Gene Regulatory Network of the Noncoding RNA Qrr5 Involved in the Cytotoxicity of Vibrio parahaemolyticus during Infection
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Plasmid, and Growth Conditions
2.2. Mutant and Complementary Strain Construction
2.3. RNA Secondary Structure Prediction
2.4. Measurement of Growth Curves
2.5. Measurement of Cytotoxicity and Adherence
2.6. Measurement of Biofilm Formation
2.7. RNA-Seq of V. parahaemolyticus after Infection of Caco-2 Cells
2.8. Reading Mapping and Analysis
2.9. Temporal Analysis
2.10. Weighted Gene Co-Expression Network Analysis (WGCNA)
2.11. Quantitative Real-Time PCR (qRT-PCR)
2.12. Statistical Analysis
3. Results and Discussion
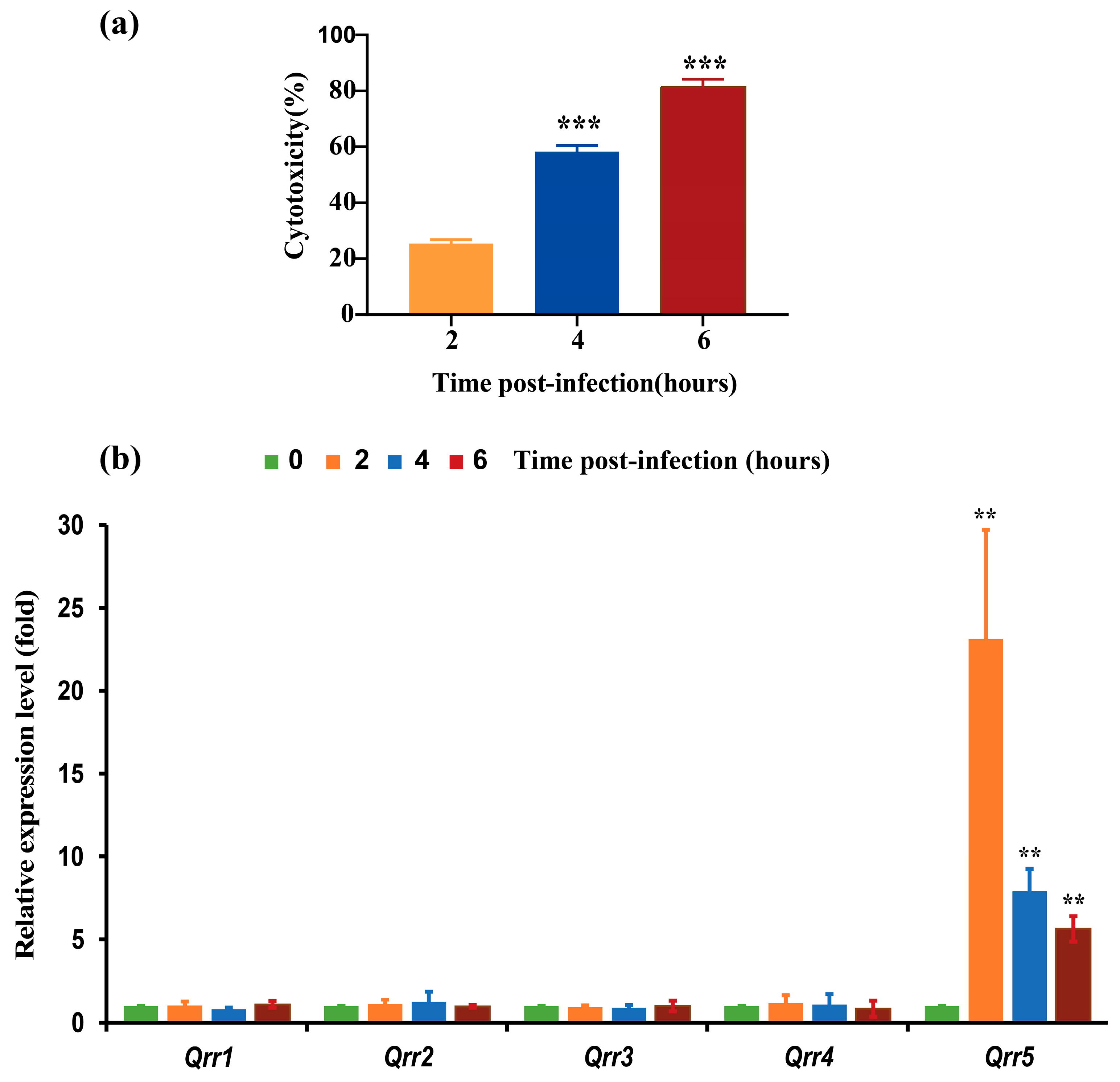
3.1. Expression of Different Qrrs during the Infection of V. parahaemolyticus RIMD2210633 to Caco-2 Cells
3.2. Qrr5 Is Required for Biofilm Formation, Cytotoxicity, and Adhesion to Caco-2 Cells in V. parahaemolyticus RIMD2210633
3.3. Untargeted Analysis of Transcriptome Data of ΔQrr5-Mutant V. parahaemolyticus RIMD2210633
3.4. Targeted Transcriptome Analysis of ΔQrr5-Mutant V. parahaemolyticus RIMD2210633
3.5. WGCNA of Transcriptome Data of ΔQrr5-Mutant V. parahaemolyticus RIMD2210633
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Waters, L.S.; Storz, G. Regulatory RNAs in Bacteria. Cell 2009, 136, 615–628. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hébrard, M.; Kröger, C.; Srikumar, S.; Colgan, A.; Händler, K.; Hinton, J.C.D. sRNAs and the virulence of Salmonella enterica serovar Typhimurium. RNA Biol. 2012, 9, 437–445. [Google Scholar] [CrossRef] [Green Version]
- Djapgne, L.; Oglesby, A.G. Impacts of Small RNAs and Their Chaperones on Bacterial Pathogenicity. Front. Cell. Infect. Microbiol. 2021, 11, 604511. [Google Scholar] [CrossRef] [PubMed]
- Jia, T.; Liu, B.; Mu, H.; Qian, C.; Wang, L.; Li, L.; Lu, G.; Zhu, W.; Guo, X.; Yang, B.; et al. A Novel Small RNA Promotes Motility and Virulence of Enterohemorrhagic Escherichia coli O157:H7 in Response to Ammonium. mBio 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Lenz, D.H.; Mok, K.C.; Lilley, B.N.; Kulkarni, R.V.; Wingreen, N.S.; Bassler, B.L. The Small RNA Chaperone Hfq and Multiple Small RNAs Control Quorum Sensing in Vibrio harveyi and Vibrio cholerae. Cell 2004, 118, 69–82. [Google Scholar] [CrossRef] [Green Version]
- Bardill, J.P.; Zhao, X.; Hammer, B.K. The Vibrio cholerae quorum sensing response is mediated by Hfq-dependent sRNA/mRNA base pairing interactions. Mol. Microbiol. 2011, 80, 1381–1394. [Google Scholar] [CrossRef] [PubMed]
- Vincent, H.A.; Henderson, C.A.; Stone, C.M.; Cary, P.D.; Gowers, D.M.; Sobott, F.; Taylor, J.E.; Callaghan, A.J. The low-resolution solution structure of Vibrio cholerae Hfq in complex with Qrr1 sRNA. Nucleic Acids Res. 2012, 40, 8698–8710. [Google Scholar] [CrossRef]
- Tu, K.C.; Bassler, B.L. Multiple small RNAs act additively to integrate sensory information and control quorum sensing in Vibrio harveyi. Genes Dev. 2007, 21, 221–233. [Google Scholar] [CrossRef] [Green Version]
- Lu, Y.; Rafiq, A.; Zhang, Z.; Aslani, F.; Fijak, M.; Lei, T.; Wang, M.; Kumar, S.; Klug, J.; Bergmann, M.; et al. Uropathogenic Escherichia coli virulence factor hemolysin A causes programmed cell necrosis by altering mitochondrial dynamics. FASEB J. 2018, 32, 4107–4120. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Qiu, Y.; Gao, H.; Sun, J.; Li, X.; Zhang, M.; Xue, X.; Yang, W.; Ni, B.; Hu, L.; et al. OpaR Controls the Metabolism of c-di-GMP in Vibrio parahaemolyticus. Front. Microbiol. 2021, 12, 676436. [Google Scholar] [CrossRef]
- Feng, L.; Rutherford, S.T.; Papenfort, K.; Bagert, J.D.; Van Kessel, J.C.; Tirrell, D.A.; Wingreen, N.S.; Bassler, B.L. A Qrr noncoding RNA deploys four different regulatory mechanisms to optimize quorum-sensing dynamics. Cell 2015, 160, 228–240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaparian, R.R.; Ball, A.S.; van Kessel, J.C. Hierarchical Transcriptional Control of the LuxR Quorum-Sensing Regulon of Vibrio harveyi. J. Bacteriol. 2022, 202, e00047-20. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Kim, I.H.; Kim, K.-S. Iron- and Quorum-sensing Signals Converge on Small Quorum-regulatory RNAs for Coordinated Regulation of Virulence Factors in Vibrio vulnificus*. J. Biol. Chem. 2016, 291, 14213–14230. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, K.-W.; Wen, Y.; Park, N.-Y.; Kim, K.-S. Quorum sensing and iron-dependent coordinated control of autoinducer-2 production via small RNA RyhB in Vibrio vulnificus. Sci. Rep. 2022, 12, 831. [Google Scholar] [CrossRef]
- Tague, J.G.; Regmi, A.; Gregory, G.J.; Boyd, E.F. Fis Connects Two Sensory Pathways, Quorum Sensing and Surface Sensing, to Control Motility in Vibrio parahaemolyticus. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Letchumanan, V.; Chan, K.-G.; Lee, L.-H. Vibrio parahaemolyticus: A review on the pathogenesis, prevalence, and advance molecular identification techniques. Front. Microbiol. 2014, 5, 705. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Orth, K. Virulence determinants for Vibrio parahaemolyticus infection. Curr. Opin. Microbiol. 2013, 16, 70–77. [Google Scholar] [CrossRef]
- Wang, J.; Zhan, Y.; Sun, H.; Fu, X.; Kong, Q.; Zhu, C.; Mou, H. Regulation of Virulence Factors Expression During the Intestinal Colonization of Vibrio parahaemolyticus. Foodborne Pathog. Dis. 2022, 19, 169–178. [Google Scholar] [CrossRef]
- Li, L.; Meng, H.; Gu, D.; Li, Y.; Jia, M. Molecular mechanisms of Vibrio parahaemolyticus pathogenesis. Microbiol. Res. 2019, 222, 43–51. [Google Scholar] [CrossRef]
- Park, K.-S.; Ono, T.; Rokuda, M.; Jang, M.; Okada, K.; Iida, T.; Honda, T. Functional Characterization of Two Type III Secretion Systems of Vibrio parahaemolyticus. Infect. Immun. 2004, 72, 6659–6665. [Google Scholar] [CrossRef]
- Piñeyro, P.; Zhou, X.; Orfe, L.H.; Friel, P.J.; Lahmers, K.; Call, D.R. Development of Two Animal Models To Study the Function of Vibrio parahaemolyticus Type III Secretion Systems. Infect. Immun. 2010, 78, 4551–4559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plaza, N.; Urrutia, I.M.; Garcia, K.; Waldor, M.K.; Blondel, C.J. Identification of a Family of Vibrio Type III Secretion System Effectors That Contain a Conserved Serine/Threonine Kinase Domain. mSphere 2022, 6, e00599-21. [Google Scholar] [CrossRef]
- Erhardt, M.; Namba, K.; Hughes, K.T. Bacterial nanomachines: The flagellum and type III injectisome. Cold Spring Harb. Perspect. Biol. 2010, 2, a000299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Hu, L.; Qiu, Y.; Osei-Adjei, G.; Tang, H.; Zhang, Y.; Zhang, R.; Sheng, X.; Xu, S.; Yang, W.; et al. QsvR integrates into quorum sensing circuit to control Vibrio parahaemolyticus virulence. Environ. Microbiol. 2019, 21, 1054–1067. [Google Scholar] [CrossRef] [PubMed]
- Cornelis, G.R. The type III secretion injectisome. Nat. Reviews. Microbiol. 2006, 4, 811–825. [Google Scholar] [CrossRef]
- Wang, L.; Ling, Y.; Jiang, H.; Qiu, Y.; Qiu, J.; Chen, H.; Yang, R.; Zhou, D. AphA is required for biofilm formation, motility, and virulence in pandemic Vibrio parahaemolyticus. Int. J. Food Microbiol. 2013, 160, 245–251. [Google Scholar] [CrossRef]
- Zeb, S.; Shah, M.A.; Yasir, M.; Awan, H.M.; Prommeenate, P.; Klanchui, A.; Wren, B.W.; Thomson, N.; Bokhari, H. Type III secretion system confers enhanced virulence in clinical non-O1/non-O139 Vibrio cholerae. Microb. Pathog. 2019, 135, 103645. [Google Scholar] [CrossRef]
- Nydam, S.; Shah, D.; Call, D. Transcriptome analysis of Vibrio parahaemolyticus in type III secretion system 1 inducing conditions. Front. Cell. Infect. Microbiol. 2014, 4, 1. [Google Scholar] [CrossRef]
- Lian, L.; Li, W.; Xue, T.; Ren, J.; Tang, F.; Liu, Y.; Xue, F.; Dai, J. Comparative transcriptomic analysis provides insights into transcription mechanisms of Vibrio parahaemolyticus T3SS during interaction with HeLa cells. Braz. J. Microbiol. 2022, 53, 289–301. [Google Scholar] [CrossRef]
- Liu, H.; Liu, W.; He, X.; Chen, X.; Yang, J.; Wang, Y.; Li, Y.; Ren, J.; Xu, W.; Zhao, Y. Characterization of a cell density-dependent sRNA, Qrr, and its roles in the regulation of the quorum sensing and metabolism in Vibrio alginolyticus. Appl. Microbiol. Biotechnol. 2020, 104, 1707–1720. [Google Scholar] [CrossRef]
- Azimi, S.; Klementiev, A.D.; Whiteley, M.; Diggle, S.P. Bacterial Quorum Sensing During Infection. Annu. Rev. Microbiol. 2020, 74, 201–219. [Google Scholar] [CrossRef] [PubMed]
- Suresh, S.; Alva, P.P.; Premanath, R. Modulation of quorum sensing-associated virulence in bacteria: Carbohydrate as a key factor. Arch. Microbiol. 2021, 203, 1881–1890. [Google Scholar] [CrossRef]
- Zhang, Y.; Qiu, Y.; Tan, Y.; Guo, Z.; Yang, R.; Zhou, D. Transcriptional Regulation of opaR, qrr2–4 and aphA by the Master Quorum-Sensing Regulator OpaR in Vibrio parahaemolyticus. PLoS ONE 2012, 7, e34622. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Shah, D.; Konkel, M.; Call, D. Type III secretion system 1 genes in Vibrio parahaemolyticus are positively regulated by ExsA and negatively regulated by ExsD. Mol. Microbiol. 2008, 69, 747–764. [Google Scholar] [CrossRef] [Green Version]
- O’Toole, G.A.; Kolter, R. Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: A genetic analysis. Mol. Microbiol. 1998, 28, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Wingett, S.; Andrews, S. FastQ Screen: A tool for multi-genome mapping and quality control. F1000Research 2018, 7, 1338. [Google Scholar] [CrossRef]
- Wang, L.; Zhang, W.; Li, Q. AssocTests: An R Package for Genetic Association Studies. J. Stat. Softw. 2020, 94, 1–26. [Google Scholar] [CrossRef]
- Ernst, J.; Bar-Joseph, Z. STEM: A tool for the analysis of short time series gene expression data. BMC Bioinform. 2006, 7, 191. [Google Scholar] [CrossRef] [Green Version]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Livny, J.; Zhou, X.; Mandlik, A.; Hubbard, T.; Davis, B.M.; Waldor, M.K. Comparative RNA-Seq based dissection of the regulatory networks and environmental stimuli underlying Vibrio parahaemolyticus gene expression during infection. Nucleic Acids Res. 2014, 42, 12212–12223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos, J.C.; Broz, P. Sensing of invading pathogens by GBPs: At the crossroads between cell-autonomous and innate immunity. J. Leukoc. Biol. 2018, 104, 729–735. [Google Scholar] [CrossRef]
- Netea, M.G.; Schlitzer, A.; Placek, K.; Joosten, L.A.B.; Schultze, J.L. Innate and Adaptive Immune Memory: An Evolutionary Continuum in the Host’s Response to Pathogens. Cell Host Microbe 2019, 25, 13–26. [Google Scholar] [CrossRef] [Green Version]
- Zhang, W.; Xie, R.; Zhang, X.D.; Lee, L.T.O.; Zhang, H.; Yang, M.; Peng, B.; Zheng, J. Organism dual RNA-seq reveals the importance of BarA/UvrY in Vibrio parahaemolyticus virulence. FASEB J. 2020, 34, 7561–7577. [Google Scholar] [CrossRef] [Green Version]
- Ning, X.; Sun, L. Gene network analysis reveals a core set of genes involved in the immune response of Japanese flounder (Paralichthys olivaceus) against Vibrio anguillarum infection. Fish Shellfish. Immunol. 2020, 98, 800–809. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Fang, L.; Zhang, Y.; Sheng, H.; Fang, W. VgrG2 of type VI secretion system 2 of Vibrio parahaemolyticus induces autophagy in macrophages. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef] [Green Version]
- MacIntyre, D.L.; Miyata, S.T.; Kitaoka, M.; Pukatzki, S. The Vibrio cholerae type VI secretion system displays antimicrobial properties. Proc. Natl. Acad. Sci. USA 2010, 107, 19520–19524. [Google Scholar] [CrossRef] [Green Version]
- Babitzke, P.; Romeo, T. CsrB sRNA family: Sequestration of RNA-binding regulatory proteins. Curr. Opin. Microbiol. 2007, 10, 156–163. [Google Scholar] [CrossRef]
- Chen, J.; Morita, T.; Gottesman, S. Regulation of Transcription Termination of Small RNAs and by Small RNAs: Molecular Mechanisms and Biological Functions. Front. Cell. Infect. Microbiol. 2019, 9, 201. [Google Scholar] [CrossRef]
- Gordon, W.; Sofia, B.A.; Adriana, B.; Giulia, T.; Gaëtan, D.M.; Thomas, H.; Hoskisson, P.A.; Ulrich, Z.; Maria, M.A.; Mélanie, B.; et al. Coexistence of Ammonium Transporter and Channel Mechanisms in Amt-Mep-Rh Twin-His Variants Impairs the Filamentation Signaling Capacity of Fungal Mep2 Transceptors. mBio 2022, 13, e02913-21. [Google Scholar] [CrossRef]
- Plano, G.; Joseph, S. The SycN/YscB chaperone-binding domain of YopN is required for the calcium-dependent regulation of Yop secretion by Yersinia pestis. Front. Cell. Infect. Microbiol. 2013, 3. [Google Scholar]
- Yi, L.L.; Pin, N.; Bing, Y.H.; Xia, X.H. Regulation of Type III Secretion of Translocon and Effector Proteins by the EsaB/EsaL/EsaM Complex in Edwardsiella tarda. Infect. Immun. 2022, 85, e00322-17. [Google Scholar] [CrossRef] [Green Version]
- Plano, G.V.; Schesser, K. The Yersinia pestis type III secretion system: Expression, assembly and role in the evasion of host defenses. Immunol. Res. 2013, 57, 237–245. [Google Scholar] [CrossRef] [PubMed]





| Strain or Plasmid | Characteristics | Reference |
|---|---|---|
| V. parahaemolyticus RIMD 2210633 | Parent strain, serotype O3:K6 | [33] |
| ΔQrr5 mutant | RIMD 2210633 ΔQrr5 | This study |
| ΔQrr5::Qrr5 | pBAD33-Qrr5 into ΔQrr5 mutant | This study |
| E. coli SM17 λpir | TpR SmR recA, thi, pro, hsdR-M+RP4:2-Tc:Mu:Km Tn7λpir | [33] |
| pDS132 | R6K ori, mobRP4 SacB Cmr | [33] |
| pBAD33 | pBR322 ori, Ampr | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zha, F.; Pang, R.; Huang, S.; Zhang, J.; Wang, J.; Chen, M.; Xue, L.; Ye, Q.; Wu, S.; Yang, M.; et al. Gene Regulatory Network of the Noncoding RNA Qrr5 Involved in the Cytotoxicity of Vibrio parahaemolyticus during Infection. Microorganisms 2022, 10, 2084. https://doi.org/10.3390/microorganisms10102084
Zha F, Pang R, Huang S, Zhang J, Wang J, Chen M, Xue L, Ye Q, Wu S, Yang M, et al. Gene Regulatory Network of the Noncoding RNA Qrr5 Involved in the Cytotoxicity of Vibrio parahaemolyticus during Infection. Microorganisms. 2022; 10(10):2084. https://doi.org/10.3390/microorganisms10102084
Chicago/Turabian StyleZha, Fei, Rui Pang, Shixuan Huang, Jumei Zhang, Juan Wang, Moutong Chen, Liang Xue, Qinghua Ye, Shi Wu, Meiyan Yang, and et al. 2022. "Gene Regulatory Network of the Noncoding RNA Qrr5 Involved in the Cytotoxicity of Vibrio parahaemolyticus during Infection" Microorganisms 10, no. 10: 2084. https://doi.org/10.3390/microorganisms10102084





