Simple Summary
Opsin contributes to the phototaxis of Asian citrus psyllid (ACP, Diaphorina citri), which is the primary vector of Candidatus Liberibacter asiaticus, causing Citrus Huanglongbing (HLB). The objective of this study was to explore the phototransduction pathway of D. citri with a view to identifying elicitors that could enhance the phototaxis of D. citri in the future. To that end, 87 genes were identified via transcriptome sequencing. Of these genes, 71 were linked to the phototransduction-fly pathway. In addition, when LW-opsin gene expression was inhibited, Gqβ and ninaC were down-regulated. The findings of our study provide a foundation for further research into the phototransduction pathway in ACP, as well as a basis for further investigation into the potential of elicitors to enhance the phototaxis of ACP.
Abstract
Opsin plays a regulatory role in phototaxis of Diaphorina citri, functioning as the initial station in the phototransduction cascade. Our study aimed to explore the D. citri phototransduction pathway to identify elicitors that may enhance D. citri phototaxis in the future. The RNAi technique was employed to inhibit LW-opsin gene expression, followed by RNA-Seq analysis to identify phototransduction genes. Finally, RT-qPCR was performed to validate whether genes in the phototransduction pathway were affected by the inhibition of LW-opsin expression. A total of 87 genes were identified within the transcriptome as involved in phototransduction based on KEGG functional annotation. Of these, 71 genes were identified as enriched in the phototransduction-fly pathway. These genes encode key proteins in this process, including Gqα, Gqβ, Gqγ, phospholipase C β (PLCβ), the cation channel transient receptor potential (TRP), and TRP-like (TRPL), among others. Moreover, the LOC103513214 (Gqβ) and LOC103518375 (ninaC) genes exhibited reduced expression when LW-opsin gene expression was suppressed. Our results provide a basis for further investigation of phototransduction in D. citri.
1. Introduction
Diaphorina citri Kuwayama (Asian citrus psyllid, Hemiptera, Psyllidae) is one of the most significant pests of citrus. It is the primary vector of Candidatus Liberibacter asiaticus, the causal agent of Citrus Huanglongbing (HLB) [1,2]. HLB is a destructive disease, often referred to as the “cancer of the citrus industry”, which represents a substantial threat to the global citrus industry and poses a considerable risk to the industry’s sustainability [3,4,5]. The most effective method for controlling HLB is the management of the Asian citrus psyllid through chemical, biological, and physical control measures. Among these, physical control, which leverages the insect’s phototactic behavior, is widely employed in field production due to its high efficacy and environmental compatibility. The Asian citrus psyllid displays positive phototaxis, exhibiting a preference for ultraviolet, blue, and green light [6,7,8]. It has been demonstrated that opsin regulates phototaxis in ACP [8]; however, the underlying molecular mechanism is not fully understood.
Opsin is a member of the G-protein-coupled receptor family, comprising seven transmembrane domains [9,10]. Based on the type of photoreceptor cell, opsins are classified into two major groups: rhabdomeric-type opsin (r-opsin) and ciliary-type opsin (c-opsin) [11,12,13]. In general, r-opsin is the main visual pigment in the rhabdomeres of insect compound eyes. Based on the spectral range to which opsins are sensitive, r-opsins of insects are classified into three categories: (1) long-wavelength-sensitive opsin (LW-opsin) with the peak absorbance more than 500 nm; (2) blue-sensitive opsin (BL-opsin) with the peak absorbance at 400–500 nm; and (3) ultraviolet-sensitive opsin (UV-opsin) with the peak absorbance at 300–400 nm [14,15,16]. Drosophila has 7 opsins (Rh1–7), of which Rh1, Rh2, and Rh5 are sensitive to blue light (absorption maxima of 486, 418, and 442 nm), Rh3 and Rh4 are sensitive to UV light (331 and 355 nm), and Rh6 is sensitive to green light (515 nm) [17,18,19,20]. Rh7 functions as a light detector and is involved in the regulation of the circadian rhythm [21]. It should be noted that different insect species vary in the number of opsins that they possess. For example, BL-opsin gene loss occurs in some species, such as Tribolium castaneum (red flour beetle, Coleoptera: Tenebrionidae), aphids (Aphidoidea), and planthoppers (Fulgoroidea) [22,23].
Opsins form covalent bonds with small molecular chromophores, thereby creating light-sensitive photopigments known as rhodopsins, which act as the primary activators in the phototransduction cascade [24]. The phototransduction pathway in Drosophila is most intensively studied. Light activates rhodopsin into metarhodopsin with chromophore conformational change, which initiates the G protein-mediated phototransduction cascade, leading to calcium and sodium ion exchange and the translation of light stimulus into an electrical signal [25,26,27]. A mutation in LW-opsin has been shown to alter gene expression in the phototransduction pathway in Plutella xylostella (diamondback moth, Lepidoptera: Plutellidae) [28]. Gene expression associated with the phototransduction pathway in the head of Mythimna separata (oriental armyworm, Lepidoptera: Noctuidae) was significantly altered following exposure to various light environments [29]. Thus, it was hypothesized that genes within the phototransduction pathway may enhance the phototaxis of the Asian citrus psyllid, making it essential to understand this pathway in D. citri.
Four visual photo-sensing opsins were identified in ACP: Dc-UV (UV-opsin), clustering with Drosophila Rh3 and Rh4; Dc-BW (BL-opsin), clustering with Rh5; Dc-LW (LW-opsin), clustering with Rh6; and Dc-UV-like, clustering with Rh7 [8]. The inhibition of UV-opsin, BL-opsin, and LW-opsin resulted in a reduction in phototactic response rates to the corresponding light source in ACP [8]. The expression of LW-opsin was observed to be the highest in ACP adults, in comparison to the other opsins [8]. Furthermore, only the expression of LW-opsin was found to be significantly up-regulated in response to white light treatment in comparison to dark treatment in our previous study (data not supplied). Accordingly, LW-opsin was selected as the target gene for further experimentation. To investigate the phototransduction pathway in D. citri, we first employed the RNA interference (RNAi) technique to inhibit LW-opsin gene expression, followed by RNA-Seq analysis to investigate the complete set of genes involved in phototransduction. Ultimately, inhibition of LW-opsin expression was observed to influence the mRNA expression levels of specific phototransduction genes, as determined by real-time quantitative PCR (RT-qPCR). This study advances our understanding of the phototransduction pathway in D. citri and provides theoretical guidance for identifying elicitors that could enhance D. citri phototaxis in the future.
2. Materials and Methods
2.1. Insects
The D. citri were reared on healthy Murraya paniculata within insect-rearing cages (60 × 40 × 75 cm) in an artificial greenhouse at the Institute of Plant Protection, Fujian Academy of Agricultural Sciences. The greenhouse was maintained under controlled conditions: temperature 26 ± 1 °C, relative humidity 60% ± 10%, and a 14:10 h (light:dark) photoperiod. To ensure a uniform age of the insects, newly molted adults were collected and placed in a cage (30 × 30 × 35 cm) with a plant for further experiments.
2.2. RNA Extraction and dsRNA Synthesis
The total RNA was extracted using the FreeZol reagent (Vazyme, Nanjing, China), purified with RNA Extraction Buffer (Beyotime, Shanghai, China), and precipitated with isopropanol. Then, 500 ng of total RNA was reverse transcribed into complementary DNA (cDNA) using the HiScript® II 1st Strand cDNA Synthesis Kit (Vazyme), following the manufacturer’s protocol. The primers for the synthesis of double-stranded RNA (dsRNA) of LW-opsin and EGFP were designed according to Li et al. [8] and were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China). The sequence fragment templates of LW-opsin and EGFP were amplified using the Phanta® Super-Fidelity DNA Polymerase (Vazyme), according to the manufacturer’s protocol. The PCR products were purified using a FastPure® Gel DNA Extraction Mini Kit (Vazyme), following the manufacturer’s protocol. A total of 2 μg of purified PCR product was employed for dsRNA synthesis in vitro using the T7 Ribomax™ Express large-scale production system (Promega, Madison, WI, USA), according to the manufacturer’s protocol.
2.3. RNA Interference of LW-Opsin
Injections of 200 ng of dsRNA were given to 10-day-old female D. citri adults using a Nanoject III microsyringe (Drummond Scientific Company, Broomall, PA, USA). After injection, the insects were reared on healthy M. paniculata in insect-rearing cages under the same conditions previously described. A total of 30 individuals were collected at 12, 24, 36, and 72 h post-injection, respectively. The dsLW-opsin and dsEGFP treatments were performed in triplicate at each of the four time points. Subsequently, the expression levels of LW-opsin were quantified via RT-qPCR to confirm the efficacy of the RNAi. The RT-qPCR amplification was conducted using the GoTaq® qPCR Master Mix (Promega) with an initial step at 95 °C for 10 min, followed by 40 cycles at 95 °C for 15 s and 60 °C for 30 s. A final cycle was carried out at 95 °C for 15 s, 60 °C for 60 s, and 95 °C for 15 s. The gene β-actin of D. citri was employed as the reference gene. The primers for RT-qPCR were designed following Li et al. [8], and they were synthesized by Sangon Biotech Co.Ltd. (Shanghai, China). The comparative Ct method (2−ΔΔCT) was employed for the calculation of the transcript level [30].
2.4. Transcriptome Sequencing
RNA-Seq was conducted on samples collected at time points where LW-opsin expression levels were suppressed. The RNA integrity number (RIN) was assessed using an RNA 6000 Nano Lab Chip Kit and Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). Samples with an RIN value of 8.0 or higher were submitted to the Shanghai Meiji Biological Company (Shanghai, China) for library construction and sequencing using the Illumina Novaseq 6000 platform (San Diego, CA, USA). Adapter trimming and low-quality filtering of the raw reads were performed using fastp (https://github.com/OpenGene/fastp, accessed on 18 July 2024). The clean reads were then aligned to the D. citri genome (https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_000475195.1/, accessed on 18 July 2024) using Bowtie2 (Version 2.4.1) and HISAT2 (http://ccb.jhu.edu/software/hisat2/index.shtml, accessed on 18 July 2024) for sequence analysis. The mapped reads were assembled and spliced with StringTie (http://ccb.jhu.edu/software/stringtie/, accessed on 18 July 2024) in accordance with the D. citri genome. New transcripts were identified by comparing them with known transcripts using the Gffcompare tool (Version 0.9.8).
2.5. Gene Function Annotation and Phototransduction Gene Enrichment
A BLASTx search was conducted in protein databases, including NR (non-redundant), Swiss-Prot, Pfam, EggNOG, GO (Gene Ontology), and KEGG (Kyoto Encyclopedia of Genes and Genomes), to determine functional annotation.
A comprehensive search of the transcriptome was performed with the keywords “phototransduction” to identify all genes related to the phototransduction pathway, based on the functional annotation of all protein databases described above. Subsequently, an enrichment analysis of the KEGG pathway was conducted via the Majorbio platform (https://www.majorbio.com/web/www/index, accessed on 31 July 2024). The annotation of the genes’ functions within the most enriched pathway was analyzed further.
2.6. Validation of Phototransduction Gene Expression by RT-qPCR
The objective was to validate whether the inhibition of LW-opsin expression affects the expression of genes involved in the phototransduction pathway. Accordingly, a subset of phototransduction genes was selected for further analysis. Primers for the selected genes are listed in Table 1. The RT-qPCR amplification was conducted following the previously described protocol.

Table 1.
Primers used to validate the expression of phototransduction genes affected by inhibition of LW-opsin expression.
2.7. Statistical Analysis
All statistical analyses were performed using the IBM SPSS software (version 22.0, IBM, Armonk, NY, USA). An independent t-test was used to assess significant differences in LW-opsin gene expression or phototransduction gene expression between the dsEGFP and dsLW-opsin treatment groups.
3. Results
3.1. RNA Interference of LW-Opsin
Following the injection of dsRNA (dsLW-opsin and dsEGFP), the relative expression levels of LW-opsin were quantified through RT-qPCR (Figure 1). The expression of LW-opsin had decreased by 46.82% at the 36 h time point and by 38.65% at the 72 h time point. Thus, the 36 h time point was considered optimal for conducting further RNA-Seq analysis.
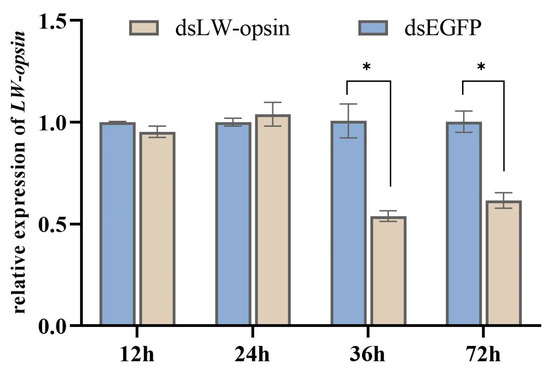
Figure 1.
The interference efficiency of RNAi at the transcription level of LW-opsin. * indicates p < 0.05.
3.2. Illumina Sequencing Analysis
A total of 40.97 Gb of clean data was obtained through transcriptome sequencing, with all samples exceeding 6.33 Gb of clean data. A total of 43,268 expressed genes were detected, comprising 23,604 known genes and 19,664 novel genes. In this paper, novel genes and novel transcripts are designated as “MSTRG”. Additionally, 83,232 transcripts were characterized, including 25,321 known transcripts and 57,911 novel transcripts. Approximately 74.38% of the high-quality clean data was retained for assembly and analysis. Furthermore, the percentage of Q30 bases exceeded 95.68% (Supplementary Data File S1), indicating a high level of base accuracy. The sequences of all transcripts were provided as supplementary material (Supplementary Data File S2).
3.3. Phototransduction Genes
A total of 87 genes, including 9 novel genes, were identified within the transcriptome as involved in phototransduction, based on KEGG functional annotation (Table 2). The 87 genes were assigned 19 distinct KO IDs. Of the 87 genes identified, 17 were associated with K08834 (MYO3, myosin III), 15 with K02183 (CALM, calmodulin), 10 with K04255 (Rh2_7, r-opsin), and 6 with K00910 (GRK, G-protein-dependent receptor kinase). Four genes were assigned to K04967 (TRPC4, transient receptor potential cation channel subfamily C member 4), four genes to K05858 (PLCB, phosphatidylinositol phospholipase C, beta), three genes to K02677 (PRKCA, classical protein kinase C alpha type), three genes to K04515 (CAMK2, calcium/calmodulin-dependent protein kinase II), and three genes to K04952 (CNGB1, cyclic nucleotide gated channel beta 1). Two genes belong to K04536 (GNB1, guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1), two to K13803 (TRPL, transient-receptor-potential-like protein), two genes to K13806 (DAGL, sn1-specific diacylglycerol lipase), one to K04547 (GNG13, guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-13), and one to K04634 (GNAQ, guanine nucleotide-binding protein G(q) subunit alpha). Additionally, one gene belongs to K04958 (ITPR1, inositol 1,4,5-triphosphate receptor type 1), one to K07972 (GNB, guanine nucleotide-binding protein subunit beta), one to K12322 (GUCY2F, guanylate cyclase 2F), and one to K13805 (ARR2, arrestin2).

Table 2.
Genes involved in phototransduction identified through KEGG functional annotation.
3.4. KEGG Pathway Enrichment of Phototransduction Genes
The phototransduction-fly (map04745, rich factor = 1), phototransduction (map04744, rich factor = 1), and circadian entrainment (map04713, rich factor = 0.26) pathways, which are associated with light perception, were significantly enriched (Figure 2, Supplementary Data File S3). A total of 71 genes were enriched in the phototransduction-fly pathway, 21 genes were enriched in the phototransduction pathway, and 30 genes were enriched in the circadian entrainment pathway.
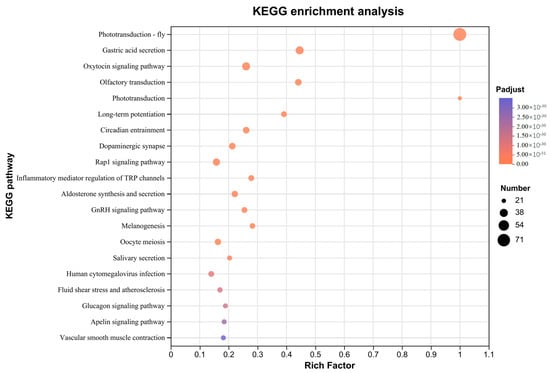
Figure 2.
Top 20 of KEGG pathway enrichment of phototransduction genes.
3.5. Gene Function Annotation Based on Phototransduction-Fly Pathway
A total of 71 genes were found to function in accordance with the phototransduction-fly pathway, as illustrated in Figure 3 and detailed in Table 3. The genes LOC103516584, LOC103513214, and LOC103507998 were identified as encoding the α-, β-, and γ-subunits of the heterotrimeric Gq protein, respectively. Four genes were identified as encoding phospholipase C β (PLCβ), four as encoding the cation channel TRP (transient receptor potential), and two as encoding the cation channel TRPL (transient receptor potential like). Three genes were identified as encoding protein kinase C (PKC), fifteen as encoding CaM (calmodulin), and one as encoding Arr2 (arrestin2). Seventeen genes were identified as encoding NINAC (neither inactivation nor afterpotential C), ten as encoding actin, and one as encoding IP3R (inositol 1,4,5-trisphosphate, InsP3). Two genes were identified as encoding DAGL (diacylglycerol lipase), three as encoding CaMKII (CaM kinase II), and six as encoding RK (G protein-coupled receptor kinase 1). To confirm the precise details of these genes, further phylogenetic analysis, gene cloning, and characterization will be required.
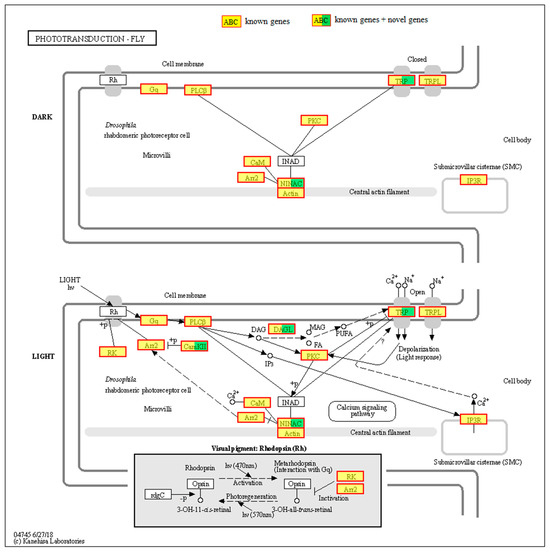
Figure 3.
Gene function annotation based on phototransduction-fly pathway. Rh, rhodopsin. Gq, heterotrimeric Gq protein. PLCβ, phospholipase C β. TRP, transient receptor potential. TRPL, transient receptor potential like. PKC, protein kinase C. CaM, calmodulin. Arr2, arrestin2. NINAC, neither inactivation nor afterpotential C. IP3R, inositol 1,4,5-trisphosphate. DAGL, diacylglycerol lipase. CaMKII, CaM kinase II. RK, G protein-coupled receptor kinase 1.

Table 3.
Genes associated with the phototransduction-fly pathway.
It is noteworthy that ten genes (LOC103506067, LOC103506845, LOC103516743, LOC103516911, LOC103516912, LOC103517452, LOC103517454, LOC103521071, LOC103524061, LOC113473058) classified within the seven-transmembrane G protein-coupled receptor superfamily were not identified as opsins. Additionally, INAD (inactivation nor afterpotential D) and rdgC (retinal degeneration C) were not identified in our transcriptome.
3.6. Knockdown of LW-Opsin Gene Changes the Gene Expression of the Phototransduction Pathway
A total of nine genes were selected for validation to determine whether the knockdown of the LW-opsin gene alters the expression of genes involved in the phototransduction pathway. The results demonstrated that only the LOC103513214 (Gqβ) and LOC103518375 (ninaC) genes exhibited a reduction in expression when LW-opsin gene expression was inhibited (Figure 4).
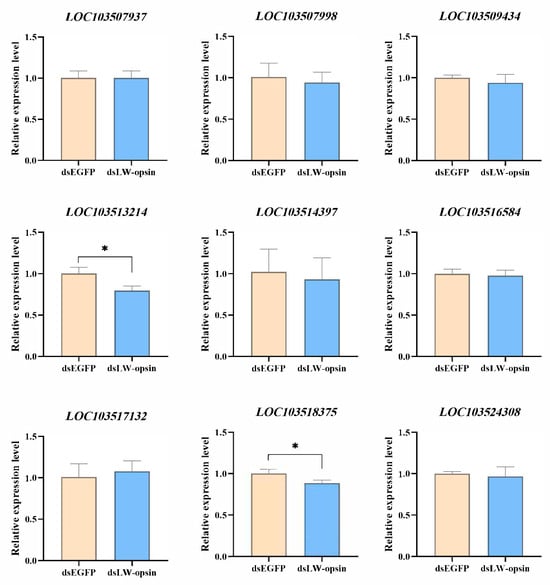
Figure 4.
Verification of the 9 phototransduction genes whose expression is influenced by the knockdown of LW-opsin gene expression. * indicates p < 0.05.
4. Discussion
The Drosophila phototransduction pathway (phototransduction-fly) has been extensively studied [13,24,25,26,27]. In this study, a total of 71 genes in D. citri were identified as enriched in the phototransduction-fly pathway, including Gqα, Gqβ, Gqγ, Arr2, PLCβ, TRP, and TRPL, among others, with the exception of INAD and rdgC. This suggests that the phototransduction pathway in D. citri is comparable to that observed in Drosophila, although potential variations cannot be ruled out. Furthermore, the phototransduction pathway in non-Drosophila insects has been investigated through transcriptome-sequencing techniques. For example, the majority of phototransduction genes were found to be conserved between D. melanogaster and Lepidoptera, including Gqα, Gqβ, Gqγ, Arr1, Arr2, trp, trpl, and others [31]. In Ptomaphagus hirtus (troglobiont beetle, Coleoptera: Leiodidae), approximately 20 genes have been identified, including LW-opsin, arr1, arr2, trp, and trp [32]. Moreover, studies have demonstrated that gene expression in the phototransduction pathway is altered by environmental stimuli. For instance, eight genes, including arr2, BRh, and rdgC, have been demonstrated to exhibit seasonal adaptations in expression in Bicyclus anynana (small brown butterfly, Lepidoptera: Nymphalidae) [33]. Gene expression levels in the phototransduction pathway, including Gq, PLCβ, PKC, TRP, TRPL, INAD, Arr2, NINAC, and CamkII, change when M. separata is exposed to different light environments [29]. These studies suggest that the phototransduction pathway is relatively conserved in insects. Furthermore, the expression of the phototransduction genes may be altered in response to changes in the external environment, particularly light.
The functions of phototransduction genes observed in D. citri have been well-demonstrated in Drosophila. For example, mutations in the α-, β-, or γ-subunits of the heterotrimeric Gq protein led to reduced light sensitivity [34,35,36]. Arr2 is a key inhibitory protein responsible for rhodopsin inactivation, which subsequently causes photoreceptor cell inactivation [37]. IP3R, an InsP3 receptor that regulates calcium channels, is linked to retinal degeneration [38,39]. NINAC includes two isoforms, P132 and P174, both of which have a protein kinase structural domain that can bind the myosin head structural domain, but the isoforms have different C-termini, both isoforms are involved in regulating the transport of CaM and Arr2, with P174 mutations resulting in brief reactivation of the photoresponse even after light stimulation ceases [40,41,42]. In the future, more physiological, behavioural, and mechanistic studies are required to gain a comprehensive understanding of the phototransduction pathway in D. citri. With the increasing accessibility of molecular technologies, such as CRISPR/Cas9-based genome editing technology and RNAi technology, research on the function of phototransduction genes in D. citri will be a promising area for future exploration.
The functional relationship between opsins and phototactic behavior is firmly established [8,43,44,45,46]. Although opsins serve as the initial stage in the phototransduction pathway, few studies have explored changes in downstream genes following opsin gene knockout or knockdown. A recent study has demonstrated that the LW-opsin mutation affects the expression of genes involved in the phototransduction pathway in P. xylostella, including NINAC, rdgC, TRP, TRPL, INAD, and others [28]. In our study, only two of the nine validated genes exhibited down-regulated gene expression following the knockdown of the LW-opsin gene. This may be due to the fact that entire female bodies were used as samples in our study, as opposed to only the heads used in previous studies [28,29,31,32,33]. Further validation is required to confirm whether the expression of genes involved in the phototransduction pathway is affected when LW-opsin expression is suppressed using only the heads of D. citri.
5. Conclusions
In conclusion, a total of 71 genes in D. citri were identified as enriched in the phototransduction-fly pathway. These genes encode key proteins in this process, including Gqα, Gqβ, Gqγ, PLCβ, TRP, and TRPL, among others. Moreover, the LOC103513214 and LOC103518375 genes exhibited a reduction in expression when LW-opsin gene expression was suppressed. Our results contribute to a deeper understanding of the molecular mechanisms underlying D. citri vision. This knowledge will facilitate the identification of elicitors that could enhance the understanding of D. citri phototaxis moving forward.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/insects15120966/s1, Supplementary Data File S1: Summary of transcriptome-sequencing results generated from Diaphorina citri; Supplementary Data File S2: The sequences of all transcripts; Supplementary Data File S3: KEGG pathway enrichment of phototransduction genes.
Author Contributions
Conceptualization, R.-Z.Q., J.Z. and S.-P.C.; methodology, S.-P.C. and X.-M.C.; validation, S.-P.C., X.-M.C. and M.-X.C.; data curation, R.-Z.Q., J.Z. and S.-P.C.; writing—original draft preparation, S.-P.C. and X.-M.C.; writing—review and editing, R.-Z.Q. and J.Z.; visualization, S.-P.C., M.-X.C. and X.-M.C.; supervision, R.-Z.Q.; funding acquisition, R.-Z.Q. and S.-P.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Natural Science Foundation of Fujian Province, grant number 2022J05077, the Free Exploring Research Project of the Fujian Academy of Agricultural Sciences, grant number ZYTS2023009, the Key Project of Natural Science Foundation of Fujian Province, grant number 2023J02009, the National Key R&D Program of China, grant number 2021YFD1400800, and the “5511” Collaborative Innovation Project of High-quality Agricultural Development and Surpassment in Fujian Province (XTCXGC2021011, XTCXGC2021017).
Data Availability Statement
All data are available in publicly accessible repositories (https://dataview.ncbi.nlm.nih.gov/object/PRJNA1178351, accessed on 1 October 2024, Accession: PRJNA1178351).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Gottwald, T.R. Current epidemiological understanding of citrus Huanglongbing. Annu. Rev. Phytopathol. 2010, 48, 119–139. [Google Scholar] [CrossRef]
- Grafton-Cardwell, E.E.; Stelinski, L.L.; Stansly, P.A. Biology and management of Asian citrus psyllid, vector of the Huanglongbing pathogens. Annu. Rev. Entomol. 2013, 58, 413–432. [Google Scholar] [CrossRef] [PubMed]
- Bové, J.M. Huanglongbing: A destructive, newly-emerging, century-old disease of citrus. J. Plant Pathol. 2006, 88, 7–37. [Google Scholar] [CrossRef]
- Neupane, D.; Moss, C.B.; Van Bruggen, A.H.C. Estimating citrus production loss due to citrus Huanglongbing in Florida. In Proceedings of the Southern Agricultural Economics Association Annual Meeting, San Antonio, TX, USA, 6–9 February 2016; Available online: https://ageconsearch.umn.edu/bitstream/230093/2/Estimating%20CitrusFinal.pdf (accessed on 28 July 2024).
- Lee, J.A.; Halbert, S.E.; Dawson, W.O.; Robertson, C.J.; Keesling, J.E.; Singer, B.H. Asymptomatic spread of Huanglongbing and implications for disease control. Proc. Natl. Acad. Sci. USA 2015, 112, 7605–7610. [Google Scholar] [CrossRef] [PubMed]
- Paris, T.M.; Allan, S.A.; Udell, B.J.; Stansly, P.A. Wavelength and polarization affect phototaxis of the Asian citrus Psyllid. Insects 2017, 8, 88. [Google Scholar] [CrossRef] [PubMed]
- Paris, T.M.; Croxton, S.D.; Stansly, P.A.; Allan, S.A. Temporal response and attraction of Diaphorina citri to visual stimuli. Entomol. Exp. Appl. 2015, 155, 137–147. [Google Scholar] [CrossRef]
- Li, C.; Tian, F.; Lin, T.; Wang, Z.; Liu, J.; Zeng, X. The expression and function of opsin genes related to the phototactic behavior of Asian citrus psyllid. Pest Manag. Sci. 2020, 76, 1578–1587. [Google Scholar] [CrossRef]
- Terakita, A. The Opsins. Genome Biol. 2005, 6, 213. [Google Scholar] [CrossRef]
- Wang, T.; Montell, C. Phototransduction and retinal degeneration in Drosophila. Pflug. Arch. Eur. J. Physiol. 2007, 454, 821–847. [Google Scholar] [CrossRef]
- Arendt, D.; Tessmar-Raible, K.; Snyman, H.; Dorresteijn, A.W.; Wittbrodt, J. Ciliary photoreceptors with a vertebrate-type opsin in an invertebrate brain. Science 2004, 306, 869–871. [Google Scholar] [CrossRef]
- Ullrich-Lüter, E.M.; D’Aniello, S.; Arnone, M.I. C-opsin expressing photoreceptors in echinoderms. Integr. Comp. Biol. 2013, 53, 27–38. [Google Scholar] [CrossRef] [PubMed]
- Yau, K.W.; Hardie, R.C. Phototransduction motifs and variations. Cell 2009, 139, 246–264. [Google Scholar] [CrossRef] [PubMed]
- Briscoe, A.D.; Chittka, L. The evolution of color vision in insects. Annu. Rev. Entomol. 2001, 46, 471–510. [Google Scholar] [CrossRef] [PubMed]
- Feuda, R.; Marletaz, F.; Bentley, M.A.; Holland, P.W.H. Conservation, duplication, and divergence of five opsin genes in insect evolution. Genome Biol. Evol. 2016, 8, 579–587. [Google Scholar] [CrossRef]
- Lebhardt, F.; Desplan, C. Retinal perception and ecological significance of color vision in insects. Curr. Opin. Insect Sci. 2017, 24, 75–83. [Google Scholar] [CrossRef]
- Mcculloch, K.J.; Macias-Muñoz, A.; Briscoe, A.D. Insect opsins and evo-devo: What have we learned in 25 Years? Philos. Trans. R. Soc. Lond. B Biol. Sci. 2022, 377, 20210288. [Google Scholar] [CrossRef]
- Charlton-Perkins, M.; Cook, T.A. Building a fly eye: Terminal differentiation events of the retina, corneal lens, and pigmented epithelia. Curr. Top. Dev. Biol. 2010, 93, 129–173. [Google Scholar] [CrossRef]
- Salcedo, E.; Huber, A.; Henrich, S.; Chadwell, L.V.; Chou, W.H.; Paulsen, R.; Britt, S.G. Blue- and green-absorbing visual pigments of Drosophila: Ectopic expression and physiological characterization of the R8 photoreceptor cell-specific Rh5 and Rh6 rhodopsins. J. Neurosci. 1999, 19, 10716–10726. [Google Scholar] [CrossRef]
- Senthilan, P.R.; Helfrich-Förster, C. Rhodopsin 7-The unusual Rhodopsin in Drosophila. PeerJ 2016, 4, e2427. [Google Scholar] [CrossRef]
- Ni, J.D.; Baik, L.S.; Holmes, T.C.; Montell, C. A rhodopsin in the brain functions in circadian photoentrainment in Drosophila. Nature 2017, 545, 340–344. [Google Scholar] [CrossRef]
- Jackowska, M.; Bao, R.; Liu, Z.; Mcdonald, E.C.; Cook, T.A.; Friedrich, M. Genomic and gene regulatory signatures of cryptozoic adaptation: Loss of blue sensitive photoreceptors through expansion of long wavelength-opsin expression in the red flour beetle Tribolium castaneum. Front. Zool. 2007, 4, 24. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, M. Parallel losses of blue opsin correlate with compensatory neofunctionalization of UV-opsin gene duplicates in aphids and planthoppers. Insects 2023, 14, 774. [Google Scholar] [CrossRef] [PubMed]
- Montell, C. Drosophila visual transduction. Trends Neurosci. 2012, 35, 356–363. [Google Scholar] [CrossRef] [PubMed]
- Katz, B.; Minke, B. The Drosophila light-activated TRP and TRPL channels—Targets of the phosphoinositide signaling cascade. Prog. Retin. Eye Res. 2018, 66, 200–219. [Google Scholar] [CrossRef]
- Landry, C.R.; Castillo-Davis, C.I.; Ogura, A.; Liu, J.S.; Hartl, D.L. Systems-level analysis and evolution of the phototransduction network in Drosophila. Proc. Natl. Acad. Sci. USA 2007, 104, 3283–3288. [Google Scholar] [CrossRef] [PubMed]
- Hardie, R.C.; Juusola, M. Phototransduction in Drosophila. Curr. Opin. Neurobiol. 2015, 34, 37–45. [Google Scholar] [CrossRef]
- Chen, S.P.; Lin, X.L.; Qiu, R.Z.; Chi, M.X.; Yang, G. An LW-opsin mutation changes the gene expression of the phototransduction pathway: A cryptochrome1 mutation enhances the phototaxis of male Plutella xylostella (Lepidoptera: Plutellidae). Insects 2023, 14, 72. [Google Scholar] [CrossRef]
- Duan, Y.; Gong, Z.J.; Wu, R.H.; Miao, J.; Jiang, Y.L.; Li, T.; Wu, X.B.; Wu, Y.Q. Transcriptome analysis of molecular mechanisms responsible for light-stress response in Mythimna separata (Walker). Sci. Rep. 2017, 7, 45188. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Macias-Muñoz, A.; Rangel Olguin, A.G.; Briscoe, A.D.; Li, W.H. Evolution of phototransduction genes in Lepidoptera. Genome Biol. Evol. 2019, 11, 2107–2124. [Google Scholar] [CrossRef]
- Friedrich, M.; Chen, R.; Daines, B.; Bao, R.; Caravas, J.; Rai, P.K.; Zagmajster, M.; Peck, S.B. Phototransduction and clock gene expression in the troglobiont beetle Ptomaphagus hirtus of Mammoth cave. J. Exp. Biol. 2011, 214, 3532–3541. [Google Scholar] [CrossRef] [PubMed]
- Macias-Munoz, A.; Smith, G.; Monteiro, A.; Briscoe, A.D. Transcriptome-wide differential gene expression in Bicyclus anynana butterflies: Female vision-related genes are more plastic. Mol. Biol. Evol. 2016, 33, 79–92. [Google Scholar] [CrossRef] [PubMed]
- Scott, K.; Becker, A.; Sun, Y.; Hardy, R.; Zuker, C. Gqα protein function in vivo: Genetic dissection of its role in photoreceptor cell physiology. Neuron 1995, 15, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Dolph, P.J.; Man-Son-Hing, H.; Yarfitz, S.; Colley, N.J.; Deer, J.R.; Spencer, M.; Hurley, J.B.; Zuker, C.S. An eye-specific G beta subunit essential for termination of the phototransduction cascade. Nature 1994, 370, 59–61. [Google Scholar] [CrossRef] [PubMed]
- Schillo, S.; Belusic, G.; Hartmann, K.; Franz, C.; Kühl, B.; Brenner-weiss, G.; Paulsen, R.; Huber, A. Targeted mutagenesis of the farnesylation site of Drosophila Gγe disrupts membrane association of the G protein bγ complex and affects the light sensitivity of the visual system. J. Biol. Chem. 2004, 279, 36309–36316. [Google Scholar] [CrossRef]
- Dolph, P.J.; Ranganathan, R.; Colley, N.J.; Hardy, R.W.; Socolich, M.; Zuker, C.S. Arrestin function in inactivation of G protein-coupled receptor rhodopsin in vivo. Science 1993, 260, 1910–1916. [Google Scholar] [CrossRef]
- Srikanth, S.; Wang, Z.; Tu, H.; Nair, S.; Mathew, M.K.; Hasan, G.; Bezprozvanny, I. Functional properties of the Drosophila melanogaster inositol 1,4,5-trisphosphate receptor mutants. Biophys. J. 2004, 86, 3634–3646. [Google Scholar] [CrossRef]
- Vázquez-Martínez, O.; Loranca, A.; Palma-Tirado, L.; Wischin-Fuentes, S.; Villalobos-Leal, M.; Antaramián, A.; Riesgo-Escovar, J.; Hernández-Muñoz, R.; Díaz-Muñoz, M. Time course of retinal degeneration associated with the absence of 1, 4, 5-inositol trisphosphate receptor in Drosophila melanogaster. Exp. Biol. Med. 2010, 235, 365–372. [Google Scholar] [CrossRef]
- Porter, J.A.; Yu, M.; Doberstein, S.K.; Pollard, T.D.; Montell, C. Dependence of calmodulin localization in the retina on the NINAC unconventional myosin. Science 1993, 262, 1038–1042. [Google Scholar] [CrossRef]
- Li, H.S.; Porter, J.A.; Montell, C. Requirement for the NINAC kinase/myosin for stable termination of the visual cascade. J. Neurosci. 1998, 18, 9601–9606. [Google Scholar] [CrossRef]
- Lee, S.J.; Montell, C. Light-dependent translocation of visual arrestin regulated by the NINAC myosin III. Neuron 2004, 43, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Wakakuwa, M.; Stewart, F.; Matsumoto, Y.; Matsunaga, S.; Arikawa, K. Physiological basis of phototaxis to near-infrared light in Nephotettix cincticeps. J. Comp. Physiol. A 2014, 200, 527–536. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; Yan, S.; Shen, Z.J.; Li, Z.; Zhang, X.F.; Liu, X.M.; Zhang, Q.W.; Liu, X.X. The expression of three opsin genes and phototactic behavior of Spodoptera exigua (Lepidoptera: Noctuidae): Evidence for visual function of Opsin in phototaxis. Insect Biochem. Mol. Biol. 2018, 96, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.P.; Liu, Z.X.; Chen, Y.T.; Wang, Y.; Chen, J.Z.; Fu, S.; Ma, W.F.; Xia, S.; Liu, D.; Wu, T.; et al. CRISPR/Cas9-mediated knockout of LW-opsin reduces the efficiency of phototaxis in the diamondback moth Plutella xylostella. Pest Manag. Sci. 2021, 77, 3519–3528. [Google Scholar] [CrossRef]
- Huang, M.; Meng, J.Y.; Zhou, L.; Yu, C.; Zhang, C.Y. Expression and function of opsin genes associated with phototaxis in Zeugodacus cucurbitae Coquillett (Diptera: Tephritidae). Pest Manag. Sci. 2023, 79, 4490–4500. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




