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Nucleic Acid Amplification Tests (NAATs). Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/naats.html (accessed on 15 June 2022).
- Arnaout, R.; Lee, R.A.; Lee, G.R.; Callahan, C.; Yen, C.F.; Smith, K.P.; Arora, R.; Kirby, J.E. SARS-CoV2 Testing: The Limit of Detection Matters. bioRxiv 2020. [Google Scholar] [CrossRef]
- Sharma, K.; Aggarwala, P.; Gandhi, D.; Mathias, A.; Singh, P.; Sharma, S.; Negi, S.S.; Bhargava, A.; Das, P.; Gaikwad, U.; et al. Comparative analysis of various clinical specimens in detection of SARS-CoV-2 using rRT-PCR in new and follow up cases of COVID-19 infection: Quest for the best choice. PLoS ONE 2021, 16, e0249408. [Google Scholar] [CrossRef]
- Medeiros da Silva, R.C.; Nogueira Marinho, L.C.; de Araujo Silva, D.N.; Costa de Lima, K.; Pirih, F.Q.; Luz de Aquino Martins, A.R. Saliva as a possible tool for the SARS-CoV-2 detection: A review. Travel Med. Infect. Dis. 2020, 38, 101920. [Google Scholar] [CrossRef]
- To, K.K.; Tsang, O.T.; Yip, C.C.; Chan, K.H.; Wu, T.C.; Chan, J.M.; Leung, W.S.; Chik, T.S.; Choi, C.Y.; Kandamby, D.H.; et al. Consistent Detection of 2019 Novel Coronavirus in Saliva. Clin. Infect. Dis. 2020, 71, 841–843. [Google Scholar] [CrossRef] [Green Version]
- Becherer, L.; Borst, N.; Bakheit, M.; Frischmann, S.; Zengerle, R.; von Stetten, F. Loop-mediated isothermal amplification (LAMP)—Review and classification of methods for sequence-specific detection. Anal. Methods 2020, 12, 717–746. [Google Scholar] [CrossRef] [Green Version]
- Zou, L.; Ruan, F.; Huang, M.; Liang, L.; Huang, H.; Hong, Z.; Yu, J.; Kang, M.; Song, Y.; Xia, J.; et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N. Engl. J. Med. 2020, 382, 1177–1179. [Google Scholar] [CrossRef]
- Shrestha, L.B.; Pokharel, K. Standard Operating Procedure for Specimen Collection, Packaging and Transport for Diagnosis of SARS-COV-2. JNMA J. Nepal Med. Assoc. 2020, 58, 627–629. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.F.; Yip, C.C.; To, K.K.; Tang, T.H.; Wong, S.C.; Leung, K.H.; Fung, A.Y.; Ng, A.C.; Zou, Z.; Tsoi, H.W.; et al. Improved Molecular Diagnosis of COVID-19 by the Novel, Highly Sensitive and Specific COVID-19-RdRp/Hel Real-Time Reverse Transcription-PCR Assay Validated In Vitro and with Clinical Specimens. J. Clin. Microbiol. 2020, 58, e00310-20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, D.K.W.; Pan, Y.; Cheng, S.M.S.; Hui, K.P.Y.; Krishnan, P.; Liu, Y.; Ng, D.Y.M.; Wan, C.K.C.; Yang, P.; Wang, Q.; et al. Molecular Diagnosis of a Novel Coronavirus (2019-nCoV) Causing an Outbreak of Pneumonia. Clin. Chem. 2020, 66, 549–555. [Google Scholar] [CrossRef] [Green Version]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brunink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, Y.W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. Laboratory Diagnosis of COVID-19: Current Issues and Challenges. J. Clin. Microbiol. 2020, 58, e00512-20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dudas, G.; Hong, S.L.; Potter, B.I.; Calvignac-Spencer, S.; Niatou-Singa, F.S.; Tombolomako, T.B.; Fuh-Neba, T.; Vickos, U.; Ulrich, M.; Leendertz, F.H.; et al. Emergence and spread of SARS-CoV-2 lineage B.1.620 with variant of concern-like mutations and deletions. Nat. Commun. 2021, 12, 5769. [Google Scholar] [CrossRef]
- Erster, O.; Beth-Din, A.; Asraf, H.; Levy, V.; Kabat, A.; Mannasse, B.; Azar, R.; Shifman, O.; Lazar, S.; Mandelboim, M.; et al. SPECIFIC DETECTION OF SARS-COV-2 B.1.1.529 (OMICRON) VARIANT BY FOUR RT-qPCR DIFFERENTIAL ASSAYS. medRxiv 2021. [Google Scholar] [CrossRef]
- Bustin, S.A.; Nolan, T. RT-qPCR Testing of SARS-CoV-2: A Primer. Int. J. Mol. Sci. 2020, 21, 3004. [Google Scholar] [CrossRef]
- Kralik, P.; Ricchi, M. A Basic Guide to Real Time PCR in Microbial Diagnostics: Definitions, Parameters, and Everything. Front. Microbiol. 2017, 8, 108. [Google Scholar] [CrossRef] [Green Version]
- Higuchi, R.; Dollinger, G.; Walsh, P.S.; Griffith, R. Simultaneous amplification and detection of specific DNA sequences. Nat. Biotechnol. 1992, 10, 413–417. [Google Scholar] [CrossRef]
- Tajadini, M.; Panjehpour, M.; Javanmard, S.H. Comparison of SYBR Green and TaqMan methods in quantitative real-time polymerase chain reaction analysis of four adenosine receptor subtypes. Adv. Biomed. Res. 2014, 3, 85. [Google Scholar] [CrossRef] [PubMed]
- VanGuilder, H.D.; Vrana, K.E.; Freeman, W.M. Twenty-five years of quantitative PCR for gene expression analysis. Biotechniques 2008, 44, 619–626. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leutenegger, C.M.; Boretti, F.S.; Mislin, C.N.; Flynn, J.N.; Schroff, M.; Habel, A.; Junghans, C.; Koenig-Merediz, S.A.; Sigrist, B.; Aubert, A.; et al. Immunization of cats against feline immunodeficiency virus (FIV) infection by using minimalistic immunogenic defined gene expression vector vaccines expressing FIV gp140 alone or with feline interleukin-12 (IL-12), IL-16, or a CpG motif. J. Virol. 2000, 74, 10447–10457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuji, S.; Iguchi, Y.; Shibata, N.; Teramura, I.; Kitagawa, T.; Yamanaka, H. Real-time multiplex PCR for simultaneous detection of multiple species from environmental DNA: An application on two Japanese medaka species. Sci. Rep. 2018, 8, 9138. [Google Scholar] [CrossRef] [Green Version]
- Elnifro, E.M.; Ashshi, A.M.; Cooper, R.J.; Klapper, P.E. Multiplex PCR: Optimization and application in diagnostic virology. Clin. Microbiol. Rev. 2000, 13, 559–570. [Google Scholar] [CrossRef]
- Engelmann, I.; Alidjinou, E.K.; Ogiez, J.; Pagneux, Q.; Miloudi, S.; Benhalima, I.; Ouafi, M.; Sane, F.; Hober, D.; Roussel, A.; et al. Preanalytical Issues and Cycle Threshold Values in SARS-CoV-2 Real-Time RT-PCR Testing: Should Test Results Include These? ACS Omega 2021, 6, 6528–6536. [Google Scholar] [CrossRef]
- Caraguel, C.G.; Stryhn, H.; Gagne, N.; Dohoo, I.R.; Hammell, K.L. Selection of a cutoff value for real-time polymerase chain reaction results to fit a diagnostic purpose: Analytical and epidemiologic approaches. J. Vet. Diagn. Invest. 2011, 23, 2–15. [Google Scholar] [CrossRef] [Green Version]
- Mouliou, D.S.; Gourgoulianis, K.I. False-positive and false-negative COVID-19 cases: Respiratory prevention and management strategies, vaccination, and further perspectives. Expert Rev. Respir. Med. 2021, 15, 993–1002. [Google Scholar] [CrossRef]
- Wolfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Muller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [Green Version]
- Younes, N.; Al-Sadeq, D.W.; Al-Jighefee, H.; Younes, S.; Al-Jamal, O.; Daas, H.I.; Yassine, H.M.; Nasrallah, G.K. Challenges in Laboratory Diagnosis of the Novel Coronavirus SARS-CoV-2. Viruses 2020, 12, 582. [Google Scholar] [CrossRef]
- Xu, J.; Wu, R.; Huang, H.; Zheng, W.; Ren, X.; Wu, N.; Ji, B.; Lv, Y.; Liu, Y.; Mi, R. Computed Tomographic Imaging of 3 Patients With Coronavirus Disease 2019 Pneumonia With Negative Virus Real-time Reverse-Transcription Polymerase Chain Reaction Test. Clin. Infect. Dis. 2020, 71, 850–852. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Habli, Z.; Saleh, S.; Zaraket, H.; Khraiche, M.L. COVID-19 in-vitro Diagnostics: State-of-the-Art and Challenges for Rapid, Scalable, and High-Accuracy Screening. Front. Bioeng. Biotechnol. 2020, 8, 605702. [Google Scholar] [CrossRef] [PubMed]

| S.No. | Name of Variant | Lineage | Earliest Sample | First Outbreak | Designated | Reference |
|---|---|---|---|---|---|---|
| 1. | Epsilon | B.1.429, B.1.427 | March 2020 | United States | 5 March 2021 | [57,58] |
| 2. | Zeta | P.2 | April 2020 | Brazil | 17 March 2021 | |
| 3. | Beta | B.1.351 | May 2020 | South Africa | 18 December 2020 | [59,60] |
| 4. | Lambda | C.37 | August 2020 | Peru | 14 June 2021 | [61,62,63] |
| 5. | Alpha | B.1.1.7 | September 2020 | United Kingdom | 18 December 2020 | [64,65] |
| 6. | Delta | B.1.617.2 | October 2020 | India | 11 May 2021 | [66,67] |
| 7. | Gamma | P.1 | November 2020 | Brazil | 11 January 2021 | [68,69] |
| 8. | Lota | B.1.526 | November 2020 | United States | 24 March 2021 | [70,71] |
| 9. | Eta | B.1.525 | December 2020 | Multiple Countries | 17 March 2021 | [72,73] |
| 10. | Kappa | B.1.617.1 | December 2020 | India | 4 April 2021 | [74,75] |
| 11. | Theta | P.3 | January 2021 | Philippines | 24 March 2021 | [76] |
| 12. | Mu | B.1.621 | January 2021 | Colombia | 30 August 2021 | [77,78] |
| 13. | B.1.1.318 | GR | January 2021 | Multiple Countries | 2 June 2021 | [79] |
| 14. | C.1.2 | GR | June 2021 | South Africa | 1 September 2021 | [80] |
| 15. | B.1.640 | GH/490R | September 2021 | Multiple Countries | 22 November 2021 | [81] |
| 16. | Omicron | BA.1 | November 2021 | South Africa | 26 November 2021 | [82,83] |
| 17. | Omicron | BA.2 | November 2021 | South Africa | 26 November 2021 | [84,85] |
| 18. | Omicron | BA.3 | November 2021 | South Africa | 26 November 2021 | [86] |
| 19. | Omicron | BA.4 | January 2022 | South Africa | 12 May 2022 | [87] |
| 20. | XD | Omicron BA.1 and Delta | January 2022 | France | 9 Mar, 2021 | [88] |
| 21. | Omicron | BA.5 | February 2022 | South Africa | 12 May 2022 | [87] |
| Test | Technique | Specimen | Advantages | Disadvantages | Reference |
|---|---|---|---|---|---|
| Viral test (Molecular genetics based) | |||||
| Antigen | Lateral flow immunoluminescent assay, single or double target | NPS and ANS | Rapid, point-of-care tests | Less sensitive, and chances of false positives | [98] |
| Nucleic acid | RT–qPCR | Saliva, NPS, nasal mid-turbinate and ANS | Sensitive, specific | Expensive, requires laboratory personnel, specialized lab equipment and reagents | [98] |
| Nucleic acid | Loop-mediated isothermal amplification (LAMP) | Saliva, urine, NPS, nasal mid-turbinate and ANS | Sensitive, specific, rapid | Complicated designing of assay, chances of false positives | [99,100] |
| Nucleic acid | Recombinase polymerase amplification (RPA) | NPS and ANS | Sensitive, specific, rapid | Complicated designing of assay, expensive | [101] |
| Nucleic acid | Nicking endonuclease amplification reaction (NEAR) | NPS and ANS | Sensitive, rapid | Chances of false negatives | [100,101] |
| Nucleic acid | Transcription mediated amplification (TMA) | NPS and ANS | Sensitive, specific | Expensive and less flexible | [102] |
| Nucleic acid | Helicase-dependent amplification (HDA) | NPS and ANS | Sensitive, rapid | Chances of false positives | [100] |
| Nucleic acid | Clustered regularly interspaced short palindromic repeats (CRISPR) | AN, OPl, NP wash/aspirate and BAL | Sensitive, specific, rapid, versatile | Target sequences of the Cas proteins are restricted; multiplexing can create interferences which may lead to cross-reactivities | [100,103] |
| Nucleic acid | Strand displacement amplification (SDA) | NPS and ANS | Rapid, sensitive | Reverse transcription of virus RNA is required, shortcomings of chosen isothermal method. | [104] |
| Volatile organic compounds (VOCs) | Rapid gas chromatography-mass spectrometry (GC-MS) | Breath | Rapid | Presumptive | [98] |
| Radiological abnormalities caused by viral infection | Computed Tomography | Cross-sectional images of patient’s chest | Non-invasive, lesser expensive | Less specific because imaging features overlap with other viral pneumonia | [105] |
| Serological/Immunological test | |||||
| Antibody | Enzyme-linked immunosorbent assay (ELISA) and chemiluminescent immunoassay (CIA) | Blood and tissue specimens | Rapid, point-of-care tests, can identify previous infection | Dependent on duration of infection, false-negative results | [106] |
| Antibody | Dried blood spot (DBS) | Dried blood samples pricked from fingers | Sensitive and rapid | Storage temperature sensitive | [107] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dutta, D.; Naiyer, S.; Mansuri, S.; Soni, N.; Singh, V.; Bhat, K.H.; Singh, N.; Arora, G.; Mansuri, M.S. COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnostics 2022, 12, 1503. https://doi.org/10.3390/diagnostics12061503
Dutta D, Naiyer S, Mansuri S, Soni N, Singh V, Bhat KH, Singh N, Arora G, Mansuri MS. COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnostics. 2022; 12(6):1503. https://doi.org/10.3390/diagnostics12061503
Chicago/Turabian StyleDutta, Debashis, Sarah Naiyer, Sabanaz Mansuri, Neeraj Soni, Vandana Singh, Khalid Hussain Bhat, Nishant Singh, Gunjan Arora, and M. Shahid Mansuri. 2022. "COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2" Diagnostics 12, no. 6: 1503. https://doi.org/10.3390/diagnostics12061503
APA StyleDutta, D., Naiyer, S., Mansuri, S., Soni, N., Singh, V., Bhat, K. H., Singh, N., Arora, G., & Mansuri, M. S. (2022). COVID-19 Diagnosis: A Comprehensive Review of the RT-qPCR Method for Detection of SARS-CoV-2. Diagnostics, 12(6), 1503. https://doi.org/10.3390/diagnostics12061503










