Abstract
Although most cases are identified at a late stage, breast cancer is the most public malignancy amongst women globally. However, mammography for the analysis of breast cancer is not routinely available at all general hospitals. Prolonging the period between detection and treatment for breast cancer may raise the likelihood of proliferating the disease. To speed up the process of diagnosing breast cancer and lower the mortality rate, a computerized method based on machine learning was created. The purpose of this investigation was to enhance the investigative accuracy of machine-learning algorithms for breast cancer diagnosis. The use of machine-learning methods will allow for the classification and prediction of cancer as either benign or malignant. This investigation applies the machine learning algorithms of random forest (RF) and the support vector machine (SVM) with the feature extraction method of linear discriminant analysis (LDA) to the Wisconsin Breast Cancer Dataset. The SVM with LDA and RF with LDA yielded accuracy results of 96.4% and 95.6% respectively. This research has useful applications in the medical field, while it enhances the efficiency and precision of a diagnostic system. Evidence from this study shows that better prediction is crucial and can benefit from machine learning methods. The results of this study have validated the use of feature extraction for breast cancer prediction when compared to the existing literature.
1. Introduction
Breast cancer is a serious and frequent reproductive cancer that primarily affects women and is caused by breast tumors. A breast tumor is a nipple discharge, lump, or a change in skin texture around the nipple area caused by the irregular growth of tissues in the breast. Tumors are classified as benign or malignant. A breast lump is a worldwide virus that affects the lives of women in the age bracket of 25–50. Doctors have found hormonal, lifestyle, and environmental factors that may raise an individual’s chances of having one. More than 5% of affected breast cancer patients were connected to familial gene mutations that have been passed down through the generations and also to obesity, aging, and hormonal abnormalities after menopause [1].
As a result, there is no prevention mechanism for breast cancer, although early discovery can greatly improve the diagnosis [2]. Furthermore, the treatment expenses might be significantly reduced as a result of this. However, because cancer symptoms might be uncommon at times, early detection is challenging. Mammograms and self-breast examinations are essential for detecting any early anomalies before the malignancy progresses. As a result, tumor diagnosis requires the automation of diagnostic systems [3].
Numerous studies have sought to utilize machine-learning algorithms to detect cancer survivability in humans, and they have established that these systems are more effective in detecting cancer diagnosis. Various machine-learning approaches have since been created for the analysis and treatment of breast cancer, as well as many aided breast cancer diagnosis approaches to improve diagnostic accuracy and lessen the number of bereavements [4].
Breast cancer is a predominant ailment that primarily distresses women, and its initial detection will hasten treatment. The basic objective of breast cancer treatment is to accurately forecast the presence of cancer and define the cancer category to regulate how to treat the disease. Nevertheless, determining the kind of breast cancer is one of the typical challenges in health-related investigations. The proper classification of breast cancer would lead to its early discovery, finding, behavior, and elimination, if possible. In addition, the precise classification of benign tumors helps save patients from receiving unneeded therapies [5].
Throughout the past few years, numerous establishments have amassed huge stores of data obtained from a variety of sources, stored in various formats. These data could be utilized in various application sectors, including medical, agronomy, and climate forecasting. These ever-increasing quantities of data exceed the capacity of conventional methods for evaluating and seeking hidden patterns and information for decision-making. By utilizing machine-learning methods, it is possible to examine data retrieved from medical data repositories [6]. For the effectiveness of disease prediction, algorithms and their use in knowledge discovery from health information sources are invaluable resources. Numerous studies have used the application of machine-learning algorithms for breast cancer prediction. Machine-learning algorithms are widely used in the advancement of breast cancer prediction models to promote operative decision-making. In current years, machine learning (ML) procedures have been executed in biomedicine to aid the fight against breast cancer [7]. It is a complex and time-consuming process to extract information from information to help the scientific analysis of breast cancer. Utilizing machine learning and feature extraction methods has substantially altered the breast cancer diagnosis process [8].
These methods were successful but have encountered slight fallbacks in areas such as accuracy and efficiency. This suggests using machine-learning techniques by improving the linear discriminant analysis algorithm for fetching latent components, support vector machine and random forest classifiers are further utilized to develop the prediction model for breast cancer analysis to aid in the medical field. Evidence from this investigation shows that better prediction is crucial. The study’s results report the benefit of using feature extraction in the prediction of breast cancer, providing fresh insight into the efficiency of linear discriminant analysis (LDA) and classification for breast cancer prediction. This helps to enhance classification performance in terms of performance evaluations such as accuracy, precision, and recall.
The following five sections are discussed in this paper: In the first section, we provide an overview of the study. Research on categorizing breast cancer is presented in Section 2. The research materials and techniques are discussed in Section 3, the research results are presented in Section 4, and the conclusion, a quick overview with criticisms of the results, is presented in Section 5.
2. Related Works
Researchers have recently employed machine-learning procedures to classify breast cancer based on diverse medical datasets. Many experts utilize these algorithms to address tough tasks because they offer good categorization results. In addition, data mining is used often in the therapeutic field to predict irregular events to acquire an improved knowledge and understanding of irredeemable diseases such as breast cancer [9]. The outcomes of utilizing machine learning in breast cancer discovery classification are encouraging. This is a list of works that are related to this method:
On the Wisconsin Breast Cancer (unique) Datasets, four key techniques were proposed [10]. To identify the highest classification accuracy, they examined the effectiveness and efficiency of various algorithms considering precision, accuracy, sensitivity, and specificity. SVM achieves a 97.13 percent accuracy and so surpasses all other methods. Finally, SVM demonstrated its efficacy in the prediction and breast cancer analysis, achieving the greatest results in terms of low margin of error and precision.
A comparison of commonly used machine-learning procedures for the detection of breast cancer and also its diagnosis was proposed [11] such as random forest, support vector machines, and Bayesian networks. For the dataset, the original Wisconsin Breast Cancer Dataset was applied as a training set to assess and make a comparison of the performance of the three machine-learning classifiers considering important parameters. The findings in the research presented an outline of existing machine-learning methods for detecting breast cancer.
Classification test sensitivity, accuracy, and specificity values of machine-learning algorithms were suggested [12]. The dataset is made up of features derived from digitized imageries of FNA tests. The dataset was then partitioned in the following way for machine-learning techniques implementation: 70% for training and 30% for testing. All the classifiers’ hyper-parameters were manually assigned. On the classification problem, all the given machine-learning procedures executed well (all of them with over 90% test accuracy). With a test accuracy of 99.04%, the multilayer perceptron method stands out among the implemented methods.
Random forest, k Nearest Neighbor, and naive Bayes were utilized to compare widely using machine-learning procedures and approaches for predicting breast cancer [13]. To evaluate the performance of numerous machine-learning procedures in terms of main attributes including precision and accuracy, the Wisconsin Diagnosis Breast Cancer Dataset was used as a training set. The results are quite encouraging and can be utilized for both detection and treatment.
The breast cancer diagnostic accuracy of various machine-learning procedures was proposed [14]. There were 1879 publications assessed in total, with 11 being chosen for efficient evaluation and meta-analysis. Five different classification machine-learning approaches for predicting the threat of breast cancer were identified. The results were 90% with the SVM, putting it in the outstanding grouping. The meta-analysis found that the SVM algorithm is more accurate than other machine-learning procedures at calculating breast cancer danger.
Support vector machines and 10-fold cross-validation were utilized to generalize classifier performance and address the issue of data overfitting in a suggested Breast Cancer Diagnosis model [15]. The suggested model outperformed existing models created to diagnose using the original Wisconsin Breast Cancer Dataset, from the investigational results. The breast cancer diagnosis model was associated with other models in use, such as random forest and naive Bayes, ROC curve, lift curve, class accuracy, AUC, and a calibration plot, which were some of the evaluation measures used. The Breast Cancer Diagnosis model had the highest accuracy of the four deployed models at 98.1%.
The Wisconsin Breast Cancer Database was used to propose a methodology of an artificial neural network for diagnosing breast cancer [16]. The study’s major goal was to examine and explain how ANN provides more effective and better solution arrangements when used in conjunction with group AI calculations for diagnosing bosom malignant growth, even when the components are reduced. When compared to previous research, it was discovered that the ANN approach with calculated calculation achieves 99.00 percent precision when compared to another AI calculation.
A machine-learning method was used to try to solve the challenge of automatic breast cancer detection [17]. The system was developed in stages, with three independent trials using the breast cancer dataset. In the first test, they demonstrated that after effective configuration, the three most popular evolutionary algorithms may produce the same results. The second experiment looked at how combining feature selection approaches enhances accuracy, while the third experiment looked at how to create a machine-learning supervised classifier automatically. They used the GP method to try to address the hyperparameter problem, which is a difficult problem for machine-learning systems. Among the many configurations, the proposed algorithm chose the most appropriate algorithm. The Python library was used in all the experiments. Although the proposed method yielded notable findings by analyzing an ensemble of techniques from an exhaustive machine-learning technique, it took a much longer time to complete.
A new breast cancer detection technology called DNNS was also proposed [18]. Unlike other techniques, this one is based on a deep neural network’s support value. A normalization technique is used to enhance the performance, efficacy, and quality of photographs. Experiments have shown that the suggested DNNS is far superior to existing approaches. It was determined that the proposed algorithm is beneficial in terms of performance, efficiency, and image quality, all of which are critical in today’s medical systems.
A strategy for enhancing the accuracy of different classifiers was suggested [19]. The classifiers were also validated and compared using two benchmark datasets. Because the chance of examples being grouped into the common class is relatively high, procedures are probable to categorize new observations to the majority class in the classification phase. During this study, such challenges are resolved. A 10-fold cross-validation assesses the outcomes and evaluates each classifier’s efficiency. The experiments revealed that applying a resample filter advances the performance of the classifier, with J48 outperforming others in the breast cancer dataset and SMO outclassing others in the WBC dataset. They focused on how to employ resampling approaches to recover the classification accuracy of identifying breast cancer in imbalanced data with missing values. Three classifier techniques were used on two separate breast cancer datasets in this study. The use of the resample filter in the preprocessing step improves the performance of the classifiers, according to the findings.
Researchers employed supervised learning techniques and prediction models [20]. k Nearest Neighbor was shown to be the most precise predictor, with 91.6% accuracy. In comparison to previous research, the model can offer a higher level of precision.
A framework for applying machine-learning approaches to accurately and quickly diagnose breast cancer was proposed [21]. Random forest was used to apply the suggested methodology to the SEER dataset of breast cancer and obtained extremely substantial outcomes with an accuracy of 99.9%. Various rules were also offered in support of the A-priori algorithm’s diagnosis of breast cancer.
The Wisconsin Breast Cancer Dataset was utilized for the comparison of most of the major machine-learning procedures for detection and diagnosis [22]. Supervised learning-decision tree, random forest, multilayer perception, support vector machine, and linear regression were compared in both the classification and regression categories. The results revealed that under the classification algorithm, the support vector machine provides high accuracy; however, under the regression methodology, multilayer perception regression delivers reduced errors.
To accurately diagnose breast cancer in its early stages, it was recommended in [23] that better machine-learning models should be developed. Using a dataset of repetitive blood investigations with anthropometric dimensions for breast cancer diagnosis, the performance of several machine-learning methods was compared. A feature selection technique called neighborhood component analysis was utilized to recognize useful biomarkers for breast cancer prediction. Comparing the effectiveness of the various classifier models, two distinct data partitioning strategies were utilized. All classifiers were optimized using a Bayesian optimization technique, which resulted in the highest possible prediction precision. The effectiveness of each classifier was evaluated using a variety of systems of measurement. The results demonstrated that k Nearest Neighbor, which is based on Bayesian optimization, outperformed the other methods of machine learning tested using the hold-out data division procedure, with an accuracy of 95.833%. The findings provide fresh insight into the potential of enhanced methods for the early diagnosis of breast cancer.
Diagnosis and detection of breast cancer were suggested in a comparison study employing data visualization and machine-learning methods [23]. Several machine-learning strategies for detecting breast cancer were suggested in this paper. The results acquired using the logistic regression model with all features involved demonstrated classification accuracy (98.1%) and the strategy exhibited an improvement in accuracy.
Because the selection is carried out in tandem with the model tuning process, it typically results in a middle ground between the two feature selection techniques [24]. The most frequent feature selection methods of this type are Lasso and Ridge regression, while a decision tree develops a model utilizing a variety of feature selection approaches. “Classifiers” and “Meta Classifiers” are components of each nested ensemble classifier. There can be more than two classification methods utilized in a meta-classifier. The nested ensemble classifiers with two layers were created by them. Typically, meta-classifiers use a layered, nested ensemble of classifiers, with two or three distinct classification algorithms per layer. The Wisconsin Dataset was used in the experiments and the k-fold cross-validation method was utilized to evaluate the model’s performance. Classification accuracy, precision, recall, F1 measure, ROC, and computing times were compared between the proposed two-layer nested ensemble classifiers and those of individual classifiers (Bayes Net and naive Bayes). The results specified that the suggested two-layer nested ensemble models were superior to both single classifiers and prior attempts. Both the SVM–naive Bayes3-Meta Classifier and the SV–BayesNet3-MetaClassifier performed at 98.07 percent accuracy (K = 10). Alternatively, the SV–naive Bayes3-MetaClassifier is more productive because it requires less time to create the model.
Early breast cancer detection with machine-learning algorithms can effectively categorize breast growth as benign or malignant [25]. The dataset used was the Wisconsin Breast Cancer Dataset. This dataset was split into two parts, with 65 percent being training data and 35 percent being testing data. A comparative analysis of various classification algorithms was conducted to determine the optimum strategy. It was discovered that parameter selection is quite crucial in the classification process. According to the findings, the random forest classifier produces the best results with 96.5 percent accuracy.
All top-tier machine-learning classifiers were used on a variety of data profiles [26]. Linear-SVM applied to the data profile with features fused from the clinical, gene expression, CNA, and CNV datasets yielded the highest accuracy (88.36%) for IntClust subtyping, with Jaccard and Dice scores of 0.802 and 0.8835, respectively. However, when using Linear-SVM and E-SVM classifiers on a variety of data profiles that incorporate features from histopathology pictures, an accuracy of 97.1% was attained, with Jaccard scores ranging from 0.9439 to 0.9472, and a Dice score of 0.971. In conclusion, our research adds credence to the claim that fusing features from multiple METABRIC datasets can enhance the performance of classifiers used to categorize breast cancer subtypes. Histopathological pictures also show promise in identifying Pam50 subtypes, and their use is anticipated to boost the accuracy of IntClust subtyping when applied to a larger population.
Histopathological images as biological data are particularly scarce, contributing to the widespread perception of imbalance in breast cancer data [27]. Recent research has shown that the cascade Deep Forest ensemble model, by learning hyper-representations using cascade ensemble decision trees, can outperform other alternatives, such as the general ensemble learning methods and the conventional deep neural networks (DNNs), for imbalanced training sets. In this study, we use a cascading deep forest to classify breast cancer subtypes, IntCluster and Pam50, with a variety of multi-omics datasets and settings. Accuracy was determined to be 83.45% for five subtypes and 77.55% for 10. This work is significant because it demonstrates that the cascade deep forest classifier can be used with only gene expression data to achieve accuracy on par with other techniques that have better computational performance, with times of about 5 s for 10 subtypes and 7 s for five subtypes recorded.
Using medical screening and diagnostics, a breast cancer prediction model was established [28]. LDA extracts useful information to avoid overfitting and improve prediction accuracy. This reduces the dataset’s features. The proposed approach can develop new features from old ones and then remove them. The newly developed features summarized the original set’s information. LDA is beneficial for determining if a set of features can predict breast cancer. The LDA–SVM prediction model uses a support vector machine (SVM) for reliable prediction. The suggested model achieves 99.2% accuracy, 98.0% precision, and 99.0% recall when tested on the Wisconsin Diagnostic Breast Cancer (WDBC) Dataset from the University of California-Irvine machine learning repository. When paired with feature extraction, SVM efficiently handles classification challenges.
A multi-modular breath analysis system with machine learning to detect cancer-specific breath from sensor response curves (taxonomies of clusters) was used [29]. They evaluated 54 gastric cancer patients’ breaths and 85 controls. An analyzer with gold nanoparticles and metal oxide sensors was used. Curve forms and other comparative features were employed to examine sensor responses. These features were used to train naive Bayes classifiers, SVMs, and random forests. The trained models’ accuracy was 77.8% (sensitivity: 66.54%; specificity: 92.39%). Shape-based characteristics enhanced accuracy and sensitivity in most circumstances. This data analysis technique seems to have the potential for detecting stomach cancer-specific breath. The cluster taxonomy-based detector effective range modeling improves outcomes.
Machine-learning approaches can predict and diagnose breast cancer early and have become a research focus [30]. This study implemented five machine-learning algorithms: support vector machine (SVM), random forest, logistic regression, decision tree (C4.5), and k Nearest Neighbour (kNN) to the Breast Cancer Wisconsin Diagnostic Dataset, and then compared their performance. This study work aims to detect and diagnose breast cancer using machine-learning algorithms and determine which are the most accurate and precise. The support vector machine was the most accurate classifier (97.2%). All work is performed in Anaconda using Python and Scikit-learn.
Studies in the literature lead us to believe that breast cancer classification models are employed in machine-learning-based detection algorithms. Looking at the research published thus far, it becomes clear that numerous studies have been conducted on breast cancer. With this improved detection capability as a result of the data obtained, a new chapter was added to the literature. When applied to the Wisconsin Breast Cancer Datasets, this study combines the characteristics of machine-learning algorithms (support vector machine), random forest, and feature extraction approaches (linear discriminant analysis) to aid in prediction and decision-making. The purpose of the analysis is to be as precise as possible while making as few mistakes as possible. Accuracy, precision, and recall are used as yardsticks to evaluate the algorithm’s efficacy and efficiency.
3. Materials and Methods
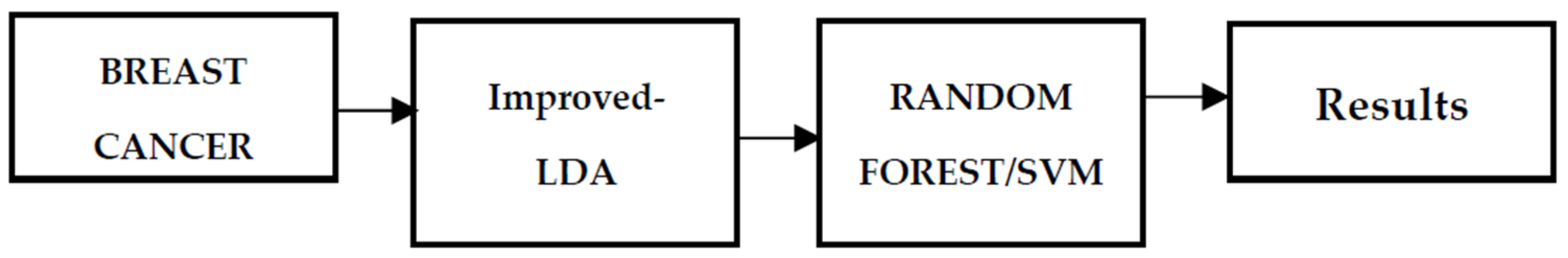
The system in this research has two primary stages: feature extraction and categorization. As an input, the dataset loads, it is then given to a feature extraction method called linear discriminant analysis, which extracts the relevant data from the dataset and classifies it using random forest and support vector machines, with the results being assessed and compared using performance metrics. Figure 1 shows the proposed overall system design of the proposed work.

Figure 1.
Proposed Workflow.
3.1. Dataset
The Wisconsin (diagnostic) Breast Cancer Dataset obtained from Kaggle will be used for this study. Breast cancer is a condition characterized by the uncontrolled growth of breast cells. Instances of breast cancer might vary greatly. Which breast cells become malignant determines the subtype of breast cancer. The Health Wisconsin Diagnostic Breast Cancer (WDBC) Dataset is used to determine if a tumor is cancerous or benign. https://www.kaggle.com/datasets/uciml/breast-cancer-wisconsin-data/discussion/62297 (accessed on 1 February 2022). The definition of attributes: Identification number; Diagnosis (1–32) (M = malignant, B = benign). There are 569 instances and 32 attributes. ID/diagnosis (30 input attributes with real-world values).
3.2. Improved Linear Discriminant Analysis
Linear discriminant analysis is a method for simplifying classification problems in supervised machine learning. Using it requires distinguishing between two or more categories; thus, it is employed for modeling set differences. It is a tool for bringing a feature from one dimension down to a lower-dimensional space. The method’s goal is to ensure optimal class separability by transforming characteristics from a high-dimensional space into a lower-dimensional space in which the proportion of between-class variation to within-class variance is maximized [31,32]. Because of its widespread use in the preprocessing of machine-learning classification applications and its ability to transform features into a lower dimensional space by minimizing the ratio of the between-class variance to the within-class variance, linear discriminant analysis (LDA) is the primary feature extraction technique used in this work. An enhancement of LDA (Algorithm 1) is presented herein by simply entering the sample’s independent variables and getting the class means and prior probability calculated. We compute the average vector dimensions for each group in the dataset. To determine the scattering matrix requires Eigenvectors (e1, e2, …, ed) and their corresponding Eigenvalues (1, 2, …, d), a dk matrix W can be created by sorting the Eigenvectors by decreasing Eigenvalues and picking the k Eigenvectors with the highest Eigenvalues. Then, the data is reconstructed in a new subspace using the Eigenvector matrix W produced above. Its representation in matrices is as concise as Y = XW. It estimates the pooled covariance matrix and computes the covariance matrices for each group. Following that, it determines the LDA discriminant function and labels the classes.
| Algorithm 1. Improved Linear Discriminant Analysis |
|
3.3. Random Forest Classifiers
Random forest classifiers are made up of numerous separate decision trees that act as a team. Every tree contained in the random forest predicts a class, with the highest votes and develops the model’s forecast. While some trees are inappropriate, numerous others will be precise, letting the forest transfer in the correct way [33]. The resulting points are the necessities for a positive random forest (Algorithm 2):
- The structures must have open signals for models formed with them to outperform random guesses.
- The separate trees’ predictions (and thus errors) must have minimal correlations.
Random forest pseudocode is divided into stages, namely, random forest generation pseudocode and prediction/forecast from the formed random forest classifier pseudocode.
| Algorithm 2. Random Forest [34] |
|
The random forest method begins by selecting “k” features at random from a total of “m” features. Then, it uses the best split strategy to discover the root node utilizing the randomly picked “k” features in the resulting phase and calculates the offspring nodes utilizing the same finest divisible method as before. Then, the first three stages need to be run until it generates a tree with a source node and the goal as a leaf node. In conclusion, phases 1–4 should then be replicated to generate “n” trees at random. The random forest is made up of these randomly produced trees.
3.4. Support Vector Machine
Support vector machine (Algorithm 3) reduces simplification errors. It is an all-inclusive and adjustable machine-learning model that can handle linear and nonlinear classification, regression, and outlier detection [35].
Support vector machine models can classify any novel text after feeding them groups of labeled training data for each class. The benefit is that more complex relationships between your data components can be kept track of without having to do complex transformations [36]. The disadvantage is that it takes much longer to train because of the increased computational expense. A support vector machine can be “taught” to detect fraudulent credit card activity by comparing thousands of records of actual and hypothetical credit card transactions. An SVM may be trained to identify handwritten digits by analyzing many scanned images of handwritten 0 s, 1 s, and other characters. Now more than ever, SVMs are finding widespread use in a wide variety of productive biological settings. Instinctive classification of gene expression profiles is another communal biomedical use of support vector machines. Theoretically, a gene expression profile derived from a peripheral fluid or tumor sample might be evaluated by a support vector machine to provide a diagnosis or prognosis. In biology, SVMs are used to categorize protein and DNA sequences, microarray expression patterns, and mass spectra [37].
| Algorithm 3. Support Vector Machine [38] |
|
3.5. Performance Evaluations
Accuracy is the number of correct forecasts provided by the model over predictions of all kinds in the categorization tasks. Accuracy is a good metric in the almost equilibrated target variable classes of the data.
Precision is a metric that shows us how much the forecasts are right.
The fraction of true positives that are accurately identified as positives is measured by sensitivity.
Specificity is defined as the percentage of genuine negatives that are accurately detected as opposed to positive, known as selectivity or real negative rate (TNR).
The F1 score measures the accuracy of a test, defined as the harmonic mean of precision and recall.
3.6. Research Tools
This study suggests developing the implementation using Python on an icore7 processor, with 1.1 GHz speed, 4 GB RAM, 20 GB hard disk, and Windows 7 OS.
4. Results and Discussions
In this study, a feature extraction technique and two machine-learning classifiers were executed on the Jupyter platform. Precisely, this section presents the outcomes of the proposed model. This study implements LDA, a dimensionality reduction technique, with the classification procedures SVM and random forest on the raw data, which is the Wisconsin Breast Cancer Dataset, and passes that raw data without LDA through the previously mentioned classifiers. The data consists of 569 instances and 32 attributes. A common practice in machine learning is to divide the available data into a train, test, and validation set. In this instance, 80% of the loaded dataset was used for training and 20% was used for testing. We then employed a technique called feature scaling, which is used to standardize the range of the data’s independent variables or features.
4.1. Feature Extraction
A technique of feature extraction, linear discriminant analysis (LDA), was implemented, which reduced the dimensionality of the data and extracted the most important columns and identified the least important columns then removed them, which meant ten (10) least important columns were dropped. After passing the data through the feature extraction technique (LDA), which reduced the data to give the most important columns of the dataset, the new data are then passed to the machine-learning classifiers, random forest, and SVM.
4.2. Random Forest Classifier
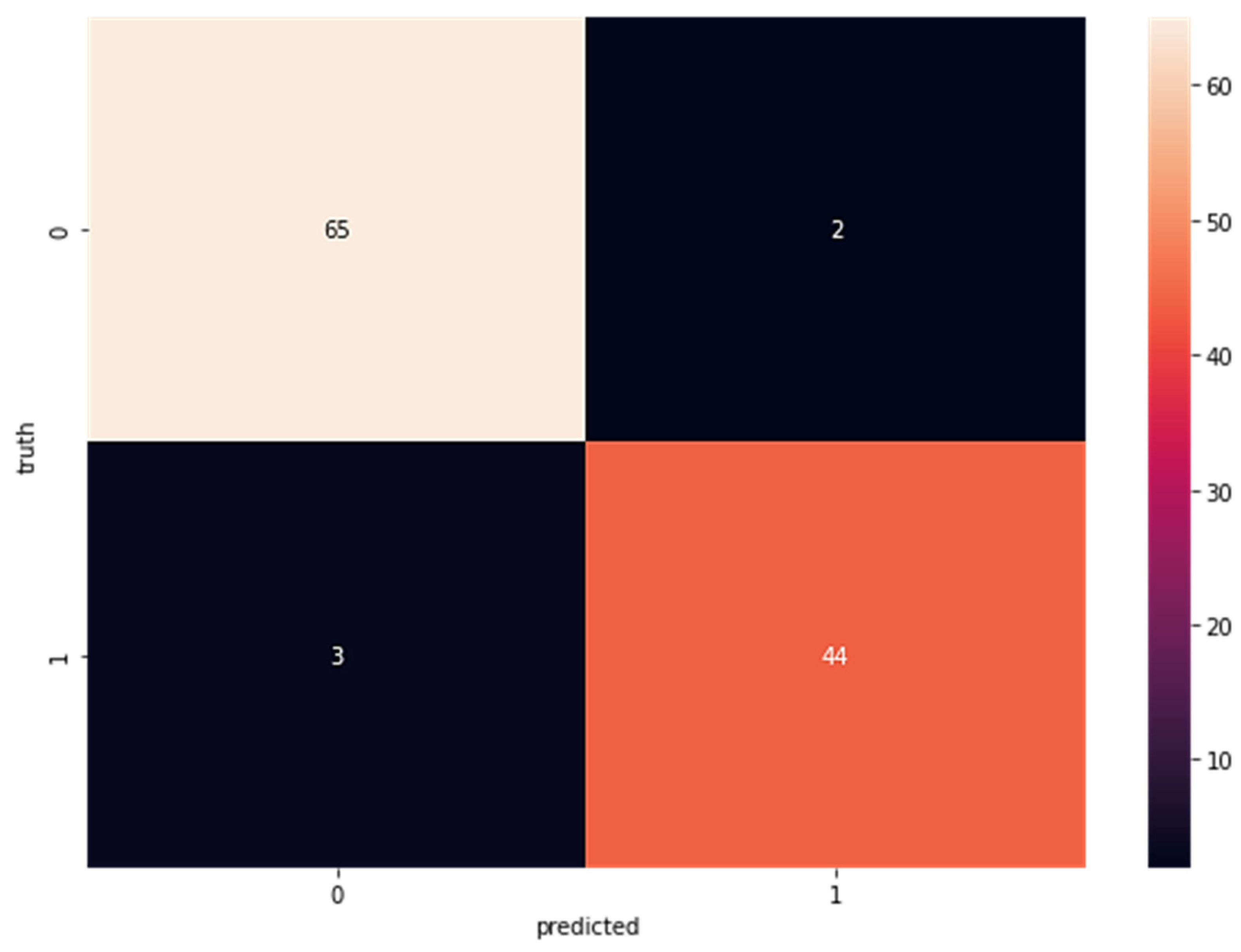
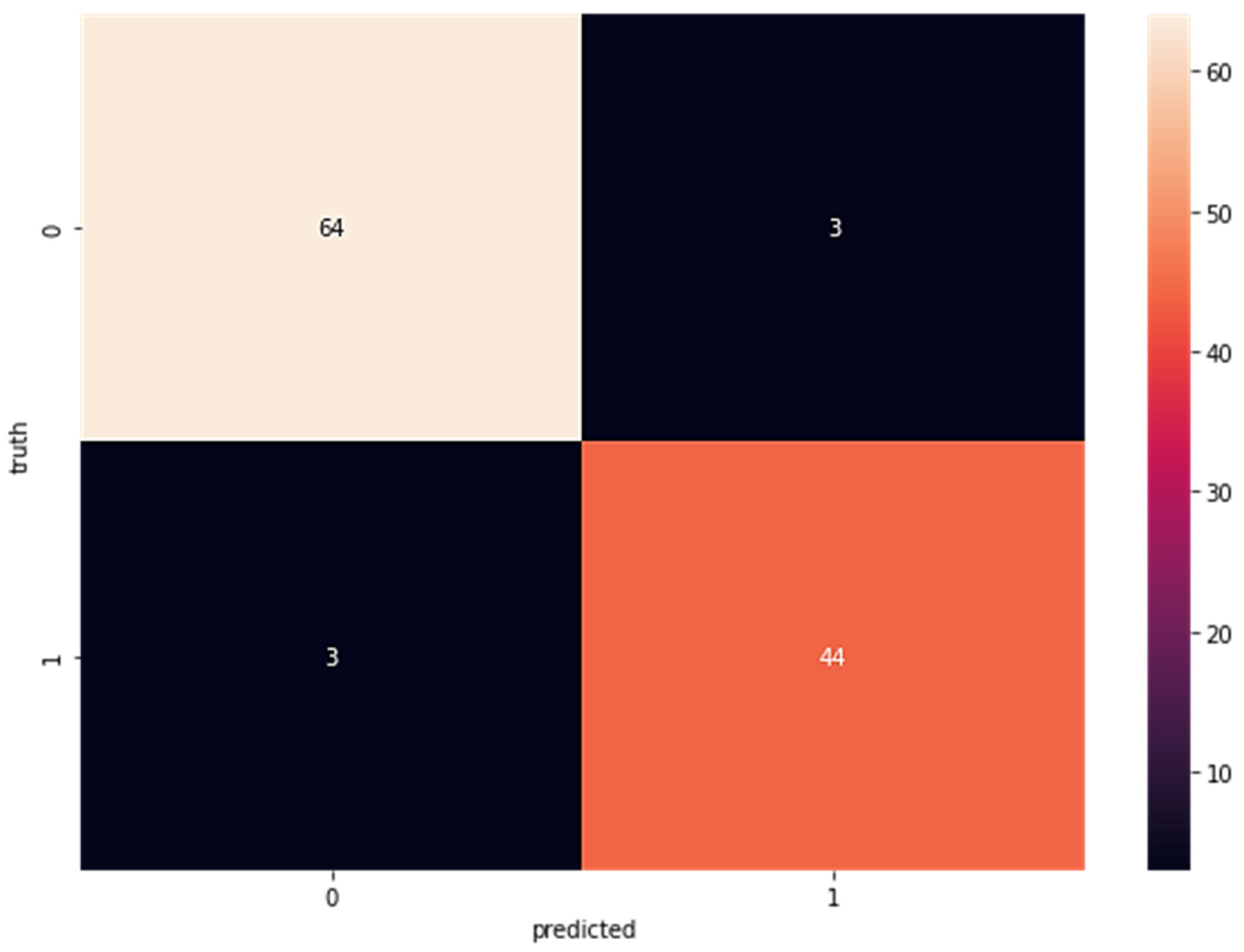
In a random forest classifier, many individual decision trees collaborate to harvest a single output. The random forest is a collection of trees, each of which makes a classification prediction; the classification with the most votes determines the model’s prediction. In the first step after LDA, this classifier is applied to the newly acquired data. Confusion matrices for the random forest categorized portion are displayed in Figure 2. The percentages add up to 65 for the true positive rate (TP), two for false positives (FP), three for false negatives (FN), and 44 for true negatives (TN). Confusion matrices for data fed directly into the random forest classifier without previously being processed by LDA are displayed in Figure 3; these matrices reveal a true positive rate (TP) of 64, a false positive rate (FP) of three, a false negative rate (FN) of three, and a true negative rate (TN) of 44.

Figure 2.
Confusion Matrix for Wisconsin Breast Cancer Dataset with LDA Using Random Forest (TP = 65, FP = 2 FN = 3 TN = 44).

Figure 3.
Confusion Matrix for Wisconsin Breast Cancer Dataset Without LDA Using Random Forest (TP = 64, FP = 3 FN = 3 TN = 44).
The results of the performance of the dataset after being passed through the random forest classifier, where the classifier achieved accuracy = 95.61, precision = 97.0, recall/sensitivity = 95.6, F1 score = 96.3, specificity = 95.7.
The decision boundary of a support vector machine is optimized to cut down on over-generalization. It is a flexible machine-learning model that handles linear and nonlinear classification, regression, and outlier detection with ease. When the LDA-refined dataset is complete, it is sent into a support vector machine classifier.
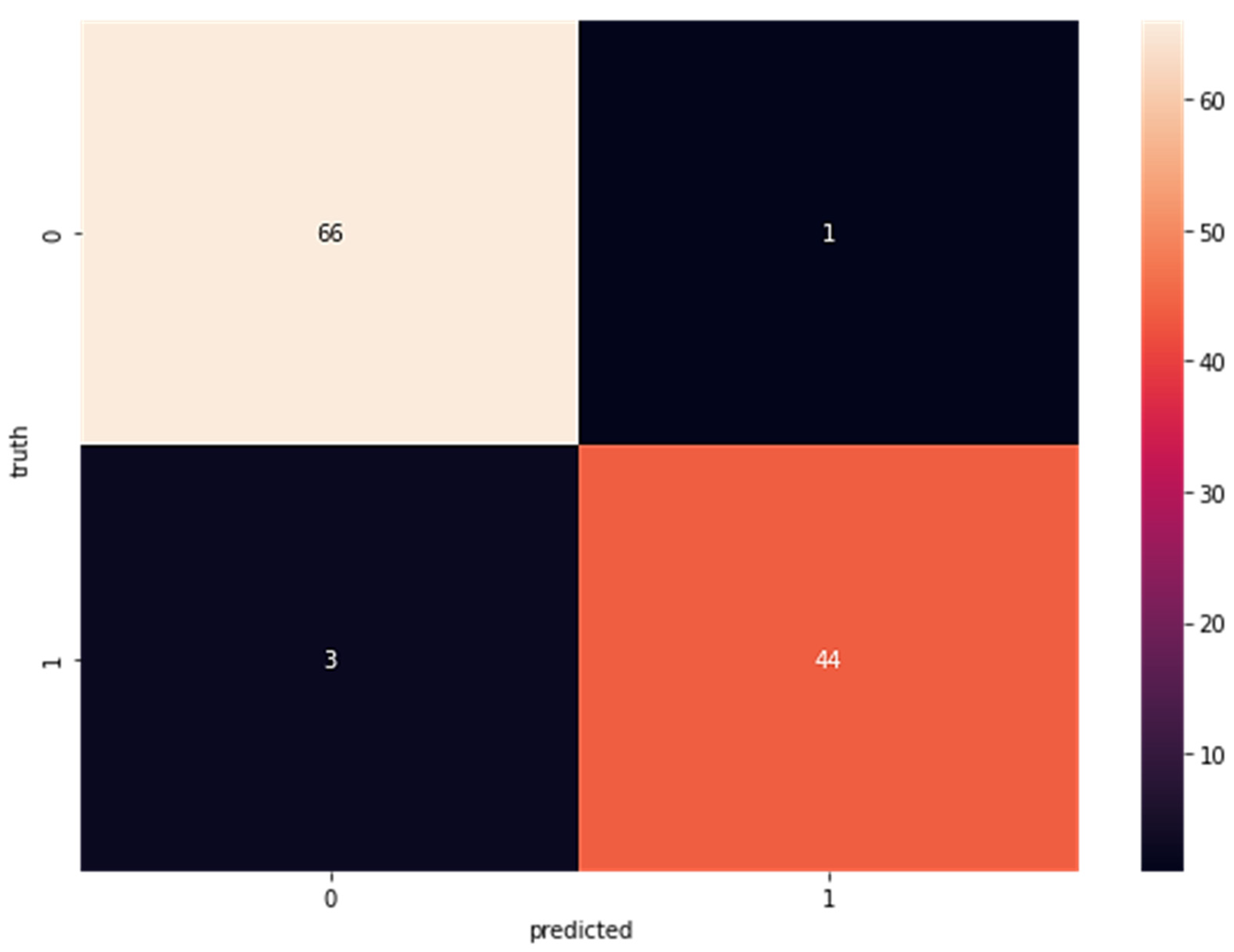
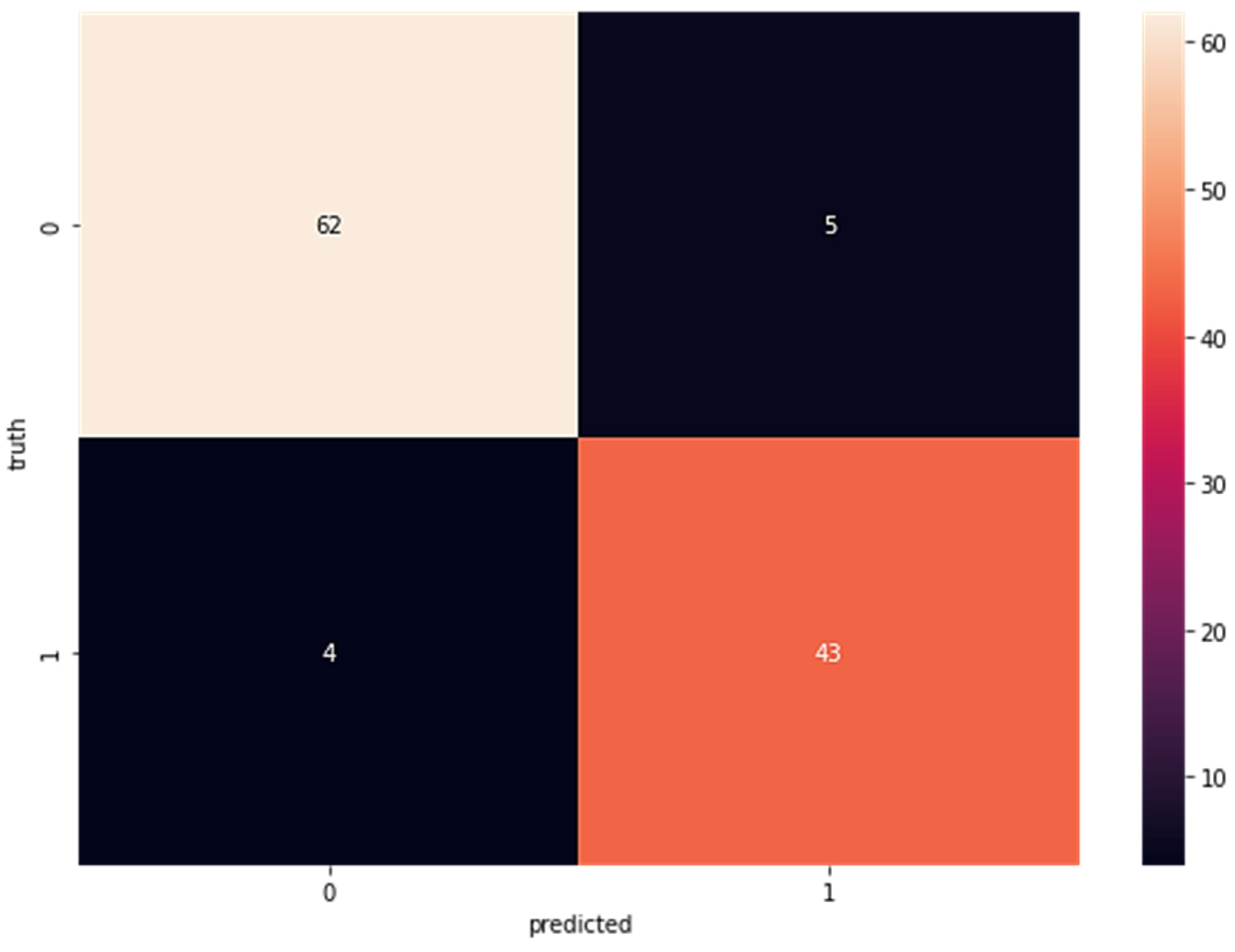
Confusion matrices for the SVM-classified subcomponent are displayed in Figure 4. The sensitivity is 66, the specificity is one, the false negatives are three, and the specificity of the real negatives is 44. As can be seen in Figure 4, when the dataset was fed directly into the SVM classifier without first being run via LDA, the results indicated a true positive rate (TP) of 63, a false positive rate (FP) of five, a false negative rate (FN) of four, and a true negative rate (TN) of 43. Figure 5 shows the results of SVM without LDA.

Figure 4.
Confusion Matrix Results for Wisconsin Breast Cancer Dataset with LDA Using SVM (TP = 66 FP = 1 FN = 3 TN = 44).

Figure 5.
Confusion Matrix for Wisconsin Breast Cancer Dataset Without LDA Using SVM (TP = 62, FP = 5 FN = 4 TN = 43).
After applying the SVM classifier to the dataset, the results are depicted in Figure 5; these show an accuracy of 97%, a precision of 99%, a recall of 96%, a sensitivity of 97%, an F1 score of 97%, and a specificity of 97%.
In this research, data classification is carried out using random forest and SVM, therefore the data are sent into these classifiers after having been subjected to feature extraction using LDA. Table 1 displays the evaluation metrics used to analyze the experimental confusion matrices.

Table 1.
Performance Metrics Table for LDA+ classifiers and classifiers without LDA.
Numerous studies were proposed in this study and Table 1 shows their evaluations, with LDA and SVM outperforming with 96.4% accuracy. Table 2 shows comparisons of the results obtained by several other related studies.

Table 2.
Comparison with Related Works.
The accuracy of this study’s prediction method was commensurate with its predecessors (as shown in Table 2). The accuracy rate in determining breast cancer was the goal of the proposed methodology and this goal was met when compared to other predictions in the literature.
5. Conclusions
In this study, the machine-learning classifiers of random forest and support vector machines were applied to a 562 instance and 39 attribute dataset of breast cancer cases in Wisconsin, obtained from Kaggle. The dataset was pre-processed and then separated into a training set and a testing set. To create a reliable predictive model, we first used the training set, which included 80% of the data, to find the best possible combination of variables, and then we used the testing set, which included the remaining 20% of the data, to conduct a fair evaluation of the final model’s performance on the training dataset. After applying linear discriminant analysis, a feature extraction technique for dimensionality reduction that selectively extracted the features needed to provide improved performance to the Wisconsin Breast Cancer Dataset, the new dataset was run through the classifiers random forest and support vector machine, with the former achieving an accuracy result of 95.6% and the latter of 96.4%; the results were then compared to prior related works. The findings of this study can aid in the detection of breast cancer. If breast cancer can be diagnosed and treated early on, it could save the lives of thousands of women and men every year. Several machine-learning methods, including naive Bayes, k Nearest Neighbor, and decision tree, are explored in this study, paving the way for further development of breast cancer diagnosis. LDA feature extraction improves breast cancer prediction systems. The suggested model builds a new dataset based on extracted features and improves performance. This paper suggests researching feature extraction strategies on datasets and computing important information to improve prediction accuracy. The LDA’s efficiency enhances the need for further feature extraction approaches to improve the model’s prediction accuracy and performance with malignant datasets. Answering these questions may yield more significant findings.
Author Contributions
Conceptualization, M.O.A. (Micheal Olaolu Arowolo) and M.O.A. (Marion Olubunmi Adebiyi); methodology, M.O.A. (Micheal Olaolu Arowolo), M.D.M., M.O.A. (Marion Olubunmi Adebiyi); software, M.O.A. (Micheal Olaolu Arowolo), M.D.M.; validation, M.O.A. (Marion Olubunmi Adebiyi), O.O.O.; formal analysis, M.O.A. (Marion Olubunmi Adebiyi), M.O.A. (Micheal Olaolu Arowolo); investigation, M.O.A. (Micheal Olaolu Arowolo), M.O.A. (Marion Olubunmi Adebiyi); resources, M.O.A. (Micheal Olaolu Arowolo), M.O.A. (Marion Olubunmi Adebiyi), M.D.M., O.O.O.; writing—original draft preparation, M.D.M., M.O.A. (Micheal Olaolu Arowolo); writing—review and editing, M.D.M., M.O.A. (Micheal Olaolu Arowolo) and M.O.A. (Marion Olubunmi Adebiyi); supervision, O.O.O.; project administration, O.O.O. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Data Availability Statement
https://www.kaggle.com/datasets/uciml/breast-cancer-wisconsin-data (accessed on 1 February 2022).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Labrèche, F.; Goldberg, M.S.; Hashim, D.; Weiderpass, E. Breast Cancer. In Occupational Cancers; Springer International Publishing: Cham, Switzerland, 2020; pp. 417–438. [Google Scholar]
- Hailu, T.; Berhe, H.; Hailu, D. Awareness of Breast Cancer and Its Early Detection Measures among Female Students, Northern Ethiopia. Int. J. Public Health Sci. 2016, 5, 213. [Google Scholar] [CrossRef]
- Akram, M.; Iqbal, M.; Daniyal, M.; Khan, A.U. Awareness and Current Knowledge of Breast Cancer. Biol. Res. 2017, 50, 33. [Google Scholar] [CrossRef]
- Kourou, K.; Exarchos, T.P.; Exarchos, K.P.; Karamouzis, M.V.; Fotiadis, D.I. Machine Learning Applications in Cancer Prognosis and Prediction. Comput. Struct. Biotechnol. J. 2015, 13, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Egwom, O.J.; Hassan, M.; Tanimu, J.J.; Hamada, M.; Ogar, O.M. An LDA–SVM Machine Learning Model for Breast Cancer Classification. BioMedInformatics 2022, 2, 345–358. [Google Scholar] [CrossRef]
- Way, G.P.; Sanchez-Vega, F.; La, K.; Armenia, J.; Chatila, W.K.; Luna, A.; Sander, C.; Cherniack, A.D.; Mina, M.; Ciriello, G.; et al. Machine Learning Detects Pan-Cancer Ras Pathway Activation in The Cancer Genome Atlas. Cell Rep. 2018, 23, 172–180.e3. [Google Scholar] [CrossRef] [PubMed]
- Banegas-Luna, A.J.; Peña-García, J.; Iftene, A.; Guadagni, F.; Ferroni, P.; Scarpato, N.; Zanzotto, F.M.; Bueno-Crespo, A.; Pérez-Sánchez, H. Towards the Interpretability of Machine Learning Predictions for Medical Applications Targeting Personalised Therapies: A Cancer Case Survey. Int. J. Mol. Sci. 2021, 22, 4394. [Google Scholar] [CrossRef]
- Fogliatto, F.S.; Anzanello, M.J.; Soares, F.; Brust-Renck, P.G. Decision Support for Breast Cancer Detection: Classification Improvement Through Feature Selection. Cancer Control 2019, 26, 107327481987659. [Google Scholar] [CrossRef]
- Aishwarja, A.I.; Eva, N.J.; Mushtary, S.; Tasnim, Z.; Khan, N.I.; Islam, M.N. Exploring the Machine Learning Algorithms to Find the Best Features for Predicting the Breast Cancer and Its Recurrence. In Proceedings of the International Conference on Intelligent Computing & Optimization, Hua Hin, Thailand, 30–31 December 2021; pp. 546–558. [Google Scholar]
- Asri, H.; Mousannif, H.; Al Moatassime, H.; Noel, T. Using Machine Learning Algorithms for Breast Cancer Risk Prediction and Diagnosis. Procedia Comput. Sci. 2016, 83, 1064–1069. [Google Scholar] [CrossRef]
- Bazazeh, D.; Shubair, R. Comparative Study of Machine Learning Algorithms for Breast Cancer Detection and Diagnosis. In Proceedings of the 2016 5th International Conference on Electronic Devices, Systems and Applications (ICEDSA), Ras Al Khaimah, United Arab Emirates, 6–8 December 2016; IEEE: Manhattan, NY, USA, 2016; pp. 1–4. [Google Scholar]
- Agarap, A.F.M. On Breast Cancer Detection. In Proceedings of the 2nd International Conference on Machine Learning and Soft Computing—ICMLSC ’18, Phu Quoc Island, Vietnam, 2–4 February 2018; ACM Press: New York, NY, USA, 2018; pp. 5–9. [Google Scholar]
- Sharma, S.; Aggarwal, A.; Choudhury, T. Breast Cancer Detection Using Machine Learning Algorithms. In Proceedings of the 2018 International Conference on Computational Techniques, Electronics and Mechanical Systems (CTEMS), Belgaum, India, 21–22 December 2018; IEEE: Manhattan, NY, USA, 2018; pp. 114–118. [Google Scholar]
- Nindrea, R.D.; Aryandono, T.; Lazuardi, L.; Dwiprahasto, I. Diagnostic Accuracy of Different Machine Learning Algorithms for Breast Cancer Risk Calculation: A Meta-Analysis. Asian Pac. J. Cancer Prev. 2018, 19, 1747–1752. [Google Scholar] [CrossRef]
- Tomar, D.; Agarwal, S. Hybrid Feature Selection Based Weighted Least Squares Twin Support Vector Machine Approach for Diagnosing Breast Cancer, Hepatitis, and Diabetes. Adv. Artif. Neural Syst. 2015, 2015, 265637. [Google Scholar] [CrossRef]
- Madhavi, B.; Reddy, R. Detection and Diagnosis of Breast Cancer Using Machine Learning Algorithm. Int. J. Adv. Sci. Technol. 2019, 28, 228–237. [Google Scholar]
- Dhahri, H.; Al Maghayreh, E.; Mahmood, A.; Elkilani, W.; Faisal Nagi, M. Automated Breast Cancer Diagnosis Based on Machine Learning Algorithms. J. Healthc. Eng. 2019, 2019, 4253641. [Google Scholar] [CrossRef] [PubMed]
- Bhise, S.; Gadekar, S.; Gaur, A.S.; Bepari, S.; Deepmala Kale, D.S.A. Breast Cancer Detection Using Machine Learning Techniques. Int. J. Eng. Res. Technol. 2021, 10. [Google Scholar] [CrossRef]
- Silva, J.; Lezama, O.B.P.; Varela, N.; Borrero, L.A. Integration of Data Mining Classification Techniques and Ensemble Learning for Predicting the Type of Breast Cancer Recurrence. In Proceedings of the International Conference on Green, Pervasive, and Cloud Computing, Uberlândia, Brazil, 26–28 May 2019; pp. 18–30. [Google Scholar]
- Jadhav, S.; Channe, H. Comparative Study of K-NN, Naive Bayes and Decision Tree Classification Techniques. Int. J. Sci. Res. 2013, 5, 1842–1845. [Google Scholar]
- Macaulay, B.O.; Aribisala, B.S.; Akande, S.A.; Akinnuwesi, B.A.; Olabanjo, O.A. Breast Cancer Risk Prediction in African Women Using Random Forest Classifier. Cancer Treat. Res. Commun. 2021, 28, 100396. [Google Scholar] [CrossRef]
- Ak, M.F. A Comparative Analysis of Breast Cancer Detection and Diagnosis Using Data Visualization and Machine Learning Applications. Healthcare 2020, 8, 111. [Google Scholar] [CrossRef]
- Vaka, A.R.; Soni, B.; Reddy, S. Breast Cancer Detection by Leveraging Machine Learning. ICT Express 2020, 6, 320–324. [Google Scholar] [CrossRef]
- Abdar, M.; Zomorodi-Moghadam, M.; Zhou, X.; Gururajan, R.; Tao, X.; Barua, P.D.; Gururajan, R. A New Nested Ensemble Technique for Automated Diagnosis of Breast Cancer. Pattern Recognit. Lett. 2020, 132, 123–131. [Google Scholar] [CrossRef]
- Kousalya, K.; Krishnakumar, B.; Shanthosh, C.I.; Sharmila, R.; Sneha, V. Diagnosis of Breast Cancer Using Machine Learning Algorithms. Int. J. Adv. Sci. Technol. 2020, 29, 970–974. [Google Scholar]
- El-Nabawy, A.; El-Bendary, N.; Belal, N.A. A Feature-Fusion Framework of Clinical, Genomics, and Histopathological Data for METABRIC Breast Cancer Subtype Classification. Appl. Soft Comput. 2020, 91, 106238. [Google Scholar] [CrossRef]
- El-Nabawy, A.; Belal, N.A.; El-Bendary, N. A Cascade Deep Forest Model for Breast Cancer Subtype Classification Using Multi-Omics Data. Mathematics 2021, 9, 1574. [Google Scholar] [CrossRef]
- Jessica, E.O.; Hamada, M.; Yusuf, S.I.; Hassan, M. The Role of Linear Discriminant Analysis for Accurate Prediction of Breast Cancer. In Proceedings of the 2021 IEEE 14th International Symposium on Embedded Multicore/Many-core Systems-on-Chip (MCSoC), Singapore, 20–23 December 2021; IEEE: Manhattan, NY, USA, 2021; pp. 340–344. [Google Scholar]
- Polaka, I.; Bhandari, M.P.; Mezmale, L.; Anarkulova, L.; Veliks, V.; Sivins, A.; Lescinska, A.M.; Tolmanis, I.; Vilkoite, I.; Ivanovs, I.; et al. Modular Point-of-Care Breath Analyzer and Shape Taxonomy-Based Machine Learning for Gastric Cancer Detection. Diagnostics 2022, 12, 491. [Google Scholar] [CrossRef] [PubMed]
- Naji, M.A.; El Filali, S.; Aarika, K.; Benlahmar, E.H.; Abdelouhahid, R.A.; Debauche, O. Machine Learning Algorithms For Breast Cancer Prediction And Diagnosis. Procedia Comput. Sci. 2021, 191, 487–492. [Google Scholar] [CrossRef]
- Tharwat, A.; Gaber, T.; Ibrahim, A.; Hassanien, A.E. Linear Discriminant Analysis: A Detailed Tutorial. AI Commun. 2017, 30, 169–190. [Google Scholar] [CrossRef]
- Zhang, D.; Jing, X.-Y.; Yang, J. Linear Discriminant Analysis. Biometric Image Discrim. Technol. 2011, 41–64. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Cateni, S.; Vannucci, M.; Vannocci, M.; Colla, V. Variable Selection and Feature Extraction Through Artificial Intelligence Techniques. Multivar. Anal. Manag. Eng. Sci. 2013, 6, 103–118. [Google Scholar] [CrossRef]
- Awad, M.; Khanna, R. Support Vector Machines for Classification. In Efficient Learning Machines; Apress: Berkeley, CA, USA, 2015; pp. 39–66. [Google Scholar]
- Cervantes, J.; Garcia-Lamont, F.; Rodríguez-Mazahua, L.; Lopez, A. A Comprehensive Survey on Support Vector Machine Classification: Applications, Challenges and Trends. Neurocomputing 2020, 408, 189–215. [Google Scholar] [CrossRef]
- Arowolo, M.O.; Adebiyi, M.O.; Nnodim, C.T.; Abdulsalam, S.O.; Adebiyi, A.A. An Adaptive Genetic Algorithm with Recursive Feature Elimination Approach for Predicting Malaria Vector Gene Expression Data Classification Using Support Vector Machine Kernels. Walailak J. Sci. Technol. 2021, 18, 9849. [Google Scholar] [CrossRef]
- Huang, M.-W.; Chen, C.-W.; Lin, W.-C.; Ke, S.-W.; Tsai, C.-F. SVM and SVM Ensembles in Breast Cancer Prediction. PLoS ONE 2017, 12, e0161501. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).