Impact of Autolysed Brewer’s Yeast and Soluble Dried Yeast Extract on Growth Performance and Mucosal Health of Atlantic Salmon (Salmo salar) Parr
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Design and Fish
2.2. Experimental Diets and Feeding
2.3. Fish Sampling
2.4. Growth Performance
2.5. Histology
2.5.1. Light Microscopy
2.5.2. Electron Microscopy
2.6. Haematology
2.6.1. Haemoglobin Concentration
2.6.2. Blood Cell Count
2.7. Gene Expression
2.7.1. RNA Extraction and cDNA Synthesis
2.7.2. Quantitative Real-Time PCR (qPCR)
2.8. Intestinal Microbiome Analysis
2.8.1. DNA Extraction
2.8.2. PCR Amplification and High-Throughput Sequencing
2.9. Statistical Analysis
3. Results
3.1. Histology
3.1.1. Light Microscopy
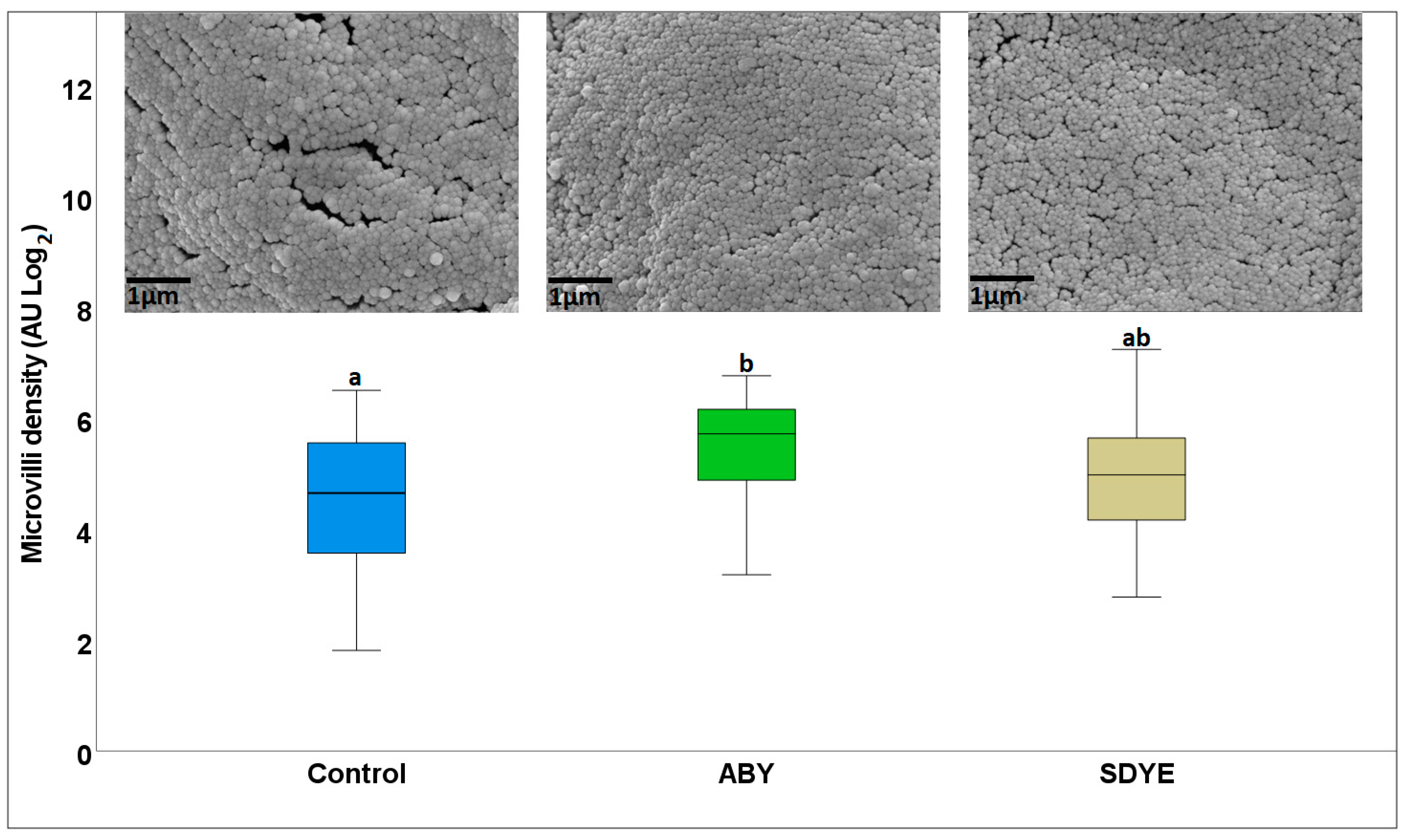
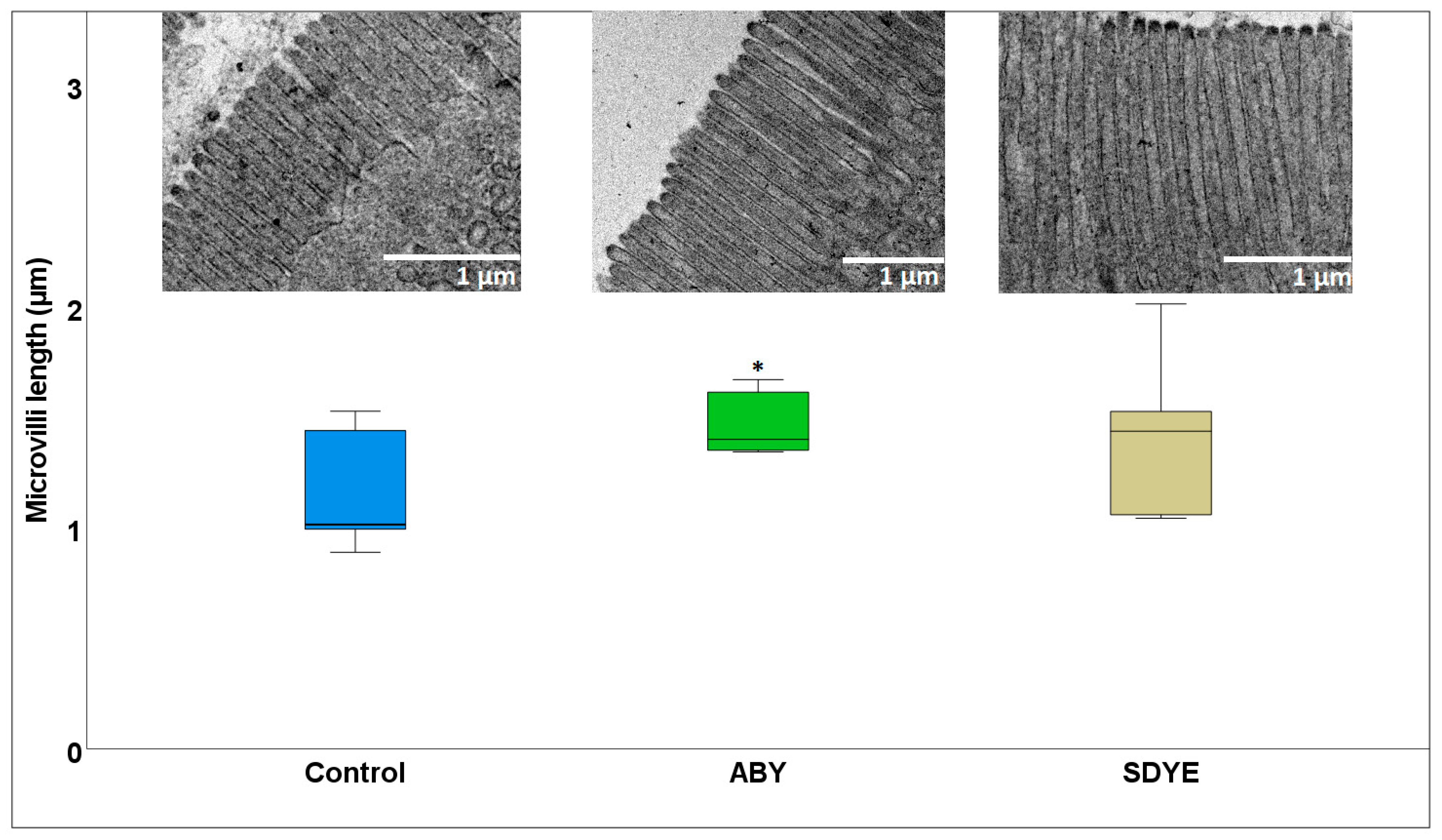
3.1.2. Electron Microscopy
3.2. Haematology
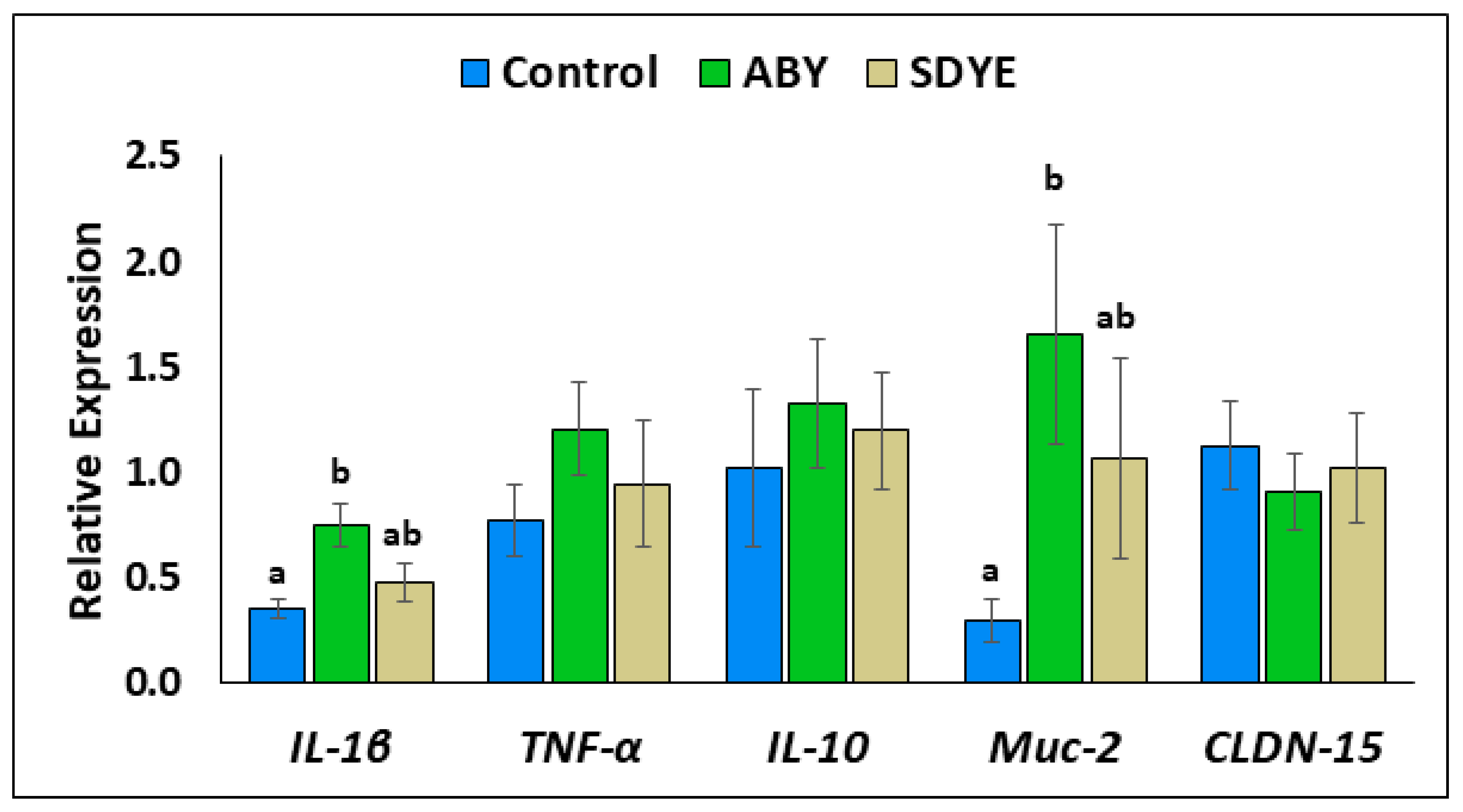
3.3. Gene Expression
3.4. Intestinal Microbiome
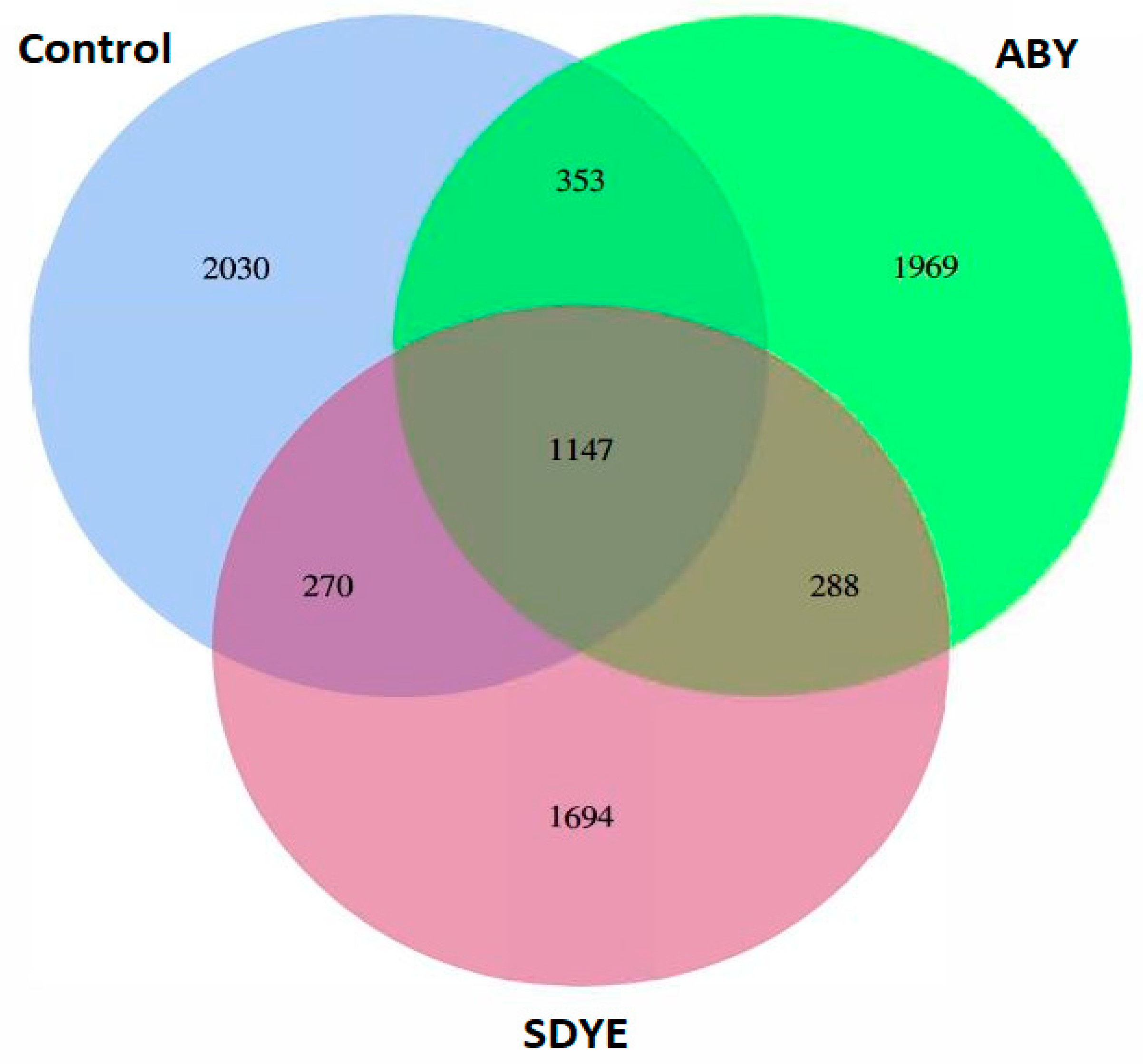
3.4.1. Sequencing and ASVs Identification
3.4.2. Diversity and Community Structure
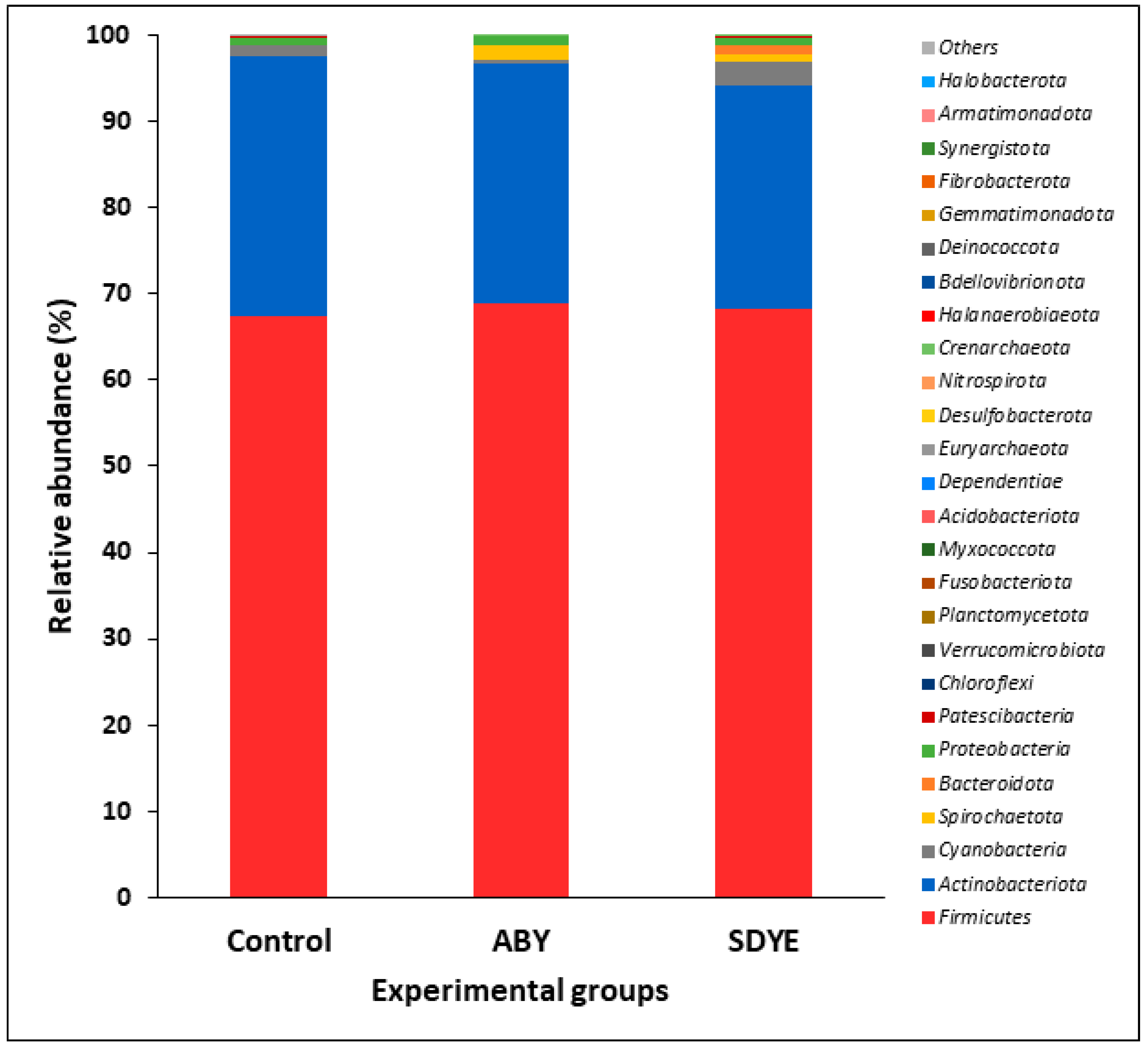
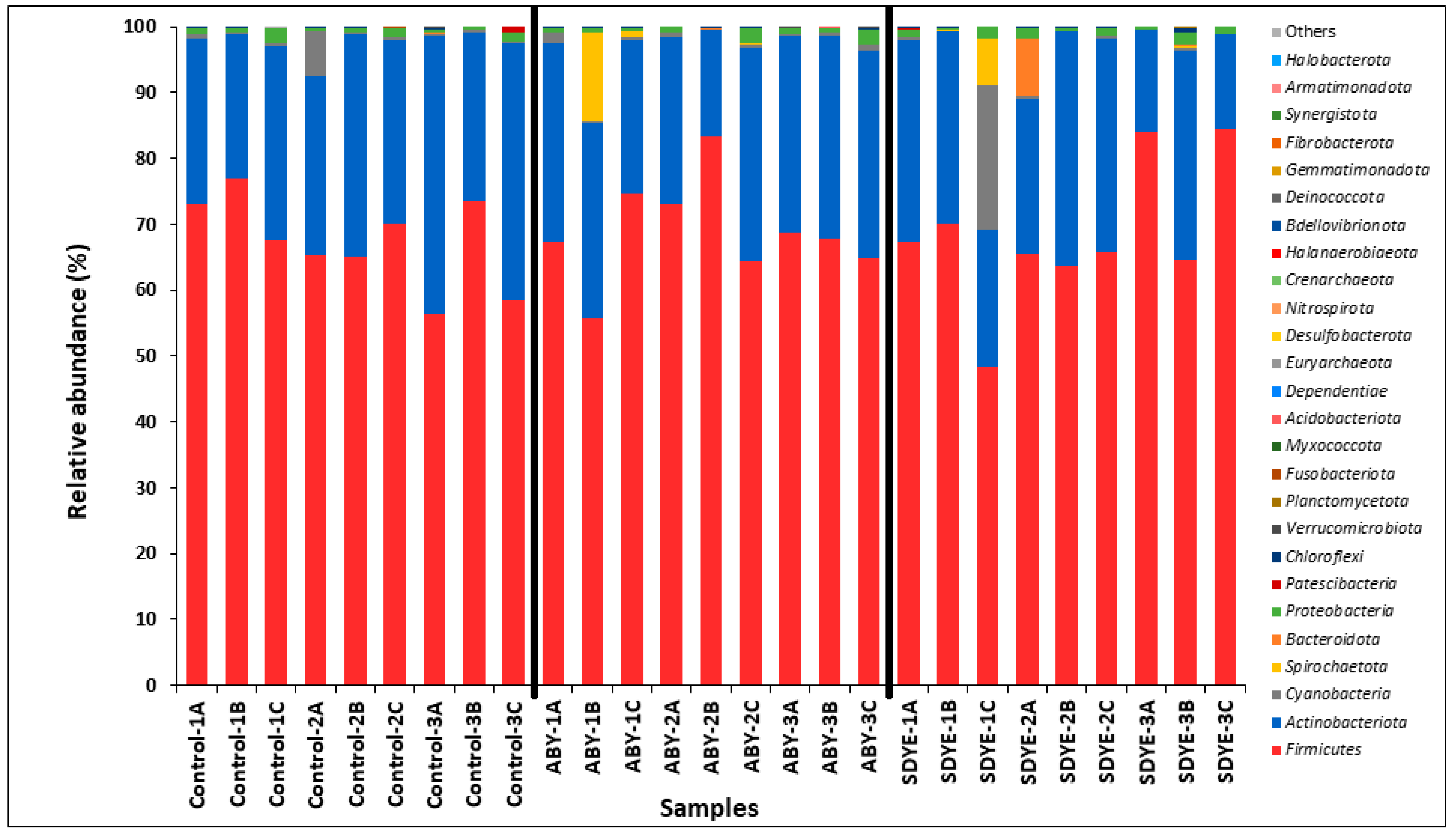
3.4.3. Bacterial Composition
3.5. Growth Performance
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- FAO. The State of World Fisheries and Aquaculture 2024—Blue Transformation in Action; FAO: Rome, Italy, 2024. [Google Scholar]
- Sprague, M.; Dick, J.R.; Tocher, D.R. Impact of sustainable feeds on omega-3 long-chain fatty acid levels in farmed Atlantic salmon, 2006–2015. Sci. Rep. 2016, 6, 21892. [Google Scholar] [CrossRef] [PubMed]
- Encarnação, P. 5—Functional feed additives in aquaculture feeds. In Aquafeed Formulation; Nates, S.F., Ed.; Academic Press: San Diego, CA, USA, 2016; pp. 217–237. [Google Scholar]
- Merrifield, D.L.; Ringø, E. Aquaculture Nutrition: Gut Health, Probiotics and Prebiotics; Wiley: Hoboken, NJ, USA, 2014. [Google Scholar]
- Machuca, C.; Méndez-Martínez, Y.; Reyes-Becerril, M.; Angulo, C. Yeast β-Glucans as Fish Immunomodulators: A Review. Animals 2022, 12, 2154. [Google Scholar] [CrossRef] [PubMed]
- Rawling, M.; Schiavone, M.; Apper, E.; Merrifield, D.L.; Castex, M.; Leclercq, E.; Foey, A. Yeast cell wall extracts from Saccharomyces cerevisiae varying in structure and composition differentially shape the innate immunity and mucosal tissue responses of the intestine of zebrafish (Danio rerio). Front. Immunol. 2023, 14, 1158390. [Google Scholar] [CrossRef] [PubMed]
- Odu-Onikosi, S.G.; Momoh, T.A.; Eynon, B.; Pontefract, N.; Kuri, V.; Kühlwein, H.; Merrifield, D.L. Autolyzed brewer’s yeast enhances growth and intestinal health in early life stages of Nile tilapia (Oreochromis niloticus L.). J. World Aquac. Soc. 2025, 56, e13120. [Google Scholar] [CrossRef]
- Gomez, D.; Oriol Sunyer, J.O.; Salinas, I. The mucosal immune system of fish: The evolution of tolerating commensals while fighting pathogens. Fish Shellfish Immunol. 2013, 35, 1729–1739. [Google Scholar] [CrossRef]
- Lazado, C.C.; Caipang, C.M.A. Mucosal immunity and probiotics in fish. Fish Shellfish Immunol. 2014, 39, 78–89. [Google Scholar] [CrossRef]
- Porter, D.; Peggs, D.; McGurk, C.; Martin, S.A.M. In-vivo analysis of Protec™ and β-glucan supplementation on innate immune performance and intestinal health of rainbow trout. Fish Shellfish Immunol. 2023, 134, 108573. [Google Scholar] [CrossRef]
- Firmino, J.P.; Galindo-Villegas, J.; Reyes-López, F.E.; Gisbert, E. Phytogenic Bioactive Compounds Shape Fish Mucosal Immunity. Front. Immunol. 2021, 12, 695973. [Google Scholar] [CrossRef]
- Refstie, S.; Baeverfjord, G.; Seim, R.R.; Elvebo, O. Effects of dietary yeast cell wall β-glucans and MOS on performance, gut health, and salmon lice resistance in Atlantic salmon (Salmo salar) fed sunflower and soybean meal. Aquaculture 2010, 305, 109–116. [Google Scholar] [CrossRef]
- Dimitroglou, A.; Reynolds, P.; Ravnoy, B.; Johnsen, F.; Sweetman, J.W.; Johansen, J.; Davies, S.J. The Effect of Mannan Oligosaccharide Supplementation on Atlantic Salmon Smolts (Salmo salar L.) Fed Diets with High Levels of Plant Proteins. J. Aquac. Res. Dev. 2011, 1 (Suppl. S11), 6. [Google Scholar]
- Leclercq, E.; Pontefract, N.; Rawling, M.; Valdenegro, V.; Aasum, E.; Andujar, L.V.; Migaud, H.; Castex, M.; Merrifield, D. Dietary supplementation with a specific mannan-rich yeast parietal fraction enhances the gut and skin mucosal barriers of Atlantic salmon (Salmo salar) and reduces its susceptibility to sea lice (Lepeophtheirus salmonis). Aquaculture 2020, 529, 735701. [Google Scholar] [CrossRef]
- Morales-Lange, B.; Hansen, J.O.; Djordjevic, B.; Mydland, L.T.; Castex, M.; Mercado, L.; Imarai, M.; Sorum, H.; Overland, M. Immunomodulatory effects of hydrolyzed Debaryomyces hansenii in Atlantic salmon (Salmo salar L): From the in vitro model to a natural pathogen challenge after seawater transfer. Aquaculture 2024, 578, 740035. [Google Scholar] [CrossRef]
- Baumgärtner, S.; James, J.; Ellison, A. The supplementation of a prebiotic improves the microbial community in the gut and the skin of Atlantic salmon (Salmo salar). Aquac. Rep. 2022, 25, 101204. [Google Scholar] [CrossRef]
- NRC. Nutrient Requirements of Fish and Shrimp; The National Academies Press: Washington, WA, USA, 2011; p. 392. [Google Scholar]
- AOAC. Official Methods of Analysis of AOAC International; Latimer, G.W., Jr., Ed.; Oxford University Press: Oxford, UK, 2023. [Google Scholar]
- Satpathy, B.B.; Ray, A.K. Effect of dietary protein and carbohydrate levels on growth, nutrient utilization and body composition in fingerling rohu, Labeo rohita (Hamilton). J. Appl. Ichthyol. 2009, 25, 728–733. [Google Scholar] [CrossRef]
- Merrifield, D.L.; Dimitroglou, A.; Bradley, G.; Baker, R.T.M.; Davies, S.J. Soybean meal alters autochthonous microbial populations, microvilli morphology and compromises intestinal enterocyte integrity of rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2009, 32, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Rawling, M.D.; Pontefract, N.; Rodiles, A.; Anagnostara, I.; Leclercq, E.; Schiavone, M.; Castex, M.; Merrifield, D.L. The effect of feeding a novel multistrain yeast fraction on European seabass (Dicentrachus labrax) intestinal health and growth performance. J. World Aquac. Soc. 2019, 50, 1108–1122. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, 10034. [Google Scholar] [CrossRef] [PubMed]
- Pontefract, N.M. Effects of Dietary Additives on Growth Performance, Immunity and Mucosal Barrier Defences of Salmonids. Ph.D. Thesis, University of Plymouth, Plymouth, UK, 2021. [Google Scholar]
- Rasmussen, R. Quantification on the LightCycler. In Rapid Cycle Real-Time PCR: Methods and Applications; Meuer, S., Wittwer, C., Nakagawara, K.-I., Eds.; Springer: Berlin/Heidelberg, Germany, 2001; pp. 21–34. [Google Scholar]
- Magoč, T.; Salzberg, S.L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011, 27, 2957–2963. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.F.; Zhou, Y.Q.; Chen, Y.R.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, 884–890. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Li, M.J.; Shao, D.T.; Zhou, J.C.; Gu, J.H.; Qin, J.J.; Chen, W.; Wei, W.Q. Signatures within esophageal microbiota with progression of esophageal squamous cell carcinoma. Chin. J. Cancer Res. 2020, 32, 755–767. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Holmes, S.P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 2017, 11, 2639–2643. [Google Scholar] [CrossRef]
- Röhmel, J. Precision intervals for estimates of the difference in success rates for binary random variables based on the permutation principle. Biom. J. 1996, 38, 977–993. [Google Scholar] [CrossRef]
- De Marco, G.; Cappello, T.; Maisano, M. Histomorphological Changes in Fish Gut in Response to Prebiotics and Probiotics Treatment to Improve Their Health Status: A Review. Animals 2023, 13, 2860. [Google Scholar] [CrossRef] [PubMed]
- Rawling, M.; Leclercq, E.; Foey, A.; Castex, M.; Merrifield, D. A novel dietary multi-strain yeast fraction modulates intestinal toll-like-receptor signalling and mucosal responses of rainbow trout (Oncorhynchus mykiss). PLoS ONE 2021, 16, e0245021. [Google Scholar] [CrossRef] [PubMed]
- Adeoye, A.A.; Obasa, S.O.; Fawole, F.J.; Wan, A.H.L.; Davies, S.J. Dietary supplementation of autolysed yeast enhances growth, liver functionality and intestinal morphology in African catfish. Aquac. Nutr. 2020, 26, 772–780. [Google Scholar] [CrossRef]
- Meena, D.K.; Das, P.; Kumar, S.; Mandal, S.C.; Prusty, A.K.; Singh, S.K.; Akhtar, M.S.; Behera, B.K.; Kumar, K.; Pal, A.K.; et al. Beta-glucan: An ideal immunostimulant in aquaculture (a review). Fish Physiol. Biochem. 2013, 39, 431–457. [Google Scholar] [CrossRef] [PubMed]
- Merrifield, D.L.; Carnevali, O. Probiotic Modulation of the Gut Microbiota of Fish. In Aquaculture Nutrition; Merrifield, D.L., Ringø, E., Eds.; Wiley: Hoboken, NJ, USA, 2014; pp. 185–222. [Google Scholar]
- Barducci, R.S.; de Abreu, V.; Santos, A.A.D.; Pacheco, L.G.; Koch, J.F.A.; Florencio, M.; Pilarski, F. Natural feed additive containing Saccharomyces cerevisiae-originated free nucleotides improves innate immunity, gut histology and disease resistance in Nile tilapia. Anim. Feed Sci. Technol. 2022, 289, 115337. [Google Scholar] [CrossRef]
- Fazio, F. Fish hematology analysis as an important tool of aquaculture: A review. Aquaculture 2019, 500, 237–242. [Google Scholar] [CrossRef]
- Kühlwein, H.; Merrifield, D.L.; Rawling, M.D.; Foey, A.D.; Davies, S.J. Effects of dietary β-(1,3)(1,6)-D-glucan supplementation on growth performance, intestinal morphology and haemato-immunological profile of mirror carp (Cyprinus carpio L.). J. Anim. Physiol. Anim. Nutr. 2014, 98, 279–289. [Google Scholar] [CrossRef]
- Neuls, L.; de Souza, V.J.; Romao, S.; Bitencourt, T.B.; Ramos, C.J.R.; Parra, J.E.G.; Cazarolli, L.H. Immunomodulatory effects of Yarrowia lipolytica as a food additive in the diet of Nile tilapia. Fish Shellfish Immunol. 2021, 119, 272–279. [Google Scholar] [CrossRef]
- Engelsma, M.Y.; Huising, M.O.; van Muiswinkel, W.B.; Flik, G.; Kwang, J.; Savelkoul, H.F.; Verburg-van Kemenade, B.M. Neuroendocrine-immune interactions in fish: A role for interleukin-1. Vet. Immunol. Immunopathol. 2002, 87, 467–479. [Google Scholar] [CrossRef] [PubMed]
- Stanforth, K.J.; Zakhour, M.I.; Chater, P.I.; Wilcox, M.D.; Adamson, B.; Robson, N.A.; Pearson, J.P. The MUC2 Gene Product: Polymerisation and Post-Secretory Organisation-Current Models. Polymers 2024, 16, 1663. [Google Scholar] [CrossRef]
- Andriamialinirina, H.J.T.; Irm, M.; Taj, S.; Lou, J.H.; Jin, M.; Zhou, Q.C. The effects of dietary yeast hydrolysate on growth, hematology, antioxidant enzyme activities and non-specific immunity of juvenile Nile tilapia, Oreochromis niloticus. Fish Shellfish Immunol. 2020, 101, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.L.; Liu, S.Y.; Zuo, F.Y.; Luo, L.; Chen, D.F.; Ou, Y.P.; Geng, Y.; Zhang, Y.F.; Lin, G.; Yang, S.Y.; et al. cMOS enhanced the mucosal immune function of skin and gill of goldfish (Carassius auratus Linnaeus) to improve the resistance to Ichthyophthirius multifiliis infection. Fish Shellfish Immunol. 2022, 126, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Yu, M. Role of Goblet Cells in Intestinal Barrier and Mucosal Immunity. J. Inflamm. Res. 2021, 14, 3171–3183. [Google Scholar] [CrossRef] [PubMed]
- Romero, J.; Ringø, E.; Merrifield, D.L. The Gut Microbiota of Fish. In Aquaculture Nutrition; Merrifield, D.L., Ringø, E., Eds.; Wiley: Hoboken, NJ, USA, 2014; pp. 75–100. [Google Scholar]
- Burr, G.; Hume, M.; Ricke, S.; Nisbet, D.; Gatlin, D., 3rd. In vitro and in vivo evaluation of the prebiotics GroBiotic-A, inulin, mannanoligosaccharide, and galactooligosaccharide on the digestive microbiota and performance of hybrid striped bass (Morone chrysops × Morone saxatilis). Microb. Ecol. 2010, 59, 187–198. [Google Scholar] [CrossRef] [PubMed]
- Dimitroglou, A.; Merrifield, D.L.; Spring, P.; Sweetman, J.; Moate, R.; Davies, S.J. Effects of mannan oligosaccharide (MOS) supplementation on growth performance, feed utilisation, intestinal histology and gut microbiota of gilthead sea bream (Sparus aurata). Aquaculture 2010, 300, 182–188. [Google Scholar] [CrossRef]
- Agboola, J.O.; Rocha, S.D.C.; Mensah, D.D.; Hansen, J.O.; Oyås, O.; Lapeña, D.; Mydland, L.T.; Arntzen, M.O.; Horn, S.J.; Overland, M. Effect of yeast species and processing on intestinal microbiota of Atlantic salmon (Salmo salar) fed soybean meal-based diets in seawater. Anim. Microbiome 2023, 5, 21. [Google Scholar] [CrossRef] [PubMed]
- Meziti, A.; Nikouli, E.; Papaharisis, L.; Kormas, K.; Mente, E. The response of gut and fecal bacterial communities of the European sea bass (Dicentrachus labrax) fed a low fish-plant meal and yeast protein supplementation diet. Sustain. Microbiol. 2024, 1, qvae005. [Google Scholar] [CrossRef]
- Spilsbury, F.; Foysal, M.J.; Tay, A.; Gagnon, M.M. Gut Microbiome as a Potential Biomarker in Fish: Dietary Exposure to Petroleum Hydrocarbons and Metals, Metabolic Functions and Cytokine Expression in Juvenile Lates calcarifer. Front. Microbiol. 2022, 13, 827371. [Google Scholar]
- Schleifer, K.-H.; Bell, J.A. Staphylococcus. In Bergey’s Manual of Systematics of Archaea and Bacteria; NIH: Bethesda, MD, USA, 2015; pp. 1–43. [Google Scholar]
- Egerton, S.; Culloty, S.; Whooley, J.; Stanton, C.; Ross, R.P. The Gut Microbiota of Marine Fish. Front. Microbiol. 2018, 9, 873. [Google Scholar] [CrossRef]
- Ringø, E.; Zhou, Z.; Vecino, J.L.G.; Wadsworth, S.; Romero, J.; Krogdahl, Å.; Olsen, R.E.; Dimitroglou, A.; Foey, A.; Davies, S.; et al. Effect of dietary components on the gut microbiota of aquatic animals. A never-ending story? Aquac. Nutr. 2016, 22, 219–282. [Google Scholar] [CrossRef]
- Carolus, H.; Van Dyck, K.; Van Dijck, P. Candida albicans and Staphylococcus Species: A Threatening Twosome. Front. Microbiol. 2019, 10, 2162. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pascual, D.; Estellé, J.; Dutto, G.; Rodde, C.; Bernardet, J.-F.; Marchand, Y.; Duchaud, E.; Przybyla, C.; Ghigo, J.-M. Growth Performance and Adaptability of European Sea Bass (Dicentrarchus labrax) Gut Microbiota to Alternative Diets Free of Fish Products. Microorganisms 2020, 8, 1346. [Google Scholar] [CrossRef] [PubMed]
- Mzula, A.; Wambura, P.N.; Mdegela, R.H.; Shirima, G.M. Present status of aquaculture and the challenge of bacterial diseases in freshwater farmed fish in Tanzania; A call for sustainable strategies. Aquac. Fish. 2021, 6, 247–253. [Google Scholar] [CrossRef]
- Pelusio, N.F.; Parma, L.; Volpe, E.; Ciulli, S.; Errani, F.; Natale, S.; De Cesare, A.; Indio, V.; Carcano, P.; Mordenti, O.; et al. Yeast-extracted nucleotides and nucleic acids as promising feed additives for European sea bass (Dicentrarchus labrax) juveniles. Front. Mar. Sci. 2023, 10, 1145660. [Google Scholar] [CrossRef]
- Ladner, J.T.; Welch, T.J.; Whitehouse, C.A.; Palacios, G.F. Genome Sequence of Weissella ceti NC36, an Emerging Pathogen of Farmed Rainbow Trout in the United States. Genome Announc. 2013, 1, 10–1128. [Google Scholar] [CrossRef]
- Abriouel, H.; Lerma, L.L.; Muñoz, M.C.C.; Montoro, B.P.; Kabisch, J.; Pichner, R.; Cho, G.S.; Neve, H.; Fusco, V.; Franz, C.M.A.P.; et al. The controversial nature of the Weissella genus: Technological and functional aspects versus whole genome analysis-based pathogenic potential for their application in food and health. Front. Microbiol. 2015, 6, 1197. [Google Scholar] [CrossRef]








| CeFi® Pro | SDYE | |
|---|---|---|
| Crude protein | 50.0% | 65.0% |
| Crude fibre | 1.0% | 1.0% |
| Crude ash | 6.6% | 9.4% |
| Crude fat | 3.0% | 0.1% |
| Moisture | 5.0% | 5.0% |
| Ingredients (g/kg) | Control | ABY | SDYE |
|---|---|---|---|
| Soybean meal 48 1 | 150.0 | 150.0 | 150.0 |
| Sunflower meal 2 | 159.4 | 157.8 | 157.8 |
| Fishmeal 2 | 70.0 | 70.0 | 70.0 |
| Corn gluten meal 2 | 220.0 | 220.0 | 220.0 |
| Wheat gluten meal 2 | 138.2 | 138.2 | 138.2 |
| Fish oil 2 | 90.0 | 90.0 | 90.0 |
| Sunflower oil 3 | 120.8 | 120.8 | 120.8 |
| Lysine 4 | 20.0 | 20.0 | 20.0 |
| DL methionine 4 | 3.2 | 3.2 | 3.2 |
| Threonine 4 | 1.7 | 1.7 | 1.7 |
| Histidine 4 | 3.1 | 3.1 | 3.1 |
| Arginine 4 | 13.6 | 13.6 | 13.6 |
| CMC binder 4 | 5.0 | 5.0 | 5.0 |
| Vitamin and mineral premix 5 | 5.0 | 5.0 | 5.0 |
| ABY 6 | - | 2.5 | - |
| SDYE 6 | - | - | 2.5 |
| Proximate composition (% wet weight basis) | |||
| Protein | 49.45 ± 0.16 | 48.73 ± 0.76 | 49.34 ± 0.07 |
| Lipid | 24.15 ± 0.07 | 23.39 ± 0.62 | 22.43 ± 1.32 |
| Moisture | 3.78 ± 0.10 | 3.93 ± 0.08 | 3.85 ± 0.13 |
| Ash | 4.46 ± 0.27 | 4.13 ± 0.15 | 4.28 ± 0.22 |
| Nitrogen-free extract | 18.16 ± 0.27 | 19.82 ± 0.45 | 20.10 ± 1.34 |
| Gross energy (kcal/g) | 5.82 ± 0.01 | 5.78 ± 0.03 | 5.73 ± 0.07 |
| Gene | Gene Annotation | NCBI Accession Number | Primer Sequence (5′–3′) |
|---|---|---|---|
| gapdh | Glyceraldehyde 3-phosphate dehydrogenase | XM_014141819.2 | Fwd—GCACCCATCGCCAAGGTTAT Rev—AGTCTTCTGTGTGGCTGTGA |
| ef1α | Elongation factor 1 alpha | NM_001141909 | Fwd—GGCTGATTGTGCTGTGCTTAT Rev—CACGAGTCTGCCCGTTCTTT |
| il-1β | Interleukin-1 beta | AY617117 | Fwd—AGGAGGGAAGCAGGGTTCA Rev—CATCAGGACCCAGCACTTGT |
| tnf-α | Tumor necrosis factor alpha | NM_001123617.1 | Fwd—GCACCGAAGACAACAAGGTTTA Rev—GCTGAACACTGCTCCCACATA |
| il-10 | Interleukin-1 beta | EF165028.1 | Fwd—ACGAAGGCATTCTACACCACTT Rev—CACCGTGTCGAGGTAGAACT |
| muc-2 | Mucin 2-like | XM_029770456.1 | Fwd—CGACTCAACGTGGATGTAGGA Rev—GCGACCACTAGCCAGAAAGA |
| cldn-15 | Claudin 15 | XM_014206890.1 | Fwd—GTCGGGATGCAGTGTTCTAAAG Rev—TGGGTGATGTTGAAAGCATACC |
| Parameters | Control | ABY | SDYE |
|---|---|---|---|
| Mucosal fold length (µm) | 203.74 ± 35.04 a | 307.41 ± 66.03 b | 375.98 ± 69.77 b |
| Muscularis thickness (µm) | 56.76 ± 13.73 | 72.97 ± 15.73 | 69.89 ± 13.11 |
| Lamina propria width (µm) | 16.81 ± 6.13 a | 11.86 ± 2.11 b | 13.31 ± 3.02 ab |
| Intestine goblet cell counts (n/100 µm) | 7.01 ± 1.71 a | 9.44 ± 1.91 b | 9.24 ± 2.44 ab |
| Skin goblet cell counts (n/200 µm) | 14.56 ± 2.85 a | 18.22 ± 2.48 b | 14.44 ± 2.66 a |
| Parameters | Control | ABY | SDYE |
|---|---|---|---|
| Haemoglobin (g/dL) | 0.03 ± 0.01 | 0.04 ± 0.01 | 0.03 ± 0.01 |
| Red blood cell count (106/µL) | 0.89 ± 0.34 | 1.19 ± 0.26 | 1.13 ± 0.20 |
| White blood cell count (103/µL) | 11.32 ± 4.36 | 14.36 ± 3.63 | 12.71 ± 5.68 |
| Treatments | Raw Paired-End Reads | Non-Chimera Reads | Number of ASVs | Sequences Assigned to ASVs |
|---|---|---|---|---|
| Control | 109,556 ± 6,279 | 100,899 ± 6,575 | 3,800 | 94,946 ± 6,458 |
| ABY | 104,255 ± 1,768 | 94,188 ± 2,059 | 3,757 | 88,385 ± 2,015 |
| SDYE | 103,025 ± 7,364 | 94,952 ± 5,462 | 3,399 | 89,588 ± 5,106 |
| Treatments | Good’s Coverage | Chao1 Index | Shannon’s Diversity Index | Simpson’s Index of Diversity |
|---|---|---|---|---|
| Control | 1.00 ± 0.00 | 880.79 ± 182.64 | 6.94 ± 0.38 | 0.96 ± 0.01 |
| ABY | 1.00 ± 0.00 | 902.04 ± 80.44 | 6.85 ± 0.60 | 0.96 ± 0.03 |
| SDYE | 1.00 ± 0.00 | 775.58 ± 193.29 | 6.53 ± 1.08 | 0.94 ± 0.08 |
| Groups | ANOSIM | ADONIS | ||
|---|---|---|---|---|
| R-Value | p-Value | F-Value | p-Value | |
| Control–ABY | 0.428 | 0.001 | 4.311 | 0.002 |
| Control–SDYE | 0.193 | 0.002 | 3.125 | 0.001 |
| ABYE–SDYE | 0.188 | 0.004 | 2.436 | 0.008 |
| Parameters | Control | ABY | SDYE |
|---|---|---|---|
| Initial body weight (g) | 37.10 ± 0.00 | 37.10 ± 0.17 | 37.03 ± 0.06 |
| Final body weight (g) | 66.64 ± 2.52 | 63.18 ± 1.58 | 67.69 ± 0.61 |
| Mean weight gain (g) | 29.54 ± 4.37 | 26.08 ± 2.58 | 30.65 ± 1.11 |
| Feed intake (g) | 27.42 ± 0.58 | 26.25 ± 0.90 | 27.45 ± 0.48 |
| Specific growth rate (%/day) | 0.93 ± 0.11 | 0.85 ± 0.06 | 0.96 ± 0.02 |
| Feed conversion ratio | 0.94 ± 0.12 | 1.01 ± 0.07 | 0.90 ± 0.02 |
| Survival (%) | 100.00 ± 0.00 | 100.00 ± 0.00 | 100.00 ± 0.00 |
| Fulton’s condition factor (K) | 1.20 ± 0.08 | 1.20 ± 0.06 | 1.17 ± 0.03 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Odu-Onikosi, S.G.; Momoh, T.A.; Abarra, S.T.; Wood, N.E.; Amulejoye, F.D.; Emery, M.; Harper, G.M.; Eynon, B.; Kuri, V.; Kühlwein, H.; et al. Impact of Autolysed Brewer’s Yeast and Soluble Dried Yeast Extract on Growth Performance and Mucosal Health of Atlantic Salmon (Salmo salar) Parr. Animals 2025, 15, 323. https://doi.org/10.3390/ani15030323
Odu-Onikosi SG, Momoh TA, Abarra ST, Wood NE, Amulejoye FD, Emery M, Harper GM, Eynon B, Kuri V, Kühlwein H, et al. Impact of Autolysed Brewer’s Yeast and Soluble Dried Yeast Extract on Growth Performance and Mucosal Health of Atlantic Salmon (Salmo salar) Parr. Animals. 2025; 15(3):323. https://doi.org/10.3390/ani15030323
Chicago/Turabian StyleOdu-Onikosi, Sheu G., Taofik A. Momoh, Sherilyn T. Abarra, Noah E. Wood, Folasade D. Amulejoye, Matthew Emery, Glenn M. Harper, Benjamin Eynon, Victor Kuri, Holger Kühlwein, and et al. 2025. "Impact of Autolysed Brewer’s Yeast and Soluble Dried Yeast Extract on Growth Performance and Mucosal Health of Atlantic Salmon (Salmo salar) Parr" Animals 15, no. 3: 323. https://doi.org/10.3390/ani15030323
APA StyleOdu-Onikosi, S. G., Momoh, T. A., Abarra, S. T., Wood, N. E., Amulejoye, F. D., Emery, M., Harper, G. M., Eynon, B., Kuri, V., Kühlwein, H., & Merrifield, D. L. (2025). Impact of Autolysed Brewer’s Yeast and Soluble Dried Yeast Extract on Growth Performance and Mucosal Health of Atlantic Salmon (Salmo salar) Parr. Animals, 15(3), 323. https://doi.org/10.3390/ani15030323






