3.1. ADG Performance, Genetic Parameters and Variance Components
The average ADG (g) was higher in the DUC (986.04 ± 125.25) than it was in the crossbred (849.43 ± 110.87) pigs (
Table S1). Duroc pigs are known to have undergone intensive artificial selection over 100 years, and have superior carcass, growth, and feed conversion efficiency traits compared with those of other breeds [
29]. KNPs, one of the parental breeds of the crossbred pigs, have undergone severe inbreeding events because of their low population size [
21], and their growth performance is also lower than that of commercial breeds. Therefore, the average ADG in the crossbred pigs was lower than it was in the DUC pigs. To better understand the genetic information of the ADG in the purebred and crossbred pigs, we estimated the direct and social genetic variance (
and
), phenotypic variance (
), total heritable variance (
), direct heritability (
), total heritability (
), and genetic correlation between the DGE and SGE (
). The variance components of these genetic parameters in DUC and crossbred pigs are given in
Table 1.
The
values for ADG were 0.16 ± 0.04 and 0.36 ± 0.05 for DUC and crossbred pigs, respectively, and the
values, which include both DGE and SGE, were slightly higher in both populations, with 0.19 ± 0.04 and 0.39 ± 0.08 for DUC and crossbred, respectively (
Table 1). However, the crossbred pigs showed little difference in heritability values (
and
) due to the negative correlation (−0.15 ± 0.27) between the direct and social genetic effects.
This finding confirmed the contribution of SGE to total heritable variance. We found that the DUC had lower heritability for ADG than the crossbred pigs, and they also had lower heritability for ADG than other Duroc populations reported previously [
30,
31,
32,
33]. These results suggest that intensive selection events for growth traits have occurred in DUC populations. Indeed, DUC pigs have been intensively selected for growth traits after they were introduced into South Korea. A recent study reported substantial genetic improvement of ADG in DUC pigs, as the estimated breeding value for ADG has increased from −5.23 g to 45.16 g since 2000 [
30]. Therefore, there may now be less chance for genetic improvement for ADG in the DUC population. The genetic correlation between SGE and DGE was neutral for DUC (0.03 ± 0.20) and weak for crossbred (−0.15 ± 0.27) (
Table 1). This result is consistent with the work of Bergsma et al. [
34], who found that the absence of conflict between an individual’s growth and a mate’s growth may be a consequence of neutral or weak social interactions.
3.2. Genetic Correlations between Purebred and Crossbred Pigs
To observe the relationship between genetic effects, we estimated the genetic correlations between the DGE and SGE for DUC and crossbred pigs (
Table 2). The genetic correlation between DGE of DUC and that of crossbred pigs was favorable (0.48 ± 0.19) and statistically significant (
Table 2).
This result is similar to that of a previous report of statistically significant and moderate genetic correlations between the DGE on ADG of a crossbred dam (F1, landrace × Yorkshire) and that of landrace (0.46 ± 0.18) or Yorkshire (0.41 ± 0.17) [
11]. The genetic correlation between the SGE of DUC and that of crossbred pigs was negative (−0.27 ± 0.25) and not statistically significant. The genetic correlation between the DGE for DUC and SGE for crossbreds was negative (−0.53 ± 0.23) and statistically significant. The estimated genetic correlation between SGE for DUC and DGE for crossbreds was the highest among the pairwise correlations (0.77 ± 0.12) and statistically significant, indicating that the SGE for DUC was highly associated with crossbred performance. Previous studies have also indicated that genomic selection of purebred animals can increase the selection response for crossbred performance [
19,
35]. Ask et al. [
11] reported a positive effect of SGE that was assessed using purebred data on the ADG traits in F1 crossbred pigs. As we found moderate (between DGE of DUC and DGE of crossbred) to high genetic correlations (between SGE of DUC and DGE of crossbred) between the DUC and crossbred pigs, we suggest that genomic selection of the purebred population, especially for social behavior, may have affected the selection response for ADG in the crossbred population.
3.3. QTLs for DGE and SGE
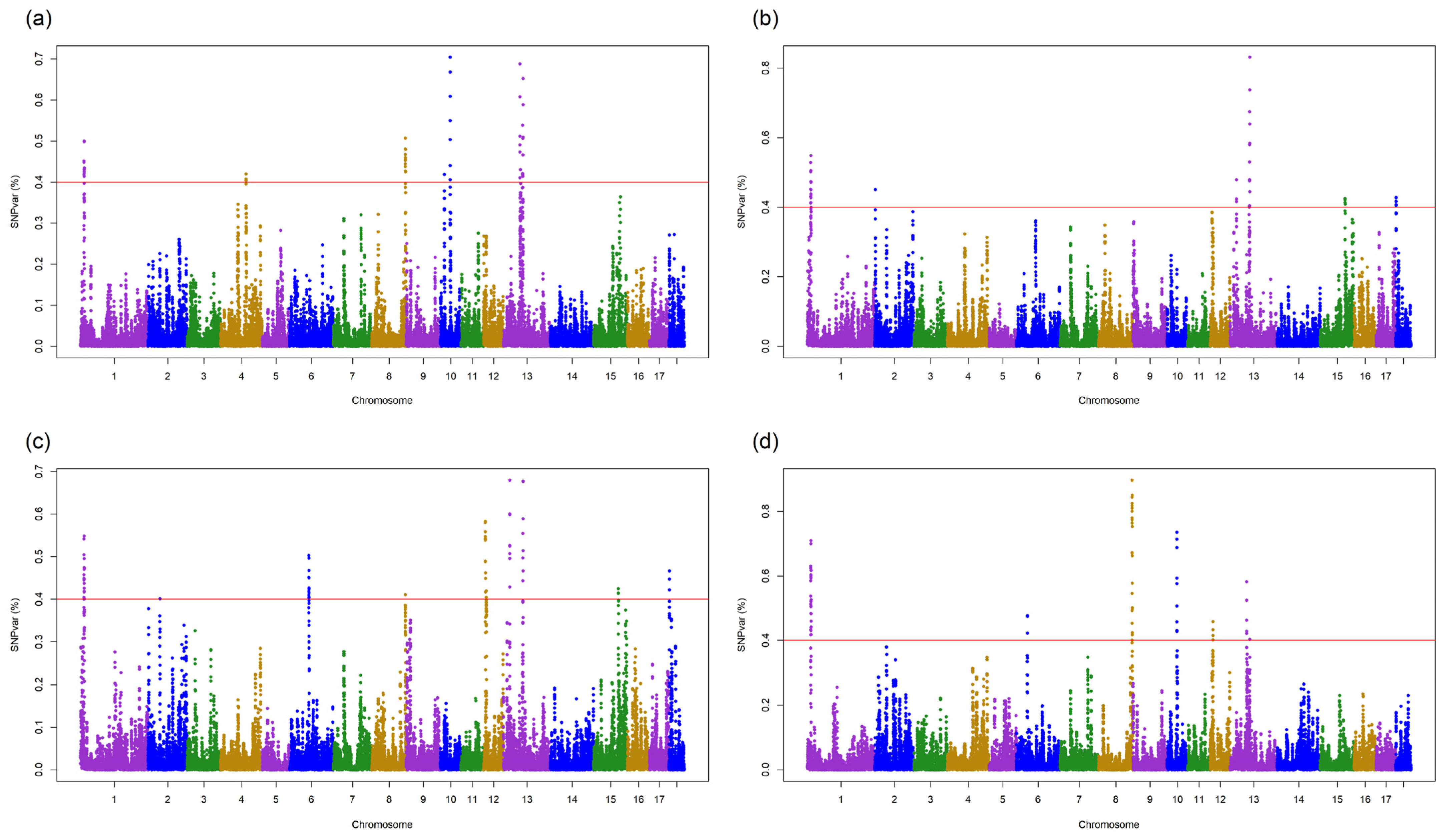
As shown in
Figure 1a and
Table S2, the GWAS identified 52 significant SNPs (explained genetic variance > 0.4%) associated with DGE in DUC pigs.
These SNPs explained 24.9% of the total genetic variance and were located on SSC1, SSC4, SSC8, SSC10, and SSC13. The gene annotations for the 1-Mb QTLs that centered those SNPs identified 97 genes associated with these QTLs (
Table S2). Because a large number of QTLs were detected in this study, we focused on the QTLs that explained the greatest genetic variance for the DGE and SGE in DUC and crossbred pigs and compared them with previously reported pig QTLs. A QTL in SSC10 (31.4–32.4 Mb) that explained the most genetic variance (0.7%) overlapped with the production QTLs that were previously reported to be associated with ADG [
36,
37], as well as with the QTLs associated with feed conversion ratio [
38] and meat-related traits [
39,
40,
41]. In the crossbred pigs, 43 significant SNPs (explained 20.5% of genetic variance) associated with DGE were located on SSC1, SSC2, SSC13, SSC15, and SSC18 (
Figure 1b), and 83 genes were annotated within the corresponding QTLs (
Table S2). We also found that the QTL (65.3–66.3 Mb) in SSC13 that explained the most genetic variance (0.8%) overlapped with the production-related QTLs that are associated with ADG [
37], as well as meat-related QTLs that are associated with backfat weight, the percentage of loin fat, and loin muscle area [
42].
For SGE in DUC, 70 significant SNPs located on SSC1, SSC2, SSC6, SSC8, SSC12, SSC13, SSC15, and SSC18 explained 33.6% of the total genetic variance (
Figure 1c,
Table S2). A QTL located on SSC13 (16.4–17.4 Mb) that explained the highest genetic variance (0.7%) overlapped with a QTL associated with the time spent socializing [
43], as well as with production-related QTLs that are associated with ADG and chest width [
36,
37]. We also detected QTLs associated with meat-related traits, such as loin muscle area, average backfat thickness, and backfat at tenth rib [
36,
44]. For SGE in crossbred pigs, 76 significant SNPs located on SSC1, SSC6, SSC8, SSC10, SSC12, and SSC13 explained 42.8% of the total genetic variance (
Figure 1d;
Table S2). The top QTLs (SSC8, 136.3–137.3 Mb), which explained 0.9% of the genetic variance, overlapped with QTLs related to coping behavior [
45] and exploration during stress [
46], as well as with QTLs associated with the percentage of lean meat, rump width, and length of humerus [
47,
48].
3.4. GO and KEGG Analyses for SGE
SGE is an established concept in behavioral ecology [
49]. SGE is not associated with one specific social interaction, but instead captures the overall effect of several social interactions between individuals on a specific trait of the recipient individual [
50]. Hong et al. [
5] identified positional candidate genes for SGE on ADG that have biological roles that are strongly associated with neuropsychiatric processes. We also identified candidate genes that may be related to the neurological disorders and behavioral changes.
We identified genes that were annotated with 17 GO terms that might be associated with the SGE in DUC (
Table 3), including
DBX1,
PAX7, and
SHH, which were annotated with the neuron fate commitment term (GO: 0048663).
DBX1 is expressed in hypothalamic progenitors and restriction of
DBX1 was found to be critical in establishing the neuronal fate of V0 and V1, which are derived from adjacent progenitor domains [
51]. The previous study revealed that
DBX1 is associated with diminished responses to feeding stressors and abnormal GABAergic neuron morphology [
51,
52]. Proskorovski-Ohayon et al. [
53] suggested that homozygous mutation in
PAX7 likely causes a neuromuscular syndrome in humans. Dysregulation of the SHH pathway in the brain was reported to be associated with neurodegenerative diseases, such as amyotrophic lateral sclerosis and Parkinson’s disease [
54].
We also identified three genes (
CCL19,
CCL21, and
SOX9) that might be associated with SGE in crossbreds. These genes were annotated with the response to interleukin-1 term (GO: 0070555). Interleukin-1 (IL-1) is a master regulator of inflammation by controlling innate immune processes [
55]. The IL-1 superfamily includes seven pro-inflammatory proteins (IL-1α, IL-1β, IL-18, IL-33, IL-36α, IL-36β, and IL-36γ). Modulation of forebrain serotonin activity by IL-1β signaling in the dorsal raphe nucleus (DRN) was reported to control aggressive behavior, and non-aggressive mice were found to have higher levels of IL-1β in DRN than aggressive mice [
56]. Therefore, we suggest that aggression behavior in pigs may be associated with SGE.
We also identified four genes (
CCL19,
CYLD,
MAS1, and
NOD2) that were annotated with GO terms associated with the regulation of NIK/NF-kappaB signaling (GO:0038061, GO:1901222, and GO:1901224) (
Table 4). Diverse external stimuli, such as the release of cytokines (TNF-alpha and IL-1), viral infections, and neurotrophic factors, lead to the activation of NF-kappaB, and genes that are regulated by NF-kappaB have key roles in stress and immune responses [
57]. Activation of NF-kappaB has been reported to be associated with human nervous system diseases, such as Huntington’s disease, Alzheimer’s disease, and Parkinson’s disease [
58,
59,
60]. A prominent behavioral symptom of these neurological disorders is apathy, which is defined as the deficit of goal-directed behavior or motivational impairment [
61,
62]. Therefore, we suggest that these genes might be also related to SGE in pigs.
Three genes (
ADCY7,
IL17RC, and
NOD2) were annotated with terms related to inflammatory response (GO:0002532, GO:0002534, and GO:1900015) (
Table 4). A previous study showed that
ADCY7 was associated with depression, using both genetically modified mice and an association study of a human population [
63].
IL17RC is essential for IL17A signaling [
64], and IL17A is generally considered to cause neurodegenerative diseases by activating glial cells [
65].
NOD2 is a positive regulator of IL-1β secretion and NF-kappaB activation [
66]. As noted above, previous studies have reported associations between IL-1β and aggression, and between activation of NF-kappaB and nervous system diseases; therefore, we suggest that these three genes may be related to SGE.
3.5. GO and KEGG Analyses for DGE
We identified the genes that were annotated with 14 and 13 significant GO terms that might be associated with the DGE in DUC and crossbred pigs, respectively (
Table 5 and
Table 6).
Four of the genes (
CAMK1,
CAV3,
FOXP1, and
SHH) were involved in the biological process of positive regulation of muscle cell differentiation (GO: 0051149) (
Table 5 and
Table 6). Growth and development of muscle are essential for the breeding of livestock species raised for meat production. Muscle formation, also called myogenesis, is a complex biological process that involves cell proliferation, differentiation, migration, myotube formation, and maturation of myofibers [
67,
68]. During postnatal growth, the increase in skeletal muscle mass is mainly due to an increase in muscle fiber size [
69].
CAMK1 and
FOXP1 were reported to be associated with increased lean body mass in mice [
70].
CAV3 null mice showed mild myopathic changes with the presence of necrotic fiber and variability in muscle fiber size [
71]. Copy number variations in
SHH were shown to have significant associations with body size traits in Chinese beef cattle breeds [
72]. Because the growth performance of pigs is closely related to the proliferation and differentiation of muscle cells, we suggest that the positive regulation of muscle cell differentiation may be associated with the DGE for ADG in DUC and crossbred pigs. The positive regulation of muscle cell differentiation term was also identified for the SGE in DUC (
Table 3). We suggest that this common GO term might be caused due to high genetic correlation (0.77 ± 0.12) between the SGE in DUC and DGE in crossbred pigs (
Table 2) or the pleiotropy phenomenon, indicating that a single gene affects two or more phenotypic traits [
73].
We also identified three genes (
FGF3,
FGF4, and
FGF19) that were annotated with terms related to the response to the fibroblast growth factor (GO: 0071774 and GO: 0044344) and receptor signaling pathway of the fibroblast growth factor (GO: 0008543) (
Table 6). Fibroblast growth factors (FGF) belong to a large protein group that is related to proliferation, migration, differentiation, and apoptosis [
74]. In zebrafish, early specification of the skull was found to be regulated by
FGF3, together with
SHH [
75].
FGF4 has been reported to be associated with axial elongation and development of mouse embryos [
76] and with Wnt signaling in mice [
77]. Benoit et al. [
78] reported several functions of
FGF19, including the regulation of skeletal muscle mass through the expansion of muscle fiber, and protection of muscle from atrophy. In mice, treatment with
FGF19 caused skeletal muscle hypertrophy, and
FGF19 increased the size of human myotubes in vitro [
78].