Methanomethylophilus alvi gen. nov., sp. nov., a Novel Hydrogenotrophic Methyl-Reducing Methanogenic Archaea of the Order Methanomassiliicoccales Isolated from the Human Gut and Proposal of the Novel Family Methanomethylophilaceae fam. nov.
Abstract
:1. Introduction
2. Materials and Methods
2.1. Medium of Enrichment, Isolation, and Maintenance
2.2. Genome Sequencing, Genomic and Phylogenomic Analyses
2.3. Microscopy
2.4. Physiological Characterization
3. Results and Discussion
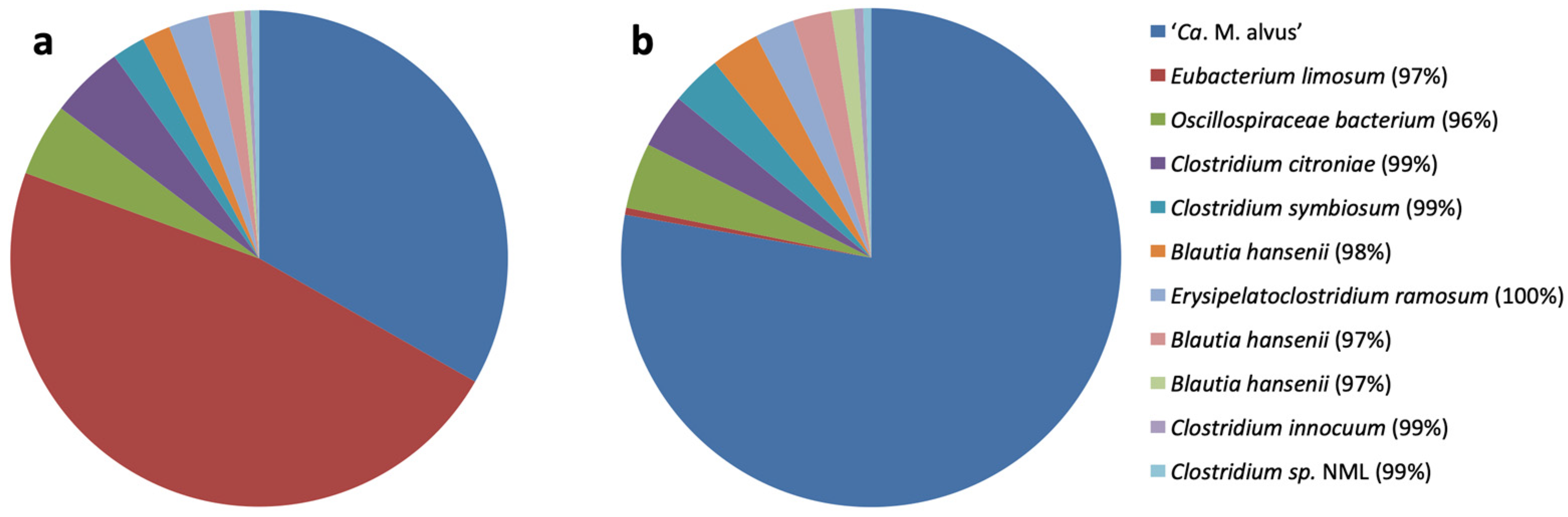
3.1. Selective Enrichment and Isolation
3.2. Phylogeny and Genome Features
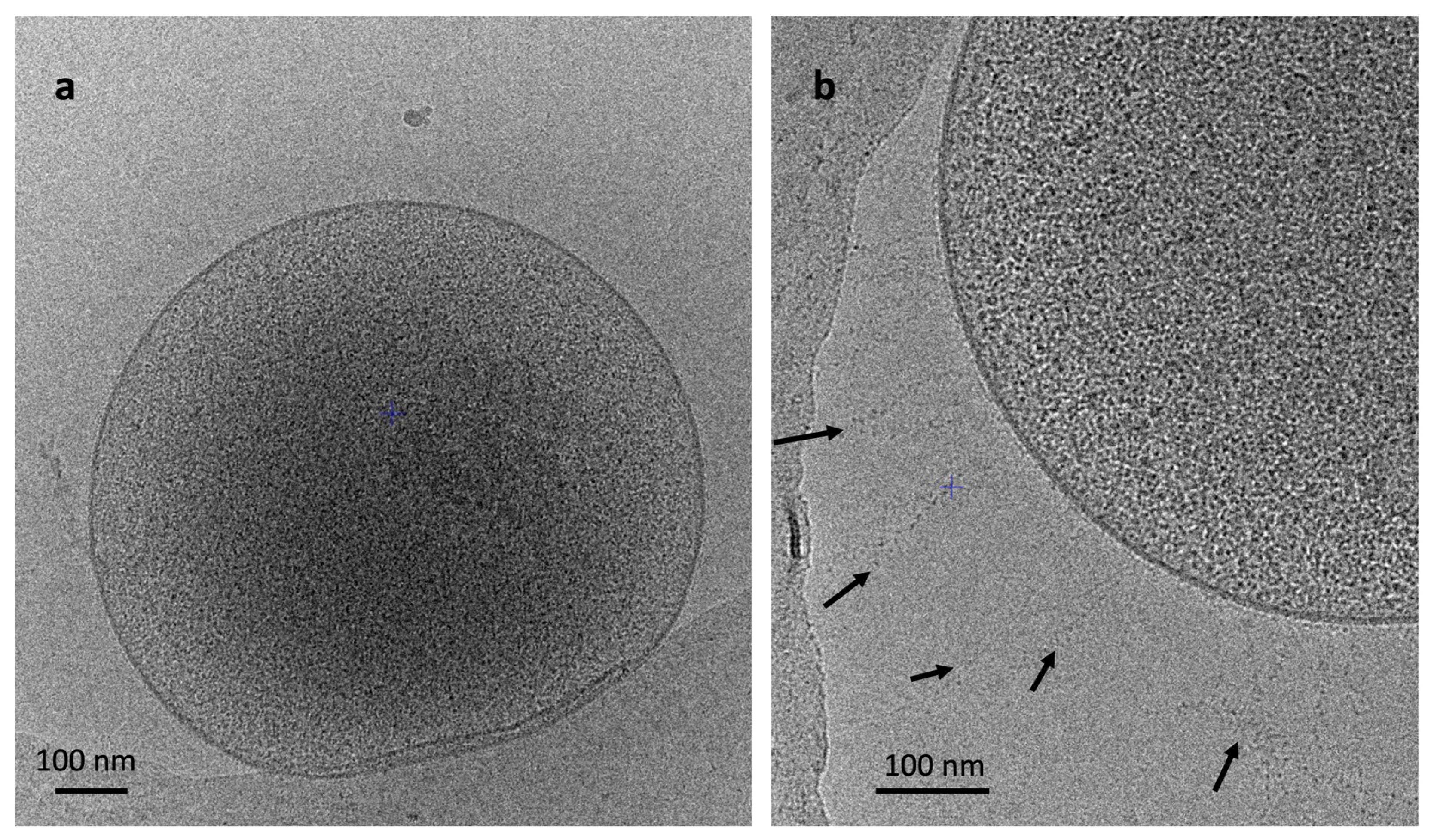
3.3. Morphology
3.4. Physiology
4. Conclusions
4.1. Description of Methanomethylophilus gen. nov.
4.2. Description of Methanomethylophilus alvi sp. nov.
4.3. Description of Methanomethylophilaceae fam. nov.
5. Patents
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Iino, T.; Tamaki, H.; Tamazawa, S.; Ueno, Y.; Ohkuma, M.; Suzuki, K.; Igarashi, Y.; Haruta, S. Candidatus Methanogranum caenicola: A Novel Methanogen from the Anaerobic Digested Sludge, and Proposal of Methanomassiliicoccaceae fam. nov. and Methanomassiliicoccales ord. nov., for a Methanogenic Lineage of the Class Thermoplasmata. Microbes Environ. 2013, 28, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Dridi, B.; Fardeau, M.L.; Ollivier, B.; Raoult, D.; Drancourt, M. Methanomassiliicoccus luminyensis gen. nov., sp. nov., a Methanogenic Archaeon Isolated from Human Faeces. Int. J. Syst. Evol. Microbiol. 2012, 62, 1902–1907. [Google Scholar] [CrossRef]
- Thomas, C.M.; Desmond-Le Quéméner, E.; Gribaldo, S.; Borrel, G. Factors Shaping the Abundance and Diversity of the Gut Archaeome across the Animal Kingdom. Nat. Commun. 2022, 13, 3358. [Google Scholar] [CrossRef]
- Söllinger, A.; Schwab, C.; Weinmaier, T.; Loy, A.; Tveit, A.T.; Schleper, C.; Urich, T. Phylogenetic and Genomic Analysis of Methanomassiliicoccales in Wetlands and Animal Intestinal Tracts Reveals Clade-Specific Habitat Preferences. FEMS Microbiol. Ecol. 2016, 92, fiv149. [Google Scholar] [CrossRef] [PubMed]
- Borrel, G.; McCann, A.; Deane, J.; Neto, M.C.; Lynch, D.B.; Brugère, J.F.; O’Toole, P.W. Genomics and Metagenomics of Trimethylamine-Utilizing Archaea in the Human Gut Microbiome. ISME J. 2017, 11, 2059–2074. [Google Scholar] [CrossRef] [PubMed]
- Chibani, C.M.; Mahnert, A.; Borrel, G.; Almeida, A.; Werner, A.; Brugère, J.-F.; Gribaldo, S.; Finn, R.D.; Schmitz, R.A.; Moissl-Eichinger, C. A Catalogue of 1,167 Genomes from the Human Gut Archaeome. Nat. Microbiol. 2022, 7, 48–61. [Google Scholar] [CrossRef]
- Cozannet, M.; Borrel, G.; Roussel, E.; Moalic, Y.; Allioux, M.; Sanvoisin, A.; Toffin, L.; Alain, K. New Insights into the Ecology and Physiology of Methanomassiliicoccales from Terrestrial and Aquatic Environments. Microorganisms 2021, 9, 30. [Google Scholar] [CrossRef]
- Chen, C.; Li, L.; Wang, Y.; Dong, X.; Zhao, F.-J. Methylotrophic Methanogens and Bacteria Synergistically Demethylate Dimethylarsenate in Paddy Soil and Alleviate Rice Straighthead Disease. ISME J. 2023, 17, 1851–1861. [Google Scholar] [CrossRef]
- Borrel, G.; Harris, H.M.B.; Parisot, N.; Gaci, N.; Tottey, W.; Mihajlovski, A.; Deane, J.; Gribaldo, S.; Bardot, O.; Peyretaillade, E.; et al. Genome Sequence of “Candidatus Methanomassiliicoccus intestinalis” Issoire-Mx1, a Third Thermoplasmatales-Related Methanogenic Archaeon from Human Feces. Genome Announc. 2013, 1, e00453-13. [Google Scholar] [CrossRef]
- Weil, M.; Hoff, K.J.; Meißner, W.; Schäfer, F.; Söllinger, A.; Wang, H.; Hagenau, L.; Kuss, A.W.; Urich, T. Full Genome Sequence of a Methanomassiliicoccales Representative Enriched from Peat Soil. Microbiol. Resour. Announc. 2021, 10, e00443-21. [Google Scholar] [CrossRef]
- Janssen, P.H.; Kirs, M. Structure of the Archaeal Community of the Rumen. Appl. Environ. Microbiol. 2008, 74, 3619–3625. [Google Scholar] [CrossRef] [PubMed]
- Borrel, G.; O’Toole, P.W.; Harris, H.M.B.; Peyret, P.; Brugère, J.F.; Gribaldo, S. Phylogenomic Data Support a Seventh Order of Methylotrophic Methanogens and Provide Insights into the Evolution of Methanogenesis. Genome Biol. Evol. 2013, 5, 1769–1780. [Google Scholar] [CrossRef] [PubMed]
- Paul, K.; Nonoh, J.O.; Mikulski, L.; Brune, A. “Methanoplasmatales,” Thermoplasmatales-Related Archaea in Termite Guts and Other Environments, Are the Seventh Order of Methanogens. Appl. Environ. Microbiol. 2012, 78, 8245–8253. [Google Scholar] [CrossRef]
- Borrel, G.; Harris, H.M.B.; Tottey, W.; Mihajlovski, A.; Parisot, N.; Peyretaillade, E.; Peyret, P.; Gribaldo, S.; O’Toole, P.W.; Brugère, J.F. Genome Sequence of “Candidatus Methanomethylophilus alvus” Mx1201, a Methanogenic Archaeon from the Human Gut Belonging to a Seventh Order of Methanogens. J. Bacteriol. 2012, 194, 6944–6945. [Google Scholar] [CrossRef]
- Oren, A.; Garrity, G.M.; Parker, C.T.; Chuvochina, M.; Trujillo, M.E. Lists of Names of Prokaryotic Candidatus Taxa. Int. J. Syst. Evol. Microbiol. 2020, 70, 3956–4042. [Google Scholar] [CrossRef] [PubMed]
- Brugère, J.F.; Borrel, G.; Gaci, N.; Tottey, W.; O’Toole, P.W.; Malpuech-Brugère, C. Archaebiotics: Proposed Therapeutic Use of Archaea to Prevent Trimethylaminuria and Cardiovascular Disease. Gut Microbes 2014, 5, 5–10. [Google Scholar] [CrossRef]
- Li, Y.; Leahy, S.C.; Jeyanathan, J.; Henderson, G.; Cox, F.; Altermann, E.; Kelly, W.J.; Lambie, S.C.; Janssen, P.H.; Rakonjac, J.; et al. The Complete Genome Sequence of the Methanogenic Archaeon ISO4-H5 Provides Insights into the Methylotrophic Lifestyle of a Ruminal Representative of the Methanomassiliicoccales. Stand. Genom. Sci. 2016, 11, 59. [Google Scholar] [CrossRef]
- Noel, S.J.; Højberg, O.; Urich, T.; Poulsen, M. Draft Genome Sequence of “Candidatus Methanomethylophilus” Sp. 1R26, Enriched from Bovine Rumen, a Methanogenic Archaeon Belonging to the Methanomassiliicoccales Order. Genome Announc. 2016, 4, e01734-15. [Google Scholar] [CrossRef]
- Lang, K.; Schuldes, J.; Klingl, A.; Poehlein, A.; Daniel, R.; Brune, A. New Mode of Energy Metabolism in the Seventh Order of Methanogens as Revealed by Comparative Genome Analysis of “Candidatus Methanoplasma Termitum”. Appl. Environ. Microbiol. 2015, 81, 1338–1352. [Google Scholar] [CrossRef]
- de la Cuesta-Zuluaga, J.; Spector, T.D.; Youngblut, N.D.; Ley, R.E. Genomic Insights into Adaptations of Trimethylamine-Utilizing Methanogens to Diverse Habitats, Including the Human Gut. mSystems 2021, 6, e00939-20. [Google Scholar] [CrossRef]
- Borrel, G.; Parisot, N.; Harris, H.M.; Peyretaillade, E.; Gaci, N.; Tottey, W.; Bardot, O.; Raymann, K.; Gribaldo, S.; Peyret, P.; et al. Comparative Genomics Highlights the Unique Biology of Methanomassiliicoccales, a Thermoplasmatales-Related Seventh Order of Methanogenic Archaea That Encodes Pyrrolysine. BMC Genom. 2014, 15, 679. [Google Scholar] [CrossRef] [PubMed]
- Kröninger, L.; Berger, S.; Welte, C.; Deppenmeier, U. Evidence for the Involvement of Two Heterodisulfide Reductases in the Energy-Conserving System of Methanomassiliicoccus luminyensis. FEBS J. 2016, 283, 472–483. [Google Scholar] [CrossRef] [PubMed]
- Kröninger, L.; Steiniger, F.; Berger, S.; Kraus, S.; Welte, C.U.; Deppenmeier, U. Energy Conservation in the Gut Microbe Methanomassiliicoccus luminyensis Is Based on Membrane-bound Ferredoxin Oxidation Coupled to Heterodisulfide Reduction. FEBS J. 2019, 286, 3831–3843. [Google Scholar] [CrossRef] [PubMed]
- Becker, K.W.; Elling, F.J.; Yoshinaga, M.Y.; Söllinger, A.; Urich, T.; Hinrichs, K.U. Unusual Butane- and Pentanetriol-Based Tetraether Lipids in Methanomassiliicoccus luminyensis, a Representative of the Seventh Order of Methanogens. Appl. Environ. Microbiol. 2016, 82, 4505–4516. [Google Scholar] [CrossRef]
- Coffinet, S.; Mühlena, L.; Lipp, J.S.; Weil, M.; Neubauer, C.; Urich, T.; Hinrichs, K.-U. Evidence for Enzymatic Backbone Methylation of the Main Membrane Lipids in the Archaeon Methanomassiliicoccus luminyensis. Appl. Environ. Microbiol. 2022, 88, e02154-21. [Google Scholar] [CrossRef]
- Brugère, J.-F.; Atkins, J.F.; O’Toole, P.W.; Borrel, G. Pyrrolysine in Archaea: A 22nd Amino Acid Encoded through a Genetic Code Expansion. Emerg. Top. Life Sci. 2018, 2, 607–618. [Google Scholar]
- Borrel, G.; Gaci, N.; Peyret, P.; O’Toole, P.W.; Gribaldo, S.; Brugère, J.F. Unique Characteristics of the Pyrrolysine System in the 7th Order of Methanogens: Implications for the Evolution of a Genetic Code Expansion Cassette. Archaea 2014, 2014, 374146. [Google Scholar] [CrossRef]
- Willis, J.C.W.; Chin, J.W. Mutually Orthogonal Pyrrolysyl-TRNA Synthetase/TRNA Pairs. Nat. Chem. 2018, 10, 831–837. [Google Scholar] [CrossRef]
- Li, S.; Wang, N.; Yu, B.; Sun, W.; Wang, L. Genetically Encoded Chemical Crosslinking of Carbohydrate. Nat. Chem. 2023, 15, 33–42. [Google Scholar] [CrossRef]
- Tang, W.H.W.; Wang, Z.; Kennedy, D.J.; Wu, Y.; Buffa, J.A.; Agatisa-Boyle, B.; Li, X.S.; Levison, B.S.; Hazen, S.L. Gut Microbiota-Dependent Trimethylamine N-Oxide (TMAO) Pathway Contributes to Both Development of Renal Insufficiency and Mortality Risk in Chronic Kidney Disease. Circ. Res. 2015, 116, 448–455. [Google Scholar] [CrossRef]
- Zhu, W.; Gregory, J.C.; Org, E.; Buffa, J.A.; Gupta, N.; Wang, Z.; Li, L.; Fu, X.; Wu, Y.; Mehrabian, M.; et al. Gut Microbial Metabolite TMAO Enhances Platelet Hyperreactivity and Thrombosis Risk. Cell 2016, 165, 111–124. [Google Scholar] [CrossRef]
- Mackay, R.J.; McEntyre, C.J.; Henderson, C.; Lever, M.; George, P.M. Trimethylaminuria: Causes and Diagnosis of a Socially Distressing Condition. Clin. Biochem. Rev. 2011, 32, 33–43. [Google Scholar]
- Fadhlaoui, K.; Arnal, M.-E.; Martineau, M.; Camponova, P.; Ollivier, B.; O’Toole, P.W.; Brugère, J.-F. Archaea, Specific Genetic Traits, and Development of Improved Bacterial Live Biotherapeutic Products: Another Face of next-Generation Probiotics. Appl. Microbiol. Biotechnol. 2020, 104, 4705–4716. [Google Scholar] [CrossRef] [PubMed]
- Widdel, F.; Pfennig, N. Studies on Dissimilatory Sulfate-Reducing Bacteria That Decompose Fatty Acids: I. Isolation of New Sulfate-Reducing Bacteria Enriched with Acetate from Saline Environments. Description of Desulfobacter postgatei Gen. Nov., Sp. Nov. Arch. Microbiol. 1981, 129, 395–400. [Google Scholar] [CrossRef]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI Prokaryotic Genome Annotation Pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Borrel, G.; Adam, P.S.; McKay, L.J.; Chen, L.-X.; Sierra-García, I.N.; Sieber, C.M.K.; Letourneur, Q.; Ghozlane, A.; Andersen, G.L.; Li, W.-J.; et al. Wide Diversity of Methane and Short-Chain Alkane Metabolisms in Uncultured Archaea. Nat. Microbiol. 2019, 4, 603–613. [Google Scholar] [CrossRef]
- Parks, D.H.; Chuvochina, M.; Rinke, C.; Mussig, A.J.; Chaumeil, P.-A.; Hugenholtz, P. GTDB: An Ongoing Census of Bacterial and Archaeal Diversity through a Phylogenetically Consistent, Rank Normalized and Complete Genome-Based Taxonomy. Nucleic Acids Res. 2022, 50, D785–D794. [Google Scholar] [CrossRef]
- Yuan, Y.; Liu, J.; Yang, T.-T.; Gao, S.-M.; Liao, B.; Huang, L.-N. Genomic Insights into the Ecological Role and Evolution of a Novel Thermoplasmata Order, “Candidatus Sysuiplasmatales”. Appl. Environ. Microbiol. 2021, 87, e01065-21. [Google Scholar] [CrossRef] [PubMed]
- Johnson, L.S.; Eddy, S.R.; Portugaly, E. Hidden Markov Model Speed Heuristic and Iterative HMM Search Procedure. BMC Bioinform. 2010, 11, 431. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Criscuolo, A.; Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): A New Software for Selection of Phylogenetic Informative Regions from Multiple Sequence Alignments. BMC Evol. Biol. 2010, 10, 210. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Hartmann, M.; Eriksson, K.M.; Pal, C.; Thorell, K.; Larsson, D.G.J.; Nilsson, R.H. METAXA2: Improved Identification and Taxonomic Classification of Small and Large Subunit rRNA in Metagenomic Data. Mol. Ecol. Resour. 2015, 15, 1403–1414. [Google Scholar] [CrossRef] [PubMed]
- Flemer, B.; Gaci, N.; Borrel, G.; Sanderson, I.R.; Chaudhary, P.P.; Tottey, W.; O’Toole, P.W.; Brugère, J.F. Fecal Microbiota Variation across the Lifespan of the Healthy Laboratory Rat. Gut Microbes 2017, 8, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (ITOL) v4: Recent Updates and New Developments. Nucleic Acids Res. 2019, 47, 256–259. [Google Scholar] [CrossRef]
- Jüttner, M.; Ferreira-Cerca, S. Looking through the Lens of the Ribosome Biogenesis Evolutionary History: Possible Implications for Archaeal Phylogeny and Eukaryogenesis. Mol. Biol. Evol. 2022, 39, msac054. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Taib, N.; Gribaldo, S.; Borrel, G. Comparative Genomic Analysis of Methanimicrococcus blatticola Provides Insights into Host-Adaptation in Archaea and the Evolution of Methanogenesis. ISME Commun. 2021, 1, 47. [Google Scholar] [CrossRef]
- Martins, M.; Aymeric, L.; Du Merle, L.; Danne, C.; Robbe-Masselot, C.; Trieu-Cuot, P.; Sansonetti, P.; Dramsi, S. Streptococcus gallolyticus Pil3 Pilus Is Required for Adhesion to Colonic Mucus and for Colonization of Mouse Distal Colon. J. Infect. Dis. 2015, 212, 1646–1655. [Google Scholar] [CrossRef]
- Medvedeva, S.; Borrel, G.; Krupovic, M.; Gribaldo, S. A Compendium of Viruses from Methanogenic Archaea Reveals Their Diversity and Adaptations to the Gut Environment. Nat. Microbiol. 2023, 8, 2170–2182. [Google Scholar] [CrossRef]
- Hocher, A.; Borrel, G.; Fadhlaoui, K.; Brugère, J.-F.; Gribaldo, S.; Warnecke, T. Growth Temperature and Chromatinization in Archaea. Nat. Microbiol. 2022, 7, 1932–1942. [Google Scholar] [CrossRef]
- Tallant, T.C.; Krzycki, J.A. Methylthiol: Coenzyme M Methyltransferase from Methanosarcina barkeri, an Enzyme of Methanogenesis from Dimethylsulfide and Methylmercaptopropionate. J. Bacteriol. 1997, 179, 6902–6911. [Google Scholar] [CrossRef] [PubMed]
- Weil, M.; Wang, H.; Zak, D.; Urich, T. Spatial and Temporal Niche Separation of Methanomassiliicoccales Phylotypes in Temperate Fens. FEMS Microbiol. Ecol. 2023, 99, fiad049. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.-J.; Pan, J.; Liu, Y.; Duan, C.-H.; Li, M. Genomic and Transcriptomic Insights into Methanogenesis Potential of Novel Methanogens from Mangrove Sediments. Microbiome 2020, 8, 94. [Google Scholar] [CrossRef] [PubMed]
- Seedorf, H.; Kittelmann, S.; Janssen, P.H. Few Highly Abundant Operational Taxonomic Units Dominate within Rumen Methanogenic Archaeal Species in New Zealand Sheep and Cattle. Appl. Environ. Microbiol. 2015, 81, 986–995. [Google Scholar] [CrossRef]
- Poulsen, M.; Schwab, C.; Jensen, B.B.; Engberg, R.M.; Spang, A.; Canibe, N.; Højberg, O.; Milinovich, G.; Fragner, L.; Schleper, C.; et al. Methylotrophic Methanogenic Thermoplasmata Implicated in Reduced Methane Emissions from Bovine Rumen. Nat. Commun. 2013, 4, 1428. [Google Scholar] [CrossRef]
- Borrel, G.; Gribaldo, S.; Brugère, J.-F.; Schmitz, R.; Moissl-Eichinger, C. The Host-Associated Archaeome. Nat. Rev. Microbiol. 2020, 18, 622–636. [Google Scholar] [CrossRef]
- Jackson, M.A.; Jeffery, I.B.; Beaumont, M.; Bell, J.T.; Clark, A.G.; Ley, R.E.; O’Toole, P.W.; Spector, T.D.; Steves, C.J. Signatures of Early Frailty in the Gut Microbiota. Genome Med. 2016, 8, 8. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, H.; Hu, L.; Zhang, G.; Lu, H.; Luo, H.; Zhao, S.; Zhu, H.; Wang, Y. Characterization of the Microbial Communities along the Gastrointestinal Tract in Crossbred Cattle. Animals 2022, 12, 825. [Google Scholar] [CrossRef]

| Characteristics | 1 | 2 |
|---|---|---|
| Source | Human feces | Human feces |
| Genome length (bp) | 1,666,767 | 2,637,810 |
| DNA G+C content (mol%) | 55.5 | 59.93 |
| Morphology | Cocci, as single cells | Cocci, as single cells |
| Diameter (µm) | 0.4–0.7 | 0.7–1.0 |
| Motility | Non-motile | Non-motile |
| Growth temperature | ||
| Range | 30–40 | 30–40 |
| Optimum | 37 | 37 |
| Growth pH | ||
| Range | 6.9–8.3 | 7.2–8.4 |
| Optimum | 7.5 | 7.6 |
| Growth NaCl (mol.L−1) | ||
| Range | 0.02–0.34 | 0.02–0.26 |
| Optimum | 0.12 | 0.17 |
| Required growth factor | Filtrate from a grown E. lenta or rumen fluid | acetate |
| Substrates for methane production | H2/methanol H2/monomethylamine H2/dimethylamine H2/trimethylamine | H2/methanol H2/monomethylamine H2/dimethylamine H2/trimethylamine |
| Lysed by osmotic shock/SDS | yes | yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borrel, G.; Fadhlaoui, K.; Ben Hania, W.; Gaci, N.; Pehau-Arnaudet, G.; Chaudhary, P.P.; Vandekerckove, P.; Ballet, N.; Alric, M.; O’Toole, P.W.; et al. Methanomethylophilus alvi gen. nov., sp. nov., a Novel Hydrogenotrophic Methyl-Reducing Methanogenic Archaea of the Order Methanomassiliicoccales Isolated from the Human Gut and Proposal of the Novel Family Methanomethylophilaceae fam. nov. Microorganisms 2023, 11, 2794. https://doi.org/10.3390/microorganisms11112794
Borrel G, Fadhlaoui K, Ben Hania W, Gaci N, Pehau-Arnaudet G, Chaudhary PP, Vandekerckove P, Ballet N, Alric M, O’Toole PW, et al. Methanomethylophilus alvi gen. nov., sp. nov., a Novel Hydrogenotrophic Methyl-Reducing Methanogenic Archaea of the Order Methanomassiliicoccales Isolated from the Human Gut and Proposal of the Novel Family Methanomethylophilaceae fam. nov. Microorganisms. 2023; 11(11):2794. https://doi.org/10.3390/microorganisms11112794
Chicago/Turabian StyleBorrel, Guillaume, Khaled Fadhlaoui, Wajdi Ben Hania, Nadia Gaci, Gérard Pehau-Arnaudet, Prem Prashant Chaudhary, Pascal Vandekerckove, Nathalie Ballet, Monique Alric, Paul William O’Toole, and et al. 2023. "Methanomethylophilus alvi gen. nov., sp. nov., a Novel Hydrogenotrophic Methyl-Reducing Methanogenic Archaea of the Order Methanomassiliicoccales Isolated from the Human Gut and Proposal of the Novel Family Methanomethylophilaceae fam. nov." Microorganisms 11, no. 11: 2794. https://doi.org/10.3390/microorganisms11112794





