Comparative Evaluation of Multiplex Real-Time PCR, Standard Urine Culture, and Rapid Nephelometric Screening in Patients with Complicated Urinary Tract Infections
Abstract
1. Introduction
2. Materials and Methods
2.1. Population, Eligibility, and Unit of Analysis
2.2. Sample Collection
2.3. Standard Urine Culture
2.4. HB&L UROQUATTRO System (Alifax, Polverara (PD), Italy)
2.5. mPCR Analysis
2.6. Statistical Analysis
3. Results
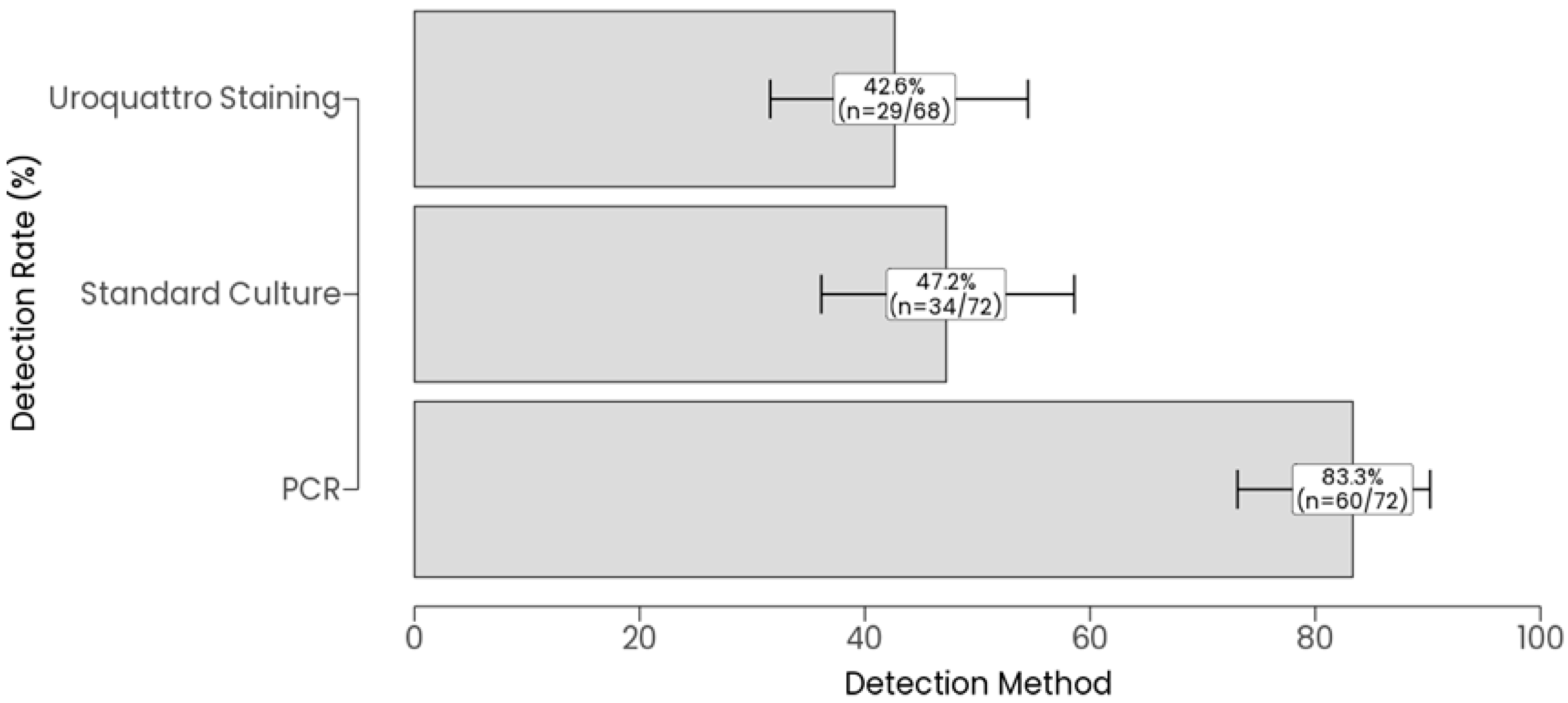
3.1. Patient Characteristics and Overall Detection Rates of All Methods
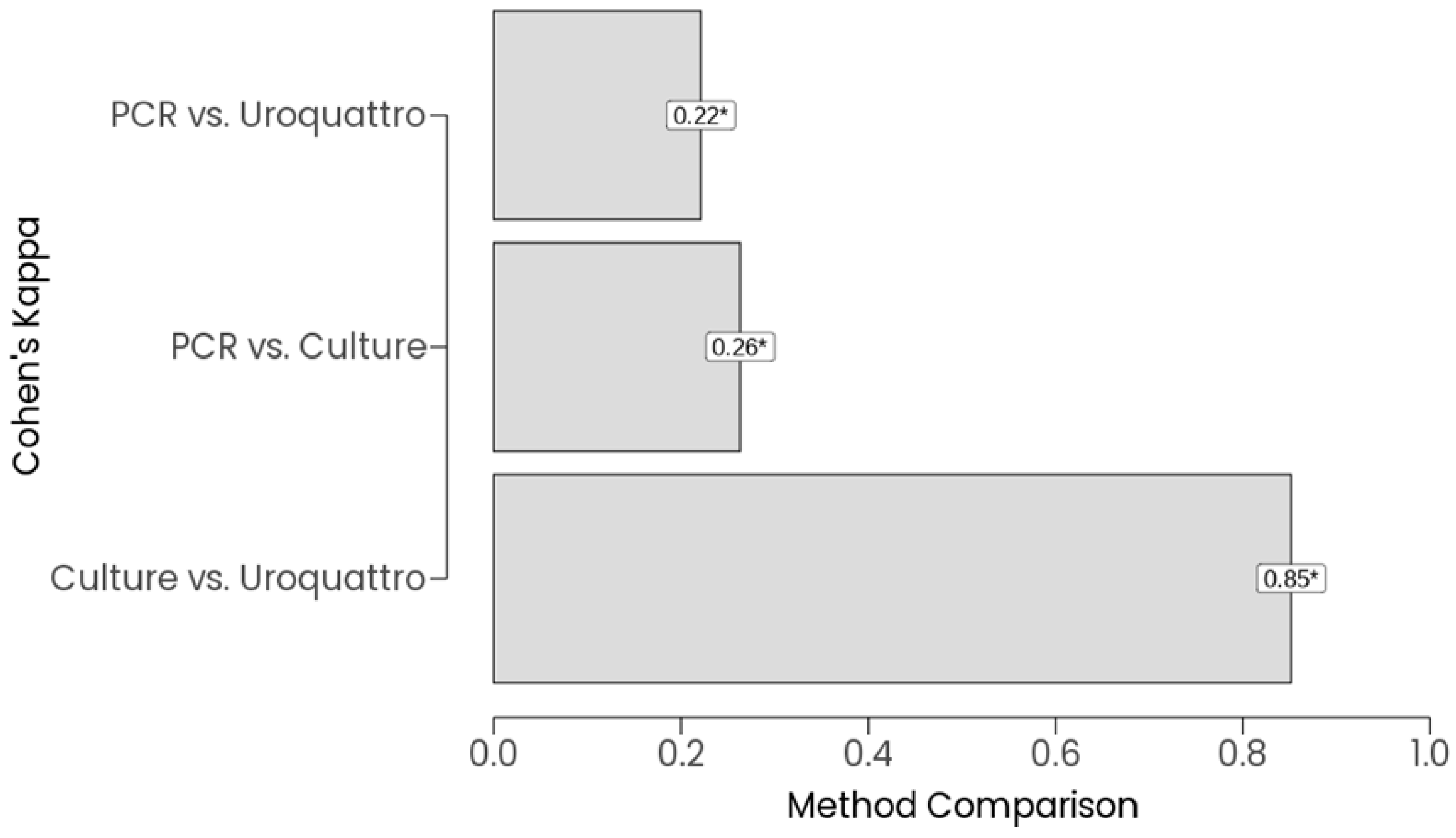
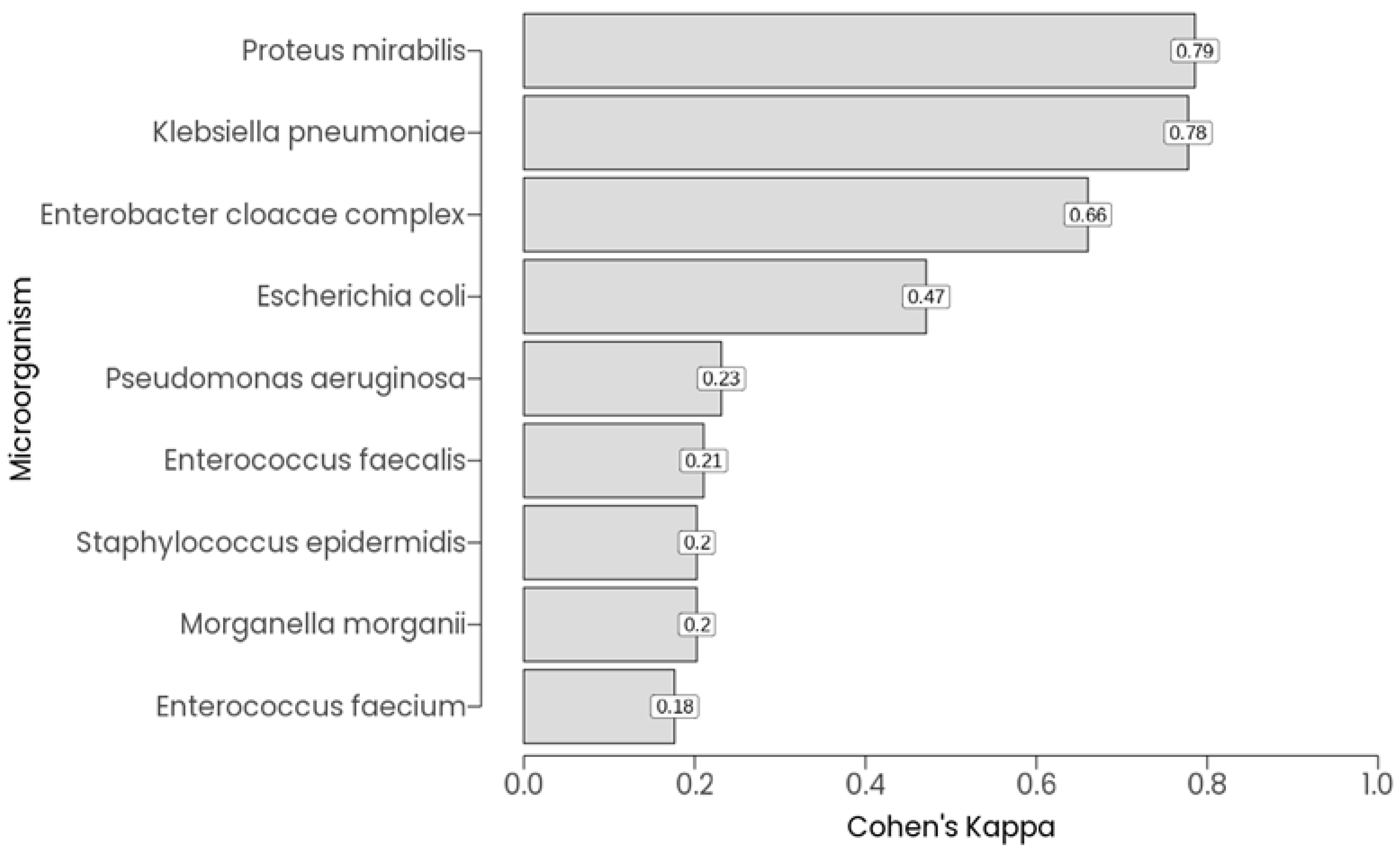
3.2. Microorganism Prevalence and Agreement

4. Discussion
4.1. Comparative Analysis of mPCR and SUC Detection Rates
4.2. Comparative Performance of Uroquattro Screening, Standard Culture, and mPCR in Diagnostic Workflows
4.3. Mechanistic Evaluation of Discordant Results and Analytical Limitations
4.4. Study Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| cUTI | Complicated Urinary Tract Infection |
| mPCR | Multiplex Real-Time PCR |
| SUC | Standard Urine Culture |
| EAU | European Association of Urology |
| IDSA | Infectious Diseases Society of America |
| MALDI-TOF | Matrix-Assisted Laser Desorption/Ionization Time-of-Flight |
| LoD | Limit of Detection |
| CFU/mL | Colony Forming Units per millilitre |
| EQUC | Extended Urine Culture protocols |
| NGS | Next-Generation Sequencing |
| Ct | Cycle threshold |
| WBC | White Blood Cells |
| RBC | Red Blood Cells |
| HGB | Hemoglobin |
| ESR | Erythrocyte Sedimentation Rate |
| CRP | C-reactive protein |
| IQR | Interquartile range |
| RIMU | Research Institute of Medical University of Plovdiv |
| IIST | Institute for Innovation and Smart Technology |
| EMB | Eosin-Methylene Blue |
| MuDT™ | Multiple Detection Temperatures |
| CI | Confidence Interval |
| κ (Kappa) | Cohen’s kappa |
| df | Degrees of freedom |
| ρ (Spearman’s ρ) | Spearman’s rank correlation coefficient |
| HIV | Human Immunodeficiency Virus |
| Cat. No. | Catalog Number |
| BA | Blood Agar |
| SD | Standard Deviation |
| r | Correlation coefficient |
| N/n | Total population size/Subsample size |
| μL | Microliter |
Appendix A
| mPCR Classification | Culture-Negative | Monomicrobial Culture | Polymicrobial Culture | Total |
|---|---|---|---|---|
| Negative | 11 | 1 | 0 | 12 |
| Monomicrobial | 6 | 7 | 0 | 13 |
| Polymicrobial | 21 | 22 | 4 | 47 |
| Total | 38 | 30 | 4 | 72 |
| SUC Positive | SUC Negative | Total | |
|---|---|---|---|
| mPCR positive | 33 (45.8%) | 27 (37.5%) | 60 (83.3%) |
| mPCR negative | 1 (1.4%) | 11 (15.3%) | 12 (16.7%) |
| Total | 34 (47.2%) | 38 (52.8%) | 72 (100%) |
| Panel | Target Organism | Gram Classification |
|---|---|---|
| Panel 1 | Escherichia coli | Gram-negative |
| Panel 1 | Klebsiella pneumoniae | Gram-negative |
| Panel 1 | Proteus mirabilis | Gram-negative |
| Panel 1 | Pseudomonas aeruginosa | Gram-negative |
| Panel 1 | Enterobacter spp. | Gram-negative |
| Panel 2 | Enterococcus faecalis | Gram-positive |
| Panel 2 | Enterococcus faecium | Gram-positive |
| Panel 2 | Staphylococcus aureus | Gram-positive |
| Panel 2 | Staphylococcus saprophyticus | Gram-positive |
| Panel 2 | Staphylococcus epidermidis | Gram-positive |
| Panel 2 | Streptococcus agalactiae (GBS) | Gram-positive |
| Panel 2 | Actinotignum (Actinobaculum) schaalii | Gram-positive |
| Panel 2 | Candida albicans | Fungal |
| Panel 2 | Candida spp. (non-albicans) | Fungal |
| Panel 3 | Acinetobacter baumannii | Gram-negative |
| Panel 3 | Aerococcus urinae | Gram-positive |
| Panel 3 | Citrobacter freundii | Gram-negative |
| Panel 3 | Citrobacter koseri | Gram-negative |
| Panel 3 | Corynebacterium urealyticum | Gram-positive |
| Panel 3 | Morganella morganii | Gram-negative |
| Panel 3 | Pantoea agglomerans | Gram-negative |
| Panel 3 | Providencia stuartii | Gram-negative |
| Panel 3 | Streptococcus anginosus | Gram-positive |
References
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary tract infections: Epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef]
- EAU Guidelines Office. EAU Guidelines on Urological Infections; European Association of Urology: Arnhem, The Netherlands, 2025. [Google Scholar]
- Medina, M.; Castillo-Pino, E. An introduction to the epidemiology and burden of urinary tract infections. Ther. Adv. Urol. 2019, 11, 1756287219832172. [Google Scholar] [CrossRef]
- Rupcheva, M.Y.; Kostadinov, K.; Kalchev, Y.; Gardzheva, P.; Hristozova, E.; Rachkovska, Z.; Lengerova, G.; Angelova, A.; Murdjeva, M.; Petrov, M.M. Urinary Tract Infections in a Single-Center Bulgarian Hospital: Trends in Etiology, Antibiotic Resistance, and the Impact of the COVID-19 Pandemic (2017–2022). Antibiotics 2025, 14, 982. [Google Scholar] [CrossRef]
- Bono, M.J.; Reygaert, W.C.; Stephens, E.L. Uncomplicated Urinary Tract Infections; StatPearls Publishing: Treasure Island, FL, USA, 2025. Available online: https://www.ncbi.nlm.nih.gov/books/NBK470195/ (accessed on 15 March 2026).
- Tandogdu, Z.; Wagenlehner, F.M. Global epidemiology of urinary tract infections. Curr. Opin. Infect. Dis. 2016, 29, 73–79. [Google Scholar] [CrossRef]
- Baimakhanova, B.; Sadanov, A.; Berezin, V.; Baimakhanova, G.; Trenozhnikova, L.; Orasymbet, S.; Seitimova, G.; Kalmakhanov, S.; Xetayeva, G.; Shynykul, Z.; et al. Emerging Technologies for the Diagnosis of Urinary Tract Infections: Advances in Molecular Detection and Resistance Profiling. Diagnostics 2025, 15, 2469. [Google Scholar] [CrossRef] [PubMed]
- Nelson, Z.; Aslan, A.T.; Beahm, N.P.; Blyth, M.; Cappiello, M.; Casaus, D.; Dominguez, F.; Egbert, S.; Hanretty, A.; Khadem, T. Guidelines for the Prevention, Diagnosis, and Management of Urinary Tract Infections in Pediatrics and Adults. JAMA Netw. Open 2024, 7, e2444495. [Google Scholar] [CrossRef] [PubMed]
- Price, T.K.; Dune, T.; Hilt, E.E.; Thomas-White, K.J.; Kliethermes, S.; Brincat, C.; Brubaker, L.; Wolfe, A.J.; Mueller, E.R.; Schreckenberger, P.C. The Clinical Urine Culture: Enhanced Techniques Improve Detection of Clinically Relevant Microorganisms. J. Clin. Microbiol. 2016, 54, 1216–1222. [Google Scholar] [CrossRef] [PubMed]
- Bank, S.; Jensen, A.; Hansen, T.M.; Søby, K.M.; Prag, J. Actinobaculum schaalii, a Common Uropathogen in Elderly Patients, Denmark. Emerg. Infect. Dis. 2010, 16, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.; Cognetti, M.; Patel, C.; Jean-Charles, N.; Tumati, A.; Burch-Smith, R.; Holton, M.; Kapoor, D.A. The Essential Role of PCR and PCR Panel Size in Comparison with Urine Culture in Identification of Polymicrobial and Fastidious Organisms in Patients with Complicated Urinary Tract Infections. Int. J. Mol. Sci. 2023, 24, 14269. [Google Scholar] [CrossRef] [PubMed]
- Hilt, E.E.; McKinley, K.; Pearce, M.M.; Rosenfeld, A.B.; Zilliox, M.J.; Mueller, E.R.; Brubaker, L.; Gai, X.; Wolfe, A.J.; Schreckenberger, P.C. Urine Is Not Sterile: Use of Enhanced Urine Culture Techniques To Detect Resident Bacterial Flora in the Adult Female Bladder. J. Clin. Microbiol. 2014, 52, 871–876. [Google Scholar] [CrossRef]
- Wolfe, A.J.; Brubaker, L. “Sterile Urine” and the Presence of Bacteria. Eur. Urol. 2015, 68, 173–174. [Google Scholar] [CrossRef]
- Ferreira, L.; Sánchez-Juanes, F.; González-Ávila, M.; Cembrero-Fuciños, D.; Herrero-Hernández, A.; González-Buitrago, J.M.; Muñoz-Bellido, J.L. Direct Identification of Urinary Tract Pathogens from Urine Samples by Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry. J. Clin. Microbiol. 2010, 48, 2110–2115. [Google Scholar] [CrossRef]
- John, G.; Mugnier, E.; Pittet, E.; Staehli, D.M.; Clerc, O.; Kenfak, A.F.; Konasch, A.; Lienhard, R.; Genné, D. Urinary culture sensitivity after a single empirical antibiotic dose for upper or febrile urinary tract infection: A prospective multicentre observational study. Clin. Microbiol. Infect. 2022, 28, 1099–1104. [Google Scholar] [CrossRef]
- Lefterova, M.I.; Suarez, C.J.; Banaei, N.; Pinsky, B.A. Next-Generation Sequencing for Infectious Disease Diagnosis and Management. J. Mol. Diagn. 2015, 17, 623–634. [Google Scholar] [CrossRef]
- Bagger, F.O.; Borgwardt, L.; Jespersen, A.S.; Hansen, A.R.; Bertelsen, B.; Kodama, M.; Nielsen, F.C. Whole genome sequencing in clinical practice. BMC Med. Genom. 2024, 17, 39. [Google Scholar] [CrossRef] [PubMed]
- Espy, M.J.; Uhl, J.R.; Sloan, L.M.; Buckwalter, S.P.; Jones, M.F.; Vetter, E.A.; Yao, J.D.C.; Wengenack, N.L.; Rosenblatt, J.E.; Cockerill, F.R., 3rd; et al. Real-Time PCR in Clinical Microbiology: Applications for Routine Laboratory Testing. Clin. Microbiol. Rev. 2006, 19, 165–256. [Google Scholar] [CrossRef] [PubMed]
- Szlachta-McGinn, A.; Douglass, K.M.; Chung, U.Y.R.; Jackson, N.J.; Nickel, J.C.; Ackerman, A.L. Molecular Diagnostic Methods Versus Conventional Urine Culture for Diagnosis and Treatment of Urinary Tract Infection: A Systematic Review and Meta-analysis. Eur. Urol. Open Sci. 2022, 44, 113–124. [Google Scholar] [CrossRef]
- Bermudez, T.; Schmitz, J.E.; Boswell, M.; Humphries, R. Novel technologies for the diagnosis of urinary tract infections. J. Clin. Microbiol. 2025, 63, e00306-24. [Google Scholar] [CrossRef]
- Kardjadj, M.; Priestly, I.P.; Chavez, R.; Derrick, D.; Huard, T.K. Clinical Symptom Resolution Following PCR-Guided vs. Culture and Susceptibility-Guided Management of Complicated UTI: How Time-To-Antibiotic Start and Antibiotic Appropriateness Mediate the Benefit of Multiplex PCR—An Ad Hoc Analysis of NCT06996301. Diagnostics 2025, 15, 3107. [Google Scholar] [CrossRef]
- Rader, T.S.; Stevens, M.P.; Bearman, G. Syndromic Multiplex Polymerase Chain Reaction (mPCR) Testing and Antimicrobial Stewardship: Current Practice and Future Directions. Curr. Infect. Dis. Rep. 2021, 23, 5. [Google Scholar] [CrossRef] [PubMed]
- Robledo, X.G.; Arcila, K.V.O.; Riascos, S.H.M.; García-Perdomo, H.A. Accuracy of molecular diagnostic techniques in patients with a confirmed urine culture: A systematic review and meta-analysis. Can. Urol. Assoc. J. 2022, 16, E484–E489. [Google Scholar] [CrossRef]
- Chang, Z.; Deng, J.; Zhang, J.; Wu, H.; Wu, Y.; Bin, L.; Li, D.; Liu, J.; Yu, R.; Lin, H.; et al. Rapid and Accurate Diagnosis of Urinary Tract Infections Using Targeted Next-Generation Sequencing: A Multicenter Comparative Study with Metagenomic Sequencing and Traditional Culture Methods. J. Infect. 2025, 90, 106459. [Google Scholar] [CrossRef]
- Queremel Milani, D.A.; Jialal, I. Urinalysis; StatPearls Publishing: Treasure Island, FL, USA, 2025. Available online: https://www.ncbi.nlm.nih.gov/books/NBK557685/ (accessed on 15 March 2026).
- Thermo Fisher Scientific. Challenging the UTI Testing Gold Standard: PCR vs. SUC. In Clinical Conversations; Thermo Fisher Scientific: Waltham, MA, USA, 2026; Available online: https://www.thermofisher.com/blog/clinical-conversations/challenging-uti-testing-gold-standard-pcr-vs-suc/ (accessed on 15 March 2026).
- Wojno, K.J.; Baunoch, D.; Luke, N.; Opel, M.; Korman, H.; Kelly, C.; Jaye, S.A.; Johnson, P.; Mallari, A.B.; Korman, A.B.; et al. Multiplex PCR Based Urinary Tract Infection (UTI) Analysis Compared to Traditional Urine Culture in Identifying Significant Pathogens in Symptomatic Patients. Urology 2020, 136, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, D.A.; Holton, M.R.; Hafron, J.; Aljundi, R.; Zwaans, B.; Hollander, M. Comparison of Polymerase Chain Reaction and Urine Culture in the Evaluation of Patients with Complex Urinary Tract Infections. Biology 2024, 13, 257. [Google Scholar] [CrossRef]
- Haley, E.; Luke, N.; Mathur, M.; Festa, R.A.; Wang, J.; Jiang, Y.; Anderson, L.; Baunoch, D. Comparison Shows that Multiplex Polymerase Chain Reaction Identifies Infection-associated Urinary Biomarker–positive Urinary Tract Infections That Are Missed by Standard Urine Culture. Eur. Urol. Open Sci. 2023, 58, 73–81. [Google Scholar] [CrossRef]
- Kline, K.A.; Lewis, A.L. Gram-Positive Uropathogens, Polymicrobial Urinary Tract Infection, and the Emerging Microbiota of the Urinary Tract. Microbiol. Spectr. 2016, 4, UTI-0012-2012. [Google Scholar] [CrossRef]
- Grigoryan, L.; Trautner, B.W.; Gupta, K. Diagnosis and Management of Urinary Tract Infections in the Outpatient Setting. JAMA 2014, 312, 1677–1684. [Google Scholar] [CrossRef]
- Lotte, R.; Lotte , L.; Ruimy, R. Actinotignum schaalii (formerly Actinobaculum schaalii): A newly recognized pathogen—Review of the literature. Clin. Microbiol. Infect. 2016, 22, 28–36. [Google Scholar] [CrossRef]
- Lleo, M.M.; Ghidini, V.; Tafi, M.C.; Castellani, F.; Trento, I.; Boaretti, M. Detecting the presence of bacterial DNA by PCR can be useful in diagnosing culture-negative cases of infection, especially in patients with suspected infection and antibiotic therapy. FEMS Microbiol. Lett. 2014, 354, 153–160. [Google Scholar] [CrossRef]
- Lehmann, L.E.; Hauser, S.; Malinka, T.; Klaschik, S.; Stüber, F.; Book, M. Real-time polymerase chain-reaction detection of pathogens is feasible to supplement the diagnostic sequence for urinary tract infections. BJU Int. 2009, 106, 114–120. [Google Scholar] [CrossRef] [PubMed]
- Chang, Z.; Chen, J.; Wang, Z.; Liang, Y.; She, S.; Wang, S.; Huang, X.; Wu, H.; Han, X.; Luo, L.; et al. Rapid and accurate preoperative diagnosis of urinary tract infections in urolithiasis patients: Clinical validation of the HB&L system. BMC Infect. Dis. 2026, 26, 247. [Google Scholar] [CrossRef] [PubMed]
- Davenport, M.; Mach, K.E.; Shortliffe, L.M.D.; Banaei, N.; Wang, T.-H.; Liao, J.C. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 2017, 14, 296–310. [Google Scholar] [CrossRef]
| Characteristic | Overall (N = 72) | No Detection (N = 11) | Detected (N = 61) | p-Value |
|---|---|---|---|---|
| Age (years) | 58 (40, 71) | 58 (25, 71) | 58 (42, 70) | >0.9 |
| Female | 53 (74%) | 9 (82%) | 44 (72%) | 0.7 |
| Diagnosis | 0.4 | |||
| Acute Pyelonephritis | 60 (83%) | 8 (73%) | 52 (85%) | |
| Acute Cystitis | 12 (17%) | 3 (27%) | 9 (15%) | |
| Symptoms | ||||
| Cloudiness | 49 (68%) | 6 (55%) | 43 (70%) | 0.3 |
| Febrile | 24 (33%) | 4 (36%) | 20 (33%) | >0.9 |
| Vomiting | 10 (14%) | 1 (9.1%) | 9 (15%) | >0.9 |
| Dysuria | 58 (81%) | 9 (82%) | 49 (80%) | >0.9 |
| Pollakiuria | 49 (68%) | 6 (55%) | 43 (70%) | 0.3 |
| Urgency | 22 (31%) | 5 (45%) | 17 (28%) | 0.3 |
| Enuresis | 1 (1.4%) | 0 (0%) | 1 (1.6%) | >0.9 |
| Haematuria | 4 (5.6%) | 1 (9.1%) | 3 (4.9%) | 0.5 |
| Suprapubic Pain | 29 (40%) | 5 (45%) | 24 (39%) | 0.7 |
| Succussion | 20 (28%) | 3 (27%) | 17 (28%) | >0.9 |
| Smell | 20 (28%) | 2 (18%) | 18 (30%) | 0.7 |
| Urine Colour Change | 59 (82%) | 8 (73%) | 51 (84%) | 0.4 |
| Congenital Abnormalities | 10 (14%) | 0 (0%) | 10 (16%) | 0.3 |
| Obstruction | 6 (8.3%) | 2 (18%) | 4 (6.6%) | 0.2 |
| Laboratory Values | ||||
| WBC (×109/L) | 8.7 (6.5, 10.8) | 9.5 (7.8, 11.1) | 8.6 (6.5, 10.8) | 0.5 |
| RBC (×1012/L) | 4.41 (3.95, 4.83) | 4.40 (3.93, 4.77) | 4.41 (3.97, 4.87) | >0.9 |
| HGB (g/L) | 125 (112, 137) | 128 (111, 140) | 124 (112, 135) | 0.5 |
| ESR (mm/h) | 26 (14, 40) | 22 (7, 37) | 27 (15, 40) | 0.5 |
| CRP (mg/L) | 8 (5, 31) | 10 (6, 87) | 8 (4, 31) | 0.4 |
| Urea (mmol/L) | 6.3 (4.3, 8.6) | 6.5 (4.1, 9.7) | 6.1 (4.3, 8.2) | 0.8 |
| Urine Specific Gravity | 1.015 (1.010, 1.025) | 1.015 (1.010, 1.020) | 1.015 (1.010, 1.025) | 0.4 |
| Urine pH | 5.50 (5.00, 6.00) | 5.50 (5.00, 6.00) | 5.50 (5.00, 6.00) | 0.8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Rupcheva, M.Y.; Kostadinov, K.; Tashev, R.; Markova, P.; Zheleva, V.; Chterev, M.; Atanasova, M.; Petrov, M.M.; Murdjeva, M. Comparative Evaluation of Multiplex Real-Time PCR, Standard Urine Culture, and Rapid Nephelometric Screening in Patients with Complicated Urinary Tract Infections. Diagnostics 2026, 16, 919. https://doi.org/10.3390/diagnostics16060919
Rupcheva MY, Kostadinov K, Tashev R, Markova P, Zheleva V, Chterev M, Atanasova M, Petrov MM, Murdjeva M. Comparative Evaluation of Multiplex Real-Time PCR, Standard Urine Culture, and Rapid Nephelometric Screening in Patients with Complicated Urinary Tract Infections. Diagnostics. 2026; 16(6):919. https://doi.org/10.3390/diagnostics16060919
Chicago/Turabian StyleRupcheva, Milena Yancheva, Kostadin Kostadinov, Radoslav Tashev, Petya Markova, Violeta Zheleva, Maritza Chterev, Mariya Atanasova, Michael M. Petrov, and Marianna Murdjeva. 2026. "Comparative Evaluation of Multiplex Real-Time PCR, Standard Urine Culture, and Rapid Nephelometric Screening in Patients with Complicated Urinary Tract Infections" Diagnostics 16, no. 6: 919. https://doi.org/10.3390/diagnostics16060919
APA StyleRupcheva, M. Y., Kostadinov, K., Tashev, R., Markova, P., Zheleva, V., Chterev, M., Atanasova, M., Petrov, M. M., & Murdjeva, M. (2026). Comparative Evaluation of Multiplex Real-Time PCR, Standard Urine Culture, and Rapid Nephelometric Screening in Patients with Complicated Urinary Tract Infections. Diagnostics, 16(6), 919. https://doi.org/10.3390/diagnostics16060919






