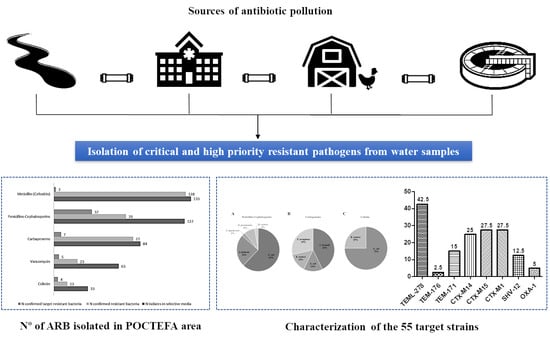
Multidrug-Resistant Bacteria Isolated from Different Aquatic Environments in the North of Spain and South of France
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Isolation and Identification of Resistant Bacteria
2.3. Phenotypic and Genotypic Characterization of Resistant Strains
2.3.1. ESBL-Producer Enterobacteriaceae and Other β-Lactamases
2.3.2. Carbapenemase-Producing Strains
2.3.3. Colistin Resistant Enterobacteriaceae
2.3.4. Methicillin Resistant Strains
2.3.5. Vancomycin Resistant Strains
2.4. Antimicrobial Resistance Patterns
2.5. Statistical Analysis
3. Results
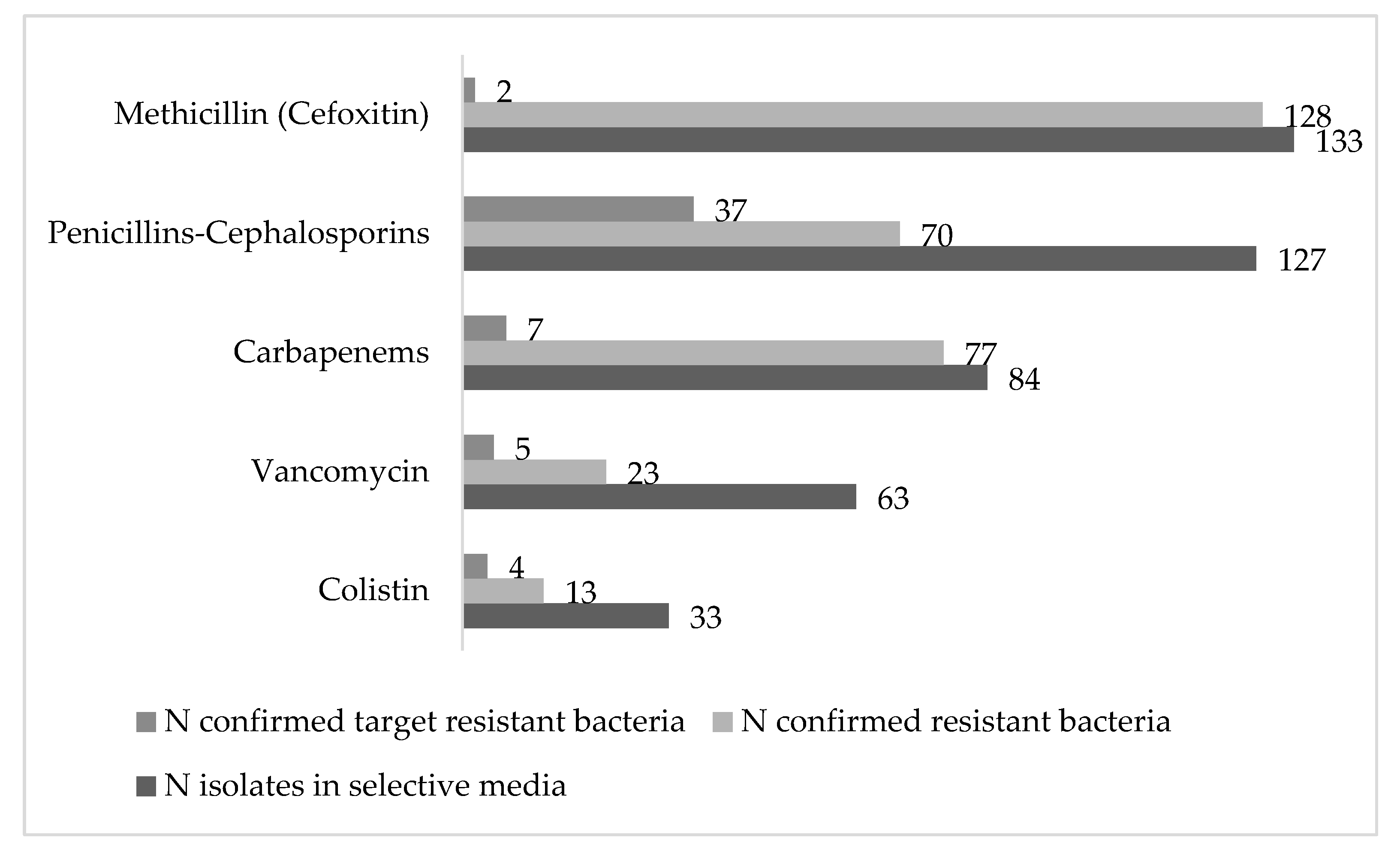
3.1. Prevalence of Resistant Bacteria in Rivers and Sewage Waters
3.2. Characteristics of the Target Isolated Pathogens
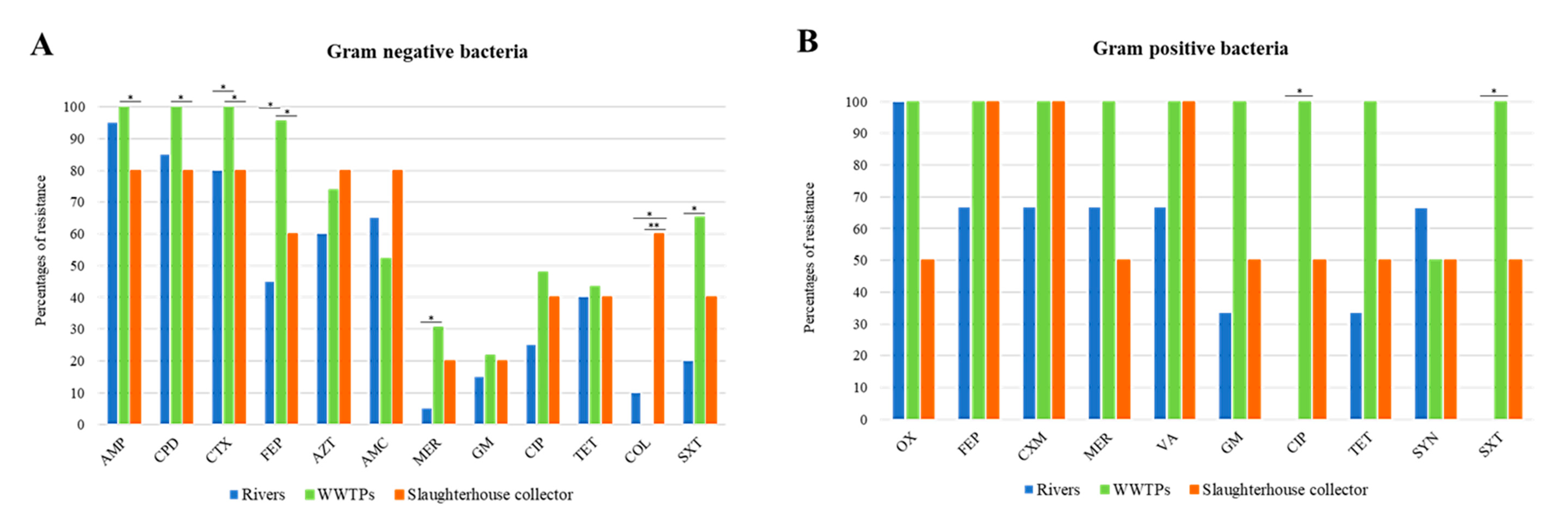
3.3. Multidrug Resistance Profiles
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Blaak, H.; Lynch, G.; Italiaander, R.; Hamidjaja, R.A.; Schets, F.M.; De Husman, A.M.R. Multidrug-resistant and extended spectrum beta-lactamase-producing Escherichia coli in dutch surface water and wastewater. PLoS ONE 2015, 10, e0127752. [Google Scholar] [CrossRef]
- Kümmerer, K. Resistance in the environment. J. Antimicrob. Chemother. 2004, 54, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Woolhouse, M. Antimicrobial resistance in humans, livestock and the wider environment. J. High Energy Phys. 2015, 6, 1575–1589. [Google Scholar] [CrossRef] [PubMed]
- Chee-Sanford, J.C.; Mackie, R.I.; Koike, S.; Krapac, I.G.; Lin, Y.-F.; Yannarell, A.C.; Maxwell, S.; Aminov, R.I. Fate and Transport of Antibiotic Residues and Antibiotic Resistance Genes following Land Application of Manure Waste. J. Environ. Qual. 2009, 38, 1086–1108. [Google Scholar] [CrossRef] [PubMed]
- García-Galán, M.J.; Díaz-Cruz, M.S.; Barceló, D. Occurrence of sulfonamide residues along the Ebro river basin. Removal in wastewater treatment plants and environmental impact assessment. Environ. Int. 2011, 37, 462–473. [Google Scholar] [CrossRef] [PubMed]
- Jäger, T.; Hembach, N.; Elpers, C.; Wieland, A.; Alexander, J.; Hiller, C.; Krauter, G.; Schwartz, T. Reduction of Antibiotic Resistant Bacteria During Conventional and Advanced Wastewater Treatment, and the Disseminated Loads Released to the Environment. Front. Microbiol. 2018, 9, 2599. [Google Scholar] [CrossRef] [PubMed]
- Ojer-Usoz, E.; González, D.; Vitas, A.I.; Leiva, J.; García-Jalón, I.; Febles-Casquero, A.; de la Soledad Escolano, M. Prevalence of extended-spectrum β-lactamase-producing Enterobacteriaceae in meat products sold in Navarra, Spain. Meat Sci. 2013, 93, 316–321. [Google Scholar] [CrossRef] [PubMed]
- Ojer-Usoz, E.; González, D.; García-Jalón, I.; Vitas, A.I. High dissemination of extended-spectrum β-lactamase-producing Enterobacteriaceae in effluents from wastewater treatment plants. Water Res. 2014, 56, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Leonard, A.F.C.; Zhang, L.; Balfour, A.J.; Garside, R.; Gaze, W.H. Human recreational exposure to antibiotic resistant bacteria in coastal bathing waters. Environ. Int. 2015, 82, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Guo, C.; Luo, Y.; Lv, J.; Zhang, Y.; Lin, H.; Wang, L.; Xu, J. Occurrence and distribution of antibiotics, antibiotic resistance genes in the urban rivers in Beijing, China. Environ. Pollut. 2016, 213, 833–840. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.T.; Yuan, Q.-B.; Yang, J. Microbial selectivity of UV treatment on antibiotic-resistant heterotrophic bacteria in secondary effluents of a municipal wastewater treatment plant. Water Res. 2013, 47, 6388–6394. [Google Scholar] [CrossRef]
- Hembach, N.; Schmid, F.; Alexander, J.; Hiller, C.; Rogall, E.T.; Schwartz, T. Occurrence of the mcr-1 colistin resistance gene and other clinically relevant antibiotic resistance genes in microbial populations at different municipal wastewater treatment plants in Germany. Front. Microbiol. 2017, 8, 1282. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics; WHO: Geneva, Switzerland, 2017. [Google Scholar]
- Falagas, M.E.; Karageorgopoulos, D.E. Extended-spectrum β-lactamase-producing organisms. J. Hosp. Infect. 2009, 73, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Zurfluh, K.; Hächler, H.; Nüesch-Inderbinen, M.; Stephan, R. Characteristics of extended-spectrum β-lactamase- and carbapenemase-producing Enterobacteriaceae isolates from rivers and lakes in Switzerland. Appl. Environ. Microbiol. 2013, 79, 3021–3026. [Google Scholar] [CrossRef] [PubMed]
- Zarfel, G.; Lipp, M.; Gürtl, E.; Folli, B.; Baumert, R.; Kittinger, C. Troubled water under the bridge: Screening of River Mur water reveals dominance of CTX-M harboring Escherichia coli and for the first time an environmental VIM-1 producer in Austria. Sci. Total Environ. 2017, 593–594, 399–405. [Google Scholar] [CrossRef]
- Stryjewski, M.E.; Corey, G.R. Methicillin-resistant Staphylococcus aureus: An evolving pathogen. Clin. Infect. Dis. 2014, 58, 10–19. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Nunes, O.C.; Manaia, C.M. Bacterial diversity and antibiotic resistance in water habitats: Searching the links with the human microbiome. FEMS Microbiol. Rev. 2014, 38, 761–778. [Google Scholar] [CrossRef]
- Porrero, M.C.; Harrison, E.; Fernández-Garayzábal, J.F.; Paterson, G.K.; Díez-Guerrier, A.; Holmes, M.A.; Domínguez, L. Detection of mecC-Methicillin-resistant Staphylococcus aureus isolates in river water: A potential role for water in the environmental dissemination. Environ. Microbiol. Rep. 2014, 6, 705–708. [Google Scholar] [CrossRef]
- Ahmed, M.O.; Baptiste, K.E. Vancomycin-Resistant Enterococci: A Review of Antimicrobial Resistance Mechanisms and Perspectives of Human and Animal Health. Microb. Drug Resist. 2018, 24, 590–606. [Google Scholar] [CrossRef]
- Lozano, C.; González-Barrio, D.; García, J.T.; Ceballos, S.; Olea, P.P.; Ruiz-Fons, F.; Torres, C. Detection of vancomycin-resistant Enterococcus faecalis ST6-vanB2 and E. faecium ST915-vanA in faecal samples of wild Rattus rattus in Spain. Vet. Microbiol. 2015, 177, 168–174. [Google Scholar] [CrossRef]
- Nakipoğlu, M.; Yilmaz, F.; Icgen, B. vanA Gene Harboring Enterococcal and Non-enterococcal Isolates Expressing High Level Vancomycin and Teicoplanin Resistance Reservoired in Surface Waters. Bull. Environ. Contam. Toxicol. 2017, 98, 712–719. [Google Scholar] [CrossRef] [PubMed]
- Lata, P.; Ram, S.; Shanker, R. Multiplex PCR based genotypic characterization of pathogenic vancomycin resistant Enterococcus faecalis recovered from an Indian river along a city landscape. Springerplus 2016, 5, 1199. [Google Scholar] [CrossRef]
- Müller, H.; Sib, E.; Gajdiss, M.; Klanke, U.; Lenz-Plet, F.; Barabasch, V.; Albert, C.; Schallenberg, A.; Timm, C.; Zacharias, N.; et al. Dissemination of multi-resistant Gram-negative bacteria into German wastewater and surface waters. FEMS Microbiol. Ecol. 2018, 94, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Guyonnet, J.; Manco, B.; Baduel, L.; Kaltsatos, V.; Aliabadi, M.H.F.S.; Lees, P. Determination of a dosage regimen of colistin by pharmacokinetic/pharmacodynamic integration and modeling for treatment of G.I.T. disease in pigs. Res. Vet. Sci. 2010, 88, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Rhouma, M.; Beaudry, F.; Letellier, A. Resistance to colistin: What is the fate for this antibiotic in pig production? Int. J. Antimicrob. Agents 2016, 48, 119–126. [Google Scholar] [CrossRef] [PubMed]
- McGann, P.; Snesrud, E.; Maybank, R.; Corey, B.; Ong, A.C.; Clifford, R.; Hinkle, M.; Whitman, T.; Lesho, E.; Schaecher, K.E. Erratum for McGann et al. Escherichia coli harboring mcr-1 and blaCTX-M on a novel IncF plasmid: First report of mcr-1 in the United States. Antimicrob. Agents Chemother. 2016, 60, 5107. [Google Scholar]
- International Organization for Standardization. Calidad del Agua. Muestreo para el Análisis Microbiológico; ISO 19458:2007; ISO: Geneva, Switzerland, 2007. [Google Scholar]
- International Organization for Standardization. Calidad del Agua. Muestreo Parte 3: Conservación y Manipulación de las Muestras de Agua; ISO 5667-3:2012; ISO: Geneva, Switzerland, 2012. [Google Scholar]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 28th ed.; CLSI: Wayne, PA, USA, 2018. [Google Scholar]
- The European Committee on Antimicrobial Susceptibility Testing (EUCAST). Breakpoint tables for interpretation of MICs and zone Diameters. Version 8.0. 2018. Available online: http://www.eucast.org (accessed on 15 September 2018).
- Jarlier, V.; Nicolas, M.; Fournier, G.; Philippon, A. Extended Broad-Spectrum β-Lactamases Conferring Transferable Resistance to Newer β- Lactam Agents in Enterobacteriaceae: Hospital Prevalence and Susceptibility Patterns. Rev. Infect. Dis. 1988, 10, 867–878. [Google Scholar] [CrossRef]
- Thean, Y.T.; Ng, L.S.Y.; He, J.; Tse, H.K.; Li, Y.H. Evaluation of screening methods to detect plasmid-mediated AmpC in Escherichia coli, Klebsiella pneumoniae, and Proteus mirabilis. Antimicrob. Agents Chemother. 2009, 53, 146–149. [Google Scholar] [CrossRef]
- Derbyshire, H.; Kay, G.; Evans, K.; Vaughan, C.; Kavuri, U.; Winstanley, T. A simple disc diffusion method for detecting AmpC and extended-spectrum β-lactamases in clinical isolates of Enterobacteriaceae. J. Antimicrob. Chemother. 2009, 63, 497–501. [Google Scholar] [CrossRef][Green Version]
- Arakawa, Y.; Shibata, N.; Shibayama, K.; Kurokawa, H.; Yagi, T.; Fujiwara, H.; Goto, M. Convenient test for screening metallo-β-lactamase-producing gram-negative bacteria by using thiol compounds. J. Clin. Microbiol. 2000, 38, 40–43. [Google Scholar]
- Pérez-pérez, F.J.; Hanson, N.D. Detection of Plasmid-Mediated AmpC β-Lactamase Genes in Clinical Isolates by Using Multiplex PCR. J. Clin. Microbiol. 2002, 40, 2153–2162. [Google Scholar] [CrossRef] [PubMed]
- Colom, K.; Pérez, J.; Alonso, R.; Fernández-Aranguiz, A.; Lariño, E.; Cisterna, R. Simple and reliable multiplex PCR assay for detection of blaTEM, bla(SHV) and blaOXA-1 genes in Enterobacteriaceae. FEMS Microbiol. Lett. 2003, 223, 147–151. [Google Scholar] [CrossRef]
- Woodford, N.; Fagan, E.J.; Ellington, M.J. Multiplex PCR for rapid detection of genes encoding CTX-M extended-spectrum β-lactamases [4]. J. Antimicrob. Chemother. 2006, 57, 154–155. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Y.; Wang, Y.; Walsh, T.R.; Yi, L.X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Xavier, B.B.; Lammens, C.; Ruhal, R.; Butaye, P.; Goossens, H. Identification of a novel plasmid-mediated colistin- resistance gene, mcr-2, in Escherichia coli, Belgium, June. Euro Surveill. 2016, 21, 6–11. [Google Scholar] [CrossRef] [PubMed]
- Yin, W.J.; Li, H.; Shen, Y.B.; Liu, Z.; Wang, S.; Shen, Z.; Zhang, R.; Timothy, R.; Walsh, J.S.; Wanga, Y. Novel Plasmid-Mediated Colistin Resistance Gene mcr-3 in Escherichia coli Wenjuan. MBio 2017, 8, e00543-17. [Google Scholar]
- Carattoli, A.; Villa, L.; Feudi, C.; Curcio, L.; Orsini, S.; Luppi, A.; Pezzotti, G.; Magistrali, C.F. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Eurosurveillance 2017, 22, 1–5. [Google Scholar] [CrossRef]
- Borowiak, M.; Fischer, J.; Hammerl, J.A.; Hendriksen, R.S.; Szabo, I.; Malorny, B. Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J. Antimicrob. Chemother. 2017, 72, 3317–3324. [Google Scholar] [CrossRef]
- Komatsu, M.; Aihara, M.; Shimakawa, K.; Iwasaki, M.; Nagasaka, Y.; Fukuda, S.; Matsuo, S.; Iwatani, Y. Evaluation of MicroScan ESBL confirmation panel for Enterobacteriaceae-producing, extended-spectrum β-lactamases isolated in Japan. Diagn. Microbiol. Infect. Dis. 2003, 46, 125–130. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Savin, M.; Bierbaum, G.; Hammerl, J.A.; Heinemann, C.; Parcina, M.; Sib, E.; Voigt, A.; Kreyenschmidt, J. ESKAPE Bacteria and Extended-Spectrum-β-Lactamase-Producing Escherichia coli Isolated from Wastewater and Process Water from German Poultry Slaughterhouses. Appl. Environ. Microbiol. 2020, 86, e02748-19. [Google Scholar] [CrossRef] [PubMed]
- Makowska, N.; Philips, A.; Dabert, M.; Nowis, K.; Trzebny, A.; Koczura, R.; Mokracka, J. Metagenomic analysis of β-lactamase and carbapenemase genes in the wastewater resistome. Water Res. 2020, 170, 115277. [Google Scholar] [CrossRef] [PubMed]
- Osińska, A.; Harnisz, M.; Korzeniewska, E. Prevalence of plasmid-mediated multidrug resistance determinants in fluoroquinolone-resistant bacteria isolated from sewage and surface water. Environ. Sci. Pollut. Res. 2016, 23, 10818–10831. [Google Scholar] [CrossRef]
- Argudín, M.A.; Deplano, A.; Meghraoui, A.; Dodémont, M.; Heinrichs, A.; Denis, O.; Nonhoff, C.; Roisin, S. Bacteria from animals as a pool of antimicrobial resistance genes. Antibiotics 2017, 6, 12. [Google Scholar] [CrossRef]
- Ovejero, C.M.; Delgado-Blas, J.F.; Calero-Caceres, W.; Muniesa, M.; Gonzalez-Zorn, B. Spread of mcr-1-carrying Enterobacteriaceae in sewage water from Spain. J. Antimicrob. Chemother. 2017, 72, 1050–1053. [Google Scholar]
- Galler, H.; Feierl, G.; Petternel, C.; Reinthaler, F.F.; Haas, D.; Habib, J.; Kittinger, C.; Luxner, J.; Zarfel, G. Multiresistant bacteria isolated from activated sludge in Austria. Int. J. Environ. Res. Public Health 2018, 15, 479. [Google Scholar] [CrossRef]
- Paulshus, E.; Kühn, I.; Möllby, R.; Colque, P.; O’Sullivan, K.; Midtvedt, T.; Lingaas, E.; Holmstad, R.; Sørum, H. Diversity and antibiotic resistance among Escherichia coli populations in hospital and community wastewater compared to wastewater at the receiving urban treatment plant. Water Res. 2019, 161, 232–241. [Google Scholar] [CrossRef]
- Lozano, C.; Gonzalez-Barrio, D.; Camacho, M.C.; Lima-Barbero, J.F.; de la Puente, J.; Höfle, U.; Torres, C. Characterization of fecal vancomycin-resistant Enterococci with acquired and intrinsic resistance mechanisms in wild animals, Spain. Microb. Ecol. 2016, 72, 813–820. [Google Scholar] [CrossRef]
- Kristich, C.J.; Rice, L.B.; Arias, C.A. Enterococcal Infection−Treatment and Antibiotic Resistance. In Enterococci: From Commensals to Leading Causes of Drug Resistant Infection; Massachusetts Eye and Ear Infirmary: Boston, MA, USA, 2014; pp. 123–185. [Google Scholar]
- Porrero, M.C.; Valverde, A.; Fernández-Llario, P.; Díez-Guerrier, A.; Mateos, A.; Lavín, S.; Cantón, R.; Fernández-Garayzabal, J.F.; Domínguez, L. Staphylococcus aureus carrying mecC gene in animals and Urban Wastewater, Spain. Emerg. Infect. Dis. 2014, 20, 899–901. [Google Scholar] [CrossRef]
- Boswihi, S.S.; Udo, E.E.; Mathew, B.; Noronha, B.; Verghese, T.; Tappa, S.B. Livestock-Associated Methicillin-Resistant Staphylococcus aureus in Patients Admitted to Kuwait Hospitals in 2016–2017. Front. Microbiol. 2020, 10, 2912. [Google Scholar] [CrossRef] [PubMed]
- EU Directive 91/271/ECC sobre el tratamiento de las aguas residuales urbanas; Diario oficial de la Unión Europea L135; European Union: Brussels, Belgium, 1991.
- Pallares-Vega, R.; Blaak, H.; van der Plaats, R.; de Roda Husman, A.M.; Hernandez Leal, L.; van Loosdrecht, M.C.M.; Weissbrodt, D.G.; Schmitt, H. Determinants of presence and removal of antibiotic resistance genes during WWTP treatment: A cross-sectional study. Water Res. 2019, 161, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Munir, M.; Wong, K.; Xagoraraki, I. Release of antibiotic resistant bacteria and genes in the effluent and biosolids of five wastewater utilities in Michigan. Water Res. 2011, 45, 681–693. [Google Scholar] [CrossRef] [PubMed]
- Jia, S.; Bian, K.; Shi, P.; Ye, L.; Liu, C.H. Metagenomic profiling of antibiotic resistance genes and their associations with bacterial community during multiple disinfection regimes in a full-scale drinking water treatment plant. Water Res. 2020, 176, 115721. [Google Scholar] [CrossRef]
- López Martínez, A. Evaluación de Tencologías en el Tratamiento de Aguas y Fangos en EDARs, para la Reducción de Microorganismos con Riesgo Sanitario y Ambiental; Universidad de Zaragoza: Zaragoza, España, 2018. [Google Scholar]



| Target | Primer | Sequence (5′–3′) | Amplicon Size (bp) | Conditions |
|---|---|---|---|---|
| blaMOXM | blaMOXM-Fw | GCTGCTCAAGGAGCACAGGAT | 520 | Initial denaturation at 94 °C for 3 min; 25 cycles of amplification: denaturation at 94 °C for 30 s, hybridization at 64 °C for 30 s and extension at 72 °C for 1 min; final elongation at 72 °C for 7 min. |
| blaMOXM-Rv | CACATTGACATAGGTGTGGTGC | |||
| blaCITM | blaCITM-Fw | TGGCCAGAACTGACAGGCAAA | 462 | |
| blaCITM-Rv | TTTCTCCTGAACGTGGCTGGC | |||
| blaDHAM | blaDHAM-Fw | AACTTTCACAGGTGTGCTGGGT | 405 | |
| blaDHAM-Rv | CCGTACGCATACTGGCTTTGC | |||
| blaACCM | blaACCM-Fw | AACAGCCTCAGCAGCCGGTTA | 346 | |
| blaACCM-Rv | TTCGCCGCAATCATCCCTAGC | |||
| blaEBC | blaEBC-Fw | TCGGTAAAGCCGATGTTGCGG | 302 | |
| blaEBC-Rv | CTTCCACTGCGGCTGCCAGTT | |||
| blaFOX | blaFOX-Fw | AACATGGGGTATCAGGGAGATG | 190 | |
| blaFOX-Rv | CAAAGCGCGTAACCGGATTGG | |||
| blaSHV | blaSHV-Fw | AGGATTGACTGCCTTTTTG | 392 | Initial denaturation at 94 °C for 5 min; 32 cycles of amplification: denaturation at 94 °C for 30 s, hybridization at 54 °C for 30 s and extension at 72 °C for 1 min; final elongation at 72 °C for 10 min. |
| blaSHV-Rv | ATTTGCTGATTTCGCTCG | |||
| blaTEM | blaTEM-Fw | ATCAGCAATAAACCAGC | 516 | |
| blaTEM-Rv | CCCCGAAGAACGTTTTC | |||
| blaOXA | blaOXA-Fw | ATATCTCTACTGTTGCATCTCC | 619 | |
| blaOXA-Rv | AAACCCTTCAAACCATCC | |||
| blaCTX-M1 | blaCTX-M1-Fw | AAAAATCACTGGCCAGTTC | 415 | Initial denaturation at 94 °C for 5 min; 30 cycles of amplification: denaturation at 94 °C for 45 s, hybridization at 55 °C for 30 s and extension at 72 °C for 1 min; final elongation at 72 °C for 6 min. |
| blaCTX-M1-Rv | AGCTTATTCATCGCCACGTT | |||
| blaCTX-M2 | blaCTX-M2-Fw | CGACGCTACCCCTGCTATT | 552 | |
| blaCTX-M2-Rv | CCAGCGTCAGATTTTTCAGG | |||
| blaCTX-M9 | blaCTX-M9-Fw | CAAAGAGAGTGCAACGGATG | 205 | |
| blaCTX-M9-Rv | ATTGGAAAGCGTTCATCACC | |||
| blaCTX-M8 | blaCTX-M8-Fw | TCGCGTTAAGCGGATGATGC | 666 | |
| blaCTX-M25 | blaCTX-M25-Fw | GCACGATGACATTCGGG | 327 | |
| blaCTX-M8/25 | blaCTX-M1-Rv | AACCCACGATGTGGGTAGC | 666/327 |
| Target | Primer | Sequence (5′–3′) | Amplicon Size (bp) | Conditions |
| mcr-1 | mcr-1-Fw | CGGTCAGTCCGTTTGTTC | 309 | 20 cycles of amplification at 94 °C for 30 s, 58 °C for 90 s, 72 °C for 1 min and final 72 °C for 10 min |
| mcr-1-Rv | CTTGGTCGGTCTGTAGGG | |||
| mcr-2 | mcr-2-Fw | TGTTGCTTGTGCCGATTGGA | 567 | 33 cycles of amplification at 95 °C for 3 min, 65 °C for 30 s, 72 °C for 1 min and final 72 °C for 10 min. |
| mcr-2-Rv | AGATGGTATTGTTGGTTGCTG | |||
| mcr-3 | mcr-3-Fw | TTGGCACTGTATTTTGCATTT | 542 | 30 cycles of amplification at 95 °C for 30 s, 50 °C for 30 s, 72 °C for 45 s and final 72 °C for 7 min. |
| mcr-3-Rv | TTAACGAAATTGGCTGGAACA | |||
| mcr-4 | mcr-4-Fw | ATTGGGATAGTCGCCTTTTT | 487 | 20 cycles of amplification at 94 °C for 30 s, 58 °C for 90 s, 72 °C for 1 min and final 72 °C for 10 min |
| mcr-4-Rv | TTACAGCCAGAATCATTATCA | |||
| mcr-5 | mcr-5-Fw | ATGCGGTTGTCTGCATTTATC | 1644 | 30 cycles of amplification at 95 °C for 30 s, 50 °C for 30 s, 72 °C for 95 s and final 72 °C for 5 min. |
| mcr-5-Rv | TCATTGTGGTTGTCCTTTTCTG |
| Sample point | CPD 1st SE | CPD 2nd SE | FOX 1st SE | FOX 2nd SE | VA 1st SE | VA 2nd SE | CARB 1st SE | CARB 2nd SE | COL 1st SE | COL 2nd SE | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1_ASE | ||||||||||||||
| 2_ASE | ||||||||||||||
| 3_ASE | ||||||||||||||
| 4_ASE | ||||||||||||||
| 5_ASE | ||||||||||||||
| 6_ASE | ||||||||||||||
| 7_ASE | ||||||||||||||
| 8_ASE | ||||||||||||||
| 9_ASE | ||||||||||||||
| 10_ASE | ||||||||||||||
| 11_ASE | ||||||||||||||
| 12_ASE | ||||||||||||||
| 13_ASE | ||||||||||||||
| 14_ASE | ||||||||||||||
| 15_ASE | ||||||||||||||
| 16_ASE | ||||||||||||||
| 17_ASE | ||||||||||||||
| 18_ASE | ||||||||||||||
| 19_ASE | ||||||||||||||
| 20_ASE | ||||||||||||||
| 21_ASC | ||||||||||||||
| 22_ASC | ||||||||||||||
| 23_ASA | ||||||||||||||
| 24_ASA | ||||||||||||||
| 25_ASA | ||||||||||||||
| 26_ASA | ||||||||||||||
| 27_ASA | ||||||||||||||
| 28_ASA | ||||||||||||||
| 29_ARD1e | ||||||||||||||
| 30_ARD1s | ||||||||||||||
| 31_ARD2e | ||||||||||||||
| 32_ARD2s | ||||||||||||||
| 33_ARD3e | ||||||||||||||
| 34_ARD3s | ||||||||||||||
| 35_ARD4e | ||||||||||||||
| 36_ARD4s | ||||||||||||||
| 37_ARH | ||||||||||||||
| 38_ARM | ||||||||||||||
| 39_ARM | ||||||||||||||
| 40_ARM |
| Antimicrobial Resistance | % Positive Rivers | % Positive WWTP/C | N Total Isolates | N of Confirmed Target ARB (%) | Target ARB | ||||
|---|---|---|---|---|---|---|---|---|---|
| 1st SE | 2nd SE | 1st SE | 2nd SE | Rivers | WWTP/C | Rivers | WWTP/C | ||
| Cefpodoxime | 75 | 50 | 83.3 | 100 | 45 | 25 | 18 (40) | 19 (75) | ESBL Enterobacteriaceae |
| Cefoxitin | 71.4 | 75 | 83.3 | 100 | 89 | 39 | 2 (2.3) | 0 | S. aureus MRSA |
| Carbapenems | 32.1 | 92.8 a | 41.6 | 91.6 a | 50 | 27 | 0 | 0 | A. baumanni |
| 0 | 2 (7.4) | P. aeruginosa | |||||||
| 0 | 5 (18.5) | Enterobacteriaceae | |||||||
| Vancomycin | 42.9 | 32.1 | 75 | 91.6 | 8 | 15 | 1 (12.5) | 4 (26.6) | E. faecium |
| Colistin | 3.6 | 53.6 b | 8.3 | 66.6 b | 8 | 5 | 2 (25) | 2 (40) | Enterobacteriaceae |
| Total | 200 | 111 | 23 (11.5) | 32 (28.9) | |||||
| Nº Strain | Samples Code | Species | Resistance Genes | AmpC | Antimicrobial Resistance | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Penicillins | Cephalosporins | Monobactams | β-lactamase Inhibitors | Carbapenems | AminogLycosides | Quinolones | Tetracyclines | Others | MDR | Extensive MDR | |||||
| 1 | 3_ASE | E. coli | TEML-278, CTX-M14 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMS | - | GM, TO | LV, CIP, MXF, NOR | TET | SXT, FOT, CHL | + | + |
| 2 | 3_ASE | E. coli | TEML-278, CTX-M14 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, FOX, FEP | AZT | AMC, AMS | - | GM, TO | LV, CIP, MXF, NOR | TET | SXT, FOT, CHL | + | + |
| 3 | 4_ASE | E. coli | TEML-278, CTX-M14 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, FOX, FEP | AZT | AMC, AMS | - | GM, TO | LV, CIP, MXF, NOR | TET | SXT, FOT, CHL | + | + |
| 4 | 6_ASE | E. coli | CTX-M15 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS | MER | - | LV, CIP, MXF, NOR | TET, TIG | SXT, CHL | + | + |
| 5 | 8_ASE | E. coli | TEML-278, CTX-M14 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMS | - | - | LV, CIP, MXF, NOR | TET | - | + | + |
| 6 | 17_ASE | E. coli | CTX-M1 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | - | TET | CHL | + | + |
| 7 | 17_ASE | E. coli | CTX-M1 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | - | TET | CHL | + | + |
| 8 | 29_ARD1e | E. coli | CTX-M1 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMS | - | - | - | TET | - | + | + |
| 9 | 30_ARD1s | E. coli | TEML-278, CTX-M14 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | - | - | - | - | LV, CIP, MXF, NOR | - | - | + | - |
| 10 | 30_ARD1s | E. coli | CTX-M14 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMS | - | - | - | - | SXT, FOT | + | + |
| 11 | 31_ARD2e | E. coli | CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | - | - | SXT | + | - |
| 12 | 32_ARD2s | E. coli | CTX-M1, SHV-12 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | LV, CIP, MXF, NOR | - | CHL | + | + |
| 13 | 32_ARD2s | E. coli | TEML-278, CTX-M1, SHV-12 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS, TZP | - | - | LV, CIP, MXF, NOR | TET | SXT, CHL | + | + |
| 14 | 33_ARD3e | E. coli | TEM-171, CTX-M1 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMS | - | - | - | - | - | + | - |
| 15 | 34_ARD3s | E. coli | TEML-278, CTX-M14 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | - | AMS | - | GM, TO | - | TET | SXT | + | + |
| 16 | 34_ARD3s | E. coli | TEML-278, CTX-M14 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | - | AMS | - | GM, TO | - | TET | SXT | + | + |
| 17 | 35_ARD4e | E. coli | TEML-278, CTX-M1 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | GM, TO | LV, CIP, MXF, NOR | TET | SXT, FOT | + | + |
| 18 | 7_ASE | E. coli | TEML-278, SHV-12 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | - | TET | + | - | |
| 19 | 30_ARD1s | E. coli | TEM-171, CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | - | - | - | - | TET | + | - | |
| 20 | 32_ARD2s | E. coli | TEML-278, CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS | - | - | - | - | FOT | + | + |
| 21 | 35_ARD4e | E. coli | TEM-176, CTX-M15, CTX-M14 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | - | - | - | - | - | - | - | - | |
| 22 | 36_ARD4s | E. coli | OXA-1, CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS | - | TO | LV, CIP, MXF, NOR | - | SXT | + | + |
| 23 | 40_ARM | E. coli | CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS | - | - | LV, CIP, MXF, NOR | TET | SXT, CHL | + | + |
| 24 | 7_ASE | S. fonticola | CTX-M1 | + | AMP, MZ | CZ, CXM, CPD, CTX, CAZ | AZT | AMC | - | - | - | - | - | + | - |
| 25 | 11_ASE | S. fonticola | - | + | AMP | CZ, CXM, CPD, CTX | - | AMC | - | - | - | - | - | + | - |
| 26 | 11_ASE | S. fonticola | CTX-M1 | + | AMP | CZ, CXM, CPD, CTX, CAZ | AZT | AMC, AMS | - | - | - | - | - | + | - |
| 27 | 11_ASE | S. fonticola | CTX-M15 | + | AMP, PIP | CZ, CXM, CPD, CTX | - | AMC, AMS | - | - | MXF | - | FOT | + | + |
| 28 | 20_ASE | S. fonticola | - | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ | AZT | AMC, AMS | - | - | - | - | - | + | - |
| 29 | 3_ASE | S. fonticola | TEM-171 | + | AMP | CZ, CXM, CPD, CTX | - | AMC | ETP | - | - | - | - | + | - |
| 30 | 5_ASE | S. fonticola | TEM-171, ACC | + | AMP, PIP | CZ, CXM, CPD, CTX | - | AMC, AMS, TZP | - | - | - | - | - | + | - |
| 31 | 11_ASE | S. fonticola | TEM-171 | + | AMP | CZ, CXM, CPD, CTX | - | AMC | - | - | - | - | - | + | - |
| 32 | 39_ARM | S. fonticola | CTX-M15, ACC | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX | AZT | AMC, AMS | - | - | - | - | COL | + | + |
| 33 | 8_ASE | S. liquefaciens | - | + | AMP | CZ, CXM, CPD | - | AMC | - | - | - | - | - | + | - |
| 34 | 33_ARD3e | K. pneumoniae | TEM-171, SHV-12, CTX-M1, DHA | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS | MER | - | LV, CIP, MXF, NOR | TET | SXT, FOT, CHL | + | + |
| 35 | 35_ARD4e | K. pneumoniae | CTX-M14 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS, TZP | MER | - | LV, CIP, MXF, NOR | TET | SXT, FOT, CHL | + | + |
| 36 | 35_ARD4e | K. pneumoniae | TEML-278, SHV-12, OXA-1, CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS | - | TO | LV, CIP, MXF, NOR | - | SXT, CHL | + | + |
| 37 | 11_ASE | K. oxytoca | - | + | AMP | CZ, FOX, FEP | AZT | AMC | - | - | - | - | - | + | - |
| 38 | 33_ARD3e | E. coli | TEML-278, KPC | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, FOX, FEP | AZT | AMC, AMS, TZP | ETP, MER, IMP | - | LV, CIP, MXF, NOR | - | SXT, FOT | + | + |
| 39 | 33_ARD3e | K. oxytoca | TEML-278, KPC | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FEP | AZT | AMC, AMS, TZP | ETP, MER, IMP | TO | LV, CIP, MXF, NOR | - | SXT, FOT | + | + |
| 40 | 36_ARD4s | C. freundii | CTX-M1 | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | - | AMC, AMS, TZP | IMP | - | MXF | - | SXT | + | + |
| 41 | 33_ARD3e | C. freundii | TEML-278, EBC, DHA, KPC | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS, TZP | ETP, MER, IMP | GM, TO | MXF | - | FOT | + | + |
| 42 | 39_ARM | C. freundii | TEML-278, EBC, KPC | + | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS, TZP | ETP, MER, IMP | GM, TO | NOR | - | FOT | + | + |
| 43 | 33_ARD3e | P. aeruginosa | TEML-278 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS | MER, IMP | AM, GM, TO | LV, CIP, NOR | TET | SXT, FOT, CHL | + | + |
| 44 | 29_ARD1e | P. aeruginosa | - | + | AMP, MZ | CZ, CXM, CPD, CTX, FOX | - | AMC, AMS | ETP, MER, IMP | - | - | TET | SXT, FOT, CHL | + | + |
| 45 | 22_ASC | E. coli | - | - | - | - | - | - | - | - | - | - | COL | - | - |
| 46 | 39_ARM | E. coli | CTX-M15 | - | AMP, PIP, MZ | CZ, CXM, CPD, CTX, CAZ, FOX, FEP | AZT | AMC, AMS, TZP | - | - | - | - | COL, FOT | + | + |
| 47 | 40_ARM | E. coli | mcr-1 | - | - | - | - | - | - | - | LV, CIP, MXF, NOR | TET | COL, SXT | + | - |
| 48 | 1_ASE | K. oxytoca | - | - | AMP | - | - | - | - | - | - | - | COL, FOT | + | - |
| Nº Strain | Samples Code | Species | Resistance Genes | Antimicrobial Resistance | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Penicillins | Cephalosporins | Carbapenems | Glycopeptides | Aminoglycosides | Macrolides | Quinolones | Tetracyclines | Others | MDR | Extensive MDR | ||||
| 49 | 14_ASE | E. faecium | vanB | OX | CXM, FEP | ETP, MER | VA | GM | ERY | - | TET | FA, Cd, FOT, SYN | + | + |
| 50 | 40_ARM | E. faecium | vanB | - | CXM, FEP | - | VA | - | ERY | - | - | FA, MUP, DAP, Cd, FOT | + | + |
| 51 | 29_ARD1e | E. faecium | vanB | OX | CXM, FEP | ETP, MER | VA | GM, TO | ERY | LV, CIP, MXF | TET | FA, SXT, Cd, FOT, SYN | + | + |
| 52 | 33_ARD3e | E. faecium | vanB | P, OX | CXM, FEP | ETP, MER | VA | GM, TO | ERY | LV, CIP, MXF | TET | FA, SXT, DAP, Cd, FOT, RA, FD, SYN | + | + |
| 53 | 40_ARM | E. faecium | vanB | OX | CXM, FEP | ETP, MER | VA | GM | ERY | LV, CIP, MXF | TET | FA, SXT, Cd, FD, SYN | + | + |
| 54 | 4_ASE | S. aureus | - | AMP, P, OX | CXM, FOX, FEP | ETP, MER, IMP | - | - | AZI, ERY | - | - | MUP, Cd, FOT | + | + |
| 55 | 15_ASE | S. aureus | - | AMP, P, OX | FOX | - | VA, TEC | - | ERY | - | - | LZ, DAP, Cd, SYN | + | + |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pérez-Etayo, L.; González, D.; Leiva, J.; Vitas, A.I. Multidrug-Resistant Bacteria Isolated from Different Aquatic Environments in the North of Spain and South of France. Microorganisms 2020, 8, 1425. https://doi.org/10.3390/microorganisms8091425
Pérez-Etayo L, González D, Leiva J, Vitas AI. Multidrug-Resistant Bacteria Isolated from Different Aquatic Environments in the North of Spain and South of France. Microorganisms. 2020; 8(9):1425. https://doi.org/10.3390/microorganisms8091425
Chicago/Turabian StylePérez-Etayo, Lara, David González, José Leiva, and Ana Isabel Vitas. 2020. "Multidrug-Resistant Bacteria Isolated from Different Aquatic Environments in the North of Spain and South of France" Microorganisms 8, no. 9: 1425. https://doi.org/10.3390/microorganisms8091425
APA StylePérez-Etayo, L., González, D., Leiva, J., & Vitas, A. I. (2020). Multidrug-Resistant Bacteria Isolated from Different Aquatic Environments in the North of Spain and South of France. Microorganisms, 8(9), 1425. https://doi.org/10.3390/microorganisms8091425