SARS-CoV-2 Spike Pseudoviruses: A Useful Tool to Study Virus Entry and Address Emerging Neutralization Escape Phenotypes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Pseudoviruses
2.2. Serum and Monoclonal Antibodies
2.3. Pseudovirus-Based Neutralization Assay
3. Results
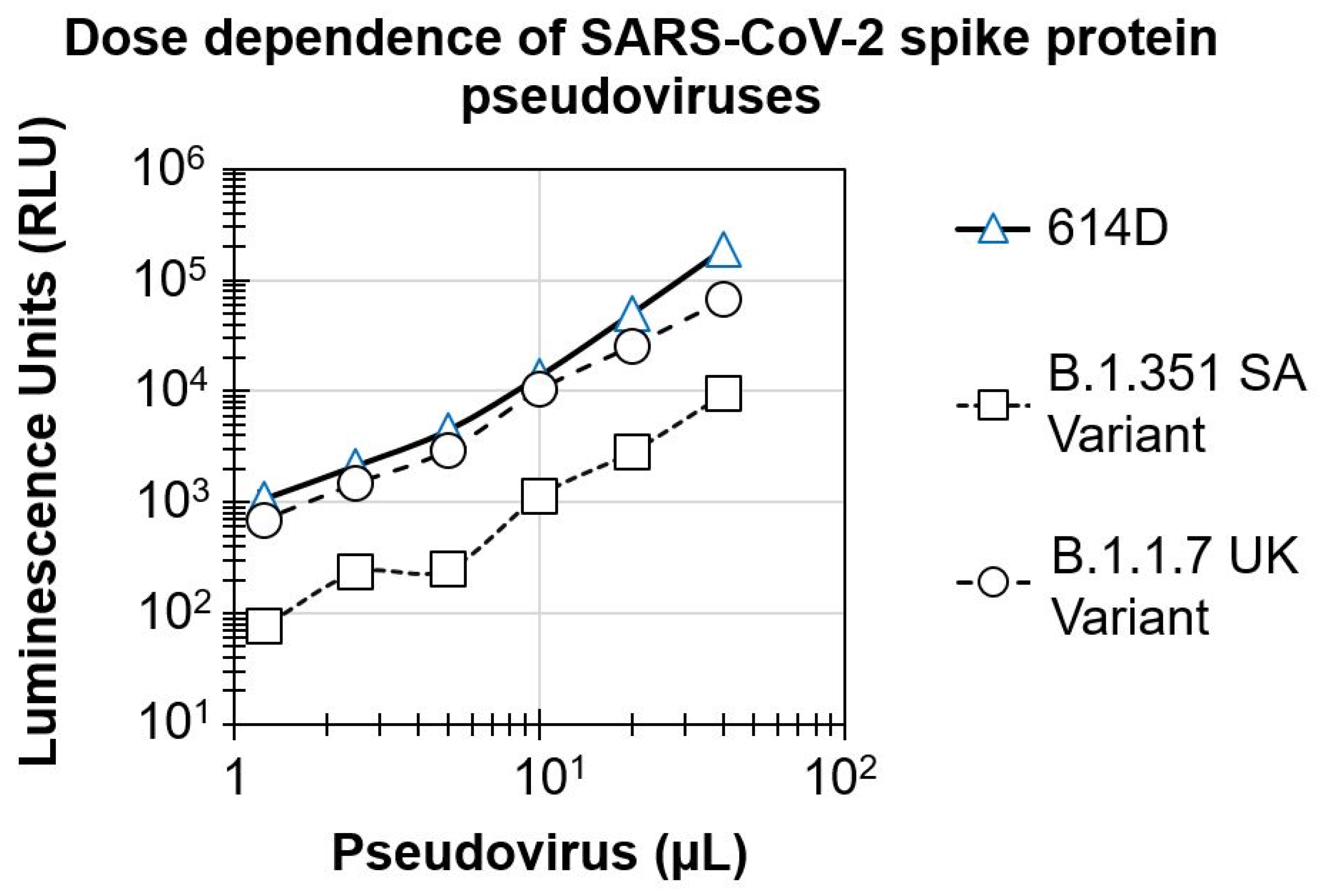
3.1. Dose Dependence of SARS-CoV-2 Spike Protein Pseudovirus Variants
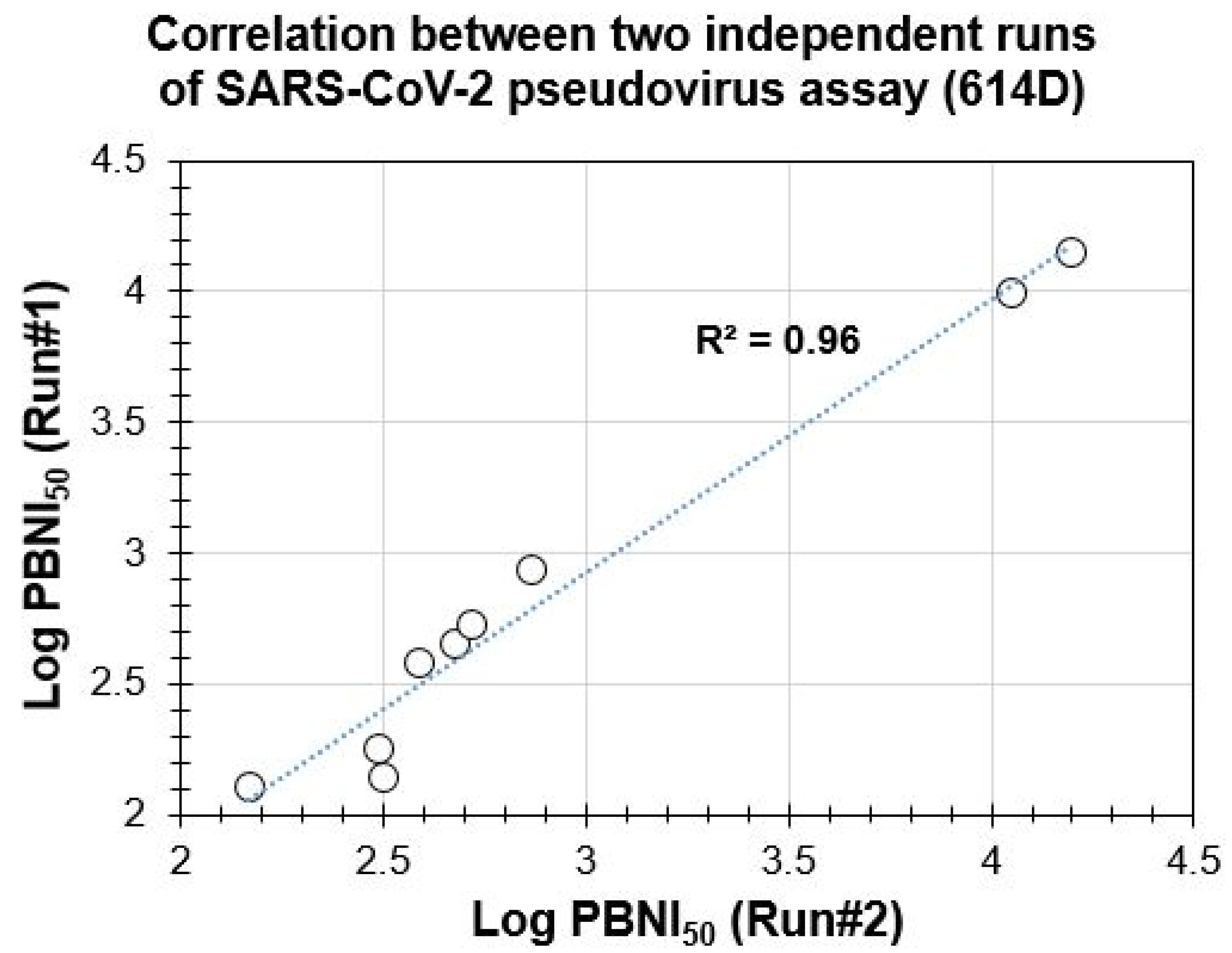
3.2. Suitability of PBNA for Evaluation of Neutralization Antibodies and Reproducibility of the SARS-CoV-2 Spike Protein Pseudovirus Assay
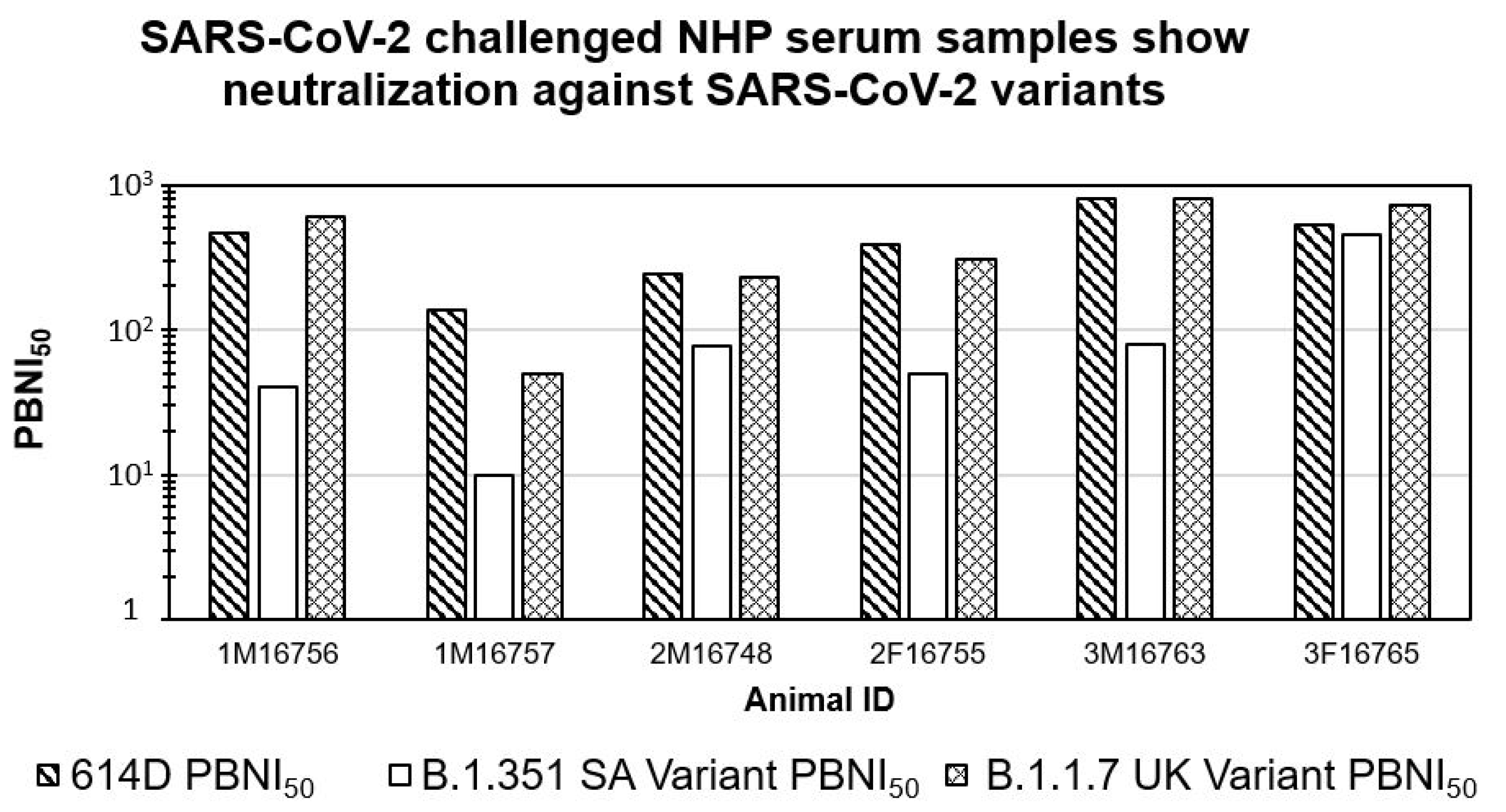
3.3. SARS-CoV-2 Challenged NHP Serum Samples Show Neutralization against SARS-CoV-2 614D and B.1.1.7 UK Variants
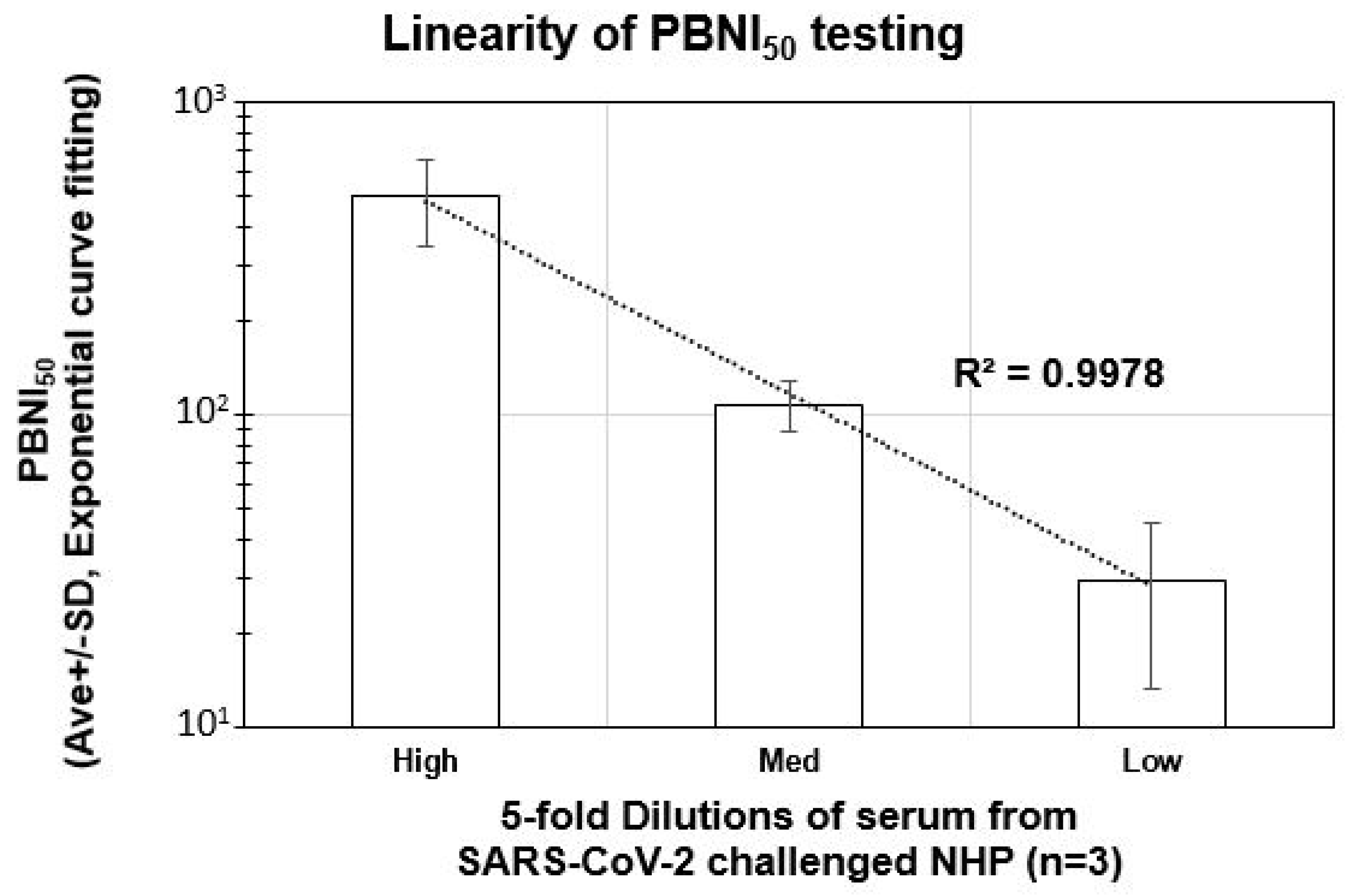
3.4. Linearity of Neutralization Index (PBNI50) Generated by the Pseudovirus Assay
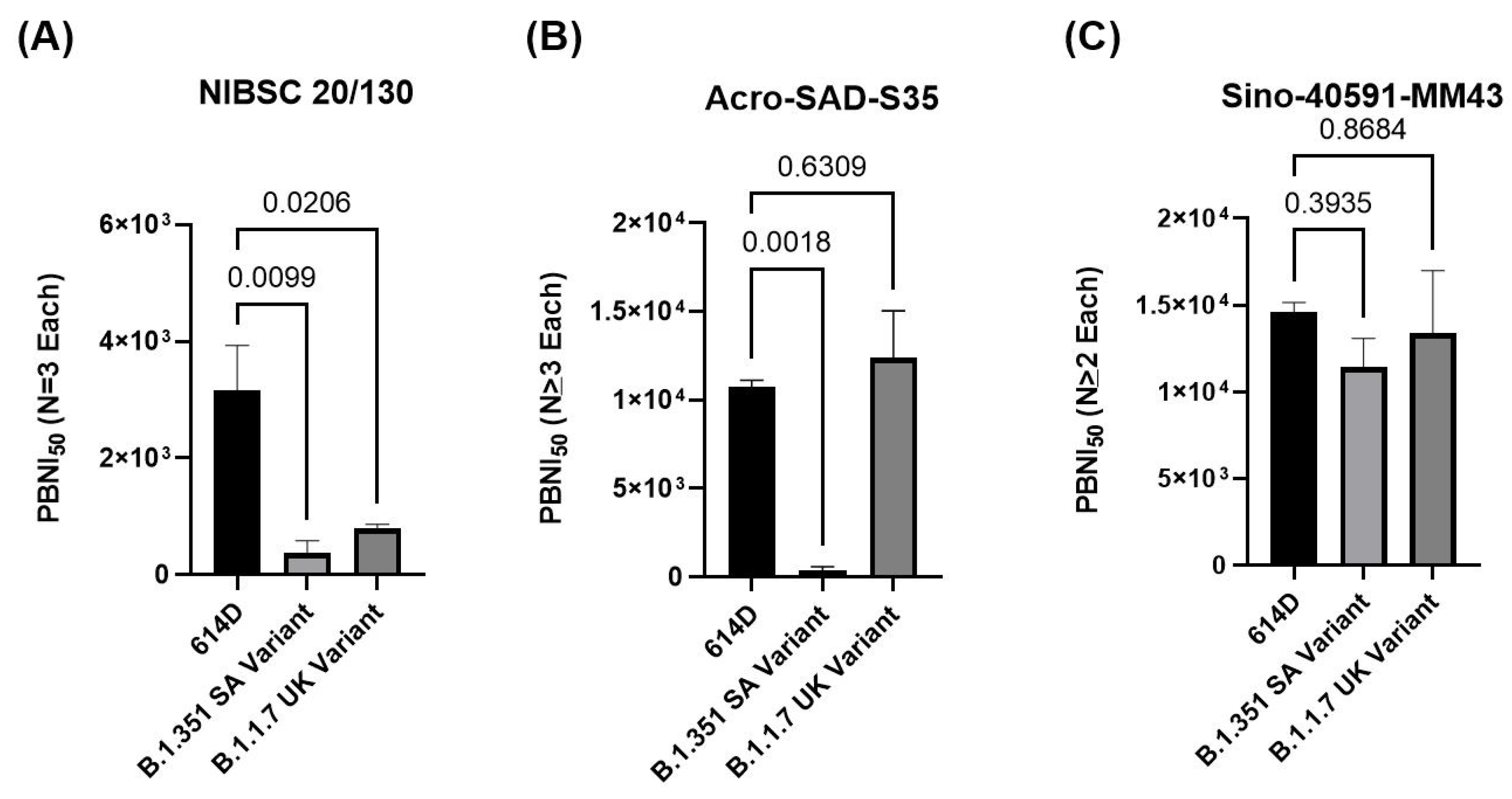
3.5. Comparison of Different Positive Controls in PBNA to Evaluate Neutralization against Variants of Concern
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 12 July 2021).
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Consortium, C.-G.U.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef]
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Dai, L.; Gao, G.F. Viral targets for vaccines against COVID-19. Nat. Rev. Immunol. 2021, 21, 73–82. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, X.; Tang, H.; McDanal, C.; Wagh, K.; Fischer, W.; Theiler, J.; Yoon, H.; Li, D.; Haynes, B.F.; Sanders, K.O.; et al. SARS-CoV-2 variant B.1.1.7 is susceptible to neutralizing antibodies elicited by ancestral spike vaccines. Cell Host Microbe 2021, 29, 529–539.e3. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F. Correlates of protection from SARS-CoV-2 infection. Lancet 2021, 397, 1421–1423. [Google Scholar] [CrossRef]
- Amanat, F.; Stadlbauer, D.; Strohmeier, S.; Nguyen, T.H.O.; Chromikova, V.; McMahon, M.; Jiang, K.; Arunkumar, G.A.; Jurczyszak, D.; Polanco, J.; et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 2020, 26, 1033–1036. [Google Scholar] [CrossRef] [PubMed]
- Caruana, G.; Croxatto, A.; Coste, A.T.; Opota, O.; Lamoth, F.; Jaton, K.; Greub, G. Diagnostic strategies for SARS-CoV-2 infection and interpretation of microbiological results. Clin. Microbiol. Infect. 2020, 26, 1178–1182. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; et al. Evaluation of Nucleocapsid and Spike Protein-Based Enzyme-Linked Immunosorbent Assays for Detecting Antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58, e00461-20. [Google Scholar] [CrossRef] [Green Version]
- Amanat, F.; White, K.M.; Miorin, L.; Strohmeier, S.; McMahon, M.; Meade, P.; Liu, W.C.; Albrecht, R.A.; Simon, V.; Martinez-Sobrido, L.; et al. An In Vitro Microneutralization Assay for SARS-CoV-2 Serology and Drug Screening. Curr. Protoc. Microbiol. 2020, 58, e108. [Google Scholar] [CrossRef] [PubMed]
- Manenti, A.; Maggetti, M.; Casa, E.; Martinuzzi, D.; Torelli, A.; Trombetta, C.M.; Marchi, S.; Montomoli, E. Evaluation of SARS-CoV-2 neutralizing antibodies using a CPE-based colorimetric live virus micro-neutralization assay in human serum samples. J. Med. Virol. 2020, 92, 2096–2104. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.A.; Mok, C.K.; Tsang, O.T.; Lv, H.; Ko, R.L.; Wu, N.C.; Yuan, M.; Leung, W.S.; Chan, J.M.; Chik, T.S.; et al. Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Euro Surveill. 2020, 25, 2000421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozcurumez, M.K.; Ambrosch, A.; Frey, O.; Haselmann, V.; Holdenrieder, S.; Kiehntopf, M.; Neumaier, M.; Walter, M.; Wenzel, F.; Wolfel, R.; et al. SARS-CoV-2 antibody testing-questions to be asked. J. Allergy Clin. Immunol. 2020, 146, 35–43. [Google Scholar] [CrossRef]
- Cantoni, D.; Mayora-Neto, M.; Temperton, N. The role of pseudotype neutralization assays in understanding SARS CoV-2. Oxf. Open Immunol. 2021, 2, iqab005. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Wu, X.; Liu, Q.; Li, Q.; Liu, S.; Lu, J.; Yang, Y.; Cao, Y.; Huang, W.; Liang, C.; et al. A Human DPP4-Knockin Mouse’s Susceptibility to Infection by Authentic and Pseudotyped MERS-CoV. Viruses 2018, 10, 448. [Google Scholar] [CrossRef] [Green Version]
- Nie, J.; Wu, X.; Ma, J.; Cao, S.; Huang, W.; Liu, Q.; Li, X.; Li, Y.; Wang, Y. Development of in vitro and in vivo rabies virus neutralization assays based on a high-titer pseudovirus system. Sci. Rep. 2017, 7, 42769. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Q.; Fan, C.; Li, Q.; Zhou, S.; Huang, W.; Wang, L.; Sun, C.; Wang, M.; Wu, X.; Ma, J.; et al. Antibody-dependent-cellular-cytotoxicity-inducing antibodies significantly affect the post-exposure treatment of Ebola virus infection. Sci. Rep. 2017, 7, 45552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.; Liu, Q.; Huang, W.; Wu, J.; Nie, J.; Wang, M.; Zhao, C.; Zhang, L.; Wang, Y. An LASV GPC pseudotyped virus based reporter system enables evaluation of vaccines in mice under non-BSL-4 conditions. Vaccine 2017, 35, 5172–5178. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Li, Q.; Liu, Q.; Huang, W.; Nie, J.; Wang, Y. A bioluminescent imaging mouse model for Marburg virus based on a pseudovirus system. Hum. Vaccin Immunother. 2017, 13, 1811–1817. [Google Scholar] [CrossRef] [Green Version]
- Crawford, K.H.D.; Eguia, R.; Dingens, A.S.; Loes, A.N.; Malone, K.D.; Wolf, C.R.; Chu, H.Y.; Tortorici, M.A.; Veesler, D.; Murphy, M.; et al. Protocol and Reagents for Pseudotyping Lentiviral Particles with SARS-CoV-2 Spike Protein for Neutralization Assays. Viruses 2020, 12, 513. [Google Scholar] [CrossRef] [PubMed]
- Hyseni, I.; Molesti, E.; Benincasa, L.; Piu, P.; Casa, E.; Temperton, N.J.; Manenti, A.; Montomoli, E. Characterisation of SARS-CoV-2 Lentiviral Pseudotypes and Correlation between Pseudotype-Based Neutralisation Assays and Live Virus-Based Micro Neutralisation Assays. Viruses 2020, 12, 1011. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.C.; Lyddon, T.D.; Suarez, R.; Salcedo, B.; LePique, M.; Graham, M.; Ricana, C.; Robinson, C.; Ritter, D.G. Optimized Pseudotyping Conditions for the SARS-COV-2 Spike Glycoprotein. J. Virol. 2020, 94, e01062-20. [Google Scholar] [CrossRef]
- Lei, C.; Qian, K.; Li, T.; Zhang, S.; Fu, W.; Ding, M.; Hu, S. Neutralization of SARS-CoV-2 spike pseudotyped virus by recombinant ACE2-Ig. Nat. Commun. 2020, 11, 2070. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, F.; Weisblum, Y.; Muecksch, F.; Hoffmann, H.H.; Michailidis, E.; Lorenzi, J.C.C.; Mendoza, P.; Rutkowska, M.; Bednarski, E.; Gaebler, C.; et al. Measuring SARS-CoV-2 neutralizing antibody activity using pseudotyped and chimeric viruses. J. Exp. Med. 2020, 217, e20201181. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Larragoite, E.T.; Williams, E.; Lama, J.; Cisneros, I.; Delgado, J.C.; Slev, P.; Rychert, J.; Innis, E.A.; Coiras, M.; et al. Neutralization assay with SARS-CoV-1 and SARS-CoV-2 spike pseudotyped murine leukemia virions. Virol. J. 2021, 18, 1. [Google Scholar] [CrossRef] [PubMed]
- Condor Capcha, J.M.; Lambert, G.; Dykxhoorn, D.M.; Salerno, A.G.; Hare, J.M.; Whitt, M.A.; Pahwa, S.; Jayaweera, D.T.; Shehadeh, L.A. Generation of SARS-CoV-2 Spike Pseudotyped Virus for Viral Entry and Neutralization Assays: A 1-Week Protocol. Front. Cardiovasc. Med. 2020, 7, 618651. [Google Scholar] [CrossRef] [PubMed]
- Planas, D.; Bruel, T.; Grzelak, L.; Guivel-Benhassine, F.; Staropoli, I.; Porrot, F.; Planchais, C.; Buchrieser, J.; Rajah, M.M.; Bishop, E.; et al. Sensitivity of infectious SARS-CoV-2 B.1.1.7 and B.1.351 variants to neutralizing antibodies. Nat. Med. 2021, 27, 917–924. [Google Scholar] [CrossRef]
- CDC. SARS-CoV-2 Variant Classifications and Definitions. Available online: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-info.html#Concern (accessed on 12 July 2021).
- Mveang Nzoghe, A.; Essone, P.N.; Leboueny, M.; Maloupazoa Siawaya, A.C.; Bongho, E.C.; Mvoundza Ndjindji, O.; Avome Houechenou, R.M.; Agnandji, S.T.; Djoba Siawaya, J.F. Evidence and implications of pre-existing humoral cross-reactive immunity to SARS-CoV-2. Immun. Inflamm. Dis. 2021, 9, 128–133. [Google Scholar] [CrossRef] [PubMed]
- Ng, K.W.; Faulkner, N.; Cornish, G.H.; Rosa, A.; Harvey, R.; Hussain, S.; Ulferts, R.; Earl, C.; Wrobel, A.G.; Benton, D.J.; et al. Preexisting and de novo humoral immunity to SARS-CoV-2 in humans. Science 2020, 370, 1339–1343. [Google Scholar] [CrossRef] [PubMed]
- Bewley, K.R.; Coombes, N.S.; Gagnon, L.; McInroy, L.; Baker, N.; Shaik, I.; St-Jean, J.R.; St-Amant, N.; Buttigieg, K.R.; Humphries, H.E.; et al. Quantification of SARS-CoV-2 neutralizing antibody by wild-type plaque reduction neutralization, microneutralization and pseudotyped virus neutralization assays. Nat. Protoc. 2021, 16, 3114–3140. [Google Scholar] [CrossRef] [PubMed]
- Grzelak, L.; Temmam, S.; Planchais, C.; Demeret, C.; Tondeur, L.; Huon, C.; Guivel-Benhassine, F.; Staropoli, I.; Chazal, M.; Dufloo, J.; et al. A comparison of four serological assays for detecting anti-SARS-CoV-2 antibodies in human serum samples from different populations. Sci. Transl. Med. 2020, 12, eabc3103. [Google Scholar] [CrossRef]
- Barouch, D.H.; Stephenson, K.E.; Sadoff, J.; Yu, J.; Chang, A.; Gebre, M.; McMahan, K.; Liu, J.; Chandrashekar, A.; Patel, S.; et al. Durable Humoral and Cellular Immune Responses 8 Months after Ad26.COV2.S Vaccination. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef] [PubMed]
- Noori, M.; Nejadghaderi, S.A.; Arshi, S.; Carson-Chahhoud, K.; Ansarin, K.; Kolahi, A.A.; Safiri, S. Potency of BNT162b2 and mRNA-1273 vaccine-induced neutralizing antibodies against severe acute respiratory syndrome-CoV-2 variants of concern: A systematic review of in vitro studies. Rev. Med. Virol. 2021, e2277. [Google Scholar] [CrossRef]
- Madhi, S.A.; Baillie, V.; Cutland, C.L.; Voysey, M.; Koen, A.L.; Fairlie, L.; Padayachee, S.D.; Dheda, K.; Barnabas, S.L.; Bhorat, Q.E.; et al. Efficacy of the ChAdOx1 nCoV-19 Covid-19 Vaccine against the B.1.351 Variant. N. Engl. J. Med. 2021, 384, 1885–1898. [Google Scholar] [CrossRef] [PubMed]
- Muik, A.; Wallisch, A.K.; Sanger, B.; Swanson, K.A.; Muhl, J.; Chen, W.; Cai, H.; Maurus, D.; Sarkar, R.; Tureci, O.; et al. Neutralization of SARS-CoV-2 lineage B.1.1.7 pseudovirus by BNT162b2 vaccine-elicited human sera. Science 2021, 371, 1152–1153. [Google Scholar] [CrossRef] [PubMed]
- Lowe, D. How Well Does the J&J Vaccine Work against the Delta Variant? Available online: https://blogs.sciencemag.org/pipeline/archives/2021/07/21/how-well-does-the-jj-vaccine-work-against-the-delta-variant (accessed on 28 July 2021).
- Westendorf, K.; Zentelis, S.; Foster, D.; Vaillancourt, P.; Wiggin, M.; Lovett, E.; Hendle, J.; Pustilnik, A.; Sauder, J.M.; Kraft, L.; et al. LY-CoV1404 potently neutralizes SARS-CoV-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- National Research Council of the National Academy of Sciences. Guide for the Care and Use of Laboratory Animals, 8th ed.; The National Academies Press: Washington, DC, USA, 2011; p. 246. [Google Scholar]
| Control | 614D Variant | B.1.351 SA Variant | B.1.1.7 UK Variant |
|---|---|---|---|
| NIBSC 20/130 | 4270 3541 1664 | 160 <800 <160 | 667.1 800 922 |
| Acro-SAD-S35 | 11,511 10,502 11,164 9881 | <160 <800 <160 | 11,192 8535 17,450 |
| Sino-40591-MM43 | 13,973 15,706 14,182 | 13,828 8182 12,243 | 9755 16,994 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kalkeri, R.; Cai, Z.; Lin, S.; Farmer, J.; Kuzmichev, Y.V.; Koide, F. SARS-CoV-2 Spike Pseudoviruses: A Useful Tool to Study Virus Entry and Address Emerging Neutralization Escape Phenotypes. Microorganisms 2021, 9, 1744. https://doi.org/10.3390/microorganisms9081744
Kalkeri R, Cai Z, Lin S, Farmer J, Kuzmichev YV, Koide F. SARS-CoV-2 Spike Pseudoviruses: A Useful Tool to Study Virus Entry and Address Emerging Neutralization Escape Phenotypes. Microorganisms. 2021; 9(8):1744. https://doi.org/10.3390/microorganisms9081744
Chicago/Turabian StyleKalkeri, Raj, Zhaohui Cai, Shuling Lin, John Farmer, Yury V. Kuzmichev, and Fusataka Koide. 2021. "SARS-CoV-2 Spike Pseudoviruses: A Useful Tool to Study Virus Entry and Address Emerging Neutralization Escape Phenotypes" Microorganisms 9, no. 8: 1744. https://doi.org/10.3390/microorganisms9081744
APA StyleKalkeri, R., Cai, Z., Lin, S., Farmer, J., Kuzmichev, Y. V., & Koide, F. (2021). SARS-CoV-2 Spike Pseudoviruses: A Useful Tool to Study Virus Entry and Address Emerging Neutralization Escape Phenotypes. Microorganisms, 9(8), 1744. https://doi.org/10.3390/microorganisms9081744






